Abstract
We sequenced the genomes of a ~7,000 year old farmer from Germany and eight ~8,000 year old hunter-gatherers from Luxembourg and Sweden. We analyzed these and other ancient genomes1–4 with 2,345 contemporary humans to show that most present Europeans derive from at least three highly differentiated populations: West European Hunter-Gatherers (WHG), who contributed ancestry to all Europeans but not to Near Easterners; Ancient North Eurasians (ANE) related to Upper Paleolithic Siberians3, who contributed to both Europeans and Near Easterners; and Early European Farmers (EEF), who were mainly of Near Eastern origin but also harbored WHG-related ancestry. We model these populations’ deep relationships and show that EEF had ~44% ancestry from a “Basal Eurasian” population that split prior to the diversification of other non-African lineages.
Near Eastern migrants played a major role in the introduction of agriculture to Europe, as ancient DNA indicates that early European farmers were distinct from European hunter-gatherers4,5 and close to present-day Near Easterners4,6. However, modelling present-day Europeans as a mixture of these two ancestral populations4 does not account for the fact that they are also admixed with a population related to Native Americans7,8. To clarify the prehistory of Europe, we sequenced nine ancient genomes (Fig. 1A; Extended Data Fig. 1): “Stuttgart” (19-fold coverage), a ~7,000 year old skeleton found in Germany in the context of artifacts from the first widespread farming culture of central Europe, the Linearbandkeramik; “Loschbour” (22-fold), an ~8,000 year old skeleton from the Loschbour rock shelter in Luxembourg, discovered in the context of hunter-gatherer artifacts (SI1; SI2); and seven ~8,000 year old samples (0.01–2.4-fold) from a hunter-gatherer burial in Motala, Sweden (the highest coverage individual was “Motala12”).
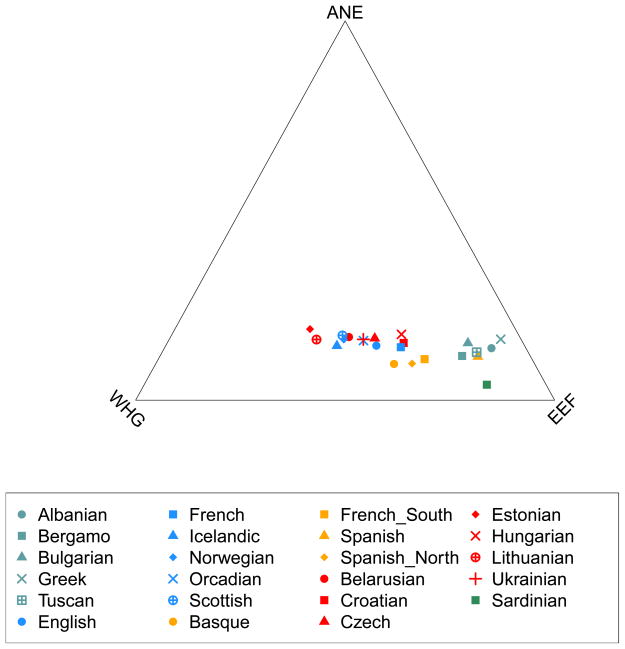
Figure 1. Map of West Eurasian populations and Principal Component Analysis.
(a) Geographical locations of analyzed samples, with color coding matching the PCA. We show all sampling locations for each population, which results in multiple points for some (e.g., Spain). (b) PCA on all present-day West Eurasians, with ancient and selected eastern non-African samples projected. European hunter-gatherers fall beyond present-day Europeans in the direction of European differentiation from the Near East. Stuttgart clusters with other Neolithic Europeans and present-day Sardinians. MA1 falls outside the variation of present-day West Eurasians in the direction of southern-northern differentiation along dimension 2.
Sequence reads from all samples revealed >20% C→T and G→A deamination-derived mismatches at the ends of the molecules that are characteristic of ancient DNA9,10 (SI3). We estimate nuclear contamination rates to be 0.3% for Stuttgart and 0.4% for Loschbour (SI3), and mitochondrial (mtDNA) contamination rates to be 0.3% for Stuttgart, 0.4% for Loschbour, and 0.01–5% for the Motala individuals (SI3). Stuttgart has mtDNA haplogroup T2, typical of Neolithic Europeans11, and Loschbour and all Motala individuals have the U5 or U2 haplogroups, typical of hunter-gatherers5,9 (SI4). Stuttgart is female, while Loschbour and five Motala individuals are male (SI5) and belong to Y-chromosome haplogroup I, suggesting that this was common in pre-agricultural Europeans (SI5).
We carried out large-scale sequencing of libraries prepared with uracil DNA glycosylase (UDG), which removes deaminated cytosines, thus reducing errors arising from ancient DNA damage (SI3). The ancient individuals had indistinguishable levels of Neanderthal ancestry when compared to each other (~2%) and to present-day Eurasians (SI6). The heterozygosity of Stuttgart (0.00074) is at the high end of present-day Europeans, while that of Loschbour (0.00048) is lower than in any present humans (SI2), reflecting a strong bottleneck in Loschbour’s ancestors as the genetic data show that he was not recently inbred (Extended Data Fig. 2). High copy numbers for the salivary amylase gene (AMY1) have been associated with a high starch diet12; our data are consistent with this finding in that the ancient hunter gatherers La Braña (from Iberia)2, Motala12, and Loschbour had 5, 6 and 13 copies respectively, whereas the Stuttgart farmer had 16 (SI7). Both Loschbour and Stuttgart had dark hair (>99% probability); and Loschbour, like La Braña and Motala12, likely had blue or intermediate-colored eyes (>75%) while Stuttgart likely had brown eyes (>99%) (SI8). Neither Loschbour nor La Braña carries the skin-lightening allele in SLC24A5 that is homozygous in Stuttgart and nearly fixed in Europeans today2, but Motala12 carries at least one copy of the derived allele, showing that this allele was present in Europe prior to the advent of agriculture.
We compared the ancient genomes to 2,345 present-day humans from 203 populations genotyped at 594,924 autosomal single nucleotide polymorphisms (SNPs) with the Human Origins array8 (SI9) (Extended Data Table 1). We used ADMIXTURE13 to identify 59 “West Eurasian” populations that cluster with Europe and the Near East (SI9 and Extended Data Fig. 3). Principal component analysis (PCA)14 (SI10) (Fig. 1B) indicates a discontinuity between the Near East and Europe, with each showing north-south clines bridged only by a few populations of mainly Mediterranean origin. We projected15 the newly sequenced and previously published1–4 ancient genomes onto the first two principal components (PCs) (Fig. 1B). Upper Paleolithic hunter-gatherers3 from Siberia like the MA1 (Mal’ta) individual project at the northern end of the PCA, suggesting an “Ancient North Eurasian” meta-population (ANE). European hunter-gatherers from Spain2, Luxembourg, and Sweden4 fall beyond present-day Europeans in the direction of European differentiation from the Near East, and form a “West European Hunter-Gatherer” (WHG) cluster including Loschbour and La Braña2, and a “Scandinavian Hunter-Gatherer” (SHG) cluster including the Motala individuals and ~5,000 year old hunter-gatherers from the Pitted Ware Culture4. An “Early European Farmer” (EEF) cluster includes Stuttgart, the ~5,300 year old Tyrolean Iceman1 and a ~5,000 year old Swedish farmer4.
Patterns observed in PCA may be affected by sample composition (SI10) and their interpretation in terms of admixture events is not straightforward, so we rely on formal analysis of f-statistics8 to document mixture of at least three source populations in the ancestry of present Europeans. We began by computing all possible statistics of the form f3(Test; Ref1, Ref2) (SI11), which if significantly negative show unambiguously8 that Test is admixed between populations anciently related to Ref1 and Ref2 (we choose Ref1 and Ref2 from 5 ancient and 192 present populations). The lowest f3-statistics for Europeans are negative (93% are >4 standard errors below 0), with most showing strong support for at least one ancient individual being one of the references (SI11). Europeans almost always have their lowest f3 with either (EEF, ANE) or (WHG, Near East) (SI11, Table 1, Extended Data Table 1), which would not be expected if there were just two ancient sources of ancestry (in which case the best references for all Europeans would be similar). The lowest f3-statistic for Near Easterners always takes Stuttgart as one of the reference populations, consistent with a Near Eastern origin for Stuttgart’s ancestors (Table 1). We also computed the statistic f4(Test, Stuttgart; MA1, Chimp), which measures whether MA1 shares more alleles with a Test population or with Stuttgart. This statistic is significantly positive (Extended Data Fig. 4, Extended Data Table 1) if Test is nearly any present-day West Eurasian population, showing that MA1-related ancestry has increased since the time of early farmers like Stuttgart (the analogous statistic using Native Americans instead of MA1 is correlated but smaller in magnitude (Extended Data Fig. 5), indicating that MA1 is a better surrogate than the Native Americans who were first used to document ANE ancestry in Europe7,8). The analogous statistic f4(Test, Stuttgart; Loschbour, Chimp) is nearly always positive in Europeans and negative in Near Easterners, indicating that Europeans have more ancestry from populations related to Loschbour than do Near Easterners (Extended Data Fig. 4, Extended Data Table 1). Extended Data Table 2 documents the robustness of key f4-statistics by recomputing them using transversion polymorphisms not affected by ancient DNA damage, and also using whole-genome sequencing data not affected by SNP ascertainment bias. Extended Data Fig. 6 shows the geographic gradients in the degree of allele sharing of present-day West Eurasians (as measured by f4-statistics) with Stuttgart (EEF), Loschbour (WHG) and MA1 (ANE).
Table 1.
Lowest f3-statistics for each West Eurasian population
| Ref1 | Ref2 | Target for which these two references give the lowest f3(X; Ref1, Ref2) |
|---|---|---|
| WHG | EEF | Sardinian*** |
| WHG | Near East | Basque, Belarusian, Czech, English, Estonian, Finnish, French_South, Icelandic, Lithuanian, Mordovian, Norwegian, Orcadian, Scottish, Spanish, Spanish_North, Ukrainian |
| WHG | Siberian | Russian |
| EEF | ANE | Abkhasian***, Albanian, Ashkenazi_Jew****, Bergamo, Bulgarian, Chechen****, Croatian, Cypriot****, Druze**, French, Greek, Hungarian, Lezgin, Maltese, Sicilian, Turkish_Jew, Tuscan |
| EEF | Native American | Adygei, Balkar, Iranian, Kumyk, North_Ossetian, Turkish |
| EEF | African | BedouinA, BedouinB†, Jordanian, Lebanese, Libyan_Jew, Moroccan_Jew, Palestinian, Saudi****, Syrian, Tunisian_Jew***, Yemenite_Jew*** |
| EEF | South Asian | Armenian, Georgian****, Georgian_Jew*, Iranian_Jew***, Iraqi_Jew*** |
Note: WHG = Loschbour or LaBraña; EEF=Stuttgart; ANE=MA1; Native American=Piapoco; African=Esan, Gambian, or Kgalagadi; South Asian=GujaratiC or Vishwabrahmin. Statistics are negative with Z<-4 unless otherwise noted: † (positive) or *, **, ***, ****, to indicate Z less than 0, −1, −2, and −3 respectively. The complete list of statistics can be found in Extended Data Table 1.
To determine the minimum number of source populations needed to explain the data for many European populations taken together, we studied the matrix of all possible statistics of the form f4(Testbase, Testi; Obase, Oj) (SI12). Testbase is a reference European population, Testi is the set of all other European Test populations, Obase is a reference outgroup, and Oj is the set of other outgroups (ancient DNA samples, Onge, Karitiana, and Mbuti). The rank of the (i, j) matrix reflects the minimum number of sources that contributed to the Test populations16,17. For a pool of individuals from 23 Test populations representing most present-day European groups, this analysis rejects descent from just two sources (P<10−12 by a Hotelling T-test17). However, three source populations are consistent with the data after excluding the Spanish who have evidence for African admixture18–20 (P=0.019, not significant after multiple-hypothesis correction), consistent with the results from ADMIXTURE (SI9), PCA (Fig. 1B, SI10) and f-statistics (Extended Data Table 1, Extended Data Fig. 6, SI11, SI12). We caution that the finding of three sources could be consistent with a larger number of mixture events. Moreover, the source populations may themselves have been mixed. Indeed, the positive f4(Stuttgart, Test; Loschbour, Chimp) statistics obtained when Test is Near Eastern (Extended Data Table 1) imply that the EEF had some WHG-related ancestry, which was greater than 0% and as high as 45% (SI13).
