Abstract
Alistipes ihumii strain AP11T sp. nov. is the type strain of A. ihumii sp. nov., a new species within the genus Alistipes. This strain, whose genome is described here, was isolated from the fecal flora of a 21-year-old French Caucasian female, suffering from a severe restrictive form of anorexia nervosa since the age of 12 years. A. ihumii is a Gram-negative anaerobic bacillus. Here we describe the features of this organism, together with the complete genome sequence and annotation. The 2,753,264 bp long genome (one chromosome but no plasmid) contains 2,254 protein-coding and 47 RNA genes, including 3 rRNA genes.
Keywords: Alistipes ihumii, genome, culturomics, taxono-genomics
Introduction
Alistipes ihumii strain AP11T (= CSUR P204 = DSM 26107) is the type strain of A. ihumii sp. nov. This bacterium is a Gram-negative, non-spore-forming, anaerobic and non-motile bacillus that was isolated from the stool of a 21-year-old French female suffering from anorexia nervosa, and is part of a “culturomics” study aiming at cultivating individually all species within human feces [1-3].
Prokaryotic taxonomy is episodically confronted with the advancement of methodological and conceptual innovations. The current classification methodology for prokaryotes is known as polyphasic taxonomy, and relies on a combination of phenotypic and genotypic characteristics [4]. The number of completely sequenced genomes is geometrically increasing with time, concurrently with the decrease in cost of such techniques. To date, more than 6,000 bacterial genomes have been published and approximately 25,000 genome sequencing projects have been announced [5]. We recently proposed to integrate genomic information in the taxonomic framework for the description of new bacterial species [6-27].
The genus Alistipes (Rautio et al. 2003) was created in 2003 [28] and is composed of strictly anaerobic Gram-negative rods that resemble the Bacteroides fragilis group in that most species are bile-resistant and indole-positive [29]. This genus is currently comprised of five species with validly published names, including A. finegoldii, A. putredinis [28], A. indistinctus [30], A. onderdonkii and A. shahii [31], to which we added three proposed new species, A. senegalensis [8], A. timonensis [9] and A. obesi [22].
Here we present a summary classification and a set of features for a new Alistipes species, A. ihumii sp. nov. strain AP11T (= CSUR P204 = DSM 26107), together with the description of the complete genomic sequence and its annotation.
Classification and features
A stool sample was collected from a 21-year-old French Caucasian female suffering from severe restrictive form of anorexia nervosa since the age of 12 years. At the time of sample collection, she was hospitalized in our hospital for recent aggravation of her medical condition (BMI: 10.4 kg/m2). The patient gave an informed and signed consent. Both this study and the assent procedure were approved by the Ethics Committee of the Institut Fédératif de Recherche IFR48, Faculty of Medicine, Marseille, France under reference 09-022. Ten other potentially new bacterial species were isolated from this patient’s stool, all of which are currently being described. Microbial culturomics also enabled the isolation of several other new bacterial species from other stool specimens [6-27]. The fecal specimen was stored at -80°C immediately after collection. Strain AP11T was isolated in November 2011 after 2 days of inoculation in anaerobic blood culture bottle with the addition of 5mL of thioglycolate and further inoculation on Columbia agar (BioMerieux, Marcy l’Etoile, France).
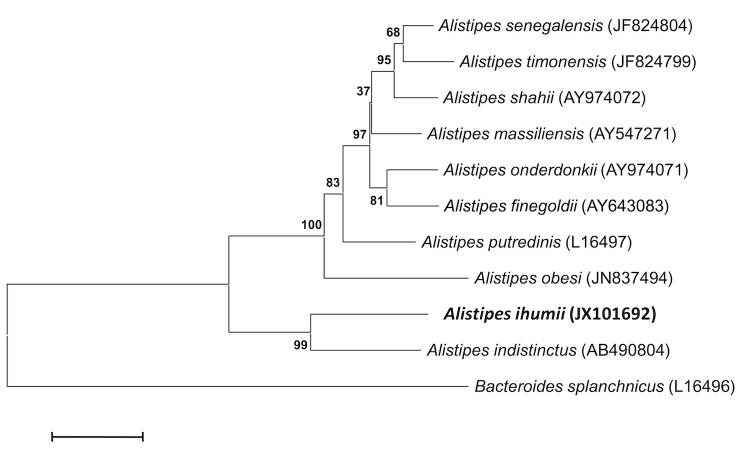
This strain exhibited a 95% 16S rRNA sequence similarity with A. indistinctus [30], the phylogenetically closest Alistipes species with a validly published name (Table 1, Figure 1), and 92% with A. onderdonkii [28] and A. putredinis [31]. This value was in the range of 16S rRNA sequence identities among species within the genus Alistipes that range from 90 to 95%, and lower than the 98.7% 16S rRNA gene sequence threshold recommended by Stackebrandt and Ebers to delineate a new species without carrying out DNA-DNA hybridization [41].
Table 1. Classification and general features of Alistipes ihumii strain AP11T according to the MIGS recommendations [32].
| MIGS ID | Property | Term | Evidence codea |
|---|---|---|---|
| Current classification | Domain Bacteria Phylum Bacteroidetes Class Bacteroidia Order Bacteroidales Family Rikenellaceae Genus Alistipes Species Alistipes ihumii Type strain AP11T |
TAS [33] TAS [34,35] TAS [34,36] TAS [34,37] TAS [34,38] TAS [28,39] IDA IDA |
|
| Gram stain | Negative | IDA | |
| Cell shape | Rod | IDA | |
| Motility | nonmotile | IDA | |
| Sporulation | nonsporulating | IDA | |
| Temperature range | mesophile | IDA | |
| Optimum temperature | 37°C | IDA | |
| MIGS-6.3 | Salinity | unknown | IDA |
| MIGS-22 | Oxygen requirement | anaerobic | IDA |
| Carbon source | unknown | ||
| Energy source | unknown | ||
| MIGS-6 | Habitat | human gut | IDA |
| MIGS-15 | Biotic relationship | free living | IDA |
| MIGS-14 | Pathogenicity Biosafety level Isolation |
unknown 2 human feces |
|
| MIGS-4 | Geographic location | France | IDA |
| MIGS-5 | Sample collection time | November 2011 | IDA |
| MIGS-4.1 | Latitude & Longitude | 43.296482 & 5.36978 | IDA |
| MIGS-4.3 | Depth | surface | IDA |
| MIGS-4.4 | Altitude | 0 m above sea level | IDA |
Evidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from the Gene Ontology project [40]. If the evidence is IDA, then the property was directly observed for a live isolate by one of the authors or an expert mentioned in the acknowledgements.
