Abstract
Activating mutations of PTPN11 (encoding the SHP2 phosphatase) are associated with Noonan syndrome, childhood leukemias, and sporadic solid tumors. Virtual screening combined with experimental assays was performed to identify inhibitors of SHP2 from a database of natural products. This effort led to the identification of Cryptotanshinone as an inhibitor of SHP2. Cryptotanshinone inhibited SHP2 with an IC50 of 22.50 μM. Fluorescence titration experiments confirmed that it directly bound to SHP2. Enzymatic kinetic analyses showed that Cryptotanshinone was a mixed-type and irreversible inhibitor. This drug was further verified for its ability to block SHP2-mediated cell signaling and cellular functions. Furthermore, mouse myeloid progenitors and patient leukemic cells with the activating mutation E76K in PTPN11 were found to be sensitive to this inhibitor. Since Cryptotanshinone is used to treat cardiovascular diseases in Asian countries, this drug has a potential to be used directly or to be further developed to treat PTPN11-associated malignancies.
Introduction
Germline or somatic mutations in protein tyrosine phosphatase (PTP) PTPN11 (SHP2) that cause hyperactivation of SHP2 catalytic activity have been identified in patients with developmental disorder Noonan syndrome (∼50%) and various childhood leukemias, including juvenile myelomonocytic leukemia (JMML) (∼35%), myelodysplastic syndrome (∼10%), B cell acute lymphoblastic leukemia/lymphoma (∼7%), and acute myeloid leukemia (AML) (∼4%) 1-5. In addition, activating mutations of PTPN11 have been found in adult AML as well as sporadic solid tumors, such as lung cancer, colon cancer, melanoma, neuroblastoma, and hepatocellular carcinoma 6, 7. In JMML, PTPN11 mutations and other JMML-associated Ras, c-CBL, or NF1 mutations are mutually exclusive 1, 8, 9. Remarkably, recent studies have demonstrated that single PTPN11 gain-of-function mutations are sufficient to induce Noonan syndrome, JMML-like myeloproliferative disease, and acute leukemias in mice 10-12. The direct connection between activating mutations of PTPN11 and these diseases makes SHP2 an attractive target for mechanism-based therapeutics.
SHP2 encoded by the PTPN11 gene is a non-receptor PTP, which contains two N-terminal Src homology 2 (SH2) domains, one PTP domain, and a C-terminal tail 13, 14. In the basal state, the N-terminal SH2 (N-SH2) domain blocks the catalytic site in the PTP domain until tyrosine-phosphorylated partners bind to SHP2 15. PTPN11 mutations found in Noonan syndrome, leukemias, and tumors disrupt the auto-inhibition between N-SH2 and PTP domains, leading to gain-of-function by allowing constitutive access to the catalytic site of the enzyme 4, 16. SHP2 is widely expressed, and involved in multiple cell signaling processes, such as the Ras-Erk, PI3K-Akt, Jak-Stat, and NF-κB pathways 17-19. Although it is a positive regulator of signaling cascades initiated by growth factors/cytokines and extracellular matrix proteins 20, 21, the mechanisms of SHP2 function are still not completely known, especially, the molecular basis for the pivotal role of its catalytic activity in cell signaling pathways remain to be further understood. Thus, specific and potent SHP2 inhibitors with pharmacological properties are greatly needed for the research on SHP2 functions and the development of new drugs that ultimately serve as treatments for PTPN11-associated malignancies.
Identification of a structurally diverse set of lead compounds could lead to the development of novel SHP2 inhibitors. The availability of three dimensional (3D) crystal structural information on SHP2 13-15 makes it possible to apply target-based computer-aided drug design (CADD) methods to find SHP2 inhibitors. As natural products have certain advantages over novel compounds for drug discovery 22, in the present study, using CADD screening against the 3D structure of SHP2 followed by experimental assays, we have screened ∼10, 000 natural products, including herbal medicine currently used in the clinic. We have identified that Cryptotanshinone inhibits SHP2 catalytic activity. Moreover, biological assays verify that it is efficacious in blocking SHP2-mediated signaling and cellular functions, especially, the growth of PTPN11 mutant mouse and human leukemia cells.
Results
CADD in silico screening and compound selection
To expedite identification of SHP2 inhibitors that may be directly used for the treatment of PTPN11-associated malignancies, we chose to screen a database of natural products using CADD screening based on the 3D structure of the SHP2 protein. A structural alignment between SHP2 and the highly related SHP1 phosphatase was performed in order to search for a potential drug-docking site in the SHP2 catalytic domain that would be specific for that protein. Not surprisingly, the catalytic pockets were almost identical in SHP2 and SHP1, except four residues, i.e. Lys358, Arg362, Lys364, and Ser 365 in SHP2, which correspond to Arg352, Lys356, Arg358, and Asn359 in SHP1 (Supplementary Fig. S1). This led to the identification of a putative site in the entrance of the SHP2 catalytic cleft that appeared to be different from that in SHP1 in both structure and amino acid composition. Ligands bind to those amino acids which in the periphery of the active center through hydrophobic stabilization or hydrogen-bond (H-bond) interactions could mask the catalytic signature residues (Cys459 and Ser460), and this represents a logical place for active site-directed screening.
In silico screening was carried out against a library of ∼10,000 small molecular weight natural products targeting the peripheral site in the entrance of the SHP2 catalytic cleft, and the primary screening should meet the following criteria: (1) The nearest distance from a compound must be within 5 Å of residues Cys459 or Ser460 in SHP2 and beyond 5 Å of residues Cys453 or Ser454 in SHP1; (2) The interaction energy between a compound and one of the hydrophobic residues in SHP2 must be more than 1KJ/mol. The obtained complex structures were refined using Embrace method of MacroModel for secondary screening. After the primary screening with Glide Module and the secondary screening with MacroModel, top 200 molecules with interaction energy less than -50 kJ/mol were selected for further analyses, such as binding modes and diversity analysis. The further selection process was mainly based on the distributions of three indexes of each compound including principle moment of inertia, dipole moment, and molecular weight. The indexes were normalized and divided into ten parts, and each dot represented a compound in the grid system of three indexes. Compounds in the same lattice were clustered into a group. One or two compounds were selected from each group. As a result, a total of 43 natural products were selected and subsequently obtained from commercial sources and subjected to experimental testing.
Identification of Cryptotanshinone as a novel SHP2 inhibitor
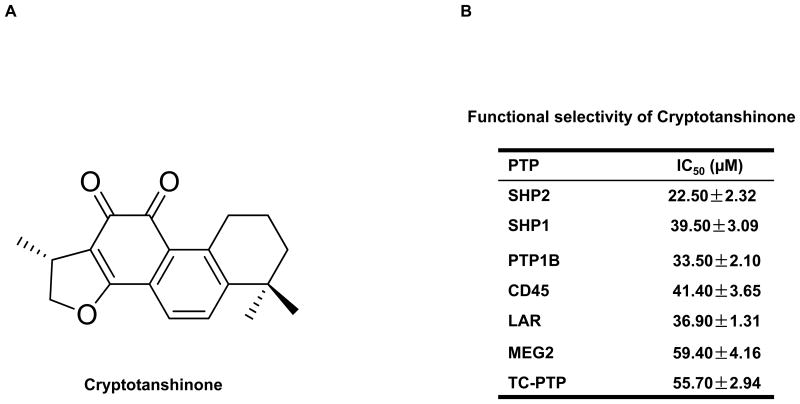
We screened the candidate compounds selected by CADD screening using SHP2 enzymatic assays. Among the 43 natural products, Cryptotanshinone (Fig. 1A and Supplementary Fig. S2), one of the major active ingredients of the traditional medicinal herbal plant Salvia miltiorrhiza Bunge (Danshen) that is used for the treatment of cardiovascular diseases in Asian countries, was able to inhibit SHP2 catalytic activity in enzymatic assays. To determine functional specificities of this compound, its inhibitory activities were assessed against a panel of mammalian PTPs, in particular those phosphatases that are highly expressed in hematopoietic cells. As shown in Fig. 1B, Cryptotanshinone inhibited the SHP2-catalyzed hydrolysis of a phospho-peptide substrate with an IC50 of 22.50 μM, which was approximately 1.76-fold selective for SHP2 over its homolog SHP1, and 1.49-, 1.84-, 1.64-, 2.64-, 2.48-fold selective for SHP2 compared to PTP1B, CD45, LAR, MEG2, and TC-PTP, respectively.
Figure 1.
