Abstract
RadA is an archaeal orthologue of human recombinase Rad51. This superfamily of recombinases, which also includes eukaryal meiosis-specific DMC1 and remotely related bacterial RecA, form filaments on single-stranded DNA in the presence of ATP and promote a strand exchange reaction between the single-stranded DNA and a homologous double-stranded DNA. Due to its feasibility of getting crystals and similarity (> 40% sequence identity) to eukaryal homologues, we have studied RadA from Methanococcus voltae (MvRadA) as a structural model for understanding the molecular mechanism of homologous strand exchange. Here we show this protein’s ATPase and strand exchange activities are minimal at pH 6.0. Interestingly, MvRadA’s pH dependence is similar to the properties of human Rad51 but dissimilar to that of the well-studied E. coli RecA. A structure subsequently determined at pH 6.0 reveals features indicative of an ATPase-inactive form with a disordered L2 loop. Comparison with a previously determined ATPase-active form at pH 7.5 implies that the stability of the ATPase-active conformation is reduced at the acidic pH. We interpret these results as further suggesting an ordered disposition of the DNA-binding L2 region, similar to what has been observed in the previously observed ATPase-active conformation, is required for promoting hydrolysis of ATP and strand exchange between single- and double-stranded DNA. His-276 in the mobile L2 region was observed to be partially responsible for the pH-dependent activities of MvRadA.
Keywords: ATPase, conformational change, DNA strand exchange, DMC1, homologous recombination, RadA, Rad51, RecA.
INTRODUCTION
Homologous strand exchange promoted by RecA orthologues enables repair of double-stranded DNA breaks which cause replication forks to stall [1-5]. This superfamily of strand exchange proteins [6] is composed of RecA from bacteria [7], RadA from archaea [8], and Rad51 [9] and meiosis-specific DMC1 [10] from eukaryota. Sequence similarity between bacterial RecA proteins and non-bacterial RadA, Rad51 and DMC1 proteins are low (<30% sequence identity). All such recombinases have an ATPase domain preceded by a strand-helix polymerization motif [11, 12]. The ATPase domain also harbours two DNA-binding loops L1 and L2 [13] located along the filament axis. This core design is flanked by a smaller domain which has been implicated in binding double-stranded DNA (dsDNA) [14, 15]. Despite low level of sequence homology, microscopic and crystallographic results have revealed a strikingly similar set of two right-handed helical assemblies: an extended “active” form and a compact “inactive” form [13, 16-22]. It is worth noting that each form includes a heterogeneous collection of filaments with a range of helical pitches. Yeast Rad51 and MvRadA have been the first two such proteins crystallized in the extended filamentous form [23, 24]. In both structures, the ATPase sites were placed between protomers, a conserved structural feature first seen in the EM-reconstructed active filament of E. coli RecA (EcRecA) [25] and later in the crystallized RecA-DNA complex [26]. The full-length MvRadA crystal structures determined thus far, though all have been crystallized in extended forms, have revealed noticeable conformational changes at the ATPase site as well as in the DNA-binding L2 region which includes the L2 loop and its elbows [27]. In the crystallized active RecA-DNA complex, this region is also ordered [26]. We asked the specific question whether homologous strand exchange and ATP hydrolysis promoted by the Rad51/DMC1/RadA proteins also requires the access to a similarly ordered conformation which has been observed in the MvRadA structure determined in the presence of a high concentration of KCl [27]. It appears so as supported by our previous finding on the potassium-dependent DNA strand exchange activity of MvRadA [28] and by several documented studies that implicate potassium dependence by eukaryal Rad51 [29, 30] and DMC1 [31]. It is worth noting that the most intensively studied EcRecA does not appear to be dependent on any monovalent cation. As such, MvRadA is a bona fide prototype of eukaryal Rad51 and DMC1 recombinases. We also noticed that pH-dependent activities are different for EcRecA and human Rad51. EcRecA has been demonstrated to have comparable strand exchange activities at pH 6-8 [32-35]. On the other hand, the strand exchange activity of human Rad51 has been observed to be attenuated in acidic pHs [36]. In order to find the molecular and structural rationalization for such a difference, we studied the pH-dependent ATPase and strand exchange activities of MvRadA and observed a pH-dependent structural difference. Both activities were observed to be minimal at pH 6.0. The RadA / AMP-PNP structure determined at pH 6.0 in the presence of high concentration of DNA-mimicking salt lacks the ordered feature in the L2 region for promoting efficient ATP hydrolysis. The correlation between the pH-dependent activities and structural changes also suggests that an ordered disposition of L2 is likely required for strand exchange.
MATERIALS AND METHODS
Cloning, Protein Preparation and Crystallization
RadA from M. voltae was cloned, expressed, purified, and crystallized as reported [24]. The optimal well solution for crystallization contained 2 mM AMP-PNP, 0.05 M MgCl2, 0.5 M NaCl, 6% PEG 3,350 and 0.05 M Tris-HCl buffer at pH 7.5.
Data Collection and Structure Determination
Crystals were transferred to stabilizing solutions composed of 2 mM AMP-PNP, 0.05 M MgCl2, 0.5 M KCl, 6% PEG 3,350, 24% glycerol and 0.1 M MES-KOH buffer or Tris-Hepes buffer at pH 6.0, soaked for 3 minutes, then transfered by a loop to a 100 K nitrogen stream generated by an Oxford CryoSystems device. The 0.4 degree oscillation images were collected and processed using a Brukers Proteum-R system as described [24]. The previously published MvRadA structure (PDB entries 1T4G) was used as the initial model for rigid body refinement. Each model was subjected to iterative rebuilding using XtalView [37] and refinement using CNS [38]. A 3.2 Å resolution anomalous difference map was analyzed for the location of potassium ions. One ordered potassium site near the triphosphate moiety of AMP-PNP was located at a major peak (~6 σ). Statistics of the X-ray diffraction data, structural refinement and geometry of the final model are shown in Table 1. The molecular figures were generated by Molscript [39] and rendered by Raster3D [40]. The structure factors and the final atomic model have been deposited in the Protein Data Bank (code 2B21).
