Abstract
Native osteochondral repair is often inadequate due to the inherent properties of the tissue and current clinical repair strategies can result in healing with a limited lifespan and donor site morbidity. This work investigates the use of polymeric gene therapy to address this problem by delivering DNA encoding for transcription factors complexed with the branched poly(ethylenimine)-hyaluronic acid (bPEI-HA) delivery vector via a porous oligo[poly(ethylene glycol) fumarate] (OPF) hydrogel scaffold. To evaluate the potential of this approach, a bilayered scaffold mimicking native osteochondral tissue organization was loaded with DNA/bPEI-HA complexes. Next, bilayered implants either unloaded or loaded in a spatial fashion with bPEI-HA and DNA encoding for either Runt-related transcription factor 2 (RUNX2) or SRY (sex determining region Y)-box 5, 6, and 9 (the SOX trio), to generate bone and cartilage tissues respectively, were fabricated and implanted in a rat osteochondral defect. At 6 weeks post-implantation, micro-computed tomography (micro-CT) analysis and histological scoring were performed on the explants to evaluate the quality and quantity of tissue repair in each group. The incorporation of DNA encoding for RUNX2 in the bone layer of these scaffolds significantly increased bone growth. Additionally, a spatially loaded combination of RUNX2 and SOX trio DNA loading significantly improved healing relative to empty hydrogels or either factor alone. Finally, the results of this study suggest that subchondral bone formation is necessary for correct cartilage healing.
Keywords: Polymeric gene delivery, OPF, bPEI-HA, SOX trio, RUNX-2
1. INTRODUCTION
Osteochondral injuries are an area of in depth research due to the inadequacy of native healing and the limited lifespan and quality of conventional treatments.1, 2 Cartilage tissue is often limited in its ability to regrow due to its heavy dependence on appropriate extracellular matrix generation and lack of blood flow. Tissue engineering is one area of research that could address these current inadequacies and provide an improved therapy for osteochondral injuries.
In particular, gene delivery is one developing and promising option for osteochondral injury repair in the context of tissue engineering. Gene delivery has the ability to utilize often otherwise unusable therapeutic proteins by directly inducing their expression within the cells of a target tissue through delivery of nucleotides into those cells. Osteochondral and chondral tissues are especially promising for polymeric gene delivery approaches because of the limited blood flow to the region, which can cause problems in DNA polymer complex delivery, and the potential for the delivered genes to induce differentiation of infiltrated mesenchymal stem cells (MSCs). By using transcription factors that regulate many downstream proteins, gene delivery could be capable of inducing more physiologically correct differentiation in target cells. This work utilized the transcription factors Runt-related transcription factor 2 (RUNX2), or CBFA1, which has been shown to induce osteogenic differentiation3, and SRY (sex determining region Y)-box 5, 6, and 9 (the SOX trio), which has been shown to induce chondrogenic differentiation.4, 5
To deliver these transcription factors into the cell, a gene delivery vector is needed. This work utilizes branched poly(ethylenimine) (bPEI), a commonly used polymer for gene delivery, modified with hyaluronic acid (HA). The resulting product, bPEI-HA has been shown to mitigate many of the negative effects associated with bPEI, such as its high cytotoxicity, while improving transfection efficiency and potentially providing cellular targeting through hyaladherins on the cell’s surface. 6–9
Studies have examined the role of the SOX trio and RUNX2 in chondrogenesis and osteogenesis both separately and in combination with each other and other bioactive factors.5, 10 It has been shown that the SOX trio is capable of inducing chondrogenic differentiation in target cells both in vitro and in vivo11 and that this combination of transcription factors has the potential to do so in vitro without the use of inductive factors.49 Further, when cells transduced with the SOX trio were implanted into a rat osteochondral defect for 8 weeks, they were found to promote defect healing.12 In other studies, PLGA scaffolds loaded with bPEI and bPEI based vectors complexed with DNA encoding for the SOX trio have been shown capable of inducing cartilage growth in vivo and in vitro.13, 14
RUNX2 has also been shown to be an effective driver of differentiation.15 When DNA encoding for RUNX2 is transduced into stem cells which are seeded onto scaffolds and implanted into animals, substantially more bone can be grown than in control situations.16–18 Additionally, when adenoviral RUNX2 is immobilized on a scaffold, it is still capable of increasing osteogenic differentiation.19 One important consideration with relation to this study is that it has been shown that these two groups, the SOX trio and RUNX2, have inhibitory effects on each other when applied to the same cell population in vitro and in vivo.20, 21 In fact, it has been shown that SOX9 is dominant and inhibitory to RUNX2 in vivo.22 This work investigates the hypothesis that by spatially loading these two complexes to mimic native tissues, distinct zones of osteogenesis and chondrogenesis can be achieved without negative interactions.
Oligo[poly(ethylene glycol) fumarate] (OPF) is a promising hydrogel for use in tissue engineering, including for applications involving bone23 and cartilage24 tissue growth, and combined osteochondral repair.25–28 This hydrogel is formed by crosslinking a linear polyester formed by condensing poly(ethylene glycol) (PEG) and fumaryl chloride. The mesh size of the crosslinked hydrogel can be controlled by varying the molecular weight of the incorporated PEG chains, and higher concentrations of fumarate ester groups inside the polymer result in higher crosslinking and more points for degradation.29, 30 OPF has numerous favorable biological properties including being biocompatible, bioinert, and non-immunogenic, in addition to having inert degradation products.31
OPF is attractive as a delivery vehicle for a number of reasons. First, it is a synthetic rather than natural polymer and therefore offers tunable properties such as mesh size, degradation rate, and mechanical properties to allow for modification for a specific application. Moreover, it is attractive for use with plasmid DNA and other nucleotides because it can be crosslinked under physiologic conditions without harsh processing that could degrade or injure the genetic material.27 The inherent hydrophilicity of the system and hydrogel fabrication process results in an ability to entrap large concentrations of polymer/DNA complexes at relatively high loading efficiencies.26–28 Furthermore, OPF has been used to deliver DNA and bPEI in previous work by either direct loading of complexes or by loading complex laden gelatin particles.26–28 These studies have explored release of plasmid DNA from OPF hydrogels both in vitro and in vivo, as well as directly from the hydrogel or from an incorporated carrier. When incorporated directly into the hydrogel during fabrication, release of plasmid/polymer complexes was found to have a favorable profile lasting several weeks.28
In the present study, we hypothesize that the regeneration of bone and cartilage tissue will be enhanced by the delivery of bPEI-HA and DNA encoding for transcription factors in a porous OPF hydrogel scaffold. The inclusion of carboxymethyl cellulose (CMC) as a porogen in the OPF scaffold is also expected to increase tissue distribution inside of the scaffolds by generating voids inside the scaffold which cells can migrate into. Additionally, the dual delivery of RUNX2 and the SOX trio is expected to improve the quality and amount of bone and cartilage generation within the defects. To examine these hypotheses, composite scaffolds were implanted in a rat osteochondral defect for 6 weeks and analyzed using micro-CT and histology.