We used the ADMIXTUREGRAPH software8,15 to fit a model (a tree structure augmented by admixture events) to the data, exploring models relating the three ancient populations (Stuttgart, Loschbour, and MA1) to two eastern non-Africans (Onge and Karitiana) and sub-Saharan Africans (Mbuti). We found no models that fit the data with 0 or 1 admixture events, but did find a model that fit with 2 admixture events (SI14). The successful model (Fig. 2A) confirms the existence of MA1-related admixture in Native Americans3, but includes the novel inference that Stuttgart is partially (44 ± 10%) derived from a lineage that split prior to the separation of eastern non-Africans from the common ancestor of WHG and ANE. The existence of such “Basal Eurasian” admixture into Stuttgart provides a simple explanation for our finding that diverse eastern non-African populations share significantly more alleles with ancient European and Upper Paleolithic Siberian hunter-gatherers than with Stuttgart (that is, f4(Eastern non-African, Chimp; Hunter-gatherer, Stuttgart) is significantly positive), but that hunter-gatherers appear to be equally related to most eastern groups (SI14). We verified the robustness of the model by reanalyzing the data using the unsupervised MixMapper7 (SI15) and TreeMix21 software (SI16), which both identified the same admixture events. The ANE/WHG split must have occurred >24,000 years ago (as it must predate the age of MA13), and the WHG/Eastern non-African split must have occurred >40,000 years ago (as it must predate the Tianyuan22 individual from China which clusters with Asians to the exclusion of Europeans). The Basal Eurasian split must be even older, and might be related to early settlement of the Levant23 or Arabia24,25 prior to the diversification of most Eurasians, or more recent gene flow from Africa26. However, the Basal Eurasian population shares much of the genetic drift common to non-African populations after their separation from Africans, and thus does not appear to represent gene flow between sub-Saharan Africans and the ancestors of non-Africans after the out-of-Africa bottleneck (SI14).
Figure 2. Modeling of West Eurasian population history.
(a) A three-way mixture model that is a fit to the data for many populations. Present-day samples are colored in blue, ancient in red, and reconstructed ancestral populations in green. Solid lines represent descent without mixture, and dashed lines represent admixture. We print mixture proportions and one standard error for the two mixtures relating the highly divergent ancestral populations. (We do not print the estimate for the “European” population as it varies depending on the population). (b) We plot the proportions of ancestry from each of three inferred ancestral populations (EEF, ANE and WHG).
Fitting present-day Europeans into the model, we find that few populations can be fit as 2-way mixtures, but nearly all are compatible with 3-way mixtures of ANE/EEF/WHG (SI14). The mixture proportions from the fitted model (Fig. 2B; Extended Data Table 3) are encouragingly consistent with those obtained from a separate method that relates European populations to diverse outgroups using f4-statistics, assuming only that MA1 is an unmixed descendent of ANE, Loschbour of WHG, and Stuttgart of EEF (SI17). We infer that EEF ancestry in Europe today ranges from ~30% in the Baltic region to ~90% in the Mediterranean, consistent with patterns of identity-by-descent (IBD) sharing27,28 (SI18) and shared haplotype analysis (chromosome painting)29 (SI19) in which Loschbour shares more segments with northern Europeans and Stuttgart with southern Europeans. Southern Europeans inherited their European hunter-gatherer ancestry mostly via EEF ancestors (Extended Data Fig. 6), while Northern Europeans acquired up to 50% of WHG ancestry above and beyond the WHG-related ancestry which they received through their EEF ancestors. Europeans have a larger proportion of WHG than ANE ancestry in general. By contrast, in the Near East there is no detectable WHG ancestry, but up to ~29% ANE in the Caucasus (SI14). A striking feature of these findings is that ANE ancestry is inferred to be present in nearly all Europeans today (with a maximum of ~20%), but was absent in both farmers and hunter-gatherers from central/western Europe during the Neolithic transition. At the same time, we infer that ANE ancestry was not completely absent from the larger European region at that time: we find that it was present in ~8,000 years old Scandinavian hunter-gatherers, since MA1 shares more alleles with Motala12 (SHG) than with Loschbour, and Motala12 fits as a mixture of 81% WHG and 19% ANE (SI14).
Two sets of European populations are poor fits for the model. Sicilians, Maltese, and Ashkenazi Jews have EEF estimates of >100% consistent with their having more Near Eastern ancestry than can be explained via EEF admixture (SI17). They also cannot be jointly fit with other Europeans (SI14), and they fall in the gap between European and Near Easterners (Fig. 1B). Finns, Mordovians and Russians (from the northwest of Russia) also do not fit (SI14; Extended Data Table 3) due to East Eurasian gene flow into the ancestors of these northeastern European populations. These populations (and Chuvash and Saami) are more related to East Asians than can be explained by ANE admixture (Extended Data Fig. 7), likely reflecting a separate stream of Siberian gene flow into northeastern Europe (SI14).
Several questions will be important to address in future ancient DNA work. Where and when did the Near Eastern farmers admix with European hunter-gatherers to produce the EEF? How did the ancestors of present-day Europeans first acquire their ANE ancestry? Discontinuity in central Europe during the late Neolithic (~4,500 years ago) associated with the appearance of mtDNA types absent in earlier farmers and hunter-gatherers30 raises the possibility that ANE ancestry may have also appeared at this time. Finally, it is important to study ancient genome sequences from the Near East to provide insights into the history of the Basal Eurasians.
Online Methods
Archeological context, sampling and DNA extraction
The Loschbour sample stems from a male skeleton excavated in 1935 at the Loschbour rock shelter in Heffingen, Luxembourg. The skeleton was AMS radiocarbon dated to 7,205 ± 50 years before present (OxA-7738; 6,220-5,990 cal BC)31. At the Palaeogenetics Laboratory in Mainz, material for DNA extraction was sampled from tooth 16 (an upper right M1 molar) after irradiation with UV-light, surface removal, and pulverization in a mixer mill. DNA extraction took place in the palaeogenetics facilities in the Institute for Archaeological Sciences at the University of Tübingen. Three extracts were made in total, one from 80 mg of powder using an established silica based protocol32 and two additional extracts from 90 mg of powder each with a protocol optimized for the recovery of short DNA molecules33.
The Stuttgart sample was taken from a female skeleton excavated in 1982 at the site Viesenhäuser Hof, Stuttgart-Mühlhausen, Germany. It was attributed to the Linearbandkeramik (5,500-4,800 BC) through associated pottery artifacts and the chronology was corroborated by radiocarbon dating of the stratigraphy34. Both sampling and DNA extraction took place in the Institute for Archaeological Sciences at the University of Tübingen. Tooth 47 (a lower right M2 molar) was removed and material from the inner part was sampled with a sterile dentistry drill. An extract was made using 40 mg of bone powder33.
The Motala individuals were recovered from the site of Kanaljorden in the town of Motala, Östergötland, Sweden, excavated between 2009 and 2013. The human remains at this site are represented by several adult skulls and one infant skeleton. All individuals are part of a ritual deposition at the bottom of a small lake. Direct radiocarbon dates on the remains range between 7,013 ± 76 and 6,701 ± 64 BP (6,361-5,516 cal BC), corresponding to the late Middle Mesolithic of Scandinavia. Samples were taken from the teeth of the nine best preserved skulls, as well as a femur and tibia. Bone powder was removed from the inner parts of the teeth or bones with a sterile dentistry drill. DNA from 100 mg of bone powder was extracted35 in the ancient DNA laboratory of the Archaeological Research Laboratory, Stockholm.
Library preparation
Illumina sequencing libraries were prepared using either double- or single-stranded library preparation protocols36,37 (SI1). For high-coverage shotgun sequencing libraries, a DNA repair step with Uracil-DNA-glycosylase (UDG) and endonuclease VIII (endo VIII) treatment was included in order to remove uracil residues38. Size fractionation on a PAGE gel was also performed in order to remove longer DNA molecules that are more likely to be contaminants37. Positive and blank controls were carried along during every step of library preparation.
Shotgun sequencing and read processing
All non-UDG-treated libraries were sequenced either on an Illumina Genome Analyzer IIx with 2×76 + 7 cycles for the Loschbour and Motala libraries, or on an Illumina MiSeq with 2×150 + 8 + 8 cycles for the Stuttgart library. We followed the manufacturer’s protocol for multiplex sequencing. Raw overlapping forward and reverse reads were merged and filtered for quality39 and mapped to the human reference genome (hg19/GRCh37/1000Genomes) using the Burrows-Wheeler Aligner (BWA)40 (SI2). For deeper sequencing, UDG-treated libraries of Loschbour were sequenced on 3 Illumina HiSeq 2000 lanes with 50-bp single-end reads, 8 Illumina HiSeq 2000 lanes of 100-bp paired-end reads and 8 Illumina HiSeq 2500 lanes of 101-bp paired-end reads. The UDG-treated library for Stuttgart was sequenced on 8 HiSeq 2000 lanes and 101-bp paired-end reads. The UDG-treated libraries for Motala were sequenced on 8 HiSeq 2000 lanes of 100-bp paired-end reads, with 4 lanes each for two pools (one of 3 individuals and one of 4 individuals). We also sequenced an additional 8 HiSeq 2000 lanes for Motala12, the Motala sample with the highest percentage of endogenous human DNA. For the Loschbour and Stuttgart high coverage individuals, diploid genotype calls were obtained using the Genome Analysis Toolkit (GATK)41.
Enrichment of mitochondrial DNA and sequencing
To test for DNA preservation and mtDNA contamination non-UDG-treated libraries of Loschbour and all Motala samples were enriched for human mitochondrial DNA using a bead-based capture approach with present-day human DNA as bait42. UDG-treatment was omitted in order to allow characterization of damage patterns typical for ancient DNA10. The captured libraries were sequenced on an Illumina Genome Analyzer IIx platform with 2 × 76 + 7 cycles and the resulting reads were merged and quality filtered39. The sequences were mapped to the Reconstructed Sapiens Reference Sequence, RSRS43, using a custom iterative mapping assembler, MIA44 (SI4).
Contamination estimates
We assessed if the sequences had the characteristics of authentic ancient DNA using four approaches. First we searched for evidence of contamination by determining whether the sequences mapping to the mitochondrial genome were consistent with deriving from more than one individual44,45. Second, for the high-coverage Loschbour and Stuttgart genomes, we used a maximum-likelihood-based estimate of autosomal contamination that uses variation at sites that are fixed in the 1000 Genomes data to estimate error, heterozygosity and contamination46 simultaneously. Third, we estimated contamination based on the rate of polymorphic sites on the X chromosome of the male Loschbour individual47 (SI3) Fourth, we analyzed non-UDG treated reads mapping to the RSRS to search for aDNA-typical damage patterns resulting in C→T changes at the 5′-end of the molecule10 (SI3).
Phylogenetic analysis of the mitochondrial genomes
All nine complete mitochondrial genomes that fulfilled the criteria of authenticity were assigned to haplogroups using Haplofind48. A Maximum Parsimony tree including present day humans and previously published ancient mtDNA sequences was generated with MEGA49. The effect of branch shortening due to a lower number of substitutions in ancient lineages was studied by calculating the nucleotide edit distance to the root for all haplogroup R sequences (SI4).
Sex determination and Y-chromosome analysis
We assessed the sex of all sequenced individuals by using the ratio of (chrY) to (chrY+chrX) aligned reads50. We downloaded a list of Y-chromosome SNPs curated by the International Society of Genetic Genealogy (ISOGG, http://www.isogg.org) v. 9.22 (accessed Feb. 18, 2014) and determined the state of the ancient individuals at positions where a single allele was observed and MAPQ≥30. We excluded C/G or A/T SNPs due to uncertainty about the polarity of the mutation in the database. The ancient individuals were assigned haplogroups based on their derived state (SI5). We also used BEAST v1.7.5151 to assess the phylogenetic position of Loschbour using 623 males from around the world with 2,799 variant sites across 500kb of non-recombining Y-chromosome sequence52 (SI5).
Estimation of Neanderthal admixture
We estimate Neanderthal admixture in ancient individuals with the f4-ratio or S-statistic8,53,54 α̂ = f4(Altai, Denisova; Test, Yoruba)/f4(Altai, Denisova; Vindija, Yoruba) which uses whole genome data from Altai, a high coverage (52×) Neanderthal genome sequence55, Denisova, a high coverage sequence37 from another archaic human population (31×), and Vindija, a low coverage (1.3×) Neanderthal genome from a mixture of three Neanderthal individuals from Vindija Cave in Croatia53.