Table 1.
X-ray diffraction data and structural refinement sta-tistics
| Diffraction data (Cu Kα radiation, wavelength = 1.5418 Å; Space group P61) | |
| PDB code | 2B21 |
| Unit cell dimensions (Å) | a = b = 83.1, c = 107.2 |
| aResolution range (Å) | 30 - 2.4 (2.5 - 2.4) |
| Observed reflections | 74520 |
| Unique reflection | 16344 |
| Completeness | 99.5% (99.1%) |
| bRsym | 0.081 (0.417) |
| I / s | 8.1 (1.8) |
| Anomalous I / σ | 3.1 (2.5) |
| Crystal structure refinement | |
| Reflection with F > 0 | 15073/ 91.8% |
| cR-factor / free R | 0.215 / 0.259 |
| Residues & Nucleotide | 300 & 1 AMP-PNP |
| Solvent molecules | 70 H2O, 2 Mg2+ & 3 K+ |
| rsmd: Bond / Angle | 0.0065 Å / 1.27° |
| Ramachandran | |
| Most favored | 89.0% |
| Disallowed | 0 |
a Values in parentheses refer to values in the highest resolution shell.
b Rsym = Σ| Ih - <I>h | / Σ Ih, where <I>h is average intensity over symmetry equivalents, h is reflection index.
c R-factor = Σ | Fobs - Fcalc | / Σ Fobs. The Free R-factor is calculated using a randomly selected 5% of the reflections set aside from the refinement.
DNA-dependent and Salt-stimulated ATPase Assays
A dye solution composed of Malachite Green, ammonium molybdate in 1.0 M HCl was used to monitor the accumulation of inorganic phosphate [41] from ATP hydrolysis. Absorbance values at 620 nm was used for phosphate quantification. The reaction solutions for the ssDNA-dependent ATPase assay is composed of 3 μM RadA, 18 μM poly-(dT)36 (in nucleotides), 5 mM ATP in a series of buffered solutions (0.05 M Tris-Hepes buffer ranging from pH 6.0 to pH 8.0, 10 mM MgCl2, 0.1 M KCl, and 0.1% v/v 2-mercaptoethanol). The salt-induced ATPase assay contained 1.0 M KCl but did not contain the ssDNA substrate poly-(dT)36. Time course of phosphate release was monitored at 5-minute interval to derive initial turnover rate.
Strand Exchange Assay
The DNA substrates (#1, 63-nt, ACAGCACCAG ATTCAGCAAT TAAGCTCTAA GCCATCCGCA AAAATGACCT CTTATCAAAA GGA; #45A, 36-nt, ACAGCACCAG ATTCAGCAAT TAAGCTCTAA GCCATG; #55A, 36-nt, CATGGCTTAG AGCTTAATTG CTGAATCTGG TGCTGT) were slightly modified from a published study [42]. Three oligonucleotides as well as the strand exchange protocol were used as we have described previously [28]. The pH values of the 50 mM Hepes-Tris buffers (pH 6.0 – 8.0) were recorded for data analysis. DNA #1 was used as the ssDNA substrate. DNA #45A and #55A at a ratio of 1.2: 1.0 was thermally annealed to form the dsDNA substrate. Thermally annealed DNA #1 and #55A was used as the hdDNA reference for quantification of strand exchange yield.
RESULTS
The ssDNA-Dependent ATPase Activity of MvRadA is Negligible at pH 6.0
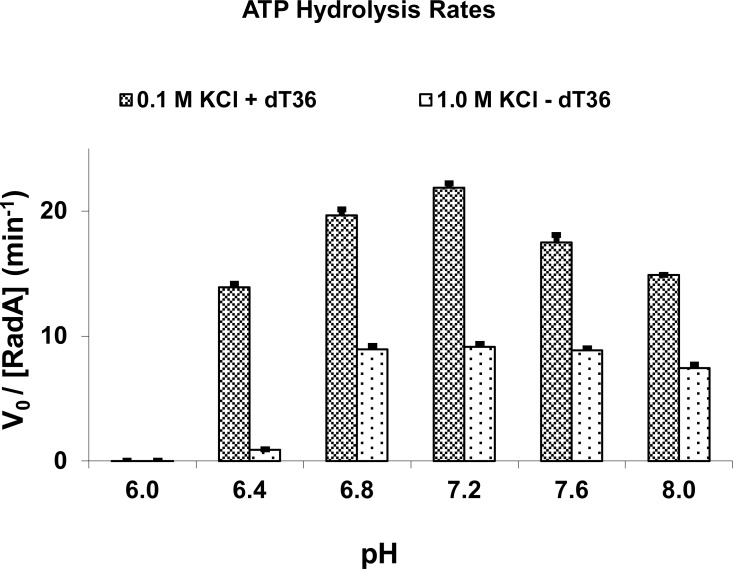
At a low concentration of salt, RecA-like DNA strand exchange proteins are well known to be DNA-dependent ATPases. The single-stranded DNA overhang generated at the site of lesion is believed to be the in vivo site in most organisms [43] for the formation of a recombinase / ssDNA filament which is active in ATP hydrolysis and homologous strand exchange with a homologous dsDNA. We first examined the ssDNA-dependent activity of MvRadA in a pH range between 6.0 and 8.0. In this assay, 3 µM MvRadA and 18 µM poly-(dT)36 (in nucleotides) were used. RecA-like recombinases are known to bind approximately 3 nucleotides per subunit. A 2-fold overdose of poly-(dT)36 was used to ensure maximal ATPase activity. The initial turnover rates were derived (Fig. 1) by monitoring the time courses of phosphate release. The turnover rate was negligible at pH 6.0 (0.013 ± 0.005 min-1). The rate abruptly rose to 13.9 ± 0.3 min-1 at pH 6.4, and stayed above 13.9 min-1 at pH 6.8 - 8.0. The peak activity was observed at pH 7.2 (21.8 ± 0.4 min-1). By setting up hanging drops over pH 5 to 9, we observed that the concentrated protein (~30 mg / ml) was soluble at pH higher than 5.2. Therefore, inactivation at pH 6.0 by protein precipitation could be ruled out.
