Abstract
Intracellular trafficking pathways, including endocytosis, autophagy and secretion, rely on directed organelle transport driven by the opposing microtubule motor proteins kinesin and dynein. Precise spatial and temporal targeting of vesicles and organelles requires the integrated regulation of these opposing motors, which are often bound simultaneously to the same cargo. Recent progress demonstrates that organelle-associated scaffolding proteins, including Milton/TRAKs, JIP1, JIP3, huntingtin, and Hook1, interact with molecular motors to coordinate activity and sustain unidirectional transport. Scaffolding proteins also bind to upstream regulatory proteins including kinases and GTPases to modulate transport in the cell. This integration of regulatory control with motor activity allows for cargo-specific changes in the transport or targeting of organelles in response to cues from the complex cellular environment.
Keywords: kinesin, dynein, dynactin, organelle transport, axonal transport, intracellular trafficking
Microtubule-Based Transport in the Cell
The spatial complexity of eukaryotic cells relies on the precise regulation of intracellular trafficking. Long-distance organelle transport depends on microtubules, which serve as polarized tracks for motility within the cell. Delivery of diverse cargos along microtubules underlies many essential cellular functions, such as protein secretion, growth and injury signaling, protein and organelle degradation, active distribution of mitochondria, and trafficking of RNA granules. Furthermore, cells precisely regulate organelle transport by coordinating motor activity with cellular demands and in response to changes in the cellular environment.
Microtubules are organized into a polarized array, with dynamic plus-ends oriented toward the cell periphery and more stable minus-ends clustered toward the center in most cell types. Anterograde transport, or motility toward plus-ends, is driven by members of the extended kinesin superfamily including kinesin-1, kinesin-2, and kinesin-3 (Box 1); retrograde transport, or motility toward minus-ends, is driven primarily by cytoplasmic dynein acting in concert with its activator dynactin (Box 2). The biophysical properties of these motors, such as velocity, run length, and stall force, have been studied in detail at the single-molecule level [1]. However, less is known about how these motors are regulated when bound to cargo in order to drive organelle-specific motility.
Box 1. Kinesins and Regulation by Autoinhibition.
The kinesin superfamily includes a diverse collection of motors that are encoded by more than 45 genes. The majority of kinesins move toward microtubule plus-ends though some move toward minus-ends (reviewed in [95]). Kinesin motors share a homologous motor domain that is fused to more diverse domains mediating specific protein and organelle associations.
Though multiple kinesin families are involved in organelle transport [95], kinesin-1, kinesin-2, and kinesin-3 motors have been best studied to date. Kinesin-1 consists of a KHC (kinesin heavy chain) dimer, which usually associates with two light chains (KLC; Box 3). Mammalian KHC is encoded by three different genes – KIF5A, KIF5B, and KIF5C; while KIF5B is ubiquitously expressed, KIF5A and KIF5C are enriched in neurons [96]. Interestingly, different KIF5’s may selectively transport certain cargos, as KIF5A preferentially binds to the GABA receptor adaptor GABARAP in yeast-two hybrid assays [97]. Kinesin-2 is also a two-headed motor formed from either homo-dimerization or heterotrimerization (with a nonmotor subunit). Kinesin-3 motors are found as either monomers or dimers [83]. Kinesins not associated with cargo can be autoinhibited by intramolecular interactions [82, 83, 98]. Kinesin autoinhibition likely has two essential cellular functions – to prevent the unnecessary crowding of kinesins on microtubules (i.e. traffic jams) and to prevent wasteful ATP hydrolysis in the absence of associated cargo [98].
For KHC, direct binding between the N-terminal motor head and a conserved basic IAK motif in the C-terminal tail results in autoinhibition [99]. In vitro motility experiments indicate that full-length KHC has little motor activity, whereas truncation of the stalk and tail leads to a constitutively active motor [100]. In early micrographs of KHC, the tail appeared to fold back onto the head via bending of the stalk [101]. Deletion of the “hinge 2” motif in the stalk blocks autoinhibition, indicating that the flexibility of this element may be pivotal for bending of the tail to meet the head [100]. Interestingly, scaffolding proteins may relieve autoinhibition by binding to both KHC stalk and tail. In vitro, the binding of a JIP1 fragment to the stalk domain is sufficient to activate KHC motility [72], though full activation in vivo may require binding of JIP1 to both the stalk and tail domains.
A single KHC tail is sufficient to autoinhibit a KHC head dimer [102]. One KHC tail peptide binds to the cleft between the two motor heads where it likely prevents ADP release by restricting the movement of the motor domains [103]. This “double lockdown” model has an important cellular implication – relief of KHC autoinhibition requires that both tails of KHC dimer be blocked from binding to the motor domains. Interestingly, scaffolding proteins known to activate KHC motility in vitro via tail binding, including JIP1 and JIP3, exist as dimers [64, 72]; thus, dimerization of scaffolding proteins may be required for full activation of KHC in the cell.
Box 2. Cytoplasmic Dynein and Dynactin.
Cytoplasmic dynein is the major minus-end-directed microtubule motor for organelle transport in eukaryotic cells. The enormous (~1.5 MDa) dynein complex consists of two dynein heavy chains (DHC; ~500 kDa), two dynein intermediate chains (DIC; ~74 kDa), two light intermediate chains (DLIC; ~33–59 kDa), and several dimers of light chains (DLC/LC7/roadblock, LC8, and TcTex1; ~10–14 kDa). The catalytic DHC is an AAA-protein that folds to form a large asymmetric ring that includes the primary ATPase domain; the microtubule-binding domain is localized to a stalk projecting from this ring [104]. While DHC is both necessary and sufficient for motor activity, additional subunits contribute to complex stability and cargo interactions.