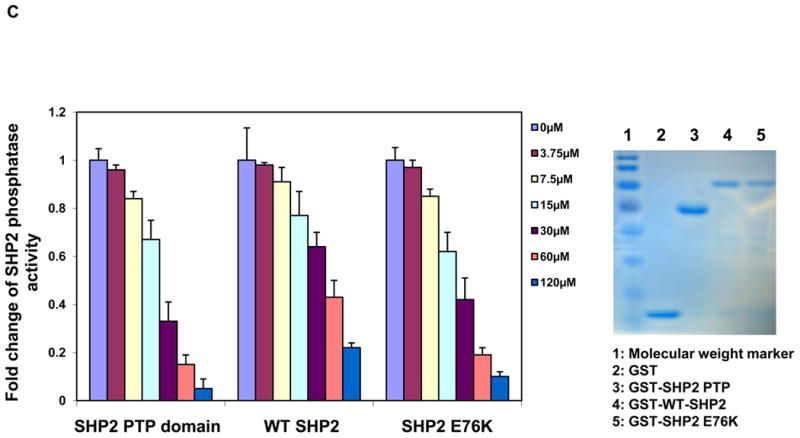
Identification of Cryptotanshinone as an inhibitor of SHP2. (A) Chemical structure of Cryptotanshinone. (B) Phosphatase assays were carried out using the indicated phosphatases as enzymes and a phospho-EGFR peptide as the substrate in the presence of various concentrations of Cryptotanshinone, as described in Experimental Section. For IC50 determinations, 6 concentrations of Cryptotanshinone were tested. Results shown are mean±S.E.M. of three independent experiments. (C) PTP activities of the SHP2 PTP domain, WT full length SHP2, and full length SHP2 E76K were determined in the presence of Cryptotanshinone at the indicated concentrations using a phospho-EGFR peptide as the substrate. Experiments were repeated three times. Similar results were obtained in each. Results shown are mean±S.E.M. of triplicates from one experiment. Shown on the right panel are purified recombinant proteins stained with Coomassie blue.
Cryptotanshinone preferentially inhibits the activated form of SHP2
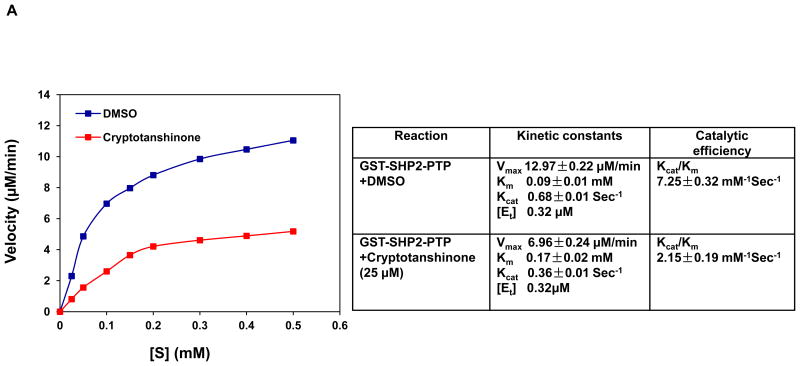
In the basal state, SHP2 is auto-inhibited by H-bonding of the N-SH2 domain loop to the deep pocket of the PTP domain. Tumor-associated mutations of PTPN11 result in amino acid changes at the interphase formed by N-SH2 and PTP domains, disrupting the inhibitory interaction, leading to hyperactivation of SHP2 catalytic activity 4, 16. The E76K mutation, localized in the N-SH2 domain is the most common and most active mutation found in leukemias 16, 23. To further characterize the acting mechanisms of Cryptotanshinone, we purified GST fusion proteins of the SHP2 PTP domain, wild-type (WT) full-length SHP2 and the full-length SHP2 E76K mutant, and compared sensitivities of these SHP2 recombinant proteins to Cryptotanshinone. This drug inhibited the SHP2 E76K mutant in a dose-dependent manner, with an IC50 of 23.90 μM that is equal to IC50 for the SHP2 PTP domain (22.50 μM). However, the IC50 of Cryptotanshinone for WT full-length SHP2 (45.18μM) was 1.89-fold and 2.01-fold higher than those for the SHP2 E76K mutant and the SHP2 PTP domain, respectively (Fig. 1C). The observation that WT full-length SHP2 is less sensitive to Cryptotanshinone than the SHP2 PTP domain and that the SHP2 E76K mutant is more sensitive than WT full-length SHP2 to this inhibitor indicates that the closed confirmation formed between N-SH2 and PTP domains interferes with Cryptotanshinone binding to SHP2 and that this compound preferentially inhibits the activated form of SHP2 with an open confirmation. To further characterize the activity of Cryptotanshinone, we also tested its effects on SHP2 catalytic activity using para-Nitrophenylphosphate (pNPP), a general small compound substrate for all PTPs. Similar effects were observed (Supplementary Fig. S3). These results suggest that Cryptotanshinone inhibits SHP2 activity likely by disrupting constitutive access to the SHP2 catalytic cleft regardless of what kinds of substrates were used. Furthermore, we performed enzymatic kinetic studies and determined Vmax, Kcat, and Km values in the absence or presence of this inhibitor. As shown in Fig. 2A, Vmax was decreased and Km was increased by this inhibitor, suggesting that Cryptotanshinone is a mixed-type inhibitor. We also tested whether the inhibition of SHP2 by Cryptotanshinone is reversible by extensive washing of the enzyme-inhibitor complex. The results obtained indicate that this drug is an irreversible inhibitor (Supplemental Fig. S4).
Figure 2.
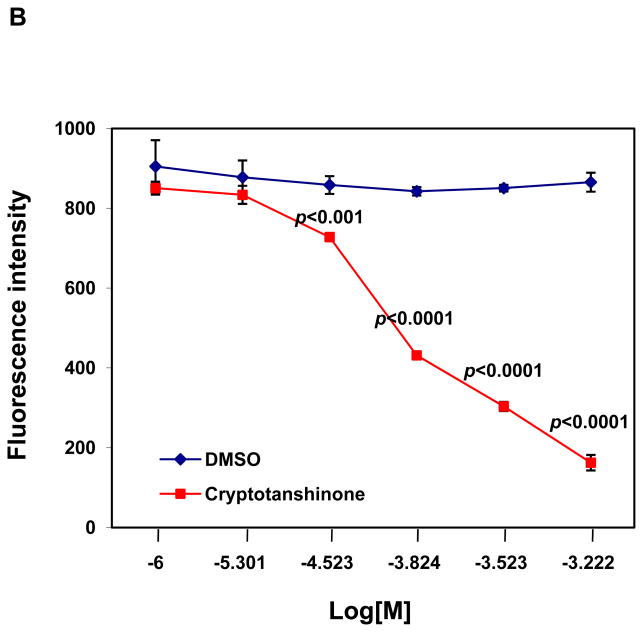
Cryptotanshinone directly binds to the SHP2 protein and functions as a mixed-type inhibitor. (A) Enzymatic kinetic analyses in the presence or absence of Cryptotanshinone (25 μM) were performed using GST-SHP2 PTP (1.0 μg) as the enzyme. Vmax, Kcat, and Km values in the absence or presence of this inhibitor were determined. Results shown in the table are mean±S.E.M. of three independent experiments. (B) Fluorescence titration of SHP2 was performed by increasing the concentrations of Cryptotanshinone while maintaining the SHP2 protein concentration (1.0 μg). The fluorescence is plotted against the log concentration in mol/L (Log [M]) for the compound. Experiments were repeated twice. Similar results were obtained in each. Data shown are mean±S.E.M. of triplicates from one representative experiment.
To validate that Cryptotanshinone was binding directly to the SHP2 protein, we examined whether binding of the inhibitor altered the fluorescence of the SHP2 PTP domain using the fluorescence quenching assay taking advantage of the four tryptophans in the PTP domain. Of these residues, Trp248 is located 15 Å from the putative pY+5 pocket and Trp423 located 8 Å from the phosphatase active site. As shown in Fig. 2B, this drug exhibited strong quenching of SHP2 fluorescence in a dose-dependent manner. These fluorescence quenching experiments imply that Cryptotanshinone binds directly to SHP2, thereby attenuating its activity toward the substrates.
