Summary
Osteogenic sarcoma (OS) is a deadly skeletal malignancy whose cause is unknown. We report here a mouse model of OS based on conditional expression of the intracellular domain of Notch1 (NICD). Expression of the NICD in immature osteoblasts was sufficient to drive the formation of bone tumors, including OS, with complete penetrance. These tumors display features of human OS, namely histopathology, cytogenetic complexity, and metastatic potential. We show that Notch activation combined with loss of p53 synergistically accelerates OS development in mice although p53-driven OS is not Rbpj-dependent, which demonstrates a dual dominance of the Notch oncogene and p53 mutation in the development of OS. Using this model, we also reveal the osteoblasts as the potential sources of OS.
Introduction
Osteogenic sarcoma, also referred to osteosarcoma (OS), is the most common primary bone cancer, comprising approximately 20% of all bone tumors and 5% of pediatric tumors overall (Gorlick and Khanna, 2010). OS affects patients of all ages but shows a substantially higher incidence in adolescents and young adults. OS patients with metastases in lungs have poor five-year survival rates, on the order of 30% or less (Jawad et al., 2011). Most cases of OS occur sporadically and our understanding of the molecular basis of the disease is still limited. Correspondingly, there have been no substantial improvements in survival rates over the past three decades, mostly owing to a lack of knowledge of the driver gene mutations and cells of origin. In addition, very few mouse models possessing a high frequency of OS are currently available (Grigoriadis et al., 1993; Harvey et al., 1993; Molyneux et al., 2010). Recently, Orkin and colleagues showed that bone-specific disruption of Trp53 and Rb leads to the formation of OS and mimics the human form of the disease (Walkley et al., 2008), supporting the hypothesis that OS may arise from mesenchymal-stem-cell-derived osteoblasts, the predominant bone-forming cells.
Evolutionarily conserved signaling pathways such as the Notch pathway are central mechanisms in embryogenesis, postnatal homeostasis, and pathological conditions including tumorigenesis (Kopan and Ilagan, 2009). Notch proteins are transmembrane receptors that are activated by physical interaction with a transmembrane ligand on adjacent cells. Once a Notch receptor is activated, it undergoes a series of intramembranous cleavages by proteolytic enzymes, including γ-secretase, which release the Notch Intracellular domain (NICD). NICD enters the nucleus and interacts with a transcriptional complex comprising Rbpj and Mastermind-like proteins to regulate expression of canonical targets such as the Hey and Hes family of transcription factors. Notch receptor mutations have been associated with several types of cancer, and current data suggest that Notch can serve as either a tumor promoter or a tumor suppressor in a context-dependent manner (Ranganathan et al., 2011). In the skeletal system, we and others have shown that Notch activation can stimulate proliferation of immature osteoblasts while inhibiting their differentiation into mature osteoblasts (Engin et al., 2008; Hilton et al., 2008; Zamurovic et al., 2004; Zanotti et al., 2008). This gain-of-function phenotype is reminiscent of osteoblastic tumors and is consistent with recent findings showing that Notch signaling is up-regulated in human OS samples and that its inhibition in vitro and in immunodeficient mice suppresses OS cell proliferation and migration (Engin et al., 2009; Tanaka et al., 2009). However, it is not known whether Notch dysregulation is involved in the initiation and/or progression of OS.
A human Notch gain-of-function mutation was discovered in cells derived from a patient with T-cell acute lymphoblastic leukemia (T-ALL) (Ellisen et al., 1991). The mutation, a chromosomal 7-chromosome 9 translocation, results in the expression of a truncated NICD-like protein from NOTCH1, which encodes one of four closely related members (Notch1–4) in mammals. Expression of this truncated protein predisposes mice to T-cell leukemia and recapitulates some aspects of the human disease (Robey et al., 1996). A subsequent study found that more than 50% of human T-ALLs have NOTCH1 gain-of-function mutations involving the extracellular hetero-dimerization domain and/or the C-terminal PEST domain, which regulates NICD degradation (Weng et al., 2004). These missense mutations and/or frame-shifting insertions or deletions result in ligand-independent cleavage and accumulation of NICD in the nucleus and/or NICD stabilization, inducing constitutively active Notch signaling. Notch gain-of-function mutations have also been found in several types of solid tumors, including lung cancer (Lawrence et al., 2014; Westhoff et al., 2009). However, somatic Notch mutations have yet to be identified in any types of mesenchyme-derived rare cancers such as OS. Based on our and other recent studies showing that activation of Notch signaling contributes to the pathogenesis of human OS, we hypothesized that osteoblast-specific activation of the Notch signaling pathway may be sufficient to initiate OS development.
Results
NICD Expression in committed osteoblasts induces malignant bone tumors that mimic human OS
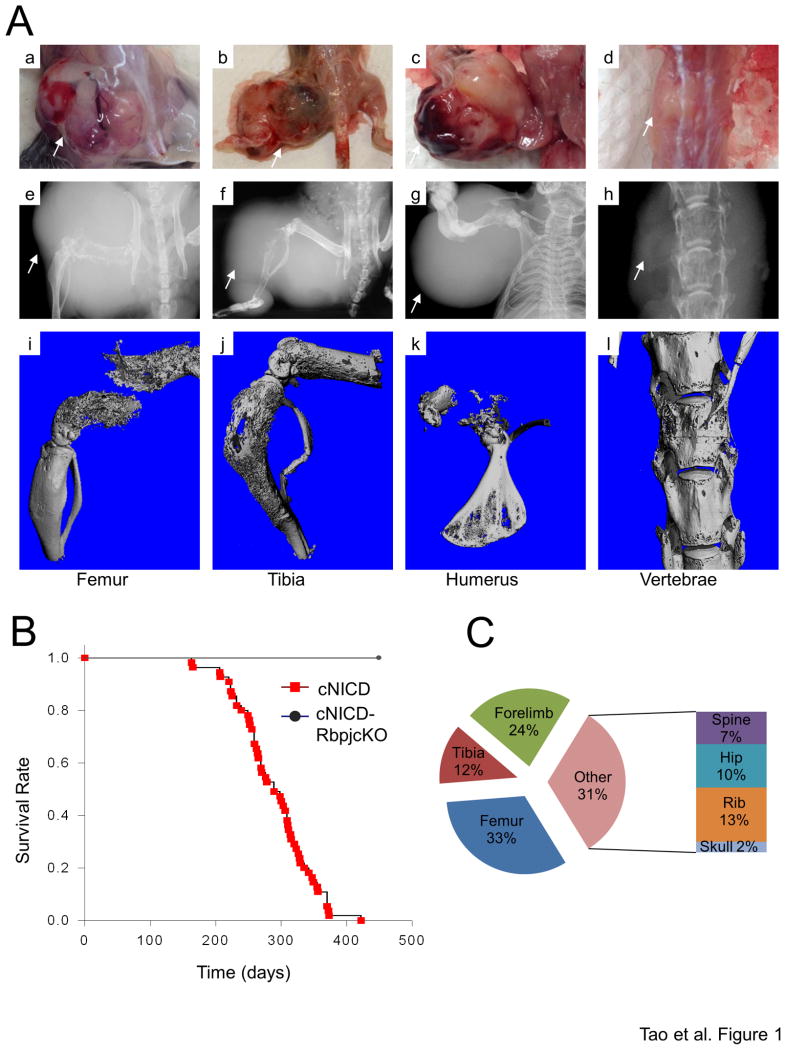
We previously applied the conditional NICD allele to allow for the tissue-restricted activation of this signaling pathway following Cre expression in bone (Tao et al., 2010b). Rosa26NICD mice express NICD-IRES-GFP bicistronic mRNA from the endogenous Rosa26 promoter, which enables monitoring of NICD expression by detecting nuclear GFP expression (Figure S1A). To achieve osteoblast-restricted expression, we utilized the Col1a1 2.3kb-Cre transgenic mouse to direct Cre expression to committed osteoblasts. The conditional Rosa26NICD mice were bred to Col1a1 2.3kb-Cre mice and the resulting Col1a1 2.3kb-Cre; Rosa26NICD (hereafter, cNICD) progenies were followed longitudinally (Figure S1B). These mice were smaller than their siblings at a young age and usually died of tumor-unrelated causes by 3 months of age. However, 100% of the surviving cNICD mice eventually caught up in growth to their littermates (Figure S1C) and spontaneously developed clinically manifested tumors between the ages of 5–14 months (Figure 1A-B), demonstrating complete penetrance in terms of tumor induction. Examination of the mice by autopsy (Figure 1Aa-d), X-ray imaging (Figure 1Ae-h), and microscopic computed tomography (micro-CT) (Figure 1Ai-l) confirmed that only the bones were affected, with no macroscopic abnormalities observed in any other tissues. The cNICD mice had monostotic or polyostotic bone tumors at all sites known to be affected in the human disease. Histological examination revealed these to be osteosarcoma or osteoma, a benign tumor that typically grows in the human skull. The most frequent sites of osteoma in cNICD mice were the maxilla and the zygomatic bone in the skull (Figure S1D-E), followed by the radius and ulna in the forelimb (Figure S1F). Since osteosarcoma was responsible for the death of most cNICD mice, all subsequent studies were performed on Notch-induced osteosarcoma (hereafter Notch OS, or NOS) (Figure S1G).
Figure 1. Osteosarcoma in the cNICD mice.
