Abstract
Strain G1T sp. nov. is the type strain of Paenibacillus gorillae a newly proposed species within the genus Paenibacillus. This strain, whose genome is described here, was isolated in France from the fecal sample of a wild western lowland gorilla from Cameroon. P. gorillae is a facultative anaerobic, Gram-negative, rod-shaped bacterium. Here we describe the features of this organism, together with the complete genome sequence and annotation. The 6,257,967 bp long genome (one chromosome but no plasmid) contains 5,856 protein-coding and 62 RNAs genes, including 60 tRNA genes.
Keywords: Paenibacillus gorillae, genome, culturomics, taxonomo-genomics
Introduction
Strain G1T (= CSUR P205 = DSM 26181) is the type strain of Paenibacillus gorillae sp. nov. This bacterium is a Gram-negative, flagellated, facultative anaerobic, indole-negative bacillus that has rounded-ends. It was isolated from the stool sample of Gorilla gorilla subsp. gorilla as part of a culturomics study aiming at cultivating bacterial species found within gorilla feces. By applying large-scale culture conditions, culturomics has previously facilitated the isolation of many new bacterial species from human stool samples [1-3].
The genus Paenibacillus was created by Ash et al. about 20 years ago [4,5]. To date, this genus comprises 145 validly published species [6] of Gram-positive, Gram-negative or variable, mostly motile and spore-forming bacteria. Members of the genus Paenibacillus are ubiquitous bacteria isolated from various environments including soil, water, rhizosphere, food, insect larvae and normal human flora [7]. Moreover, Paenibacillus species were also isolated from or involved in human infections including wound infections, bacteremia and endocarditis [8-13]. Currently, a polyphasic approach that combines proteomics by MALDI-TOF spectral analysis, genomic data and phenotypic characterization is being used as a new approach to describe bacterial species [7,14-25].
Here we present a summary classification and a set of features for P. gorillae sp. nov. strain G1T together with the description of the complete genome sequence and annotation. These characteristics support the circumscription of the species P. gorillae [26].
Classification and features
In July 2011, a fecal sample was collected from a wild western lowland gorilla near Minton, a village in the south-central part of the DJA FAUNAL Park (Cameroon). The collection of the stool sample was approved by the Ministry of Scientific Research and Innovation of Cameroon. No experimentation was conducted on this gorilla. The fecal specimen was preserved at -80°C after collection and sent to Marseille. Strain G1T (Table 1) was isolated in January 2012 by cultivation on Columbia agar with sheep blood 5% (BioMerieux, France). This strain exhibited a 98.28% 16S rRNA nucleotide sequence similarity with Paenibacillus xinjiangensis, the phylogenetically closest validly published Paenibacillus species (Figure 1). This value was lower than the 98.7% 16S rRNA gene sequence threshold recommended by Stackebrandt and Ebers to delineate a new species without carrying out DNA-DNA hybridization [42].
Table 1. Classification and general features of Paenibacillus gorillae strain G1T according to the MIGS recommendations [27].
| MIGS ID | Property | Term | Evidence codea |
|---|---|---|---|
| Domain Bacteria | TAS [28] | ||
| Phylum Firmicutes | TAS [29-31] | ||
| Class Bacilli | TAS [32,33] | ||
| Current classification | Order Bacillales | TAS [34,35] | |
| Family Paenibacillaceae | TAS [33,36] | ||
| Genus Paenibacillus | TAS [4,5,37-39] | ||
| Species Paenibacillus gorillae | IDA | ||
| Type strain G1T | IDA | ||
| Gram stain | negative | IDA | |
| Cell shape | rod-shaped | IDA | |
| Motility | motile | IDA | |
| Sporulation | Sporulating | IDA | |
| Temperature range | mesophilic | IDA | |
| Optimum temperature | 25°C | IDA | |
| MIGS-6.3 | Salinity | growth in BHI medium + 2% NaCl | IDA |
| MIGS-22 | Oxygen requirement | aerobic | IDA |
| Carbon source | varied (see Table 2) | IDA | |
| Energy source | chemoorganoheterotrophic | IDA | |
| MIGS-6 | Habitat | gorilla gut | IDA |
| MIGS-15 | Biotic relationship | free living | IDA |
| MIGS-14 | Pathogenicity Biosafety level Isolation |
unknown 2 gorilla feces |
NAS NAS IDA |
| MIGS-4 | Geographic location | Cameroon | IDA |
| MIGS-5 | Sample collection time | July 2011 | IDA |
| MIGS-4.1 | Latitude | 2.783938 | IDA |
| MIGS-4.1 | Longitude | 13.030472 | IDA |
| MIGS-4.3 | Depth | surface | IDA |
| MIGS-4.4 | Altitude | > 600 m above sea level | IDA |
Evidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from the Gene Ontology project [40]. If the evidence is IDA, then the property was directly observed for a live isolate by one of the authors or an expert mentioned in the acknowledgements.
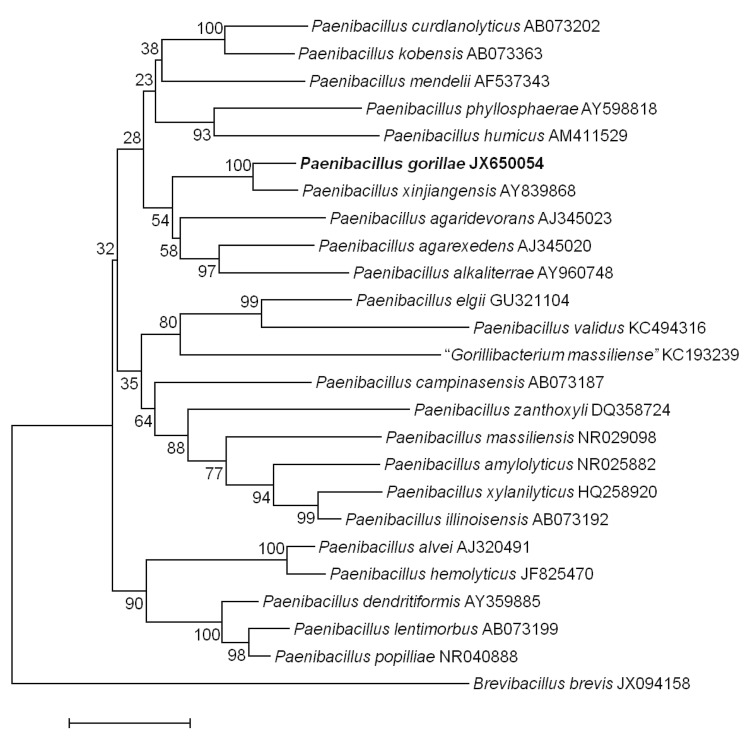
Figure 1.
