Abstract
Hyperactivity of a branch of the unfolded protein response in CD8α+ dendritic cells degrades endoplasmic reticulum–associated mRNAs, which leads to a defect in the cross-presentation of dead cell–derived antigens.
A subset of mouse dendritic cells (DCs), CD8α+ DCs, which have similarities to CD141+ human DCs, are particularly efficient at internalizing and cross-presenting dead cell–derived antigens to major histo-compatibility (MHC) class I molecules for the activation of CD8+ T cells. This process has been proposed to control infection by certain types of cytotoxic viruses and parasites, elicit antitumor immune responses and facilitate central and peripheral tolerance1. Therefore, understanding its regulation in both physiological and pathophysiological settings is an important goal. In this issue of Nature Immunology, Osorio et al. report that hyperactivity of the endoplasmic reticulum (ER) stress-reactive endonuclease IRE-1α causes a defect in the cross-presentation of dead cell–derived antigens by CD8α+ DCs2. The mechanism for this seems to involve the previously described process of regulated IRE-1α-dependent degradation (RIDD), whereby hyperactive IRE-1α, which normally functions to selectively process mRNA encoding the ER stress protein XBP-1, exerts its nuclease activity on ER-associated mRNAs other than Xpb1 mRNA3.
IRE-1α is an ER-spanning protein that is activated as part of the unfolded protein response (UPR) to ER stress. Its activation involves trans-autophosphorylation, nucleotide binding and homo-oligomerization, which are triggered by the accumulation of unfolded proteins in the ER4. In normal physiology, such modifications of IRE-1α activate its cytosolic endo-RNase domain, which selectively splices Xbp1 mRNA and thereby enables its translation into the transcriptionally active XBP-1 protein. XBP-1 induces many proteins that serve to relieve ER stress, including protein chaperones and proteins that facilitate the degradation of misfolded proteins. Treatment of Drosophila cells in vitro with drugs that activate a robust UPR results in not only the splicing of Xbp1 mRNA but also the endonuclease-mediated degradation of various mRNAs undergoing cotranslational translocation across the ER membrane3. Such mRNAs contain a consensus sequence in a stem-loop structure sensitive to IRE-1-mediated cleavage. The authors speculate that the RIDD process may work in concert with the translation-suppressing effect of the UPR to relieve ER stress3.
Studies subsequent to that initial report3 revealed that RIDD can be activated in cultured mammalian cells treated with ER stressors or chemical reagents that force the autophosphorylation and high-order oligomerization of IRE-1α. Moreover, as exemplified in the study of Osorio et al.2, IRE-1α and RIDD are hyperactivated when XBP-1, which is a negative regulator of IRE-1α, is targeted through genetic engineering5. Although the physiological relevance of such models of RIDD activation is uncertain (discussed below), studies of those models can reveal interesting biological effects. For example, in XBP-deficient plasma cells, RIDD cleaves mRNA encoding secretory immunoglobulin μ-chains, which leads to defective production of immunoglobulin M6, and in certain cell lines treated with experimental ER stress–inducing agents, RIDD-mediated degradation of microRNA has been linked to the activation of caspase-2 and apoptosis7. Evidence in vivo that RIDD is an important component of the physio logical UPR or contributes to chronic UPR-driven disease processes is lacking. However, IRE-1-mediated cleavage of individual mRNAs has been shown to have functional consequences under certain dietary conditions in mice or during development in Drosophila8,9.
Osorio et al. find that the combination of high basal IRE-1α activity in CD8α+ DCs plus the added IRE-1α-activating effect of deletion of XBP-1 leads to the hyperactivation of IRE-1α sufficient to activate RIDD2. By comparing cells in which XBP-1 is deleted with those in which IRE-1α is deleted (which also cannot generate XBP-1s protein), the investigators conclude that defective cross-presentation of antigens derived from dead cells, which is the main alteration in the phenotype of XBP-1-deficient CD8α+ DCs, is IRE-1α dependent and thus is probably due to RIDD. Interestingly, another alteration in their phenotype, expansion of the ER, is not IRE-1-α dependent and probably reflects a compensatory response to the deficiency in XBP-1-induced ER protein chaperones. Curiously, a published report has shown that DCs from lymphoid chimeric mice lacking XBP-1 have diminished survival10, but the mechanism for this has not been reported.