Single molecule studies show that mammalian dynein is a weak motor with stall force of ~1pN and a variable step size of 8–24 nm [1]. Unlike kinesin, dynein can sidestep to move laterally onto adjacent protofilaments. While single molecules of mammalian dynein can move bidirectionally along microtubules in vitro [105], in the cell dynein functions as a robust minus-end-directed motor. Dynein interacts directly with dynactin as well as other activator or adaptor complexes, including Bicaudal D (BicD), lissencephaly 1 (Lis1) and nuclear distribution protein E (NudE) or NudE-like (NudEL) [104, 106]. These activators may act in concert to enhance efficient minus-end-directed transport.
Dynactin is a large ~1-MDa complex that interacts directly with dynein via its largest subunit, p150Glued, named for its size (~150 kDa) and its homology to the Drosophila Glued gene [107]. The N-terminus of p150Glued contains the CAP-Gly microtubule-binding domain [108], followed in some isoforms by a lower affinity microtubule-binding domain enriched in basic amino acid residues [109]. p150Glued forms a dimer, and also binds directly to the DIC subunit of cytoplasmic dynein, via a coiled-coil (CC1) domain [110–112]. A second coiled-coil domain (CC2) in p150Glued interacts with actin-related protein 1 (Arp1), which forms a short actin-like filament at the base of the dynactin complex. Additional dynactin subunits include p62, dynamitin (p50), pArp11, CapZ, p27, p25 and p24 [113]. There is a cargo-binding domain (~AA 1049–1278) at the C-terminus of p150Glued that binds to a growing list of vesicular adaptors, such as the Rab7-interacting lysosomal protein (RILP [24]), huntingtin-associated protein 1 (HAP1 [45]), the retromer sortin nexin 6 (SNX6 [28, 29]), and JIP1 [72].
Recent progress on the roles of adaptor or scaffolding proteins is providing new insights into motor coordination at the cellular level. Scaffolding proteins can effectively regulate the activities of organelle-associated motors to control directionality. Scaffolding proteins also bind to upstream regulatory elements, including kinases, phosphatases, Ca2+-signaling, and G-proteins. This allows for the integration of diverse signals to yield organelle-specific responses to the local cellular environment. Here, we focus on new developments in the regulation of organelle-associated motors by scaffolding and adaptor proteins, including Milton/TRAK, Miro, RILP, huntingtin, La, JIPs1-4, and Hook. Progress to date has identified some common mechanisms, but also highlights the exquisite selectivity of regulatory control that leads to cargo-specific and regionally-specific patterns of motility within the cell.
MODELS FOR MOTOR REGULATION DURING INTRACELLULAR TRANSPORT
The simplest model to describe cargo-specific regulation of organelle motility involves the selective recruitment of either kinesin or dynein motors (Figure 1A). If kinesin is recruited, the organelle will move toward the microtubule plus-end. However, if the bound kinesins dissociate and dynein is recruited, then the cargo will move toward the microtubule minus-end. This model would be predicted to yield unidirectional, highly processive motility. In contrast to this simple model, increasing evidence from both in vitro and cellular studies suggests that opposing kinesin and dynein motors are often bound simultaneously to cargos moving along microtubules [2–7]. Two models have been put forth to describe the interactions of opposing kinesin and dynein motors bound to the same cargo, and the resulting patterns of motility.
Figure 1. Three current models for the regulation of microtubule motors bound to vesicular and organelle cargo.

(A) In the selective recruitment model, only one type of motor, either kinesin (KHC) or dynein-dynactin, are bound to cargo at time. If kinesin is bound, the cargo will move unidirectionally in the anterograde direction toward the plus-end of the microtubule. If dynein-dynactin is bound, the cargo will move unidirectionally in the retrograde direction, toward the microtubule minus-end.
(B) In the tug-of-war model, both kinesin and dynein-dynactin motors are bound to the cargo simultaneously. The cargo will move bidirectionally along the microtubule, depending on stochastic variations in the dominant motor type. Note that for simplification, this figure only illustrates one dynein-dynactin complex per vesicle, but that likely 6–8 dynein-dynactin complexes are on each vesicle to reach force balance with one kinesin.
(C) In the coordination model, kinesin and dynein are bound to the cargo simultaneously, but the activities of these motors are governed by a scaffolding protein that coordinates the engagement of dynein-dynactin with the autoinhibition of kinesin. Inset – Generalized model for the integration of upstream signaling with downstream motility by scaffolding proteins. Scaffolding proteins interact with vesicle/organelle linker proteins, upstream signaling proteins, and molecular motors, forming an integrated regulatory unit. While scaffolding proteins may also mediate the association of motors with the vesicle or organelle, this is not always the case.
Many organelles in the cell exhibit bidirectional motility, characterized by brief excursions toward either the microtubule plus- or minus-end punctuated with frequent directional switches. A tug-of-war mechanism has been proposed to explain these observations. In this scenario, the team of motors, either plus-end-directed or minus- end-directed, that produce the greater force at any given time determines the direction of transport (Figure 1B). Force-dependent detachment of the opposing motors from the microtubule track prevents unproductive stalling. Stochastic motor release from the track allows for alterations in the dominant motor type over time, leading to directional switching [8]. Experimental work in a number of systems supports this model. For example, late endosomes and lysosomes along neuronal axons exhibit bidirectional motility, characterized by short run lengths either toward or away from the cell body punctuated by frequent changes in directions. Quantitative immunoblotting of purified vesicles indicates that each organelle binds few (1–2) kinesin motors and a larger team (6–12) of dynein motors [3]. However, because kinesins generally exhibit high unitary stall forces (~5–7 pN) whereas mammalian cytoplasmic dynein has a low unitary stall force (~1 pN) [1], these opposing motors are actually present near force balance on each organelle. The resulting stochastic tug-of-war between these relatively evenly matched motor teams is thus predicted to cause frequent directional switches and low net processivity [3].