2. MATERIALS AND METHODS
2.1 In Vivo Experimental Design
Composite scaffolds consisting of OPF, CMC, and bPEI-HA/DNA complexes were examined in vivo for their ability to generate tissue in a rat knee osteochondral defect model. Groups for this study were designed to examine the interactions and efficacy of the use of DNA encoding for the transcription factors SOX 5, SOX 6, and SOX 9 (the SOX trio), and RUNX2 delivered with bPEI-HA. The groups examined here are summarized below in Table 1 and included a material control, RUNX2 DNA only, and SOX trio DNA only in order to identify the effects of each component individually as well as a combination group used to identify combinatory effects of RUNX2 and the SOX trio.
Table 1.
In vivo experimental groups for rat osteochondral defect implantation.
| Group | Top (Chondrogenic) Layer | Bottom (Osteogenic) Layer | Purpose |
|---|---|---|---|
| Empty | OPF only | OPF only | Scaffold control |
| SOX Trio | bPEI-HA/SOX trio complexes | OPF only | Elucidate the effects of SOX trio alone |
| RUNX2 | OPF only | bPEI-HA/RUNX2 complexes | Elucidate the effects of RUNX2 alone |
| Combined | bPEI-HA/SOX trio complexes | bPEI-HA/RUNX2 complexes | Show potential interactions between SOX trio and RUNX2 |
2.2 Assembly of bPEI-HA/DNA Complexes
Branched PEI-HA was synthesized as previously described,6, 9 using a reductive amination reaction to directly conjugate the hyaluronic acid fragments (6.4kDa) (LifeCore Biomedical, Chaska, MN) to the primary amines of the bPEI (Sigma-Aldrich, St. Louis, MO). The structure was verified with 1H NMR to ensure correct conjugation as has been described previously.6, 9
Plasmid DNA encoding for RUNX2, SOX5, SOX6, and SOX9 (Origene, Rockville, MD) was expanded using DNA expansion kits according to the manufacturer’s instructions (Qiagen, Venlo, Netherlands), collected, and used directly. For loading into hydrogels, bPEI-HA and DNA were combined drop wise in a constant 7.5:1 Nitrogen:Phosphate (N:P) ratio and allowed to complex in ultrapure (type 1) water (Super-Q Water Purification System, EMD Millipore, Billerica, MA) at room temperature for 30 min before use. After complexation, complexes were lyophilized for 48 hrs in preparation for use in hydrogel loading.
2.3 OPF Synthesis and Characterization
Synthesis of OPF was performed as previously described.29, 32, 33 Briefly, anhydrous dichloromethane (EMD, Billerica, MA) was obtained through refluxing in the presence of calcium hydride (Sigma Aldrich, St. Louis, MO) followed by distillation. Anhydrous PEG (Mn = 9.3 ± 0.1 kDa, Mw = 13.1 ± 0.1 kDa, n =3) (Sigma Aldrich, St. Louis, MO) was generated through distillation in toluene (Fisher Scientific, Waltham, MA) and then added to the anhydrous dichloromethane. Triethylamine (Sigma Aldrich, St. Louis, MO) and fumaryl chloride (Acros, Geel, Belgium) were added to this PEG solution drop wise and the reaction was incubated for 2 days. Purification was then performed and characterization of the product was performed through analysis with gel permeation chromatography using PEG standards and 1H NMR to verify correct structure and fumarate PEG ratios as previously described.29, 32, 33 The OPF used in this work has a Mn of 19.8± 0.3 kDa and a Mw of 89.9 ± 3.9 kDa (n = 3).
2.4 Composite Scaffold Fabrication
Scaffolds for use in the studies described below were composites consisting of an OPF hydrogel crosslinked around CMC particles (US Pharmacopeia Grade Lot #YD0567, Spectrum Chemicals and Products, Gardena, CA) up to 100 μm in diameter. For fabrication of the composites, OPF and poly(ethylene glycol)-diacrylate (PEGDA) (MW = 3,400 Da, Laysan Bio, Arab, AL) were first dissolved in PBS [2:1 OPF:PEG-DA weight ratio, 38% w/v gel in water]. Immediately before OPF crosslinking was initiated, dried DNA/bPEI-HA complexes were suspended in the OPF/PEGDA solution. For each layer, a DNA concentration of 6.2 mg/ml (after swelling) was maintained, which corresponds to levels previously shown to elicit a response in vivo.34 Once dispersed, ammonium persulfate (APS) and N,N,N′,N′-tetramethylethane-1,2-diamine (TEMED) were added to the solution [3.6 mM] and it was vigorously stirred for 30 s. At the end of these 30 s, CMC particles were added to the mixture and it was stirred for an additional 45 s before being added into custom poly(tetrafluoroethylene) (PTFE) molds. For bilayered scaffolds, the bottom osteogenic layer was allowed to crosslink for 3 min in a PTFE mold before addition of another mold directly on top was filled with crosslinking chondrogenic layer hydrogel. Once the two layers were stacked, they were allowed to finish crosslinking for 10 min prior to use.
Scaffolds for use in in vitro release and in vivo implantation were fabricated as 8 mm diameter by 1.2 mm thickness discs which were then punched with a custom punch such that the final scaffold punch dimensions were 1.2 mm in diameter by 1.2 mm thickness with each layer having a 0.6 mm thickness. These dimensions correspond to a 1.5 mm diameter by 1.5 mm thickness implant after swelling.
2.5 Surgical Procedure
32 Lewis rats (Charles River Laboratories, Wilmington, MA) 12 weeks of age for skeletal maturity were used in this study. All manipulations described followed protocols approved by the Institutional Animal Care and Use Committee at Rice University and published NIH guidelines for the care and use of laboratory animals have been observed. All implants were fabricated directly before implantation. Each composite scaffold contained CMC and OPF in an 1:1 weight ratio during fabrication as well as 8.4 μg DNA per scaffold in the combined scaffolds (4.2 μg DNA in either the RUNX2 or SOX trio only scaffolds). All animals survived the surgery and post-operative period with no signs of distress of infection for the duration of the study.
To complete the surgery, previously established procedures were followed to create defects in the right femoral condyle of the rat.35, 36 The surgical site was sterilely prepared and a lateral approach was used to expose the knee joint through blunt dissection. A lateral parapatellar incision was made and the patella dislocated medially to access the knee. The joint capsule was then opened and the defect was established along the midline and midway up the trochlear groove from the knee with a 0.9 mm drill bit and was created with a 1.5 mm drill bit. The defect was then flushed with saline and filled with the implant prepared as described above. The patella was then physically relocated and the wound closed in three layers. Animals were closely monitored for any signs of discomfort.
2.6 In Vivo Tissue Preparation
Six weeks after implantation, animals were anesthetized with 4–5% isofluorane and euthanized with carbon dioxide. A subsequent bilateral thoracotomy was performed to ensure death before the treated knees were extracted. The knees were then placed in 10% neutral buffered formalin for 1 week for fixation. Micro-CT scans were then performed on all samples before they were decalcified in 5% formic acid for 14 days and dried in an ethanol gradient from 50–100% (50, 70, 80, 90, 95, 100%) before staining. In vivo animal samples were analyzed for cartilage and bone generation through micro-CT and histology according to guidelines previously established for use in a bilayed osteochondral defect in a rabbit model.37–40
2.7 Micro-CT Imaging and Analysis
Before decalcification for histology was performed, micro-CT analysis was used to determine the mineralization as previously described.41 All images were taken with a high resolution SkyScan-1172 micro-CT imaging system. Samples were oriented so that the axis of the defect was vertical in the micro-CT chamber. The scanner resolution was set to 10.2 μm/pixel with a voltage of 100 KV and a current of 100 μA. The resolution of the camera was set to high (1280 × 1024 raw images). The defect was then reconstructed by backprojecting the raw images in a NRecon CT Reconstruction software package.
