Abstract
Background
The obligate intracellular parasite Toxoplasma gondii establishes a life-long chronic infection within any warm-blooded host. After ingestion of an encysted parasite, T. gondii disseminates throughout the body as a rapidly replicating form during acute infection. Over time and after stimulation of the host immune response, T. gondii differentiates into a slow growing, cyst form that is the hallmark of chronic infection. Global transcriptome analysis of both host and parasite during the establishment of chronic T. gondii infection has not yet been performed. Here, we conducted a dual RNA-seq analysis of T. gondii and its rodent host to better understand host and parasite responses during acute and chronic infection.
Results
We obtained nearly one billion paired-end RNA sequences from the forebrains of uninfected, acutely and chronically infected mice, then aligned them to the genomic reference files of both T. gondii and Mus musculus. Gene ontology (GO) analysis of the 100 most highly expressed T. gondii genes showed less than half were shared between acute and chronic infection. The majority of the highly expressed genes common in both acute and chronic infection were involved in transcription and translation, underscoring that parasites in both stages are actively synthesizing proteins. Similarly, most of the T. gondii genes highly expressed during chronic infection were involved in metabolic processes, again highlighting the activity of the cyst stage at 28 days post-infection. Comparative analyses of host genes using uninfected forebrain revealed over twice as many immune regulatory genes were more abundant during chronic infection compared to acute. This demonstrates the influence of parasite development on host gene transcription as well as the influence of the host environment on parasite gene transcription.
Conclusions
RNA-seq is a valuable tool to simultaneously analyze host and microbe transcriptomes. Our data shows that T. gondii is metabolically active and synthesizing proteins at 28 days post-infection and that a distinct subset of host genes associated with the immune response are more abundant specifically during chronic infection. These data suggest host and pathogen interplay is still present during chronic infection and provides novel T. gondii targets for future drug and vaccine development.
Electronic supplementary material
The online version of this article (doi:10.1186/1471-2164-15-806) contains supplementary material, which is available to authorized users.
Keywords: Toxoplasma, RNA-seq, Transcriptome, Acute infection, Chronic infection, Forebrain, Metabolism, Immune response
Background
Toxoplasma gondii is an obligate intracellular parasite that can infect any nucleated cell of warm-blooded animals. The parasite has both sexual and asexual cycles where the sexual cycle takes place in the intestinal cells of the definitive feline host and the asexual cycle occurs in all warm-blooded animals [1]. The asexual stages of T. gondii consist of the rapidly replicating tachyzoite and the slow growing encysted bradyzoite. In the host, the tachyzoite is the prominent stage during initial acute infection [2]. Once the tachyzoite is subjected to stress from the host immune response, it differentiates to the bradyzoite form and eventually establishes a chronic infection [3]. The bradyzoite persists for the lifetime of the host as intracellular cysts present in striated muscle and the central nervous system [4].
T. gondii is one of the most prominent parasites in humans with prevalence rates between 10 and 80 percent worldwide, depending on the country [5]. Complications such as hydrocephaly, retinochoroiditis, mental retardation and even death can occur in developing fetuses [6, 7]. In immunocompetent humans, infection with T. gondii is generally asymptomatic presenting flu-like symptoms in approximately 10 percent of individuals [8]. Patients with compromised immune systems, such as those infected with HIV, are at great risk of developing severe symptoms such as Toxoplasmic Encephalitis (TE) and ocular infection that may result in blindness [9, 10]. Disease is largely associated with sporadic reactivation of the latent bradyzoite back to the rapidly replicating tachyzoite [11]. Currently there are no drugs that can combat the bradyzoite form of the parasite or effective vaccines to protect against infection. These issues highlight the critical need to understand the cellular triggers that control development between the tachyzoite and bradyzoite stages.
Previous work to characterize the transcriptome of tachyzoite and bradyzoites from T. gondii has primarily used microarray technology from samples prepared in tissue culture [12–14]. Transcriptomic studies have also used a combination of in vitro and in vivo samples with in vitro tachyzoites, in vivo bradyzoites, and oocysts collected from infected felines as well as tissue culture tachyzoites and bradyzoites, developing oocysts, and bradyzoites purified from mouse brains 21 days post-infection [12, 15] While these studies have provided valuable insight into this developmental process, the information that can be extracted is limited because tissue culture conditions for tachyzoite and bradyzoite development do not precisely model animal infections. Other in vivo microarray studies have compared peritoneal-derived tachyzoites from different strain types of T. gondii from wild type and interferon-γ (IFN-γ) deleted mice, and transcriptional changes in the brain of mice eight days after T. gondii infection [16–18]. These studies have highlighted important aspects of the developmental process, but the dynamic range of microarrays is restricted and sample preparations are unable to be simultaneously processed and analyzed for both host and pathogen. One way to overcome these limitations is RNA sequencing (RNA-seq), a breakthrough molecular tool that can provide the transcript profile (transcriptome) of total cellular RNA with a large dynamic range and improved sensitivity [19]. RNA-seq has detected novel T. gondii tachyzoite transcripts and alternative splicing between strains [20]. RNA-seq of tissue culture-derived bradyzoites has shown dysregulation of bradyzoite genes in the deletion mutant of a mucin domain containing cyst wall protein CST1 [21]. Using RNA-seq, transcriptome analysis has been performed to compare mouse brains that were uninfected or infected with T. gondii for 32 days [22]. These data highlight the sensitivity and depth of knowledge that can be obtained from RNA-seq studies; however, a time course of T. gondii infection and simultaneous analysis of the parasite transcriptome has not been performed.
To provide a more comprehensive analysis of T. gondii and the host during both acute and chronic infection, we collected RNA-seq data from three experimental groups of mice: uninfected, 10 and 28 days post-infection. Because T. gondii preferentially establishes cysts in the brains of mice and reactivation of cysts is the main cause of TE, we chose to analyze the brains of mice. A novel aspect of this dataset is that parasites were not purified from the brain tissue but instead, samples were rapidly processed so that RNA-seq reads represent the "interactome" between host and pathogen during the peak of acute and chronic infection. We report that many genes involved in T. gondii transcription, translation and metabolism are highly expressed during chronic infection. For the host, we find that more genes are increased in abundance during chronic versus (vs) acute infection, attesting to a continuously active host response even at 28 days post-infection.
Results
Sequencing and mapping the T. gondii/host interactome
To study parasite-host gene expression dynamics we used RNA-seq on samples collected from forebrains of mice infected with type II strains of T. gondii (schematized in Figure 1). Type II strains of T. gondii have been detected in the mouse brain as early as 4 days post-infection and numbers continue to increase until 10 days post-infection [23, 24], the peak of acute infection [25]. Cyst structures are present in the brains of mice at 21 days post-infection [26], which is generally considered to be the beginning of chronic infection. By 28 days post-infection, cysts have stably formed in the brain while parasite numbers have decreased elsewhere in the body [23, 27]. When brains were sectioned and analyzed for parasite distribution, high numbers of T. gondii were observed in the frontal lobe at 32 days post-infection [22]. We examined mouse brains at 10 and 28 days post-infection using an In Vivo Imaging System (IVIS), which confirmed that parasites were primarily localized in the forebrains of mice (Figure 1). Therefore to maximize parasite transcripts as well as to compare the same host tissue during acute and chronic infection, we chose to collect mouse forebrain samples at 10 and 28 days post-infection as well as uninfected mice. Nine mouse forebrains were sequenced individually: three uninfected, three infected for ten days and three infected for 28 days. Nearly one billion 100 base-pair (bp) paired-end RNA sequences were generated. Between 81,000,000 and 114,000,000 reads were obtained from each forebrain sample (Table 1). Between 69-76% of the reads aligned to the M. musculus reference genome while approximately 0.1% aligned to the T. gondii TGME49 reference. Since T. gondii was not purified from the forebrains as a means to rapidly process the samples and preserve the interactome, uninfected mouse forebrain samples were mapped to the T. gondii TGME49 reference to determine the extent of false positive reads. A small number of uninfected mouse reads aligned to the T. gondii genomic reference file (Table 1). The reads aligned to T. gondii ribosomal associated RNA or small (~200-300 bp) hypothetical proteins, none of which were considered differentially expressed and were treated as background.
Figure 1.
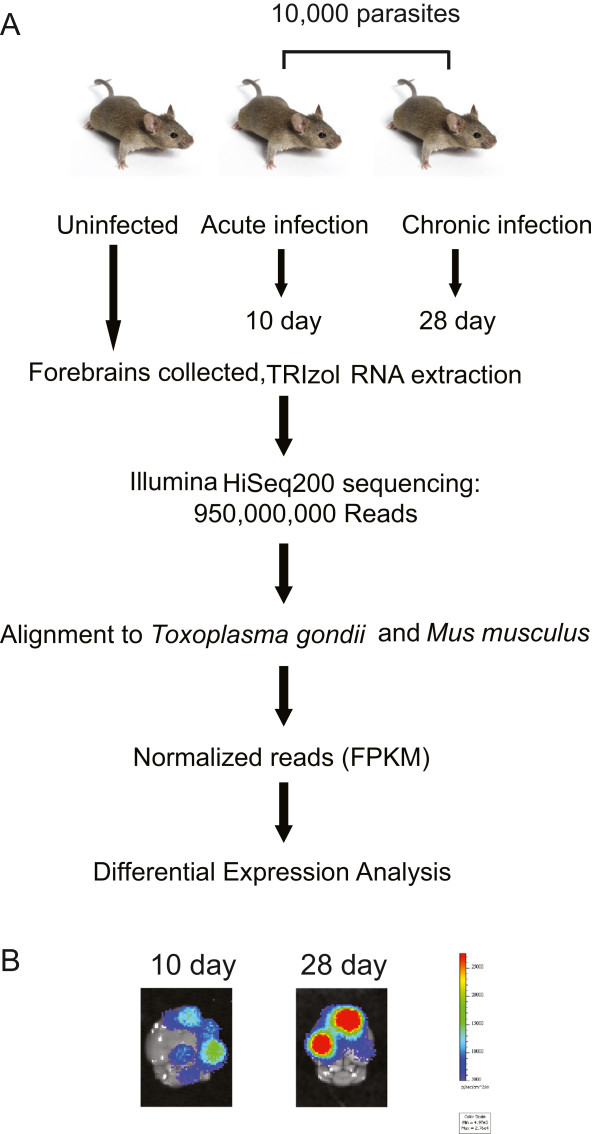
Schematic of T. gondii /host dataset generation. (A) Nine mice were divided into three experimental groups: uninfected, 10 days post-infection, and 28 days post-infection. Infected mice were given 10,000 type II ME49 parasites and sacrificed on the corresponding days. The forebrains were removed and homogenized in TRIzol, and RNA was extracted and purified. A cDNA library was generated from the RNA prior to IlluminaHiSeq2000 sequencing. Raw reads were aligned to either the T. gondii or M. musculus genomes, normalized and analyzed for differential gene expression. (B) To examine T. gondii in the brains of mice at the designated time points, mice were infected with 10,000 parasites of a bioluminescent T. gondii[62]. Shown are representative brains for 10 and 28 day post-infection mice, after the brains were soaked for 5 minutes in luciferin prior to imaging.
Table 1.