A third model postulates that opposing motors are simultaneously bound to cargo, but their motility is tightly coordinated by regulatory mechanisms. In this coordination model, both kinesin and dynein motors remain continuously bound to the organelle, but are not constitutively active (Figure 1C). Instead, motor activity may be specifically regulated by post-translational modifications or by adaptor or scaffolding proteins. This model would explain the motility of fast-moving processive cargos with few directional changes, such as autophagosomes moving along neuronal axons. These organelles stably bind both kinesin and dynein motors yet undergo primarily unidirectional transport toward the cell soma [4], indicating that the associated kinesin motors are efficiently inhibited during dynein-driven motility [9].
What is the advantage of maintaining simultaneous, stable associations with opposing motors? In the case of the unregulated tug-of-war model, one possibility is that while the stochastic directional changes produce less efficient long-distance motility, these properties may be important for developing and/or maintaining stable distributions of organelles along extended cellular processes such as the axon. In the coordination model, opposing motors may allow for quick direction changes, perhaps to avoid roadblocks or traffic jams [10, 11]. Another major advantage of the simultaneous binding of opposing motors is to allow for rapid responses to changes in the local cellular environment.
While cargo-specific regulation allows the cell to selectively modulate the motility of different organelle populations, additional mechanisms also contribute to the regulation of trafficking within the cell. There is strong data indicating track-specific regulation of organelle transport via post-translational modification of microtubules [12– 15] or through the binding of microtubule-associated proteins (MAPs) that may aid or hinder motor processivity [10, 16–18]. Additionally, regionally-specific regulation of transport may result from differences in the local organization of the cytoskeleton such as the enrichment of dynamic microtubule plus-ends near the cell cortex, microtubule-microtubule intersections [19], or the formation of dense actin filaments that can induce track switching or impede motion (reviewed in [20]). Finally, given the crowded nature of the cellular environment, there is growing evidence that many of the prominent pauses or directional changes observed during organelle motility result from collisions with other cellular organelles, much like cars colliding at rush hour on crowded highways [21, 22]. However, the strikingly different types of motility observed for distinct cargos moving through the same region of the cell indicate that cargo-specific regulation may be the dominant mechanism involved in the control of organelle motility in vivo.
SCAFFOLDING PROTEINS COORDINATE MOTOR ACTIVITY
Eukaryotic cells express a repertoire of scaffolding proteins, which act as binding platforms to mediate multiple protein-protein interactions and to integrate diverse activities (Figure 1C, inset). Scaffolding proteins involved in the regulation of molecular motors generally bind to multiple classes of proteins, including 1) membrane-associated cargo receptors; 2) components of both kinesin and dynein motor complexes; and 3) signaling proteins such as kinases or GTPases. In this manner, scaffolding proteins integrate on-board signaling proteins to locally regulate cargo motility. Scaffolding proteins are often core components of large complexes, interacting with additional accessory or adaptor proteins. Frequently, scaffolding proteins form multiple distinct interactions with motors, which may enhance the stability of the holo-complex. Here, we highlight some of the best-characterized motor-scaffolding complexes and their regulatory schemes.
Retrograde Scaffolding Proteins: RILP Complex and Retromers
The scaffolding protein RILP (Rab7-interacting lysosomal protein) facilitates the retrograde transport of late endosomes and lysosomes. As an early endosome matures into a late endosome, activated GTP-Rab7 orchestrates the recruitment of RILP and ORP1L. RILP then recruits dynactin via direct binding to the C-terminus of the p150Glued subunit of dynactin [23]. The RILP scaffolding complex makes an additional association with the late endosome via the Arp1 subunit of dynactin, which can bind to vesicle-associated βIII-spectrin [24, 25].
The stoichiometry of RILP association has been suggested to play an important role in recruiting teams of dyneins to lysosomes. In optical trapping experiments in rodent macrophage cell lines, phagosomes exhibit bidirectional movement with measurable forces generated by both kinesin and dynein [6, 26]. However, highly processive retrograde runs exhibit higher stall forces that cluster in increments of 2 pN. Since mammalian dynein has a unitary stall force of 1 pN, these results suggest that dynein motors are recruited onto vesicles in pairs [26], consistent with recruitment of dimeric RILP by Rab7 [23].
Regulation of motor association via membrane lipid composition is emerging as an important yet understudied facet of vesicular transport. The oxysterol-binding protein ORP1L, shown to be a cholesterol sensor, facilitates RILP/p150Glued binding to cholesterol-rich lysosomes, which are then transported toward the nucleus. In contrast, cholesterol-poor lysosomes remain at the cell periphery, because organelle-associated ORP1L undergoes a conformational change to recruit the ER protein VAP, leading to dissociation of p150Glued [27]. The retromer complex, which facilitates the return of endosomal transmembrane proteins to the trans-Golgi network (TGN), also relies on lipid composition to regulate release of the retrograde motor. Once the vesicle reaches the Golgi, the retromer component SNX6, which binds directly to p150Glued [28, 29], senses Golgi-enriched phosphotidylinositol-4-phosphate (PI4P) and releases dynactin [30].