1.5 mm diameter, 1.5 mm height volumes of interest were then selected within the reconstructed images at the site of the defect for each sample. To calculate the percent bone formation within these volumes of interest, a lower binarization threshold of 45 and upper binarization threshold of 255 were set to identify bone morphology within the samples. With these values, percent bone formation within the samples was defined as the bone volume within the volume of interest divided by the total volume of interest expressed as a percentage.
2.8 Histology
For histological analysis, the dried samples were embedded in paraffin and sectioned in 6 μm slices. Sections from two locations within the defect (center and medial) were obtained and stained with hematoxylin and eosin (H&E) and Safranin O/Fast Green. Safranin O was selected for its ability to stain glycosaminoglycans (GAGs) and give an indication of the extent of chondrogenesis. 37, 39 Each of these sections was independently analyzed by three evaluators according to procedures described previously.37, 39 Briefly, the region of the defect corresponding to the native cartilage thickness was analyzed for cartilage tissue generation and the lower portion corresponding to native bone was analyzed for bone regeneration. Osteochondral repair was evaluated using 13 predetermined measures of regeneration which are listed in Table 2 and cover the quantity and quality of the regeneration of tissues, scaffold degradation, and integration of new tissues with surrounding native tissues. It is important to note that while the micro-CT percent bone formation was calculated digitally, all histological filling scores were completed blindly and independently by 3 investigators.
Table 2.
| Overall Defect Evaluation | |
| Percent Filling with New Tissue | |
| Percent | Score |
| 90–100% | 3 |
| 50–90% | 2 |
| 10–50% | 1 |
| 0–10% | 0 |
| Percent Implant Degradation | |
| 90–100% | 3 |
| 50–90% | 2 |
| 10–50% | 1 |
| 0–10% | 0 |
| Subchondral Tissue Evaluation | |
| Percent Filling with New Bone | |
| Percent | Score |
| 90–100% | 3 |
| 50–90% | 2 |
| 10–50% | 1 |
| 0–10% | 0 |
| Subchondral Tissue Morphology | |
| Mostly trabecular bone | 4 |
| Mostly compact bone | 3 |
| Mostly cartilage | 2 |
| Mostly fibrous tissue | 1 |
| Only fibrous tissue or no tissue | 0 |
| Extent of New Bone Bonding with Adjacent Bone | |
| Complete on both edges | 3 |
| Complete on one edge | 2 |
| Partial on both edges | 1 |
| No continuity on either edge | 0 |
| Chondral Tissue Evaluation | |
| Morphology of New Surface Tissue | |
| Exclusively Articular Cartilage | 4 |
| Mainly Hyaline Cartilage | 3 |
| Fibrocartilage (spherical morphology in >75% of cells) | 2 |
| Mostly Fibrous Tissue (spherical morphology in <75% cells) | 1 |
| No tissue | 0 |
| Morphology of New Cartilage | |
| Exclusively Articular Cartilage | 3 |
| Mainly Hyaline Cartilage | 2 |
| Mainly Fibrocartilage | 1 |
| Only Fibrous Tissue/ No Tissue | 0 |
| Thickness of New Cartilage | |
| Similar to surrounding cartilage | 3 |
| Greater than surrounding cartilage | 2 |
| Less than surrounding cartilage | 1 |
| No cartilage | 0 |
| Joint Surface Regularity | |
| Smooth, intact surface | 3 |
| Surface fissures (<25% new surface thickness) | 2 |
| Deep fissures (25–99% new surface thickness) | 1 |
| Complete disruption of the new surface | 0 |
| Chondrocyte Distribution | |
| Columnar | 3 |
| Mixed Columnar-clusters | 2 |
| Clusters | 1 |
| Individual or disorganized cells | 0 |
| Chondrocyte Cellularity | |
| Similar number of chondrocytes | 3 |
| More chondrocytes | 2 |
| Fewer chondrocytes | 1 |
| No Chondrocytes | 0 |
| Safranin O Staining | |
| Similar staining intensity | 4 |
| Stronger Staining intensity | 3 |
| Moderate staining intensity | 2 |
| Poor staining intensity | 1 |
| Little or no staining intensity | 0 |
| Chondrocyte and GAG Content of Adjacent Cartilage | |
| Normal cellularity with normal GAG content | 3 |
| Normal cellularity with moderate GAG content | 2 |
| Clearly less cells with poor GAG content | 1 |
| Few cells with little or no GAGs or no cartilage | 0 |
2.9 Statistics
Statistical analysis was performed on the data collected for in vitro release and micro-CT analysis using two-way ANOVA with a priori value of 0.05. Post hoc analysis was performed via Tukey-Kramer HSD to identify statistical significance (p<0.05) between each of the groups. Each of the in vivo histological scoring parameters was analyzed using the Kruskal Wallis test of variance and the Wilcoxon rank sum test for difference of groups with a significance level of p<0.05. All data are presented as means ± standard deviation, with the exception of histological results which presented the total number of each score attained for each group. The in vitro studies had an n of 4 while in vivo studies had a n of 8.
3. RESULTS
3.1 Gross Observation
During the tissue harvest, no signs of gross infection or any other adverse tissue responses were observed and all animals were ambulatory for the duration of the study.
3.2 Micro-CT
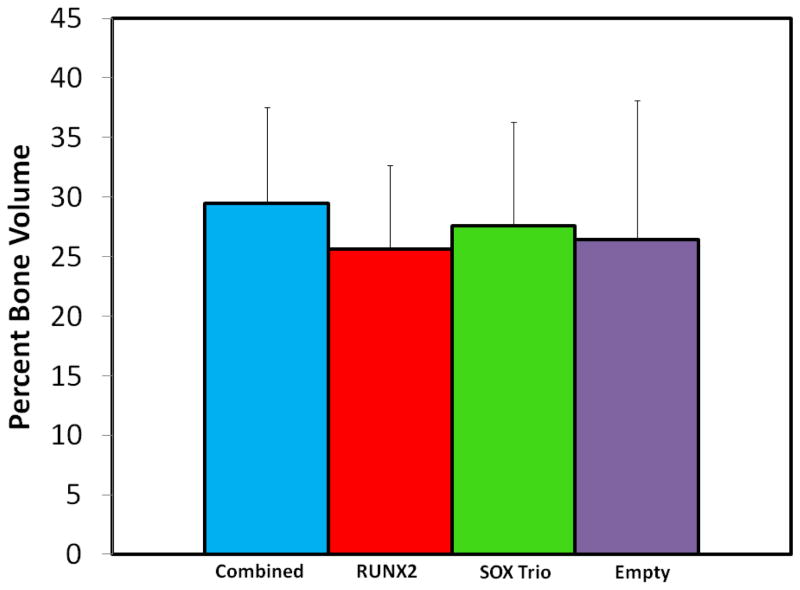
Micro-CT images were taken for each sample and analyzed to determine the percentage of bone present within the defect volume. There was no significant difference in percent regenerated bone volume between groups, as shown in Figure 1, but the analysis demonstrated measureable bone growth in all in vivo study groups with a high percentage formation of 29.5±8.0% found in the combination treatment group.