Mapped paired-end reads of individual mouse forebrain samples
| Days post-infection | Sample number | Total number of reads | mapped T. gondiipaired reads | mapped mouse paired reads |
|---|---|---|---|---|
| Uninfected | 1 | 112075860 | 10752 | 71901643 |
| Uninfected | 2 | 112948998 | 7272 | 72770895 |
| Uninfected | 3 | 103209252 | 12175 | 69295619 |
| 10 | 1 | 102581171 | 51734 | 68040784 |
| 10 | 2 | 125546828 | 51788 | 81691030 |
| 10 | 3 | 81630704 | 21234 | 47283570 |
| 28 | 1 | 103765423 | 90414 | 65891584 |
| 28 | 2 | 113094439 | 61481 | 69756643 |
| 28 | 3 | 114177191 | 89520 | 71828085 |
The first column is the time-point during T. gondii infection for the mouse forebrain samples used for RNA extraction. As three replicates were used for each time point, the second column is the sample number. The third column is the total number of reads obtained from Illumina HiSeq sequencing. The fourth column is the total number of paired-end reads from each mouse forebrain sample that mapped to the T. gondii genomic reference file. The fifth column is the total number of paired-end reads from each experimental group that mapped to the M. musculus genomic reference file.
Transcript abundance of T. gondiiduring acute and chronic infection in mice
Abundance estimates for each T. gondii gene were calculated and the three biological replicates for each time point were averaged before differential expression analysis was performed using Cuffdiff, a Cufflinks program. Cuffdiff calculates the fold change to determine which genes are differentially regulated between time points. Cuffdiff also calculates a p-value and q-value to determine if the fold change is significant. Each forebrain was treated as a biological replicate in Cuffdiff and therefore variation between replicates was considered when assigning a p-value. A p-value and q-value <0.05 were considered significant. It is important to note that parasites were not purified from mouse forebrains prior to RNA extraction. As a result, the concentration of T. gondii RNA could not be normalized between time points. To address the potential difference in parasite numbers between experimental time points quantitative PCR was performed using genomic DNA extracted from the forebrains at the time of RNA extraction. Using a standard curve generated from serial dilutions of genomic DNA extracted from a known number of parasites, we found approximately 500 parasites per 350 ng of genomic DNA in 10 day post-infection samples and approximately 1500 parasites per 350 ng of genomic DNA in the forebrains of mice infected for 28 days. These numbers are in agreement with the average number of reads generated from each time point with approximately twice as many reads aligning to the T. gondii genome in the 28 day samples than the 10 day samples (Table 1). To further examine differences in parasite numbers between acute and chronic infection samples, the FPKM fold change of housekeeping genes α-tubulin, actin, glyceraldehyde 3-phosphate dehydrogenase 1 and 2 (GAPDH 1 and 2), and hexokinase were examined between acute and chronic time points (Table 2). The fold change of these housekeeping genes between chronic and acute time points were 0.6-1.8, suggesting that global parasite transcript levels in our acute and chronic infection samples do not dramatically change.
Table 2.
Fold change between chronic and acute infection for previously characterized T. gondii genes
| Gene ID | Description | Fold change: chronic/acute |
|---|---|---|
| TGME49_316400 | α tubulin | 0.6 |
| TGME49_209030 | actin | 1.2 |
| TGME49_289690 | GAPDH 1 | 1.8 |
| TGME49_269190 | GAPDH 2 | 1 |
| TGME49_265450 | hexokinase | 0.7 |
| TGME49_291890 | MIC1 | 0.024 |
| TGME49_233460 | SAG1 | 0.0055 |
| TGME49_259020 | BAG1 | 48 |
| TGME49_268860 | ENO1 | 38 |
First column is the gene number from ToxoDB.org. The middle column is gene description where α-tubulin, actin, hexokinase, GAPDH-1 and 2 are housekeeping genes, tachyzoite specific genes are SAG1 and MIC1, and bradyzoite specific genes are BAG1 and ENO1. The third column is the average FPKM value for chronic genes divided by the average FPKM value for acute genes.
When comparing the fold change of previously characterized tachyzoite-specific genes, surface antigen 1 (SAG1) and microneme protein (MIC) 1, and bradyzoite specific genes bradyzoite antigen 1 (BAG1) and enolase 1 (ENO1), we saw large differential expression between acute and chronic infected mice (Table 2). These results along with qPCR performed on genomic DNA and quantitation of the reads that map to the T. gondii genome (Table 1) suggest that the significant differential expression of genes between time points is not due to an overabundance of transcripts in chronically infected mice compared to acutely infected mice. This data is supported in previous work that determined there is minimal increase in parasite numbers in the brain between 10 and 20 days [28]. To account for minimal variability of parasite numbers at 28 days post-infection differentially expressed T. gondii genes with a fold-change of >5 were considered for further analysis. The low abundance of SAG-1 and high abundance of BAG-1 between 28 day post-infection and 10 day post infection time points also suggests that tachyzoites are the primary stage present at 10 days post-infection, while the majority of parasites at 28 days post-infection are in the bradyzoite stage.
To explore the similarities in expression of T. gondii genes during acute and chronic infection, the 100 most highly expressed genes from acute and chronic time points were compared. Forty-two of the most highly expressed T. gondii genes during the acute stage were also among the 100 most highly expressed transcripts during the chronic stage (Figure 2A). To help interpret the biological functions of these genes, statistically over-represented GO terms were compiled (Figure 2B) using the Blast2GO program, a GO term analysis program for non-model organisms [29]. The GO term categories for the genes with the greatest abundance in both acute and chronic samples were transcription, translation, macromolecule biosynthesis and cellular metabolism.
Figure 2.
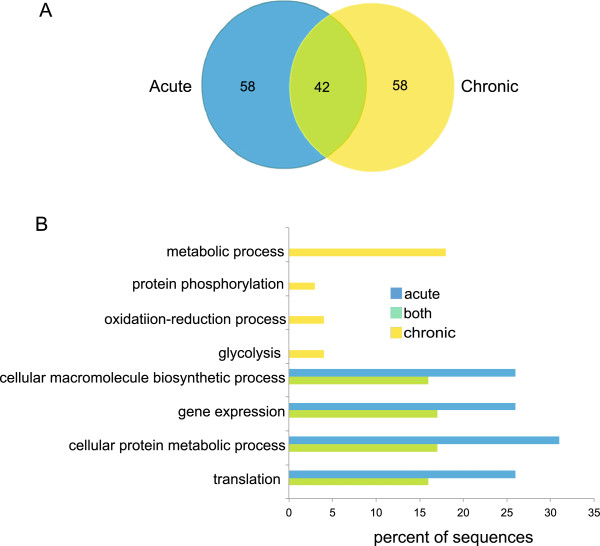
Top 100 T. gondii genes from acute and chronic infection. (A) Venn diagram compares the 100 T. gondii genes with the greatest FPKM value from acute and chronic time points. Of the top 100 genes for each stage, 42 genes were similarly abundant (green), and 58 were only abundant in acute (blue) or chronic (yellow) infection. (B) GO terms were assigned to the top 100 T. gondii genes from acute and chronic infection. Genes were grouped based on whether they are similar or different between time points prior to GO term analysis.
Differential expression of T. gondiibetween acute and chronic infection in mice
To further investigate the transcriptome of T. gondii during acute and chronic infection, differential expression analysis was performed. Fold change and significance values were calculated for each of the ~8900 annotated T. gondii genes (Additional file 1). From this analysis, we found 547 significantly differentially expressed genes (DEGs, with p-value and q-value <0.05) between acute and chronic infection. Of these, the DEGs with a fold change of >5 are presented in Tables 3 and 4.
Table 3.
T. gondii DEGs that were more abundant >5-fold in acute vs chronic infection
| Gene ID | Description | Fold change: acute/chronic |
|---|---|---|
| TGME49_233480 | SAG-related sequence SRS29C (SRS29C) | 305 |
| TGME49_233460 | SAG-related sequence SRS29B (SAG1) | 181 |
| TGME49_291890 | microneme protein MIC1 (MIC1) | 41 |
| TGME49_241240 | hypothetical protein | 40 |
| TGME49_224460 | aminopeptidase | 26 |
| TGME49_262050 | rhoptry kinase family protein ROP39 | 23 |
| TGME49_297880 | dense granule protein DG32 | 19 |
| TGME49_271050 | SAG 34A/SAG/2 | 18 |
| TGME49_294200 | glucose-6-phosphate 1-dehydrogenase | 13 |
| TGME49_230160 | hypothetical protein | 12 |
| TGME49_261740 | hypothetical protein | 12 |
| TGME49_291960 | rhoptry kinase family protein ROP40 | 10 |
| TGME49_268850 | enolase 2 | 9 |
| TGME49_277490 | hypothetical protein | 9 |
| TGME49_215960 | hypothetical protein | 9 |
| TGME49_222170 | dense-granule antigen DG32 | 8 |
| TGME49_293430 | hypothetical protein | 8 |
| TGME49_262730 | rhoptry protein ROP16 (ROP16) | 8 |
| TGME49_314400 | pyruvate dehydrogenase E1 component | 8 |
| TGME49_315320 | SAG-related sequence SRS52A (SRS52A) | 7 |
| TGME49_200360 | hypothetical protein | 7 |
| TGME49_285870 | SAG-related sequence SRS20A (SRS20A) | 7 |
| TGME49_269950 | hypothetical protein | 7 |
| TGME49_247280 | hypothetical protein | 7 |
| TGME49_294800 | elongation factor 1-alpha (EF-1-ALPHA) | 7 |
| TGME49_213050 | hypothetical protein | 6 |
| TGME49_249180 | bifunctional dihydrofolate reductase-thymidylate synthase | 6 |
| TGME49_226710 | hypothetical protein | 6 |
| TGME49_237880 | hypothetical protein | 6 |
| TGME49_250115 | hypothetical protein | 6 |
| TGME49_254720 | dense granule protein GRA8 (GRA8) | 6 |
| TGME49_253930 | GCC2 and GCC3 domain-containing protein | 6 |
| TGME49_299780 | hypothetical protein | 6 |
| TGME49_275860 | hypothetical protein | 6 |
| TGME49_310780 | dense granule protein GRA4 (GRA4) | 6 |
| TGME49_277080 | microneme protein MIC5 (MIC5) | 6 |
| TGME49_226960 | phosphofructokinase PFKII (PFKII) | 6 |
| TGME49_243730 | rhoptry protein ROP9 (ROP9) | 6 |
| TGME49_207400 | hypothetical protein | 6 |
| TGME49_259240 | ribosomal protein RPS21 (RPS21) | 6 |
| TGME49_205340 | ribosomal protein RPS12 (RPS12) | 6 |
| TGME49_209150 | non-proton pumping type-II NADH dehydrogenase I (NDH2-I) | 6 |
| TGME49_253690 | hypothetical protein | 5 |
| TGME49_229670 | ribosomal protein RPS23 (RPS23) | 5 |
| TGME49_244690 | hypothetical protein | 5 |
| TGME49_212290 | ribosomal protein RPS19 (RPS19) | 5 |
| TGME49_288360 | tryptophanyl-tRNA synthetase (TrpRS2) | 5 |
| TGME49_251810 | translation initiation factor eIF-5A | 5 |
| TGME49_200350 | subtilisin SUB3 (SUB3) | 18* |
| TGME49_250955 | KRUF family protein | 21* |
| TGME49_279350 | hypothetical protein | 23* |
| TGME49_313250 | hypothetical protein | 26* |
| TGME49_307760 | Tubulin-tyrosine ligase family protein | 27* |
| TGME49_315740 | SAG-related sequence SRS54 | 27* |
| TGME49_243700 | hypothetical protein | 29* |
| TGME49_293210 | hypothetical protein | 30* |
| TGME49_276110 | cytochrome b5 family heme/steroid binding | 34* |
| TGME49_218740 | membrane protein | 65* |
| TGME49_294805 | hypothetical protein | 77* |
| TGME49_294990 | hypothetical protein | 106* |
| TGME49_216770 | hypothetical protein | 125* |
| TGME49_230180 | hypothetical protein | 129* |
| TGME49_305050 | calmodulin | 295* |
First column is the gene number from ToxoDB.org. The second column is the gene description. The far right column is the average FPKM value from acute infection divided by the average FPKM value from chronic infection, called the fold change. * in the fold change column indicates the average FPKM value during chronic infection was 0, and could not be divided by the average FPKM value in acute samples. A p-value and q-value of <0.05 was considered to be statistically significant and only genes that met these standards were included on this table.