Mitochondrial Scaffolding Proteins: Milton/TRAKs and Miro
In neuronal axons, the majority of mitochondria (~60%) are either stationary or move bidirectionally with frequent directional changes, while the remainder move robustly in either the anterograde or retrograde directions [31] [32, 33]. In Drosophila, the Milton/Miro complex mediates the binding of microtubule-based motors to mitochondria. Miro (Mitochondrial Rho GTPase) associates with the outer mitochondrial membrane via a transmembrane domain and recruits Milton, which binds directly to KHC in a KLC-independent manner [34]; see Box 3.
Box 3. Kinesin Light Chain – Autoinhibitory Clamp or Cargo Adaptor?
Cytoplasmic kinesin-1 is typically depicted as an inactive heterotetrameric complex with two KHCs bound to two KLCs. The tetratricopeptide repeat (TPR) domain of KLC binds to C-terminal KHC (AA 771–810) in a region distinct from the autoinhibitory KHC tail [114, 115]. Experiments with full-length proteins demonstrate that KHC homodimers are more robustly autoinhibited in the presence of KLC. Indeed, KHC/KLC heterotetramers have decreased microtubule-binding ability compared to KHC homodimers [114]. Moreover, when compared to the infrequent runs made by KHC, addition of KLC leads to ~30% decrease in run frequency and velocity and ~60% decrease in run length [100], consistent with a role for KLC in maintaining KHC autoinhibition.
Cargo-associated KHC may not always be associated with KLC. Sucrose density gradient centrifugation has identified three separate pools of kinesin – a low-density pool containing only KLC, a high-density pool containing KHC and KLC, and a higher-density pool containing KHC alone [116]. Recent experiments also found that a pool of naked KHC in high-density sucrose gradient fractions [64], consistent with the ability of many scaffolding proteins or cargo to bind to KHC in a KLC-independent manner, including JIP3 [64], Milton [34], and RNA granules in Drosophila [117]. In contrast, KLC acts as an adaptor that mediates the binding kinesin-1 to other cargos, such as bacteria-containing lysosomes [118, 119] and the neuronal cargo calsyntenin [120].
We propose a unifying hypothesis whereby soluble kinesin-1 exists as a KHC/KLC heterotetramer that is robustly autoinhibited. Activation of this complex involves relief of KHC autoinhibition, likely by a scaffolding protein. For some cargos, KLC is not necessary for this activation step. For others cargos though, KLC may function as an obligate cargo adaptor by binding to cargo-associated proteins.
At neuronal synapses, which have high local energy demands, elevated calcium levels disrupt the association of KHC with mitochondria, leading to the effective local sequestration of mitochondria. Miro has two calcium-sensing EF hand domains and is suggested to regulate this response; two molecular mechanisms have been proposed. In one model, high levels of calcium induce the binding of the KHC motor head to Miro, which effectively prevents the head from processing along microtubules [35]. In another model, calcium binding to Miro results in the release of KHC from Milton, likely by altering the binding of Milton and Miro [36]. In addition to regulation by calcium sensing, Miro can be phosphorylated by the PINK1 kinase upon mitochondrial damage or depolarization, and then targeted for degradation by the ubiquitin ligase Parkin. This mechanism results in the removal of the Milton/Miro complex and its associated motors from damaged mitochondria; the resulting arrested motility has been proposed to facilitate clearance by mitophagy [37].
Recently, the mammalian homologues of Milton, TRAK1 and TRAK2, were shown to associate with the retrograde motor complex via the p150Glued subunit of dynactin. Interestingly, N-terminal TRAK binds to both dynactin and KHC (Figure 2A), which may mediate bidirectional transport of mitochondria. Consistent with this possibility, artificially induced recruitment of an N-terminal fragment of TRAK to peroxisomes results in bidirectional motility [38]. In contrast, depletion of TRAKs results in arrest of the anterograde- and retrograde-moving pools of mitochondria in both axons and dendrites, suggesting that TRAKs mediate processive transport in a regulated manner that remains to be fully elucidated.
Figure 2. Schematic of binding interactions between bidirectional scaffolding proteins with motor and regulatory proteins.
(A-D) One feature of bidirectional scaffolding proteins is that binding domains of anterograde and retrograde motors often are located in close proximity or are overlapping. TRAK binds to Miro, KHC, Miro and p150Glued dynactin. Huntingtin binds to dynein intermediate chain (DIC), optineurin, HAP40, and HAP1, which binds to KLC and p150Glued and can be phosphorylated at S421. JIP3 binds to JNK, ARF6, KHC tail, KLC, and p150Glued. JIP1 binds to JNK, KHC tail and stalk, KLC, and p150Glued. LZ: leucine zipper, JBD: JNK-binding domain; SH3: src homology domain; PTB: phosphotyrosine-binding domain.
Mitochondrial motility is also regulated by the mitochondrial docking protein syntaphilin, which anchors stationary mitochondria along microtubules [39]. Consistent with the idea that mitochondrial capture occurs at sites of high energetic demand, the LKB1/NUAK1 kinase pathway, which is necessary for axon branching, was recently shown to regulate mitochondrial capture at presynaptic sites by syntaphilin [40]. It remains to be established whether LKB1/NUAK1 phosphorylates syntaphilin directly or acts indirectly via intermediate effectors. Moreover, syntaphilin may compete with Milton for binding to mitochondrial-associated kinesin. At high calcium concentrations, KHC tail dissociates from Milton and binds to syntaphilin. Unlike other adaptor proteins that bind to KHC tail in order to relieve autoinhibition and activate KHC activity (Box 1), the binding of KHC tail to syntaphilin abrogates the ability of the motor domain to hydrolyze ATP, although the mechanism is not yet clear [41].