(A) Necropsy photos (a-d), X-ray images (e-h), and micro-CT scans (i-l) of representative NOS in the femur (a, e, i), tibia (b, f, j), humerus (c, g, k), and vertebrae (d, h, l) with white arrows pointing to the tumor.
(B) Kaplan-Meier tumor-free survival plots in cNICD (n=50) and cNICD-RbpjcKO (n=16) mice.
(C) Location and frequency of NOS in cNICD mice.
See also Figure S1.
The most frequent site of NOS was the femur, a major site for human OS, followed by the forelimb/tibia/ribs and hip and spine (Figure 1C). A peak incidence in the long bones closely approximates that seen in human OS (Unni and Inwards, 2009). All OS tumors in the 41 cNICD mice appeared remarkably similar in morphology. Macroscopically, these tumors were firm, rounded, displayed a white-to-grayish cut surface, and often contained areas of hemorrhage and cystic degeneration (Figures 1Aa-d and S1H). In most cases, the underlying cortical bone was disintegrated and the medullary cavity was filled with tumor cells (Figures 1Ae-l, S1H and S1I).
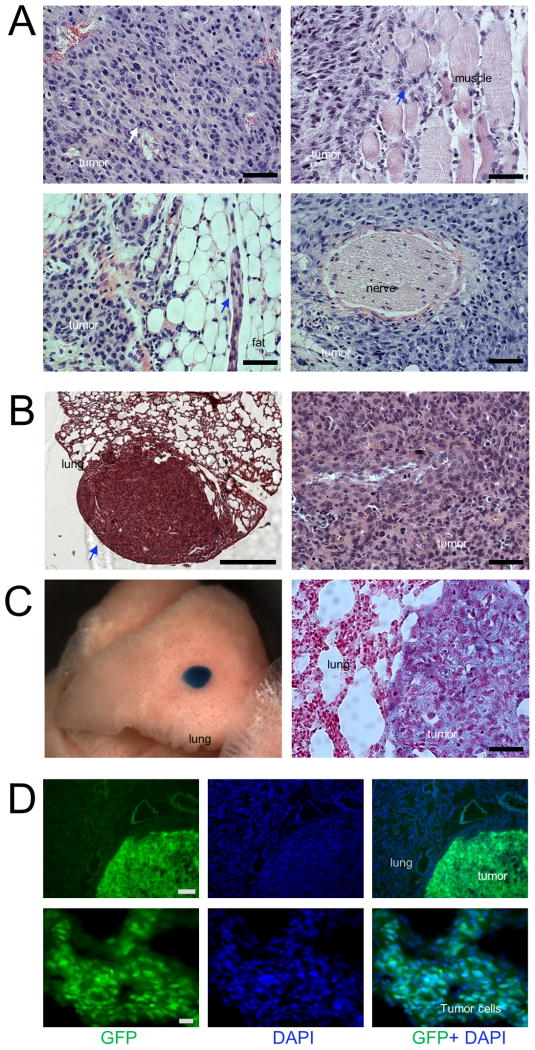
Examination of individual NOS tumors from different animals revealed uniform histopathological features: poorly differentiated or undifferentiated hypercellular lesions with small deposits of osteoid (bone matrix prior to calcification), scattered mitoses and clear invasion of the adjacent muscle, fat tissues but not nerves (Figures 2A and S2A). Together, these tumors displayed histological characteristics typical of the most aggressive type of human OS. Metastases are a major cause of death in patients with OS, with the lung being the most common site. We observed that a significant fraction of the OS lesions metastasized to lung in approximately 40% of the examined NOS-bearing mice. Some metastatic lesions were detected macroscopically (Figures 2B and S2B) whereas others could be only observed by microscopy (Figures 2C and S2C). We did not detect metastases in other organs. Expression of LacZ (Figures 2C and S2D) or nuclear GFP (Figures 2D and S2E) in these metastatic lung tumors and primary tumors, but not in adjacent tissues, demonstrates their osteoblastic origin. Thus, NOS closely resembles the human disease in terms of skeletal distribution, radiographic and histologic features, and in the development of spontaneous lung metastases. Our NOS model therefore constitutes a valuable resource for the study of this cancer.
Figure 2. Histopathological features of primary and metastatic NOS.
(A) Histology of representative NOS tumors stained with hematoxylin and eosin (H&E) showing poorly differentiated OS with high cellularity and scattered mitoses (white arrow, top left), and invasive infiltration of tumor cells into surrounding muscle tissue (blue arrow, top right) and fat (blue arrow, bottom left), but not nerves (bottom right). Scale bars, 50 μm.
(B) H&E-stained sections of representative lung metastatic OS mass (blue arrow) showing high cellularity at high magnification (right). Scale bars, 500 μm (left) and 50 μm (right).
(C) Photo of a mouse lung lobe (left), which was harvested from a tri-transgenic mouse (Rosa26NICD; Col1a1 2.3kb-Cre; Rosa26LacZ) with whole mount X-gal staining shows a metastatic lesion (blue) that could not be detected macroscopically. Lung section counter-stained with eosin (right) showed that LacZ was only expressed in tumor cells (blue) but not in lung cells. Scale bars, 50 μm.
(D) Immunofluorescent images stained with GFP (green) and DAPI (Blue) in lung metastatic tumor at low (top panels, scale bars, 200 μm) and high (bottom panels, 20 μm) magnification.
See also Figure S2
Cell lines derived from NOS form aggressive high-grade–type tumors in nude mice
To further characterize these tumors, cell lines were derived from primary tumors collected from a variety of locations from different cNICD mice: one of the tumors (T14) was from the femur, one (T6) from the tibia, one (T12) from the humerus and one (T3) from the scapula. All tumors yielded fibroblastic-like cells, and immunostaining confirmed GFP expression in tumor cells (data not shown). The resulting cell lines (called NOS-T3, NOS-T6, NOS-T12, and NOS-T14) were fully immortalized. In order to investigate their tumorigenic potential, we subcutaneously injected the NOS cells into immuno-compromised mice. All yielded substantial masses between 14 and 28 days. The resulting tumors closely resembled the parental osteosarcomas and were poorly differentiated or undifferentiated in nature (Figure 3A). Macroscopically, we did not observe lung metastases in the transplanted mice, perhaps due to the extremely rapid tumor development in these mice, which may not allow for metastatic disease to become evident. We also performed intra-tibial injection of NOS-T14 into nude mice (Figure 3A) (n=4), which induced the formation of masses in the injected leg (Figure S3A-B). Altogether, the NOS cell lines retained their ability to form high-grade–type osteosarcoma in vivo.
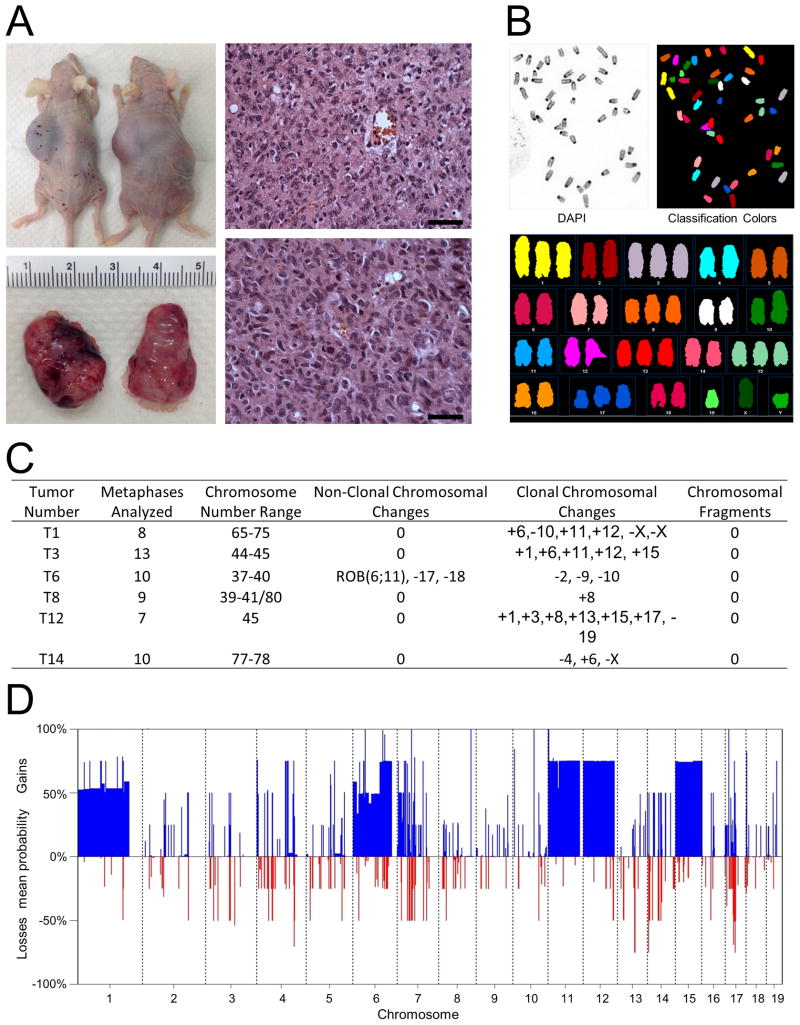
Figure 3. Transplanted tumors and genomic instability in NOS.
(A) Photos of representative nude mice (top left) and subcutaneous tumors (bottom left) obtained from the mice 14 days after a single s.c. injection with the NOS-T12 cell line into their flanks and H&E-stained section of primary OS T12 tumor (top right) and transplanted tumors (bottom right) derived from the injected mice. Scale bars, 50 μm.