Figure 1.
Phylogenetic tree highlighting the position of Alistipes ihumii strain AP11T relative to other type strains within the genus Alistipes. GenBank accession numbers are indicated in parentheses. Sequences were aligned using CLUSTALW, and phylogenetic inferences obtained using the maximum-likelihood method within the MEGA software. Numbers at the nodes are percentages of bootstrap values obtained by repeating the analysis 500 times to generate a majority consensus tree. Bacteroides splanchnicus was used as the outgroup. The scale bar represents a 2% nucleotide sequence divergence.
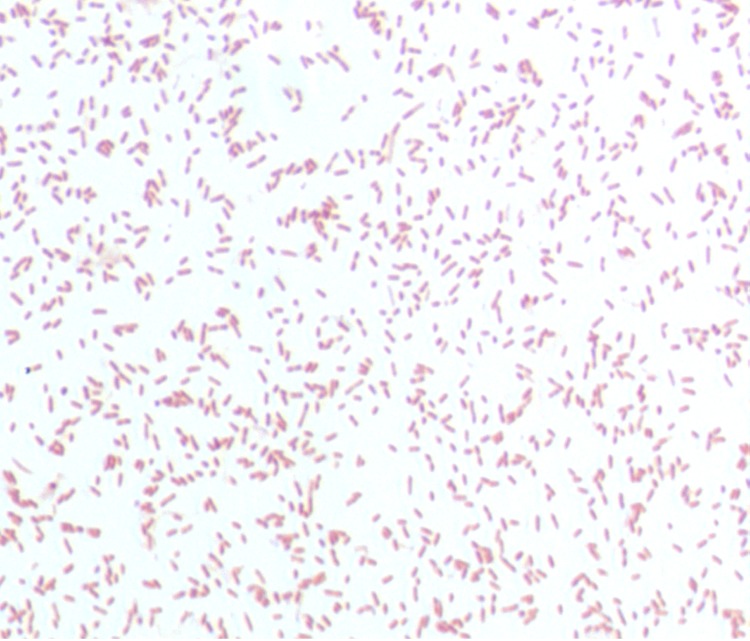
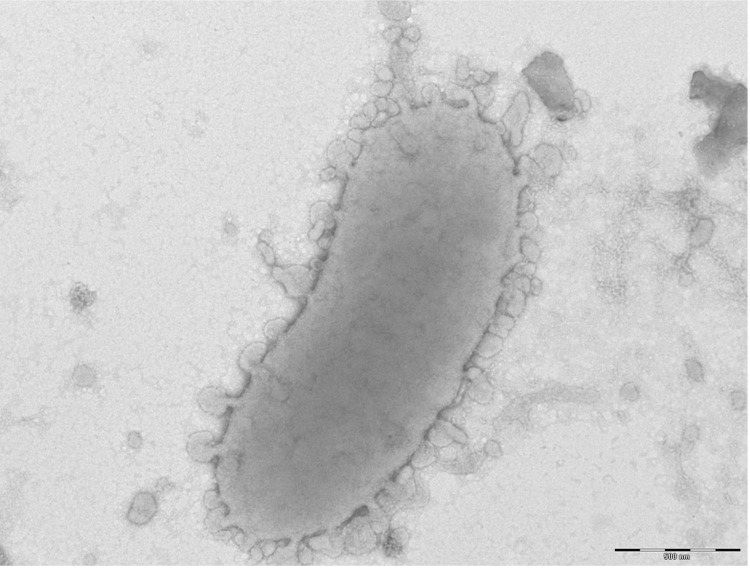
Different growth temperatures (25, 30, 37, 45°C) were tested. Growth was observed between 25 and 45°C, with optimal growth at 37°C after 24 hours of inoculation. Colonies were about 0.2 mm in diameter, transparent, and exhibited a ß-hemolytic activity on blood-enriched Columbia agar. Growth of the strain was tested on 5% sheep blood agar, under anaerobic and microaerophilic conditions using the GENbag anaer and GENbag microaer systems, respectively (BioMerieux), and under aerobic conditions with or without 5% CO2. Optimal growth of this strain was obtained anaerobically, weak growth was observed under microaerophilic conditions, and no growth was observed under aerobic atmosphere. The motility test was negative. Cells grown on agar are Gram-negative rods (Figure 2) and have mean diameter and length of 0.72 and 1.69 µm, respectively, as determined using electron microscopy (Figure 3).
Figure 2.
Gram stain of A. ihumii strain AP11T
Figure 3.
Transmission electron microscopy of A. ihumii strain AP11T, using a Morgani 268D (Philips) at an operating voltage of 60kV. The scale bar represents 500 nm.
Strain AP11T exhibited oxidase but no catalase activities. Using API 50CH (BioMérieux), we observed that strain AP11T was asaccharolytic. Using API 32A (BioMérieux), positive reactions were obtained for α-glucosidase, β-glucosidase, N-acetyl-β-glucosaminidase, mannose and raffinose fermentation, alkaline phosphatase, leucyl glycine arylamidase, alanine arylamidase, and glutamyl glutamic acid arylamidase. Weak reactions were observed for α-galactosidase and glutamic acid decarboxilase. Negative reactions were obtained for urease, arginine dihydrolase, β-galactosidase, 6 phospho-β-galactosidase , α-arabinosidase, β-glucuronidase, α-fucosidase, nitrate reduction, indole production, arginine arylamidase, proline arylamidase, phenylalanine arylamidase, leucine arylamidase, pyroglutamic acid arylamidase, tyrosine arylamidase, glycine arylamidase, histidine arylamidase, and serine arylamidase. A. ihumii is susceptible to amoxicillin, imipenem, and clindamycin, but resistant to vancomycin. When compared with representative species from the genus Alistipes, strain AP11T exhibited the phenotypic differences detailed in Table 2.