Phylogenetic tree highlighting the position of Paenibacillus gorillae strain G1T relative to other type strains within the Paenibacillus genus. GenBank accession numbers are indicated in parentheses. Sequences were aligned using CLUSTAL X (V2), and phylogenetic inferences obtained using the maximum-likelihood method within the MEGA 5 software [41]. Numbers at the nodes are percentages of bootstrap values obtained by repeating the analysis 1,000 times to generate a majority consensus tree. Brevibacillus brevis was used as outgroup. The scale bar represents a 2% nucleotide sequence divergence.
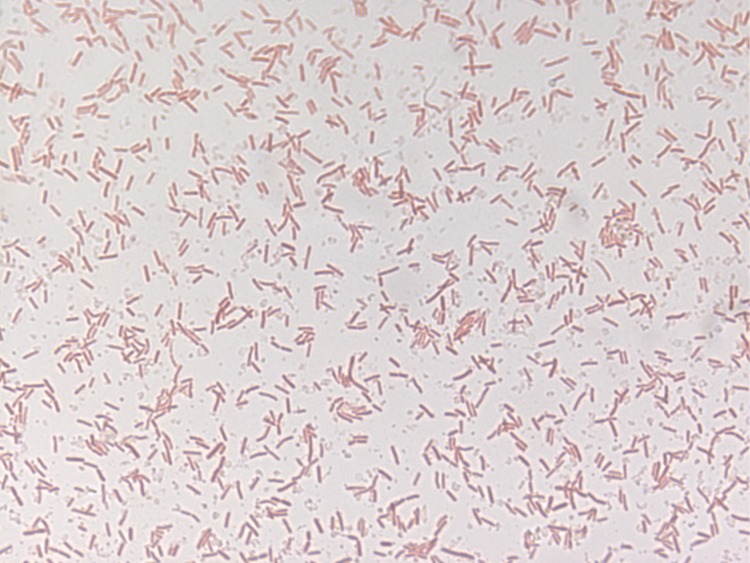
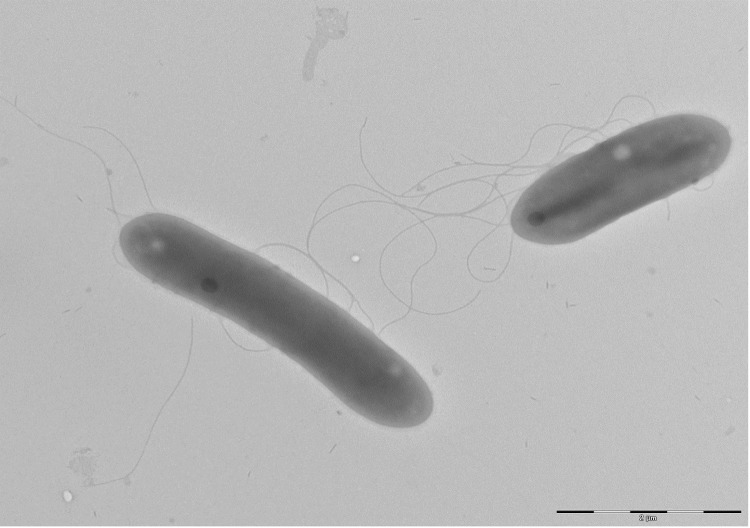
Different growth temperatures (25, 30, 37, 45°C) were tested. Growth occurred for the temperatures (25°C-37°C), but the optimal growth was observed at 25°C. Colonies were 2-8 mm in diameter on Columbia agar, appear whitish in color at 25°C and produce a clear liquid. Growth of the strain was tested under anaerobic and microaerophilic conditions using the GENbag anaer and GENbag microaer systems, respectively (BioMérieux), and in aerobic conditions, with or without 5% CO2. Growth was achieved under aerobic (with and without CO2), microaerophilic and anaerobic conditions. Gram staining showed Gram-negative bacilli (Figure 2). A motility test was positive. Cells grown on agar sporulate and the rods have a length ranging from 2.5 to 3.97 µm (mean 3.2 µm) and a diameter ranging from 0.76 to 0.83 µm (mean 0.79 µm) as determined by negative staining transmission electron microscopy (Figure 3).
Figure 2.
Gram staining of P. gorillae strain G1T
Figure 3.
Transmission electron microscopy of P. gorillae strain G1T, using a Morgani 268D (Philips) at an operating voltage of 60kV. The scale bar represents 2 µm.
Strain G1T exhibited oxidase activity but not catalase activity. Using the API 50CH system (BioMerieux), a positive reaction was observed for D-mannose, amygdalin, L-arabinose, cellobiose, lactose, D-xylose, D-glucose, mannitol, arabinose, xylose, glycerol, D-galactose, N-acetylglucosamine, arbutin, aesculin, D-sorbitol, D-maltose, D-saccharose, D-trehalose, D-tagatose, L-rhamnose, salicin, adonitol, D-melibiose, D-raffinose, D-ribose, D-fructose and hydrolysis of starch. Negative reactions were observed for potassium gluconate, potassium 2-cetogluconate, inulin, D-melezitose, Glycogen, β-gentiobiose, D-turanose, methyl- αD-mannopyranoside and methyl- αD-glucopyranoside. Using the API ZYM system, negative reactions were observed for lipase (C14), a-chymotrypsin, esterase (C4), esterase lipase (C8), naphthyl-AS-BI-phosphohydrolase, phenylalanine arylamidase, leucine arylamidase, cystine arylamidase, valine arylamidase, glycine arylamidase, arginine arylamidase and β-glucosidase. Using the API Coryne system, positive reactions were observed for β-glucuronidase, alkaline phosphatase, α-glucosidase, α-galactosidase and N-acetyl-β-glucosaminidase activities. The urease reaction, nitrate reduction and indole production were negative. P. gorillae is susceptible to imipenem, rifampicin, gentamycin, nitrofurantoin and vancomycin, but resistant to metronidazole, trimethoprim/sulfamethoxazole, ceftriaxone, ciprofloxacin and amoxicillin.