Figure. (1).
PH-dependent ATPase Activities. The turnover rates were derived by monitoring the time course of phosphate release. All reaction solutions contained 5 mM ATP, 10 mM MgCl2, and 50 mM of Hepes-Tris buffer at indicated pH. ThessDNA-dependent ATP hydrolysis were carried out in the presence of 3 mM MvRadA and 18 mM single-stranded poly-(dT)36 (in nucleotides) and 0.1 M of KCl. The salt-stimulated ATP hydrolysis was carried out in the presence of 3 mM MvRadA and 1.0 M KCl. The activities at pH 6.0 are too low to show. Standard deviations from multiple experiments are shown as error bars.
The Strand Exchange Activity of MvRadA is Negligible at pH 6.0
The strand exchange activity promoted by this archaeal protein was then examined in the same pH range in the presence of 3 mM ATP and 3 mM MgCl2 (Fig. 2A and 2C). A higher concentration of MvRadA (20 µM) and approximately 3 nucleotides / base pairs of DNA substrates per RadA subunit were used to enable facile fluorescence detection on the acrylamide gel. The trend was similar to that of the ATPase activity. At pH 6.0, only ~3% of the 63-nt ssDNA substrate was converted into the 63-nt / 36-nt hdDNA. This level of activity was comparable to basal level activities in the absence of ATP or in the presence of ADP (1-5%, data not shown). The yield abruptly rose to ~23% at pH 6.4 and remained steady around 45% at pH 6.8 - 8.0. In both the ATPase and strand exchange assays, activities close to optimal (within 2 fold) were observed at pH 6.4 - 8.0 while drastically reduced activities were observed at pH 6.0.
Figure. (2).
PH-dependent Strand Exchange Activities. A scheme of the reaction is shown on the top right corner. The acrylamide gels were stained by ethidium bromide. Strand exchange yields were quantified by intensities of fluorescence at bands corresponding to the 36-nt / 63-nt heteroduplex DNA (hdDNA). The pH values are shown atop the gel lanes. A. Strand exchange activity in the presence of an ATP-regenerating system. B. Strand exchange activity in the presence of AMP-PNP. C. Quantified stand exchange yields from A and B.
The High Salt-Stimulated ATPase is Negligible at pH 6.0
Another property share by many RecA homologues is the partial activation of ATPase activity by high concentrations of inorganic salts as ionic substitutes for ssDNA [30, 44, 45]. For MvRadA, KCl has been identified as the substitute for ssDNA, and the ATPase-active filament has been determined [27]. The high salt-stimulated ATPase activity in 1.0 M KCl (Fig. 1), though noticeably lower than the ssDNA-stimulated activity, was observed to somewhat correlate with the stand exchange and ssDNA-dependent ATPase activities. A minimal turnover rate was observed at pH 6.0 (0.03 ± 0.009 min-1). A less dramatic rise to 0.90 ± 0.03 min-1) was observed at pH 6.4. The high salt-stimulated ATPase activities were near constant (7.5-9.1 min-1) at pH 6.8 - 8.0.
The Strand Exchange Activities in the Presence of AMP-PNP is Minimal at pH 6.0
Strand exchange promoted by RecA-like recombinases is known to require the presence of magnesium and nucleoside triphosphate cofactors but not ATP hydrolysis per se. We further studied strand exchange activities of MvRadA in the presence of AMP-PNP, a stable ATP analogue. In the presence of 3 mM AMP-PNP, however, a higher concentration of MgCl2 (10 mM) was found to be required for optimal yield. Higher yields were observed than those in the presence of ATP (Fig. 2A-C). Approximately 20% of ssDNA was converted into hdDNA at pH 6.0, which is significantly higher than the basal level activity. Nevertheless, at pH 6.4-6.8, the yields were at least two folds as high as that of pH 6.0.
Crystal Structure of MvRadA at pH 6.0
Based on the similar trends of these assays, we reasoned that there may be a common structural property that governs not only ATP hydrolysis but also DNA strand exchange. It is worth noting that similar pH dependence, though not as pronounced, was observed in the presence of AMP-PNP (Fig. 2B), a stable ATP analogue for crystallographic study. We further reasoned that this structural property could be addressed by a feasible experiment of incubating MvRadA crystals in a high salt solution, as a DNA mimicking agent, at pH 6.0.