Huntingtin Scaffolds Kinesin-1 and Dynein Motors in Neurons
Huntingtin, best known for its causative role in Huntington’s disease when mutated, is a large (350-kDa) organelle-associated protein that scaffolds both minus-end-directed and plus-end-directed motors. Huntingtin binds directly to the intermediate chain of cytoplasmic dynein, DIC [42]. Huntingtin also binds to the adaptor protein HAP1, which binds to KHC [43] and to KLC [44]. HAP1 in turn interacts with the p150Glued subunit of dynactin [45, 46], thus forming a secondary interaction between the huntingtin complex and the dynein motor (Figure 2B).
Huntingtin has been shown to facilitate dynein-mediated organelle trafficking of organelles including late endosomes/lysosomes in HeLa cells [42, 47]. In neurons, huntingtin depletion inhibits the axonal transport of cargos including synaptic vesicles [48], BDNF- and APP-positive vesicles [49, 50], and autophagosomes [51]. Huntingtin also mediates the dendritic transport of β-actin mRNA [52] and TrkB receptors [53]. The binding of huntingtin and HAP1 to dynein and dynactin may enhance the stability of the active motor complex and thus enhance processive motility, while HAP1 may modulate kinesin autoinhibition [51], although the specific molecular mechanisms involved remain to be determined. Phosphorylation of huntingtin has been proposed to act as a molecular switch to regulate direction of transport on microtubules, as the phosphorylation of huntingtin at S421 by the Akt kinase promotes the anterograde transport of BDNF-positive vesicles [50].
Huntingtin may be uniquely poised to integrate adaptors for both microtubule-based and actin-based motors. Huntingtin binds to the myosin VI adaptor optineurin [54] and knockdown experiments in HeLa cells suggest that huntingtin may be necessary for the transition of endosomes and lysosomes from actin tracks to microtubule tracks at the cell periphery [47]. Indeed, another huntingtin-binding adaptor, HAP40, also binds to the early endosomal marker Rab5; thus it has been proposed that huntingtin alternates between two states – a HAP40/optineurin/myosin complex that facilitates actin-based motility of early endosomes and a HAP1/kinesin/dynein complex that facilitates microtubule-based transport of late endosomes and lysosomes [55].
Importantly, pathogenic polyglutamine expansion of N-terminal huntingtin causative for Huntington’s disease disrupts the transport of multiple organelle cargos in neurons [43, 48, 49, 51, 56]. These observations may link defects in the scaffolding of motor proteins to the pathogenesis of neurodegenerative disease. Motor activity may also be altered more directly in pathological conditions. Mutant huntingtin induces the aberrant activation of neuronal-specific JNK3, a kinase that can phosphorylate KIF5C at S176 in vitro [57, 58]. Studies on phosphomutants (at the conserved KIF5B S175 site) indicate that while ATPase activity, microtubule-binding, and processivity are not altered, there is some enhanced autoinhibition of the phosphorylated kinesin motor that may favor minus-end-directed over plus-end-directed motility [59]. Thus, direct phosphorylation of molecular motors may also contribute to the tuning of transport, and this tuning may be altered in disease.
La Scaffolding Protein in RNA Transport
Another bidirectional scaffolding protein involved in RNA trafficking is the RNA-binding protein La, which facilitates the binding of anterograde and retrograde motors to RNA granules in axons. La can be covalently modified at residue K41 by addition of small ubiquitin-like modifying polypeptide (SUMO). Sumoylated La associates with DIC but not with KHC, suggesting that sumoylation of La may preferentially promote retrograde RNA transport. Indeed, live-cell imaging shows that wild type La-GFP moves in both anterograde and retrograde directions along axons whereas La-GFP containing a K41R mutation only moves in the anterograde direction, indicative of failure to associate with dynein in the absence of sumoylation [60]. Importantly, these results imply that SUMO ligases may be enriched or preferentially activated in the distal axon. This work was one of the first clear demonstrations that the post-translational modification of a scaffolding protein results in a switch in transport direction for an intracellular cargo, in support of the coordination model proposed above (Figure 1).
JNK Binding Scaffolds: JIP3 and JIP4
JNK-interacting proteins (JIPs) are scaffolds identified through their ability to bind to multiple kinases in the c-jun N-terminal kinase (JNK) pathway. The four mammalian JIPs all share a similar JNK-binding domain. However, the domain structure common to JIP1 and JIP2 is distinct from that shared by JIP3 and JIP4 [61]. JIP1 and JIP3/JIP4 have been shown to function as scaffolding proteins for microtubule motors, but these proteins exhibit distinct regulatory mechanisms and modulate different cellular pathways.
JIP3, originally identified as Sunday Driver in Drosophila, was subsequently characterized as an adaptor that associates with dynein/dynactin during the transport of axonal injury signals in neurons [62]. Mass spectrometry of isolated JIP3-positive vesicles identified two distinct classes – an endocytic population of large vesicles likely involved in initial retrograde injury signaling to the cell body and a pool of small vesicles containing synaptic proteins suggested to play a role in axonal outgrowth and guidance [63]. JIP3 is a bidirectional scaffolding protein capable of binding to both kinesin-1 and dynein motor complexes. Binding of JIP3 to the tail of KHC relieves autoinhibition (Box 1) and activates KHC motility in single molecule assays [64]. The JIP3 homologue in C. elegans, UNC16, has also been shown to bind to both kinesin-1 and dynein light intermediate chain (DLIC; [65]).