(B) Representative spectral karyotyping (SKY) evaluation of harvested cells from NOS-T12 cell line.
(C) Summary of SKY results for the indicated tumors.
(D) Summary of percentages of gains (positive axis) and losses (negative axis) for chromosomes 1–19, as determined by analysis of copy number variation (CNV), compared tumor to non-tumor controls using whole genome sequencing (CNV-seq).
See also Figure S3.
Notch-induced tumors exhibit cytogenetic complexity with recurrent chromosomal changes
To examine whether the Notch-induced tumors develop genomic instability, we first performed spectral karyotyping (SKY) on early passage cultures derived from six independent primary tumors (T3, T6, T8, T12, T14, and T29). NOS primary cells consistently presented high levels of aneuploidy, a characteristic of human OS (Figure 3B). Chromosomal aberrations were found in all samples as clonal and nonclonal, nonrecurrent alterations (Figure 3C), and also included nonrecurrent unbalanced translocations (Figure S3C). As with human OS, none of the translocations were recurrent. Two samples were found to be nearly tetraploid, a state also observed in human OS (Figure S3D). We did not detect small chromosomes nor fragments. Next, we examined DNA copy number variants (CNV) using CNV-sequencing (Daines et al., 2009). Genomic imbalances were detected in all tumors examined (T1, T6, T14, T29), with gains and losses ranging from segmental to focal, along with microamplifications and deletions. The frequency of chromosomal numerical abnormalities (CNAs) along each chromosome was calculated, revealing many recurrent events in the primary tumors (Figures 3D and S3E). Somatic CNAs arising in cancer can mediate rate limiting steps in tumorigenesis through the gain of oncogene function and/or the loss of tumor suppressors. The substantial changes included recurrent gains of chromosomes 1, 6, 11, 12, and 15 and loss of chromosomes 7, 14 and 17, all present in at least 50% of cases (Figure 3D). Some of the gains identified in primary tumors, namely chromosomes 6 and 11, are homologous to aberrations seen in human chromosomes 7 and 17, respectively; these chromosomes are frequently gained and recurrently altered in human OS (Yen et al., 2009). Altogether, cytogenetic complexity with recurrent chromosomal changes suggests that genomic instability plays a key role in the development of Notch-induced tumors, and that the NOS lesions from cNICD mice share certain key downstream target genes and pathways for tumor initiation and progression.
Rbpj is required for NOS initiation in cNICD mice
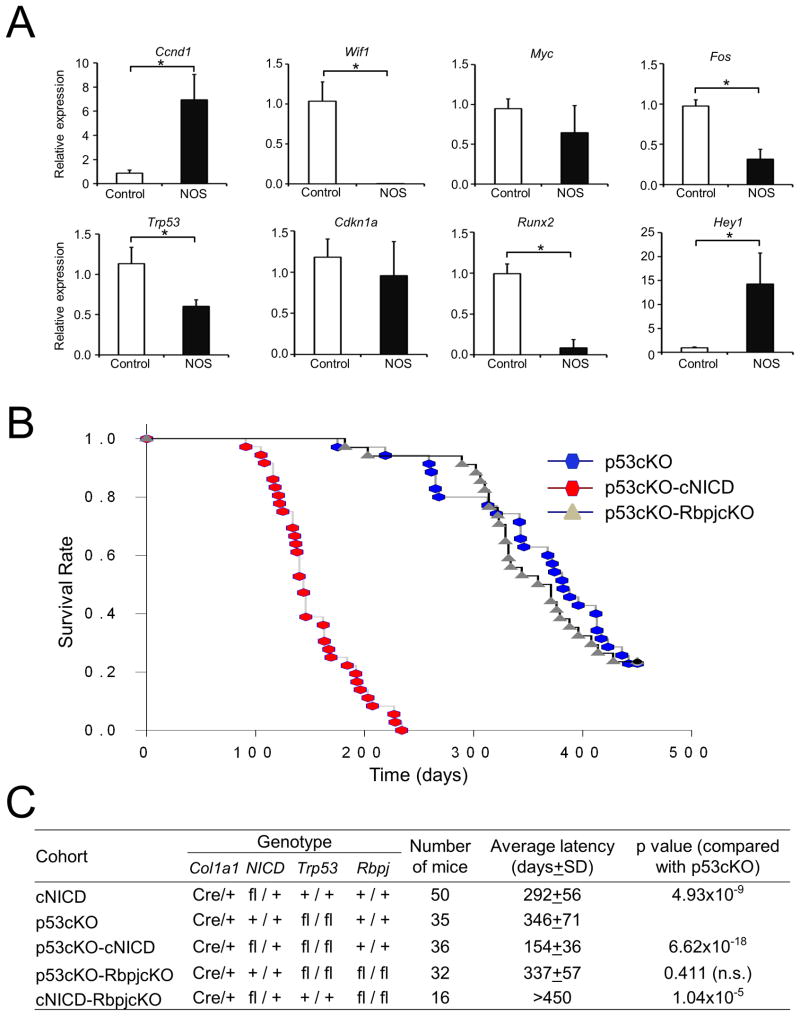
Our previous work showed that the osteosclerotic phenotype manifested in cNICD mice was dramatically reversed when the effect of the Notch gain of function was completely rescued by breeding these mice onto a RbpjFlox/Flox background (Col1a1 2.3kb-Cre; Rosa26NICD ;RbpjFlox/Flox, thereafter cNICD-RbpjcKO) (Tao et al., 2010b). This suggests that the effects of gain of Notch function in osteoblasts primarily occurs via canonical Notch signaling and is completely dependent on Rbpj function. We further investigated this by following these mice for a longer period of time to detect OS formation. We observed that the cNICD-RbpjcKO mice had a 0% incidence of bone tumors by 450 days of age compared to all cNICD mice, which developed bone tumors and died on average between 163 and 442 days (a mean of 290 days) (Figure 1B). These data suggest that Rbpj and its targets are specifically responsible for initiating the downstream events in the development of OS in vivo. Mechanistically, as previously shown, this pathological Notch gain of function likely acts through its targets, including Hes and Hey proteins, to positively regulate pro-proliferation factors such as cyclin D1 and negatively regulate pro-differentiation factors such as the Runx2 transcription factor (Engin et al., 2008; Hilton et al., 2008; Zamurovic et al., 2004; Zanotti et al., 2008). Consistent with this phenomenon, we found that Hey1 and Ccnd1 were continually up-regulated to high levels in osteosarcomas from cNICD mice, whereas Runx2 was significantly suppressed (Figure 4A). This is in agreement with recent reports showing that cyclin D1 and Runx2 are significantly involved in chromosomal instability and tumorigenesis in mice (Casimiro et al., 2012; Zaidi et al., 2007). Also, low level of expression of Runx2 is frequently observed in poorly-differentiated OS (Lengner et al., 2006; Mutsaers et al., 2013). We examined the expression of additional genes associated with human OS and found reduced expression of Trp53, a tumor suppressor gene. Interestingly, we also found highly decreased expression of Wnt inhibitory factor 1 (Wif1), a recently identified tumor suppressor gene in OS (Figure 4A). Reduction of WIF1/Wif1 expression has been associated with epigenetic silencing in human OS and accelerated progression of OS in mice (Kansara et al., 2009). We did not detect increased expression of Cdkn1a and proto-oncogenes Fos and Myc in NOS; overexpression of these proto-oncogenes results in the development of osteogenic tumors in mice and has been frequently detected in human OS (Grigoriadis et al., 1993; Shimizu et al., 2010). Altogether, these data support our hypothesis that Notch activation in committed osteoblastic lineage cells may be a dominant mechanism in OS initiation and progression.
Figure 4. Genetic interaction of Notch activation and p53 loss of function in OS development.
(A) Quantitative RT-PCR analysis of genes associated with human OS comparing Notch-induced OS with normal bone (mean ± SD of three mice per group). *p<0.05 (Student's t test).
(B) Kaplan-Meier survival plots of p53cKO, p53cKO-cNICD and p53cKO-RbpjcKO mice.
(C) Summary of genetic studies from each cohort of mice. Mice without bone tumors were sacrificed by 450 days of age in cohorts of p53cKO (8 of 35), p53cKO-RbpjcKO (8 of 34), and cNICD-RbpjcKO (16 of 16). The p values were calculated using SigmaPlot through all Pairwise Multiple Comparison Procedures (Holm-Sidak method). fl, Flox; n.s, not significant.