When compared to other Paenibacillus species [43-46] and Brevibacillus brevis [47], P. gorillae sp. nov. strain G1T exhibited the phenotypic differences detailed in Table 2.
Table 2. Differential phenotypic characteristics between P. gorillae sp. nov. strain G1T and phylogenetically close Paenibacillus species.
| Characteristic | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| Gram stain | - | - | var | var | + | +/var |
| Production of | ||||||
| Catalase | - | + | + | + | + | + |
| Oxidase | + | - | - | + | - | + |
| Nitrate reductase | - | + | + | - | + | + |
| Urease | - | + | + | na | na | - |
| Indole | - | - | + | + | na | - |
| Utilization of | ||||||
| D-mannose | + | + | + | - | + | - |
| Amygdalin | + | - | - | - | + | - |
| L-Arabinose | + | w | - | - | + | - |
| Cellobiose | + | + | + | - | + | + |
| D-lactose | + | + | + | na | + | - |
| D-xylose | + | + | var | - | - | - |
| D-glucose | + | + | + | + | + | - |
| D-Mannitol | + | - | + | - | + | + |
| D-Arabinose | + | - | na | na | - | - |
| Glycerol | + | - | var | - | + | - |
| D-Galactose | + | - | + | na | + | + |
| Starch | + | + | + | + | + | - |
| N-acetylglucosamine | + | + | + | na | - | + |
| Arbutin | + | - | na | + | + | na |
| Aesculin | + | + | + | na | + | + |
| D-sorbitol | + | - | na | na | - | - |
| D-maltose | + | + | + | na | + | - |
| D-saccharose | + | + | na | - | + | - |
| D-trehalose | + | + | + | - | + | - |
| D-tagatose | + | - | na | na | - | - |
| Potassium gluconate | - | - | + | - | - | w |
| L-rhamnose | + | - | na | na | - | - |
| Salicin | + | + | - | - | + | - |
| Adonitol | + | - | na | + | - | - |
| D-melibiose | + | + | na | + | + | - |
| D-raffinose | + | + | na | + | + | - |
| D-ribose | + | - | + | na | + | - |
| D-fructose | + | + | w | na | + | - |
| Habitat | Gorilla gut | Gorilla gut | Roots of Perilla frutescens | Honeybee larvae | Human: blood culture | Environment: soil |
Strain: 1, Paenibacillus gorillae G1T; 2, “Gorillibacterium massiliense” G5T; 3, Paenibacillus elgii SD17T; 4, Paenibacillus alvei BCRC 11220T; 5, Paenibacillus massiliensis 2301065 T; 6, Brevibacillus brevis NBRC 15304T.
var: variable, +: positive result, -: negative result, na: data not available, w: weak positive result
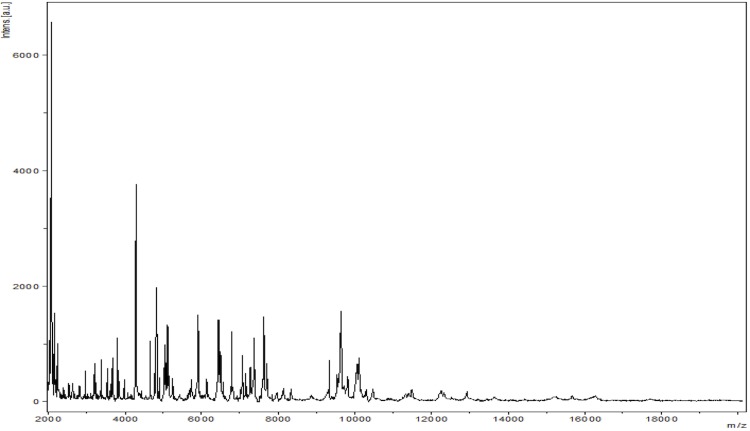
Matrix-assisted laser-desorption/ionization time-of-flight (MALDI-TOF) MS protein analysis was carried out as previously described [14] using a Microflex spectrometer (Bruker Daltonics, Leipzig, Germany). Twelve distinct deposits were made for strain G1T from 12 isolated colonies. The 12 G1T spectra were imported into the MALDI BioTyper software (version 2.0, Bruker) and analyzed by standard pattern matching (with default parameter settings) against 6,252 bacterial spectra including 123 spectra from 67 Paenibacillus species, used as reference data, in the BioTyper database. Interpretation of scores was as follows: a score > 2 to a validly published species enabled the identification at the species level, a score > 1.7 but < 2 enabled the identification at the genus level; and a score < 1.7 did not enable any identification. For strain G1T, the obtained scores ranged from 1.177 to 1.343, thus suggesting that our isolate was not a member of a known species. We incremented our database with the spectrum from strain G1T (Figure 4). Spectrum differences with other of Paenibacillus species are shown in Figure 5.
Figure 4.
Reference mass spectrum from P. gorillae strain G1T. Spectra from 12 individual colonies were compared and a reference spectrum was generated.
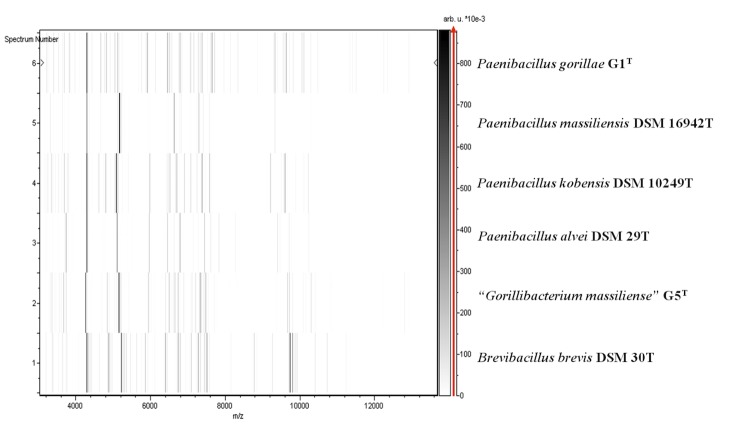
Figure 5.