While JIP3 has been most clearly implicated in injury signaling in neurons, growth factors such as nerve growth factor (NGF) or brain-derived neurotrophic factor (BDNF) are also actively transported from the distal axon back to the soma. This trafficking proceeds via the endosomal pathway, in which endosomes mature from Rab5-positive early endosomes to Rab7-positive late endosomes [66]. One downstream effector of the activated NGF receptor TrkB is extracellular signal-regulated kinase (ERK), which may directly phosphorylate dynein intermediate chain (DIC) at the S80 site to efficiently recruit dyneins for the retrograde transport of signaling endosomes [67]. Interestingly, TrkB can directly bind to the scaffolding protein JIP3 [68]. This raises the possibility that phosphorylation of DIC at S80 may enhance dynein recruitment to signaling endosomes in a scaffolding-protein-dependent manner.
Both of the homologous proteins JIP3 and JIP4 are effectors of the endosomal membrane protein ADP-ribosylation factor 6 (ARF6). While JIP3 is primarily expressed in neuronal cells, JIP4, is more widely expressed, and has been shown to interact with ARF6, Kinesin-1, and dynactin to regulate the trafficking of recycling endosomes during cytokinesis [69]. GTP-ARF6 binds directly to the LZII (leucine zipper II) region of JIP3 and JIP4, where it blocks the association with the TPR domain of KLC and thus favors binding to dynactin [69]. However, KHC tail binds to JIP3 at an N-terminal region distinct from the KLC-binding domain (Figure 2C), and thus it remains unclear whether the binding of ARF6 to JIP3 effectively inhibits kinesin motor activation.
JIP1 Regulates Highly Processive Organelle Transport
The scaffolding protein JIP1 mediates the axonal transport of organelles that move processively toward both microtubule plus-ends and minus-ends, including mitochondria, synaptic vesicles [70], and APP-positive vesicles [71, 72]. JIP1 also regulates the transport of a more unidirectional cargo, the autophagosome [9]. JIP1 is thus a bidirectional scaffolding protein that interacts directly with both kinesin and dynein motor complexes [72, 73]. JIP1 also binds to transmembrane cargo protein APP [74, 75] and to upstream kinases and phosphatases, including JNK [76, 77].
JIP1 regulates Kinesin-1 motor function via distinct binding interactions with both the stalk and tail domains of KHC (Figure 2D). JIP1 binding relieves KHC autoinhibition, leading to activation of kinesin motility in single molecule studies in vitro. JIP1 also interacts with the dynein motor complex via direct binding to the C-terminal domain of p150Glued. Because JIP1 cannot simultaneously bind to both KHC and dynactin, it assembles into two mutually exclusive complexes that mediate either plus-end-directed or minus-end-directed transport [72]. This mechanism ensures processive transport in either direction.
In addition to direct interactions with KHC, JIP1 also binds directly to KLC, which may act as an additional autoinhibitory clamp on the Kinesin-1 motor (Box 2). Activation of KHC in the presence of KLC requires the KHC-binding protein FEZ1, which is also found in a complex with JIP1 [78]. Thus, in addition to sustaining transport, JIP1 may facilitate the initial formation of an active transport complex for APP-positive vesicles. Consistent with this possibility, depletion of JIP1 leads to a significant decrease in the number of APP vesicles in the axon [72].
Phosphorylation of JIP1 by JNK kinase acts as a directional switch for APP transport in axons. Phosphorylation at S421, within the KHC-tail-binding domain of JIP1, stabilizes the JIP1-kinesin complex and preferentially enhances plus-end-directed APP transport [72]. In contrast, expression of a nonphosphorylatable mutant of JIP1 (S421A) favors the minus-end-directed motility of APP vesicles in axons. JIP1 can also bind to the MKP phosphatase [77], suggesting that a complete regulatory mechanism for directional control is integrated by this scaffolding protein.
JIP1 can homodimerize via its SH3 (src homology) domain [79] and additionally can heterodimerize with JIP2 and JIP3. It is unclear if JIP1 and JIP3 binding forms a heterodimer or a heterotetramer, but the JIP1/JIP3 complex can be co-transported [80] and may function to facilitate APP transport. Immunostaining and coimmunoprecipitation experiments suggest that JIP1 preferentially associates with phosphorylated APP [71] and the additional association of JIP3 with this complex may sustain phosphorylation of APP [81].
SCAFFOLDS AND ADPATORS REGULATING OTHER KINESINS
While Kinesin-1 is a major motor for intracellular transport, other members of the extended kinesin superfamily also have established roles in vesicle trafficking (Box 1). Both Kinesin-2 and Kinesin-3 are regulated by autoinhibition [82, 83], but much less is known about how the activity of these kinesins is coordinated with that of opposing dynein motors.
The tail domain of the Kinesin-2 motor KIF17 interacts with the PDZ domain of the scaffolding protein Mint1. Mint1 in turn binds to NMDA receptor subunit 2B to mediate trafficking in dendrites [84]. Ca2+/calmodulin-dependent protein kinase (CaMKII) directly phosphorylates KIF17 tail to disrupt its association with Mint1, thus releasing the cargo from the motor [85]. Modulation of this interaction contributes to synaptic plasticity, and targeted disruption of this interaction in mice leads to deficits in both behavior and memory [86]. Kinesin-2 has also been shown to bind directly to dynactin [87, 88], raising the interesting possibility that dynactin may serve to coordinate bidirectional motility, but more work is required to unravel the mechanisms involved.
Kinesin-3 interacts with the DENN/MADD protein, a Rab3 GDP-GTP exchange factor (GEF) that binds to the stalk region of the Kinesin-3 motors KIF1A and KIF1 Bβ. This regulates the axonal transport of Rab3-positive synaptic vesicle precursors [89]. Though it has not been demonstrated that direct binding of DENN/MADD is sufficient to activate KIF1B motility in vitro, DENN/MADD likely act as a scaffolding protein that mediates the regulated activity of motors on cargos in the cell.