Table 4.
T. gondii DEGs that were more abundant >5 fold in chronic vs acute infection
| Gene ID | Description | Fold change: chronic/acute |
|---|---|---|
| TGME49_224630 | zinc finger (CCCH type) protein | 86 |
| TGME49_259020 | bradyzoite antigen BAG1 (BAG1) | 48 |
| TGME49_202020 | DnAK-TPR | 44 |
| TGME49_278080 | Toxoplasma gondii family A protein (SUSA-1) | 32 |
| TGME49_291040 | lactate dehydrogenase LDH2 (LDH2) | 31 |
| TGME49_200250 | microneme protein MIC17A (MIC17A) | 29 |
| TGME49_260190 | microneme protein MIC13 (MIC13) | 29 |
| TGME49_267680 | microneme protein MIC12 (MIC12) | 28 |
| TGME49_262970 | hypothetical protein | 26 |
| TGME49_245530 | hypothetical protein | 26 |
| TGME49_204420 | oocyst wall protein OWP1 (OWP1) | 23 |
| TGME49_318880 | hypothetical protein | 18 |
| TGME49_289370 | hypothetical protein | 18 |
| TGME49_207210 | hypothetical protein | 18 |
| TGME49_209985 | cAMP-dependent protein kinase | 18 |
| TGME49_309930 | melibiase subfamily protein | 17 |
| TGME49_320260 | hypothetical protein | 15 |
| TGME49_293780 | hypothetical protein | 14 |
| TGME49_216140 | tetratricopeptide repeat-containing protein | 14 |
| TGME49_280570 | SAG-related sequence SRS35A (SRS35A) | 13 |
| TGME49_320190 | SAG-related sequence SRS16B (SRS16B) | 11 |
| TGME49_250940 | hypothetical protein | 11 |
| TGME49_306620 | AP2 domain transcription factor AP2IX-9 (AP2IX9) | 11 |
| TGME49_209755 | hypothetical protein | 11 |
| TGME49_207160 | SAG-related sequence SRS49D (SRS49D) | 10 |
| TGME49_202030 | hypothetical protein | 10 |
| TGME49_312600 | heat shock protein HSP21 (HSP21) | 9 |
| TGME49_290000 | hypothetical protein | 9 |
| TGME49_256760 | pyruvate kinase PyK1 (PYKI) | 8 |
| TGME49_225290 | GDA1/CD39 (nucleoside phosphatase) | 7 |
| TGME49_269670 | hypothetical protein | 7 |
| TGME49_253330 | Rhoptry kinase family protein | 7 |
| TGME49_225540 | hypothetical protein | 7 |
| TGME49_282130 | hypothetical protein | 6 |
| TGME49_207710 | phosphatidylinositol synthase | 6 |
| TGME49_205680 | hypothetical protein | 6 |
| TGME49_276200 | hypothetical protein | 6 |
| TGME49_283780 | glucose-6-phosphate isomerase GPI (GPI) | 6 |
| TGME49_285980 | glucosephosphate-mutase GPM1 (GPM1) | 6 |
| TGME49_264420 | lipoprotein | 6 |
| TGME49_226420 | peptidase family M3 protein | 6 |
| TGME49_290980 | glycine C-acetyltransferase | 6 |
| TGME49_275320 | penicillin amidase | 5 |
| TGME49_201840 | aspartyl protease ASP1 (ASP1) | 5 |
| TGME49_246080 | NAD dependent epimerase/dehydratase | 5 |
| TGME49_222370 | SAG-related sequence SRS13 (SRS13) | 5 |
| TGME49_315760 | AP2 domain transcription factor AP2XI-4 (AP2XI4) | 5 |
| TGME49_294400 | hypothetical protein | 5 |
| TGME49_215910 | hypothetical protein | 5 |
| TGME49_224950 | calcium-dependent protein kinase CDPK5 (CDPK5) | 5 |
| TGME49_256060 | nucleosome assembly protein (nap) protein | 5 |
| TGME49_205750 | histone deacetylase complex subunit Sin3 (SIN3) | 5 |
| TGME49_253320 | hypothetical protein | 15* |
| TGME49_254330 | lipase | 16* |
| TGME49_309790 | hypothetical protein | 16* |
| TGME49_269300 | lipase | 17* |
| TGME49_223258 | hypothetical protein | 17* |
| TGME49_207875 | GCC2 and GCC3 domain-containing protein | 17* |
| TGME49_269020 | hypothetical protein | 18* |
| TGME49_261200 | TBC domain-containing protein | 18* |
| TGME49_254150 | hypothetical protein | 18* |
| TGME49_306500 | hypothetical protein | 18* |
| TGME49_245440 | hypothetical protein | 19* |
| TGME49_207980 | PIG-P protein | 19* |
| TGME49_220150 | 50S ribosomal protein L16 | 19* |
| TGME49_320720 | hypothetical protein | 19* |
| TGME49_268765 | hypothetical protein | 19* |
| TGME49_260530 | Sel1 repeat-containing protein | 22* |
| TGME49_269320 | hypothetical protein | 23* |
| TGME49_215300 | hypothetical protein | 24* |
| TGME49_308096 | rhoptry kinase family protein | 25* |
| TGME49_240470 | hypothetical protein | 25* |
| TGME49_460810 | ribosomal RNA | 25* |
| TGME49_310045 | hypothetical protein | 26* |
| TGME49_224180 | hypothetical protein | 26* |
| TGME49_205210 | hypothetical protein | 26* |
| TGME49_204040 | hypothetical protein | 27* |
| TGME49_297850 | Branched-chain-amino-acid aminotransferase | 27* |
| TGME49_215130 | adaptor-related protein complex 3 | 27* |
| TGME49_207600 | tubulin/FtsZ family, GTPase | 27* |
| TGME49_200230 | microneme protein MIC17C (MIC17C) | 28* |
| TGME49_231125 | hypothetical protein | 30* |
| TGME49_219610 | hypothetical protein | 31* |
| TGME49_240480 | cpw-wpc domain-containing protein | 31* |
| TGME49_315520 | calcium binding egf domain-containing protein | 32* |
| TGME49_232430 | hypothetical protein | 33* |
| TGME49_244412 | hypothetical protein | 33* |
| TGME49_223855 | RNA recognition motif-containing protein | 35* |
| TGME49_260325 | hypothetical protein | 37* |
| TGME49_234625 | EGF family domain-containing protein | 37* |
| TGME49_255460 | hypothetical protein | 40* |
| TGME49_321710 | IgA-specific serine endopeptidase | 43* |
| TGME49_319090 | proteasome maturation factor ump1 protein | 50* |
| TGME49_209090 | hypothetical protein | 52* |
| TGME49_210682 | hypothetical protein | 53* |
| TGME49_213480 | hypothetical protein | 53* |
| TGME49_210095 | hypothetical protein | 57* |
| TGME49_313890 | hypothetical protein | 58* |
| TGME49_271450 | hypothetical protein | 59* |
| TGME49_321700 | hypothetical protein | 80* |
| TGME49_219742 | hypothetical protein | 87* |
| TGME49_266600 | Kazal-type serine protease inhibitor | 89* |
| TGME49_257970 | hypothetical protein | 89* |
| TGME49_258370 | rhoptry kinase family protein ROP28 (ROP28) | 103* |
| TGME49_295662 | hypothetical protein | 110* |
| TGME49_278882 | GDA1/CD39 (nucleoside phosphatase) | 113* |
| TGME49_264150 | hypothetical protein | 141* |
First column is the gene number from ToxoDB.org. The far right column is the fold change calculated from the average FPKM value from chronic infection divided by the average FPKM value from acute infection. * in the fold change column indicates the average FPKM value during acute infection was 0, and could not be divided by the average FPKM value in chronic infection. A p-value and q-value of <0.05 was considered to be statistically significant and only genes that met these standards were included on this table.
Sixty-three T. gondii genes were >5 fold more abundant in acute compared to chronic infection (Table 3). SAG-related sequence (SRS) are a family of GPI-anchored surface antigens related to the first characterized T. gondii surface antigen, SAG1 [30]. SRS2/SRS29C was the most differently expressed gene in acute compared to chronic infection, 305-fold. Five additional SRS genes were >5 fold more abundant: SRS20A, SAG1/SRS29B, SAG2/SRS34A, SRS54 and SRS52A. Four genes for rhoptry proteins (ROP) were >5 fold more abundant during acute infection: ROP9, ROP16, ROP39, and ROP40. ROP16 is involved in decreased synthesis of cytokines in mouse bone marrow-derived macrophages [31]. ROP9 is a tachyzoite-specific protein with no known function [32]. ROP39 and ROP40 have homology to ROP2, but the functions have yet to be elucidated. Because of the extensive study of tachyzoites, most of the DEGs highly abundant in acute compared to chronic infection (Table 3) have been previously identified as tachyzoite-specific markers. Twenty-six of the 63 acute infection DEGs encode hypothetical proteins, with no homology to any annotated protein in the BLAST database. Characterizing these hypothetical proteins could be vital to understanding parasites during acute infection.
Fifty-one of the 107 T. gondii DEGs associated with chronic infection with a >5 fold change are annotated as hypothetical (Table 4). Among the chronic infection DEGs, four microneme proteins were identified MIC12, MIC13, MIC17A and MIC17C. MIC12 and MIC13 were previously shown to be bradyzoite specific [12], but our data revealed MIC17A and MIC17C abundant specifically during chronic infection. Another interesting group of chronic infection DEGs are those involved in glycolysis: glucose-6-phosphate isomerase, pyruvate kinase, lactate dehydrogenase 2, and glucosephosphate mutase. Previous data showed that tachyzoites and bradyzoites use the glycolytic pathway differently with lactate dehydrogenase 2 and pyruvate kinase being up-regulated during the bradyzoite stage [33, 34]. Our data shows that in addition to these two previously described bradyzoite-specific glycolytic enzymes, glucose-6-phosphate isomerase and glucosephosphate mutase are also more abundant in chronic vs acute infection. These results strengthen the idea that bradyzoites do not have an active TCA cycle because the transcripts of the key enzymes in the TCA cycle were less abundant in chronic infection compared to acute. Four SRS genes were also identified as more abundant in chronic compared to acute infection. SRS35A, also known as SAG4 and P18, is a long-known bradyzoite-specific marker [35]. SAG2C/SRS49D and SRS9/SRS16B are previously identified bradyzoite-specific genes that are important for persistence of infection [36–39]. SRS13 was identified as up-regulated in a microarray analysis comparing tissue culture tachyzoites to mouse-derived bradyzoites, but the function has yet to be determined [12]. Several novel chronic infection DEGs were found including DnAK-TPR, heat-shock protein 21, calcium dependent protein kinase CDPK5 (Table 4). The high number of novel and hypothetical DEGs highlights the fact that much is still unknown about T. gondii during animal infection. These hypothetical proteins have no known homology to proteins in the host and therefore could be essential and specific to parasite function during infection. Identification of novel DEGs that could play critical roles in T. gondii infection is the first step in elucidating potential targets for both vaccine and drug development.
To highlight the accuracy and sensitivity of RNA-seq, q-PCR was performed on a family of CCCH zinc fingers in T. gondii that are more abundant in our data set during chronic infection. TGME49_224630, TGME49_262970, and TGME49_311100 contain CCCH zinc finger domains and had a fold change of 86, 26, and 4.7, respectively (Table 4 excluding TGME49_311100 which did not meet the >5 fold cut-off). These genes were chosen based on the possible similarity in function and the range of transcript differential expression between chronic and acute infection. The increase in abundance of these transcripts between 10 day and 28 day post-infection was observed using qPCR when normalized to the house keeping gene tub1a. The fold change between chronic and acute infection for TGME49_224630, TGME49_262970, and TGME49_311100 were 53, 79, and 6.1 respectively. Not only does this data demonstrate the range and accuracy of RNA-seq, but also confirms the validity of the differential expression analysis.