Our previous soaking and co-crystallization experiments with MvRadA / AMP-PNP crystals in an ATPase-activating dose of a potassium salt at a pH range between 7.0 and 8.0 had reproducibly resulted in an ATPase-active conformation (PDB code 1XU4, Fig. 3A) with the highest quality data determined at 2.0 Å resolution [27, 46]. We were unable to get sizeable crystals at pH lower than 7.0, though showers of micro-crystals were observed. As a result, a soaking experiment using crystals grown at pH 7.5 was carried out. The ATPase activity of MvRadA in the presence of 0.4 M KCl is observed to be over 90% of the maximum reached at 0.8 – 1.6 M KCl. The crystals, however, could not survive soaking in stabilizing solutions with over 0.8 M KCl. Structures determined after soaking at pH 6.0 in the presence of 0.5 M KCl were also reproducible. Soaking in both Tris-Hepes (pH 6.0) and Mes-KOH (pH 6.0) buffers produced essentially identical structures except for differences in resolution from crystal to crystal. The resulting ATPase site is shown in (Fig. 3B). The best data set of pH 6.0 MvRadA / AMP-PNP complex soaked with Mes-KOH buffer was refined to 2.4 Å resolution. Compared with the ATPase-active filament at pH 7.5, the helical filament pitch of this pH 6.0 form is noticeably elongated from 105.4 to 107.2 Å. In both soaked crystals, the ATP analogue was tethered between MvRadA protomers. One subunit (yellow subunit, Fig. 3) binds the ATP analogue and an octahedral Mg2+ largely through its conserved P-loop (residues Gly-105 to Thr-112) [47] and the base-stacking Arg-158. The adjacent subunit (gray subunit, Fig. 3) contributes the ATP cap (residues Asp-302 to Asp-308) and the C-terminal portion of the L2 region (residues Asn-256 to Arg-285). In this manner, ATP bridges the filamentous assembly of MvRadA. The major differences lie in the L2 region, solvent entities near the triphosphate and positions of catalytic residues Glu-151 [13] and Gln-257 [48]. In the previously determined ATPase-active form at pH 7.5, two potassium ions and His-280 directly contact the γ-phosphate (2.6 – 2.9 Å), and both catalytic residues Glu-151 and Gln-257 form hydrogen bonds with a candidate for the nucleophilic water (big green sphere, Fig. 3A). We have interpreted these features as essential elements for promoting ATP hydrolysis. In the pH 6.0 structure, however, only one potassium ion (purple spheres, Fig. 3B) was located by anomalous signals between the γ-phosphate of the ATP analogue and the side chain of Asp-302. And the K+ ion was found too distant (3.4 Å) from the γ-phosphate to adequately polarize the phosphate for hydrolysis. The candidate for the nucleophilic water bound by Glu-151 was further away from the γ-phosphorous atom, with distance rising from 3.7 Å to 4.5 Å. It also lost hydrogen bond contact with the terminal phosphate. Unlike the largely ordered L2 region in the ATPase-active form, the entire L2 region in the pH 6.0 form was noticeably more disordered. There were no electron densities for a long stretch of residues from Ala-260 to Val-278. Notably, His-280 was located far apart from the ATP analogue. As such, the polarizing effect by His-280 is also lost. The catalytic side chains of Glu-151 and Gln-257 were found in different conformations. The Gln-257 side chain is too distant to form a hydrogen bond with the nucleophile candidate (big green sphere, Fig. 3B). The root-mean-square difference between the 300 ordered Cα atoms of the pH 6.0 and pH 7.5 structures is 0.65 Å. The deviation of the pH 6.0 structure from previously determined ATPase-inactive forms fall between 0.17 and 0.25 Å. Even for the ordered parts of the most flexible L2 region, the differences are within 1.0 Å. We conclude that the pH 6.0 conformation closely resemble the recurrent ATPase-inactive conformation determined in the presence of ADP or in the absence of a high concentration of a potassium salt [46]. Therefore, the pH 6.0 structure does not represent a novel conformation. Rather, we interpret the structural finding as suggesting the ATPase-active conformation is destabilized at pH 6.0.
Figure. (3).
ATPase Site in Stereo. Two recombinase subunits are coloured in yellow and grey, respectively. K+ ions, Mg2+ ions and water molecules are coloured in purple, red and green, respectively. The putative hydrolysis water in each structure is shown in a bigger sphere. Selected hydrogen bonds and metal-coordination bonds are shown in dashed lines in brown. A. ATPase site previously seen in a KCl-soaked crystal at pH 7.5. B. ATPase site seen in KCl-soaked crystals at pH 6.0. At the acidic pH, the direct contacts between the g-phosphate moiety of the ATP analogue and the electron-withdrawing potassium ions and His-280 side chain are lost.
Properties of H280N and H276N Mutant Proteins
The well-studied E. coli RecA has been demonstrated to have comparable strand exchange activities at pH 6 - 8 [32-35]. In these studies, mutations at two residues (G160N and H163A) within the DNA-binding L1 loop of E. coli RecA has been shown to exhibit pH dependencies opposite to that of MvRadA with attenuated strand exchange activity at higher rather than lower pH. The strand exchange activity of human Rad51, on the other hand, has been shown to exhibit pH dependence similar to that of MvRadA [36]. Finally, we ask which chargeable group in MvRadA may be responsible for their similar pH dependencies.
Though we could not rule out other protonatable residues, the His-280 and His276, as the only two His residues in the mobile L2 region, are primary candidates for affecting the pH dependence. We have previously made an H280N mutant protein [46]. Despite its ability to fold properly and even form micro-crystals, the H280N mutant protein showed neither detectable ATPase activity nor strand exchange activity [46]. A previous mutagenesis study at an analogous site of yeast Rad51 [23] has also demonstrated that an Ala substitution for the equivalent His abolishes DNA binding. The sensitivity to mutation is not surprising since this ATP-contacting His residue is invariable in all archaeal and eukaryal orthologues.