Figure 1.
Average percent bone generation identified through micro-CT. Error bars represent the standard deviation of each group for n=8.
3.3 Histology
Histological analysis was performed on the collected sections from the medial edge and the center of each defect as described above to determine the regeneration and tissue response within the created defects. Histology was selected as the main metric for tissue generation rather than gene expression due to the transient nature of expression of genes delivered with polymeric gene delivery vectors.42 The results for each parameter outlined in Table 2 are displayed in Figures 2, 3, and 4 and numerous statistical differences were observed. Representative histological results from the center of each group are included as in Figures 5, 6, 7, and 8.
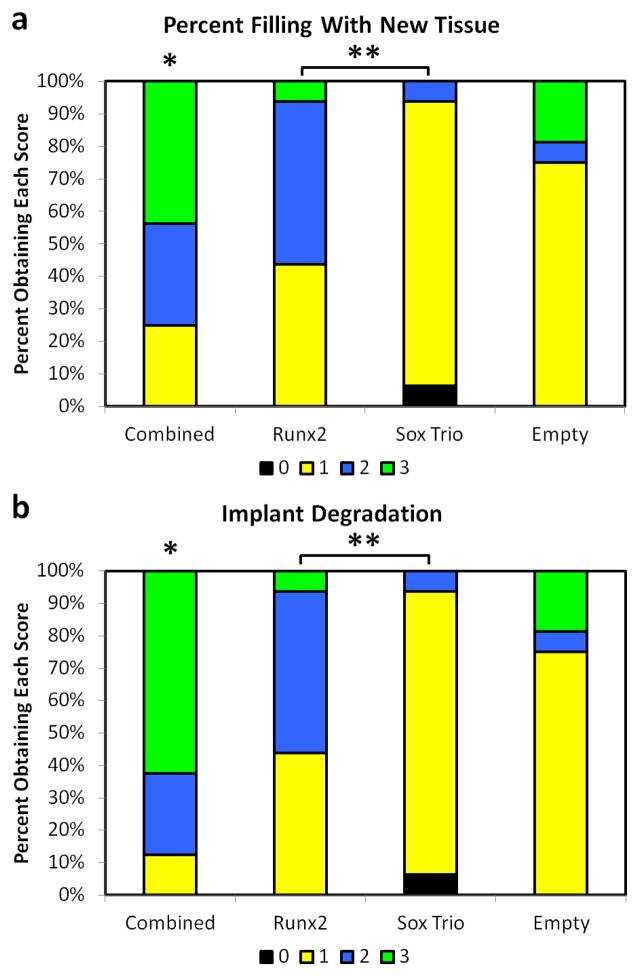
Figure 2.
Overall defect histological analysis where (a) corresponds to percent filling with new tissue score and (b) corresponds to percent implant degradation. 0, 1, 2, and 3 correspond to histology scores described in Table 2. Groups indicated by a “*” are statistically different from all other groups while groups connected by lines and a “**” are significantly different from each other.
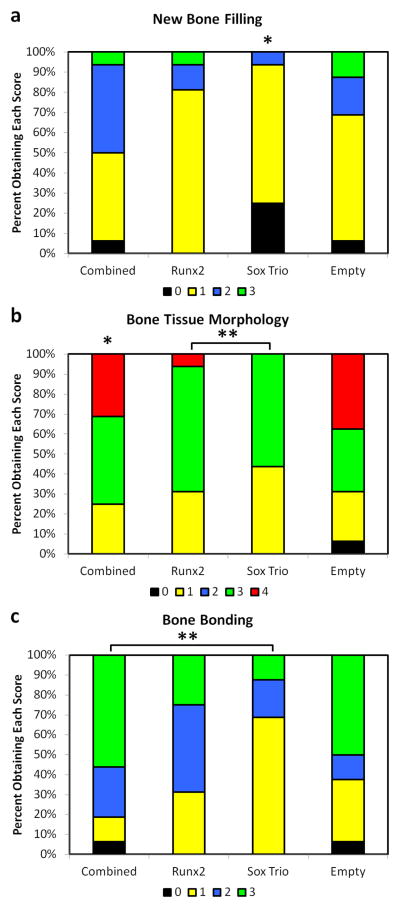
Figure 3.
Bone tissue histological analysis where (a) corresponds to filling with new bone, (b) corresponds to subchondral bone morphology, and (c) corresponds to bone bonding with surrounding tissue. 0, 1, 2, 3, and 4 correspond to histology scores described in Table 2. Groups indicated by a “*” are statistically different from all other groups while groups connected by lines and a “**” are significantly different from each other.
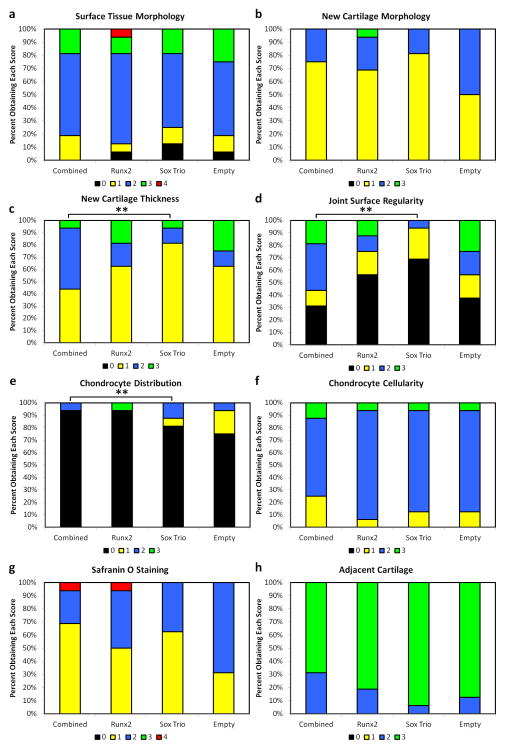
Figure 4.
Cartilage tissue histological analysis where (a) corresponds to the morphology of the new surface tissue, (b) is the morphology of the generated cartilage, (c) is the thickness of new cartilage, (d) is the joint surface regularity, (e) is the chondrocyte distribution, (f) is the chondrocyte cellularity, (g) is the safranin O staining, and (h) is the quality of the surrounding cartilage. 0, 1, 2, 3, and 4 correspond to histology scores described in Table 2. Groups indicated by a “*” are statistically different from all other groups while groups connected by lines and a “**” are significantly different from each other.
Figure 5.