Differential expression of host genes during T. gondiiinfection
To understand the transcriptional changes of the host during stages of T. gondii infection, differential expression analysis was conducted. Differential expression was determined between acute vs uninfected, chronic vs uninfected, and acute vs chronic time points. Genes were considered differentially expressed if the p-value and q-value was <0.05 and the fold change between time points was >2-fold. The host underwent extensive transcriptional changes during T. gondii infection (Figure 3A). When comparing acute T. gondii infected mice with uninfected, 1004 mouse genes were more abundant during acute infection and 143 were less abundant (Figure 3A). Over twice as many mouse genes, 2510, were more abundant in mice with a chronic T. gondii infection compared to uninfected while only 132 genes were less abundant. Finally, 1872 mouse genes were more abundant and 190 were less abundant in chronically vs. acutely infected mice. This increase of differentially regulated host genes during chronic infection is illustrated by Venn diagram analysis of DEGs >2-fold (Figure 3B). To understand the similarities and differences between DEGs more abundant during acute vs uninfected and chronic vs uninfected time points, we identified genes that had increased abundance at both time points as well as those found only in acute or chronic (Additional files 2, 3 and 4). Out of the 1004 DEGs more abundant in acute vs uninfected, 902 were also abundant in the chronic vs uninfected group. More mouse genes, 1608, were increased in abundance in the chronic vs uninfected time point that were not considered differentially expressed in acute. This data suggests genes activated during the peak of acute infection are maintained during chronic infection and that an entirely new subset of transcripts are expressed during chronic infection. The increase in differentially expressed genes during chronic infection could also be due to recruitment of cells to the site of infection.
Figure 3.
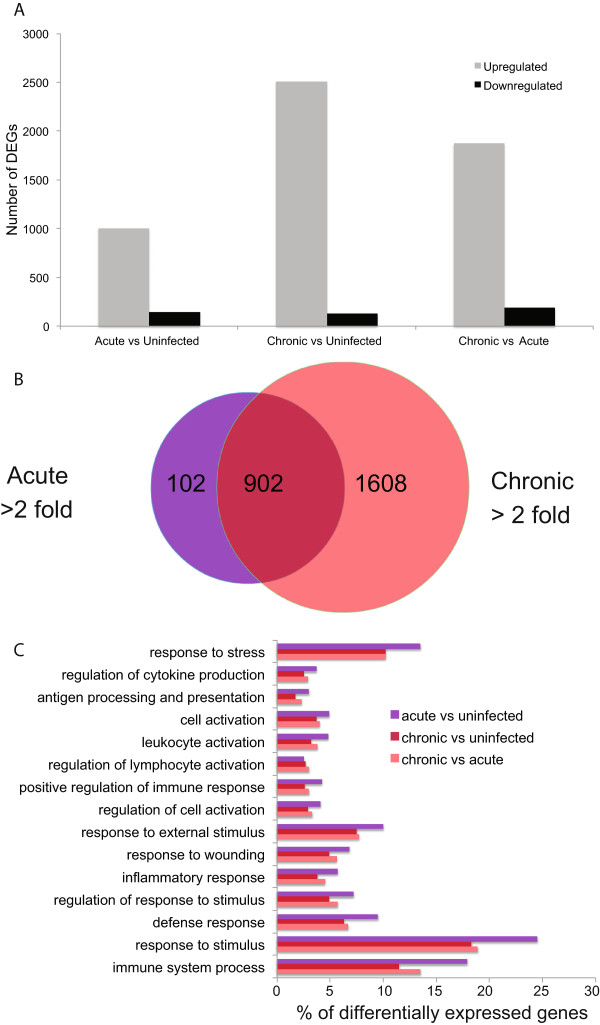
More host genes have increased abundance during chronic infection. (A) DEGs in the mouse with a fold change >2 were grouped based on increased abundance (grey) and decreased abundance (black) between acute vs uninfected, chronic vs uninfected, and chronic vs acute time points. (B) A Venn diagram was created to compare DEGs with increased abundance in acute vs uninfected (purple) and chronic vs uninfected time points (red). Of the 1004 more abundant DEGs in acute vs uninfected, 902 were also more abundant in the chronic vs uninfected group (magenta). (C) To explore the function of DEGs analyzed in the Venn diagram, a GO term enrichment analysis was performed.
GO term enrichment analysis of DEGs in mice during T. gondiiinfection
To understand the functions of DEGs in mice during acute and chronic T. gondii infection and to characterize the overlap between DEGs at these time points (Figure 3B) we performed GO term enrichment analyses. Among the host genes more abundant during both acute and chronic infection stages, overrepresented GO terms were related to stress and immune responses (Figure 3C). This suggests infection with T. gondii stimulates the immune response and/or immune cell recruitment into the brain even when the majority of parasites are in the encysted stage, as seen by the increased abundance of BAG1 and decreased abundance of SAG1 (Table 2). It also suggests a specific subset of host genes are responsible for immune stimulation during chronic infection that are distinct from acute, although many acute infection associated genes are still activated.
Only a small number of host DEGs were less abundant during both acute and chronic infection (Figure 4A). Go term analysis of DEGs with decreased abundance showed little functional overlap between the different stages of infection. The few commonalities in GO categories included secondary metabolism, membrane organization and phagocytosis (Figure 4B). The proposed functions of the DEGs that were less abundant specifically during acute infection were different from those less abundant during chronic infection. GO terms for genes with decreased abundance specifically during acute infection were proteolysis, protein metabolic process, and anatomical structure, while GO terms specific to chronic infection were cell-cell communication and primary metabolic function. The enrichment of metabolism-associated processes among less abundant DEGs suggests a link between T. gondii infection and host metabolism, possibly as a means of restricting parasite growth.
Figure 4.
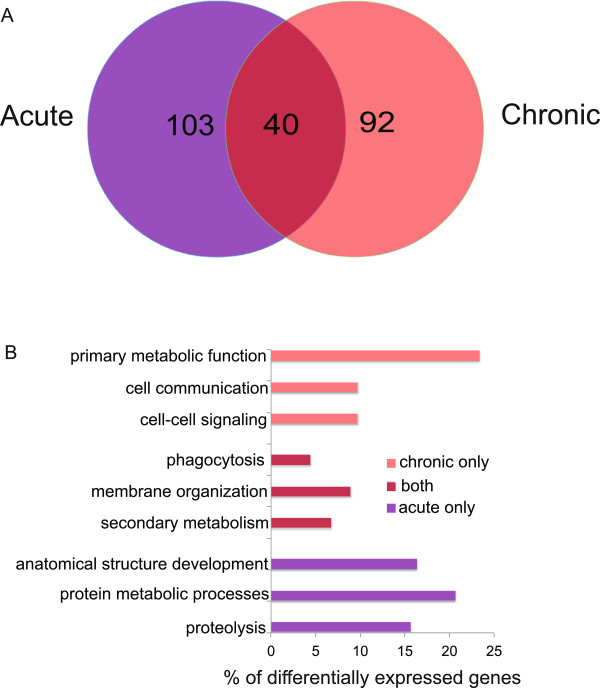
Analysis of host genes with decreased abundance during T. gondii infection. (A) A Venn diagram was created to compare DEGs >2-fold decreased abundance in acute vs uninfected (purple) and chronic vs uninfected (red) mice. Forty host genes were less abundant during both acute and chronic infection (magenta). (B) To explore the function of DEGs analyzed in the Venn diagram, a GO term enrichment analysis was performed. Functionally enriched GO terms display little similarity between time points.
Analysis of the mouse genes with increased abundance during T. gondiiinfection
Given that more host transcripts had >2-fold increased abundance during chronic infection (Figure 3B), further examination of the most abundant mouse genes during acute and chronic time points was performed to enhance the understanding of the host response. DEGs with a FPKM fold change >20 were compared between acute vs uninfected and chronic vs uninfected mice (Figure 5A). 155 genes met this cut-off in the acute vs uninfected group while 540 genes had 20-fold or higher FPKM values in chronic vs uninfected time points. Of these more abundant DEGs, 146 were shared between acute vs uninfected and chronic vs uninfected groups. Only 9 more abundant DEGs had a fold change >20 in acute vs uninfected that were not highly abundant in chronic vs uninfected time points. Several of these acute-infection specific host genes belong to the family of guanylate-binding proteins, which are GTPases that are induced by interferon-γ (Table 5). Conversely, 394 DEGs had a fold change of >20 in chronic vs uninfected that did not meet this cut-off in acute vs uninfected. The majority of genes increased in abundance during acute infection are maintained into chronic infection. These results show that few host genes are specifically increased during acute T. gondii infection. It also suggests a unique set of host genes are differentially expressed during chronic infection.
Figure 5.
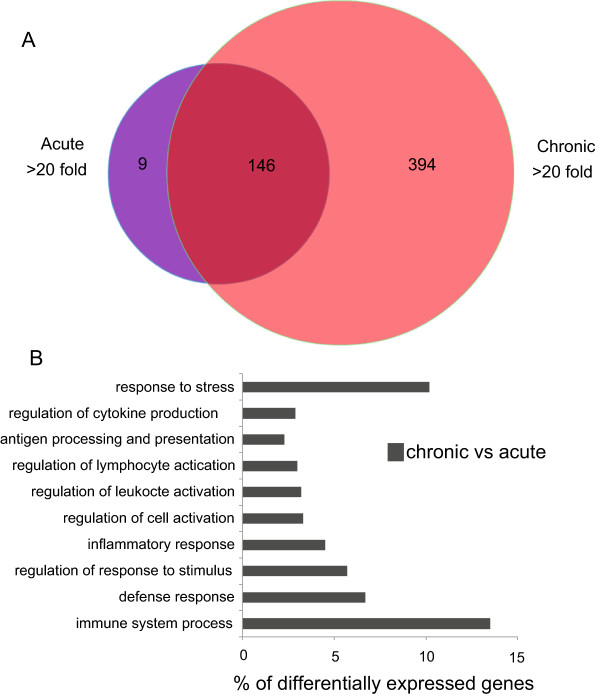
Few highly expressed DEGs are specific to acute infection. (A) To better characterize abundant DEGs, a Venn diagram of the more abundant DEGs in the mouse with a fold change >20 between acute vs uninfected (purple) and chronic vs uninfected (red) time points. Of these more abundant DEGs, 146 were shared between acute and chronic infection (magenta). Only 9 DEGs had a fold change >20 in acute vs uninfected that were also not highly abundant in chronic vs uninfected time points. (B) To analyze the function of the host more abundant DEGs in chronic vs acute time points, GO term enrichment analysis was performed. Many genes differentially expressed in chronic infection are associated with immune regulation and stress response.
Table 5.