See also Figure S4
Tumor initiation and progression in cNICD mice are accelerated by loss of p53, but loss of Rbpj does not prevent tumorigenesis in p53 null mice
To assess the functional link between Notch and other molecules associated with human OS such as p53, and the role of Notch signaling in an established spontaneous model of OS, we employed genetically engineered mice harboring the p53 mutation. In human OS, deficiency of p53 may serve as an OS driver given that a small percentage of patients with Li-Fraumeni syndrome carrying TP53 germline mutations develop OS (Li and Fraumeni, 1969). In mice, several studies have demonstrated that osteoblast-specific deletion of Trp53 leads to a high incidence of OS (Berman et al., 2008; Lengner et al., 2006; Lin et al., 2009; Walkley et al., 2008). We observed that approximately 80% of Col1a1 2.3kb-Cre;Trp53Flox/Flox (thereafter p53cKO) mice developed OS and died within 450 days (a mean of 370 days) (Figure 4B). Our results are consistent with a previous report of this mouse model (Lin et al., 2009). Compared to cNICD mice, p53cKO mice began developing OS around 175 days, but its progression was significantly slower (Figures 4B-C and S4A-B). To examine whether Notch gain of function modulates the tumorigenic process driven by p53 inactivation, we bred mice with the conditional NICD allele into an osteoblast-specific p53 null background. Adult Col1a1 2.3kb-Cre;Rosa26NICD; Trp53Flox/Flox (thereafter, p53cKO-cNICD) mice died between 91 and 234 days (a mean of 154 days), a time frame significantly shorter than that observed in p53cKO or cNICD mice (Figures 4B-C and S4A-B). We observed that 30 of p53cKO-cNICD mice spontaneously developed OS (Figure S4C-E). Malignant tumors from these mice closely resembled NOS in cNICD mice with regard to macroscopic appearance (Figure S4Ea-d), skeletal location and radiographic features (Figures S4C and S4Ee-I), rate of lung metastasis (approximately 50% of the examined mice) (Figure S4F-G), as well as cellular morphology and distribution of targeted GFP-positive cells (Figure S4H). Notably, we found a significantly higher incidence of OS in the axial skeleton, namely the spine of the p53cKO-cNICD mice (Figures S4C and S4Ec, g, k). Regarding treatment of human patients with primary tumors in the axial skeleton, the difficulty in achieving complete surgical resection contributes to the higher risk of local recurrence and to a dismal survival outcome (Jawad et al., 2011).
These data suggest that the p53 deletion in the cNICD mice significantly influences not only tumor initiation and progression but also a preference for tumor location. On the other hand, to examine whether the Rbpj or Notch canonical pathway is required for tumorigenesis induced by p53 loss of function, we bred mice carrying the Notch loss-of-function allele (RbpjFlox) onto the osteoblast-specific p53-null background. Approximately 80% of Col1a1 2.3kb-Cre;Trp53Flox/Flox;RbpjFlox/Flox (hereafter, p53cKO-RbpjcKO) mice died within 450 days (a mean of 378 days) (Figure 4B-C). This cohort of mice spontaneously developed OS at a rate similar to that observed in the cohort of p53cKO mice. The lack of a significant effect of loss of Rpbj on a p53-null background could be due to the contribution from Rbpj-independent or non-canonical Notch signaling. Non-canonical signaling, primarily reported in Drosophila but still poorly characterized in mammalian systems, may be activated on a p53-null background (Kopan and Ilagan, 2009). Our data show that Notch activation can independently initiate OS and also act synergistically with loss of p53 in tumor progression, suggesting that the genetic interaction between Notch and p53 promotes OS initiation and progression, although Rbpj or canonical Notch function is dispensable for p53-induced tumorigenesis in the context examined.
The transcriptional profile of murine NOS is similar to that of human OS
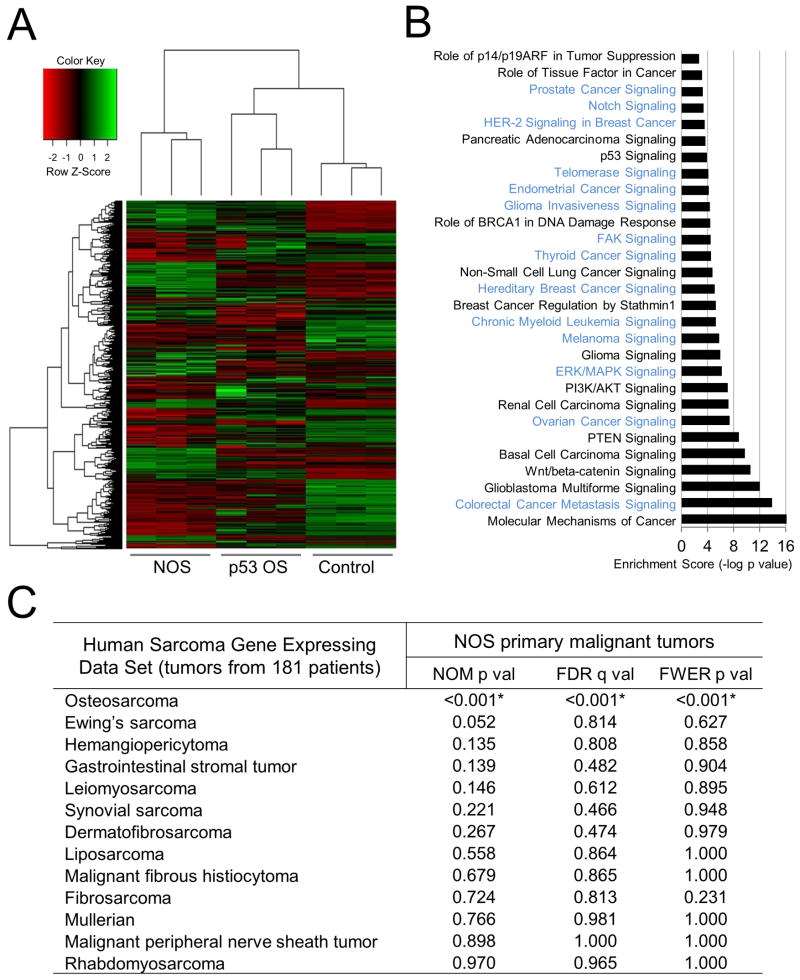
To determine whether the similarity between NOS and OS extends to the transcriptomic level, we performed transcriptional profiling analysis on NOS tumors and control bone samples from wild-type mice. For this purpose, total RNAs were extracted from NOS tumors with similar histological features from the same anatomic locations and then subjected to whole transcriptome shotgun sequencing (RNA-seq) analysis. The samples generated an average of 48.8±9.8 million mapped reads, which aligned to 14,495 ± 313 genes at ≥0.1 reads per kilobase per million (RPKM). Since murine OS induced by inactivation of p53 exhibits transcriptional profiles comparable to human OS (Walkley et al., 2008), we compared the expression profiles of tumors from both models. Clustering analysis demonstrated a highly similar expression signature among osteosarcomas induced by either Notch or loss of p53. An unsupervised hierarchical clustering and heat map of the samples based on differentially expressed genes is representatively shown in Figure 5A and Table S1-2. Ingenuity Pathway Analysis (IPA) of the significantly regulated genes in the mouse tumors compared to control bone tissues highlighted that the two types of tumors shared frequently altered canonical signaling pathways, including Wnt/beta-catenin, PTEN, PI3K/AKT and p53, as well as molecular mechanisms of cancer (Figure 5B and Table S3-4). We also identified a few signaling pathways differentially regulated in NOS, including Notch, ERK/MAPK and Sonic Hedgehog, which play critical roles in tumorigenesis and targeted therapies (Dorsam and Gutkind, 2007; Ng and Curran, 2011; Ranganathan et al., 2011). In addition, RNA-seq analysis of NOS tumors also revealed dysregulation of many components in Notch and p53 signaling pathways, including Notch1, Notch3, Il6, Cdkn2a, and Mdm2 (Figure S5A). Up-regulation of Notch receptors and other components of the Notch signaling pathway were observed in several types of tumors, including T-ALL and human OS (Engin et al., 2009; Koch and Radtke, 2007; Zender et al., 2013), although we do not know why they are up-regulated in NOS. Overall, we observed conservation and alteration in gene expression profiling from OS induced by different drivers, which may provide insight into the causative roles of either Notch gain of function or p53 loss of function in tumorigenesis.
Figure 5. Expression signature of NOS compared to p53-loss-induced OS.
(A) Unsupervised hierarchical clustering of mouse samples and heat map of RNA-sequencing data between Notch OS or p53-loss-induced tumors (p53 OS)
(B) Ingenuity Pathways Analysis (IPA) of cancer canonical signaling pathways associated with significantly regulated genes (p<0.05) in Notch OS versus control. Significantly changed pathways only observed in NOS are colored in light blue, and those changed in both NOS and p53-induced OS are shown in black.
(C) GSEA results for each of the indicated human sarcomas for NOS primary malignant tumors. Asterisks indicate statistical significance. NOM p val, nominal p value; FDR, false discovery rate; FWER, family-wise error rate. See also Figure S5, Tables S1, S2, S3, and S4.
To further explore the extent of the molecular similarity between human OS and the tumors induced by Notch activation in mice, we compared our RNA-seq expression profiles of NOS tumors with microarray-derived expression profiles from several published human sarcoma expression data sets (Baird et al., 2005; Henderson et al., 2005). To pursue this, we employed Gene Set Enrichment Analysis (GSEA) for interpreting gene expression data. The murine tumor and control expression data in units of FPKM were normalized across samples and converted to orthologous human genes. Enrichment of a given gene set was defined as having p value <0.05, a false discovery rate (FDR) q value <0.15, and a family-wise error rate (FWER) p value <0.05. Notably, primary NOS tumors showed a significant enrichment for human OS, but not other human sarcomas, across entire human tumor data sets from the Baird (Figure 5C) and the Henderson data (data not shown).