Effects of Cryptotanshinone on SHP2-mediated cell signaling and cellular function
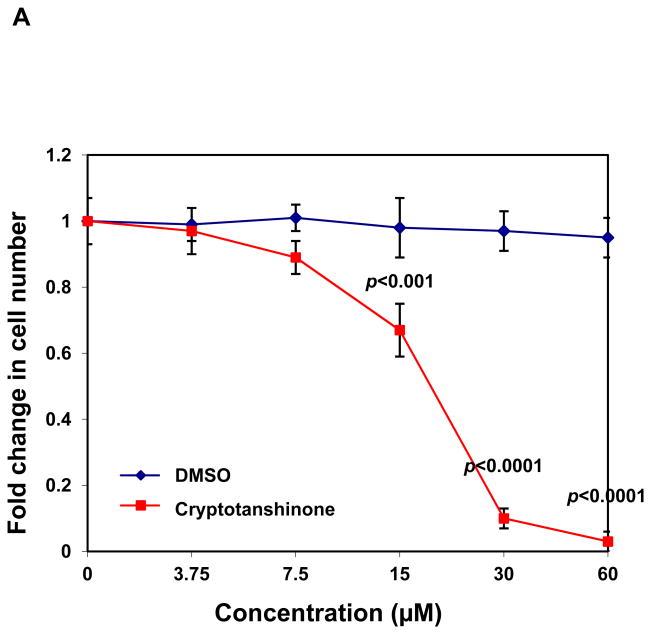
Our previous studies have demonstrated that SHP2 promotes IL-3-stimulated cell proliferation through activation of Erk, Akt, and Jak2 24, and JMML patient or mouse bone marrow cells with activating mutations of PTPN11 are hypersensitive to GM-CSF and IL-3 25, 26,10, 12, 27. To characterize the effects of Cryptotanshinone on SHP2-mediated cellular function, IL-3-dependent Ba/F3 cells were therefore used. As shown in Fig.3A, this compound displayed significant growth inhibitory effect in Ba/F3 cells with an IC50 of 17.22 μM, consistent with the overall positive role of SHP2 catalytic activity in cellular response to IL-3. To evaluate whether Cryptotanshinone functions through interfering with SHP2-mediated cell signaling, we examined the effects of this inhibitor on IL-3-induced activation of Erk, Akt, and Jak2. As shown in Fig. 3B and 3C, IL-3-induced activation of Erk, Akt, Jak2, and Stat5 determined by their phosphorylation levels was greatly suppressed by Cryptotanshinone, and the drug functioned in a dose-dependent manner, in line with the notion that SHP2 catalytic activity is required for optimal activation of these signaling pathways triggered by IL-3 24. Additionally, we found that tyrosine-phosphorylation of Gab2 (carried out by Jak2 kinase), an important adaptor protein for the PI3K-Akt pathway 28, 29, and Gab2-SHP2 interaction in Ba/F3 cells were markedly decreased by Cryptotanshinone treatment (Supplementary Fig. S5). Taken together, these results imply that IL-3 signaling processes including Ras-Erk, PI3K-Akt, and Jak2-Stat5 pathways were down-regulated following the treatment with Cryptotanshinone through a mechanism involving the inhibition of SHP2 catalytic activity.
Figure 3.
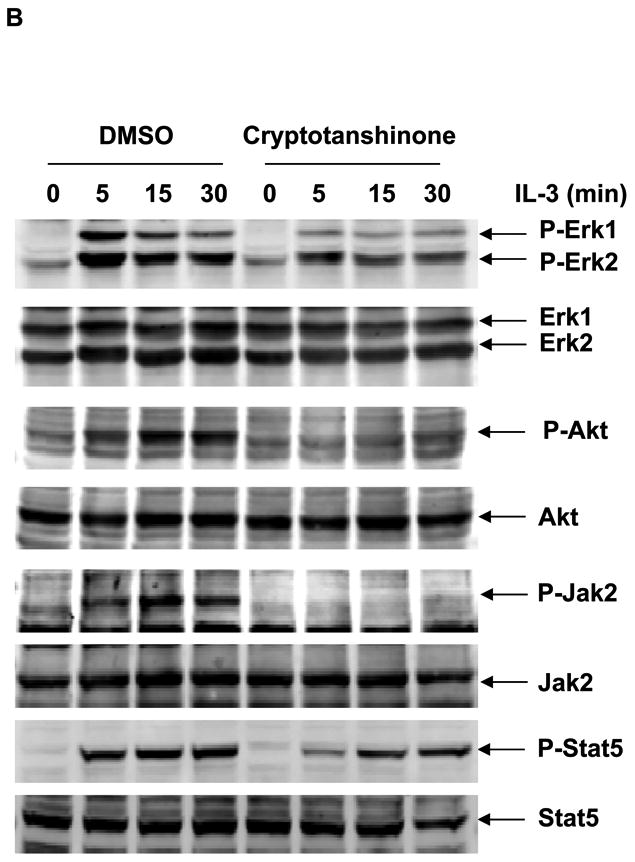
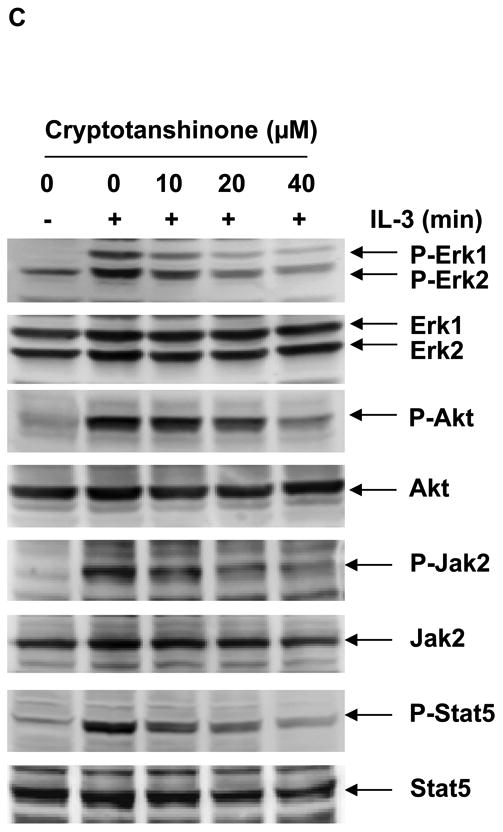
Cryptotanshinone inhibits SHP2-mediated IL-3 signaling and IL-3-induced cellular responses. (A) Ba/F3 cells were cultured in IL-3 (1.0 ng/mL) containing medium supplemented with Cryptotanshinone at the indicated concentrations or control DMSO. Cell numbers were determined 48 hours later using the One Solution Cell Proliferation Assay kit. Experiments were performed three times with similar results obtained in each. Data shown are mean±S.E.M. of triplicates from one representative experiment. (B) Ba/F3 cells were deprived of IL-3 overnight. Cells were treated with Cryptotanshinone (20 μM) for 3 hours and then stimulated with IL-3 (1.0 ng/mL) for the indicated times. (C) Ba/F3 cells were deprived of IL-3 overnight. Cells were treated with Cryptotanshinone at the indicated concentrations for 3 hours and then stimulated with IL-3 (1.0 ng/mL) for 10 min. Whole cell lysates were prepared. The levels of p-Erk, p-Akt, p-Jak2, and p-Stat5 were determined by immunoblotting analyses. Blots were striped and reprobed with anti-Erk, anti-Akt, anti-Jak2, and anti-Stat5 antibodies to check for protein loading. Experiments were repeated three times. Representative results from one experiment are shown.
Effects of Cryptotanshinone are decreased in SHP2 knockdown cells
To validate that the effects of Cryptotanshinone on cell growth were mediated through inhibiting SHP2, we tested Cryptotanshinone in SHP2 knockdown cells. Hela cells were transfected with SHP2-specific siRNA and then treated with Cryptotanshinone or DMSO for 24 hours. As shown in Fig. 4, proliferation of control cells was decreased by the SHP2 inhibitor, consistent with the positive role of SHP2 in growth factor signaling. SHP2-depleted cells were much less sensitive to this drug, suggesting that SHP2 is a main target of Cryptotanshinone. However, since the growth of SHP2-depleted cells was still decreased by Cryptotanshinone, these experiments additionally indicate that Cryptotanshinone may have additional targets.
Figure 4.
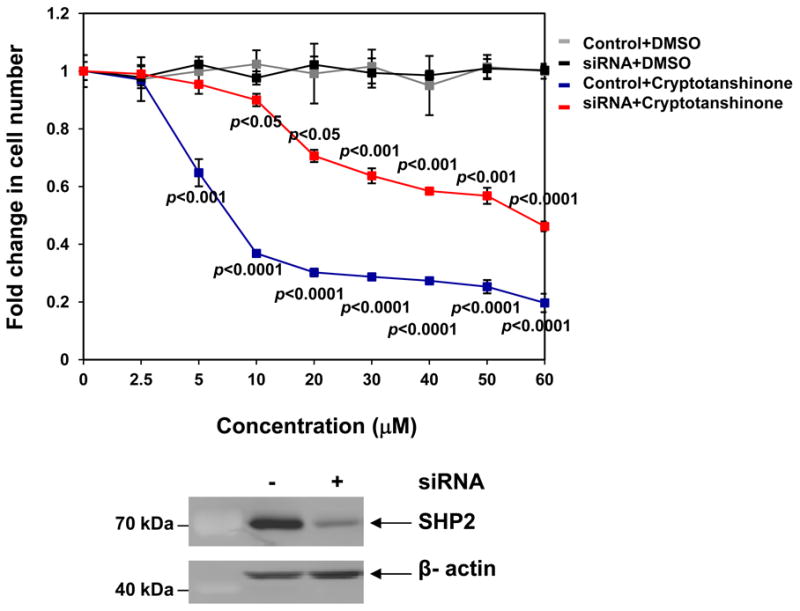
Effects of Cryptotanshinone are decreased in SHP2 knockdown cells. HeLa cells were transfected with control or SHP2 siRNA (400 pmoles/8.0×105 cells). Forty-eight hours later, transfected cells were treated with Cryptotanshinone at the indicated concentrations. Cell numbers were determined 24 hours after the treatment. Cryptotanshinone or DMSO-treated knock-down and control cells were normalized against untreated knock-down and controls cells, respectively. SHP2 levels in knockdown and control cells were analyzed by immunoblotting analyses with anti-Shp2 antibody. Blots were stripped and reprobed with anti β-actin antibody to monitor protein loading. Experiments were performed three times. Similar results were obtained in each. Data shown are mean±S.E.M. of triplicates from one representative experiment.