Mouse DEGs more abundant in acute, but not chronic infection
| Gene ID | Gene name | Description | Fold change (Acute/uninfected) | Fold change (Chronic/uninfected) |
|---|---|---|---|---|
| ENSMUSG00000088071 | Gm22818 | predicted gene, 22818 | 276 | 1 |
| ENSMUSG00000002831 | Gbp10 | guanylate-binding protein 10 | 183 | No transcripts in Chronic |
| ENSMUSG00000002833 | Gbp6 | guanylate binding protein 6 | 130 | No transcripts in Chronic |
| ENSMUSG00000024334 | Gbp4* | guanylate binding protein 4 | 61 | No transcripts in Chronic |
| ENSMUSG00000075010 | Gbp8* | guanylate-binding protein 9 | 61 | No transcripts in Chronic |
| ENSMUSG00000085377 | Gbp9* | guanylate-binding protein 8 | 61 | No transcripts in Chronic |
| ENSMUSG00000098049 | BC042782# | cDNA sequence BC042782 | 52 | 1.8 |
| ENSMUSG00000079362 | n-R5s189# | nuclear encoded rRNA 5S 189 | 52 | 1.8 |
| ENSMUSG00000005800 | Plin4 | perilipin 4 | 27 | 1 |
The first column is the ensemble gene ID and the second is the gene name. The third column is the official gene symbol from ensemble.org, and the fourth column is the fold change as average acute FPKM value was divided by uninfected. The fifth column is the fold change as average chronic FPKM value was divided by uninfected. Only genes with a p-value and q-value <0.05 were considered differentially expressed. * and # mark genes located in the same contiguous loci and thus reads mapping to this area were combined.
Elevation of immune response genes during chronic infection
To assess the function of the mouse genes increased in abundance specifically during chronic infection, GO term enrichment was performed on genes with a fold change >20 in chronic vs acute infection. GO terms for genes highly increased in abundance during chronic infection were related to stress and immune responses (Figure 5B). Table 6 shows the top 50 DEGs more abundant during chronic infection. Many of these genes are immunoglobulin heavy chain variable regions, which share sequence similarity and can be difficult to differentiate between. The variable regions of immunoglobulins are responsible for specificity of antibodies to antigens, suggesting a different subset of antibodies being produced in response to T. gondii antigens that may not be abundant during acute infection. Another highly differentially expressed set of genes are H2-EB2 and H2-M2, both of which are involved in antigen presentation. H2-EB2 is an MHC class I membrane-associated protein while H2-M2 is an MHC class II membrane-associated protein. These antigen-presenting proteins are key players in the continued stimulation of the immune system at later time points. This increased in abundance of immune genes during chronic infection indicates a unique set of DEGs may be involved in the chronic infection immune response and/or a novel population of immune cells are recruited into the brain after acute infection.
Table 6.
The top 50 mouse DEGs more abundant in chronic vs. acute infection
| Gene ID | Description | Gene symbol | Fold change (chronic vs acute) |
|---|---|---|---|
| ENSMUSG00000076652 | immunoglobulin heavy variable 7-3 | Ighv7-3 | 672* |
| ENSMUSG00000076653 | immunoglobulin heavy variable 7-2 | Ighv7-2 | 672* |
| ENSMUSG00000095571 | immunoglobulin heavy variable 5-17 | Ighv5-17 | 672* |
| ENSMUSG00000076564 | immunoglobulin kappa chain variable 12-46 | Igkv12-46 | 656* |
| ENSMUSG00000096422 | immunoglobulin kappa variable 12-44 | Igkv12-44 | 656* |
| ENSMUSG00000076604 | immunoglobulin kappa joining 1 | Igkj1 | 353* |
| ENSMUSG00000076605 | immunoglobulin kappa joining 2 | Igkj2 | 353* |
| ENSMUSG00000076607 | immunoglobulin kappa joining 4 | Igkj4 | 353* |
| ENSMUSG00000076608 | immunoglobulin kappa joining 5 | Igkj5 | 353* |
| ENSMUSG00000076612 | immunoglobulin heavy constant gamma 2C | Ighg2c | 23* |
| ENSMUSG00000076617 | immunoglobulin heavy constant | Ighm | 23* |
| ENSMUSG00000076618 | immunoglobulin heavy joining 4 | Ighj4 | 23* |
| ENSMUSG00000076619 | immunoglobulin heavy joining 3 | Ighj3 | 23* |
| ENSMUSG00000076621 | immunoglobulin heavy joining 1 | Ighj1 | 23* |
| ENSMUSG00000094028 | immunoglobulin heavy diversity 4-1 | Ighd4-1 | 23* |
| ENSMUSG00000095079 | immunoglobulin heavy constant alpha | Igha | 23* |
| ENSMUSG00000095007 | immunoglobulin kappa chain variable 12-41 | Igkv12-41 | 356 |
| ENSMUSG00000076609 | immunoglobulin kappa chain, constant region | Igkc | 353 |
| ENSMUSG00000016283 | histocompatibility 2, M region locus 2 | H2-M2 | 111 |
| ENSMUSG00000002992 | apolipoprotein C-II | Apoc2 | 70 |
| ENSMUSG00000074336 | apolipoprotein C-IV | Apoc4 | 70 |
| ENSMUSG00000019987 | arginase, liver | Arg1 | 69 |
| ENSMUSG00000071716 | apolipoprotein L 7e | Apol7e | 60 |
| ENSMUSG00000058216 | cDNA sequence BC021614 | BC021614 | 45 |
| ENSMUSG00000042262 | chemokine (C-C motif) receptor 8 | Ccr8 | 44 |
| ENSMUSG00000014453 | B lymphoid kinase | Blk | 38 |
| ENSMUSG00000050063 | kallikrein related-peptidase 6 | Klk6 | 38 |
| ENSMUSG00000082976 | predicted gene 15056; similar to beta-defensin 52 | Gm15056 | 38 |
| ENSMUSG00000053977 | CD8 antigen, alpha chain | Cd8a | 37 |
| ENSMUSG00000030577 | CD22 antigen; hypothetical protein LOC100047973 | Cd22 | 36 |
| ENSMUSG00000020017 | histidine ammonia lyase | Hal | 35 |
| ENSMUSG00000027863 | CD2 antigen | Cd2 | 35 |
| ENSMUSG00000067149 | immunoglobulin joining chain | Igj | 35 |
| ENSMUSG00000030724 | CD19 antigen | Cd19 | 34 |
| ENSMUSG00000067341 | histocompatibility 2, class II antigen E beta2 | H2-Eb2 | 34 |
| ENSMUSG00000068129 | cystatin F (leukocystatin) | Cst7 | 33 |
| ENSMUSG00000079293 | C-type lectin domain family 7, member a | Clec7a | 32 |
| ENSMUSG00000096594 | immunoglobulin kappa variable 8-19 | Igkv8-19 | 32 |
| ENSMUSG00000002033 | CD3 antigen, gamma polypeptide | Cd3g | 31 |
| ENSMUSG00000050232 | chemokine (C-X-C motif) receptor 3 | Cxcr3 | 28 |
| ENSMUSG00000053044 | CD8 antigen, beta chain 1 | Cd8b1 | 28 |
| ENSMUSG00000054672 | RIKEN cDNA 5830411 N06 gene | 5830411N06Rik | 28 |
| ENSMUSG00000094738 | predicted gene, 26177 | Gm26177 | 28 |
| ENSMUSG00000005947 | integrin alpha E, epithelial-associated | Itgae | 27 |
| ENSMUSG00000022657 | CD96 antigen | Cd96 | 27 |
| ENSMUSG00000024669 | CD5 antigen | Cd5 | 27 |
| ENSMUSG00000024910 | cathepsin W | Ctsw | 27 |
| ENSMUSG00000031933 | folate receptor 4 (delta) | Folr4 | 27 |
| ENSMUSG00000026070 | interleukin 18 receptor 1 | Il18r1 | 26 |
| ENSMUSG00000035042 | chemokine (C-C motif) ligand 5 | Ccl5 | 25 |
The first column is the ensemble gene ID and the second is the gene name. The third column is the official gene symbol from ensemble.org, and the fourth column is the fold change as average chronic FPKM value was divided by acute. Only genes with a p-value and q-value <0.05 were considered differentially expressed. * in the fold change column indicates the average FPKM value during acute infection was 0, and could not be divided by the average FPKM value in chronic infection.
Discussion
Infection with T. gondii is often asymptomatic in immune competent individuals, but presents serious health risks if acquired congenitally or if a person becomes immune-compromised. Rising evidence is showing complications, such as psychiatric disorders and increased rates of suicide, occur in people with healthy immune systems who have a chronic T. gondii infection [40, 41]. Currently, drug treatment is only effective against the acute stage of infection and there are no therapeutics interventions available to target the encysted form during chronic infection. There are also no vaccines against T. gondii approved for use in humans. This lack of therapeutic intervention highlights the need to better understand the biological differences between the acute rapidly replicating form of the parasite and the chronic associated cyst stage.
Our data provides a list of candidate genes that could be targeted for novel therapeutics or gene deletion to create a non-persistent vaccine strain (Tables 3 and 4). Many of these genes highly differentially regulated are hypothetical proteins with no known orthologs in other organisms. Hypothetical T. gondii genes not found in the mammalian host could be excellent pathogen specific drug targets. The fact that multiple T. gondii microneme proteins are more abundant in chronic infection stages raises the question of whether parasites during chronic infection are actively invading cells. If parasites are invading at this time, this could explain the continued stimulation of the host response so late in infection. Although microarray analysis of human fibroblasts showed that tissue culture derived bradyzoites stimulated a weaker immune response than tachyzoites after two days [42]. An additional T. gondii gene that is more abundant in chronic infection is cAMP-dependent protein kinase. cAMP-dependent protein kinase is crucial for growth of tachyzoites and is proposed to be critical in tachyzoite to bradyzoite stage conversion [43, 44], but the mechanism is unknown. Another potentially interesting T. gondii gene that is more abundant in chronic infection is calcium dependent protein kinase 5, CDPK5. The T. gondii paralog, TgCDPK1, was shown to be necessary for tachyzoite motility, invasion and egress [45]. Studying the role of CDPK5 in bradyzoite development could be pivotal in understanding the biology of this stage.
Our study also provides crucial insight into host response to the parasite during both acute and chronic infection. It shows that, at least in the beginning stages of chronic infection, the host immune system is still actively combating infection. Chronic infection of T. gondii is typically thought of as a period in which the parasite transitions to the encysted, less immune stimulatory form resulting in a dampening of the immune response. Another study assessing the mouse transcriptome showed many immune associated genes are still expressed at 32 days post T. gondii infection [22]. To assess the overarching function of genes increased in abundance between acute and chronic infection Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was performed (Additional file 5). The KEGG database is a bioinformatics tool that assembles large-scale molecular datasets, such as gene lists, into biological pathway maps (example Additional file 6). Analysis of our dataset suggests active NK cells are recruited to the brain during chronic infection by the increased abundance of perforin, granzymes A and B, and IL-10 (additional files 3 and 4). NK cells have long been known to be essential for the control of acute T. gondii infection [46], but their role in chronic infection maintenance has yet to be elucidated. NK cells are a significant source of IFN-γ during acute T. gondii infection [47, 48] and IFN-γ is also necessary to maintain chronic infection [49], but producers of IFN-γ during chronic infection have not been determined. In mice with an established chronic T. gondii infection, NK cells are a major source of IFN-γ essential to combat infection with H5N1 influenza virus [50]. Chronic T. gondii infection was equally effective to protect against lethal influenza virus whether the mice had been infected with T. gondii for 1 month or 4 months, suggesting that NK cells are active in late stages of chronic infection. Similarly, NK are elicited in peritoneal exudate 6 months after T. gondii infection [51]. Our data suggests that NK cells play a role in chronic T. gondii infection maintenance, which will be the focus of future studies in understanding infection persistence.