Pending the arrival of data to support the proposal of the physiological relevance of RIDD, findings obtained with experimental RIDD can be exploited to delineate various mechanisms of cell biology. In the present study, the selectivity of the defect in the cross-presentation of dead cell–derived antigens versus the cross-presentation of soluble antigens or presentation of endogenous antigens on MHC class I is very interesting. DCs can acquire dead cell–derived antigens by phagocytosis of apoptotic cells through a process called ‘efferocytosis’, which involves the recognition of apoptotic cells by cell-surface receptors, followed by the phagocytosis of those apoptotic cells and their degradation in phagolysosomes. DCs can also internalize cells that have undergone post-apoptotic necrosis or primary necrosis, but the mechanisms are usually different from those of efferocytosis. The proposal of the functional importance of DC-mediated cross-presentation of dead cell– derived antigens in host defense is supported by a variety of in vitro and in vivo studies, but definitive evidence of a molecular-genetic cause obtained by actual in vivo infection models is limited. Progress has been made in this area through the identification of a receptor, DNGR-1, that recognizes necrotic cells11 and a multiprotein complex of the receptor tyrosine kinase Axl, the lipoprotein receptor–related protein LRP1 (CD91) and the GTP-binding protein RanBP9 that mediates efferocytosis by CD8α+ DCs in vivo12. In both cases, virus-infected mice lacking those molecules show defective cross-presentation of viral antigen and greater susceptibility to viral infection.
Osorio et al. study the uptake, by DCs, of mouse embryonic fibroblasts (MEFs) treated with ultraviolet irradiation2, which causes apoptosis or necrosis depending on the time of exposure. That distinction may have mechanistic implications for how RIDD functions in CD8α+ DCs, because if the defect were relatively specific for efferocytosis, one set of candidates for RIDD substrates would be mRNAs encoding efferocytosis receptors and/or their adaptors. However, if cross-presentation of antigens from both apoptotic and necrotic cells were affected, the focus would be on molecules common to both processes, such as phagocytosis, or on molecules involved in the key downstream steps of cross-presentation, which include antigen processing, loading onto MHC class I and transport of antigen–MHC class I complexes to the cell surface13.
Investigators have proposed that DCs can use two cross-presentation pathways depending on the source of antigen, DC subtype and immunological setting. In the ‘cytosolic’ pathway, internalized and partially degraded antigens are exported to the cytosol, followed by proteasomal processing and loading onto MHC class I in the ER or in the endosomes (Fig. 1). Transport into the ER may be mediated by TAP (‘transporter associated with antigen processing’), which effects this function in the endogenous MHC class I pathway. In the ‘vacuolar’ (or ‘endocytic’) pathway, both antigen processing and loading on MHC class I occur within endosomes or phagosomes in a proteasome- and TAP-independent manner, with MHC class I recycled from the plasma membrane. In reality, this dichotomy is simplistic, and ‘hybrid’ pathways have been observed in certain experimental settings.
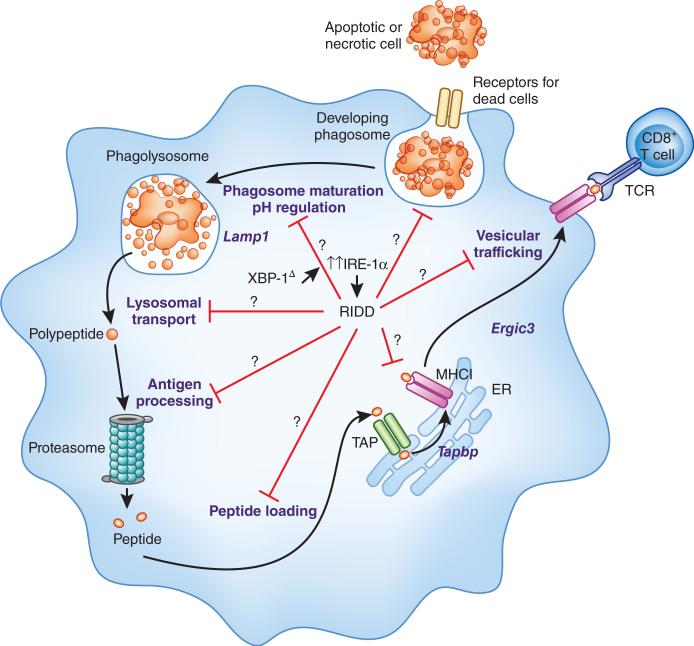
Figure 1.
Possible mechanisms by which RIDD compromises the cross-presentation of dead cell–derived antigens in CD8α+ DCs. Deletion of XBP-1 (XBP-1Δ) in CD8α+ DCs leads to hyperactivation of IRE-1α, which leads to the cleavage of multiple mRNAs by endonucleases (RIDD). The main consequence observed by Osorio et al. is defective cross-presentation of dead cell–derived antigens2. In theory, one or more mRNAs encoding proteins involved in the various stages of antigen cross-presentation (bold purple font) could be relevant targets of RIDD. Osorio et al. have identified several mRNA substrates of RIDD that could be involved2, such as Lamp1, Tapbp and Ergic3, but the roles of their products remain to be demonstrated. Moreover, the specificity for dead cell–derived antigens rather than soluble or endogenous antigens raises the possibility that one or more molecules involved in the uptake and/or processing of dead cells may be the relevant targets of RIDD. TCR, T cell antigen receptor; MHCI, MHC class I.