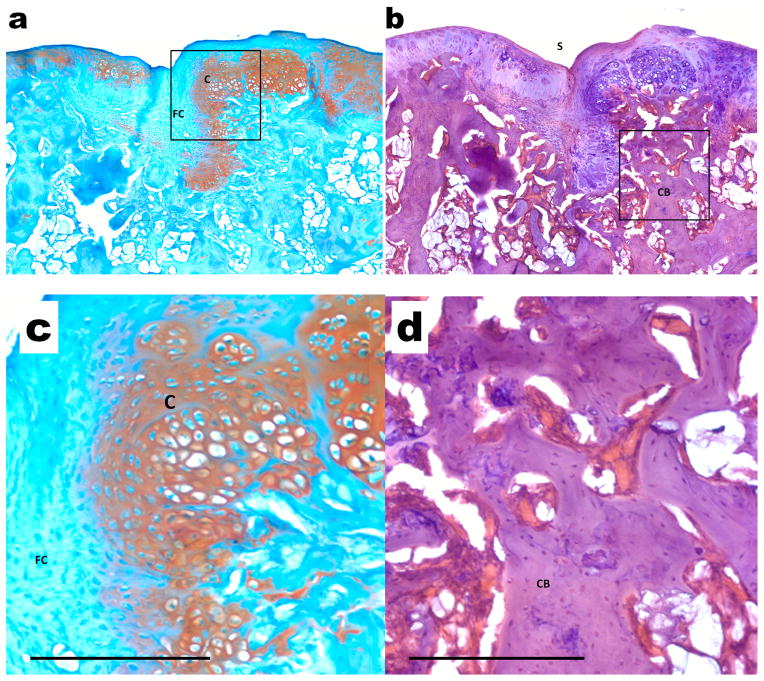
Representative histological sections of osteochondral tissue formation after 6 weeks of implantation of composite scaffolds in the Combined treatment group which had bPEI-HA and DNA encoding for the SOX trio in the top layer and RUNX2 in the bottom layer. Sections were stained with (a) Safranin-O/Fast Green and (b) hematoxylin and eosin (H&E). Scale bars = 500 μm. Magnified images indicate (c) a thick layer with a mixture of fibrocartilage (FC) and mostly hyaline cartilage (C) and (d) underlying regenerated subchondral cortical bone (CB) and near complete degradation of the composite scaffold. Boxed regions are shown in higher magnification with scale bars = 250 μm.
Figure 6.
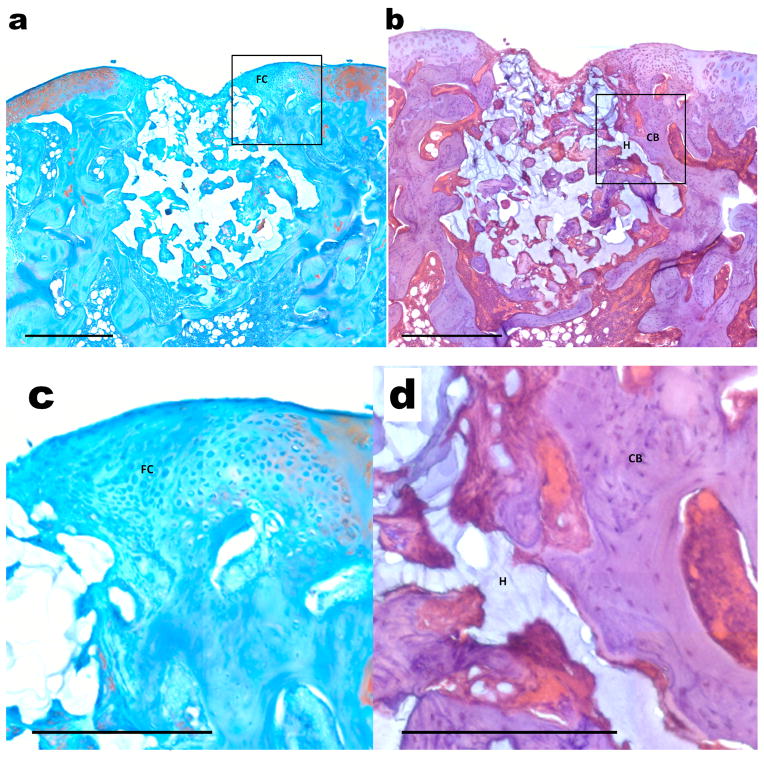
Representative histological sections of osteochondral tissue formation after 6 weeks of implantation of composite scaffolds in the RUNX2 treatment group which had bPEI-HA and DNA encoding for RUNX2 in the bottom layer. Sections were stained with (a) Safranin-O/Fast Green and (b) hematoxylin and eosin (H&E). Scale bars = 500 μm. Images with higher magnification indicate (b) regions of fibrocartilage (FC) with little safranin-o staining and (c) regions with remaining hydrogel (H) indicated by voids in contact with newly formed cortical bone tissue (CB).
Figure 7.
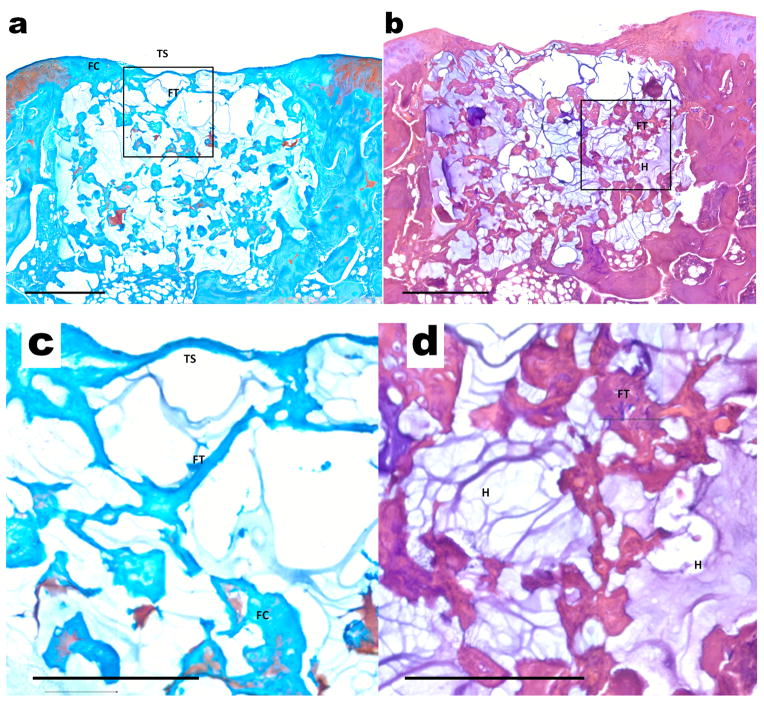
Representative histological sections of osteochondral tissue formation after 6 weeks of implantation of composite scaffolds in the SOX trio treatment group which had bPEI-HA and DNA encoding for the SOX trio in the top layer. Sections were stained with (a) Safranin-O/Fast Green and (b) hematoxylin and eosin (H&E). Scale bars = 500 μm. Images with higher magnification indicate (c) regions with very thin surface (TS), fibrous tissue (FT) and some little fibrocartilage with minor safranin-o staining (FC) and (d) subchondral regions with significant hydrogel remaining (H) and poor bone regeneration with fibrous tissue present (FT).
Figure 8.