The other His residue in the L2 region of MvRadA, His-276, (Fig. 3) is conserved in approximately three-fourths of known RadA / Rad51 / DMC1 sequences. The remaining ~1/4 sequences have an Asn at the equivalent positions. We therefore made an H276N mutant RadA. This mutant protein’s peak ssDNA-dependent ATPase activity (Fig. 4A) was observed to be 19.9 ± 0.1 min-1, approximately 90% of the wild-type MvRadA (21.8 ± 0.4 min-1). Interestingly, the ssDNA-dependent ATPase activity of the H276N protein appeared less dependent on pH. Even at pH 6.0, it recorded a turnover rate (8.22 ± 0.08 min-1) ~40% of its peak rate at pH 7.2. In contrast, the wild-type protein exhibited a negligible turnover rate. The salt-stimulated ATPase activity in the presence of 1.0 M KCl (Fig. 4A) also peaked at pH 7.2 (8.3 ± 0.2 min-1), again ~90% of the wild-type protein (9.1 ± 0.3 min-1). At pH 6.0, the wild-type and the H276N proteins both exhibited turnover rates less than 0.2 min-1, a minimum level for our assay to precisely quantify. At pH 6.4, however, the salt-stimulated ATPase activity of the mutant protein was ~3 fold as active as the wild-type protein (3.0 versus 0.9 min-1). It appears that the salt-stimulated ATPase activity of the H276N protein is also less dependent on pH than the wild-type protein. This mutant protein was also active in promoting strand exchange in the presence of either ATP or the non-hydrolysable AMP-PNP (Fig. 4B). In the presence of AMP-PNP, however, the existence of multiple bands on the agarose gel appears to indicate that the DNA strand exchange reaction was partially stalled before resolving various ssDNA-dsDNA joint molecules. Meaningful quantification was therefore infeasible for reactions in the presence of AMP-PNP. As observed for the ATPase activity of this mutant protein, the strand exchange yields in the presence of ATP (Fig. 4C and 4D) also exhibited less pronounced pH dependence. Though only about half as high as the wild-type RadA on average, the strand exchange yields promoted by the H276N protein at pH 6.0 and 6.4 were nearly 50% higher than those at pH 6.8 to 8.0. In contrast, the wild-type protein was largely inefficient at pH 6.0. Collectively, we interpret the attenuated pH dependence of the H276N protein as suggesting ionization at the His-276 side chain is partially responsible for causing the pH dependence of the wild-type MvRadA.
Figure. (4).
PH-dependent Activities of H276N Protein. Activities were measured and shown for the H276N protein as in Figure 1 and 2. A. ATP hydrolysis rates in the presence of poly-(dT)36 or high concentration of KCl. B. Strand exchange activities in the presence of ATP and AMP-PNP at pH 7.2. The reaction seems stalled in the presence of AMP-PNP. C. Strand exchange activities in the presence of an ATP-regenerating system. D. Quantified stand exchange yields from C as well as from Figure 2A. The H276N protein exhibited attenuated pH dependence.
DISCUSSION
RecA recombinase and its orthologs promote DNA strand exchange in the presence of ATP and magnesium ion. The DNA bound along the axis of such recombinase filaments is in an extended state competent for DNA strand exchange [26, 49]. The formation of such nucleoprotein filaments possibly requires the participation of both L1 and L2 regions. The ATPase-active conformation appears to have well dispositioned L1 and L2 regions along the filament axis (Fig. 5A), a feature suitable for optimal interaction with DNA. We have previously observed that the ATPase and strand exchange activities of MvRadA are both K+-dependent and correlated with accessibility to the ATPase-active conformation [28]. Our structural and functional studies on the pH dependence of MvRadA reveal a similar correlation. At pH 6.0, activities of MvRadA are much lower and the crystallized filament is in an ATPase-inactive conformation with a disordered L2 loop. Consistent with known requirements for cofactors but not ATP hydrolysis per se, the ordered L2 region is likely required in both polarizing ATP and promoting DNA strand exchange. The correlation between pH-dependent activities and conformational differences further supports the notion that L2 play an important role in binding DNA and / or promoting strand exchange.
Figure. (5).
DNA-Binding Site in Stereo. A. A crystallized MvRadA filament with subunits shown in alternating colours. The ATP analogues are shown in cyan. Three DNA strands (structure unknown) are depicted along the helical groove near the axis. B. A stereo view of one MvRadA protomer. The L1 region is highlighted in green. Ordered parts of the L2 regions are highlighted (magenta at pH 6.0 and blue at pH 7.5). The filament axis lies vertically. Two AMP-PNP molecules and the side chains of Arg-218, Arg-224, Arg-230, His-276 and His-280 are shown in ball-and-stick model. Residues 260 to 278 in the L2 region of the pH 6.0 structure were disordered in the crystal. C. An enlarged view of B. Both His-276 and His-280 are located in a short helix within the mobile L2 region.
The DNA-binding L1 and L2 regions in ATPase-active form of MvRadA enclose a cleft where three Arg residues (218, 224 and 230) in the L1 region are located (Fig. 5). Unlike RecA which does not have a conserved Lys or Arg in its L1 region, the three Arg residues in the L1 region of MvRadA are invariable in the Rad51 / DMC1 / RadA group. The equivalent residues of MvRadA’s Arg-224 in two hyperthermophiles have been shown to be important for binding dsDNA and promoting strand exchange [12]. Mutations at the counterparts of all three Arg residues in human DMC1 have also been shown to be defective in promoting strand exchange [50]. As such, the axial groove lined with the L1 and L2 regions enclose a DNA-binding site. In the ATPase-active conformation observed at pH 7.5, the helix (residues 275 to 282) harbouring His-280 and His-276 appears to bridge ATP at one end and anchor a largely ordered L2 loop at the other. Though the pH 6.0 structure does not represent a novel conformation, the recurrence of an ATPase-inactive conformation during repeated crystal soaking experiment appears to suggest that the ATPase-active conformation is destabilized at pH 6.0. The minimal activities of MvRadA at pH 6.0, along with a disordered L2 loop in the pH 6.0 structure, supports the notion that a well-dispositioned L2 loop may be required for promoting strand exchange as reviewed [51]. As seen in the ATPase-active conformation of MvRadA, the positive end of the dipole moment caused by the short helix in the L2 region is pointing to the axial groove along with the three L1 Arg residues (residues 218, 224, 230, Fig. 5B, 5C). Such a spatial arrangement of conserved L1 and L2 residues has a clear electrostatic feature suitable for clamping DNA in between. The positive charges near the N-terminus of this helix is also a destabilizing factor for the formation of the helix in the absence of anionic DNA or high concentration of salt. Protonation at lower pH would add excessive positive charge to this location and possibly further destabilize this helix. In the absence of K+ or at pH 6.0, the short helix unfolds and the strand exchange activity of MvRadA is much lower and a largely disordered L2 region has been observed.