In this context, Osorio et al. first show that the export of an endocytosed, soluble reporter molecule to the cytosol is intact2, but it remains unknown what would happen if that molecule were delivered to the DC inside a dead cell; i.e., the condition in which cross-presentation becomes defective in XBP-1-deficient CD8α+ DCs. In a promising step forward, the authors find mRNA substrates for RIDD in CD8α+ DCs that encode proteins involved in antigen uptake, lysosomal processing and transport (Fig. 1). Examples include the cell-surface antigen transporter CD207 (langerin), the lysosomal protein LAMP-1, and TAP-binding protein, which mediates the interaction between TAP and MHC class I. Future studies of molecular-genetic causes will be needed to determine if loss of those or any other of the many mRNA substrates for RIDD identified in XBP-1-deficient CD8α+ DCs encode molecules with a role in defective cross-presentation. If molecules that mediate the uptake of dead cells are not the relevant substrates of RIDD in XBP-1-deficient CD8α+ DCs (as discussed above), future studies should determine why these cells have a selective defect in the cross-presentation of antigens derived from dead cells, because the candidate molecules mentioned by Osorio et al. have roles in the acquisition and cross- presentation of antigens in general2. Another important issue is whether the coexisting XBP-1-dependent and RIDD-independent ER-expansion phenotype functions in concert with RIDD to effect defective cross-presentation.
In closing, we return to the most important issue surrounding the RIDD field in general, and the study by Osorio et al.2 in particular: the physiological or patho physiological relevance of RIDD in natural settings in vivo. It is essential that future studies of RIDD address this issue, perhaps by investigating disease processes characterized by chronic, robust ER stress, such as certain types of neurodegenerative disease and advanced atherosclerosis. The type of hyperactivation of IRE-1α necessary for RIDD may be present in such settings. Moreover, it is possible that RIDD is activated in cells infected with certain types of viruses, in which IRE-1α is activated but the transcriptional activity of XBP-1 is blocked14. If future studies do identify a role for RIDD in natural disease settings in vivo, innovative work on strategies that disable RIDD in favor of Xbp1 mRNA splicing15 could have therapeutic benefit, including bolstering CD8α+ DC–mediated host defense based on the work of Osorio et al.2 presented here.
Footnotes
COMPETING FINANCIAL INTERESTS
The authors declare no competing financial interests.
References
- 1.Joffre OP, Segura E, Savina A, Amigorena S. Nat. Rev. Immunol. 2012;12:557–569. doi: 10.1038/nri3254. [DOI] [PubMed] [Google Scholar]
- 2.Osorio F, et al. Nat. Immunol. 2014;15:248–257. doi: 10.1038/ni.2808. [DOI] [PubMed] [Google Scholar]
- 3.Hollien J, Weissman JS. Science. 2006;313:104–107. doi: 10.1126/science.1129631. [DOI] [PubMed] [Google Scholar]
- 4.Ron D, Hubbard SR. Cell. 2008;132:24–26. doi: 10.1016/j.cell.2007.12.017. [DOI] [PubMed] [Google Scholar]
- 5.So JS, et al. Cell Metab. 2012;16:487–499. doi: 10.1016/j.cmet.2012.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Benhamron S, et al. Eur. J. Immunol. 2013 Nov 18; doi:10.1002/eji.201343953. [Google Scholar]
- 7.Upton JP, et al. Science. 2012;338:818–822. doi: 10.1126/science.1226191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Iqbal J, et al. Cell Metab. 2008;7:445–455. doi: 10.1016/j.cmet.2008.03.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Coelho DS, et al. Cell Rep. 2013;5:791–801. doi: 10.1016/j.celrep.2013.09.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Iwakoshi NN, Pypaert M, Glimcher LH. J. Exp. Med. 2007;204:2267–2275. doi: 10.1084/jem.20070525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Iborra S, et al. J. Clin. Invest. 2012;122:1628–1643. doi: 10.1172/JCI60660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Subramanian M, et al. J. Clin. Invest. (in the press) [Google Scholar]
- 13.Segura E, Villadangos JA. Traffic. 2011;12:1677–1685. doi: 10.1111/j.1600-0854.2011.01254.x. [DOI] [PubMed] [Google Scholar]
- 14.Tardif KD, Mori K, Kaufman RJ, Siddiqui AJ. Biol. Chem. 2004;279:17158–17164. doi: 10.1074/jbc.M312144200. [DOI] [PubMed] [Google Scholar]
- 15.Han D, et al. Cell. 2009;138:562–575. doi: 10.1016/j.cell.2009.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]