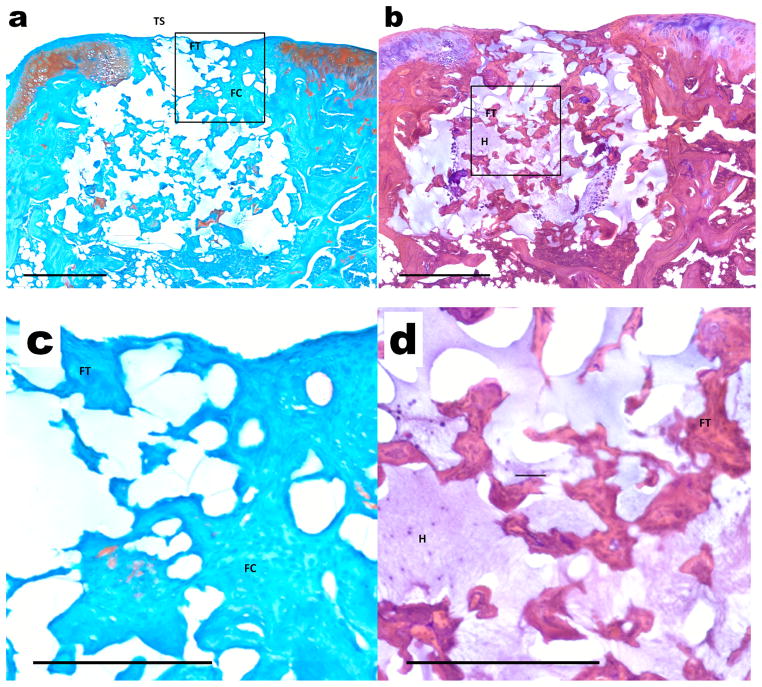
Representative histological sections of osteochondral tissue formation after 6 weeks of implantation of composite scaffolds in the Empty hydrogel treatment group which had no incorporated bPEI-HA/DNA complexes. Sections were stained with (a) Safranin-O/Fast Green and (b) hematoxylin and eosin (H&E). Scale bars = 500 μm. Images with higher magnification indicate regions with (c) a thin chondral layer (TS) made of fibrous tissue (FT) and some fibrocartilage (FC) and (d) regions within the subchondral bone layer with significant hydrogel remaining (H) with ingrowing fibrous tissue (FT).
In all of the groups a portion of the composite scaffold remained in the defect region. The groups with either an empty hydrogel or with only the SOX trio delivered had significantly more hydrogel present and significantly less new tissue growth into the defect as seen in Figures 2, 7, and 8. The combined treatment group had significantly higher tissue filling and less hydrogel present than all other groups as shown in Figures 2 and 5. Finally, the RUNX2 treatment group statistically fell between the best case combined treatment group and the more negative Empty and SOX trio only groups for tissue filling and implant degradation as shown in Figure 2.
For bone growth and quality, the SOX trio only group scored significantly lower in many cases. For percent filling with bone the SOX trio treatment group grew significantly less bone than all other test groups as demonstrated in Figures 3 and 7. For subchondral tissue morphology, the SOX trio scored significantly lower than all groups except for the blank hydrogels, signifying a poorer quality of the generated tissue. These results can be seen in the bone excerpts from the representative histological sections, Figures 5–8. While RUNX2 outperformed the SOX trio in bone quality and quantity, it was not significantly different from the empty gels and scored significantly worse than the combined group which was significantly better than all other test groups as seen in Figures 3a, 3b, and 5. These differences also are reflected in the example histological sections. Finally as seen in Figure 3c, there were fewer statistical differences in the bone bonding scoring, but the SOX trio did score significantly lower than the combined treatment group.
In the cartilage scoring portion, significant differences were found. For joint surface regularity and the thickness of the new cartilage shown in Figures 4c and 4d, the combined treatment group significantly outperformed the SOX trio group, signifying an improvement in the surface healing of the defect which is clearly seen in the representative histological sections in Figures 5 and 8. Conversely, for chondrocyte distribution the SOX trio group was found to have a significantly better distribution than the combined treatment group, shown in Figure 4e. Similar trends as were found for the bone histological scoring were present in the cartilage scoring categories which were listed in this paragraph, but did not achieve levels of significance.
4. DISCUSSION
Composite scaffolds were designed and fabricated to facilitate cellular infiltration and tissue ingrowth. OPF was selected due to its established ability to support regrowth of cartilage and bone while remaining non-toxic to target tissues immediately after implantation and during and after degradation. Additionally, by using a hydrogel, loading of high amounts of DNA in a spatially bilayered approach was possible while retaining the ability to release the genetic material and polymer delivery vector simultaneously. Because previous work has shown that implanting porous hydrogels into a defect can result in improved tissue ingrowth, this work utilized CMC as a porogen.43, 44 It was shown that in all groups a significant level of tissue ingrowth within the scaffold was achieved.
The primary aim of this study was to determine if DNA encoding for transcription factors delivered by bPEI-HA to a rat osteochondral defect via an OPF/CMC composite scaffold is capable of improving tissue generation in an in vivo situation. To achieve this goal, first, bilayered OPF/CMC hydrogels loaded with DNA encoding for eGFP complexed with bPEI-HA as representative complexes were fabricated and their release kinetics studied in vitro to characterize the system, as shown in the supplementary data. eGFP was selected as a representative gene for use in in vitro studies due to its similar size to the plasmids used in this study (4.7 kb vs. 4.7–5.8 kb for Runx2 and the Sox trio) and due to its use as an analog in the literature.9, 45 Once release was characterized, implants were fabricated incorporating bPEI-HA and DNA encoding for the SOX trio, DNA encoding for RUNX2, or a spatially controlled combination of both and implanted into a rat osteochondral defect for 6 weeks, a timepoint established in the literature. 46–48
The results presented above indicate that in this approach, the RUNX2 treatment group is more capable than the SOX trio treatment group at achieving the desired tissue growth of either bone or cartilage respectively. While the micro-CT analysis was unable to identify significant differences between groups, RUNX2 containing groups did significantly better in bone and general histological scoring than groups without RUNX2. This disparity between tests is interesting, but the data produced by the histological sections is superior due to its direct visualization of the morphology and integration of the generated tissue rather than detection of mineralization within a pre-set volume.
The SOX trio only group proved incapable of generating significant cartilage growth within the cartilage layer. It did, however score significantly better with respect to the distribution of the chondrocytes that did form, as seen in Figure 4e. This could indicate that the SOX trio is having an effect on the target cells. The most probable reason for this disparity in appropriate tissue generation is the inherent burst release associated with this system. Previous work has shown that subchondral bone growth is necessary to support cartilage growth.49 If it is assumed that the bioactive factors in this system are released quickly, the DNA encoding for the SOX trio attempted to grow cartilage in this situation without the mechanical support of subchondral bone. It is possible that in future work, if release of the cartilage treatment factors was delayed, that more complete cartilage healing could be observed.
It was found that combination therapy consisting of RUNX2 and SOX trio resulted in greater tissue filling, implant degradation, and bone generation and quality. Several previous studies have found that the combination of RUNX2 and the SOX trio results in negative bone and cartilage regrowth, and it has been shown that they can counteract each other’s action.20, 21 In fact, if they were transfecting the same cells, it would be expected from previous work that the SOX trio would be dominant.22 As for the lack of interaction, we hypothesize the spatial aspect of the loading excluded the infiltrated cells from being transfected with both transcription factors. Because the bPEI-HA/DNA complexes were entrapped in the OPF mesh network at the time of composite scaffold fabrication and implantation, the two DNA treatments were kept separate. Once implanted, the natural confines of the defect could limit diffusion of the complexes and the ability of RUNX2 and the SOX trio to interact.