Table 2. Differential characteristics of Alistipes strains†.
| Properties | A. ihumii | A. senegalensis | A. timonensis |
A. putredinis |
A. indistinctus |
A. shahii |
A. obesi |
A. finegoldii |
|---|---|---|---|---|---|---|---|---|
| Cell diameter (µm) | 0.72 | 0.56 | 0.62 | 0.40 | 0.60 | 0.15 | 0.44-0.76 | 0.8-2.0 |
| Oxygen requirement | anaerobic | anaerobic | anaerobic | anaerobic | anaerobic | anaerobic | anaerobic | anaerobic |
| Pigment production | – | + | + | – | + | + | + | + |
| Gram stain | – | – | – | – | – | – | – | – |
| Salt requirement | na | + | – | – | – | – | – | na |
| Motility | – | – | – | – | – | – | + | – |
| Endospore formation | – | – | – | – | – | – | na | – |
| Production of | ||||||||
| Alkaline phosphatase | na | na | na | + | w | + | + | + |
| Catalase | – | + | + | + | + | – | + | – |
| Oxidase | + | – | – | – | – | – | – | na |
| Nitrate reductase | – | na | na | – | – | – | – | – |
| Urease | – | na | na | – | – | + | – | na |
| β-galactosidase | – | w | + | – | – | + | + | + |
| N-acetyl-glucosamine | + | na | W | – | + | + | + | + |
| Indole | – | w | W | + | – | + | – | + |
| Activity for | ||||||||
| Leucyl glycine arylamidase | + | + | + | + | – | + | + | – |
| Glutamic acid decarboxylase | w | na | + | + | – | – | – | na |
| Glycine arylamidase | – | + | + | na | – | – | – | na |
| Chymotrypsin | na | na | na | – | – | – | na | na |
| Acid from | ||||||||
| L-Arabinose | na | na | na | – | + | na | na | na |
| Raffinose | + | na | – | – | + | + | – | na |
| Mannose | + | + | – | – | + | + | – | na |
| Mannitol | na | na | na | na | + | na | na | na |
| Sucrose | na | na | na | – | + | + | na | na |
| D-glucose | na | na | na | – | + | + | na | na |
| D-fructose | na | na | na | – | + | + | na | na |
| D-maltose | na | na | na | – | + | + | na | na |
| D-lactose | na | na | na | – | + | + | na | na |
| Hydrolysis of gelatin | na | na | na | + | + | – | na | + |
| G+C content (mol%) | 57.90 | 58.40 | 58.82 | 55.3 | 55.2 | 57.20 | 58.60 | 56.65 |
| Habitat | human gut | human gut | human gut | appendix of children | human gut | human gut | human gut | human appendix tissue |
na = data not available; w = weak
†Alistipes ihumii strain AP11T, A. senegalensis strain JC50T, A. timonensis strain JC136T, A. putredinis strain ATCC29800 T, A. indistinctus strain YIT12060T, A. shahii strain WAL 8301T, A. obesi strain ph8T and A. finegoldii AHN2437T
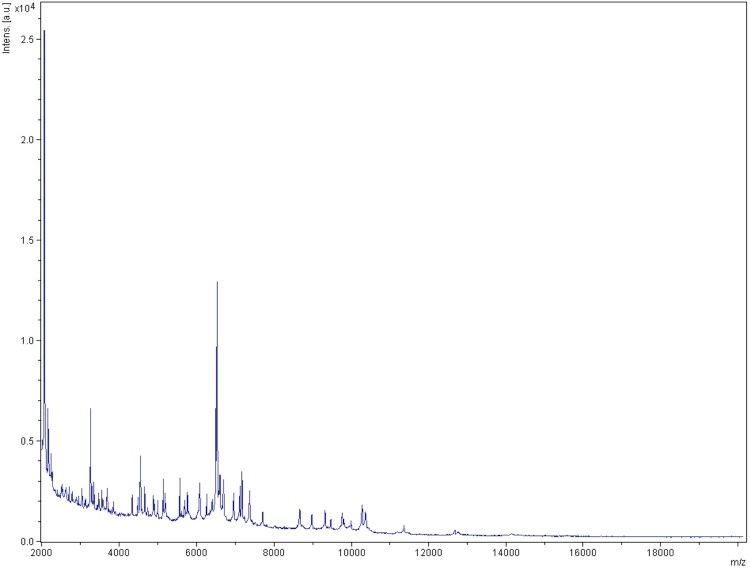
Matrix-assisted laser-desorption/ionization time-of-flight (MALDI-TOF) MS protein analysis was carried out as previously described [42] using a Microflex spectrometer (Bruker Daltonics, Leipzig, Germany). Twelve individual colonies were deposited on a MTP 384 MALDI-TOF target plate (Bruker). The twelve AP11T spectra were imported into the MALDI BioTyper software (version 2.0, Bruker) and analyzed by standard pattern matching (with default parameter settings) against the main spectra of 4,706 bacteria, including spectra from A. finegoldii, A. onderdonkii, A. shahii, A. senegalensis, A. obesi and A. timonensis, used as reference data in the BioTyper database. The output score enabled the presumptive identification and discrimination of the tested species from those in the database: a score > 2 with a validated species identifies a strain at the species level; and a score < 1.7 indicates a species-level match was not made. For strain AP11T, no significant score was obtained, suggesting that our isolate was not a member of any known species (Figures 4 and 5). We added the spectrum from strain AP11T to our database.
Figure 4.
Reference mass spectrum from A. ihumii strain AP11T. Spectra from 12 individual colonies were compared and a reference spectrum was generated.