Gel view comparing Paenibacillus gorillae G1T spectra with other members of the Paenibacillus genus (P. massiliensis, P. kobensis, and P. alvei) and with Brevibacillus brevis and “Gorillibacterium massiliense”. The Gel View displays the raw spectra of all loaded spectrum files arranged in a pseudo-gel like look. The x-axis records the m/z value. The left y-axis displays the running spectrum number originating from subsequent spectra loading. The peak intensity is expressed by a Gray scale scheme code. The color bar and the right y-axis indicate the relation between the color a peak is displayed with and the peak intensity in arbitrary units.
Genome sequencing information
Genome project history
The organism was selected for sequencing on the basis of its phylogenetic position and 16S rRNA similarity to other members of the genus Paenibacillus, and is part of a “culturomics” study of the gorilla flora which aims to isolate all bacterial species within gorilla feces. It is the 44th genome of a Paenibacillus species sequenced and the first genome of Paenibacillus gorillae sp. nov. sequenced. A summary of the project information is shown in Table 3. The Genbank accession number is CBVJ000000000 and consists of 167 contigs (150 large contigs). Table 3 shows the project information and its association with MIGS version 2.0 compliance [48].
Table 3. Project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | High-quality draft |
| MIGS-28 | Libraries used | 454 paired-end 3- kb libraries |
| MIGS-29 | Sequencing platform | 454 GS FLX Titanium |
| MIGS-31.2 | Sequencing coverage | 17.2× |
| MIGS-30 | Assemblers | Newbler version 2.5.3 |
| MIGS-32 | Gene calling method | Prodigal |
| EMBL Date of Release | November 26, 2013 | |
| EMBL ID | CBVJ000000000 | |
| MIGS-13 | Project relevance | Study of the gorilla gut microbiome |
Growth conditions and DNA isolation
P. gorillae sp. nov. strain G1T, CSUR P205, DSM 26181, was grown aerobically on 5% sheep blood-enriched Columbia agar at 25°C. Four Petri dishes were spread and resuspended in 3×500µl of TE buffer and stored at 80°C. Then, 500µl of this suspension were thawed, centrifuged 3 minutes at 10,000 rpm and resuspended in 3×100µL of G2 buffer (EZ1 DNA Tissue kit, Qiagen). A first mechanical lysis was performed by glass powder on the Fastprep-24 device (Sample Preparation system, MP Biomedicals, USA) using 2×20 seconds cycles. DNA was then treated with 2.5µg/µL lysozyme (30 minutes at 37°C) and extracted using the BioRobot EZ1 Advanced XL (Qiagen). The DNA was then concentrated and purified using the Qiamp kit (Qiagen). The yield and the concentration was measured by the Quant-it Picogreen kit (Invitrogen) on the Genios Tecan fluorometer at 50ng/µl.
Genome sequencing and assembly
A shotgun and a 3 kb paired end library were pyrosequenced on the 454_Roche_Titanium. This project was loaded on a 1/4 region for each application on PTP Picotiterplates. The shotgun library was constructed with 500ng of DNA as describes by the manufacturer Roche with the Rapid library Preparation kit for XL+. The concentration of the shotgun library was measured with a TBS fluorometer and determined to be 2.89E+09 molecules/µL. The paired-end library was prepared with 5 µg of bacterial DNA using the DNA fragmentation on a Covaris S-Series (S2) instrument (Woburn, Massachusetts, USA) with an enrichment size at 3.2kb. The DNA fragmentation was visualized with an Agilent 2100 BioAnalyzer on a DNA labchip 7500. The library was constructed according to the 454 GS FLX Titanium paired-end protocol (Roche). Circularization and nebulization were performed and generated a pattern with an optimum at 591bp. After PCR amplification through 17 cycles followed by double size selection, the single stranded paired-end library was quantified using the Quant-it Ribogreen kit (Invitrogen) on a Genios Tecan fluorometer at 691 pg/µL. The library concentration equivalence was calculated as 1.07E+10 molecules/µL. The library was stored at -20°C until further use.
The shotgun XL+ library was clonally amplified with 6 cpb in 2 emPCR reactions. The paired-end library was clonally amplified with 0.5 cpb in 3 emPCR reactions with the GS Titanium SV emPCR Kit (Lib-L) v2 (Roche). The yields of the emPCR were 16.9% and 8.61% respectively, and within the expected yield range of 5 to 20%, as recommended by the Roche procedure.
A total of 790,000 beads for each ¼ region per application were loaded on the GS Titanium PicoTiterPlate PTP Kit 70×75 and sequenced with the GS FLX Titanium Sequencing Kit XLR70 (Roche). The run was performed overnight and then analyzed on the cluster through the gsRunBrowser and Newbler assembler (Roche). A total of 339,189 passed filter wells were obtained and generated 108.46 Mb of sequences with a length average of 330 bp. The passed filter sequences were assembled using Newbler with 90% identity and 40-bp as overlap. The final assembly identified 11 scaffolds with 150 large contigs (>1.5kb) generating a genome size of 6.22 Mb corresponding to a genome coverage of 17.2×.
Genome annotation
Open Reading Frames (ORFs) were predicted using Prodigal [49] with default parameters but the predicted ORFs were excluded if they spanned a sequencing gap region. The predicted bacterial protein sequences were searched against the GenBank database [50] and the Clusters of Orthologous Groups (COG) databases using BLASTP. The tRNAScanSE tool [51] was used to find tRNA genes, whereas ribosomal RNAs were found by using RNAmmer [52] and BLASTn against the GenBank database. ORFans were identified if their BLASTP E-value was lower than 1e-03 for alignment length greater than 80 amino acids. If alignment lengths were smaller than 80 amino acids, we used an E-value of 1e-05.
To estimate the mean level of nucleotide sequence similarity at the genome level between P. gorillae sp nov. strain G1T and other Paenibacillaceae species, we use the Average Genomic Identity of orthologous gene Sequences (AGIOS) program. Briefly, this software combines the Proteinortho software [53] to detect orthologous proteins between genomes compared on a pair-wise basis, then retrieves the corresponding genes and determines the mean percentage of nucleotide sequence identity among orthologous ORFs using the Needleman-Wunsch global alignment algorithm.