Production of nitric oxide (NO) is crucial for control of T. gondii growth [52, 53], and triggers differentiation of tachyzoites into bradyzoites in tissue culture [54]. Mice deficient in the inducible nitric oxide synthetase gene succumb to non-lethal doses of T. gondii, but only during chronic infection [52, 53]. One host gene differentially expressed during chronic infection is arginase-1 (ARG-1), which had a fold change of nearly 70 in chronic vs. acute samples (Table 6). Arginine is not only a substrate for NO production, but it is an essential amino acid for T. gondii [55]. ARG-1 depletes host cell arginine, possibly as means to starve the parasite, but in type I strains, T. gondii initiates expression of ARG-1 via ROP16, potentially to preserve infected tissue [31]. While NO is detrimental for parasite growth, it also results in inflammation and subsequent destruction of host tissue. In the brain, microglial cells are the main producers of NO and have the potential to cause neuronal degradation. T. gondii-infected astrocytes secrete factors that decrease NO production by microglial cells, thus preserving both host and pathogen during latent infection [56]. It is unclear whether the increase in ARG-1 transcripts is induced by T. gondii or the host as ROP16 is more abundant in T. gondii during acute infection (Table 3), and ARG-1 is more abundant in the host during chronic infection. Furthermore, ROP16 is polymorphic and in type II strains, such as ME49 used in this study, does not maintain STAT3/6 activation and may not initiate ARG-1 expression [57]. An alternative mechanism may be responsible for the increased abundance of ARG-1 during chronic infection. Another highly differentiated host genes during chronic infection is kallikrein-6 peptidase. Kallikrein peptidases have been implicated in infection through involvement in vasodilation and permeability [58], but in the context of bacterial infections, kallikreins are also involved in the generation of NO [58, 59]. Specifically, kallikrein-6 has been shown to be up-regulated in the CNS during inflammation, possibly as a means to promote lymphocyte survival [60, 61]. Our data suggest kallikrein proteases could be involved in the parasite’s transition from the rapidly replicating form to the encysted form in the host. Together these data suggest that regulation of NO production during chronic infection is of vital importance for both the host and the parasites, and will be a future avenue of research.
Conclusions
The depth of RNA-seq coverage allowed, for the first time, simultaneous sampling of both host and microbe during acute and chronic stages of animal infection. In this study, we show the majority of highly expressed T. gondii genes common to both acute and chronic infection are involved in transcription and translation, underscoring that parasites in both stages are actively synthesizing proteins. Similarly, most of the T. gondii genes highly expressed during chronic infection are involved in metabolism, highlighting the metabolic activity of the cyst at 28 days post-infection. For the host, analysis of transcripts at 10 and 28 days post-infection compared to uninfected mice showed that more immunity associated host genes are increased in abundance at 28 days post-infection vs 10 days post-infection. The increase in abundance of T. gondii genes during chronic infection is in conjunction with the heightened host response; indicative of the constant battle for survival between the host and the parasite. Discussed here are only a few examples of hypotheses that can be generated from this transcriptome data set. This dataset is novel because information from the host and pathogen is provided at multiple time points, allowing for the interplay between both to be studied. Many platforms, such as KEGG pathways and DAVID, are available for the research community to further investigate these data and cater to their scientific interests. This data provides the potential to elucidate mechanisms required for Apicomplexan parasites to maintain a relationship with their hosts, which will lead to better therapeutics, vaccines and diagnostic methods.
Methods
Ethics statement
Animals were housed under conventional, specific-pathogen-free conditions and were treated in compliance with guidelines set by the Institutional Animal Care and Use Committee of the University of Wisconsin School of Medicine and Public Health (IACUC), according to IACUC approved protocol number M01545. This protocol adheres to the regulations and guidelines set by the National Research Council. The University of Wisconsin is accredited by the International Association for Assessment and Accreditation of Laboratory Animal Care.
IVIS detection of T. gondiiin the mouse forebrain at 10 days and 28 days post-infection
The ME49 strain of T. gondii with a deletion of the gene HPT and an insertion of the coding region for firefly luciferase, as previously described [62], was used for these experiments. 6–8 week old BALB/C mice (National Cancer Institute, Charles River Laboratories, Frederick, MD) received an intraperitoneal (i.p.) inoculation of 104 freshly lysed tachyzoites. Mice were imaged using IVIS (PerkinElmer) at 10 days and 28 days post-infection. Mice were anesthetized with isoflurane and intravenous (i.v.) injected with 3 mg of luciferin, the substrate for luciferase, and imaged ventrally, dorsally, and then sacrificed. The brains of the mice were removed and soaked in luciferin for 5 minutes prior to imaging.
Generation of mRNA and RNA-seq
A ME49 strain of T. gondii that was recently passaged through the sexual cycle was used to inoculate mice for RNA-seq analysis. ME49 was maintained as tachyzoites in monolayers of Human Foreskin Fibroblasts in Dulbecco’s Modified Eagle’s Medium supplemented with 10% FBS, 2 mM L-glutamine, and 1% penicillin-streptomycin. 6–8 week old CBA/J mice (National Cancer Institute, Charles River Laboratories, Frederick, MD) were either left uninfected or i.p. injected with 104 parasites and were sacrificed at 10 days and 28 days post-infection. Uninfected mice were sacrificed along with the 28 day post-infection group. We selected T. gondii infected mice that were healthy and not displaying any signs of disease, so samples would not contain host transcripts involved with inappetence, dehydration or general malaise to confound our analyses. To minimize changes to the transcriptome, the forebrains were rapidly and precisely sectioned at the intersection of the optic nerves using a mouse brain matrix (Zivic Instruments) with less than one minute between animal sacrifice to forebrain homogenization in 3 mL of TRIzol. Total RNA was isolated according to manufacturer’s protocol. RNA was purified using Promega SV total RNA isolation system according to manufacturer’s protocol. RNA was submitted to the University of Wisconsin Biotechnology Center for purity analysis using the Agilent 2100 Bioanalyzer and sequencing using the Illumina HiSeq2000. Sequencing was performed on each individual mouse and samples were not pooled. Infection was quantified in the un-used hindbrains collected at 28 days post-infection and stained with fluorescein labeled Dolichos biflorus agglutinin (Vector Laboratories) for cyst detection. All 28 day post-infection hindbrains contained a minimum of 10,000 cysts.
Determination of T. gondiiparasite numbers in mouse forebrain samples
Genomic DNA was extracted from each mouse forebrain at the time of RNA extraction using TRIzol according to manufactures instructions. DNA was purified by phenol/chloroform extraction followed by ethanol precipitation. Genomic DNA was used as the template for quantitative PCR using T. gondii primers for the housekeeping gene alpha-tubulin (TUB1A). Tub1A Forward primer 5’-GACGACGCCTTCAACACCTTCTTT-3’, Tub1A Rev 5’-AGTTGTTCGCAGCATCCTCTTTCC-3’. Primer efficiency for TUB1A was 2.002 with an R2 value of .99 using T. gondii genomic DNA. To determine parasite burden in the mouse forebrain samples, a standard curve was generated using a genomic DNA preparation of known parasite numbers. Quantitative PCR was performed on serial dilutions of parasite genomic DNA, using TUB1A primers, ranging from 10 to 1×106 parasites. A best-fit logarithmic line was generated with an R2 of 0.999. The equation of the line along with Ct values obtained from qPCR of TUB1A on genomic DNA from each forebrain sample was used to extrapolate parasite numbers. qPCR was performed on each sample in duplicate using BIO-RAD iTaq Universal SYBR Green Supermix product number 172–5121.
Quantitative PCR of T. gondiiCCCH zinc fingers
Sequences for TGME49_224630, TGME49_269270, and TGME49_311100 were obtained from ToxoDB.org. Sequences were run through BLASTp to confirm presence of CCCH zinc finger motifs. cDNA was generated from the same RNA samples used for RNA sequencing with Invitrogen Superscript III Reverse Transcriptase cDNA synthesis kit. All CCCH zinc fingers were normalized to the T. gondii house keeping gene tub1A. Efficiencies were determined using in vitro bradyzoite cDNA. RNA was extracted from 5 day bradyzoites grown under low CO2 and high pH conditions using TRIzol. cDNA was generated using the Invitrogen Superscript III Reverse Transcriptase cDNA synthesis kit. Efficiencies were calculated using the slopes of a 1:10 dilution series (neat through 104) and the formula E = 10[-1/slope]. Efficiencies for tub1A, TGME49_224630, TGME49_269270, and TGME49_311100 were 1.96, 1.89, 2.14, and 2.04 (Between 95-107% efficient). Quantitative PCR was performed using Bio-Rad iTaq Universal SYBR Green Supermix on an Applied Biosystems StepOnePlus Real-Time PCR system. Primers were used at a 300 nM concentration and an extension temperature of 60°C for 60 seconds for all primer sets except TGME49_224630 which was run at 56.5°C for 60 seconds. Relative quantification was calculated using Pfaffl’s method [63]. The three biological replicates were used and conducted in duplicate. Wells with only one melt curve and temperature were used, and duplicate Ct values were all at or below a 0.25 difference in cycle threshold value. Primers: Tub1A, same as previously described above
TGME49_224630 Forward 5’-GCGAGGATGAGTGTGGG-3’
Reverse 5’- AGGCGTCACCGTTTGG-3’
TGME49_269270 Forward 5’- GCTTACCGAGGATGACCTGCT-3’
Reverse 5’-CCGTACACTGGTGGCGATCAT-3’
TGME49_311100 Forward 5’- TTTGCCCACACAGCCGAAGAA-3’
Reverse 5’- GCCACAGATGCCTTCCGTAAC-3’.
RNA-seq, mapping and differential expression analysis
Approximately 950,000,000 paired end 100 bp reads were generated from Illumina HiSeq2000 sequencing. Aligning RNA-seq data for eukaryotic organisms becomes difficult when mapping to a genomic reference because of the presence of introns in the reference and polyadenylated transcripts in the data. If an RNA-seq read spans an exon-exon junction or a polyadenylated region of a transcript it will be "unmappable" to the reference genome and is discarded. Bioinformatics software, such as TopHat, has been created to consider exon-exon boundaries during the mapping process [64]. Raw reads were uploaded onto the Galaxy platform [65–67]. Reads were filtered by Sanger quality score using FASTQ Groomer v. 1.0.4 and paired end reads were aligned against the genomes of T. gondii (TGME49 version 9.0; ToxoDB.org) and M. musculus (GRCm38 version 74.38; ensemble.org/Mus_musculus) references uploaded into Galaxy using TopHat2 [68]. Parameters for TopHat2: Max edit distance of 2, final read mismatch of 2, anchor length of 8, minimum intron length of 70, maximum intron length of 500000, max insertion and deletion length of 3, number of mismatches allowed of 2, and a minimum length of read segments of 25. Reads were not treated as strand specific as they were paired end reads. The total numbers of reads were as followed: Uninfected mouse 1 was 112075860, Uninfected mouse 2 was 112948998, Uninfected mouse 3 was 103209252, 10 day post-infection mouse 1 was 102581171, 10 day post-infection mouse 2 was 125546828, 10 day post-infection mouse 3 was 81630704, 28 day post-infection mouse 1 was 103765423, 28 day post-infection mouse 2 was 113094439, and 28 day post-infection mouse 3 was 114177191. The number of reads that mapped to the T. gondii genomic reference file and M. musculus genomic reference files are listed in Table 1.
The program Cufflinks [64] was used to convert aligned reads of BAM files, generated from Tophat2, into relative expression values for each gene represented as FPKM (fragments per kilobase of exon per million mapped reads). Cuffdiff was used to detect significant changes in differential expression between the experimental groups. When running Cuffdiff, a GTF file obtained from ToxoDB.org and ensemble.org/Mus_musculus was used as a guide and the TopHat2 aligned BAM files from each biological replicate were used as the source of comparison between experimental groups. A geometric library normalization and a pooled cross-replicate dispersion estimation method was used when comparing differential expression between each experimental group. Genes with a p-value, for statistical significance, and q-value, to detect the false discovery rate, of <0.05 were considered differentially expressed. The "gene differential testing" output file from Cuffdiff was used to identify differentially expressed genes.