Finally, we ask which conserved chargeable group in RadA, Rad51 and DMC1 may be responsible for their similar pH dependencies and whether sequence variation can explain the difference with EcRecA. Though we could not rule out other protonatable groups such as His-280, the His-276 side chain appears partially responsible. This His residue sits at the N-terminus of the helical segment within the mobile L2 region. A positively charged His-276 at low pH should destabilize this short helix.
CONCLUSION
RecA-like strand exchange proteins are fascinating systems for studying the correlation between assembly, conformation and activity. Full-length and truncated MvRadA crystallizes in helical filaments of different pitches [24, 52] and in ring-shaped hexamers [53]. The conformation of MvRadA remains mobile in the crystallized filaments, thus providing us a feasible platform to correlate structure with function. We have previously observed noticeable conformational changes especially in the ordering of the L2 region in response to activating dose of potassium or calcium ions [27, 54]. Here we demonstrate lack of such ordering of the L2 region at an acidic pH where the protein is inactive. The correlation between pH-dependent activities and structural changes provided further support to the notion that the ATPase-active and strand exchange-active conformations resemble each other and are more ordered than the inactive conformation.
ACKNOWLEDGEMENTS
We thank Drs. Gabriele Schatte and Wilson Quail at the Saskatchewan Structural Sciences Centre for their assistances to use the X-ray facility. We also thank Xinguo Qian, Yujiong He, Xinfeng Ma for their technical support.
ABBREVIATIONS
- MvRadA
= RadA recombinase from Methanococcus voltae
- EcRecA
= RecA recombinase from E. coli
- ATP
= adenosine 5'-triphosphate
- ssDNA
= single-stranded DNA
- dsDNA
= double-stranded DNA
- hdDNA
= heteroduplex DNA
- AMP-PNP
= 5'-adenylyl imidodiphosphate
CONFLICT OF INTEREST
This work was supported by Saskatchewan Health Research Foundation (SHRF) Establishment Grant 1425, Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant 161981-03, Canadian Institutes of Health Research (CIHR) Operating Grant 63860, and SHRF Phase 3 Team Grant to Molecular Design Research Group at University of Saskatchewan which provided financial support to first author Dr. Rao.
REFERENCES
- 1.Cox MM. A broadening view of recombinational DNA repair in bacteria. Genes Cells. 1998;3(2):65–78. doi: 10.1046/j.1365-2443.1998.00175.x. [DOI] [PubMed] [Google Scholar]
- 2.Cox MM, Goodman MF, Kreuzer KN, Sherratt DJ, Sandler SJ, Marians KJ. The importance of repairing stalled replication forks. Nature. 2000;404(6773):37–41. doi: 10.1038/35003501. [DOI] [PubMed] [Google Scholar]
- 3.Courcelle J, Ganesan AK, Hanawalt PC. Therefore, what are recombination proteins there for?. Bioessays. 2001;23(5):463–70. doi: 10.1002/bies.1065. [DOI] [PubMed] [Google Scholar]
- 4.Lusetti SL, Cox MM. The bacterial RecA protein and the recombinational DNA repair of stalled replication forks. Annu. Rev. Biochem. 2002;71:71–100. doi: 10.1146/annurev.biochem.71.083101.133940. [DOI] [PubMed] [Google Scholar]
- 5.Kowalczykowski SC. Initiation of genetic recombination and recombination-dependent replication. Trends Biochem Sci. 2000;25(4):156–65. doi: 10.1016/s0968-0004(00)01569-3. [DOI] [PubMed] [Google Scholar]
- 6.Seitz EM, Kowalczykowski SC. The DNA binding and pairing preferences of the archaeal RadA protein demonstrate a universal characteristic of DNA strand exchange proteins. Mol. Microbiol. 2000;37(3):555–60. doi: 10.1046/j.1365-2958.2000.02009.x. [DOI] [PubMed] [Google Scholar]
- 7.Clark AJ, Margulies AD. Isolation and Characterization of Recombination-Deficient Mutants of Escherichia Coli K12. Proc. Natl. Acad. Sci. USA. 1965;53:451–9. doi: 10.1073/pnas.53.2.451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sandler SJ, Satin LH, Samra HS, Clark AJ. recA-like genes from three archaean species with putative protein products similar to Rad51 and Dmc1 proteins of the yeast Saccharomyces cerevisiae. Nucleic Acids Res. 1996;24(11):2125–32. doi: 10.1093/nar/24.11.2125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Shinohara A, Ogawa H, Ogawa T. Rad51 protein involved in repair and recombination in S.cerevisiae is a RecA-like protein. Cell. 1992;69(3):457–70. doi: 10.1016/0092-8674(92)90447-k. [DOI] [PubMed] [Google Scholar]
- 10.Bishop DK, Park D, Xu L, Kleckner N. DMC1: a meiosis-specific yeast homolog of E.coli recA required for recombintion synaptonemal complex formation, and cell cycle progression. Cell. 1992;69(3):439–56. doi: 10.1016/0092-8674(92)90446-j. [DOI] [PubMed] [Google Scholar]
- 11.Pellegrini L, Yu DS, Lo T, Anand S, Lee M, Blundell TL, Venkitaraman AR. Insights into DNA recombination from the structure of a RAD51-BRCA2 complex. Nature. 2002;420(6913):287–93. doi: 10.1038/nature01230. [DOI] [PubMed] [Google Scholar]