Genome properties
The genome 6,257,967 bp long (1 chromosome, but no plasmid) with a 48,80% G+C content (Figure 6 and Table 4). It is composed of 167 contigs (150 large contigs, 11 scaffolds). Of the 5,918 predicted genes, 5,856 were protein-coding genes and 62 were RNAs (1 gene is 16S rRNA, 1 gene is 23S rRNA and 60 are tRNA genes). A total of 4,296 genes (73.36%) were assigned a putative function (by COGS or by NR blast) and 304 genes were identified as ORFans (5.19%). The remaining genes were annotated as hypothetical proteins (917 genes, 15.66%). The distribution of genes into COGs functional categories is presented in Table 5. The properties and statistics of the genome are summarized in Tables 4 and 5.
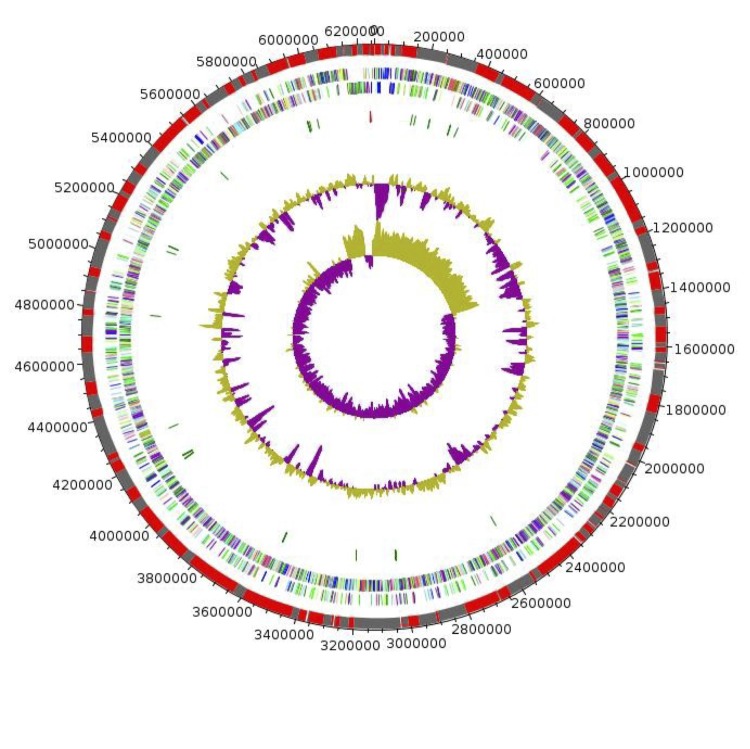
Figure 6.
Graphical circular map of the chromosome. From outside to the center: Genes on the forward strand colored by COG categories (only genes assigned to COG), genes on the reverse strand colored by COG categories (only gene assigned to COG), RNA genes (tRNAs green, rRNAs red), G+C content and GC skew. Purple and olive indicating negative and positive values, respectively.
Table 4. Nucleotide content and gene count levels of the genome.
| Attribute | Value | % of totala |
|---|---|---|
| Genome size (bp) | 6,257,967 | 100 |
| DNA G+C content (bp) | 3,053,870 | 48.80 |
| DNA coding region (bp) | 5,416,322 | 86.55 |
| Total genes | 5,918 | 100 |
| RNA genes | 62 | 1.05 |
| Protein-coding genes | 5,856 | 98.95 |
| Genes with function prediction | 4,296 | 73.36 |
| Genes assigned to COGs | 4,305 | 73.51 |
| Genes with peptide signals | 809 | 13.81 |
| Genes with transmembrane helices | 1,440 | 24.59 |
a The total is based on either the size of the genome in base pairs or the total number of protein coding genes in the annotated genome
Table 5. Number of genes associated with the 25 general COG functional categories.
| Code | Value | %agea | Description |
|---|---|---|---|
| J | 195 | 3.33 | Translation |
| A | 0 | 0 | RNA processing and modification |
| K | 570 | 9.73 | Transcription |
| L | 186 | 3.18 | Replication, recombination and repair |
| B | 1 | 0.02 | Chromatin structure and dynamics |
| D | 31 | 0.53 | Cell cycle control, mitosis and meiosis |
| Y | 0 | 0 | Nuclear structure |
| V | 101 | 1.72 | Defense mechanisms |
| T | 354 | 6.05 | Signal transduction mechanisms |
| M | 247 | 4.22 | Cell wall/membrane biogenesis |
| N | 76 | 1.30 | Cell motility |
| Z | 2 | 0.03 | Cytoskeleton |
| W | 1 | 0.02 | Extracellular structures |
| U | 59 | 1.01 | Intracellular trafficking and secretion |
| O | 127 | 2.17 | Posttranslational modification, protein turnover, chaperones |
| C | 189 | 3.23 | Energy production and conversion |
| G | 591 | 10.09 | Carbohydrate transport and metabolism |
| E | 414 | 7.07 | Amino acid transport and metabolism |
| F | 96 | 1.64 | Nucleotide transport and metabolism |
| H | 125 | 2.13 | Coenzyme transport and metabolism |
| I | 144 | 2.46 | Lipid transport and metabolism |
| P | 365 | 6.23 | Inorganic ion transport and metabolism |
| Q | 147 | 2.51 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 735 | 12.55 | General function prediction only |
| S | 339 | 5.79 | Function unknown |
| - | 1,551 | 26.49 | Not in COGs |
a The total is based on the total number of protein coding genes in the annotated genome
Genomic comparison of P. gorillae and other members of the family Paenibacillaceae.
Here, we compared the genome of P. gorillae strain G1T with those of “G. massiliense” strain G5T, P. elgii strain B69, P. alvei strain DSM 29, P. massiliensis strain DSM 16942 and B. brevis strain NBRC 100599 (Table 6). The draft genome of P. gorillae is larger in size than that of “G. massiliense” (6.25 vs 5.54 Mb) and smaller in size than those of P. elgii, P. alvei, P. massiliensis and B. brevis (6.25 vs 7.96, 6.83, 6.39 and 6.3 Mb respectively). P. gorillae has a lower G+C content than those of “G. massiliense” and P. elgii (48.8% vs 50.39% and 52.6% respectively) but higher than those of P. alvei and B. brevis (48.8% vs 45.9% and 47.3% respectively) and slightly higher than P. massiliensis (48.8% vs 48.5%). The protein content of P. gorillae is lower than that of P. elgii, P. alvei and B. brevis (5,856 vs 7,597, 6,823 and 5,946 respectively) but higher than that of “G. massiliense” and P. massiliensis (5,856 vs 5,146 and 5,496 respectively) (Table 6). In addition, P. gorillae shares 1,987, 2,380, 2,055, 2,121 and 1,935 orthologous genes with “G. massiliense”, P. elgii, P. alvei, P. massiliensis and B. brevis, respectively (Table 7). The nucleotide sequence identity of orthologous genes ranges from 66.3 to 68.7% among previously published genomes, and from 65.7 to 68.6% between P. gorillae and the other studied genomes (Table 7), thus confirming its status as a new species. Table 7 summarizes the number of orthologous genes and the average percentage of nucleotide sequence identity between the different genomes studied.