To test whether mouse NOS tumors display altered expression of genes contained within the regions cytogenetically defined in human OS, we employed cytogenetic region enrichment analysis (CREA) (Walkley et al., 2008). We found that NOS displayed significant enrichment at chromosomal region 4Q27-33 (Figure S5B). Additionally, to examine the extent of Notch activation in human OS we performed expression analysis of the established Notch target gene HEY1 using a recently published microarray data set of 34 patient samples (Paoloni et al., 2009). Interestingly, we found high HEY1 expression levels in a portion of OS patients (Figure S5C), consistent with our previous finding that Notch signaling is significantly up-regulated in human OS samples (Engin et al., 2009). Altogether, comparisons with human tumor expression data highlight the similarity between our mouse tumor model and human OS.
Committed osteoblasts are a potential source of tumor cells in osteosarcomagenesis
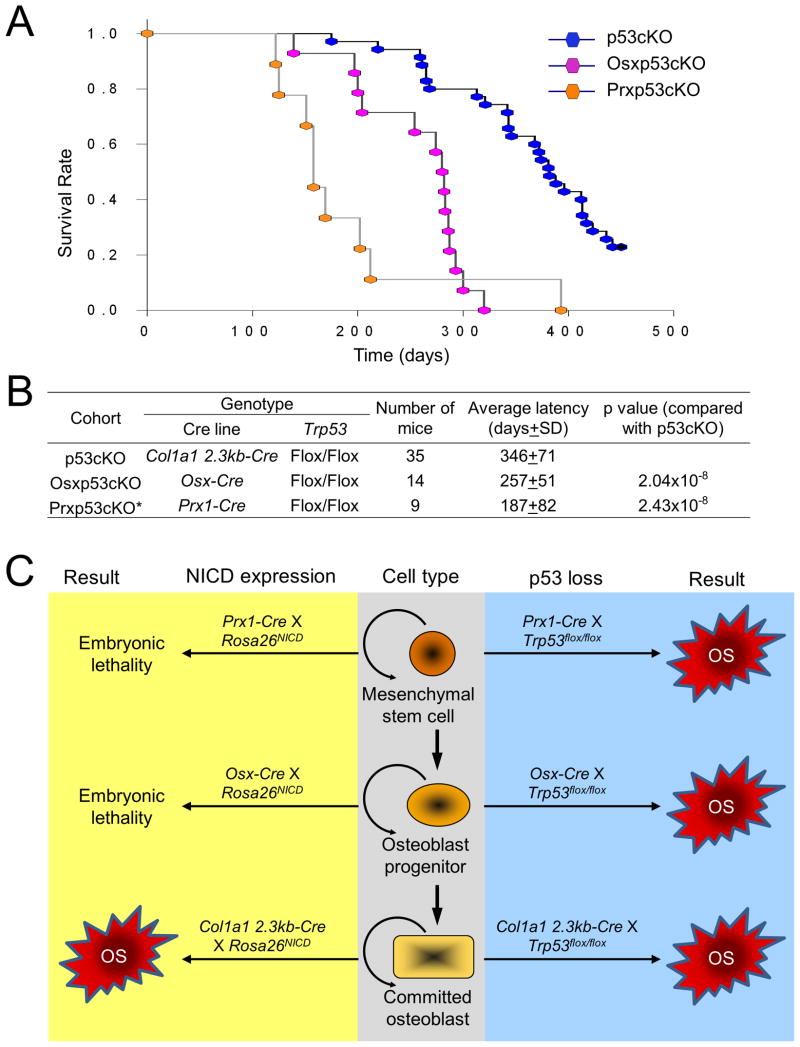
Noting that p53-loss-induced OS cells can arise from a variety of stages of mesenchymal/osteoblast differentiation, we investigated whether Notch activation at any stage, if expressed early, could induce OS. Also, to compare the cells of origin in two murine models, Notch-induced and p53-loss-induced OS, we first bred conditional Trp53Flox/Flox mice to either Prx1-Cre or Osx-Cre mice. Prx1-Cre is widely expressed in mesenchymal stem cells (MSC) or progenitors of the developing mammalian limbs and head, whereas Osx-Cre is expressed in osteoblast precursors in a transition stage from a multi/bipotential precursor to the osteoblast lineage (Hilton et al., 2008; Walkley et al., 2008). We observed that Prx1-Cre;Trp53Flox/Flox (thereafter Prxp53cKO) mice (n=9) developed OS or other tumors and died within 393 days (a mean of 187 days) and that the Osx-Cre;Trp53Flox/Flox (thereafter Osxp53cKO) mice (n=14) developed OS and died within 320 days (a mean of 257 days) (Figure 6A). Our observations are consistent with previous reports of these mouse models (Lin et al., 2009; Walkley et al., 2008). Compared to our p53cKO mice, Prxp53cKO and Osxp53cKO mice developed OS in a significantly shorter time frame (Figure 6B). This suggests that OS-originating cells come from different stages of differentiation, spanning early to late stages (Prx1-Cre>Osx-Cre>Col1a1 2.3kb-Cre). We next investigated if NICD expression at early stages of mesenchymal/osteoblast differentiation leads to OS. We started by breeding the conditional Rosa26NICD mice to either Prx1-Cre, or Osx-Cre mice. Surprisingly, no Prx1-Cre;Rosa26NICD, or Osx-Cre;Rosa26NICD pups were generated; all compound mutant embryos were abnormal and significantly smaller compared to their littermates between E15.5 and E18.5, indicating that embryonic lethality (Figure 6C and data not shown). This suggests that expression of NICD was not tolerated or was toxic to cells at early stages, likely due to suppression of MSC and progenitor differentiation. The above results suggest that the two murine models have common cells of origin in a population of committed osteoblasts, but the tumors induced by p53 loss have a wider range of cellular origin (Figure 6C).
Figure 6. Candidate cells of origin in murine OS models.
(A) Kaplan-Meier survival plots of p53cKO, Osxp53cKO and Prxp53cKO mice.
(B) Summary of genetic studies from each cohort of mice. The p values were calculated using SigmaPlot (Holm-Sidak method). *There is no significant difference (p=0.153) between Osxp53cKO and Prxp53cKO.
(C) Summary of results from various genetic experiments. Loss of function of p53 can induce OS at the early and late stage of MSC differentiation, but Notch gain of function can only induce OS at a late stage.
Discussion
In this study we model bone tumors in a genetically engineered mouse and show that Notch activation in osteoblasts is sufficient to initiate OS-like solid tumors recapitulating the defining features of human OS. We also show a robust synergy between Notch and p53 in tumorigenesis, indicating that Notch gain of function greatly promotes progression of OS initiated by loss of p53. Our transgenic models may provide valuable platforms for probing cancer genetics and targeted therapeutics.
Expression of an oncogene in a Cre/loxP-based conditional approach from the Rosa26 locus has been recently used for establishing a few mouse genetic models of cancers derived from mesenchymal cells, including synovial sarcoma (SS) expressing SS18-SSX2 (Haldar et al., 2007), and clear cell sarcoma (CCS) expressing EWS-ATF1 fusion protein (Straessler et al., 2013). We applied the same targeted strategy and generated a Notch OS mouse model by expressing the truncated protein NICD. A comparison of the three models highlights fully penetrant tumorigenesis when initiated by a cell-type-specific Cre expression. Expression of either SS18-SSX2 or EWS-ATF1, or NICD, in most cells in vivo was toxic. The later onset of detectable malignant tumors in the Notch OS mouse model relative to the SS and CCS models suggests that additional hits may be required to foster tumor progression. This may also represent a fundamental difference between the transformation ability of the two types of oncogenes (translocation-associated fusion versus truncated protein mutation). Human cancers have been modeled in mice by cell-specific expression of an activating truncated form of the Notch receptor (Allen et al., 2011; Robey et al., 1996; Zender et al., 2013). Gain-of-function mutations in NOTCH1 have been found in approximately 10% of cases of non-small-cell lung cancer (Lawrence et al., 2013; Westhoff et al., 2009), the leading cause of cancer-related deaths. Notably, the Notch pathway is very active in primary tumors and in cell lines from patients with liver cancer and OS (Engin et al., 2009; Zender et al., 2013); however, Notch mutations in low-frequency sarcoma remain underappreciated owing to scarce resources. Interestingly, somatic point mutations in NOTCH1 have been recently found in most of the 21 tumor types analyzed in exome sequences from 4,742 human cancers (Lawrence et al., 2014), suggesting that OS and other types of cancers of mesenchymal origin may contain these mutations.
In agreement with our model, Molyneux et al. showed that osteoblast-specific expression of simian virus 40 (SV40) large T antigen (TAg) oncoprotein induces a range of osteoblastic lesions, from benign osteoma to poorly differentiated OS with aggressive metastatic features (Molyneux et al., 2010). Transgenic mice expressing the oncogene c-fos developed osteoblastic and chondroblastic OS (Grigoriadis et al., 1993). Commonly, oncogene-induced OS tumors have a long latency period. In our mouse model, once visible, OS tumors grow rapidly and cause death within a month, whereas osteomas grow very slowly, frequently until death occurs. Interestingly, OS lesions and osteomas seldom occur at the same anatomic site. It is currently unknown why the same oncogene leads to two types of tumors, but the transforming ability of Notch mutations may be dependent on the cell types in which the gene is expressed. This is evident from the fact that lung-specific expression of Notch truncated protein induces only benign tumors, but fails to cause progression to lung cancer (Allen et al., 2011), whereas liver-specific expression definitely results in malignant tumors (Zender et al., 2013). Alternatively, we speculate that a different hit induced by micro-environmental cues may promote different downstream events, as seen in the TAg-induced model, in which additional loss of Prkar1a defines a subclass of high-grade–type OS (Molyneux et al., 2010).