One potential explanation for the faster implant degradation and tissue ingrowth in the combination treatment group could be the presence and amount of bPEI-HA in the defect. In combination treatment groups, more bPEI-HA was present because of the bilayered loading. OPF degrades via hydrolysis and the incorporation of an acid into the system accelerates the rate of this hydrolysis.50 The faster degradation of the hydrogel would allow more space for cellular infiltration and allow more cells to receive treatment with RUNX2 before the complexes were released and cleared from the defect. It is important to note here though that the degradation effects supplied by the bPEI-HA cannot be solely responsible for the tissue growth observed as the SOX trio and RUNX2 groups showed significant differences in tissue type and quality while incorporating identical amounts of bPEI-HA.
The primary limitation of the current study is its scope and additional studies could be considered to attempt to refine the general approach presented here. The most obvious approach would involve creating a different composite scaffold capable of delaying release from the chondrogenic layer until appropriate subchondral bone has been formed. With that said, the research presented here has shown that DNA encoding transcription factors delivered with bPEI-HA is capable of improving tissue healing in vivo. Additionally, spatial loading of two different treatments in a hydrogel scaffold is superior in this system to either treatment alone and improves tissue ingrowth and implant degradation.
5. CONCLUSIONS
The experiments described above show the potential of a bilayered approach in a rat osteochondral defect. Spatial design and loading of the described implants resulted in measurable improvements in overall tissue generation and quality, especially in the bone layer. This improvement in healing is especially true when a combination of the SOX trio and RUNX2 are used, though the inclusion of DNA encoding for RUNX2 also had a significant effect on tissue growth. The results here demonstrate the ability to apply polymeric gene therapy and transcription factors in vivo directly without implanting transfected or transduced cells and in spatial combination with each other in an osteochondral defect.
Supplementary Material
Acknowledgments
We acknowledge support from the National Institutes of Health (R21 AR56076 and R01 AR57083). SRS would like to acknowledge the Baylor College of Medicine Medical Scientist Training Program. Also, we would like to thank Dr. Limin Wang for his help in developing the OPF/CMC formulations.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errorsmaybe discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Trippel SB, Ghivizzani SC, Nixon AJ. Gene Ther. 2004;11(4):351–359. doi: 10.1038/sj.gt.3302201. [DOI] [PubMed] [Google Scholar]
- 2.Alford JW, Cole BJ. The American Journal of Sports Medicine. 2005;33(2):295–306. doi: 10.1177/0363546504273510. [DOI] [PubMed] [Google Scholar]
- 3.de Crombrugghe B, Lefebvre V, Nakashima K. Current Opinion in Cell Biology. 2001;13(6):721–728. doi: 10.1016/s0955-0674(00)00276-3. [DOI] [PubMed] [Google Scholar]
- 4.Ikeda Toshiyuki, Kamekura Satoru, Mabuchi Akihiko, Kou Ikuyo, Seki Shoji, Takato Tsuyoshi, Nakamura Kozo, Kawaguchi Hiroshi, Ikegawa Shiro, Chung Ung-il. Arthritis & Rheumatism. 2004;50(11):3561–3573. doi: 10.1002/art.20611. [DOI] [PubMed] [Google Scholar]
- 5.Ikeda T, Kamekura S, Mabuchi A, Kou I, Seki S, Takato T, Nakamura K, Kawaguchi H, Ikegawa S, Chung U-i. Arthritis & Rheumatism. 2004;50(11):3561–3573. doi: 10.1002/art.20611. [DOI] [PubMed] [Google Scholar]
- 6.Saraf A, Hacker MC, Sitharaman B, Grande-Allen KJ, Barry MA, Mikos AG. Biomacromolecules. 2008;9(3):818–827. doi: 10.1021/bm701146f. [DOI] [PubMed] [Google Scholar]
- 7.Ito T, Iida-Tanaka N, Niidome T, Kawano T, Kubo K, Yoshikawa K, Sato T, Yang Z, Koyama Y. Journal of Controlled Release. 2006;112(3):382–388. doi: 10.1016/j.jconrel.2006.03.013. [DOI] [PubMed] [Google Scholar]
- 8.Ito T, Yoshihara C, Hamada K, Koyama Y. Biomaterials. 2010;31(10):2912–2918. doi: 10.1016/j.biomaterials.2009.12.032. [DOI] [PubMed] [Google Scholar]
- 9.Needham CJ, Williams AK, Chew SA, Kasper FK, Mikos AG. Biomacromolecules. 2012;13(5):1429–1437. doi: 10.1021/bm300145q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lefebvre V, Smits P. Birth Defects Research Part C: Embryo Today: Reviews. 2005;75(3):200–212. doi: 10.1002/bdrc.20048. [DOI] [PubMed] [Google Scholar]
- 11.Yang HN, Park JS, Woo DG, Jeon SY, Do HJ, Lim HY, Kim SW, Kim JH, Park KH. Biomaterials. 2011;32(30):7695–7704. doi: 10.1016/j.biomaterials.2011.06.059. [DOI] [PubMed] [Google Scholar]
- 12.Lee JM, Im GI. Biomaterials. 2012;33(7):2016–2024. doi: 10.1016/j.biomaterials.2011.11.050. [DOI] [PubMed] [Google Scholar]
- 13.Im GI, Kim HJ, Lee JH. Biomaterials. 2011;32(19):4385–4392. doi: 10.1016/j.biomaterials.2011.02.054. [DOI] [PubMed] [Google Scholar]
- 14.Park JS, Yang HN, Woo DG, Jeon SY, Do HJ, Lim HY, Kim JH, Park KH. Biomaterials. 2011;32(14):3679–3688. doi: 10.1016/j.biomaterials.2011.01.063. [DOI] [PubMed] [Google Scholar]
- 15.Komori T. Frontiers in Bioscience : A Journal and Virtual Library. 2008;13:898–903. doi: 10.2741/2730. [DOI] [PubMed] [Google Scholar]
- 16.Yang S, Wei D, Wang D, Phimphilai M, Krebsbach PH, Franceschi RT. Journal of Bone and Mineral Research. 2003;18(4):705–715. doi: 10.1359/jbmr.2003.18.4.705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhao Z, Zhao M, Xiao G, Franceschi RT. Molecular Therapy. 2005;12(2):247–253. doi: 10.1016/j.ymthe.2005.03.009. [DOI] [PubMed] [Google Scholar]
- 18.Byers BA, Guldberg RE, García AsJ. Tissue Engineering. 2004;10(11–12):1757–1766. doi: 10.1089/ten.2004.10.1757. [DOI] [PubMed] [Google Scholar]