Mutant cells harboring hyperactivated SHP2 are more sensitive to Cryptotanshinone
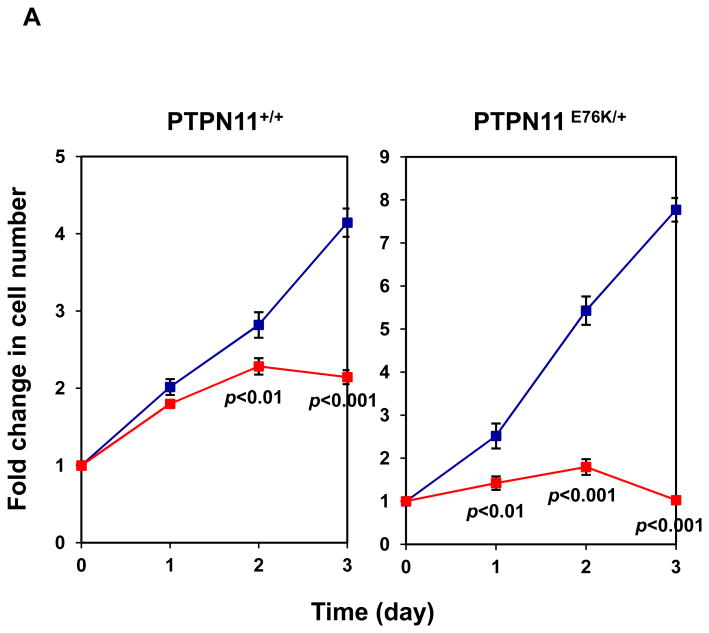
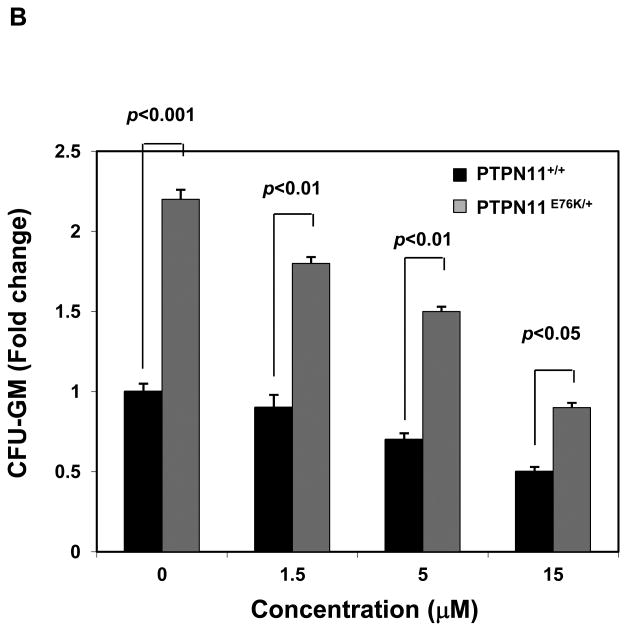
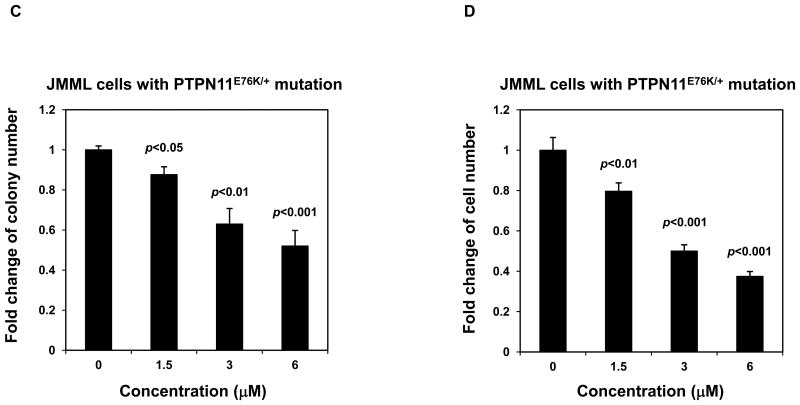
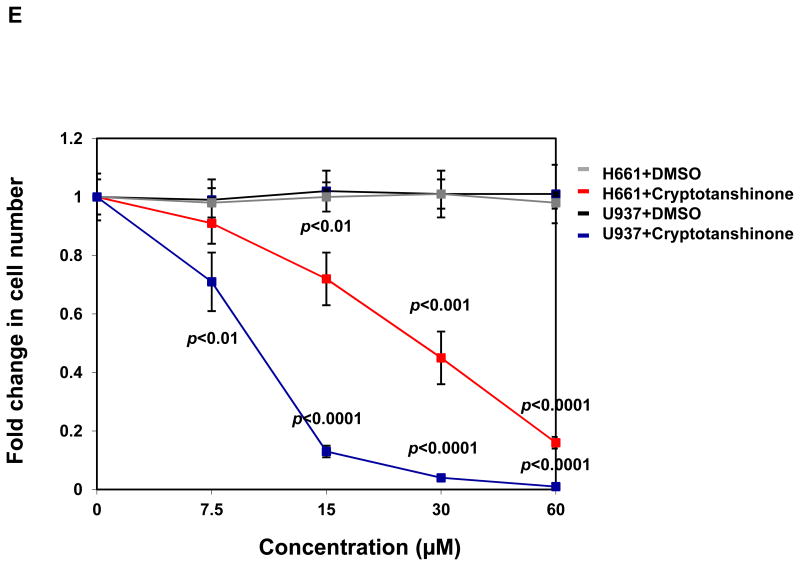
We next assessed effects of Cryptotanshinone on mutant cells with a tumor-associated PTPN11 activating mutation (PTPN11E76K/+). As illustrated in Fig. 5A, treatment of both WT and PTPN11E76K/+ mouse embryonic fibroblasts with the compound decreased cell growth. Intriguingly, PTPN11E76K/+ mutant cells were more sensitive to SHP2 inhibition. Previous studies have shown that bone marrow cells from JMML patients or mice with activating mutations of PTPN11 display a characteristic hypersensitive growth pattern, resulting in greater production of granulocyte-macrophage colonies in response to GM-CSF in CFU assays 25, 26. We next assessed whether Cryptotanshinone could inhibit colony formation and proliferation of mutant hematopoietic cells with PTPN11 activating mutations. As previously reported, compared to those of WT counterparts, myeloid progenitors from PTPN11E76K/+ mice showed greatly increased responses to GM-CSF 12. However, colony-forming capabilities of mutant SHP2-expressing progenitors were significantly reduced over a range of Cryptotanshinone concentrations (Fig. 5B). Furthermore, colony formation of myeloid blasts from JMML patients with the PTPN11E76K/+ mutation was exquisitely sensitive to Cryptotanshinone (Fig. 5C). Consistent with this data, the total cell number of JMML cells cultured in GM-CSF-containing medium was greatly decreased by the treatment of this inhibitor (Fig. 5D). These findings suggest that a low concentration of Cryptotanshinone is sufficient to overcome the dominant effects of SHP2 gain-of-function mutations in primary myeloid progenitors, confirming that human JMML might be addicted to oncogenic PTPN11 mutations. Additionally, we found that treatment of U973 human lymphoma cells harboring an activating mutation (G60R) of PTPN11, and H661 human lung cancer cells carrying another activating mutation (N58S) of PTPN11 with the compound decreased cell proliferation in a dose-dependent manner (Fig. 5E). A robust induction of apoptosis was observed in Cryptotanshinone-treated U937 cells (Fig. 5F) while Cryptotanshinone-treated H661 cells showed G1-S arrest (Supplemental Fig. S6), consistent with the positive role of SHP2 in the cell cycle progression and cell survival. The difference in the responses to SHP2 inhibition in different cell lines might be due to different other genetic mutations in these cell lines. Overall, these results support that Cryptotanshinone sensitively inhibits mutant cells harboring activating mutations of PTPN11 and suggest that this drug may be useful for the treatment of PTPN11 mutation-associated malignancies.
Figure 5.