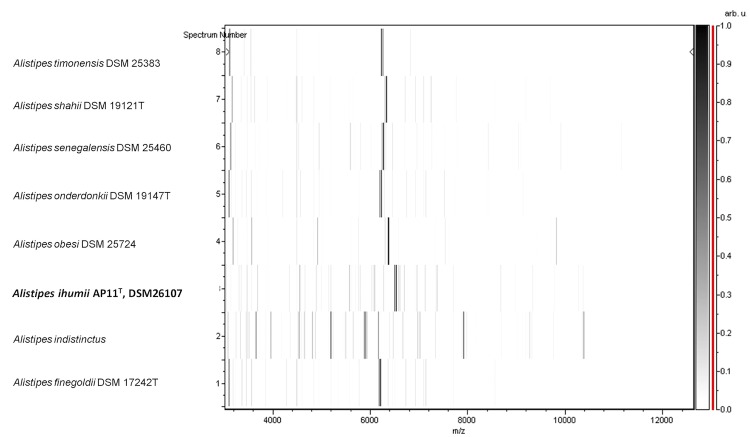
Figure 5.
Gel view comparing spectra from Alistipes ihumii strain AP11T and other members of the genus Alistipes (A. obesi, A. timonensis, A. senegalensis, A. shahii, A. onderdonkii and A. finegoldii). The Gel View displays the raw spectra of all loaded spectrum files arranged in a pseudo-gel like look, with each peak displayed as a band or bar. The peak intensity is reflected by the intensity of the gray color. The right y-axis shows the relationship between the shades of gray and the peak intensity in arbitrary units. The x-axis records the m/z value. The left y-axis displays the running spectrum number originating from subsequent spectra loading.
Genome sequencing information
Genome project history
The organism was selected for sequencing on the basis of its phylogenetic position and 16S rRNA similarity to other members of the Alistipes genus, and is part of a “culturomics” study of the human digestive flora aiming at isolating all bacterial species within human feces. It was the eighth sequenced genome from an Alistipes species and the first from Alistipes ihumii sp. nov. A summary of the project information is shown in Table 3. The Genbank accession number is CAPH00000000 and consists of 60 contigs. Table 3 shows the project information and its association with MIGS version 2.0 compliance [43].
Table 3. Project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | High-quality draft |
| MIGS-28 | Libraries used | One 454 paired end 3-kb library |
| MIGS-29 | Sequencing platforms | 454 GS FLX Titanium |
| MIGS-31.2 | Fold coverage | 35× |
| MIGS-30 | Assemblers | Newbler version 2.5.3 |
| MIGS-32 | Gene calling method | Prodigal |
| Genbank ID | CAPH00000000 | |
| Genbank Date of Release | November 28, 2012 | |
| Gold ID | Gi20720 | |
| MIGS-13 | Project relevance | Study of the human gut microbiome |
Growth conditions and DNA isolation
A. ihumii sp. nov. strain AP11T, (= CSURP204 = DSM 26107), was grown aerobically on 5% sheep blood agar medium at 37°C. Five Petri dishes were spread and resuspended in 3x100µl of G2 buffer (EZ1 DNA Tissue kit, Qiagen). A first mechanical lysis was performed by glass powder on the Fastprep-24 device (Sample Preparation system, MP Biomedicals, USA) for 2×20 seconds. DNA was treated with 2.5 µg/µL of lysozyme (30 minutes at 37°C) and extracted using the BioRobot EZ 1 Advanced XL (Qiagen). The DNA was concentrated and purified on a Qiamp kit (Qiagen). The yield and the concentration of DNA was 70.7 ng/µl as measured by using Quant-it Picogreen kit (Invitrogen) on the Genios Tecan fluorometer.
Genome sequencing and assembly
A 3kb paired-end sequencing strategy (Roche, Meylan, France) was used. DNA (5 µg) was mechanically fragmented for the paired-end sequencing, using a Covaris device (Covaris Inc., Woburn, MA,USA) with an enrichment size of 3-4 kb. The DNA fragmentation was visualized through an Agilent 2100 BioAnalyzer on a DNA Labchip 7500 which yielded an optimal size of 2.3 kb. The library was constructed using the 454 GS FLX Titanium paired-end rapid library protocol. Circularization and nebulization were performed which generated a pattern of optimal size of 457 bp. PCR amplification was performed for 17 cycles followed by double size selection. The single-stranded paired-end library was quantified using a Quant-it Ribogreen Kit (Invitrogen) and the Genios Tecan fluorometer. The library concentration equivalence was calculated as 1.94× 1010 molecules/µL. The library was stored at -20°C until further use.
The paired-end library was clonally amplified with 0.5 and 1 cpb in 2 emPCR reactions with the GS Titanium SV emPCR Kit (Lib-L) v2 (Roche). The yield of the shotgun emPCR reactions was 6.24 and 16.24% respectively for the two kinds of paired- end emPCR reactions according to the quality expected (range of 5 to 20%) from the Roche procedure. Two libraries were loaded on the GS Titanium PicoTiterPlates (PTP Kit 70x75, Roche) and pyrosequenced with the GS Titanium Sequencing Kit XLR70 and the GS FLX Titanium sequencer (Roche). The run was performed overnight and then analyzed on the cluster through the gsRunBrowser and Newbler assembler (Roche). A total of 260,838 passed filter wells were obtained and generated 96.3 Mb with an average length of 369 bp. The passed filter sequences were assembled using Newbler with 90% identity and 40 bp as overlap. The final assembly identified 9 scaffolds and 60 contigs (> 1,500 bp) and generated a genome size of 2.75 Mb which corresponds to a coverage of 35× genome equivalent.
Genome annotation
Open Reading Frames (ORFs) were predicted using Prodigal [44] with default parameters but the predicted ORFs were excluded if they were spanning a sequencing gap region. The predicted bacterial protein sequences were searched against the GenBank database [45] and the Clusters of Orthologous Groups (COG) databases using BLASTP. The tRNAScan-SE tool [46] was used to find tRNA genes, whereas ribosomal RNAs were found by using RNAmmer [47] and BLASTn against the GenBank database. Lipoprotein signal peptides and numbers of transmembrane helices were predicted using SignalP [48] and TMHMM [49] respectively. ORFans were identified if their BLASTP E-value was lower than 1e-03 for alignment length greater than 80 amino acids. If alignment lengths were smaller than 80 amino acids, we used an E-value of 1e-05. Such parameter thresholds have already been used in previous works to define ORFans.