Inference of demographic history and inbreeding
We used the Pairwise Sequentially Markovian Coalescent (PSMC)56 to infer the size of the ancestral population of Stuttgart and Loschbour. This analysis requires high quality diploid genotype calls and cannot be performed in the low-coverage Motala samples. To determine whether the low effective population size inferred for Loschbour is due to recent inbreeding, we plotted the time-to-most-recent common ancestor (TMRCA) along each of chr1-22 to detect runs of low TMRCA.
Analysis of segmental duplications and copy number variants
We built read-depth based copy number maps for the Loschbour, Stuttgart and Motala12 genomes in addition to the Denisova and Altai Neanderthal genome and 25 deeply sequenced modern genomes55 (SI7). We built these maps by aligning reads, subdivided into their non-overlapping 36-bp constituents, against the reference genome using the mrsFAST aligner57, and renormalizing read-depth for local GC content. We estimated copy numbers in windows of 500 unmasked base pairs slid at 100 bp intervals across the genome. We called copy number variants using a scale space filter algorithm. We genotyped variants of interest and compared the genotypes to those from individuals sequenced as part of the 1000 Genomes Project58.
Phenotypic inference
We inferred likely phenotypes (SI8) by analyzing DNA polymorphism data in the VCF format59 using VCFtools (http://vcftoools.sourceforge.net/). For the Loschbour and Stuttgart individuals, we included data from sites not flagged as LowQuality, with genotype quality (GQ) of ≥30, and SNP quality (QUAL) of ≥50. For Motala12, which is of lower coverage, we included sites having at least 2× coverage and that passed visual inspection of the local alignment using samtools tview (http://samtools.sourceforge.net)60
Human Origins dataset curation
The Human Origins array consists of 14 panels of SNPs for which the ascertainment is well known8,61. All population genetics analysis were carried out on a set of 594,924 autosomal SNPs, after restricting to sites that had >90% completeness across 7 different batches of sequencing, and that had >97.5% concordance with at least one of two subsets of samples for which whole genome sequencing data was also available. The total dataset consists of 2,722 individuals, which we filtered to 2,345 individuals (203 populations) after removing outlier individuals or relatives based on visual inspection of PCA plots14,62 or model-based clustering analysis13. Whole genome amplified (WGA) individuals were not used in analysis, except for a Saami individual who we included because of the special interest of this population for Northeastern European population history (Extended Data Fig. 7).
ADMIXTURE analysis
We merged all Human Origins genotype data with whole genome sequencing data from Loschbour, Stuttgart, MA1, Motala12, Motala_merge, and LaBrana. We then thinned the resulting dataset to remove SNPs in linkage-disequilibrium with PLINK 1.0763, using a window size of 200 SNPs advanced by 25 SNPs and an r2 threshold of 0.4. We ran ADMIXTURE 1.2313,64 for 100 replicates with different starting random seeds, default 5-fold cross-validation, and varying the number of ancestral populations K between 2 and 20. We assessed clustering quality using CLUMPP65. We used the ADMIXTURE results to identify a set of 59 “West Eurasian” (European/Near Eastern) populations based on values of a “West Eurasian” ancestral population at K=3 (SI9). We also identified 15 populations for use as “non-West Eurasian outgroups” based on their having at least 10 individuals and no evidence of European or Near Eastern admixture at K=11, the lowest K for which Near Eastern/European-maximized ancestral populations appeared consistently across all 100 replicates.
Principal Components Analysis
We used smartpca14 (version: 10210) from EIGENSOFT62,66 5.0.1 to carry out Principal Components Analysis (PCA) (SI10). We performed PCA on a subset on individuals and then projected others using the lsqproject: YES option that gives an unbiased inference of the position of samples even in the presence of missing data (especially important for ancient DNA).
f3-statistics
We use the f3-statistic8 , where ti, r1,i and r2,i are the allele frequencies for the ith SNP in populations Test, Ref1, Ref2, respectively, to determine if there is evidence that the Test population is derived from admixture of populations related to Ref1 and Ref2 (SI11). A significantly negative statistic provides unambiguous evidence of mixture in the Test population8. We allow Ref1 and Ref2 to be any Human Origins population with 4 or more individuals, or Loschbour, Stuttgart, MA1, Motala12, LaBrana. We assess significance of the f3-statistics using a block jackknife67 and a block size of 5cM. We report significance as the number of standard errors by which the statistic differs from zero (Z-score). We also perform an analysis in which we constrain the reference populations to be (i) EEF (Stuttgart) and WHG (Loschbour or LaBrana), (ii) EEF and a Near Eastern population, (iii) EEF and ANE (MA1), or (iv) any two present-day populations, and compute a Zdiff score between the lowest f3-statistic observed in the dataset, and the f3-statistic observed for the specified pair.
f4-statistics
We analyze f4-statistics8 of the form to assess if populations A, B are consistent with forming a clade in an unrooted tree with respect to C, D. If they form a clade, the allele frequency differences between the two pairs should be uncorrelated and the statistic has an expected value of 0. We set the outgroup D to be a sub-Saharan African population or Chimpanzee. We systematically tried all possible combinations of the ancient samples or 15 “non-West Eurasian outgroups” identified by ADMIXTURE analysis as A, B, C to determine their genetic affinities (SI14). Setting A as a present-day test population and B as either Stuttgart or BedouinB, we documented relatedness to C=(Loschbour or MA1) or C=(MA1 and Karitiana) or C=(MA1 or Han) (Extended Data Figs. 4, 5, 7). Setting C as a test population and (A, B) a pair from (Loschbour, Stuttgart, MA1) we documented differential relatedness to ancient populations (Extended Data Fig. 6). We computed D-statistics53 using transversion polymorphisms in whole genome sequence data55 to confirm robustness to ascertainment and ancient DNA damage (Extended Data Table 2).
Minimum number of source populations for Europeans
We used qpWave16,17 to study the minimum number of source populations for a designated set of Europeans (SI12). We use f4-statistics of the form X(l, r) = f4(l0, l; r0, r) where l0,r0 are arbitrarily chosen “base” populations, and l, r are other populations from two sets L and R respectively. If X(l, r) has rank r and there were n waves of immigration into R with no back-migration from R to L, then r+1 ≤ n. We set L to include Stuttgart, Loschbour, MA1, Onge, Karitiana, Mbuti and R to include 23 modern European populations who fit the model of SI14 and had admixture proportions within the interval [0,1] for the method with minimal modeling assumptions (SI17).
Admixture proportions for Stuttgart in the absence of a Near Eastern ancient genome
We used Loschbour and BedouinB as surrogates for “Unknown hunter-gatherer” and Near Eastern (NE) farmer populations that contributed to Stuttgart (SI13). Ancient Near Eastern ancestry in Stuttgart is estimated by the f4-ratio8,15 f4(Outgroup, X; Loschbour, Stuttgart)/f4(Outgroup, X; Loschbour, NE). A complication is that BedouinB is a mixture of NE and African ancestry. We therefore subtracted17 the effects of African ancestry using estimates of the BedouinB African admixture proportion from ADMIXTURE (SI9) or ALDER68.
Admixture graph modeling
We used ADMIXTUREGRAPH8 (version 3110) to model population relationships between Loschbour, Stuttgart, Onge, and Karitiana using Mbuti as an African outgroup. We assessed model fit using a block jackknife of differences between estimated and fitted f-statistics for the set of included populations (we expressed the fit as a Z score). We determined that a model failed if |Z|>3 for at least one f-statistic. A basic tree model failed and we manually amended the model to test all possible models with a single admixture event, which also failed. Further manual amendment to include 2 admixture events resulted in 8 successful models, only one of which could be amended to also fit MA1 as an additional constraint. We successfully fit both the Iceman and LaBrana into this model as simple clades and Motala12 as a 2-way mixture. We also fit present-day West Eurasians as clades, 2-way mixtures, or 3-way mixtures in this basic model, achieving a successful fit for a larger number of European populations (n=26) as 3-way mixtures. We estimated the individual admixture proportions from the fitted model parameters. To test if fitted parameters for different populations are consistent with each other, we jointly fit all pairs of populations A and B by modifying ADMIXTUREGRAPH to add a large constant (10,000) to the variance term f3(A0, A, B). By doing this, we can safely ignore recent gene flow within Europe that affects statistics that include both A and B.
Ancestry estimates from f4-ratios
We estimate EEF ancestry using the f4-ratio8,15 f4(Mbuti, Onge; Loschbour, European)/f4(Mbuti, Onge; Loschbour, Stuttgart), which produces consistent results with ADMIXTUREGRAPH (SI14). We use f4(Stuttgart, Loschbour; Onge MA1)/f4(Mbuti, MA1; Onge, Loschbour) to estimate Basal Eurasian admixture into Stuttgart. We use f4(Stuttgart, Loschbour; Onge Karitiana)/f4(Stuttgart, Loschbour; Onge MA1) to estimate ANE mixture in Karitiana (Fig. 2B). We use f4(Test, Stuttgart; Karitiana, Onge)/f4(MA1, Stuttgart; Karitiana, Onge) to lower bound ANE mixture into North Caucasian populations.
MixMapper analysis
We carried out MixMapper 2.07 analysis, a semi-supervised admixture graph fitting technique. First, we infer a scaffold tree of populations without strong evidence of mixture relative to each other (Mbuti, Onge, Loschbour and MA1). We do not include European populations in the scaffold as all had significantly negative f3-statistics indicating admixture. We then ran MixMapper to infer the relatedness of the other ancient and present-day samples, fitting them onto the scaffold as 2- or 3-way mixtures. The uncertainty in all parameter estimates is measured by block bootstrap resampling of the SNP set (100 replicates with 50 blocks).
TreeMix analysis
We applied TreeMix21 to Loschbour, Stuttgart, Motala12, and MA13, LaBrana2 and the Iceman1, along with the present-day samples of Karitiana, Onge and Mbuti. We restricted the analysis to 265,521 Human Origins array sites after excluding any SNPs where there were no-calls in any of the studied individuals. The tree was rooted with Mbuti and standard errors were estimated using blocks of 500 SNPs. We repeated the analysis on whole-genome sequence data, rooting with Chimp and replacing Onge with Dai since we did not have Onge whole genome sequence data55. We varied the number of migration events (m) between 0 and 5.
Inferring admixture proportions with minimal modeling assumptions
We devised a method to infer ancestry proportions from three ancestral populations (EEF, WHG, and ANE) without strong phylogenetic assumptions (SI17). We rely on 15 “non-West Eurasian” outgroups and study f4(European, Stuttgart; O1, O2) which equals αβ f4(Loschbour, Stuttgart; O1, O2) + α(1−β) f4(MA1, Stuttgart; O1, O2) if European has 1−a ancestry from EEF and β, 1−β ancestry from WHG and ANE respectively. This defines a system of equations with unknowns αβ, α(1−β), which we solve with least squares implemented in the function lsfit in R to obtain estimates of α and β. We repeated this computation 22 times dropping one chromosome at a time20 to obtain block jackknife67 estimates of the ancestry proportions and standard errors, with block size equal to the number of SNPs per chromosome. We assessed consistency of the inferred admixture proportions with those derived from the ADMIXTUREGRAPH model based on the number of standard errors between the two (Extended Data Table 1).
Haplotype-based analyses
We used RefinedIBD from BEAGLE 427 with the settings ibdtrim=20 and ibdwindow=25 to study IBD sharing between Loschbour and Stuttgart and populations from the POPRES dataset69. We kept all IBD tracts spanning at least 0.5 centimorgans (cM) and with a LOD score >3 (SI18). We also used ChromoPainter29 to study haplotype sharing between Loschbour and Stuttgart and present-day West Eurasian populations (SI19). We identified 495,357 SNPs that were complete in all individuals and phased the data using Beagle 427 with parameters phase-its=50 and impute-its=10. We did not keep sites with missing data to avoid imputing modern alleles into the ancient individuals. We used both unlinked (-k 1000) and linked modes (estimating -n and -M by sampling 10% of individuals). We combined ChromoPainter output for chromosomes 1-22 using ChromoCombine29. We carried out a PCA of the co-ancestry matrix using fineSTRUCTURE29.
Extended Data
Extended Data Figure 1.
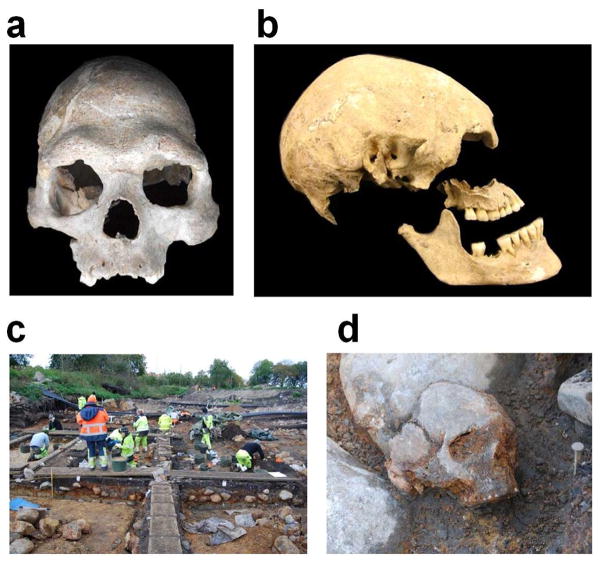
Photographs of analyzed ancient samples.