- 19.Zhang Y, Deng X, Scheller EL, Kwon TG, Lahann J, Franceschi RT, Krebsbach PH. Biomaterials. 2010;31(12):3231–3236. doi: 10.1016/j.biomaterials.2010.01.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cao L, Yang F, Liu G, Yu D, Li H, Fan Q, Gan Y, Tang T, Dai K. Biomaterials. 2011;32(16):3910–3920. doi: 10.1016/j.biomaterials.2011.02.014. [DOI] [PubMed] [Google Scholar]
- 21.Jeon SY, Park JS, Yang HN, Woo DG, Park KH. Biomaterials. 2012;33(17):4413–4423. doi: 10.1016/j.biomaterials.2012.02.051. [DOI] [PubMed] [Google Scholar]
- 22.Zhou G, Zheng Q, Engin F, Munivez E, Chen Y, Sebald E, Krakow D, Lee B. Proceedings of the National Academy of Sciences. 2006;103(50):19004–19009. doi: 10.1073/pnas.0605170103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Temenoff JS, Park H, Jabbari E, Conway DE, Sheffield TL, Ambrose CG, Mikos AG. Biomacromolecules. 2003;5(1):5–10. doi: 10.1021/bm030067p. [DOI] [PubMed] [Google Scholar]
- 24.Park H, Temenoff JS, Tabata Y, Caplan AI, Mikos AG. Biomaterials. 2007;28(21):3217–3227. doi: 10.1016/j.biomaterials.2007.03.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hui J, Ren X, Afizah M, Chian K, Mikos A. Clinical Orthopaedics and Related Research. 2013;471(4):1174–1185. doi: 10.1007/s11999-012-2487-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kasper FK, Jerkins E, Tanahashi K, Barry MA, Tabata Y, Mikos AG. J Biomed Mater Res Part A. 2006;78A(4):823–835. doi: 10.1002/jbm.a.30736. [DOI] [PubMed] [Google Scholar]
- 27.Kasper FK, Kushibiki T, Kimura Y, Mikos AG, Tabata Y. Journal of Controlled Release. 2005;107(3):547–561. doi: 10.1016/j.jconrel.2005.07.005. [DOI] [PubMed] [Google Scholar]
- 28.Kasper FK, Seidlits SK, Tang A, Crowther RS, Carney DH, Barry MA, Mikos AG. Journal of Controlled Release. 2005;104(3):521–539. doi: 10.1016/j.jconrel.2005.03.008. [DOI] [PubMed] [Google Scholar]
- 29.Kinard LA, Kasper FK, Mikos AG. Nature protocols. 2012;7(6):1219–1227. doi: 10.1038/nprot.2012.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Temenoff JS, Athanasiou KA, Lebaron RG, Mikos AG. Journal of Biomedical Materials Research. 2002;59(3):429–437. doi: 10.1002/jbm.1259. [DOI] [PubMed] [Google Scholar]
- 31.Shin H, Temenoff JS, Mikos AG. Biomacromolecules. 2003;4(3):552–560. doi: 10.1021/bm020121m. [DOI] [PubMed] [Google Scholar]
- 32.Jo S, Shin H, Shung AK, Fisher JP, Mikos AG. Macromolecules. 2001;34(9):2839–2844. [Google Scholar]
- 33.Kinard L, Chu C-Y, Tabata Y, Kasper FK, Mikos A. Pharmaceutical Research. 30(9):2332–2343. doi: 10.1007/s11095-013-1077-5. [DOI] [PubMed] [Google Scholar]
- 34.Chew SA, Kretlow JD, Spicer PP, Edwards AW, Baggett LS, Tabata Y, Kasper FK, Mikos AG. Tissue Eng Part A. 2010;17(5–6):751–763. doi: 10.1089/ten.tea.2010.0496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Coburn JM, Gibson M, Monagle S, Patterson Z, Elisseeff JH. Proceedings of the National Academy of Sciences. 2012;109(25):10012–10017. doi: 10.1073/pnas.1121605109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hwang NS, Varghese S, Lee HJ, Zhang Z, Ye Z, Bae J, Cheng L, Elisseeff J. Proceedings of the National Academy of Sciences. 2008;105(52):20641–20646. doi: 10.1073/pnas.0809680106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Guo X, Park H, Young S, Kretlow JD, van den Beucken JJ, Baggett LS, Tabata Y, Kasper FK, Mikos AG, Jansen JA. Acta Biomaterialia. 2010;6(1):39–47. doi: 10.1016/j.actbio.2009.07.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Holland TA, Bodde EWH, Baggett LS, Tabata Y, Mikos AG, Jansen JA. J Biomed Mater Res Part A. 2005;75A(1):156–167. doi: 10.1002/jbm.a.30379. [DOI] [PubMed] [Google Scholar]
- 39.Holland TA, Bodde EWH, Cuijpers VMJI, Baggett LS, Tabata Y, Mikos AG, Jansen JA. Osteoarthritis and Cartilage. 2007;15(2):187–197. doi: 10.1016/j.joca.2006.07.006. [DOI] [PubMed] [Google Scholar]
- 40.Kim K, Lam J, Lu S, Spicer PP, Lueckgen A, Tabata Y, Wong ME, Jansen JA, Mikos AG, Kasper FK. Journal of Controlled Release. 2013;168(2):166–178. doi: 10.1016/j.jconrel.2013.03.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Young S, Patel ZS, Kretlow JD, Murphy MB, Mountziaris PM, Baggett LS, Ueda H, Tabata Y, Jansen JA, Wong M, Mikos AG. Tissue engineering Part A. 2009;15(9):2347–2362. doi: 10.1089/ten.tea.2008.0510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Godbey WT, Wu KK, Mikos AG. Journal of Controlled Release. 1999;60(2–3):149–160. doi: 10.1016/s0168-3659(99)00090-5. [DOI] [PubMed] [Google Scholar]
- 43.Stachowiak AN, Bershteyn A, Tzatzalos E, Irvine DJ. Advanced Materials. 2005;17(4):399–403. [Google Scholar]
- 44.Loebsack A, Greene K, Wyatt S, Culberson C, Austin C, Beiler R, Roland W, Eiselt P, Rowley J, Burg K. Journal of Biomedical Materials Research. 2001;57(4):575–581. doi: 10.1002/1097-4636(20011215)57:4<575::aid-jbm1204>3.0.co;2-9. [DOI] [PubMed] [Google Scholar]
- 45.Chew SA, Hacker MC, Saraf A, Raphael RM, Kasper FK, Mikos AG. Biomacromolecules. 2010;11(3):600–609. doi: 10.1021/bm901147k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Fan H, Hu Y, Qin L, Li X, Wu H, Lv R. J Biomed Mater Res Part A. 2006;77A(4):785–794. doi: 10.1002/jbm.a.30647. [DOI] [PubMed] [Google Scholar]
- 47.Lee KH, Song SU, Hwang TS, Yi Y, Oh IS, Lee JY, Choi KB, Choi MS, Kim SJ. Human gene therapy. 2001;12(14):1805–1813. doi: 10.1089/104303401750476294. [DOI] [PubMed] [Google Scholar]
- 48.Rutherford RB, Moalli M, Franceschi RT, Wang D, Gu K, Krebsbach PH. Tissue Engineering. 2002;8(3):441–452. doi: 10.1089/107632702760184709. [DOI] [PubMed] [Google Scholar]
- 49.Qiu YS, Shahgaldi BF, Revell WJ, Heatley FW. Osteoarthritis and Cartilage. 2003;11(11):810–820. doi: 10.1016/s1063-4584(03)00164-x. [DOI] [PubMed] [Google Scholar]
- 50.Larson RA, Weber EJ. Reaction Mechanisms in Environmental Organic Chemistry. CRC press; 1994. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.