Orthologous gene sets composed of one gene from A. ihumii compared to each of A. obesi strain ph8T (GenBank accession number CAHA00000000), A. finegoldii strain AHN 2437 (CP003274), A. indistinctus strain YIT 12060 (ADLD00000000), A. putredinis strain DSM 17216 (ABFK00000000), A. senegalensis strain JC50T (CAHI00000000), A. shahii strain WAL 8301 (FP929032), and A. timonensis strain JC136T (CAEG00000000) were identified using the Proteinortho software (version 1.4) [50] using a 30% protein identity and an E-value of 1e-05. The average percentage of nucleotide sequence identity of each orthologous set was determined using the Needleman-Wunsch algorithm global alignment technique. Artemis [51] was used for data management and DNA Plotter [52] was used for visualization of genomic features. The Mauve alignment tool was used for multiple genomic sequence alignment and visualization [53].
Genome properties
The genome of A. ihumii strain AP11T is 2,753,264 bp long (1 chromosome, but no plasmid) with a 57.90% G + C content (Figure 6 and Table 4). Of the 2,301 predicted genes, 2,254 were protein-coding genes, and 47 were RNAs. One rRNA operon (one 16S rRNA, one 23S rRNA and one 5S rRNA) and 44 predicted tRNA genes were identified in the genome. A total of 1,465 genes (63.66%) were assigned a putative function. Two hundred thirty-seven genes were identified as ORFans (10.29%). The remaining genes were annotated as hypothetical proteins. The properties and the statistics of the genome are summarized in Tables 4 and 5. The distribution of genes into COGs functional categories is presented in Table 5.
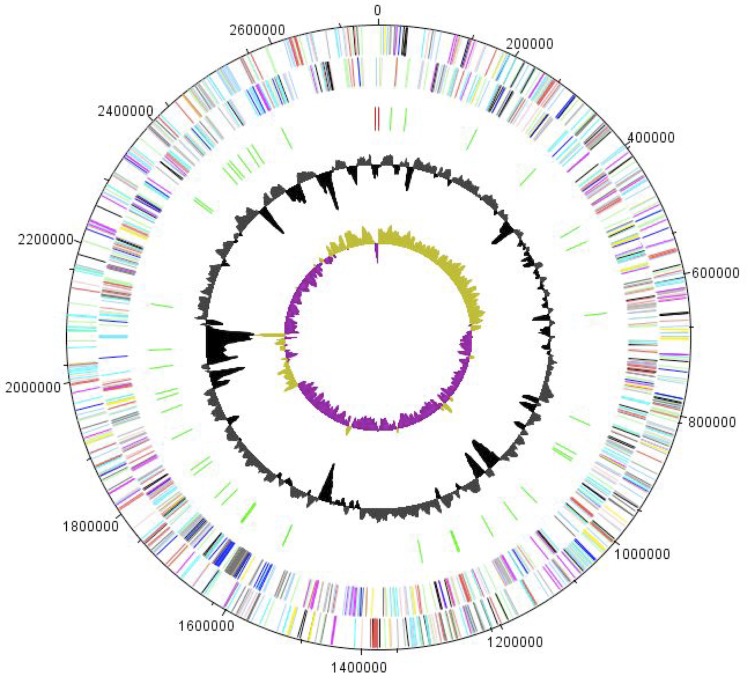
Figure 6.
Graphical circular map of the chromosome. From the outside in, the outer two circles show open reading frames oriented in the forward and reverse (colored by COG categories) directions, respectively. The third circle marks the rRNA gene operon (red) and tRNA genes (green). The fourth circle shows the G+C% content plot. The inner-most circle shows GC skew, purple and olive indicating negative and positive values, respectively.
Table 4. Nucleotide content and gene count levels of the genome.
| Attribute | Value | % of totala |
|---|---|---|
| Genome size (bp) | 2,753,264 | |
| DNA coding region (bp) | 2,320,878 | 84.29 |
| DNA G+C content (bp) | 1,594,140 | 57.90 |
| Number of replicons | 1 | |
| Extrachromosomal elements | 0 | |
| Total genes | 2,301 | 100 |
| RNA genes | 47 | 2.04 |
| rRNA operons | 1 | |
| Protein-coding genes | 2,254 | 97.95 |
| Genes with function prediction | 1,540 | 66.92 |
| Genes assigned to COGs | 1,465 | 63.66 |
| Protein coding genes assigned Pfam domains | 1,834 | 79.70 |
| Genes with peptide signals | 296 | 12.86 |
| Genes with transmembrane helices | 457 | 19.86 |
| CRISPR repeats | 1 |
a The total is based on either the size of the genome in base pairs or the total number of protein coding genes in the annotated genome.
Table 5. Number of genes associated with the 25 general COG functional categories.
| Code | Value | %agea | Description |
|---|---|---|---|
| J | 143 | 6.34 | Translation |
| A | 0 | 0 | RNA processing and modification |
| K | 88 | 3.90 | Transcription |
| L | 113 | 5.01 | Replication, recombination and repair |
| B | 0 | 0 | Chromatin structure and dynamics |
| D | 19 | 0.84 | Cell cycle control, mitosis and meiosis |
| Y | 0 | 0 | Nuclear structure |
| V | 28 | 1.24 | Defense mechanisms |
| T | 38 | 1.69 | Signal transduction mechanisms |
| M | 161 | 7.14 | Cell wall/membrane biogenesis |
| N | 6 | 0.27 | Cell motility |
| Z | 0 | 0 | Cytoskeleton |
| W | 0 | 0 | Extracellular structures |
| U | 32 | 142 | Intracellular trafficking and secretion |
| O | 60 | 2.66 | Posttranslational modification, protein turnover, chaperones |
| C | 111 | 4.92 | Energy production and conversion |
| G | 106 | 4.70 | Carbohydrate transport and metabolism |
| E | 131 | 5.81 | Amino acid transport and metabolism |
| F | 52 | 2.31 | Nucleotide transport and metabolism |
| H | 75 | 3.33 | Coenzyme transport and metabolism |
| I | 49 | 2.17 | Lipid transport and metabolism |
| P | 72 | 3.19 | Inorganic ion transport and metabolism |
| Q | 21 | 0.93 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 235 | 10.43 | General function prediction only |
| S | 91 | 4.04 | Function unknown |
| - | 790 | 35.05 | Not in COGs |
aThe total is based on the total number of protein coding genes in the annotated genome.