Recent work provides insight on the coordinate regulation of kinesin-3 motors and dynein during endosomal transport in filamentous fungi. In both Ustilago maydis and Aspergillus nidulans, a Hook protein is required for the transport of early endosomes. While the C-terminal domain of Hok1/HookA mediates binding to endosomes, the N-terminal domain is required to recruit dynein and dynactin to endosomes to drive minus end-directed motility along the microtubule [90, 91]}, that can counteract the activity of the plus end-directed kinesin-3 motor [91]. It remains to be determined if motors bind to Hok1/HookA directly or if additional binding partners mediate these interactions.
In mammalian cells, three Hook proteins are expressed and differentially localized within the cell; all three can associate with microtubules either directly or indirectly via a conserved N-terminal domain [92]. Hook1 has been shown to associate with tubular endosomes via its C-terminal domain and is required for cargo sorting into tubular endosomes in conjunction with Rab22 [93], while Hook2 is implicated in ciliogenesis and Hook3 is enriched at the Golgi. All three mammalian Hook proteins have been implicated in vesicle trafficking and/or fusion [94], but it is not yet clear if these proteins interact with kinesin or dynein motors to regulate organelle transport along microtubules.
CONCLUDING REMARKS
Live cell imaging continues to reveal the complex dynamics of intracellular trafficking though many mechanistic questions remain (Outstanding Questions Box). Distinct organelle populations move with characteristic patterns of motility, some highly processive, others displaying frequent directional switching. This diversity is enabled by cargo-specific regulation of motility, integrated via organelle-bound scaffolding proteins. By binding to molecular motors either directly or via adaptor proteins, scaffolding proteins coordinate the activities of opposing motors. The additional interactions of scaffolding proteins with regulatory proteins such as kinases and GTPases favor the formation of an organelle-specific regulatory module. This compartmentalized integration of signaling and motility may be essential for the sustained transport of organelles over long distances, such as transport along the axons of human motor neurons that can reach a meter in length. Vesicle and organelle motility also varies with the local cellular environment and in response to extracellular stimuli. The ability of scaffolding proteins to fully integrate upstream signaling with downstream responses such as activation of motility, or induction of directional switching allows rapid and effective responses to these stimuli.
Outstanding Questions Box.
Can multiple scaffolding complexes associate with a single cargo?
Depletion or mutation of one scaffolding protein often leads to transport defects in many different organelles. How does one adaptor associate with multiple cargos and what accounts for cargo-specific motile characteristics such as directionality, speed, run length?
Since many scaffolding proteins and most molecular motors are dimers, what is the binding stoichiometry of scaffolding proteins to motors (e.g. 1:1 or 1:2)?
Are motors recruited from a cytoplasmic pool upon scaffolding protein activation or are resident vesicle-associated motors activated upon scaffolding protein association? Alternatively, might dormant cargo-associated scaffolding proteins become activated and then bind to motors (Figure II)?
How do upstream pathways (e.g. kinases) spatially regulate the initiation of transport? Is this step different from sustaining transport?
How is sustained transport effectively regulated over long distances (>100 um)?
A comparison of the diverse mechanisms by which scaffolding proteins function suggests some common themes, such as the simultaneous association but selective activation of opposing motors, and the important role for autoinhibition of kinesins. However, each type of cargo is likely governed by distinct mechanisms that provide specific regulatory control over organelle transport in the cell. Multi-protein binding platforms such as scaffolding proteins thus facilitate both integrated and specific regulation of intracellular motility.
Figure I (inset, Box #2). Schematic of binding domains in the p150Glued subunit of dynactin.

Several scaffolding proteins share a binding site on C-terminal p150Glued.
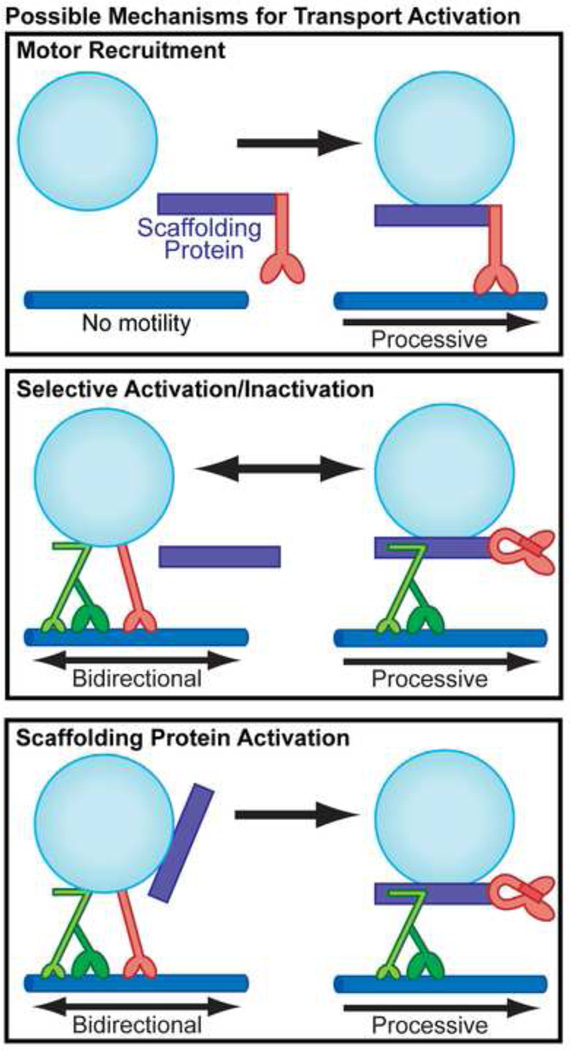
Figure II (inset, Outstanding Questions Box). Scaffolding proteins may regulate organelle motility by multiple different mechanisms.