(A) Loschbour skull; (B) Stuttgart skull, missing the lower right M2 we sampled; (C) excavation at Kanaljorden in Motala, Sweden; (D) Motala 1 in situ.
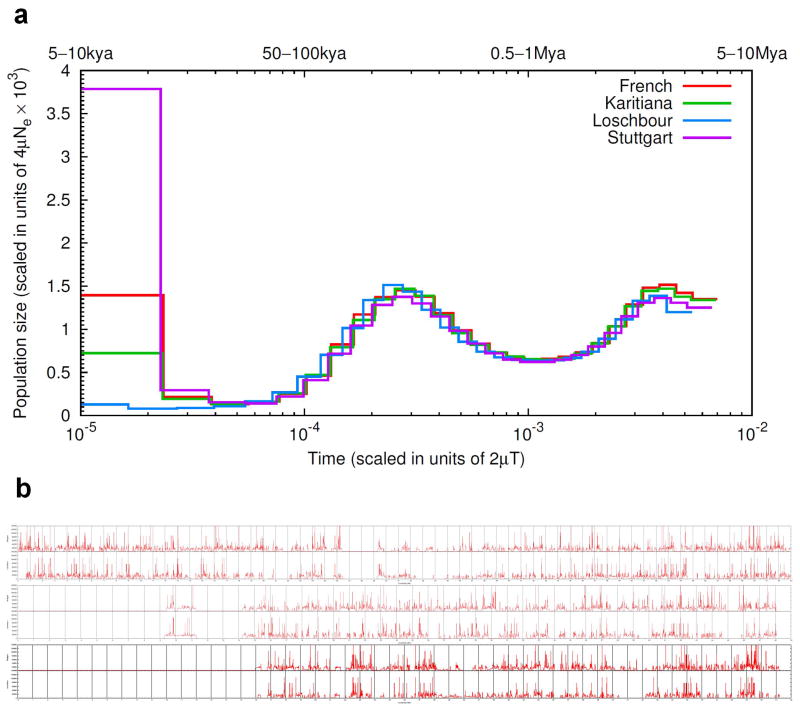
Extended Data Figure 2.
Pairwise Sequential Markovian Coalescent (PSMC) analysis.
(A) Inference of population size as a function of time, showing a very small recent population size over the most recent period in the ancestry of Loschbour (at least the last 5–10 thousand years). (B) Inferred time since the most recent common ancestor from the PSMC for chromosomes 20, 21, 22 (top to bottom); Stuttgart is plotted on top and Loschbour at bottom.
Extended Data Figure 3.
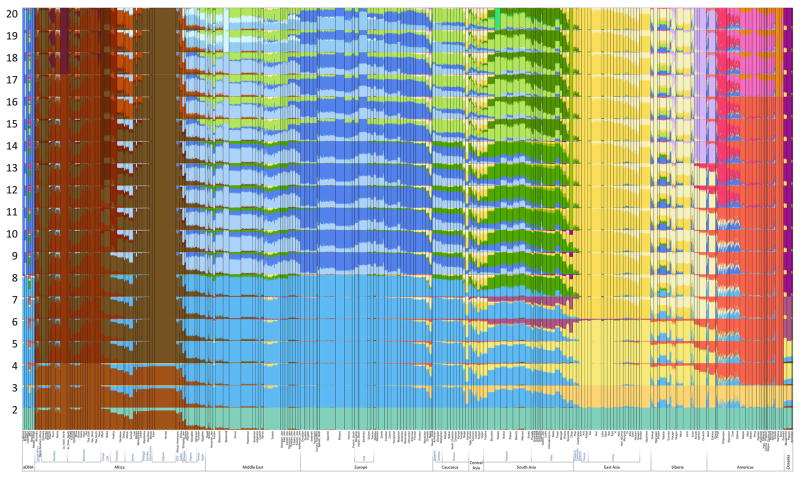
ADMIXTURE analysis (K=2 to K=20).
Ancient samples (Loschbour, Stuttgart, Motala_merge, Motala12, MA1, and LaBrana) are at left.
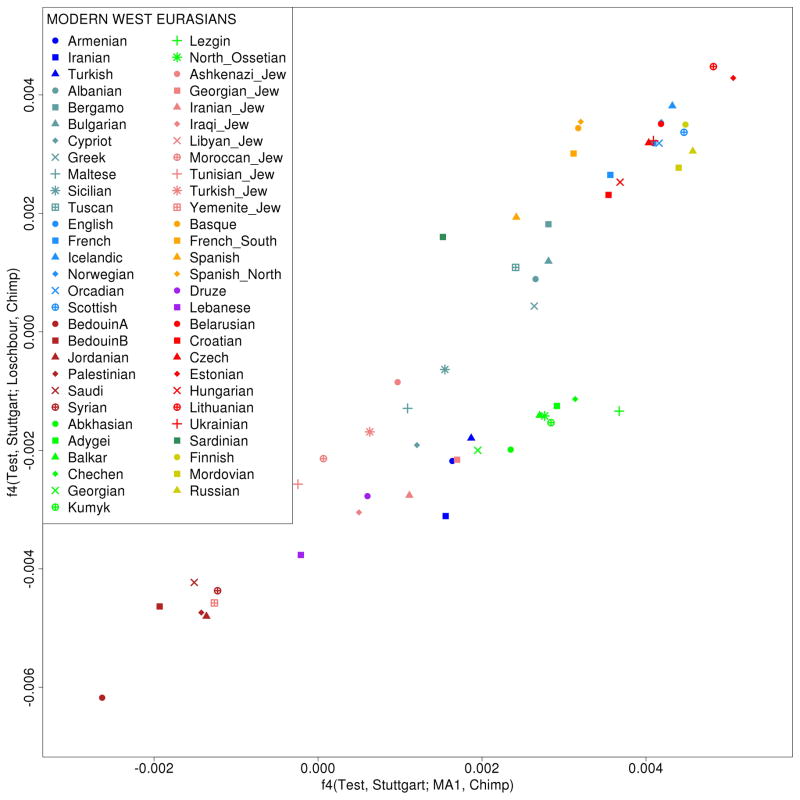
Extended Data Figure 4.
ANE ancestry is present in both Europe and the Near East but WHG ancestry is restricted to Europe, which cannot be due to a single admixture event.
(x-axis) We computed the statistic f4(Test, Stuttgart; MA1, Chimp), which measures where MA1 shares more alleles with a test population than with Stuttgart. It is positive for most European and Near Eastern populations, consistent with ANE (MA1-related) gene flow into both regions. (y-axis) We computed the statistic f4(Test, Stuttgart; Loschbour, Chimp), which measures whether Loschbour shares more alleles with a test sample than with Stuttgart. Only European populations show positive values of this statistic, providing evidence of WHG (Loschbour-related) admixture only in Europeans.
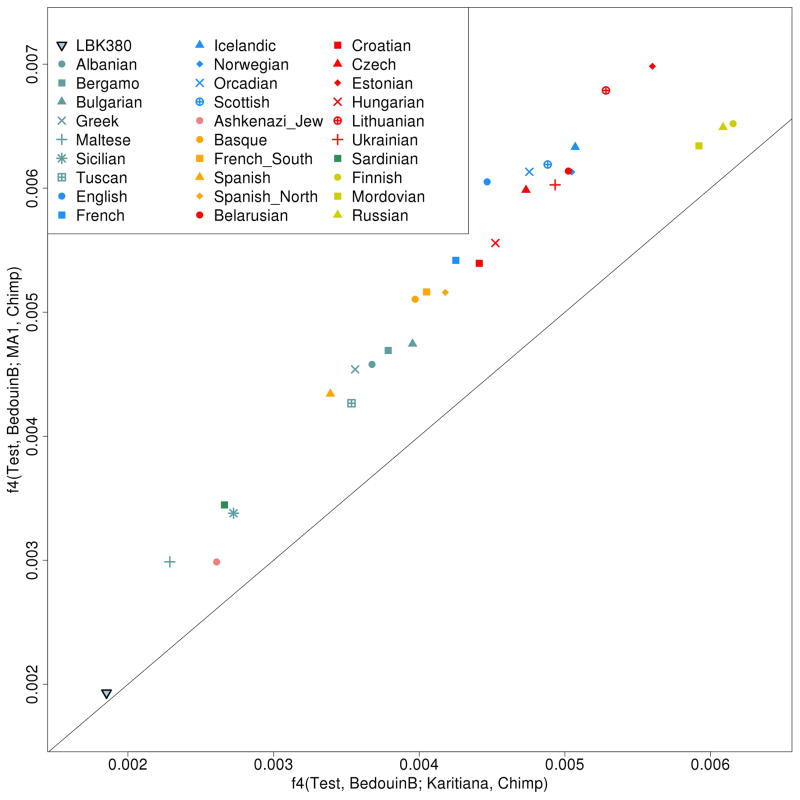
Extended Data Figure 5.
MA1 is the best surrogate for ANE for which we have data.
Europeans share more alleles with MA1 than with Karitiana, as we see from the fact that in a plot of f4(Test, BedouinB; MA1, Chimp) and f4(Test, BedouinB; Karitiana, Chimp), the European cline deviates in the direction of MA1, rather than Karitiana (the slope is >1 and European populations are above the line indicating equality of these two statistics).
Extended Data Figure 6.
The differential relatedness of West Eurasians to Stuttgart (EEF), Loschbour (WHG), and MA1 (ANE) cannot be explained by two-way mixture.
We plot on a West Eurasian map the statistic f4(Test, Chimp; A1, A2), where A1 and A2 are a pair of the three ancient samples representing the three ancestral populations of Europe. (A) In both Europe and the Near East/Caucasus, populations from the south have more relatedness to Stuttgart than those from the north where ANE influence is also important. (B) Northern European populations share more alleles with Loschbour than with Stuttgart, as they have additional WHG ancestry beyond what was already present in EEF. (C) We observe a striking contrast between Europe west of the Caucasus and the Near East in degree of relatedness to WHG. In Europe, there is a much higher degree of allele sharing with Loschbour than with MA1, which we ascribe to the 60–80% WHG/(WHG+ANE) ratio in most Europeans that we report in SI14. In contrast, the Near East has no appreciable WHG ancestry but some ANE ancestry, especially in the northern Caucasus. (Jewish populations are marked with a square in this figure to assist in interpretation as their ancestry is often anomalous for their geographic regions.)
Extended Data Figure 7.
Evidence for Siberian gene flow into far northeastern Europe.
Some northeastern European populations (Chuvash, Finnish, Russian, Mordovian, Saami) share more alleles with Han Chinese than with other Europeans who are arrayed in a cline from Stuttgart to Lithuanians/Estonians in a plot of f4(Test, BedouinB; Han, Mbuti) against f4(Test, BedouinB; MA1, Mbuti).
Extended Data Table 1.