Table 6. Genomic comparison of P. gorillae sp. nov., strain G1T with four other members of the family Paenibacillaceae†.
| Species | Strain | NCBI accession number | Genome size (Mb) | G+C content |
|---|---|---|---|---|
| Paenibacillus gorillae | G1T | CBVJ000000000 | 6.25 | 48.8 |
| “Gorillibacterium massiliense” | G5T | CBQR000000000 | 5.54 | 50.39 |
| Paenibacillus elgii | B69 | AFHW00000000 | 7.96 | 52.6 |
| Paenibacillus alvei | DSM 29 | AMBZ00000000 | 6.83 | 45.9 |
| Paenibacillus massiliensis | DSM 16942 | ARIL00000000 | 6.39 | 48.5 |
| Brevibacillus brevis | NBRC 100599 | AP008955 | 6.3 | 47.3 |
†Species and strain names, GenBank genome accession numbers, sizes and G+C contents
Table 7. Genomic comparison of P. gorillae sp. nov., strain G1T with four other members of the family Paenibacillaceae. †.
| P. gorillae | “G. massiliense" | P. elgii | P. alvei | P. massiliensis | B. brevis | |
|---|---|---|---|---|---|---|
| P. gorillae | 5,856 | 67.8 | 68.6 | 68 | 67.8 | 65.7 |
| “G. massiliense” | 1,987 | 5,146 | 68.7 | 66.7 | 66.9 | 65.3 |
| P. elgii | 2,380 | 2,122 | 7,597 | 67.6 | 67 | 66.4 |
| P. alvei | 2,055 | 1,846 | 2,336 | 6,823 | 67.9 | 66 |
| P. massiliensis | 2,121 | 1,902 | 2,296 | 1,994 | 5,496 | 65.3 |
| B. brevis | 1,935 | 1,716 | 2,278 | 1,936 | 1,872 | 5,946 |
†numbers of orthologous proteins shared between genomes (lower left triangle), average percentage similarity of nucleotides corresponding to orthologous proteins shared between genomes (upper right triangle). Bold numbers indicate numbers of proteins per genome.
Conclusion
On the basis of phenotypic (Table 2), phylogenetic and genomic analyses (taxonogenomics) (Table 7), we formally propose the creation of Paenibacillus gorillae sp. nov. that contains the strain G1T. This strain has been found in gorilla stool sample collected from Cameroon.
Description of Paenibacillus gorillae sp. nov.
Paenibacillus gorillae (gor.il.lae, N.L. gen fem, of the gorilla from which the stool sample was obtained).
P. gorillae is a facultative aerobic Gram-negative. Optimal growth is achieved aerobically. A weak growth is observed in microaerophilic or anaerobic conditions. Growth occurs between 25 and 37°C, with optimal growth observed at 25°C. Cells stain Gram-negative, are rod-shaped, endospore-forming and motile with a mean diameter of 0.79 µm (range 0.76 to 0.83 µm) and a mean length of 3.2 µm (range 2.5 to 3.97 µm). Colonies are whitish and 2-8 mm in diameter on blood-enriched Columbia agar.
Catalase negative, oxidase positive. Using the API 50CH system (BioMerieux), a positive reaction is obtained for D-mannose, amygdalin, L-arabinose, cellobiose, lactose, D-xylose, D-glucose, mannitol, arabinose, xylose, glycerol, D-galactose, N-acetylglucosamine, arbutin, aesculin, D-sorbitol, D-maltose, D-saccharose, D-trehalose, D-tagatose, L-rhamnose, salicin, adonitol, D-melibiose, D-raffinose, D-ribose, D-fructose and hydrolysis of starch. Negative reactions are obtained for potassium gluconate, potassium 2-cetogluconate, inulin, D-melezitose, Glycogen, β-gentiobiose, D-turanose, Methyl- αD-mannopyranoside and Methyl- αD-glucopyranoside. Using the API ZYM system, negative reactions are obtained for lipase (C14), a-chymotrypsin, esterase (C4), esterase lipase (C8), naphthyl-AS-BI-phosphohydrolase, phenylalanine arylamidase, leucine arylamidase, cystine arylamidase, valine arylamidase, glycine arylamidase, arginine arylamidase and β-glucosidase. Using the API Coryne system, positive reactions are observed for β-glucuronidase, alkaline phosphatase, α-glucosidase, α-galactosidase and N-acetyl-β-glucosaminidase activities. The urease reaction, nitrate reduction and indole production were negative. P. gorillae is susceptible to imipenem, rifampicin, gentamycin, nitrofurantoin and vancomycin, but resistant to metronidazole, trimethoprim/sulfamethoxazole, ceftriaxone, ciprofloxacin and amoxicillin.
The G+C content of the genome is 48.8%. The 16S rRNA and genome sequences are deposited in GenBank under accession numbers JX650054 and CBVJ000000000, respectively. The type strain G1T (= CSUR P205 = DSM 26181) was isolated from the fecal flora of a Gorilla gorilla gorilla from Cameroon.
Acknowledgements
Fadi Bittar was supported by a Chair of Excellence IRD provided by the Institut de Recherche pour le Développement / Méditerranée-Infection Foundation. Mamadou Bhoye Keita was funded by the Méditerranée-Infection foundation. The authors thank the Xegen company for automating the genome annotation process.