Genome comparison with other Alistipes species
Here, we compared the genome of A. ihumii strain AP11T to those of A. obesi strain ph8T (GenBank accession number CAHA00000000), A. finegoldii strain AHN 2437 (CP003274), A. indistinctus strain YIT 12060 (ADLD00000000), A. putredinis strain DSM 17216 (ABFK00000000), A. senegalensis strain JC50T (CAHI00000000), A. shahii strain WAL 8301 (FP929032), and A. timonensis strain JC136T (CAEG00000000). The draft genome of A. ihumii is larger than that of A. putredinis (2.75 and 2.55 Mb, respectively) but smaller than those of A. indistinctus, A. obesi, A. timonensis, A. finegoldii, A. shahii and A. senegalensis (2.85, 3.16, 3.49, 3.73, 3.76, and 4.01 Mb, respectively). The G+C content of A. ihumii is comparable to that of A. shahii (57.90 and 57.60%, respectively), lower than those of A. timonensis and A. senegalensis (58.8 and 58.4%, respectively) and higher than those of A. putredinis, A. indistinctus and A. finegoldii (53.30, 54.80 and 56.60%, respectively). A. ihumii has a smaller gene content than those of A. putredinis, A. indistinctus, A. obesi, A. timonensis, A. shahii, A. senegalensis, and A. finegoldii (2,301, 2,335, 2,342, 2,619, 2,709, 3,132, 3,161, and 3,231, respectively). The ratio of genes per MB of A. ihumii is higher than those of A. timonensis, A. senegalensis, A. indistinctus, and A. obesi (836, 776, 788, 821, and 828, respectively), comparable to that of A. shahii (833) and smaller than those of A. finegoldii and A. putredinis (866 and 915, respectively).
The average genomic nucleotide sequence identity between A. ihumii and other Alistipes species ranged from 70.23 to 74.37%, whereas values ranged from 69.70 to 90.98% among other Alistipes species (Table 6).
Table 6. Numbers of orthologous proteins shared between genomes.
| AIH | ASE | AT | AS | AF | AP | AO | AIN | |
|---|---|---|---|---|---|---|---|---|
| A. ihumii | 2,254 | 1,190 | 1,164 | 958 | 1,150 | 1,055 | 1,130 | 1,147 |
| A. senegalensis | 71.16 | 3,161 | 1,764 | 1,739 | 1,660 | 1,277 | 1,405 | 1,218 |
| A.timonensis | 70.90 | 90.98 | 2,709 | 1,650 | 1,585 | 1,238 | 1,377 | 1,210 |
| A.shahii | 71.19 | 86.33 | 80.03 | 3,132 | 1,674 | 1,270 | 1,166 | 1,155 |
| A.finegoldii | 71.62 | 82.04 | 81.14 | 82.90 | 3,231 | 1,303 | 1,385 | 1,202 |
| A.putrenidis | 70.23 | 75.32 | 75.21 | 75.50 | 76.23 | 2,335 | 1,182 | 1,038 |
| A.onderdonkii | 71.26 | 76.42 | 76.23 | 77.06 | 76.31 | 74.45 | 2,619 | 1,137 |
| A.indistinctus | 74.37 | 70.02 | 70.05 | 70.00 | 69.91 | 69.70 | 69.91 | 2,342 |
Upper right triangle- numbers of orthologous proteins shared between genomes
Lower left triangle- average percentage of nucleotide identity between orthologous gene sets shared between genomes
Bold- numbers of proteins per genome
AIH- A. ihumii, ASE- A. senegalensis, AT- A. timonensis, AS- A. shahii, AF- A. finegoldii, AP- A. putredinis, AO- A. obesi, AIN- A. indistinctus
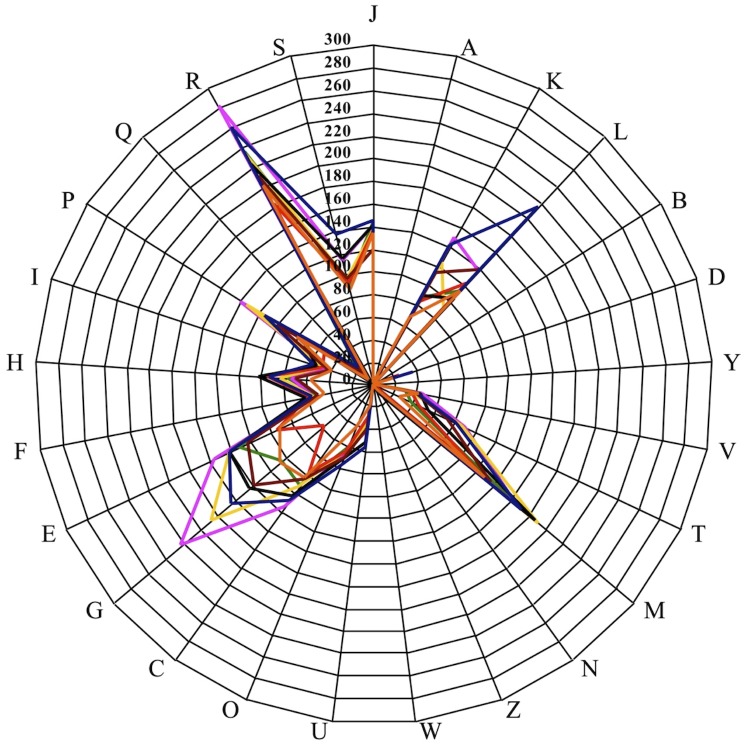
However, the distribution of genes into COG categories was not entirely similar in all eight compared genomes (Figure 7).
Figure 7.
Distribution of functional classes of predicted genes in Alistipes ihumii (colored in green), A. senegalensis (pink), A. timonensis (yellow), A. shahii (brown), A. finegoldii (blue), A. putredinis (red), A. obesi (orange) and A. indistinctus (black) chromosomes according to the clusters of orthologous groups of proteins.
Conclusion
On the basis of phenotypic, phylogenetic and genomic analyses, we formally propose the creation of Alistipes ihumii sp. nov. that contains strain AP11T. This bacterial strain has been isolated from the fecal flora of a patient suffering from anorexia nervosa living in Marseille, France. Several other new bacterial species were also cultivated from this patient as well as fecal samples from other patients using microbial culturomics [6-27], thus suggesting that the human fecal flora from human remains partially unknown.