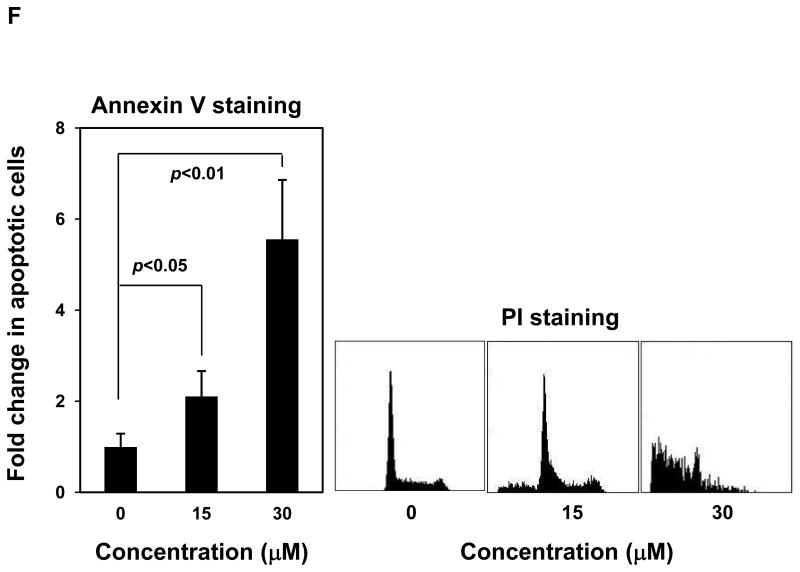
Mutant cells harboring hyperactivated SHP2 are more sensitive to Cryptotanshinone. (A) WT and PTPN11E76K/+ MEFs were cultured in 10% FBS-containing DMEM supplemented with Cryptotanshinone (20 μM). Cell numbers were determined using the One Solution Cell Proliferation Assay kit. Experiments were performed three times. Similar results were obtained in each. Data shown are mean±S.E.M. of triplicates from one representative experiment. (B) Bone marrow cells harvested from PTPN11E76K/+/Mx1-Cre+ and PTPN11+/+/Mx1-Cre+ mice were plated in methylcellulose medium containing GM-CSF (1.0 ng/mL) and Cryptotanshinone at the indicated concentrations or control DMSO. Colonies were enumerated 7 days later and normalized against the number of colonies derived from WT control cells treated with control DMSO. Representative results from two independent experiments are shown. Data are presented as mean±S.E.M. of triplicates. (C) Bone marrow cells or splenocytes from JMML patients with the PTPN11E76K/+ mutation were plated in methylcellulose medium containing GM-CSF (1.0 ng/mL) and Cryptotanshinone at the indicated concentrations or control DMSO. Colonies were enumerated 14 days later and normalized against the number of colonies derived from the cells treated with DMSO. Three patient samples were tested in three independent experiments. Similar results were obtained in each. Data are presented as mean±S.E.M. of triplicates from one patient sample. (D) Bone marrow cells or splenocytes from JMML patients with the PTPN11E76K/+ mutation were cultured in RPMI 1640 medium containing GM-CSF (1.0 ng/mL) and Cryptotanshinone at the indicated concentrations or control DMSO. Total cell numbers were determined and normalized against the number of the cells treated with DMSO. Three patient samples were tested in three independent experiments. Similar results were obtained in each. Data are presented as mean±S.E.M. of triplicates from one patient sample. (E) Human lymphoma cell line U937 with the PTPN11G60R/+ mutation and human lung cancer cell line H661 with the PTPN11N58S/+ mutation were incubated with Cryptotanshinone at the indicated concentrations for 48 hours. DMSO-treated cells were included as negative controls. Cell numbers were determined using the One Solution Cell Proliferation Assay kit. Experiments were repeated three times. Similar results were obtained in each. Data are presented as mean±S.E.M. of triplicates from one representative experiment. (F) U937 cells were incubated with Cryptotanshinone at the indicated concentrations. Apoptotic cells were quantified 24 hours later by FACS using an Annexin V staining kit following the instructions provided by the manufacturer. Cell cycle changes were determined by propidium iodide staining followed by FACS analyses 48 hours later. Experiments were performed three times. Data presented on the left panel are mean±S.E.M. of three experiments. Cell cycle profiles shown on the right panel are representatives of three independent experiments.
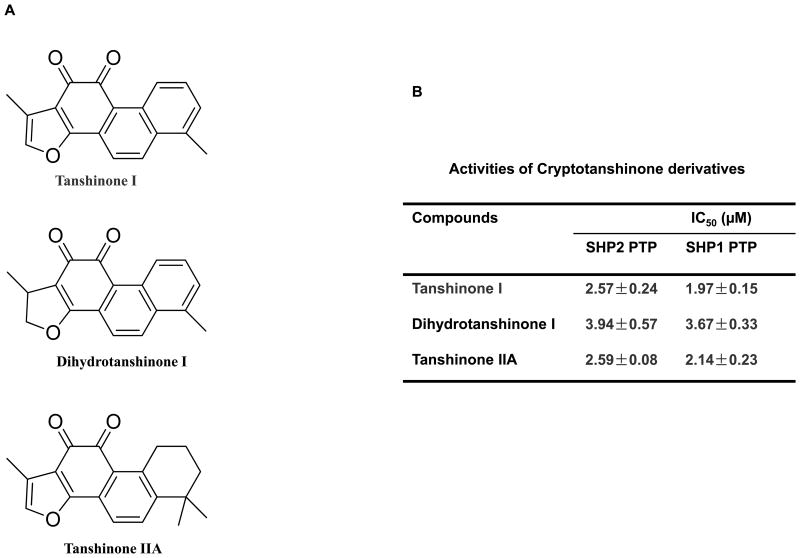
Cryptotanshinone functioned much more effectively in primary leukemic cells from human patients even at the concentrations lower than its in vitro IC50 (Fig. 5C and 5D). This unexpected observation indicates that this drug may have other targets that are also important for cell proliferation. Another possibility is that metabolites of Cryptotanshinone may have better activities. To test the second possibility, we determined potential effects of Cryptotanshinone derivatives, i.e. Tanshinone I, Dihydrotanshinone, and Tanshinone IIA (Fig. 6A) on SHP2 activity. As demonstrated in Fig. 6B, indeed these three derivatives potently inhibited SHP2. The IC50 of Tanshinone I, Dihydrotanshinone, and Tanshinone IIA was 8.75, 5.71, 8.69-fold lower than that of Cryptotanshinone, even though these compounds showed no selectivity between SHP2 and SHP1.
Figure 6.
Inhibition of SHP2 and SHP1 by Cryptotanshinone derivatives. (A) Chemical structures of Tanshinone I, Dihydrotanshinone, Tanshinone IIA. (B) Phosphatase assays were carried out using the indicated phosphatases as enzymes and a phospho-EGFR peptide as the substrate in the presence of various concentrations of the compounds, as described in Fig. 1B. For IC50 determinations, 5 concentrations of the compounds were tested. Results shown are mean±S.E.M. of three independent experiments.
Cryptotanshinone likely binds to the entrance of the active site of SHP2
To evaluate the binding mode between Cryptotanshinone and the SHP2 PTP domain, computer docking was performed. Several docked poses with similar interaction energies were inspected. The pose with the lowest interaction energy is shown in Supplemental Fig. S7A and S7B, the two carbonyl oxygen atoms of Cryptotanshinone direct to the H-bonding surface, and the cycloalkane and the two methyl groups are surrounded by the hydrophobic surface. This molecular model predicts that Cryptotanshinone binds to a specific site where the two carbonyl oxygen atoms form hydrogen bonds with Lys364 and Lys366 at the periphery of the catalytic cleft. The two H-bond lengths (O…H) are 1.850 Å and 1.738 Å, respectively. Also, the benzene ring structure of Cryptotanshinone is almost parallel to the ring structure of Tyr279 and this aromatic π-π stacking reinforces the complex structure. These findings help explain the molecular basis of SHP2 inhibition by Cryptotanshinone.
Discussion
In this study, we have identified a known natural product as a SHP2 inhibitor. SHP2 is an important signaling component downstream of growth factor/cytokine receptors and cell adhesion molecules. Germline and somatic activating mutations of PTPN11 (encoding SHP2) are associated with human diseases, such as Noonan syndrome, childhood leukemias, and solid tumors 1, 4-6. More importantly, these mutations play a causative role in the pathogenesis of Noonan syndrome and leukemias 10-12, 27, 30, 31. Thus, identification of inhibitors of the SHP2 phosphatase would greatly facilitate the development of therapeutic drugs for these diseases. Despite a great need, progress in developing selective SHP2 inhibitors is limited due to the high homology of the catalytic sites in protein tyrosine phosphatases. Using CADD database screening in combination with experimental assays, we have now identified the natural product Cryptotanshinone as a SHP2 inhibitor, although it shows moderate selectivity for SHP2 versus SHP1 and other PTPs.