References
- 1.Lagier JC, Armougom F, Million M, Hugon P, Pagnier I, Robert C, Bittar F, Fournous G, Gimenez G, Maraninchi M, et al. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin Microbiol Infect 2012; 18:1185-1193 [DOI] [PubMed] [Google Scholar]
- 2.Dubourg G, Lagier JC, Armougom F, Robert C, Hamad I, Brouqui P, Raoult D. The gut microbiota of a patient with resistant tuberculosis is more comprehensively studied by culturomics than by metagenomics. Eur J Clin Microbiol Infect Dis 2013; 32:637-645 10.1007/s10096-012-1787-3 [DOI] [PubMed] [Google Scholar]
- 3.Pfleiderer A, Lagier JC, Armougom F, Robert C, Vialettes B, Raoult D. Culturomics identified 11 new bacterial species from a single anorexia nervosa stool sample. Eur J Clin Microbiol Infect Dis 2013; 32:1471-1481 10.1007/s10096-013-1900-2 [DOI] [PubMed] [Google Scholar]
- 4.Ash C, Priest FG, Collins MD. Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Proposal for the creation of a new genus Paenibacillus. Antonie van Leeuwenhoek 1993; 64:253-260 10.1007/BF00873085 [DOI] [PubMed] [Google Scholar]
- 5.Judicial Commission of the International Committee on Systematics of Prokaryotes The type species of the genus Paenibacillus Ash et al. 1994 is Paenibacillus polymyxa. Opinion 77. Int J Syst Evol Microbiol 2005; 55:513 10.1099/ijs.0.63546-0 [DOI] [PubMed] [Google Scholar]
- 6.Abstract for the genus Paenibacillus NamesforLife, LLC. Retrieved Novembre 1, 2013.
- 7.Mishra AK, Lagier JC, Rivet R, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Paenibacillus senegalensis sp. nov. Stand Genomic Sci 2012; 7:70-81 10.4056/sigs.3056450 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Glaeser SP, Falsen E, Busse HJ, Kämpfer P. Paenibacillus vulneris sp. nov., isolated from a necrotic wound. Int J Syst Evol Microbiol 2013; 63:777-782 10.1099/ijs.0.041210-0 [DOI] [PubMed] [Google Scholar]
- 9.Anikpeh YF, Keller P, Bloemberg GV, Grünenfelder J, Zinkernagel AS. Spacecraft bacterium, Paenibacillus pasadenensis, causing wound infection in humans. BMJ Case Rep 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rieg S, Martin Bauer T, Peyerl-Hoffmann G, Held J, Ritter W, Wagner D, Kern WV, Serr A. Paenibacillus larvae bacteremia in injection drug users. Emerg Infect Dis 2010; 16:487-489 10.3201/eid1603.091457 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ouyang J, Pei Z, Lutwick L, Dalal S, Yang L, Cassai N, Sandhu K, Hanna B, Wieczorek RL, Bluth M, Pincus MR. Case report: Paenibacillus thiaminolyticus: a new cause of human infection, inducing bacteremia in a patient on hemodialysis. Ann Clin Lab Sci 2008; 38:393-400 [PMC free article] [PubMed] [Google Scholar]
- 12.Teng JL, Woo PC, Leung KW, Lau SK, Wong MK, Yuen KY. Pseudobacteraemia in a patient with neutropenic fever caused by a novel paenibacillus species: Paenibacillus hongkongensis sp. nov. Mol Pathol 2003; 56:29-35 10.1136/mp.56.1.29 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ferrand J, Hadou T, Selton-Suty C, Goehringer F, Sadoul N, Alauzet C, Lozniewski A. Cardiac Device-Related Endocarditis Caused by Paenibacillus glucanolyticus. J Clin Microbiol 2013; 51:3439-3442 10.1128/JCM.00864-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Keita MB, Diene S, Robert C, Raoult D, Fournier PE, Bittar F. Non-contiguous finished genome sequence and description of Bacillus massiliogorillae sp. nov. Stand Genomic Sci 2013; 9:93-105 10.4056/sigs.4388124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mishra AK, Lagier JC, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Clostridium senegalense sp. nov. Stand Genomic Sci 2012; 6:386-395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lagier JC, Armougom F, Mishra AK, Ngyuen TT, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Alistipes timonensis sp. nov. Stand Genomic Sci 2012; 6:315-324 10.4056/sigs.2685971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lagier JC, El Karkouri K, Nguyen TT, Armougom F, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Anaerococcus senegalensis sp. nov. Stand Genomic Sci 2012; 6:116-125 10.4056/sigs.2415480 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Roux V, El Karkouri K, Lagier JC, Robert C, Raoult D. Non-contiguous finished genome sequence and description of Kurthia massiliensis sp. nov. Stand Genomic Sci 2012; 7:221-232 10.4056/sigs.3206554 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mishra AK, Lagier JC, Robert C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Peptoniphilus timonensis sp. nov. Stand Genomic Sci 2012; 7:1-11 10.4056/sigs.2956294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hugon P, Ramasamy D, Lagier JC, Rivet R, Couderc C, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of Alistipes obesi sp. nov. Stand Genomic Sci 2013; 7:427-439 10.4056/sigs.3336746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ramasamy D, Lagier JC, Nguyen TT, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of Dielma fastidiosa gen. nov., sp. nov., a new member of the Family Erysipelotrichaceae. Stand Genomic Sci 2013; 8:336-351 10.4056/sigs.3567059 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mishra AK, Lagier JC, Robert C, Raoult D, Fournier PE. Genome sequence and description of Timonella senegalensis gen. nov., sp. nov., a new member of the suborder Micrococcinae. Stand Genomic Sci 2013; 8:318-335 10.4056/sigs.3476977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mishra AK, Hugon P, Lagier JC, Nguyen TT, Couderc C, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of Enorma massiliensis gen. nov., sp. nov., a new member of the Family Coriobacteriaceae. Stand Genomic Sci 2013; 8:290-305 10.4056/sigs.3426906 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ramasamy D, Lagier JC, Gorlas A, Raoult D, Fournier PE. Non contiguous-finished genome sequence and description of Bacillus massiliosenegalensis sp. nov. Stand Genomic Sci 2013; 8:264-278 10.4056/sigs.3496989 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hugon P, Mishra AK, Lagier JC, Nguyen TT, Couderc C, Raoult D, Fournier PE. Non-contiguous finished genome sequence and description of Brevibacillus massiliensis sp. nov. Stand Genomic Sci 2013; 8:1-14 10.4056/sigs.3466975 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sentausa E, Fournier PE. Advantages and limitations of genomics in prokaryotic taxonomy. Clin Microbiol Infect. 2013 doi: 10.1111/1469-0691.12181. [DOI] [PubMed] [Google Scholar]