Description of Alistipes ihumii sp. nov.
Alistipes ihumii (i.hum.i’i. N.L. gen. n. ihumii, based on the acronym IHUMI, the Institut Hospitalo-Universitaire Méditerranée-Infection, where the type strain was isolated).
Colonies are 0.2 mm in diameter and are translucent on blood-enriched Columbia agar. Cells are rod-shaped with a mean diameter of 0.72 µm and a mean length of 1.69 µm. Optimal growth is achieved anaerobically. No growth is obtained aerobically but weak growth is observed in microaerophilic conditions. Growth occurs between 25°C and 45°C, with an optimal growth observed at 37°C.
Cells stain Gram-negative, are non motile and are asaccharolytic. Activities present are α-glucosidase, β-glucosidase, N-acetyl-β-glucosaminidase, mannose and rafinnose fermentation, alkaline phosphatase, leucyl glycine arylamidase, alanine arylamidase, and glutamyl glutamic acid arylamidase. Cells are negative for urease, arginine dihydrolase, β-galactosidase, 6-phospho-β-galactosidase, α-arabinosidase, β-glucuronidase, α-fucosidase, nitrate reduction, indole production, arginine arylamidase, proline arylamidase, phenylalanine arylamidase, leucine arylamidase, pyroglutamic acid arylamidase, tyrosine arylamidase, glycine arylamidase, histidine arylamidase, and serine arylamidase. Cells are susceptible to amoxicillin, imipenem, and clindamycin, but resistant to vancomycin. The G+C content of the genome is 57.90%. The 16S rRNA and genome sequences are deposited in Genbank under accession numbers JX101692 and CAPH00000000, respectively.
The type strain AP11T (= CSUR P204 = DSM 26107) was isolated from the fecal flora of a 21-year-old French Caucasian female suffering from severe anorexia nervosa.
Acknowledgements
The authors thank the Xegen Company (www.xegen.fr) for automating the genomic annotation process. This study was funded by the Mediterranee-Infection Foundation.
References
- 1.Lagier JC, Armougom F, Million M, Hugon P, Pagnier I, Robert C, Bittar F, Fournous G, Gimenez G, Maraninchi M, et al. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin Microbiol Infect 2012; 18:1185-1193 [DOI] [PubMed] [Google Scholar]
- 2.Dubourg G, Lagier JC, Armougom F, Robert C, Hamad I, Brouqui P. The gut microbiota of a patient with resistant tuberculosis is more comprehensively studied by culturomics than by metagenomics. Eur J Clin Microbiol Infect Dis 2013; 2013:637-645. [DOI] [PubMed]
- 3.Pfleiderer A, Lagier JC, Armougom F, Robert C, Vialettes B, Raoult D. Culturomics identified 11 new bacterial species from a single anorexia nervosa stool sample. [Epub ahead of print]. Eur J Clin Microbiol Infect Dis 2013. [DOI] [PubMed]
- 4.Tindall BJ, Rossello-Mora R, Busse HJ, Ludwig W, Kampfer P. Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 2010; 60:249-266 10.1099/ijs.0.016949-0 [DOI] [PubMed] [Google Scholar]
- 5.Genome Online Database http://www.genomesonline.org/cgi-bin/GOLD/index.cgi
- 6.Kokcha S, Mishra AK, Lagier JC, Million M, Leroy Q, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Bacillus timonensis sp. nov. Stand Genomic Sci 2012; 6:346-355 10.4056/sigs.2776064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lagier JC, El Karkouri K, Nguyen TT, Armougom F, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Anaerococcus senegalensis sp. nov. Stand Genomic Sci 2012; 6:116-125 10.4056/sigs.2415480 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Mishra AK, Gimenez G, Lagier JC, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Alistipes senegalensis sp. nov. Stand Genomic Sci 2012; 6:304-314 10.4056/sigs.2625821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lagier JC, Armougom F, Mishra AK, Ngyuen TT, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Alistipes timonensis sp. nov. Stand Genomic Sci 2012; 6:315-324 10.4056/sigs.2685971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Mishra AK, Lagier JC, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Clostridium senegalense sp. nov. Stand Genomic Sci 2012; 6:386-395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mishra AK, Lagier JC, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Peptoniphilus timonensis sp. nov. Stand Genomic Sci 2012; 7:1-11 10.4056/sigs.2956294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mishra AK, Lagier JC, Rivet R, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Paenibacillus senegalensis sp. nov. Stand Genomic Sci 2012; 7:70-81 10.4056/sigs.3056450 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lagier JC, Gimenez G, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Herbaspirillum massiliense sp. nov. Stand Genomic Sci 2012; 7:200-209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Roux V, El Karkouri K, Lagier JC, Robert C, Raoult D. Non-contiguous finished genome sequence and description of Kurthia massiliensis sp. nov. Stand Genomic Sci 2012; 7:221-232 10.4056/sigs.3206554 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kokcha S, Ramasamy D, Lagier JC, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Brevibacterium senegalense sp. nov. Stand Genomic Sci 2012; 7:233-245 10.4056/sigs.3256677 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ramasamy D, Kokcha S, Lagier JC, N’Guyen TT, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Aeromicrobium massilense sp. nov. Stand Genomic Sci 2012; 7:246-257 10.4056/sigs.3306717 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lagier JC, Ramasamy D, Rivet R, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Cellulomonas massiliensis sp. nov. Stand Genomic Sci 2012; 7:258-270 10.4056/sigs.3316719 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lagier JC, El Karkouri K, Rivet R, Couderc C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Senegalemassilia anaerobia sp. nov. Stand Genomic Sci 2013; 7:343-356 10.4056/sigs.3246665 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mishra AK, Hugon P, Lagier JC, Nguyen TT, Robert C, Couderc C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Peptoniphilus obesi sp. nov. Stand Genomic Sci 2013; 7:357-369 10.4056/sigs.32766871 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mishra AK, Lagier JC, Nguyen TT, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Peptoniphilus senegalensis sp. nov. Stand Genomic