It is not unexpected that in our p53-loss–driven OS model, adding a “second hit” Notch activation mutation significantly shortens tumor latency and aggressiveness, suggesting a critical role for Notch signaling in OS progression. Other tumor models based on Notch activation show similar results. Transgenic mice expressing NICD specifically in T cells developed T-cell leukemia at a frequency of about 20% by five months of age (Robey et al., 1996), but once a second mutation was added penetrance of leukemia increased to 100% and tumor latency decreased to 2 months (Beverly and Capobianco, 2003). By supplementing Myc expression, lung-specific expression of the Notch truncated protein enabled progression to lung cancer and metastases (Allen et al., 2011). Additionally, in our cNICD model, p53 deletion can dramatically accelerate tumor progression, suggesting that p53 certainly constitutes a “second hit”. We propose that the second hit is the factor that defines whether the tumor is benign or malignant in the cNICD mice.
Using RNA-seq and IPA, we demonstrated a high degree of conservation of changes in gene expression and signaling pathways, including Wnt, between that model and ours. Consistent with a recent report (Mutsaers et al., 2013), we found that expression of Wnt secreted antagonists, including Dkk1, Sost, and Wif1, was significantly decreased in NOS, implicating Wnt activation in this context. This “Notch on-Wnt on” model has been supported by a recent study showing that Notch activation in cells of the osteoblastic lineage up-regulates Wnt signaling through suppression of Dkk1 and Sost (Canalis et al., 2013a). Moreover, Wif1 is often epigenetically silenced in human OS, and its targeted disruption accelerates OS development in mice (Kansara et al., 2009). It is currently unknown whether the epigenetic mechanism is accountable for Wif1 silencing in NOS tumors and whether ectopic Wif1 expression inhibits NOS cell growth. On the one hand, aberrant activation of the WNT pathway is associated with numerous cancers including OS (Gorlick and Khanna, 2010). Elevated levels of nuclear localized β-catenin, a critical mediator of the canonical Wnt pathway, have been detected in the majority of OS tumors, as well as sporadic mutations of β-catenin (Wagner et al., 2011). On the other hand, several studies suggest that down-regulation of Wnt signaling has an important role in OS pathogenesis (Cai et al., 2009). So far it is unclear whether up- or down-regulation of Wnt in vivo promotes OS formation and metastasis. In addition, increasing evidence supports that Wnt and Notch signaling interact with each other in both synergistic and antagonistic manners in a context-dependent fashion (Tao et al., 2010a), but it is not clear how they cooperate to initiate development of OS tumor cells in the skeletal system.
Ages of patients with OS present a bimodal pattern, with the first peak in adolescents and the second in the elderly (Jawad et al., 2011). In the latter group OS frequently arises under chronic conditions, such as Paget's disease resulting from an inflammatory process. Although our cNICD mice exhibit generalized osteosclerotic disease (Tao et al., 2010b), OS induction does not require additional inflammation. Nonetheless, we found that interleukin-6 (IL-6), a major mediator of inflammation, is greatly increased in tumor tissues. Notably, IL-6 is a downstream effector of Notch signaling and its expression is mediated by Hey1 in osteoblasts (Sethi et al., 2011). In addition, this cascade of Notch1-Hey1-IL-6 released from osteoblasts directly activates osteoclastogenesis, which acts a “Vicious Cycle” of cancer osteolytic metastases (Tao et al., 2011). It is conceivable that the severe osteolytic phenotype observed in the tumors from cNICD mice may partially result from this “Vicious Cycle”.
Demarest et al. showed that expression of the Notch truncated protein retains its requirement for maintenance of Notch-induced T-ALL (Demarest et al., 2011). It will be interesting to know whether continuous presence of NICD is obligatory for NOS progression. This possibility warrants further investigation for therapeutic purposes. A major challenge in OS is that no specific treatment is available for patients unsuitable for intensive chemotherapy and surgical regimens. We and others have shown that inhibition of Notch signaling impedes human OS growth (Engin et al., 2009; Tanaka et al., 2009). However, this study shows that blockade of the Notch canonical pathway in OS cells fails to halt tumor progression. We speculate that the effect of Notch inhibition by γ-secretase inhibitor (GSIs) on OS growth emanates from the Notch non-canonical pathway in a non-cell-autonomous fashion. Sethi et al. support our proposition with their findings that cancer progression is impeded by disrupting Notch signaling in normal neighboring cells rather than in tumor cells (Sethi et al., 2011). By contrast, GSIs are effective in blocking all Notch signals in tumor tissues, but their application as anti-cancer drugs is unlikely due to severe on-target intestinal toxicity. Notably, newly developed humanized antibodies against Notch ligands have been tested in mice without limiting adverse effects (Tran et al., 2013). These findings open new perspectives for the continued development of Notch inhibitors for cancer therapy.
The cell of origin in human OS remains elusive. Much of our understanding of cells of origin has derived from the study of OS mouse models generated by cell-type-specific loss of p53 (Berman et al., 2008; Lengner et al., 2006; Lin et al., 2009; Walkley et al., 2008). Despite the difference in stages of Cre expression, all those models develop OS with clinical features very similar to those of human OS, although latencies vary among them. These data imply that the source of OS cells may be MSC, osteoblast progenitors, or committed osteoblasts. We generated NOS tumors within the immature osteoblast lineage expressing Col1a1 2.3kb-Cre, suggesting that the committed osteoblasts are favored candidate cells of origin in our model. However, our data do not exclude Prx1-Cre or Osx-Cre-expressing cells as potential sources of OS. Somatic mutations may sporadically occur in only some of these cells at earlier stages in human tissues. By breeding NICD mice with Prx1-Cre or Osx-Cre mice, the Notch mutated protein is practically present in all Cre-expressing cells and their lineage, thereby potentially overburdening the organism and leading to embryonic death. Moreover, Prx1-Cre or Osx-Cre-expressing cells are not exclusive to the osteoblast lineage and may serve important roles in other type of cells (Hilton et al., 2008; Walkley et al., 2008). We cannot exclude that osteoblasts at a more differentiated stage than cells expressing Col1a1 2.3kb-Cre are also capable of initiating OS. Indeed, recent studies have shown that Notch activation in different population cells at later stages of osteoblastic differentiation results in overlapping, but distinct, pathological bone phenotypes (Canalis et al., 2013b). Altogether, these data support that committed osteoblasts are the favorite candidate for cells of origin of OS.
In summary, we have revealed the critical role of Notch gain-of-function mutations as a cause of OS in mice and advanced our understanding of the genetic factors and signaling pathways in pathogenesis of OS. Our study also warrants future screening and identification of patients with mesenchyme-derived tumors affected by mutations in Notch receptors, and in other components of the pathway, for developing targeted treatment strategies. Importantly, by studying gain- and loss-of-function genetic models in vivo, we have generated robust insights into how signaling pathways such as Notch and p53 contribute to osteosarcomagenesis, which may eventually provide a rational strategy for devising specific therapies targeting pathways leading to OS. Finally, our study supports the hypothesis that Notch activating mutations can act as a common triggering mechanism in cells of mesenchymal origin such as committed osteoblasts.
Experimental Procedures
Animal husbandry
Conditional knockout mice (RbpjFlox/Flox, Trp53Flox/Flox) and transgenic mice (Col1a1 2.3kb-Cre, Rosa26NICD) have been previously described (Dacquin et al., 2002; Han et al., 2002; Jonkers et al., 2001; Murtaugh et al., 2003). Prx1-Cre, Osx-Cre, and Rosa26LacZ reporter mice were purchased from The Jackson Laboratory (Bar Harbor, ME, USA). Genotyping by PCR was performed as described. The mice were maintained on a hybrid 129 X C57BL/6 background and housed in a specific pathogen-free facility under controlled conditions of light, temperature and humidity. These studies were conducted in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals and approved by the Baylor College of Medicine Institutional Animal Care and Use Committee.
Radiography, histological analysis, immunostaining, qRT-PCR, spectral karyotyping analysis, and statistical analyses
Mouse tumors were extracted after asphyxiation and were fixed overnight in 10% formalin solution prior to paraffin embedding. Skeletal preparations, radiography, histological analysis, immunostaining, qRT-PCR, spectral karyotyping analysis, and statistical analyses are described in the Supplemental Experimental Procedures.
RNA sequencing, whole-genome sequencing, and copy number variation analysis
Sequencing was performed on a HiSeq 2000 platform at the Human Genome Sequencing Center of Baylor College of Medicine according to the manufacturer's protocol. For RNA sequencing, libraries were prepared according to the dUTP protocol. Illumina paired-end whole genome shotgun libraries were prepared from tumor genomic DNA or non-tumor control tissue, which was used as reference genome. For a detailed description of sample preparation and bioinformatics analyses for high-throughput data, see the Supplemental Experimental Procedures.
Human OS gene expression and GSEA
Human data from Paoloni et al. were downloaded from the Gene Expression Omnibus gateway (http://www-ncbi-nlm-nih-gov.ezproxyhost.library.tmc.edu/geo/) using accession numbers GSE16091. Human sarcoma expression data sets were downloaded from the Baird and Henderson studies (Baird et al., 2005; Henderson et al., 2005). Microarray data normalization and statistical analyses were performed using the “limma: Linear Models for Microarray Data” of the R statistical package. GSEA was performed as described (http://www.broad.mit.edu/gsea). We applied default algorithm settings and each data set underwent 1000 permutations. A nominal p value <0.05 and an FDR q value <0.15 were considered significant.