Cryptotanshinone is one of the major active ingredients of the traditional medicinal herbal plant Salvia miltiorrhiza Bunge (Danshen). This drug has multiple functions and is commonly used in Asian countries to treat coronary heart disease, hyperlipidemia, acute ischemic stroke, hepatitis, chronic renal failure, and Alzheimer's disease 32-36. Recently, Cryptotanshinone has become increasingly important in cancer therapy 37. It has been found to inhibit cancer cell growth through blocking the dimerization of Stat3 and by suppressing mTOR-mediated cyclin D1 expression and Rb phosphorylation 38, 39. Besides downregulating anti-apoptotic protein expression survivin and Mcl-1, Cryptotanshinone induction of reactive oxygen species (ROS) activates p38/JNK and inhibits Erk1/2, leading to caspase-independent cell death in tumor cells 40. Our study in this report provides the first evidence that Cryptotanshinone is an inhibitor of the SHP2 phosphatase. Given the positive roles that SHP2 phosphatase plays in the signal transduction of growth factors/cytokines and cell adhesion molecules, it is quite possible that some of the functions of this drug, such as antitumor activity and anti-thrombosis activity, might be mediated through suppressing the catalytic activity of SHP2.
It is interesting that Cryptotanshinone preferentially inhibits the activated form of SHP2, which can result either from activating mutations such as E76K or from SHP2 binding to N-SH2 domain docking proteins. Since Cryptotanshinone inhibits SHP2 catalytic activity toward both the pY-peptide substrate and the small molecule substrate pNPP, it is likely that this drug functions as designed by restricting the access of these substrates to the catalytic pocket of SHP2. This notion is supported by the mode of interaction revealed by computational modeling in which Cryptotanshinone interacts with Lys364, Lys366, and Tyr279 residues at the periphery of the active site within the PTP domain. These residues most likely contribute to the binding affinity of Cryptotanshinone with SHP2. Lys364 is unique to SHP2 and this residue has been shown to be critical for the binding of another inhibitor of SHP2 41.
The discovery of SHP2-specific inhibitors is complicated by the extremely high homology of the catalytic core of SHP2 with those of SHP1 and other PTPs. Our initial structure-based approach was based on targeting a non-homologous pocket in the pY-peptide binding cleft in SHP2 adjacent to the homologous catalytic site and having unique features in both amino acid composition and 3D structure 42. A few novel compounds were identified as selective inhibitors of SHP2. Using the same strategy, we also screened a database of Food and Drug Administration -approved known drugs. Unfortunately, none of the 35 drugs identified in the CADD screening were able to inhibit SHP2 (data not shown). In the present study, we chose to target the catalytic site of SHP2 and screened a database of natural products that have advantages over novel compounds for drug development. This effort led us to the identification of Cryptotanshinone as a SHP2 inhibitor. The IC50 of Cryptotanshinone for SHP2 is only 2-fold lower than that for SHP1. This is because the active site of SHP2 is highly homologous to that of SHP1. Nevertheless, Cryptotanshinone may be still useful for future studies of SHP2 in solid tumors since SHP2 is ubiquitously expressed whereas SHP1 expression is restricted to hematopoietic cells. Further medicinal chemistry studies of this natural product is necessary to establish independent intellectual property positions that will allow pharmacological validation of its effect of SHP2 inhibition for the development of analogs for pre-clinical and clinical studies. Additionally, co-crystallization of this inhibitor and SHP2 is required to reveal the structural bases of the binding interactions of the compound with SHP2. This will be essential for developing a SHP2-specific inhibitor in our future effort.
Experimental Section
CADD in Silico Screening
Briefly, the 3D crystal structure of SHP2 was downloaded from Protein Data Bank (www.rcsb.org) with the ID code 2SHP. The receptor structure was then prepared using Protein Prepare wizard by assigning bond orders, adding hydrogen, and deleting water. The N-SH2 domain of SHP2, which blocks the catalytic site, was removed before the molecular docking. The PTP domain was minimized with Module MacroModel using default parameters. The force field used was OPLS 2005. The solvent used was water with dielectric constant 1.0. The minimization method was PRCG with 500 maximal iterations. The docking site of the catalytic domain of SHP2 was chosen by Module sitemap and then grid parameter files were produced by default method in Glide. A database of natural products (∼10,000 compounds) was used for computational calculation. The structure of compounds was sketched in Maestro and was processed using LigPrep. The chirality of each molecule was preserved. Partial charges were from OPLS 2005. Molecular docking computation was then carried out using default parameters of Glide Module. The docked pose, score, interaction energy and distance to residues of each molecule were recorded. After primary screening, the obtained complex structure was refined using Embrace method of MacroModel. The force field we used was OPLS 2005 and the convergence threshold was set to 0.05 kJ/mol. The interaction energy of optimized pose was used for further consideration. All these calculations were performed using Schrödinger 2009 software.
Reagents and Chemicals
RPMI 1640, DMEM, IMDM, and the penicillin-streptomycin stock solution were purchased from Thermo Scientific/HyClone Laboratories (Logan, UT, USA). Fetal bovine serum (FBS) was obtained from Invitrogen (Grand Island, NY, USA). The CellTiter 96® AQueous One Solution Cell Proliferation Assay kit was purchased from Promega (Madison, WI, USA). Antibodies specific for p-Erk (E-4), Erk (C-16), Jak2 (C-20), and SHP2 (C-18) were purchased from Santa Cruz Biotechnology, Inc (Santa Cruz, CA, USA). Antibodies against p-Akt (Ser473), Akt, and p-Stat5 (Y694) were obtained from Cell Signaling Technology (Beverly, MA, USA). Antibody against Stat5 was purchased from BD Biosciences (San Jose, CA, USA). Antibodies specific for p-Jak2 (Tyr1007/1008) and phospho-tyrosine (pY) were purchased from Millipore Corporation (Temecula, CA, USA). Goat-anti-rabbit and anti-mouse IgG horseradish peroxidase (HRP) conjugates were purchased from Jackson ImmunoResearch Laboratories, Inc. (West Grove, PA, USA). Dimethyl sulfoxide (DMSO) and other chemicals used in buffer solutions were provided by Fisher Scientific (Pittsburgh, PA, USA). Cryptotanshinone, Tanshinone I, Dihydrotanshinone, Tanshinone IIA were obtained from Xi'an HaoXuan Bio-Tech Co., Ltd (Xi'an, China), and dissolved in DMSO to prepare stock solutions (20 mM) for subsequent experiments. All test compounds possess a purity of ≥98% by HPLC according to the manufacturer.
In Vitro Phosphatase Activity Assay
Glutathione S-transferase (GST) fusion proteins of SHP2 purified in-house were used as the enzyme and a phospho-peptide corresponding to the surrounding sequence of pTyr10,18 in the epidermal growth factor receptor (EGFR, Asp-Ala-Asp-Glu-Tyr[PO3H2]-Leu-Ile-Pro-Gln-Gln-Gly) was used as the substrate. The assay determines free phosphate generated by dephosphorylation of the substrate using the Malachite Green reagent (Sigma, St. Louis, MO, USA). 0.5 μg GST-SHP2 PTP was used in the phosphatase assay for screening of SHP2 inhibitors. Initial rate conditions (0.2 mM substrate and 10 min incubation time) (Vmax concentration of substrate is approximately 0.3 mM) were determined following the standard enzymatic kinetic assay and used in subsequent screening. In brief, 0.5 μg of GST-SHP2 PTP was incubated in 40 μL assay buffer (25 mM Tris-HCl, pH 7.4, 50 mM NaCl, 5 mM DTT, and 2.5 mM EDTA) with test compounds at various concentrations at room temperature for 30 min. The substrate was then added to a final concentration of 0.2 mM. The reaction system was incubated at 30°C for 30 min. Finally, 50 μL of Malachite Green solution was added and OD620 was measured after 10 min. The protocols for the phosphatase assays for SHP1, PTP1B, CD45, LAR, MEG2, and TC-PTP were similar, with the exception that GST-CD45 cytoplasmic domain, GST-LAR, GST-MEG2, and GST-TC-PTP enzymes purchased from Biomol International, L.P. (Plymouth Meeting, PA, USA), were used in the respective assays.
Fluorescence Titrations
For all experiments, purified SHP2 PTP domain GST-fusion protein was diluted into 20 mM Tris-HCl, pH 7.5. Fluorescence spectra were recorded with a Luminescence Spectrometer LS50 (Perkin-Elmer, Boston, MA, USA). Titrations were performed by increasing the test compound concentrations while maintaining the SHP2 protein concentration at 3 μM. Contributions from background fluorescence of the inhibitor were accounted for by subtracting the fluorescence of the inhibitor alone from the protein-inhibitor solution. The excitation wavelength was 295 nm and fluorescence was monitored from 360 to 500 nm. All reported fluorescence intensities were relative values and were not corrected for wavelength variations in detector response.
Cell Proliferation Assay
Briefly, Ba/F3 cells were seeded at a density of 5×103 cells/well in 96-well plates in RPMI-1640 containing 10% FBS plus recombinant mouse IL-3 (1.0 ng/mL). Cells were grown overnight and then treated with either the test compound or the same concentrations of DMSO. After incubation for two days, the number of viable cells was determined using a CellTiter 96® AQueous One Solution Cell Proliferation Assay kit (Promega, Madison, WI, USA).