West Eurasians genotyped on the Human Origins array and key f-statistics.
| Sampling Location | Lowest f3(X; Ref1, Ref2) | Lowest f3(X; EEF,
WHG) (Z<0 and Zdiff<3 reported) |
Lowest f3(X; Near
East, WHG) (Z<0 and Zdiff<3 reported) |
Lowest f3(X; EEF,
ANE) (Z<0 and Zdiff<3 reported) |
f4(Stuttgart,
X; Loschbour, Chimp) |
f4(Stuttgart,
X; MA1, Chimp) |
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| X | N | Lat. | Long. | Ref1 | Ref2 | statistic | Z | Ref1 | Ref2 | statistic | Z | Zdiff | Ref1 | Ref2 | statistic | Z | Zdiff | Ref1 | Ref2 | statistic | Z | Zdiff | statistic | Z | statistic | Z |
| Abkhasian | 9 | 43 | 41.02 | Stu | MA1 | −0.0053 | −2.9 | Georgian | LaB | −0.0004 | −0.5 | 2.6 | Stu | MA1 | −0.0053 | −2.9 | 0.0 | 0.0020 | 4.2 | −0.0023 | −4.7 | |||||
| Adygei | 17 | 44 | 39 | Piapoco | Stu | −0.0073 | −5.9 | Stu | MA1 | −0.0067 | −4.1 | 0.3 | 0.0013 | 2.6 | −0.0029 | −6.0 | ||||||||||
| Albanian | 6 | 41.33 | 19.83 | Stu | MA1 | −0.0121 | −7.0 | Iraqi_Jew | Los | −0.0090 | −9.1 | 1.7 | Stu | MA1 | −0.0121 | −7.0 | 0.0 | −0.0009 | −1.8 | −0.0027 | −5.4 | |||||
| Armenian | 10 | 40.19 | 44.55 | GujaratiC | Stu | −0.0070 | −8.2 | Stu | MA1 | −0.0068 | −4.1 | 0.1 | 0.0022 | 4.5 | −0.0016 | −3.3 | ||||||||||
| Ashkenazi_Jew | 7 | 52.23 | 21.02 | Stu | MA1 | −0.0057 | −3.4 | Iraqi_Jew | Los | −0.0042 | −4.7 | 1.0 | Stu | MA1 | −0.0057 | −3.4 | 0.0 | 0.0008 | 1.7 | −0.0010 | −2.0 | |||||
| Balkar | 10 | 43.48 | 43.62 | Piapoco | Stu | −0.0113 | −8.9 | Stu | MA1 | −0.0092 | −5.5 | 1.1 | 0.0014 | 2.9 | −0.0027 | −5.6 | ||||||||||
| Basque | 29 | 43.04 | −0.65 | Iraqi_Jew | Los | −0.0083 | −10.3 | Stu | Los | −0.0061 | −3.8 | 1.3 | Iraqi_Jew | Los | −0.0083 | −10.3 | 0.0 | Stu | MA1 | −0.0041 | −2.4 | 2.2 | −0.0034 | −7.2 | −0.0032 | −6.7 |
| BedouinA | 25 | 31 | 35 | Esan | Stu | −0.0162 | −18.2 | 0.0062 | 13.0 | 0.0026 | 5.4 | |||||||||||||||
| BedouinB | 19 | 31 | 35 | Esan | Stu | 0.0089 | 7.8 | 0.0046 | 9.3 | 0.0019 | 3.9 | |||||||||||||||
| Belarusian | 10 | 53.92 | 28.01 | Georgian | Los | −0.0133 | −17.6 | Georgian | Los | −0.0133 | −17.6 | 0.0 | Stu | MA1 | −0.0102 | −6.1 | 1.9 | −0.0035 | −6.9 | −0.0042 | −8.6 | |||||
| Bergamo | 12 | 46 | 10 | Stu | MA1 | −0.0106 | −6.2 | Stu | Los | −0.0068 | −4.2 | 1.7 | Iraqi_Jew | Los | −0.0100 | −11.9 | 0.3 | Stu | MA1 | −0.0106 | −6.2 | 0.0 | −0.0018 | −3.9 | −0.0028 | −5.8 |
| Bulgarian | 10 | 42.16 | 24.74 | Stu | MA1 | −0.0130 | −8.2 | Stu | LaB | −0.0074 | −4.5 | 2.8 | Iraqi_Jew | Los | −0.0106 | −12.4 | 1.5 | Stu | MA1 | −0.0130 | −8.2 | 0.0 | −0.0012 | −2.5 | −0.0028 | −5.9 |
| Chechen | 9 | 43.33 | 45.65 | Stu | MA1 | −0.0056 | −3.2 | Georgian | Los | −0.0002 | −0.3 | 2.8 | Stu | MA1 | −0.0056 | −3.2 | 0.0 | 0.0011 | 2.3 | −0.0031 | −6.2 | |||||
| Croatian | 10 | 43.51 | 16.45 | Stu | MA1 | −0.0114 | −6.7 | Stu | Los | −0.0065 | −3.8 | 2.1 | Iraqi_Jew | Los | −0.0112 | −13.0 | 0.2 | Stu | MA1 | −0.0114 | −6.7 | 0.0 | −0.0023 | −4.7 | −0.0035 | −7.4 |
| Cypriot | 8 | 35.13 | 33.43 | Stu | MA1 | −0.0057 | −3.2 | Yemenite_Jew | Los | −0.0013 | −1.5 | 2.5 | Stu | MA1 | −0.0057 | −3.2 | 0.0 | 0.0019 | 3.9 | −0.0012 | −2.5 | |||||
| Czech | 10 | 50.1 | 14.4 | Georgian | Los | −0.0137 | −17.9 | Stu | Los | −0.0088 | −5.3 | 3.0 | Georgian | Los | −0.0137 | −17.9 | 0.0 | Stu | MA1 | −0.0121 | −7.2 | 0.9 | −0.0032 | −6.6 | −0.0040 | −8.2 |
| Druze | 39 | 32 | 35 | Stu | MA1 | −0.0024 | −1.5 | Stu | MA1 | −0.0024 | −1.5 | 0.0 | 0.0028 | 5.9 | −0.0006 | −1.3 | ||||||||||
| English | 10 | 50.75 | −2.09 | Iraqi_Jew | Los | −0.0129 | −14.8 | Stu | Los | −0.0090 | −5.5 | 2.2 | Iraqi_Jew | Los | −0.0129 | −14.8 | 0.0 | Stu | MA1 | −0.0125 | −7.4 | 0.1 | −0.0032 | −6.5 | −0.0041 | −8.5 |
| Estonian | 10 | 58.54 | 24.89 | Abkhasian | Los | −0.0124 | −15.1 | Abkhasian | Los | −0.0124 | −15.1 | 0.0 | Stu | MA1 | −0.0094 | −5.6 | 1.9 | −0.0043 | −8.5 | −0.0051 | −10.1 | |||||
| Finnish | 7 | 60.2 | 24.9 | Abkhasian | Los | −0.0102 | −11.3 | Abkhasian | Los | −0.0102 | −11.3 | 0.0 | Stu | MA1 | −0.0078 | −4.4 | 1.4 | −0.0035 | −6.9 | −0.0045 | −9.1 | |||||
| French | 25 | 46 | 2 | Stu | MA1 | −0.0131 | −8.4 | Stu | Los | −0.0098 | −6.3 | 1.5 | Iraqi_Jew | Los | −0.0129 | −16.8 | 0.2 | Stu | MA1 | −0.0131 | −8.4 | 0.0 | −0.0027 | −5.6 | −0.0036 | −7.7 |
| French_South | 7 | 43.44 | −0.62 | Iraqi_Jew | Los | −0.0095 | −9.5 | Stu | LaB | −0.0089 | −5.0 | 0.3 | Iraqi_Jew | Los | −0.0095 | −9.5 | 0.0 | Stu | MA1 | −0.0086 | −4.8 | 0.4 | −0.0030 | −6.2 | −0.0031 | −6.2 |
| Georgian | 10 | 42.5 | 41.85 | GujaratiC | Stu | −0.0036 | −4.0 | Stu | MA1 | −0.0036 | −2.1 | −0.2 | 0.0020 | 4.2 | −0.0019 | −3.9 | ||||||||||
| Georgian_Jew | 7 | 41.72 | 44.78 | GujaratiC | Stu | −0.0009 | −0.9 | Stu | MA1 | −0.0002 | −0.1 | 0.3 | 0.0022 | 4.3 | −0.0017 | −3.4 | ||||||||||
| Greek | 20 | 39.84 | 23.17 | Stu | MA1 | −0.0118 | −7.4 | Iraqi_Jew | Los | −0.0080 | −11.1 | 2.3 | Stu | MA1 | −0.0118 | −7.4 | 0.0 | −0.0004 | −0.9 | −0.0026 | −5.6 | |||||
| Hungarian | 20 | 47.49 | 19.08 | Stu | MA1 | −0.0133 | −8.4 | Stu | Los | −0.0087 | −5.6 | 2.2 | Iraqi_Jew | Los | −0.0127 | −15.9 | 0.4 | Stu | MA1 | −0.0133 | −8.4 | 0.0 | −0.0025 | −5.3 | −0.0037 | −7.8 |
| Icelandic | 12 | 64.13 | −21.93 | Abkhasian | Los | −0.0121 | −15.6 | Stu | Los | −0.0078 | −4.8 | 2.7 | Abkhasian | Los | −0.0121 | −15.6 | 0.0 | Stu | MA1 | −0.0097 | −5.9 | 1.5 | −0.0038 | −7.7 | −0.0043 | −8.9 |
| Iranian | 8 | 35.59 | 51.46 | Piapoco | Stu | −0.0094 | −7.2 | Stu | MA1 | −0.0087 | −5.2 | 0.4 | 0.0031 | 6.3 | −0.0016 | −3.2 | ||||||||||
| Iranian_Jew | 9 | 35.7 | 51.42 | GujaratiC | Stu | −0.0018 | −2.0 | Stu | MA1 | −0.0012 | −0.6 | 0.2 | 0.0028 | 5.7 | −0.0011 | −2.2 | ||||||||||
| Iraqi_Jew | 6 | 33.33 | 44.42 | Vishwabrahmin | Stu | −0.0026 | −2.6 | Stu | MA1 | −0.0009 | −0.5 | 0.9 | 0.0030 | 6.1 | −0.0005 | −1.0 | ||||||||||
| Jordanian | 9 | 32.05 | 35.91 | Esan | Stu | −0.0145 | −14.3 | 0.0048 | 9.6 | 0.0014 | 2.8 | |||||||||||||||
| Kumyk | 8 | 43.25 | 46.58 | Piapoco | Stu | −0.0111 | −8.2 | Stu | MA1 | −0.0109 | −6.5 | 0.1 | 0.0015 | 3.1 | −0.0028 | −5.7 | ||||||||||
| Lebanese | 8 | 33.82 | 35.57 | Esan | Stu | −0.0105 | −9.4 | Stu | MA1 | −0.0068 | −3.9 | 1.9 | 0.0038 | 7.7 | 0.0002 | 0.4 | ||||||||||
| Lezgin | 9 | 42.12 | 48.18 | Stu | MA1 | −0.0100 | −6.0 | Stu | MA1 | −0.0100 | −6.0 | 0.0 | 0.0013 | 2.7 | −0.0037 | −7.5 | ||||||||||
| Libyan_Jew | 9 | 32.92 | 13.18 | Esan | Stu | −0.0051 | −4.4 | Stu | MA1 | 0.0000 | 0.0 | 2.7 | 0.0030 | 6.2 | 0.0004 | 0.9 | ||||||||||
| Lithuanian | 10 | 54.9 | 23.92 | Abkhasian | Los | −0.0119 | −14.9 | Abkhasian | Los | −0.0119 | −14.9 | 0.0 | Stu | MA1 | −0.0069 | −3.9 | 2.8 | −0.0045 | −9.0 | −0.0048 | −9.9 | |||||
| Maltese | 8 | 35.94 | 14.38 | Stu | MA1 | −0.0086 | −4.9 | Yemenite_Jew | Los | −0.0051 | −6.0 | 2.0 | Stu | MA1 | −0.0086 | −4.9 | 0.0 | 0.0013 | 2.7 | −0.0011 | −2.3 | |||||
| Mordovian | 10 | 54.18 | 45.18 | Abkhasian | Los | −0.0115 | −14.4 | Abkhasian | Los | −0.0115 | −14.4 | 0.0 | Stu | MA1 | −0.0113 | −6.6 | 0.3 | −0.0028 | −5.5 | −0.0044 | −9.0 | |||||
| Moroccan_Jew | 6 | 34.02 | −6.84 | Esan | Stu | −0.0062 | −5.2 | Yemenite_Jew | Los | −0.0021 | −2.2 | 2.9 | Stu | MA1 | −0.0032 | −1.7 | 1.4 | 0.0021 | 4.3 | −0.0001 | −0.1 | |||||
| North_Ossetian | 10 | 43.02 | 44.65 | Piapoco | Stu | −0.0093 | −7.2 | Stu | MA1 | −0.0076 | −4.4 | 1.0 | 0.0014 | 2.9 | −0.0028 | −5.6 | ||||||||||
| Norwegian | 11 | 60.36 | 5.36 | Georgian | Los | −0.0120 | −14.8 | Georgian | Los | −0.0120 | −14.8 | 0.0 | Stu | MA1 | −0.0093 | −5.4 | 1.4 | −0.0035 | −7.3 | −0.0042 | −8.7 | |||||