- 12.Shin DS, Pellegrini L, Daniels DS, Yelent B, Craig L, Bates D, Yu DS, Shivji MK, Hitomi C, Arvai AS, Volkmann N, Tsuruta H, Blundell TL, Venkitaraman AR, Tainer JA. Full-length archaeal Rad51 structure and mutants: mechanisms for RAD51 assembly and control by BRCA2. Embo J. 2003;22(17):4566–76. doi: 10.1093/emboj/cdg429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Story RM, Weber IT, Steitz TA. The structure of the E.coli recA protein monomer and polymer. Nature. 1992;355(6358):318–25. doi: 10.1038/355318a0. [DOI] [PubMed] [Google Scholar]
- 14.Aihara H, Ito Y, Kurumizaka H, Yokoyama S, Shibata T. The N-terminal domain of the human Rad51 protein binds DNA: structure and a DNA binding surface as revealed by NMR. J. Mol. Biol. 1999;290(2):495–504. doi: 10.1006/jmbi.1999.2904. [DOI] [PubMed] [Google Scholar]
- 15.Aihara H, Ito Y, Kurumizaka H, Terada T, Yokoyama S, Shibata T. An interaction between a specified surface of the C-terminal domain of RecA protein and double-stranded DNA for homologous pairing. J. Mol. Biol. 1997;274(2):213–21. doi: 10.1006/jmbi.1997.1403. [DOI] [PubMed] [Google Scholar]
- 16.Ogawa T, Yu X, Shinohara A, Egelman EH. Similarity of the yeast RAD51 filament to the bacterial RecA filament. Science. 1993;259(5103):1896–9. doi: 10.1126/science.8456314. [DOI] [PubMed] [Google Scholar]
- 17.Yang S, VanLoock MS, Yu X, Egelman EH. Comparison of bacteriophage T4 UvsX and human Rad51 filaments suggests that RecA-like polymers may have evolved independently. J. Mol. Biol. 2001;312(5):999–1009. doi: 10.1006/jmbi.2001.5025. [DOI] [PubMed] [Google Scholar]
- 18.Yang S, Yu X, Seitz EM, Kowalczykowski SC, Egelman EH. Archaeal RadA protein binds DNA as both helical filaments and octameric rings. J. Mol. Biol. 2001;314(5):1077–85. doi: 10.1006/jmbi.2000.5213. [DOI] [PubMed] [Google Scholar]
- 19.Yu X, Jacobs SA, West SC, Ogawa T, Egelman EH. Domain structure and dynamics in the helical filaments formed by RecA and Rad51 on DNA. Proc. Natl. Acad. Sci. USA. 2001;98(15):8419–24. doi: 10.1073/pnas.111005398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Datta S, Prabu MM, Vaze MB, Ganesh N, Chandra NR, Muniyappa K, Vijayan M. Crystal structures of Mycobacterium tuberculosis RecA and its complex with ADP-AlF(4): implications for decreased ATPase activity and molecular aggregation. Nucleic Acids Res. 2000;28(24):4964–73. doi: 10.1093/nar/28.24.4964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Datta S, Krishna R, Ganesh N, Chandra NR, Muniyappa K, Vijayan M. Crystal structures of Mycobacterium smegmatis RecA and its nucleotide complexes. J. Bacteriol. 2003;185(14):4280–4. doi: 10.1128/JB.185.14.4280-4284.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Rajan R, Bell CE. Crystal structure of RecA from Deinococcus radiodurans: insights into the structural basis of extreme radioresistance. J. Mol. Biol. 2004;344(4):951–63. doi: 10.1016/j.jmb.2004.09.087. [DOI] [PubMed] [Google Scholar]
- 23.Conway AB, Lynch TW, Zhang Y, Fortin GS, Fung CW, Symington LS, Rice PA. Crystal structure of a Rad51 filament. Nat. Struct. Mol. Biol. 2004;11(8):791–6. doi: 10.1038/nsmb795. [DOI] [PubMed] [Google Scholar]
- 24.Wu Y, He Y, Moya IA, Qian X, Luo Y. Crystal Structure of Archaeal Recombinase Rad A Snapshot of Its Extended Conformation. Mol. Cell. 2004;15(3):423–435. doi: 10.1016/j.molcel.2004.07.014. [DOI] [PubMed] [Google Scholar]
- 25.VanLoock MS, Yu X, Yang S, Lai AL, Low C, Campbell MJ, Egelman EH. ATP-mediated conformational changes in the RecA filament. Structure (Camb): 2003;11(2):187–96. doi: 10.1016/s0969-2126(03)00003-0. [DOI] [PubMed] [Google Scholar]
- 26.Chen Z, Yang H, Pavletich NP. Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures. Nature. 2008;453(7194):489–4. doi: 10.1038/nature06971. [DOI] [PubMed] [Google Scholar]
- 27.Wu Y, Qian X, He Y, Moya IA, Luo Y. Crystal structure of an ATPase-active form of Rad51 homolog from Methanococcus voltae.Insights into potassium dependence. J. Biol. Chem. 2005;280 (1):722–8. doi: 10.1074/jbc.M411093200. [DOI] [PubMed] [Google Scholar]
- 28.Qian X, He Y, Wu Y, Luo Y. Asp302 Determines Potassium Dependence of a RadA Recombinase from Methanococcus voltae. J. Mol. Biol. 2006;360(3):537–47. doi: 10.1016/j.jmb.2006.05.058. [DOI] [PubMed] [Google Scholar]
- 29.Rice KP, Eggler AL, Sung P, Cox MM. DNA pairing and strand exchange by the Escherichia coli RecA and yeast Rad51 proteins without ATP hydrolysis on the importance of not getting stuck. J. Biol Chem. 2001;276(42):38570–81. doi: 10.1074/jbc.M105678200. [DOI] [PubMed] [Google Scholar]
- 30.Liu Y, Stasiak AZ, Masson JY, McIlwraith MJ, Stasiak A, West SC. Conformational changes modulate the activity of human RAD51 protein. J. Mol Biol. 2004;337(4):817–27. doi: 10.1016/j.jmb.2004.02.022. [DOI] [PubMed] [Google Scholar]
- 31.Sehorn MG, Sigurdsson S, Bussen W, Unger VM, Sung P. Human meiotic recombinase Dmc1 promotes ATP-dependent homologous DNA strand exchange. Nature. 2004;429(6990):433–7. doi: 10.1038/nature02563. [DOI] [PubMed] [Google Scholar]