Sci 2013; 7:370-381 10.4056/sigs.3366764 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lagier JC, El Karkouri K, Mishra AK, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Enterobacter massiliensis sp. nov. Stand Genomic Sci 2013; 7:399-412 10.4056/sigs.3396830 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hugon P, Ramasamy D, Lagier JC, Rivet R, Couderc C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Alistipes obesi sp. nov. Stand Genomic Sci 2013; 7:427-439 10.4056/sigs.3336746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mishra AK, Hugon P, Robert C, Couderc C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Peptoniphilus grossensis sp. nov. Stand Genomic Sci 2012; 7:320-330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mishra AK, Hugon P, Nguyen TT, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of Enorma massiliensis gen. nov., sp. nov., a new member of the Family Coriobacteriaceae. Stand Genomic Sci 2013; 8:290-305 10.4056/sigs.3426906 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ramasamy D, Lagier JC, Gorlas A, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of Bacillus massiliosenegalensis sp. nov. Stand Genomic Sci 2013; 8:264-278 10.4056/sigs.3496989 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ramasamy D, Lagier JC, Nguyen TT, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of of Dielma fastidiosa gen. nov., sp. nov., a new member of the Family Erysipelotrichaceae. Stand Genomic Sci 2013; 8:336-351 10.4056/sigs.3567059 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mishra AK, Lagier JC, Robert C, Raoult D, Fournier PE. Genome sequence and description of Timonella senegalensis gen. nov., sp. nov., a new member of the suborder Micrococcinae. Stand Genomic Sci 2013; 8:318-335 10.4056/sigs.3476977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Rautio M, Eerola E, Väisänen-Tunkelrott ML, Molitoris D, Lawson P, Collins MD, Jousimies-Somer H. Reclassification of Bacteroides putredinis (Weinberg et al., 1937) in a new genus Alistipes gen. nov., as Alistipes putredinis comb. nov., and description of Alistipes finegoldii sp. nov., from human sources. Syst Appl Microbiol 2003; 26:182-188 10.1078/072320203322346029 [DOI] [PubMed] [Google Scholar]
- 29.Tyrrell KL, Warren YA, Citron DM, Goldstein EJC. Re-assessment of phenotypic identifications of Bacteroides putredinis to Alistipes species using molecular methods. Anaerobe 2011; 17:130-134 10.1016/j.anaerobe.2011.04.002 [DOI] [PubMed] [Google Scholar]
- 30.Nagai F, Morotomi M, Watanabe Y, Sakon H, Tanaka R. Alistipes indistinctus sp. nov. and Odoribacter laneus sp. nov., common members of the human intestinal microbiota isolated from faeces. Int J Syst Evol Microbiol 2009; 60:1296-1302 10.1099/ijs.0.014571-0 [DOI] [PubMed] [Google Scholar]
- 31.Song Y, Könönen E, Rautio M, Liu C, Bryk A, Eerola E, Finegold SM. Alistipes onderdonkii sp. nov. and Alistipes shahii sp. nov., of human origin. Int J Syst Evol Microbiol 2006; 56:1985-1990 10.1099/ijs.0.64318-0 [DOI] [PubMed] [Google Scholar]
- 32.12. Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547. [DOI] [PMC free article] [PubMed]
- 33.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Validation List No 143. Int J Syst Evol Microbiol 2012; 62:1-4 10.1099/ijs.0.039487-0 [DOI] [Google Scholar]
- 35.Krieg NR, Ludwig W, Euzéby J, Whitman WB. Phylum XIV. Bacteroidetes phyl. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011, p. 25. [Google Scholar]
- 36.Krieg NR. Class I. Bacteroidia class. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011, p. 25. [Google Scholar]
- 37.Krieg NR. Order I. Bacteroidales ord. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011, p. 25. [Google Scholar]
- 38.Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB. Family III. Rikenellaceae fam. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011, p. 54. [Google Scholar]
- 39.Validation List no. 94. Validation of publication of new names and new combinations previously effectively published outside the IJSEM. Int J Syst Evol Microbiol 2003; 53:1701-1702 10.1099/ijs.0.03001-0 [DOI] [PubMed] [Google Scholar]
- 40.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Stackebrandt E, Ebers J. Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 2006; 33:152-155 [Google Scholar]
- 42.Seng P, Drancourt M, Gouriet F, La Scola B, Fournier PE, Rolain JM, Raoult D. Ongoing revolution in bacteriology: routine identification of bacteria by matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Clin Infect Dis 2009; 49:543-551 10.1086/600885 [DOI] [PubMed] [Google Scholar]
- 43.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Prodigal. http://prodigal.ornl.gov
- 45.GenBank database. http://www.ncbi.nlm.nih.gov/genbank
- 46.Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 1997; 25:955-964 10.1093/nar/25.5.0955 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lagesen K, Hallin P, Rodland EA, Staerfeldt HH, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 2007; 35:3100-3108 10.1093/nar/gkm160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 2004; 340:783-795 10.1016/j.jmb.2004.05.028 [DOI] [PubMed] [Google Scholar]
- 49.Krogh A, Larsson B, von Heijne G, Sonnhammer EL. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 2001; 305:567-580 10.1006/jmbi.2000.4315 [DOI] [PubMed] [Google Scholar]
- 50.Lechner M, Findeib S, Steiner L, Marz M, Stadler PF, Prohaska SJ. Proteinortho: Detection of (Co-)orthologs in large-scale analysis. BMC Bioinformatics 2011; 12:124 10.1186/1471-2105-12-124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Rutherford K, Parkhill J, Crook J, Horsnell T, Rice P, Rajandream MA, Barrell B. Artemis: sequence visualization and annotation. Bioinformatics 2000; 16:944-945 10.1093/bioinformatics/16.10.944 [DOI] [PubMed] [Google Scholar]
- 52.Carver T, Thomson N, Bleasby A, Berriman M, Parkhill J. DNAPlotter: circular and linear interactive genome visualization. Bioinformatics 2009; 25:119-120 10.1093/bioinformatics/btn578 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Darling AC, Mau B, Blattner FR, Perna NT. Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 2004; 14:1394-1403 10.1101/gr.2289704 [DOI] [PMC free article] [PubMed] [Google Scholar]