Supplementary Material
Highlights.
Notch 1 intracellular domain induces bone tumors in mice with 100% penetrance
Mouse osteosarcoma initiated by Notch activation mimics the human disease
Notch activation and p53 loss independently and synergistically induce osteosarcoma
Committed osteoblasts are a potential source of tumor cells in osteosarcomagenesis
Significance.
Human OS is an aggressive mesenchyme-derived tumor associated with high mortality that affects mostly adolescents. The survival rate for OS patients has not improved substantially over the past decades. Our findings reveal a critical role for Notch gain of function as a cause of OS and advance our understanding of the pathogenesis of this disease as well as its cells of origin. Our study also supports the hypothesis that Notch-activating mutations can act as an OS-initiating mechanism in cells of mesenchymal lineage such as committed osteoblasts. Moreover, our OS models constitute valuable platforms for probing cancer genetics and developing more effective therapies for OS.
Acknowledgments
We thank C. Lietman, S. Chen, Y. Bae, P. Campeau, L. Wang, J. Yustein, K. Scott, L. Kurenbekova, and members of the Lee lab for technical assistance and/or discussions. We thank P. Fonseca for editorial assistance. This work was supported by the NIH [AR061565 to J.T., HD022657 and DE016990 to B.L.] and the Cancer Prevention and Research Institute of Texas [RP101017 to B.L.]. This work was also supported by the BCM Intellectual and Developmental Disabilities Research Center [HD024064] from the Eunice Kennedy Shriver National Institute of Child Health & Human Development.
Footnotes
Accession Numbers: The raw sequence data from RNA-sequencing and whole-genome sequencing generated in this study have been deposited in NCBI SRA database under accession number SRP036832 and SRP036865.
Supplemental Information: It includes 5 figures, 4 tables, and Supplemental Experimental Procedures.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Allen TD, Rodriguez EM, Jones KD, Bishop JM. Activated Notch1 induces lung adenomas in mice and cooperates with Myc in the generation of lung adenocarcinoma. Cancer research. 2011;71:6010–6018. doi: 10.1158/0008-5472.CAN-11-0595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baird K, Davis S, Antonescu CR, Harper UL, Walker RL, Chen Y, Glatfelter AA, Duray PH, Meltzer PS. Gene expression profiling of human sarcomas: insights into sarcoma biology. Cancer research. 2005;65:9226–9235. doi: 10.1158/0008-5472.CAN-05-1699. [DOI] [PubMed] [Google Scholar]
- Berman SD, Calo E, Landman AS, Danielian PS, Miller ES, West JC, Fonhoue BD, Caron A, Bronson R, Bouxsein ML, et al. Metastatic osteosarcoma induced by inactivation of Rb and p53 in the osteoblast lineage. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:11851–11856. doi: 10.1073/pnas.0805462105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beverly LJ, Capobianco AJ. Perturbation of Ikaros isoform selection by MLV integration is a cooperative event in Notch(IC)-induced T cell leukemogenesis. Cancer cell. 2003;3:551–564. doi: 10.1016/s1535-6108(03)00137-5. [DOI] [PubMed] [Google Scholar]
- Cai Y, Mohseny AB, Karperien M, Hogendoorn PC, Zhou G, Cleton-Jansen AM. Inactive Wnt/beta-catenin pathway in conventional high-grade osteosarcoma. The Journal of pathology. 2009;220:24–33. doi: 10.1002/path.2628. [DOI] [PubMed] [Google Scholar]
- Canalis E, Adams DJ, Boskey A, Parker K, Kranz L, Zanotti S. Notch signaling in osteocytes differentially regulates cancellous and cortical bone remodeling. The Journal of biological chemistry. 2013a;288:25614–25625. doi: 10.1074/jbc.M113.470492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Canalis E, Parker K, Feng JQ, Zanotti S. Osteoblast lineage-specific effects of notch activation in the skeleton. Endocrinology. 2013b;154:623–634. doi: 10.1210/en.2012-1732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casimiro MC, Crosariol M, Loro E, Ertel A, Yu Z, Dampier W, Saria EA, Papanikolaou A, Stanek TJ, Li Z, et al. ChIP sequencing of cyclin D1 reveals a transcriptional role in chromosomal instability in mice. The Journal of clinical investigation. 2012;122:833–843. doi: 10.1172/JCI60256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dacquin R, Starbuck M, Schinke T, Karsenty G. Mouse alpha1(I)-collagen promoter is the best known promoter to drive efficient Cre recombinase expression in osteoblast. Dev Dyn. 2002;224:245–251. doi: 10.1002/dvdy.10100. [DOI] [PubMed] [Google Scholar]
- Daines B, Wang H, Li Y, Han Y, Gibbs R, Chen R. High-throughput multiplex sequencing to discover copy number variants in Drosophila. Genetics. 2009;182:935–941. doi: 10.1534/genetics.109.103218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Demarest RM, Dahmane N, Capobianco AJ. Notch is oncogenic dominant in T-cell acute lymphoblastic leukemia. Blood. 2011;117:2901–2909. doi: 10.1182/blood-2010-05-286351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorsam RT, Gutkind JS. G-protein-coupled receptors and cancer. Nature reviews. 2007;7:79–94. doi: 10.1038/nrc2069. [DOI] [PubMed] [Google Scholar]
- Ellisen LW, Bird J, West DC, Soreng AL, Reynolds TC, Smith SD, Sklar J. TAN-1, the human homolog of the Drosophila notch gene, is broken by chromosomal translocations in T lymphoblastic neoplasms. Cell. 1991;66:649–661. doi: 10.1016/0092-8674(91)90111-b. [DOI] [PubMed] [Google Scholar]
- Engin F, Bertin T, Ma O, Jiang MM, Wang L, Sutton RE, Donehower LA, Lee B. Notch signaling contributes to the pathogenesis of human osteosarcomas. Human Molecular Genetics. 2009;18:1464–1470. doi: 10.1093/hmg/ddp057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engin F, Yao Z, Yang T, Zhou G, Bertin T, Jiang MM, Chen Y, Wang L, Zheng H, Sutton RE, et al. Dimorphic effects of Notch signaling in bone homeostasis. NatMed. 2008;14:299–305. doi: 10.1038/nm1712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorlick R, Khanna C. Osteosarcoma. J Bone Miner Res. 2010;25:683–691. doi: 10.1002/jbmr.77. [DOI] [PubMed] [Google Scholar]
- Grigoriadis AE, Schellander K, Wang ZQ, Wagner EF. Osteoblasts are target cells for transformation in c-fos transgenic mice. The Journal of Cell Biology. 1993;122:685–701. doi: 10.1083/jcb.122.3.685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haldar M, Hancock JD, Coffin CM, Lessnick SL, Capecchi MR. A conditional mouse model of synovial sarcoma: insights into a myogenic origin. Cancer cell. 2007;11:375–388. doi: 10.1016/j.ccr.2007.01.016. [DOI] [PubMed] [Google Scholar]
- Han H, Tanigaki K, Yamamoto N, Kuroda K, Yoshimoto M, Nakahata T, Ikuta K, Honjo T. Inducible gene knockout of transcription factor recombination signal binding protein-J reveals its essential role in T versus B lineage decision. International immunology. 2002;14:637–645. doi: 10.1093/intimm/dxf030. [DOI] [PubMed] [Google Scholar]
- Harvey M, McArthur MJ, Montgomery CA, Jr, Butel JS, Bradley A, Donehower LA. Spontaneous and carcinogen-induced tumorigenesis in p53-deficient mice. Nature genetics. 1993;5:225–229. doi: 10.1038/ng1193-225. [DOI] [PubMed] [Google Scholar]
- Henderson SR, Guiliano D, Presneau N, McLean S, Frow R, Vujovic S, Anderson J, Sebire N, Whelan J, Athanasou N, et al. A molecular map of mesenchymal tumors. Genome biology. 2005;6:R76. doi: 10.1186/gb-2005-6-9-r76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hilton MJ, Tu X, Wu X, Bai S, Zhao H, Kobayashi T, Kronenberg HM, Teitelbaum SL, Ross FP, Kopan R, Long F. Notch signaling maintains bone marrow mesenchymal progenitors by suppressing osteoblast differentiation. Nature medicine. 2008;14:306–314. doi: 10.1038/nm1716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jawad M, Cheung M, Clarke J, Koniaris L, Scully S. Osteosarcoma: improvement in survival limited to high-grade patients only. Journal of Cancer Research and Clinical Oncology. 2011;137:597–607. doi: 10.1007/s00432-010-0923-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jonkers J, Meuwissen R, van der Gulden H, Peterse H, van der Valk M, Berns A. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nature genetics. 2001;29:418–425. doi: 10.1038/ng747. [DOI] [PubMed] [Google Scholar]