- 32.Pinsince JM, Muench KA, Bryant FR, Griffith JD. Two mutant RecA proteins possessing pH-dependent strand exchange activity exhibit pH-dependent presynaptic filament formation. J. Mol. Biol. 1993;233(1):59–66. doi: 10.1006/jmbi.1993.1484. [DOI] [PubMed] [Google Scholar]
- 33.Meah YS, Bryant FR. Activation of a recombinase-deficient mutant recA protein with alternate nucleoside triphosphate cofactors. J. Biol. Chem. 1993;268(32):23991–6. [PubMed] [Google Scholar]
- 34.Muench KA, Bryant FR. Disruption of an ATP-dependent isomerization of the recA protein by mutation of histidine 163. J. Biol. Chem. 1991;266(2):844–50. [PubMed] [Google Scholar]
- 35.Muench KA, Bryant FR. An obligatory pH-mediated isomerization on the [Asn-160]recA protein-promoted DNA strand exchange reaction pathway. J. Biol. Chem. 1990;265(20):11560–6. [PubMed] [Google Scholar]
- 36.Baumann P, West SC. The human Rad51 protein: polarity of strand transfer and stimulation by hRP-A. Embo J. 1997;16(17):5198–206. doi: 10.1093/emboj/16.17.5198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.McRee DE. XtalView/Xfit-A versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 1999;125(2-3):156–65. doi: 10.1006/jsbi.1999.4094. [DOI] [PubMed] [Google Scholar]
- 38.Brunger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang JS, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 1998;54 (Pt 5):905–21. doi: 10.1107/s0907444998003254. [DOI] [PubMed] [Google Scholar]
- 39.Kraulis P. MOLSCRIPT:a program to produce both detailed and schematic plots of protein structures. J. Appl. Cryst. 1991;24:946–950. [Google Scholar]
- 40.Bacon DJ, Anderson WF. A Fast Algorithm for Rendering Space-Filling Molecule Pictures'. J. Molec. Graphics. 1988;6:219–220. [Google Scholar]
- 41.Itaya K, Ui M. A new micromethod for the colorimetric determination of inorganic phosphate. Clin. Chim. Acta. 1966;14(3):361–6. doi: 10.1016/0009-8981(66)90114-8. [DOI] [PubMed] [Google Scholar]
- 42.Mazin AV, Zaitseva E, Sung P, Kowalczykowski SC. Tailed duplex DNA is the preferred substrate for Rad51 protein-mediated homologous pairing. Embo J. 2000;19(5):1148–56. doi: 10.1093/emboj/19.5.1148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kim JI, Cox MM. The RecA proteins of Deinococcus radiodurans and Escherichia coli promote DNA strand exchange via inverse pathways. Proc. Natl. Acad. Sci. USA. 2002;99(12):7917–21. doi: 10.1073/pnas.122218499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Pugh BF, Cox MM. High salt activation of recA protein ATPase in the absence of DNA. J. Biol. Chem. 1988;263(1):76–83. [PubMed] [Google Scholar]
- 45.Tombline G, Fishel R. Biochemical characterization of the human RAD51 protein.I. ATP hydrolysis. J. Biol. Chem. 2002;277(17):14417–25. doi: 10.1074/jbc.M109915200. [DOI] [PubMed] [Google Scholar]
- 46.Qian X, Wu Y, He Y, Luo Y. Crystal Structure of Methanococcus voltae RadA in Complex with ADP: Hydrolysis-Induced Conformational Change. Biochemistry. 2005;44(42):13753–61. doi: 10.1021/bi051222i. [DOI] [PubMed] [Google Scholar]
- 47.Saraste M, Sibbald PR, Wittinghofer A. The P-loop--a common motif in ATP- and GTP-binding proteins. Trends Biochem. Sci. 1990;15(11):430–4. doi: 10.1016/0968-0004(90)90281-f. [DOI] [PubMed] [Google Scholar]
- 48.Kelley JA, Knight KL. Allosteric regulation of RecA protein function is mediated by Gln194. J. Biol. Chem. 1997;272(41):25778–82. doi: 10.1074/jbc.272.41.25778. [DOI] [PubMed] [Google Scholar]
- 49.Egelman E H. Does a stretched DNA structure dictate the helical geometry of RecA-like filaments?. J. Mol. Biol. 2001;309(3):539–42. doi: 10.1006/jmbi.2001.4686. [DOI] [PubMed] [Google Scholar]
- 50.Kinebuchi T, Kagawa W, Enomoto R, Tanaka K, Miyagawa K, Shibata T, Kurumizaka H, Yokoyama S. Structural basis for octameric ring formation and DNA interaction of the human homologous-pairing protein Dmc1. Mol. Cell. 2004;14(3):363–74. doi: 10.1016/s1097-2765(04)00218-7. [DOI] [PubMed] [Google Scholar]
- 51.Bell CE. Structure and mechanism of Escherichia coli RecA ATPase. Mol. Microbiol. 2005;58(2):358–66. doi: 10.1111/j.1365-2958.2005.04876.x. [DOI] [PubMed] [Google Scholar]
- 52.Galkin VE, Wu Y, Zhang XP, Qian X, He Y, Yu X, Heyer WD, Luo Y, Egelman EH. The Rad51/RadA N-terminal domain activates nucleoprotein filament ATPase activity. Structure. 2006;14(6):983–92. doi: 10.1016/j.str.2006.04.001. [DOI] [PubMed] [Google Scholar]
- 53.Du L, Luo Y. Structure of a hexameric form of RadA recombinase from Methanococcus voltae. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2012;68(Pt 5):511–6. doi: 10.1107/S1744309112010226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Qian X, He Y, Ma X, Fodje MN, Grochulski P, Luo Y. Calcium stiffens archaeal Rad51 recombinase from Methanococcus voltae for homologous recombination. J. Biol. Chem. 2006;281(51):39380–7. doi: 10.1074/jbc.M607785200. [DOI] [PubMed] [Google Scholar]