GO term analysis
FASTA sequences of the most abundant T. gondii genes from different time points during infection were loaded into the program Blast2Go [29]. T. gondii is not an available organism on many GO term analysis programs, making Blast2Go ideal for uncommon models. Sequences were run with the blastx program against the nr database. Aligned sequences were mapped and assigned GO term annotations. Combined graphs were generated representing the most enriched GO terms in the provided gene list. A score was assigned to determine significance of enrichment. For analysis of M. musculus, a gene list was generated from each experimental time point based on gene names in column C of Additional files 2, 3, and 4. Ensembl gene IDs were obtained from these gene names, exported from BioMart and uploaded to the functional annotation tool. A functional annotation chart of the enriched GO terms was generated using GO terms associated with biological process. GO terms were assigned a p-value to indicate significance of enrichment [69]. Only GO terms with a p-value <0.05 were used to represent functional enrichment. For further analysis of M. musculus, the online Database for Annotation, Visualization, and Integrated Discovery (DAVID) was used (david.abcc.ncifcrf.gov).
Availability
Raw short read RNA-seq data has been submitted to ToxoDB and to NCBI with SRA: SRS550800, http://www.ncbi.nlm.nih.gov/biosample/2615816.
Electronic supplementary material
Additional file 1: Differential expression analysis of T. gondii genes between acute and chronic time points generated by Cuffdiff. (XLS 2 MB)
Additional file 2: Differential expression analysis of M. musculus genes between acute vs. uninfected time points generated by Cuffdiff. (XLS 16 MB)
Additional file 3: Differential expression analysis of M. musculus genes between chronic vs. uninfected time points generated by Cuffdiff. (XLSX 9 MB)
Additional file 4: Differential expression analysis of M. musculus genes chronic divided by acute time points generated by Cuffdiff. (XLS 17 MB)
Additional file 5: Top 5 significant KEGG pathways during chronic vs acute infection. Genes with a >2 fold abundance between chronic and acute infection were uploaded to the KEGG pathway database. Displayed above are the most 5 most significant pathways enriched during chronic infection. The first column is the KEGG pathway description. Second column is the number of genes that fall into each category. The third column is the p-value designating the significance of each category. (PDF 151 KB)
Additional file 6: NK cell mediated cytotoxicity KEGG pathway. Schematic representation of NK cell mediated cytotoxicity provided by Kyoto Encyclopedia of Genes and Genomes ( http://www.genome.jp/kegg/). Red stars indicate genes that are more abundant between chronic and acute infection in the mouse forebrain. For description of features on the map visit http://www.genome.jp/kegg/document/help_pathway.html. (PDF 295 KB)
Acknowledgements
We sincerely thank Colin Dewey for assistance with data analysis, David Sibley for low-passage ME49 and Kyle Boldon for assistance with tissue processing. This research was supported by the University of Wisconsin Food Research Institute and the American Cancer Society #RSG-07-202-01-MBC (L.J.K.), and an NSF fellowship number DGE-0718123 (K.J.P.).
Footnotes
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
MTA and LJK designed and performed the mouse studies and collected the RNA. KJP performed all of the data analysis and prepared all of the figures. KJP and LJK wrote the manuscript. All authors have read and edited the manuscript.
Contributor Information
Kelly J Pittman, Email: kjpittman@wisc.edu.
Matthew T Aliota, Email: mtaliota@wisc.edu.
Laura J Knoll, Email: ljknoll@wisc.edu.
References
- 1.Dubey JP. Advances in the life cycle of Toxoplasma gondii. Int J Parasitol. 1998;28(7):1019–1024. doi: 10.1016/S0020-7519(98)00023-X. [DOI] [PubMed] [Google Scholar]
- 2.Black MW, Boothroyd JC. Lytic cycle of Toxoplasma gondii. Microbiol Mol Biol Rev. 2000;64(3):607–623. doi: 10.1128/MMBR.64.3.607-623.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Skariah S, McIntyre MK, Mordue DG. Toxoplasma gondii: determinants of tachyzoite to bradyzoite conversion. Parasitol Res. 2010;107(2):253–260. doi: 10.1007/s00436-010-1899-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Weiss LM, Kim K. The development and biology of bradyzoites of Toxoplasma gondii. Front Biosci. 2000;5:D391–D405. doi: 10.2741/Weiss. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Robert-Gangneux F, Darde ML. Epidemiology of and diagnostic strategies for toxoplasmosis. Clin Microbiol Rev. 2012;25(2):264–296. doi: 10.1128/CMR.05013-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Roizen N, Swisher CN, Stein MA, Hopkins J, Boyer KM, Holfels E, Mets MB, Stein L, Patel D, Meier P, Withers S, Remington J, Mack D, Heydemann PT, Patton D, McLeod R. Neurologic and developmental outcome in treated congenital toxoplasmosis. Pediatrics. 1995;95(1):11–20. [PubMed] [Google Scholar]
- 7.McAuley J, Boyer KM, Patel D, Mets M, Swisher C, Roizen N, Wolters C, Stein L, Stein M, Schey W, Remington J, Meier P, Johnson D, Heydemann P, Holfels E, Withers S, Mack D, Brown C, Patton D, McLeod R. Early and longitudinal evaluations of treated infants and children and untreated historical patients with congenital toxoplasmosis: the Chicago Collaborative Treatment Trial. Clin Infect Dis. 1994;18(1):38–72. doi: 10.1093/clinids/18.1.38. [DOI] [PubMed] [Google Scholar]
- 8.Montoya JG, Liesenfeld O. Toxoplasmosis. Lancet. 2004;363(9425):1965–1976. doi: 10.1016/S0140-6736(04)16412-X. [DOI] [PubMed] [Google Scholar]
- 9.Ong EL. Common AIDS-associated opportunistic infections. Clin Med. 2008;8(5):539–543. doi: 10.7861/clinmedicine.8-5-539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Luft BJ, Remington JS. Toxoplasmic encephalitis in AIDS. Clin Infect Dis. 1992;15(2):211–222. doi: 10.1093/clinids/15.2.211. [DOI] [PubMed] [Google Scholar]
- 11.Feustel SM, Meissner M, Liesenfeld O. Toxoplasma gondii and the blood–brain barrier. Virulence. 2012;3(2):182–192. doi: 10.4161/viru.19004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Buchholz KR, Fritz HM, Chen X, Durbin-Johnson B, Rocke DM, Ferguson DJ, Conrad PA, Boothroyd JC. Identification of tissue cyst wall components by transcriptome analysis of in vivo and in vitro Toxoplasma gondii bradyzoites. Eukaryot Cell. 2011;10(12):1637–1647. doi: 10.1128/EC.05182-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bahl A, Davis PH, Behnke M, Dzierszinski F, Jagalur M, Chen F, Shanmugam D, White MW, Kulp D, Roos DS. A novel multifunctional oligonucleotide microarray for Toxoplasma gondii. BMC Genomics. 2010;11:603. doi: 10.1186/1471-2164-11-603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cleary MD, Singh U, Blader IJ, Brewer JL, Boothroyd JC. Toxoplasma gondii asexual development: identification of developmentally regulated genes and distinct patterns of gene expression. Eukaryot Cell. 2002;1(3):329–340. doi: 10.1128/EC.1.3.329-340.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fritz HM, Buchholz KR, Chen X, Durbin-Johnson B, Rocke DM, Conrad PA, Boothroyd JC. Transcriptomic analysis of toxoplasma development reveals many novel functions and structures specific to sporozoites and oocysts. PLoS One. 2012;7(2):e29998. doi: 10.1371/journal.pone.0029998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Skariah S, Mordue DG. Identification of Toxoplasma gondii genes responsive to the host immune response during in vivo infection. PLoS One. 2012;7(10):e46621. doi: 10.1371/journal.pone.0046621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hill RD, Gouffon JS, Saxton AM, Su C. Differential gene expression in mice infected with distinct Toxoplasma strains. Infect Immun. 2012;80(3):968–974. doi: 10.1128/IAI.05421-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jia B, Lu H, Liu Q, Yin J, Jiang N, Chen Q. Genome-wide comparative analysis revealed significant transcriptome changes in mice after Toxoplasma gondii infection. Parasit Vectors. 2013;6:161. doi: 10.1186/1756-3305-6-161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.McGettigan PA. Transcriptomics in the RNA-seq era. Curr Opin Chem Biol. 2013;17(1):4–11. doi: 10.1016/j.cbpa.2012.12.008. [DOI] [PubMed] [Google Scholar]
- 20.Hassan MA, Melo MB, Haas B, Jensen KD, Saeij JP. De novo reconstruction of the Toxoplasma gondii transcriptome improves on the current genome annotation and reveals alternatively spliced transcripts and putative long non-coding RNAs. BMC Genomics. 2012;13:696. doi: 10.1186/1471-2164-13-696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tomita T, Bzik DJ, Ma YF, Fox BA, Markillie LM, Taylor RC, Kim K, Weiss LM. The Toxoplasma gondii cyst wall protein CST1 is critical for cyst wall integrity and promotes bradyzoite persistence. PLoS Pathog. 2013;9(12):e1003823. doi: 10.1371/journal.ppat.1003823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tanaka S, Nishimura M, Ihara F, Yamagishi J, Suzuki Y, Nishikawa Y. Transcriptome Analysis of Mouse Brain Infected with Toxoplasma gondii. Infect Immun. 2013;81(10):3609–3619. doi: 10.1128/IAI.00439-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Derouin F, Garin YJ. Toxoplasma gondii: blood and tissue kinetics during acute and chronic infections in mice. Exp Parasitol. 1991;73(4):460–468. doi: 10.1016/0014-4894(91)90070-D. [DOI] [PubMed] [Google Scholar]
- 24.Mordue DG, Monroy F, La Regina M, Dinarello CA, Sibley LD. Acute toxoplasmosis leads to lethal overproduction of Th1 cytokines. J Immunol. 2001;167(8):4574–4584. doi: 10.4049/jimmunol.167.8.4574. [DOI] [PubMed] [Google Scholar]
- 25.Saeij JP, Boyle JP, Grigg ME, Arrizabalaga G, Boothroyd JC. Bioluminescence imaging of Toxoplasma gondii infection in living mice reveals dramatic differences between strains. Infect Immun. 2005;73(2):695–702. doi: 10.1128/IAI.73.2.695-702.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ferguson DJ, Hutchison WM. An ultrastructural study of the early development and tissue cyst formation of Toxoplasma gondii in the brains of mice. Parasitol Res. 1987;73(6):483–491. doi: 10.1007/BF00535321. [DOI] [PubMed] [Google Scholar]
- 27.Di Cristina M, Marocco D, Galizi R, Proietti C, Spaccapelo R, Crisanti A. Temporal and spatial distribution of Toxoplasma gondii differentiation into Bradyzoites and tissue cyst formation in vivo. Infect Immun. 2008;76(8):3491–3501. doi: 10.1128/IAI.00254-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Cannella D, Brenier Pinchart MP, Braun L, Van Rooyen JM, Bougdour A, Bastien O, Behnke MS, Curt RL, Curt A, Saeij JP, Sibley LD, Pelloux H, Hakimi MA. miR-146a and miR-155 delineate a MicroRNA fingerprint associated with Toxoplasma persistence in the host brain. Cell Rep. 2014;6(5):928–937. doi: 10.1016/j.celrep.2014.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gotz S, Garcia-Gomez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talon M, Dopazo J, Conesa A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008;36(10):3420–3435. doi: 10.1093/nar/gkn176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jung C, Lee CY, Grigg ME. The SRS superfamily of Toxoplasma surface proteins. Int J Parasitol. 2004;34(3):285–296. doi: 10.1016/j.ijpara.2003.12.004. [DOI] [PubMed] [Google Scholar]