- Kansara M, Tsang M, Kodjabachian L, Sims NA, Trivett MK, Ehrich M, Dobrovic A, Slavin J, Choong PFM, Simmons PJ, et al. Wnt inhibitory factor 1 is epigenetically silenced in human osteosarcoma, and targeted disruption accelerates osteosarcomagenesis in mice. The Journal of clinical investigation. 2009;119:837–851. doi: 10.1172/JCI37175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koch U, Radtke F. Notch and cancer: a double-edged sword. Cell Mol Life Sci. 2007;64:2746–2762. doi: 10.1007/s00018-007-7164-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kopan R, Ilagan MX. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 2009;137:216–233. doi: 10.1016/j.cell.2009.03.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence MS, Stojanov P, Mermel CH, Robinson JT, Garraway LA, Golub TR, Meyerson M, Gabriel SB, Lander ES, Getz G. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature. 2014;505:495–501. doi: 10.1038/nature12912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence MS, Stojanov P, Polak P, Kryukov GV, Cibulskis K, Sivachenko A, Carter SL, Stewart C, Mermel CH, Roberts SA, et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature. 2013;499:214–218. doi: 10.1038/nature12213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lengner CJ, Steinman HA, Gagnon J, Smith TW, Henderson JE, Kream BE, Stein GS, Lian JB, Jones SN. Osteoblast differentiation and skeletal development are regulated by Mdm2-p53 signaling. The Journal of Cell Biology. 2006;172:909–921. doi: 10.1083/jcb.200508130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li FP, Fraumeni JF., Jr Soft-tissue sarcomas, breast cancer, and other neoplasms. A familial syndrome? Annals of internal medicine. 1969;71:747–752. doi: 10.7326/0003-4819-71-4-747. [DOI] [PubMed] [Google Scholar]
- Lin PP, Pandey MK, Jin F, Raymond AK, Akiyama H, Lozano G. Targeted mutation of p53 and Rb in mesenchymal cells of the limb bud produces sarcomas in mice. Carcinogenesis. 2009;30:1789–1795. doi: 10.1093/carcin/bgp180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molyneux SD, Di Grappa MA, Beristain AG, McKee TD, Wai DH, Paderova J, Kashyap M, Hu P, Maiuri T, Narala SR, et al. Prkar1a is an osteosarcoma tumor suppressor that defines a molecular subclass in mice. The Journal of clinical investigation. 2010;120:3310–3325. doi: 10.1172/JCI42391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murtaugh LC, Stanger BZ, Kwan KM, Melton DA. Notch signaling controls multiple steps of pancreatic differentiation. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:14920–14925. doi: 10.1073/pnas.2436557100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mutsaers AJ, Ng AJ, Baker EK, Russell MR, Chalk AM, Wall M, Liddicoat BJ, Ho PW, Slavin JL, Goradia A, et al. Modeling distinct osteosarcoma subtypes in vivo using Cre:lox and lineage-restricted transgenic shRNA. Bone. 2013;55:166–178. doi: 10.1016/j.bone.2013.02.016. [DOI] [PubMed] [Google Scholar]
- Ng JM, Curran T. The Hedgehog's tale: developing strategies for targeting cancer. Nature reviews. 2011;11:493–501. doi: 10.1038/nrc3079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paoloni M, Davis S, Lana S, Withrow S, Sangiorgi L, Picci P, Hewitt S, Triche T, Meltzer P, Khanna C. Canine tumor cross-species genomics uncovers targets linked to osteosarcoma progression. BMC genomics. 2009;10:625. doi: 10.1186/1471-2164-10-625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ranganathan P, Weaver KL, Capobianco AJ. Notch signalling in solid tumours: a little bit of everything but not all the time. Nature reviews. 2011;11:338–351. doi: 10.1038/nrc3035. [DOI] [PubMed] [Google Scholar]
- Robey E, Chang D, Itano A, Cado D, Alexander H, Lans D, Weinmaster G, Salmon P. An activated form of Notch influences the choice between CD4 and CD8 T cell lineages. Cell. 1996;87:483–492. doi: 10.1016/s0092-8674(00)81368-9. [DOI] [PubMed] [Google Scholar]
- Sethi N, Dai X, Winter CG, Kang Y. Tumor-derived JAGGED1 promotes osteolytic bone metastasis of breast cancer by engaging notch signaling in bone cells. Cancer cell. 2011;19:192–205. doi: 10.1016/j.ccr.2010.12.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimizu T, Ishikawa T, Sugihara E, Kuninaka S, Miyamoto T, Mabuchi Y, Matsuzaki Y, Tsunoda T, Miya F, Morioka H, et al. c-MYC overexpression with loss of Ink4a/Arf transforms bone marrow stromal cells into osteosarcoma accompanied by loss of adipogenesis. Oncogene. 2010;29:5687–5699. doi: 10.1038/onc.2010.312. [DOI] [PubMed] [Google Scholar]
- Straessler KM, Jones KB, Hu H, Jin H, van de Rijn M, Capecchi MR. Modeling clear cell sarcomagenesis in the mouse: cell of origin differentiation state impacts tumor characteristics. Cancer cell. 2013;23:215–227. doi: 10.1016/j.ccr.2012.12.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka M, Setoguchi T, Hirotsu M, Gao H, Sasaki H, Matsunoshita Y, Komiya S. Inhibition of Notch pathway prevents osteosarcoma growth by cell cycle regulation. British journal of cancer. 2009;100:1957–1965. doi: 10.1038/sj.bjc.6605060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tao J, Chen S, Lee B. Alteration of Notch signaling in skeletal development and disease. Ann N Y Acad Sci. 2010a;1192:257–268. doi: 10.1111/j.1749-6632.2009.05307.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tao J, Chen S, Yang T, Dawson B, Munivez E, Bertin T, Lee B. Osteosclerosis owing to Notch gain of function is solely Rbpj-dependent. J Bone Miner Res. 2010b;25:2175–2183. doi: 10.1002/jbmr.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tao J, Erez A, Lee B. One NOTCH Further: Jagged 1 in Bone Metastasis. Cancer cell. 2011;19:159–161. doi: 10.1016/j.ccr.2011.01.043. [DOI] [PubMed] [Google Scholar]
- Tran IT, Sandy AR, Carulli AJ, Ebens C, Chung J, Shan GT, Radojcic V, Friedman A, Gridley T, Shelton A, et al. Blockade of individual Notch ligands and receptors controls graft-versus-host disease. The Journal of clinical investigation. 2013;123:1590–1604. doi: 10.1172/JCI65477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Unni KK, Inwards CY. Dahlin's Bone Tumors: General Aspects and Data on 10,165 Cases. 6. Philadelphia: Lippincott Williams & Wilkins; 2009. [Google Scholar]
- Wagner ER, Luther G, Zhu G, Luo Q, Shi Q, Kim SH, Gao JL, Huang E, Gao Y, Yang K, et al. Defective osteogenic differentiation in the development of osteosarcoma. Sarcoma. 2011;2011:325238. doi: 10.1155/2011/325238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walkley CR, Qudsi R, Sankaran VG, Perry JA, Gostissa M, Roth SI, Rodda SJ, Snay E, Dunning P, Fahey FH, et al. Conditional mouse osteosarcoma, dependent on p53 loss and potentiated by loss of Rb, mimics the human disease. Genes and Development. 2008;22:1662–1676. doi: 10.1101/gad.1656808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weng AP, Ferrando AA, Lee W, Morris JPt, Silverman LB, Sanchez-Irizarry C, Blacklow SC, Look AT, Aster JC. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science (New York, NY. 2004;306:269–271. doi: 10.1126/science.1102160. [DOI] [PubMed] [Google Scholar]
- Westhoff B, Colaluca IN, D'Ario G, Donzelli M, Tosoni D, Volorio S, Pelosi G, Spaggiari L, Mazzarol G, Viale G, et al. Alterations of the Notch pathway in lung cancer. Proceedings of the National Academy of Sciences of the United States of America. 2009;106:22293–22298. doi: 10.1073/pnas.0907781106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yen CC, Chen WM, Chen TH, Chen WY, Chen PC, Chiou HJ, Hung GY, Wu HT, Wei CJ, Shiau CY, et al. Identification of chromosomal aberrations associated with disease progression and a novel 3q13.31 deletion involving LSAMP gene in osteosarcoma. International journal of oncology. 2009;35:775–788. doi: 10.3892/ijo_00000390. [DOI] [PubMed] [Google Scholar]
- Zaidi SK, Pande S, Pratap J, Gaur T, Grigoriu S, Ali SA, Stein JL, Lian JB, van Wijnen AJ, Stein GS. Runx2 deficiency and defective subnuclear targeting bypass senescence to promote immortalization and tumorigenic potential. Proceedings of the National Academy of Sciences of the United States of America. 2007;104:19861–19866. doi: 10.1073/pnas.0709650104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zamurovic N, Cappellen D, Rohner D, Susa M. Coordinated activation of notch, Wnt, and transforming growth factor-beta signaling pathways in bone morphogenic protein 2-induced osteogenesis. Notch target gene Hey1 inhibits mineralization and Runx2 transcriptional activity. The Journal of biological chemistry. 2004;279:37704–37715. doi: 10.1074/jbc.M403813200. [DOI] [PubMed] [Google Scholar]
- Zanotti S, Smerdel-Ramoya A, Stadmeyer L, Durant D, Radtke F, Canalis E. Notch inhibits osteoblast differentiation and causes osteopenia. Endocrinology. 2008;149:3890–3899. doi: 10.1210/en.2008-0140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zender S, Nickeleit I, Wuestefeld T, Sorensen I, Dauch D, Bozko P, El-Khatib M, Geffers R, Bektas H, Manns MP, et al. A critical role for notch signaling in the formation of cholangiocellular carcinomas. Cancer cell. 2013;23:784–795. doi: 10.1016/j.ccr.2013.04.019. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.