| Orcadian | 13 | 59 | −3 | Armenian | Los | −0.0102 | −13.4 | Stu | Los | −0.0059 | −3.6 | 2.5 | Armenian | Los | −0.0102 | −13.4 | 0.0 | Stu | MA1 | −0.0098 | −5.9 | 0.5 | −0.0032 | −6.7 | −0.0042 | −8.6 |
| Palestinian | 38 | 32 | 35 | Esan | Stu | −0.0120 | −13.2 | 0.0047 | 10.2 | 0.0014 | 3.1 | |||||||||||||||
| Russian | 22 | 61 | 40 | Chukchi | Los | −0.0119 | −11.3 | Abkhasian | Los | −0.0119 | −17.1 | 0.0 | Stu | MA1 | −0.0106 | −6.6 | 0.8 | −0.0030 | −6.2 | −0.0046 | −9.4 | |||||
| Sardinian | 27 | 40 | 9 | Stu | LaB | −0.0044 | −2.6 | Stu | LaB | −0.0044 | −2.6 | 0.0 | Iraqi_Jew | Los | −0.0033 | −4.2 | 0.0 | Stu | MA1 | −0.0035 | −2.1 | 0.3 | −0.0016 | −3.4 | −0.0015 | −3.3 |
| Saudi | 8 | 18.49 | 42.52 | Kgalagadi | Stu | −0.0042 | −3.6 | 0.0042 | 8.6 | 0.0015 | 3.1 | |||||||||||||||
| Scottish | 4 | 56.04 | −3.94 | Iraqi_Jew | Los | −0.0103 | −8.3 | Iraqi_Jew | Los | −0.0103 | −8.3 | 0.0 | Stu | MA1 | −0.0090 | −4.7 | 0.7 | −0.0034 | −6.4 | −0.0045 | −8.7 | |||||
| Sicilian | 11 | 37.59 | 13.77 | Stu | MA1 | −0.0108 | −6.5 | Yemenite_Jew | Los | −0.0066 | −8.1 | 2.4 | Stu | MA1 | −0.0108 | −6.5 | 0.0 | 0.0006 | 1.3 | −0.0015 | −3.2 | |||||
| Spanish | 53 | 40.43 | −2.83 | Iraqi_Jew | Los | −0.0126 | −17.8 | Stu | Los | −0.0104 | −6.8 | 1.4 | Iraqi_Jew | Los | −0.0126 | −17.8 | 0.0 | Stu | MA1 | −0.0120 | −7.6 | 0.3 | −0.0019 | −4.2 | −0.0024 | −5.2 |
| Spanish_North | 5 | 42.8 | −2.7 | Iraqi_Jew | Los | −0.0112 | −9.9 | Stu | Los | −0.0102 | −5.4 | 0.5 | Iraqi_Jew | Los | −0.0112 | −9.9 | 0.0 | Stu | MA1 | −0.0082 | −4.4 | 1.3 | −0.0035 | −6.9 | −0.0032 | −6.4 |
| Syrian | 8 | 35.13 | 36.87 | Esan | Stu | −0.0101 | −8.7 | 0.0044 | 8.6 | 0.0012 | 2.4 | |||||||||||||||
| Tunisian_Jew | 7 | 36.8 | 10.18 | Gambian | Stu | −0.0026 | −2.0 | 0.0026 | 5.2 | 0.0002 | 0.5 | |||||||||||||||
| Turkish | 56 | 39.22 | 32.66 | Piapoco | Stu | −0.0129 | −11.3 | Stu | MA1 | −0.0106 | −6.9 | 1.3 | 0.0018 | 3.8 | −0.0019 | −4.0 | ||||||||||
| Turkish_Jew | 8 | 41.02 | 28.95 | Stu | MA1 | −0.0075 | −4.3 | Yemenite_Jew | Los | −0.0049 | −5.8 | 1.4 | Stu | MA1 | −0.0075 | −4.3 | 0.0 | 0.0017 | 3.6 | −0.0006 | −1.3 | |||||
| Tuscan | 8 | 43 | 11 | Stu | MA1 | −0.0109 | −6.4 | Stu | Los | −0.0055 | −3.2 | 2.3 | Iraqi_Jew | Los | −0.0092 | −10.1 | 0.9 | Stu | MA1 | −0.0109 | −6.4 | 0.0 | −0.0011 | −2.2 | −0.0024 | −5.0 |
| Ukrainian | 9 | 50.29 | 31.56 | Georgian | Los | −0.0134 | −16.7 | Georgian | Los | −0.0134 | −16.7 | 0.0 | Stu | MA1 | −0.0114 | −6.6 | 1.3 | −0.0032 | −6.4 | −0.0041 | −8.5 | |||||
| Yemenite_Jew | 8 | 15.35 | 44.2 | Esan | Stu | −0.0027 | −2.4 | 0.0046 | 9.1 | 0.0013 | 2.6 | |||||||||||||||
Note: Zdiff is the number of standard errors of the difference between the lowest f3-statistic over all reference pairs and the lowest f3-statistic for a subset of reference pairs.
Abbreviations used: Stu: Stuttgart; Los: Loschbour; LaB: LaBrana.
Extended Data Table 2.
Confirmation of key findings on transversions and on whole genome sequence data.
| Interpretation | D(A, B; C, D) on Human Origins genotype data | D(A, B; C, D) on whole genome sequence data transversions | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
| ||||||||||||||
| A | B | C | D | 594,924 SNPs | 110,817 transversions | A | B | C | D | statistic | Z | |||
| statistic | Z | statistic | Z | |||||||||||
| Stuttgart has Near Eastern ancestry | Stuttgart | Armenian | Loschbour | Chimp | 0.0219 | 4.5 | 0.0189 | 2.9 | ||||||
|
| ||||||||||||||
| Europeans have more WHG-related ancestry than Stuttgart | Stuttgart | French | Loschbour | Chimp | −0.0266 | −5.7 | −0.031 | −5.0 | Stuttgart | French2 | Loschbour | Chimp | −0.03 | −4.7 |
| Lithuanian | Stuttgart | Loschbour | Chimp | 0.0446 | 9.1 | 0.0477 | 7.2 | |||||||
|
| ||||||||||||||
| West Eurasians have more ANE-related ancestry than Stuttgart | French | Stuttgart | MA1 | Chimp | 0.0367 | 7.7 | 0.0386 | 5.5 | French2 | Stuttgart | MA1 | Chimp | 0.037 | 6.4 |
| Lezgin | Stuttgart | MA1 | Chimp | 0.0372 | 7.6 | 0.0409 | 5.6 | |||||||
|
| ||||||||||||||
| MA1 is a better surrogate of ANE ancestry than Karitiana | French | Chimp | MA1 | Karitiana | 0.0207 | 4.5 | 0.0214 | 2.8 | French2 | Chimp | MA1 | Karitiana2 | 0.026 | 3.8 |
|
| ||||||||||||||
| Eastern non-Africans closer to WHG/ANE/SHG than to EEF | Loschbour | Stuttgart | Onge | Chimp | 0.0196 | 3.5 | 0.0202 | 2.5 | ||||||
| Loschbour | Stuttgart | Papuan | Chimp | 0.0142 | 2.6 | 0.0127 | 1.5 | Loschbour | Stuttgart | Papuan2 | Chimp | 0.017 | 2.7 | |
| Loschbour | Stuttgart | Dai | Chimp | 0.0164 | 3.2 | 0.021 | 2.8 | Loschbour | Stuttgart | Dai2 | Chimp | 0.018 | 2.9 | |
| MA1 | Stuttgart | Papuan | Chimp | 0.0139 | 2.2 | 0.0103 | 1.0 | MA1 | Stuttgart | Papuan2 | Chimp | 0.018 | 2.8 | |
| MA1 | Stuttgart | Dai | Chimp | 0.0174 | 3.0 | 0.016 | 1.7 | MA1 | Stuttgart | Dai2 | Chimp | 0.028 | 4.3 | |
| Motala12 | Stuttgart | Papuan | Chimp | 0.0182 | 3.2 | 0.011 | 1.1 | Motala12 | Stuttgart | Papuan2 | Chimp | 0.023 | 3.7 | |
| Motala12 | Stuttgart | Dai | Chimp | 0.0156 | 2.8 | 0.0149 | 1.6 | Motala12 | Stuttgart | Dai2 | Chimp | 0.02 | 3.2 | |
| LaBrana | Stuttgart | Papuan | Chimp | 0.0123 | 2.3 | 0.0101 | 1.1 | LaBrana | Stuttgart | Papuan2 | Chimp | 0.02 | 3.2 | |
| LaBrana | Stuttgart | Dai | Chimp | 0.0149 | 2.9 | 0.0228 | 2.5 | LaBrana | Stuttgart | Dai2 | Chimp | 0.024 | 3.7 | |
|
| ||||||||||||||
| Native Americans closer to ANE than to WHG | Karitiana | Chimp | MA1 | Loschbour | 0.0467 | 7.1 | 0.0467 | 4.4 | Karitiana2 | Chimp | MA1 | Loschbour | 0.052 | 7.1 |
|
| ||||||||||||||
| West Eurasians closer to Native Americans than to other Eastern non-Africans | Stuttgart | Chimp | Karitiana | Papuan | 0.0559 | 10.9 | 0.0474 | 6.6 | Stuttgart | Chimp | Karitiana2 | Papuan2 | 0.052 | 7.6 |
| Stuttgart | Chimp | Karitiana | Onge | 0.0237 | 5.1 | 0.0179 | 2.6 | |||||||
|
| ||||||||||||||
| Ancient Eurasian hunter-gatherers equally related to Eastern non-Africans other than Native Americans | Loschbour | MA1 | Dai | Chimp | −0.0015 | −0.2 | 0.0016 | 0.2 | Loschbour | MA1 | Dai2 | Chimp | −0.013 | −1.9 |
| Loschbour | MA1 | Papuan | Chimp | 0.0002 | 0.0 | 0.0012 | 0.1 | Loschbour | MA1 | Papuan2 | Chimp | −0.003 | −0.4 | |
| Loschbour | Motala12 | Dai | Chimp | 0.0024 | 0.4 | 0.009 | 0.9 | Loschbour | Motala12 | Dai2 | Chimp | −0.002 | −0.3 | |
| Loschbour | Motala12 | Papuan | Chimp | −0.0028 | −0.4 | 0.0046 | 0.5 | Loschbour | Motala12 | Papuan2 | Chimp | −0.004 | −0.6 | |
| MA1 | Motala12 | Dai | Chimp | 0.0026 | 0.4 | 0.0047 | 0.4 | MA1 | Motala12 | Dai2 | Chimp | 0.01 | 1.5 | |
| MA1 | Motala12 | Papuan | Chimp | −0.0047 | −0.7 | −0.001 | −0.1 | MA1 | Motala12 | Papuan2 | Chimp | −0.004 | −0.5 | |
|
| ||||||||||||||
| LaBrana and Loschbour are a clade | LaBrana | Loschbour | Dai | Chimp | −0.0028 | −0.5 | 0.0024 | 0.3 | LaBrana | Loschbour | Dai2 | Chimp | 0.007 | 1.1 |
| LaBrana | Loschbour | Papuan | Chimp | −0.0031 | −0.5 | −0.0012 | −0.1 | LaBrana | Loschbour | Papuan2 | Chimp | 0.002 | 0.3 | |
| LaBrana | Loschbour | MA1 | Chimp | −0.006 | −0.8 | 0.0101 | 0.7 | LaBrana | Loschbour | MA1 | Chimp | 0.005 | 0.7 | |
|
| ||||||||||||||
| SHG closer to ANE than to WHG | Motala12 | Loschbour | MA1 | Chimp | 0.0425 | 5.3 | 0.0353 | 2.6 | Motala12 | Loschbour | MA1 | Chimp | 0.042 | 5.9 |
| Motala12 | LaBrana | MA1 | Chimp | 0.0465 | 5.8 | 0.0347 | 2.4 | Motala12 | LaBrana | MA1 | Chimp | 0.038 | 5.4 | |
|
| ||||||||||||||
| LaBrana and Loschbour equally related to Stuttgart | LaBrana | Loschbour | Stuttgart | Chimp | −0.0176 | −2.6 | −0.0106 | −1.0 | LaBrana | Loschbour | Stuttgart | Chimp | −0.012 | −1.8 |
Extended Data Table 3.