Western Blot Analysis
Ba/F3 cells were starved in serum and cytokine-free RPMI1640 overnight before treatment with the compound and stimulation with IL-3. Stimulated cells were harvested and lysed on ice with RIPA buffer containing 50 mM Tris-HCl, pH 7.4; 1% NP-40; 0.25% Na-deoxycholate; 150 mM NaCl; 1 mM EDTA; 1 mM NaF; 1 mM Na3VO4; 1 mM PMSF and protease inhibitor cocktail (Roche, Indianapolis, IN, USA). Equivalent amounts of protein (50 μg) were resolved on 10% SDS-PAGE and transferred to nitrocellulose membranes (Millipore, Bedford, MA, USA). Membranes were blocked with 2% BSA in TBS-T [20 mM Tris-HCl (pH 7.4), 150 mM NaCl, and 0.1% Tween 20] for 1 hour at room temperature and probed with primary antibodies overnight at 4°C. Blots were washed with TBS-T and exposed to HRP conjugated goat-anti-mouse or goat-anti-rabbit secondary antibodies for 1 hour at room temperature. Immunoreactive bands were detected by using ECL Plus Reagents (GE Healthcare, Piscataway, NJ, USA).
Colony-Forming Unit Assay
Freshly harvested mouse bone marrow cells or patient splenocytes (5×104 cells/mL) were assayed for colony forming units (CFUs) in 0.9% methylcellulose IMDM containing 30% FBS, glutamine (10-4 M), β-mercaptoethanol (3.3×10-5 M), GM-CSF (1.0 ng/mL), and varying doses of Cryptotanshinone or the same concentrations of DMSO. After 7 days (mouse bone marrow cells) or 14 days (patient cells) of culture at 37°C in a humidified 5% CO2 incubator, myeloid colonies (CFU-GM and CFU-M) were counted under an inverted microscope.
Statistical Analysis
Data are presented as mean±S.E.M. Statistical significance was determined using unpaired two-tailed Student's t test. p values <0.05 were considered to be significant.
Supplementary Material
Acknowledgments
This work was supported by the National Institutes of Health grants HD070716 and HL068212 (to C.K.Q.).
Abbreviations Used
- SHP2
Src homology 2 domain-containing tyrosine phosphatase 2
- PTP
Protein tyrosine phosphatase
- pY
Phospho-tyrosine
- JMML
Juvenile myelomonocytic leukemia
- CADD
Computer-aided drug design
- FBS
Fetal bovine serum
Footnotes
W. Liu, B. Yu, and G. Xu contributed equally to this paper.
Author Contributions: W.L. and B.Y. contributed equally to the work.
Notes: The authors declare no competing financial interest.
Supporting Information Available: Supplementary Figures S1-S7. This material is available free of charge via the Internet at http://pubs.acs.org.
References
- 1.Loh ML, Vattikuti S, Schubbert S, Reynolds MG, Carlson E, Lieuw KH, Cheng JW, Lee CM, Stokoe D, Bonifas JM, Curtiss NP, Gotlib J, Meshinchi S, Le Beau MM, Emanuel PD, Shannon KM. Mutations in PTPN11 implicate the SHP-2 phosphatase in leukemogenesis. Blood. 2004;103:2325–2331. doi: 10.1182/blood-2003-09-3287. [DOI] [PubMed] [Google Scholar]
- 2.Loh ML, Reynolds MG, Vattikuti S, Gerbing RB, Alonzo TA, Carlson E, Cheng JW, Lee CM, Lange BJ, Meshinchi S. PTPN11 mutations in pediatric patients with acute myeloid leukemia: results from the Children's Cancer Group. Leukemia. 2004;18:1831–1834. doi: 10.1038/sj.leu.2403492. [DOI] [PubMed] [Google Scholar]
- 3.Tartaglia M, Martinelli S, Cazzaniga G, Cordeddu V, Iavarone I, Spinelli M, Palmi C, Carta C, Pession A, Arico M, Masera G, Basso G, Sorcini M, Gelb BD, Biondi A. Genetic evidence for lineage-related and differentiation stage-related contribution of somatic PTPN11 mutations to leukemogenesis in childhood acute leukemia. Blood. 2004;104:307–313. doi: 10.1182/blood-2003-11-3876. [DOI] [PubMed] [Google Scholar]
- 4.Tartaglia M, Mehler EL, Goldberg R, Zampino G, Brunner HG, Kremer H, van der Burgt I, Crosby AH, Ion A, Jeffery S, Kalidas K, Patton MA, Kucherlapati RS, Gelb BD. Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat Genet. 2001;29:465–468. doi: 10.1038/ng772. [DOI] [PubMed] [Google Scholar]
- 5.Tartaglia M, Niemeyer CM, Fragale A, Song X, Buechner J, Jung A, Hahlen K, Hasle H, Licht JD, Gelb BD. Somatic mutations in PTPN11 in juvenile myelomonocytic leukemia, myelodysplastic syndromes and acute myeloid leukemia. Nat Genet. 2003;34:148–150. doi: 10.1038/ng1156. [DOI] [PubMed] [Google Scholar]
- 6.Bentires-Alj M, Paez JG, David FS, Keilhack H, Halmos B, Naoki K, Maris JM, Richardson A, Bardelli A, Sugarbaker DJ, Richards WG, Du J, Girard L, Minna JD, Loh ML, Fisher DE, Velculescu VE, Vogelstein B, Meyerson M, Sellers WR, Neel BG. Activating mutations of the noonan syndrome-associated SHP2/PTPN11 gene in human solid tumors and adult acute myelogenous leukemia. Cancer Res. 2004;64:8816–8820. doi: 10.1158/0008-5472.CAN-04-1923. [DOI] [PubMed] [Google Scholar]
- 7.Miyamoto D, Miyamoto M, Takahashi A, Yomogita Y, Higashi H, Kondo S, Hatakeyama M. Isolation of a distinct class of gain-of-function SHP-2 mutants with oncogenic RAS-like transforming activity from solid tumors. Oncogene. 2008;27:3508–3515. doi: 10.1038/sj.onc.1211019. [DOI] [PubMed] [Google Scholar]
- 8.Loh ML, Sakai DS, Flotho C, Kang M, Fliegauf M, Archambeault S, Mullighan CG, Chen L, Bergstraesser E, Bueso-Ramos CE, Emanuel PD, Hasle H, Issa JP, van den Heuvel-Eibrink MM, Locatelli F, Stary J, Trebo M, Wlodarski M, Zecca M, Shannon KM, Niemeyer CM. Mutations in CBL occur frequently in juvenile myelomonocytic leukemia. Blood. 2009;114:1859–1863. doi: 10.1182/blood-2009-01-198416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Muramatsu H, Makishima H, Jankowska AM, Cazzolli H, O'Keefe C, Yoshida N, Xu Y, Nishio N, Hama A, Yagasaki H, Takahashi Y, Kato K, Manabe A, Kojima S, Maciejewski JP. Mutations of an E3 ubiquitin ligase c-Cbl but not TET2 mutations are pathogenic in juvenile myelomonocytic leukemia. Blood. 2010;115:1969–1975. doi: 10.1182/blood-2009-06-226340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Araki T, Mohi MG, Ismat FA, Bronson RT, Williams IR, Kutok JL, Yang W, Pao LI, Gilliland DG, Epstein JA, Neel BG. Mouse model of Noonan syndrome reveals cell type- and gene dosage-dependent effects of Ptpn11 mutation. Nat Med. 2004;10:849–857. doi: 10.1038/nm1084. [DOI] [PubMed] [Google Scholar]
- 11.Chan G, Kalaitzidis D, Usenko T, Kutok JL, Yang W, Mohi MG, Neel BG. Leukemogenic Ptpn11 causes fatal myeloproliferative disorder via cell-autonomous effects on multiple stages of hematopoiesis. Blood. 2009;113:4414–4424. doi: 10.1182/blood-2008-10-182626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Xu D, Liu X, Yu WM, Meyerson HJ, Guo C, Gerson SL, Qu CK. Non-lineage/stage-restricted effects of a gain-of-function mutation in tyrosine phosphatase Ptpn11 (Shp2) on malignant transformation of hematopoietic cells. J Exp Med. 2011;208:1977–1988. doi: 10.1084/jem.20110450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Barford D, Neel BG. Revealing mechanisms for SH2 domain mediated regulation of the protein tyrosine phosphatase SHP-2. Structure. 1998;6:249–254. doi: 10.1016/s0969-2126(98)00027-6. [DOI] [PubMed] [Google Scholar]
- 14.Eck MJ, Pluskey S, Trub T, Harrison SC, Shoelson SE. Spatial constraints on the recognition of phosphoproteins by the tandem SH2 domains of the phosphatase SH-PTP2. Nature. 1996;379:277–280. doi: 10.1038/379277a0. [DOI] [PubMed] [Google Scholar]