- 31.Butcher BA, Fox BA, Rommereim LM, Kim SG, Maurer KJ, Yarovinsky F, Herbert DR, Bzik DJ, Denkers EY. Toxoplasma gondii rhoptry kinase ROP16 activates STAT3 and STAT6 resulting in cytokine inhibition and arginase-1-dependent growth control. PLoS Pathog. 2011;7(9):e1002236. doi: 10.1371/journal.ppat.1002236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Reichmann G, Dlugonska H, Fischer HG. Characterization of TgROP9 (p36), a novel rhoptry protein of Toxoplasma gondii tachyzoites identified by T cell clone. Mol Biochem Parasitol. 2002;119(1):43–54. doi: 10.1016/S0166-6851(01)00397-8. [DOI] [PubMed] [Google Scholar]
- 33.Denton H, Roberts CW, Alexander J, Thong KW, Coombs GH. Enzymes of energy metabolism in the bradyzoites and tachyzoites of Toxoplasma gondii. FEMS Microbiol Lett. 1996;137(1):103–108. doi: 10.1111/j.1574-6968.1996.tb08090.x. [DOI] [PubMed] [Google Scholar]
- 34.Al-Anouti F, Tomavo S, Parmley S, Ananvoranich S. The expression of lactate dehydrogenase is important for the cell cycle of Toxoplasma gondii. J Biol Chem. 2004;279(50):52300–52311. doi: 10.1074/jbc.M409175200. [DOI] [PubMed] [Google Scholar]
- 35.Odberg-Ferragut C, Soete M, Engels A, Samyn B, Loyens A, Van Beeumen J, Camus D, Dubremetz JF. Molecular cloning of the Toxoplasma gondii sag4 gene encoding an 18 kDa bradyzoite specific surface protein. Mol Biochem Parasitol. 1996;82(2):237–244. doi: 10.1016/0166-6851(96)02740-5. [DOI] [PubMed] [Google Scholar]
- 36.Lekutis C, Ferguson DJ, Boothroyd JC. Toxoplasma gondii: identification of a developmentally regulated family of genes related to SAG2. Exp Parasitol. 2000;96(2):89–96. doi: 10.1006/expr.2000.4556. [DOI] [PubMed] [Google Scholar]
- 37.Saeij JP, Arrizabalaga G, Boothroyd JC. A cluster of four surface antigen genes specifically expressed in bradyzoites, SAG2CDXY, plays an important role in Toxoplasma gondii persistence. Infect Immun. 2008;76(6):2402–2410. doi: 10.1128/IAI.01494-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Van TT, Kim SK, Camps M, Boothroyd JC, Knoll LJ. The BSR4 protein is up-regulated in Toxoplasma gondii bradyzoites, however the dominant surface antigen recognised by the P36 monoclonal antibody is SRS9. Int J Parasitol. 2007;37(8–9):877–885. doi: 10.1016/j.ijpara.2007.02.001. [DOI] [PubMed] [Google Scholar]
- 39.Kim SK, Karasov A, Boothroyd JC. Bradyzoite-specific surface antigen SRS9 plays a role in maintaining Toxoplasma gondii persistence in the brain and in host control of parasite replication in the intestine. Infect Immun. 2007;75(4):1626–1634. doi: 10.1128/IAI.01862-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wong WK, Upton A, Thomas MG. Neuropsychiatric symptoms are common in immunocompetent adult patients with Toxoplasma gondii acute lymphadenitis. Scand J Infect Dis. 2013;45(5):357–361. doi: 10.3109/00365548.2012.737017. [DOI] [PubMed] [Google Scholar]
- 41.Godwin R. Toxoplasma gondii and elevated suicide risk. Vet Rec. 2012;171(9):225. doi: 10.1136/vr.e5832. [DOI] [PubMed] [Google Scholar]
- 42.Fouts AE, Boothroyd JC. Infection with Toxoplasma gondii bradyzoites has a diminished impact on host transcript levels relative to tachyzoite infection. Infect Immun. 2007;75(2):634–642. doi: 10.1128/IAI.01228-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kirkman LA, Weiss LM, Kim K. Cyclic nucleotide signaling in Toxoplasma gondii bradyzoite differentiation. Infect Immun. 2001;69(1):148–153. doi: 10.1128/IAI.69.1.148-153.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kurokawa H, Kato K, Iwanaga T, Sugi T, Sudo A, Kobayashi K, Gong H, Takemae H, Recuenco FC, Horimoto T, Akashi H. Identification of Toxoplasma gondii cAMP dependent protein kinase and its role in the tachyzoite growth. PLoS One. 2011;6(7):e22492. doi: 10.1371/journal.pone.0022492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lourido S, Shuman J, Zhang C, Shokat KM, Hui R, Sibley LD. Calcium-dependent protein kinase 1 is an essential regulator of exocytosis in Toxoplasma. Nature. 2010;465(7296):359–362. doi: 10.1038/nature09022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gazzinelli RT, Hieny S, Wynn TA, Wolf S, Sher A. Interleukin 12 is required for the T-lymphocyte-independent induction of interferon gamma by an intracellular parasite and induces resistance in T-cell-deficient hosts. Proc Natl Acad Sci U S A. 1993;90(13):6115–6119. doi: 10.1073/pnas.90.13.6115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Denkers EY, Gazzinelli RT, Martin D, Sher A. Emergence of NK1.1+ cells as effectors of IFN-gamma dependent immunity to Toxoplasma gondii in MHC class I-deficient mice. J Exp Med. 1993;178(5):1465–1472. doi: 10.1084/jem.178.5.1465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hunter CA, Subauste CS, Van Cleave VH, Remington JS. Production of gamma interferon by natural killer cells from Toxoplasma gondii-infected SCID mice: regulation by interleukin-10, interleukin-12, and tumor necrosis factor alpha. Infect Immun. 1994;62(7):2818–2824. doi: 10.1128/iai.62.7.2818-2824.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Suzuki Y. Host resistance in the brain against Toxoplasma gondii. J Infect Dis. 2002;185(Suppl 1):S58–S65. doi: 10.1086/337999. [DOI] [PubMed] [Google Scholar]
- 50.O'Brien KB, Schultz-Cherry S, Knoll LJ. Parasite-mediated upregulation of NK cell-derived gamma interferon protects against severe highly pathogenic H5N1 influenza virus infection. J Virol. 2011;85(17):8680–8688. doi: 10.1128/JVI.05142-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hauser WE, Jr, Sharma SD, Remington JS. Natural killer cells induced by acute and chronic toxoplasma infection. Cell Immunol. 1982;69(2):330–346. doi: 10.1016/0008-8749(82)90076-4. [DOI] [PubMed] [Google Scholar]
- 52.Khan IA, Schwartzman JD, Matsuura T, Kasper LH. A dichotomous role for nitric oxide during acute Toxoplasma gondii infection in mice. Proc Natl Acad Sci U S A. 1997;94(25):13955–13960. doi: 10.1073/pnas.94.25.13955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Scharton-Kersten TM, Yap G, Magram J, Sher A. Inducible nitric oxide is essential for host control of persistent but not acute infection with the intracellular pathogen Toxoplasma gondii. J Exp Med. 1997;185(7):1261–1273. doi: 10.1084/jem.185.7.1261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Bohne W, Heesemann J, Gross U. Reduced replication of Toxoplasma gondii is necessary for induction of bradyzoite-specific antigens: a possible role for nitric oxide in triggering stage conversion. Infect Immun. 1994;62(5):1761–1767. doi: 10.1128/iai.62.5.1761-1767.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Fox BA, Gigley JP, Bzik DJ. Toxoplasma gondii lacks the enzymes required for de novo arginine biosynthesis and arginine starvation triggers cyst formation. Int J Parasitol. 2004;34(3):323–331. doi: 10.1016/j.ijpara.2003.12.001. [DOI] [PubMed] [Google Scholar]
- 56.Rozenfeld C, Martinez R, Figueiredo RT, Bozza MT, Lima FR, Pires AL, Silva PM, Bonomo A, Lannes-Vieira J, De Souza W, Moura-Neto V. Soluble factors released by Toxoplasma gondii-infected astrocytes down-modulate nitric oxide production by gamma interferon-activated microglia and prevent neuronal degeneration. Infect Immun. 2003;71(4):2047–2057. doi: 10.1128/IAI.71.4.2047-2057.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Saeij JP, Coller S, Boyle JP, Jerome ME, White MW, Boothroyd JC. Toxoplasma co-opts host gene expression by injection of a polymorphic kinase homologue. Nature. 2007;445(7125):324–327. doi: 10.1038/nature05395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Maeda H, Wu J, Okamoto T, Maruo K, Akaike T. Kallikrein-kinin in infection and cancer. Immunopharmacology. 1999;43(2–3):115–128. doi: 10.1016/S0162-3109(99)00104-6. [DOI] [PubMed] [Google Scholar]
- 59.Zhang X, Scicli GA, Xu X, Nasjletti A, Hintze TH. Role of endothelial kinins in control of coronary nitric oxide production. Hypertension. 1997;30(5):1105–1111. doi: 10.1161/01.HYP.30.5.1105. [DOI] [PubMed] [Google Scholar]
- 60.Scarisbrick IA, Blaber SI, Lucchinetti CF, Genain CP, Blaber M, Rodriguez M. Activity of a newly identified serine protease in CNS demyelination. Brain. 2002;125(Pt 6):1283–1296. doi: 10.1093/brain/awf142. [DOI] [PubMed] [Google Scholar]
- 61.Scarisbrick IA, Epstein B, Cloud BA, Yoon H, Wu J, Renner DN, Blaber SI, Blaber M, Vandell AG, Bryson AL. Functional role of kallikrein 6 in regulating immune cell survival. PLoS One. 2011;6(3):e18376. doi: 10.1371/journal.pone.0018376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tobin CM, Knoll LJ. A patatin-like protein protects Toxoplasma gondii from degradation in a nitric oxide-dependent manner. Infect Immun. 2012;80(1):55–61. doi: 10.1128/IAI.05543-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29(9):e45. doi: 10.1093/nar/29.9.e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28(5):511–515. doi: 10.1038/nbt.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Goecks J, Nekrutenko A, Taylor J. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 2010;11(8):R86. doi: 10.1186/gb-2010-11-8-r86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M, Nekrutenko A, Taylor J. Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol. 2010;19(19):11–21. doi: 10.1002/0471142727.mb1910s89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Giardine B, Riemer C, Hardison RC, Burhans R, Elnitski L, Shah P, Zhang Y, Blankenberg D, Albert I, Taylor J, Miller L, Kent WJ, Nekrutenko A. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 2005;15(10):1451–1455. doi: 10.1101/gr.4086505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14(4):R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi: 10.1038/nprot.2008.211. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Differential expression analysis of T. gondii genes between acute and chronic time points generated by Cuffdiff. (XLS 2 MB)
Additional file 2: Differential expression analysis of M. musculus genes between acute vs. uninfected time points generated by Cuffdiff. (XLS 16 MB)
Additional file 3: Differential expression analysis of M. musculus genes between chronic vs. uninfected time points generated by Cuffdiff. (XLSX 9 MB)
Additional file 4: Differential expression analysis of M. musculus genes chronic divided by acute time points generated by Cuffdiff. (XLS 17 MB)
Additional file 5: Top 5 significant KEGG pathways during chronic vs acute infection. Genes with a >2 fold abundance between chronic and acute infection were uploaded to the KEGG pathway database. Displayed above are the most 5 most significant pathways enriched during chronic infection. The first column is the KEGG pathway description. Second column is the number of genes that fall into each category. The third column is the p-value designating the significance of each category. (PDF 151 KB)
Additional file 6: NK cell mediated cytotoxicity KEGG pathway. Schematic representation of NK cell mediated cytotoxicity provided by Kyoto Encyclopedia of Genes and Genomes ( http://www.genome.jp/kegg/). Red stars indicate genes that are more abundant between chronic and acute infection in the mouse forebrain. For description of features on the map visit http://www.genome.jp/kegg/document/help_pathway.html. (PDF 295 KB)