Admixture proportions for European populations. The estimates from the model with minimal assumptions are from SI17. The estimates from the full modeling are from SI14 either by single population analysis or co-fitting population pairs and averaging over fits (these averages are the results plotted in Fig. 2B). Populations that do not fit the models are not reported.
| Full modeling of population relationships (individual fits) |
Full modeling of population relationships (averaged fits) |
Modeling of
population relationships with minimal assumptions |
Model-based (averaged) - Model with minimal assumptions (Z-score) |
||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EEF | WHG | ANE | EEF | WHG | ANE | EEF | WHG | ANE | EEF | WHG | ANE | ||||
| Mean | Range | Mean | Range | Mean | Range | ||||||||||
| Albanian | 0.781 | 0.092 | 0.127 | 0.781 | 0.772–0.819 | 0.082 | 0.032–0.098 | 0.137 | 0.129–0.158 | 0.595 ± 0.112 | 0.353 ± 0.150 | 0.052 ± 0.049 | 1.658 | −1.807 | 1.741 |
| Ashkenazi_Jew | 0.931 | 0 | 0.069 | 0.938 ± 0.146 | −0.021 ± 0.185 | 0.083 ± 0.049 | |||||||||
| Basque | 0.593 | 0.293 | 0.114 | 0.569 | 0.527–0.616 | 0.335 | 0.255–0.392 | 0.096 | 0.076–0.129 | 0.569 ± 0.091 | 0.315 ± 0.124 | 0.115 ± 0.041 | −0.001 | 0.165 | −0.472 |
| Belarusian | 0.418 | 0.431 | 0.151 | 0.426 | 0.397–0.464 | 0.408 | 0.338–0.443 | 0.167 | 0.150–0.199 | 0.272 ± 0.094 | 0.554 ± 0.131 | 0.174 ± 0.047 | 1.637 | −1.118 | −0.158 |
| Bergamo | 0.715 | 0.177 | 0.108 | 0.721 | 0.704–0.793 | 0.163 | 0.061–0.189 | 0.117 | 0.104–0.147 | 0.644 ± 0.125 | 0.248 ± 0.170 | 0.108 ± 0.053 | 0.615 | −0.503 | 0.162 |
| Bulgarian | 0.712 | 0.147 | 0.141 | 0.718 | 0.707–0.778 | 0.132 | 0.047–0.151 | 0.151 | 0.138–0.175 | 0.556 ± 0.110 | 0.328 ± 0.143 | 0.116 ± 0.043 | 1.469 | −1.372 | 0.804 |
| Croatian | 0.561 | 0.293 | 0.145 | 0.564 | 0.548–0.586 | 0.285 | 0.242–0.310 | 0.151 | 0.137–0.172 | 0.453 ± 0.122 | 0.407 ± 0.159 | 0.140 ± 0.046 | 0.911 | −0.768 | 0.238 |
| Czech | 0.495 | 0.338 | 0.167 | 0.489 | 0.460–0.531 | 0.348 | 0.273–0.382 | 0.163 | 0.145–0.196 | 0.402 ± 0.117 | 0.400 ± 0.162 | 0.198 ± 0.050 | 0.744 | −0.322 | −0.698 |
| English | 0.495 | 0.364 | 0.141 | 0.503 | 0.476–0.536 | 0.353 | 0.296–0.382 | 0.144 | 0.130–0.169 | 0.475 ± 0.091 | 0.357 ± 0.125 | 0.168 ± 0.043 | 0.304 | −0.028 | −0.561 |
| Estonian | 0.322 | 0.495 | 0.183 | 0.323 | 0.293–0.345 | 0.49 | 0.451–0.520 | 0.187 | 0.172–0.205 | 0.072 ± 0.121 | 0.778 ± 0.176 | 0.150 ± 0.064 | 2.070 | −1.636 | 0.584 |
| French | 0.554 | 0.311 | 0.135 | 0.563 | 0.537–0.601 | 0.297 | 0.230–0.328 | 0.14 | 0.126–0.169 | 0.498 ± 0.097 | 0.359 ± 0.127 | 0.142 ± 0.039 | 0.672 | −0.487 | −0.060 |
| French_South | 0.675 | 0.195 | 0.13 | 0.636 | 0.589–0.738 | 0.256 | 0.111–0.323 | 0.108 | 0.088–0.151 | 0.636 ± 0.116 | 0.225 ± 0.165 | 0.140 ± 0.057 | −0.003 | 0.189 | −0.558 |
| Greek | 0.792 | 0.058 | 0.151 | 0.791 | 0.780–0.816 | 0.048 | 0.019–0.060 | 0.161 | 0.150–0.171 | 0.658 ± 0.098 | 0.255 ± 0.127 | 0.086 ± 0.039 | 1.357 | −1.627 | 1.915 |
| Hungarian | 0.558 | 0.264 | 0.179 | 0.548 | 0.520–0.590 | 0.279 | 0.199–0.313 | 0.174 | 0.156–0.210 | 0.391 ± 0.109 | 0.454 ± 0.153 | 0.155 ± 0.050 | 1.437 | −1.145 | 0.371 |
| Icelandic | 0.394 | 0.456 | 0.15 | 0.409 | 0.386–0.424 | 0.448 | 0.409–0.473 | 0.143 | 0.126–0.170 | 0.342 ± 0.102 | 0.476 ± 0.137 | 0.182 ± 0.045 | 0.654 | −0.204 | −0.861 |
| Lithuanian | 0.364 | 0.464 | 0.172 | 0.352 | 0.327–0.384 | 0.488 | 0.433–0.527 | 0.16 | 0.135–0.184 | 0.248 ± 0.117 | 0.548 ± 0.163 | 0.205 ± 0.052 | 0.886 | −0.367 | −0.864 |
| Maltese | 0.932 | 0 | 0.068 | 1.298 ± 0.185 | −0.509 ± 0.248 | 0.211 ± 0.079 | |||||||||
| Norwegian | 0.411 | 0.428 | 0.161 | 0.417 | 0.388–0.438 | 0.423 | 0.383–0.450 | 0.16 | 0.140–0.181 | 0.273 ± 0.115 | 0.557 ± 0.161 | 0.170 ± 0.055 | 1.252 | −0.831 | −0.185 |
| Orcadian | 0.457 | 0.385 | 0.158 | 0.465 | 0.439–0.493 | 0.378 | 0.329–0.403 | 0.157 | 0.140–0.179 | 0.395 ± 0.088 | 0.437 ± 0.122 | 0.168 ± 0.041 | 0.798 | −0.487 | −0.264 |
| Sardinian | 0.817 | 0.175 | 0.008 | 0.818 | 0.791–0.874 | 0.141 | 0.058–0.182 | 0.041 | 0.026–0.068 | 0.883 ± 0.128 | 0.075 ± 0.166 | 0.042 ± 0.048 | −0.510 | 0.400 | −0.024 |
| Scottish | 0.39 | 0.428 | 0.182 | 0.408 | 0.387–0.424 | 0.421 | 0.384–0.448 | 0.171 | 0.149–0.201 | 0.286 ± 0.112 | 0.532 ± 0.156 | 0.182 ± 0.053 | 1.091 | −0.712 | −0.210 |
| Sicilian | 0.903 | 0 | 0.097 | 1.012 ± 0.149 | −0.131 ± 0.199 | 0.119 ± 0.060 | |||||||||
| Spanish | 0.809 | 0.068 | 0.123 | 0.759 | 0.736–0.804 | 0.126 | 0.066–0.170 | 0.115 | 0.091–0.151 | 0.856 ± 0.126 | −0.015 ± 0.165 | 0.160 ± 0.049 | −0.769 | 0.855 | −0.922 |
| Spanish_North | 0.713 | 0.125 | 0.163 | 0.612 | 0.561–0.660 | 0.292 | 0.214–0.365 | 0.096 | 0.072–0.126 | 0.581 ± 0.120 | 0.298 ± 0.158 | 0.121 ± 0.046 | 0.254 | −0.038 | −0.533 |
| Tuscan | 0.746 | 0.136 | 0.118 | 0.751 | 0.737–0.806 | 0.123 | 0.047–0.145 | 0.126 | 0.114–0.150 | 0.734 ± 0.118 | 0.153 ± 0.160 | 0.113 ± 0.054 | 0.141 | −0.188 | 0.249 |
| Ukrainian | 0.462 | 0.387 | 0.151 | 0.463 | 0.445–0.491 | 0.376 | 0.322–0.399 | 0.16 | 0.148–0.187 | 0.259 ± 0.123 | 0.596 ± 0.173 | 0.145 ± 0.057 | 1.661 | −1.269 | 0.269 |
| Finnish | −0.299 ± 0.204 | 1.194 ± 0.296 | 0.105 ± 0.105 | ||||||||||||
| Mordovian | −0.255 ± 0.173 | 1.151 ± 0.246 | 0.104 ± 0.090 | ||||||||||||
| Russian | −0.303 ± 0.211 | 1.230 ± 0.301 | 0.072 ± 0.106 | ||||||||||||
Supplementary Material
Acknowledgments
We are grateful to Cynthia Beall, Neil Bradman, Amha Gebremedhin, Damian Labuda, Mari Nelis and Anna Di Rienzo for sharing DNA samples; to Detlef Weigel, Christa Lanz, Verena Schünemann, Peter Bauer and Olaf Riess for support and access to DNA sequencing facilities; to Philip Johnson for advice on contamination estimation; to Garrett Hellenthal for help with the ChromoPainter software; and to Pontus Skoglund for sharing graphics software. We thank Kenneth Nordtvedt for alerting us to newly discovered Y-chromosome SNPs. We downloaded the POPRES data from dbGaP at http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000145.v4.p2 through dbGaP accession number phs000145.v1.p2. We thank all the volunteers who donated DNA. We thank the staff of the Unità Operativa Complessa di Medicina Trasfusionale, Azienda Ospedaliera Umberto I, Siracusa, Italy for assistance in sample collection; and The National Laboratory for the Genetics of Israeli Populations for facilitating access to DNA. We thank colleagues at the Applied Genomics at the Children’s Hospital of Philadelphia, especially Hakon Hakonarson, Cecilia Kim, Kelly Thomas, and Cuiping Hou, for genotyping samples on the Human Origins array. JKr is grateful for support from DFG grant # KR 4015/1-1, the Carl-Zeiss Foundation and the Baden Württemberg Foundation. SP, GR, QF, CF, KP, SC and JKe acknowledge support from the Presidential Innovation Fund of the Max Planck Society. GR was supported by an NSERC fellowship. JGS acknowledges use of the Extreme Science and Engineering Discovery Environment (XSEDE), which is supported by NSF grant number OCI-1053575. EB and OB were supported by RFBR grants 13-06-00670, 13-04-01711, 13-04-90420 and by the Molecular and Cell Biology Program of the Presidium, Russian Academy of Sciences. BM was supported by grants OTKA 73430 and 103983. ASaj was supported by a Finnish Professorpool (Paulo Foundation) Grant. The Lithuanian sampling was supported by the LITGEN project (VP1-3.1-ŠMM-07-K-01-013), funded by the European Social Fund under the Global Grant Measure. AS was supported by Spanish grants SAF2011-26983 and EM 2012/045. OU was supported by Ukrainian SFFS grant F53.4/071. SAT was supported by NIH Pioneer Award 8DP1ES022577-04 and NSF HOMINID award BCS-0827436. KT was supported by an Indian CSIR Network Project (GENESIS: BSC0121). LS was supported by an Indian CSIR Bhatnagar Fellowship. RV, MM, JP and EM were supported by the European Union Regional Development Fund through the Centre of Excellence in Genomics to the Estonian Biocentre and University of Tartu and by an Estonian Basic Research grant SF0270177As08. MM was additionally supported by Estonian Science Foundation grant #8973. JGS and MS were supported by NIH grant GM40282. PHS and EEE were supported by NIH grants HG004120 and HG002385. DR and NP were supported by NSF HOMINID award BCS-1032255 and NIH grant GM100233. DR and EEE are Howard Hughes Medical Institute investigators. This project has been funded in part with federal funds from the National Cancer Institute, National Institutes of Health, under contract HHSN26120080001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government. This Research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research.
Footnotes
Supplementary Information is linked to the online version of the paper at www.nature.com/nature. The fully public version of the Human Origins dataset can be found at http://genetics.med.harvard.edu/reichlab/Reich_Lab/Datasets.html. The full version of the dataset (including additional samples) is available to researchers who send a signed letter to DR indicating that they will abide by specified usage conditions (SI9).
Author contributions
BB, EEE, JBu, MS, SP, JKe, DR and JKr supervised the study. IL, NP, AM, GR, SM, KK, PHS, JGS, SC, ML, QF, HL, CdF, KP, WH, MMet, MMey and DR analyzed genetic data. FH, EF, DD, MF, J-MG, JW, AC and JKr obtained human remains. AM, CE, RBo, KB, SS, CP, NR and JKr processed ancient DNA. IL, NP, SN, NR, GA, HAB, GBa, EB, OB, RBa, GBe, HB-A, JBe, FBe, CMB, FBr, GBJB, FC, MC, DECC, DCor, LD, GvD, SD, J-MD, SAF, IGR, MG, MH, BH, TH, UH, ARJ, SK-Y, RKh, EK, RKi, TK, WK, VK, AK, LL, SL, TL, RWM, BM, EM, JMol, JMou, KN, DN, TN, LO, JP, FP, OLP, VR, FR, IR, RR, HS, ASaj, ASal, EBS, ATar, DT, ST, IU, OU, RVa, MVi, MVo, CW, LY, PZ, TZ, CC, MGT, AR-L, SAT, LS, KT, RVi, DCom, RS, MMet, SP and DR assembled the genotyping dataset. IL, NP, DR and JKr wrote the manuscript with help from all co-authors.