- 15.Hof P, Pluskey S, Dhe-Paganon S, Eck MJ, Shoelson SE. Crystal structure of the tyrosine phosphatase SHP-2. Cell. 1998;92:441–450. doi: 10.1016/s0092-8674(00)80938-1. [DOI] [PubMed] [Google Scholar]
- 16.Keilhack H, David FS, McGregor M, Cantley LC, Neel BG. Diverse biochemical properties of Shp2 mutants. Implications for disease phenotypes. J Biol Chem. 2005;280:30984–30993. doi: 10.1074/jbc.M504699200. [DOI] [PubMed] [Google Scholar]
- 17.Xu D, Qu CK. Protein tyrosine phosphatases in the JAK/STAT pathway. Front Biosci. 2008;13:4925–4932. doi: 10.2741/3051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chan G, Kalaitzidis D, Neel BG. The tyrosine phosphatase Shp2 (PTPN11) in cancer. Cancer Metastasis Rev. 2008;27:179–192. doi: 10.1007/s10555-008-9126-y. [DOI] [PubMed] [Google Scholar]
- 19.Tonks NK. Protein tyrosine phosphatases: from genes, to function, to disease. Nat Rev Mol Cell Biol. 2006;7:833–846. doi: 10.1038/nrm2039. [DOI] [PubMed] [Google Scholar]
- 20.Neel BG, Gu H, Pao L. The ‘Shp’ing news: SH2 domain-containing tyrosine phosphatases in cell signaling. Trends Biochem Sci. 2003;28:284–293. doi: 10.1016/S0968-0004(03)00091-4. [DOI] [PubMed] [Google Scholar]
- 21.Dance M, Montagner A, Salles JP, Yart A, Raynal P. The molecular functions of Shp2 in the Ras/Mitogen-activated protein kinase (ERK1/2) pathway. Cell Signal. 2008;20:453–459. doi: 10.1016/j.cellsig.2007.10.002. [DOI] [PubMed] [Google Scholar]
- 22.Newman DJ. Natural products as leads to potential drugs: an old process or the new hope for drug discovery? J Med Chem. 2008;51:2589–2599. doi: 10.1021/jm0704090. [DOI] [PubMed] [Google Scholar]
- 23.Tartaglia M, Martinelli S, Stella L, Bocchinfuso G, Flex E, Cordeddu V, Zampino G, Burgt I, Palleschi A, Petrucci TC, Sorcini M, Schoch C, Foa R, Emanuel PD, Gelb BD. Diversity and functional consequences of germline and somatic PTPN11 mutations in human disease. Am J Hum Genet. 2006;78:279–290. doi: 10.1086/499925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yu WM, Hawley TS, Hawley RG, Qu CK. Catalytic-dependent and -independent roles of SHP-2 tyrosine phosphatase in interleukin-3 signaling. Oncogene. 2003;22:5995–6004. doi: 10.1038/sj.onc.1206846. [DOI] [PubMed] [Google Scholar]
- 25.Emanuel PD, Bates LJ, Castleberry RP, Gualtieri RJ, Zuckerman KS. Selective hypersensitivity to granulocyte-macrophage colony-stimulating factor by juvenile chronic myeloid leukemia hematopoietic progenitors. Blood. 1991;77:925–929. [PubMed] [Google Scholar]
- 26.Lyubynska N, Gorman MF, Lauchle JO, Hong WX, Akutagawa JK, Shannon K, Braun BS. A MEK inhibitor abrogates myeloproliferative disease in Kras mutant mice. Sci Transl Med. 2011;3:76ra27. doi: 10.1126/scitranslmed.3001069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chan RJ, Leedy MB, Munugalavadla V, Voorhorst CS, Li Y, Yu M, Kapur R. Human somatic PTPN11 mutations induce hematopoietic-cell hypersensitivity to granulocyte-macrophage colony-stimulating factor. Blood. 2005;105:3737–3742. doi: 10.1182/blood-2004-10-4002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nishida K, Hirano T. The role of Gab family scaffolding adapter proteins in the signal transduction of cytokine and growth factor receptors. Cancer Sci. 2003;94:1029–1033. doi: 10.1111/j.1349-7006.2003.tb01396.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gu H, Neel BG. The “Gab” in signal transduction. Trends Cell Biol. 2003;13:122–130. doi: 10.1016/s0962-8924(03)00002-3. [DOI] [PubMed] [Google Scholar]
- 30.Schubbert S, Lieuw K, Rowe SL, Lee CM, Li X, Loh ML, Clapp DW, Shannon KM. Functional analysis of leukemia-associated PTPN11 mutations in primary hematopoietic cells. Blood. 2005;106:311–317. doi: 10.1182/blood-2004-11-4207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yu WM, Daino H, Chen J, Bunting KD, Qu CK. Effects of a leukemia-associated gain-of-function mutation of SHP-2 phosphatase on interleukin-3 signaling. J Biol Chem. 2006;281:5426–5434. doi: 10.1074/jbc.M507622200. [DOI] [PubMed] [Google Scholar]
- 32.Zhou LM, Zuo Z, Chow MSS. Danshen: An overview of its chemistry, pharmacology, pharmacokinetics, and clinical use. Journal of Clinical Pharmacology. 2005;45:1345–1359. doi: 10.1177/0091270005282630. [DOI] [PubMed] [Google Scholar]
- 33.Lu Y, Foo LY. Polyphenolics of Salvia--a review. Phytochemistry. 2002;59:117–140. doi: 10.1016/s0031-9422(01)00415-0. [DOI] [PubMed] [Google Scholar]
- 34.Stickel F, Brinkhaus B, Krahmer N, Seitz HK, Hahn EG, Schuppan D. Antifibrotic properties of botanicals in chronic liver disease. Hepatogastroenterology. 2002;49:1102–1108. [PubMed] [Google Scholar]
- 35.Wojcikowski K, Johnson DW, Gobe G. Herbs or natural substances as complementary therapies for chronic kidney disease: ideas for future studies. Journal of Laboratory and Clinical Medicine. 2006;147:160–166. doi: 10.1016/j.lab.2005.11.011. [DOI] [PubMed] [Google Scholar]
- 36.Yu XY, Lin SG, Chen X, Zhou ZW, Liang J, Duan W, Chowbay B, Wen JY, Chan E, Cao J, Li CG, Zhou SF. Transport of cryptotanshinone, a major active triterpenoid in Salvia miltiorrhiza Bunge widely used in the treatment of stroke and Alzheimer's disease, across the blood-brain barrier. Current Drug Metabolism. 2007;8:365–377. doi: 10.2174/138920007780655441. [DOI] [PubMed] [Google Scholar]
- 37.Chen W, Lu Y, Chen G, Huang S. Molecular Evidence of Cryptotanshinone for Treatment and Prevention of Human Cancer. Anticancer Agents Med Chem. 2013;13:979–987. doi: 10.2174/18715206113139990115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Shin DS, Kim HN, Shin KD, Yoon YJ, Kim SJ, Han DC, Kwon BM. Cryptotanshinone Inhibits Constitutive Signal Transducer and Activator of Transcription 3 Function through Blocking the Dimerization in DU145 Prostate Cancer Cells. Cancer Research. 2009;69:193–202. doi: 10.1158/0008-5472.CAN-08-2575. [DOI] [PubMed] [Google Scholar]
- 39.Chen W, Luo Y, Liu L, Zhou H, Xu B, Han X, Shen T, Liu Z, Lu Y, Huang S. Cryptotanshinone inhibits cancer cell proliferation by suppressing Mammalian target of rapamycin-mediated cyclin D1 expression and Rb phosphorylation. Cancer Prev Res (Phila) 2010;3:1015–1025. doi: 10.1158/1940-6207.CAPR-10-0020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Chen W, Liu L, Luo Y, Odaka Y, Awate S, Zhou H, Shen T, Zheng S, Lu Y, Huang S. Cryptotanshinone activates p38/JNK and inhibits Erk1/2 leading to caspase-independent cell death in tumor cells. Cancer Prev Res (Phila) 2012;5:778–787. doi: 10.1158/1940-6207.CAPR-11-0551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhang X, He Y, Liu S, Yu Z, Jiang ZX, Yang Z, Dong Y, Nabinger SC, Wu L, Gunawan AM, Wang L, Chan RJ, Zhang ZY. Salicylic acid based small molecule inhibitor for the oncogenic Src homology-2 domain containing protein tyrosine phosphatase-2 (SHP2) J Med Chem. 2010;53:2482–2493. doi: 10.1021/jm901645u. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Yu WM, Guvench O, Mackerell AD, Qu CK. Identification of small molecular weight inhibitors of Src homology 2 domain-containing tyrosine phosphatase 2 (SHP-2) via in silico database screening combined with experimental assay. J Med Chem. 2008;51:7396–7404. doi: 10.1021/jm800229d. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.