- 27.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archae, Bacteria, and Eukarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gibbons NE, Murray RGE. Proposals concerning the higher taxa of Bacteria. Int J Syst Bacteriol 1978; 28:1-6 10.1099/00207713-28-1-1 [DOI] [Google Scholar]
- 30.Garrity GM, Holt JG. The Road Map to the Manual. In: Garrity GM, Boone DR, Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 1, Springer, New York, 2001, p. 119-169. [Google Scholar]
- 31.Murray RGE. The Higher Taxa, or, a Place for Everything...? In: Holt JG (ed), Bergey's Manual of Systematic Bacteriology, First Edition, Volume 1, The Williams and Wilkins Co., Baltimore, 1984, p. 31-34. [Google Scholar]
- 32.Ludwig W, Schleifer KH, Whitman WB. Class I. Bacilli class nov. In: De Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 3, Springer-Verlag, New York, 2009, p. 19-20. [Google Scholar]
- 33.List of new names and new combinations previously effectively, but not validly, published. List no. 132. Int J Syst Evol Microbiol 2010; 60:469-472 10.1099/ijs.0.022855-0 [DOI] [PubMed] [Google Scholar]
- 34.Skerman VBD, McGowan V, Sneath PHA. Approved Lists of Bacterial Names. Int J Syst Bacteriol 1980; 30:225-420 10.1099/00207713-30-1-225 [DOI] [PubMed] [Google Scholar]
- 35.Prévot AR. In: Hauderoy P, Ehringer G, Guillot G, Magrou. J., Prévot AR, Rosset D, Urbain A (eds), Dictionnaire des Bactéries Pathogènes, Second Edition, Masson et Cie, Paris, 1953, p. 1-692 [Google Scholar]
- 36.De Vos P, Ludwig W, Schleifer KH, Whitman WB. Family IV. Paenibacillaceae fam. nov. In Bergey's Manual of Systematic Bacteriology, 2nd Edition, vol 3 (The Firmicutes), Springer; New York, 2009, p. 269. [Google Scholar]
- 37.Validation List no. 51. Validation of the publication of new names and new combinations previously effectively published outside the IJSB. Int J Syst Bacteriol 1994; 44:852 10.1099/00207713-44-4-852 [DOI] [Google Scholar]
- 38.Shida O, Takagi H, Kadowaki K, Nakamura LK, Komagata K. Transfer of Bacillus alginolyticus, Bacillus chondroitinus, Bacillus curdlanolyticus, Bacillus glucanolyticus, Bacillus kobensis, and Bacillus thiaminolyticus to the genus Paenibacillus and emended description of the genus Paenibacillus. Int J Syst Bacteriol 1997; 47:289-298 10.1099/00207713-47-2-289 [DOI] [PubMed] [Google Scholar]
- 39.Behrendt U, Schumann P, Stieglmeier M, Pukall R, Augustin J, Spröer C, Schwendner P, Moissl-Eichinger C, Ulrich A. Characterization of heterotrophic nitrifying bacteria with respiratory ammonification and denitrification activity – Description of Paenibacillus uliginis sp. nov., an inhabitant of fen peat soil and Paenibacillus purispatii sp. nov., isolated from a spacecraft assembly clean room. Syst Appl Microbiol 2010; 33:328-336 10.1016/j.syapm.2010.07.004 [DOI] [PubMed] [Google Scholar]
- 40.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol 2011; 28:2731-2739 10.1093/molbev/msr121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Stackebrandt E, Ebers J. Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 2006; 33:152-155 [Google Scholar]
- 43.Kim DS, Bae CY, Jeon JJ, Chun SJ, Oh HW, Hong SG, Baek KS, Moon EY, Bae KS. Paenibacillus elgii sp. nov., with broad antimicrobial activity. Int J Syst Evol Microbiol 2004; 54:2031-2035 10.1099/ijs.0.02414-0 [DOI] [PubMed] [Google Scholar]
- 44.Ueda J, Yamamoto S, Kurosawa N. Paenibacillus thermoaerophilus sp. nov., a moderately thermophilic bacterium isolated from compost. Int J Syst Evol Microbiol 2013; 63:3330-3335 10.1099/ijs.0.048090-0 [DOI] [PubMed] [Google Scholar]
- 45.Lee FL, Kuo HP, Tai CJ, Yokota A, Lo CC. Paenibacillus taiwanensis sp. nov., isolated from soil in Taiwan. Int J Syst Evol Microbiol 2007; 57:1351-1354 10.1099/ijs.0.64764-0 [DOI] [PubMed] [Google Scholar]
- 46.Roux V, Raoult D. Paenibacillus massiliensis sp. nov., Paenibacillus sanguinis sp. nov. and Paenibacillus timonensis sp. nov., isolated from blood cultures. Int J Syst Evol Microbiol 2004; 54:1049-1054 10.1099/ijs.0.02954-0 [DOI] [PubMed] [Google Scholar]
- 47.Takebe F, Hirota K, Nodasaka Y, Yumoto I. Brevibacillus nitrificans sp. nov., a nitrifying bacterium isolated from a microbiological agent for enhancing microbial digestion in sewage treatment tanks. Int J Syst Evol Microbiol 2012; 62:2121-2126 10.1099/ijs.0.032342-0 [DOI] [PubMed] [Google Scholar]
- 48.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Prodigal. http://prodigal.ornl.gov
- 50.GenBank database. http://www.ncbi.nlm.nih.gov/genbank
- 51.Lowe TM, Eddy SR. t-RNAscan-SE: a program for improved detection of transfer RNA gene in genomic sequence. Nucleic Acids Res 1997; 25:955-964 10.1093/nar/25.5.0955 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lagesen K, Hallin P, Rodland EA, Staerfeldt HH, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 2007; 35:3100-3108 10.1093/nar/gkm160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Lechner M, Findeib S, Steiner L, Marz M, Stadler PF, Prohaska SJ. Proteinortho: Detection of (Co-)orthologs in large-scale analysis. BMC Bioinformatics 2011; 12:124 10.1186/1471-2105-12-124 [DOI] [PMC free article] [PubMed] [Google Scholar]