The aligned sequences are available through the European Nucleotide Archive under accession number PRJEB6272.
The authors declare competing financial interests: UH is an employee of Illumina, TL is an employee of AMGEN, and JM is an employee of 23andMe.
References
- 1.Keller A, et al. New insights into the Tyrolean Iceman’s origin and phenotype as inferred by whole-genome sequencing. Nat Commun. 2012;3:698. doi: 10.1038/ncomms1701. [DOI] [PubMed] [Google Scholar]
- 2.Olalde I, et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature. 2014;507:225–228. doi: 10.1038/nature12960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Raghavan M, et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature. 2014;505:87–91. doi: 10.1038/nature12736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Skoglund P, et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science. 2012;336:466–469. doi: 10.1126/science.1216304. [DOI] [PubMed] [Google Scholar]
- 5.Bramanti B, et al. Genetic discontinuity between local hunter-gatherers and Central Europe’s first farmers. Science. 2009;326:137–140. doi: 10.1126/science.1176869. [DOI] [PubMed] [Google Scholar]
- 6.Haak W, et al. Ancient DNA from European early Neolithic farmers reveals their Near Eastern affinities. PLoS Biol. 2010;8:e1000536. doi: 10.1371/journal.pbio.1000536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lipson M, et al. Efficient moment-based inference of admixture parameters and sources of gene flow. Mol Biol Evol. 2013;30:1788–1802. doi: 10.1093/molbev/mst099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Patterson N, et al. Ancient admixture in human history. Genetics. 2012;192:1065–1093. doi: 10.1534/genetics.112.145037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Krause J, et al. A complete mtDNA genome of an early modern human from Kostenki, Russia. Curr Biol. 2010;20:231–236. doi: 10.1016/j.cub.2009.11.068. [DOI] [PubMed] [Google Scholar]
- 10.Sawyer S, Krause J, Guschanski K, Savolainen V, Pääbo S. Temporal patterns of nucleotide misincorporations and DNA fragmentation in ancient DNA. PLoS ONE. 2012;7:e34131. doi: 10.1371/journal.pone.0034131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Haak W, et al. Ancient DNA from the first European farmers in 7500-Year-old Neolithic sites. Science. 2005;310:1016–1018. doi: 10.1126/science.1118725. [DOI] [PubMed] [Google Scholar]
- 12.Perry GH, et al. Diet and the evolution of human amylase gene copy number variation. Nat Genet. 2007;39:1256–1260. doi: 10.1038/ng2123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Alexander DH, Novembre J, Lange K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 2009;19:1655–1664. doi: 10.1101/gr.094052.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS Genet. 2006;2:e190. doi: 10.1371/journal.pgen.0020190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Reich D, Thangaraj K, Patterson N, Price AL, Singh L. Reconstructing Indian population history. Nature. 2009;461:489–494. doi: 10.1038/nature08365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Moorjani P, et al. Genetic evidence for recent population mixture in India. Am J Hum Genet. 2013;93:422–438. doi: 10.1016/j.ajhg.2013.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Reich D, et al. Reconstructing Native American population history. Nature. 2012;488:370–374. doi: 10.1038/nature11258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Botigué LR, et al. Gene flow from North Africa contributes to differential human genetic diversity in southern Europe. Proceedings of the National Academy of Sciences. 2013 doi: 10.1073/pnas.1306223110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Cerezo M, et al. Reconstructing ancient mitochondrial DNA links between Africa and Europe. Genome Res. 2012;22:821–826. doi: 10.1101/gr.134452.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Moorjani P, et al. The history of African gene flow into southern Europeans, Levantines, and Jews. PLoS Genet. 2011;7:e1001373. doi: 10.1371/journal.pgen.1001373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Pickrell JK, Pritchard JK. Inference of population splits and mixtures from genome-wide Allele frequency data. PLoS Genet. 2012;8:e1002967. doi: 10.1371/journal.pgen.1002967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fu Q, et al. DNA analysis of an early modern human from Tianyuan Cave, China. Proc Natl Acad Sci USA. 2013;110:2223–2227. doi: 10.1073/pnas.1221359110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Bar-Yosef O. The chronology of the Middle Paleolithic of the Levant. New York: Plenum Press; 1998. pp. 39–56. [Google Scholar]
- 24.Armitage SJ, et al. The southern route “Out of Africa”: evidence for an early expansion of modern humans into Arabia. Science. 2011;331:453–456. doi: 10.1126/science.1199113. [DOI] [PubMed] [Google Scholar]
- 25.Rose JI, et al. The Nubian Complex of Dhofar, Oman: an African middle stone age industry in Southern Arabia. PLoS ONE. 2011;6:e28239. doi: 10.1371/journal.pone.0028239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Brace CL, et al. The questionable contribution of the Neolithic and the Bronze Age to European craniofacial form. Proc Natl Acad Sci U S A. 2006;103:242–247. doi: 10.1073/pnas.0509801102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Browning BL, Browning SR. Improving the Accuracy and Efficiency of Identity-by-Descent Detection in Population Data. Genetics. 2013;194:459–471. doi: 10.1534/genetics.113.150029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ralph P, Coop G. The geography of recent genetic ancestry across Europe. PLoS Biol. 2013;11:e1001555. doi: 10.1371/journal.pbio.1001555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lawson DJ, Hellenthal G, Myers S, Falush D. Inference of Population Structure using Dense Haplotype Data. PLoS Genet. 2012;8:e1002453. doi: 10.1371/journal.pgen.1002453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Brandt G, et al. Ancient DNA reveals key stages in the formation of central European mitochondrial genetic diversity. Science. 2013;342:257–261. doi: 10.1126/science.1241844. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Delsate D, Guinet JM, Saverwyns S. De l’ocre sur le crâne mésolithique (haplogroupe U5a) de Reuland-Loschbour (Grand-Duché de Luxembourg) ? Bull Soc Préhist Luxembourgeoise. 2009;31:7–30. [Google Scholar]
- 32.Rohland N, Hofreiter M. Ancient DNA extraction from bones and teeth. Nat Protocols. 2007;2:1756–1762. doi: 10.1038/nprot.2007.247. [DOI] [PubMed] [Google Scholar]
- 33.Dabney J, et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proceedings of the National Academy of Sciences. 2013;110:15758–15763. doi: 10.1073/pnas.1314445110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Stäuble HSfV-uFdUF. Häuser und absolute Datierung der Ältesten Bandkeramik. Habelt: 2005. [Google Scholar]
- 35.Yang DY, Eng B, Waye JS, Dudar JC, Saunders SR. Improved DNA extraction from ancient bones using silica-based spin columns. Am J Phys Anthropol. 1998;105:539–543. doi: 10.1002/(SICI)1096-8644(199804)105:4<539::AID-AJPA10>3.0.CO;2-1. [DOI] [PubMed] [Google Scholar]
- 36.Meyer M, Kircher M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb Protoc. 2010;2010 doi: 10.1101/pdb.prot5448. pdb prot5448. [DOI] [PubMed] [Google Scholar]
- 37.Meyer M, et al. A High-Coverage Genome Sequence from an Archaic Denisovan Individual. Science. 2012;338:222–226. doi: 10.1126/science.1224344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Briggs AW, et al. Removal of deaminated cytosines and detection of in vivo methylation in ancient DNA. Nucleic Acids Res. 2010;38:e87–e87. doi: 10.1093/nar/gkp1163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kircher M. Methods Mol Biol Vol. 840 Methods in Molecular Biology. 2012:197–228. doi: 10.1007/978-1-61779-516-9_23. [DOI] [PubMed] [Google Scholar]
- 40.Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–1760. doi: 10.1093/bioinformatics/btp324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.McKenna A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–1303. doi: 10.1101/gr.107524.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Maricic T, Whitten M, Pääbo S. Multiplexed DNA Sequence Capture of Mitochondrial Genomes Using PCR Products. PLoS ONE. 2010;5:e14004. doi: 10.1371/journal.pone.0014004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Behar Doron M, et al. A Copernican Reassessment of the Human Mitochondrial DNA Tree from its Root. Am J Hum Genet. 2012;90:675–684. doi: 10.1016/j.ajhg.2012.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Green RE, et al. A Complete Neandertal Mitochondrial Genome Sequence Determined by High-Throughput Sequencing. Cell. 2008;134:416–426. doi: 10.1016/j.cell.2008.06.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Fu Q, et al. A Revised Timescale for Human Evolution Based on Ancient Mitochondrial Genomes. Curr Biol. 2013;23:553–559. doi: 10.1016/j.cub.2013.02.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Fu Q. 2014 (in preparation) [Google Scholar]
- 47.Rasmussen M, et al. An Aboriginal Australian Genome Reveals Separate Human Dispersals into Asia. Science. 2011;334:94–98. doi: 10.1126/science.1211177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Vianello D, et al. HAPLOFIND: a new method for high-throughput mtDNA haplogroup assignment. Hum Mutat. 2013;34:1189–1194. doi: 10.1002/humu.22356. [DOI] [PubMed] [Google Scholar]
- 49.Tamura K, et al. MEGA5: Molecular Evolutionary Genetics Analysis using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol. 2011;28:2731–2739. doi: 10.1093/molbev/msr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Skoglund P, Storå J, Götherström A, Jakobsson M. Accurate sex identification of ancient human remains using DNA shotgun sequencing. J Archaeol Sci. 2013;40:4477–4482. [Google Scholar]
- 51.Drummond A, Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007;7:214. doi: 10.1186/1471-2148-7-214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Lippold S, et al. Human paternal and maternal demographic histories: insights from high-resolution Y chromosome and mtDNA sequences. bioRxiv. 2014 doi: 10.1101/001792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Green RE, et al. A Draft Sequence of the Neandertal Genome. Science. 2010;328:710–722. doi: 10.1126/science.1188021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Reich D, et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature. 2010;468:1053–1060. doi: 10.1038/nature09710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Prufer K, et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature. 2014;505:43–49. doi: 10.1038/nature12886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Li H, Durbin R. Inference of human population history from individual whole-genome sequences. Nature. 2011;475:493–496. doi: 10.1038/nature10231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Hach F, et al. mrsFAST: a cache-oblivious algorithm for short-read mapping. Nat Meth. 2010;7:576–577. doi: 10.1038/nmeth0810-576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.An integrated map of genetic variation from 1, 092 human genomes. Nature. 2012;491:56–65. doi: 10.1038/nature11632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Danecek P, et al. The variant call format and VCFtools. Bioinformatics. 2011;27:2156–2158. doi: 10.1093/bioinformatics/btr330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Li H. The sequence alignment/map (SAM) format and SAMtools. Bioinformatics. 2009;25:2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Keinan A, Mullikin JC, Patterson N, Reich D. Measurement of the human allele frequency spectrum demonstrates greater genetic drift in East Asians than in Europeans. Nat Genet. 2007;39:1251–1255. doi: 10.1038/ng2116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Price AL, et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 2006;38:904–909. doi: 10.1038/ng1847. [DOI] [PubMed] [Google Scholar]
- 63.Purcell S, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Alexander D, Lange K. Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinformatics. 2011;12:246. doi: 10.1186/1471-2105-12-246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Jakobsson M, Rosenberg NA. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics. 2007;23:1801–1806. doi: 10.1093/bioinformatics/btm233. [DOI] [PubMed] [Google Scholar]
- 66.Price AL, Zaitlen NA, Reich D, Patterson N. New approaches to population stratification in genome-wide association studies. Nat Rev Genet. 2010;11:459–463. doi: 10.1038/nrg2813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Busing FTA, Meijer E, Leeden R. Delete-m Jackknife for Unequal m. Statistics and Computing. 1999;9:3–8. [Google Scholar]
- 68.Loh PR, et al. Inferring Admixture Histories of Human Populations Using Linkage Disequilibrium. Genetics. 2013;193:1233–1254. doi: 10.1534/genetics.112.147330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Nelson MR, et al. The Population Reference Sample, POPRES: a resource for population, disease, and pharmacological genetics research. Am J Hum Genet. 2008;83:347–358. doi: 10.1016/j.ajhg.2008.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.