The scaffolding proteins known to regulate organelle motility are diverse, and may function through diverse mechanisms. In the motor recruitment model, a soluble scaffolding protein may mediate the association of the motor with its cargo. In a selective activation/inactivation model, scaffolding proteins do not recruit motors to cargo, but function instead to regulate motor activity, selectively activating or inactivating one of the vesicle-bound motors. In a third scaffolding protein activation model, vesicle- bound scaffolding proteins become activated to tightly control activity of vesicle-bound motors.
Table 1. Summary chart of scaffolding proteins and their associated motors and regulatory schemes.
DLIC: dynein light intermediate chain.
| Scaffolding Protein | Kinesin Association | Dynein/Dynactin Association | Regulatory Scheme | Additional Adaptors in Complex | Organelles |
|---|---|---|---|---|---|
| RILP | - | Dynactin (p150Glued) | GTPase (Rab7) Cholesterol sensing (ORP1L) |
Rab7, ORP1L, βIII-spectrin | Late endosomes |
| SNX6 | - | Dynactin (p150Glued) | Lipid sensing (PI4P) | SNX1, SNX4, other retromers | Retromer-associated vesicles |
| Milton/TRAK | Kinesin-1 (KHC) | Dynactin (p150Glued) | Calcium sensing (Miro) Phosphorylation/degradation (PINK/Parkin) |
Miro | Mitochondria |
| Huntingtin | Kinesin-1 (KHC, KLC via HAP1) | Dynein (DIC) Dynactin (p150Glued) via HAP1 |
Phosphorylation (Akt) | HAP1, Optineurin, HAP40 | Endosomes, lysosomes, autophagosomes, APP-positive vesicles, BDNF-positive vesicles, GABA receptors, RNA |
| La | Kinesin-1 (KHC) | Dynein (DIC) | Sumoylation | - | RNA granules |
| JIP3/JIP4 | Kinesin-1 (KHC tail and KLC) | Dynactin (p150Glued) Dynein (DLIC) |
GTPase (ARF6) | JIP1, JNK | Signaling endosomes, synaptic vesicles |
| JIP1 | Kinesin-1 (KHC stalk, KHC tail, and KLC) |
Dynactin (p150Glued) | Phosphorylation (JNK) Degradation | FEZ1, JIP3 | APP-positive vesicles, autophagosomes, synaptic vesicles, mitochondria |
| Mint1 | Kinesin-2 (KIF17 tail) | - | Phosphorylation of KIF17 tail (CaMKII) | CASK, Velis | NMDA receptors |
| DENN/MADD | Kinesin-3 (KIF1B stalk) | - | GTPase (Rab3) | - | Synaptic vesicles |
| Hook | Kinesin-3 | Dynein | - | - | Early endosomes |
Highlights.
Organelle-bound scaffolding proteins regulate opposing kinesin and dynein motors.
Scaffolding proteins also bind upstream regulatory kinases or G-proteins.
Scaffolding proteins integrate signaling with coordinate control of motor activity.
These modules allow for sustained transport and rapid response to local environment.
Acknowledgments
We thank Alison Twelvetrees, Armen Moughamian, and Yvette Wong for critical reading of the manuscript. This work was supported by NIH grants F31NS073262 to MMF and GM48661 ELFH.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Mallik R, et al. Teamwork in microtubule motors. Trends Cell Biol. 2013;23:575–582. doi: 10.1016/j.tcb.2013.06.003. [DOI] [PubMed] [Google Scholar]
- 2.Encalada SE, et al. Stable kinesin and dynein assemblies drive the axonal transport of mammalian prion protein vesicles. Cell. 2011;144:551–565. doi: 10.1016/j.cell.2011.01.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hendricks AG, et al. Motor coordination via a tug-of-war mechanism drives bidirectional vesicle transport. Curr Biol. 2010;20:697–702. doi: 10.1016/j.cub.2010.02.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Maday S, et al. Autophagosomes initiate distally and mature during transport toward the cell soma in primary neurons. J Cell Biol. 2012;196:407–417. doi: 10.1083/jcb.201106120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Soppina V, et al. Tug-of-war between dissimilar teams of microtubule motors regulates transport and fission of endosomes. Proc Natl Acad Sci U S A. 2009;106:19381–19386. doi: 10.1073/pnas.0906524106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hendricks AG, et al. Force measurements on cargoes in living cells reveal collective dynamics of microtubule motors. Proc Natl Acad Sci U S A. 2012;109:18447–18452. doi: 10.1073/pnas.1215462109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Szpankowski L, et al. Subpixel colocalization reveals amyloid precursor protein-dependent kinesin-1 and dynein association with axonal vesicles. Proc Natl Acad Sci U S A. 2012;109:8582–8587. doi: 10.1073/pnas.1120510109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Muller MJ, et al. Tug-of-war as a cooperative mechanism for bidirectional cargo transport by molecular motors. Proc Natl Acad Sci U S A. 2008;105:4609–4614. doi: 10.1073/pnas.0706825105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Fu M-M, Holzbaur ELF. LC3 binding to the scaffolding protein JIP1 regulates processive dynein-driven transport of autophagosomes. Dev Cell. 2014:29. doi: 10.1016/j.devcel.2014.04.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dixit R, et al. Differential regulation of dynein and kinesin motor proteins by tau. Science. 2008;319:1086–1089. doi: 10.1126/science.1152993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Blehm BH, et al. In vivo optical trapping indicates kinesin’s stall force is reduced by dynein during intracellular transport. Proc Natl Acad Sci U S A. 2013;110:3381–3386. doi: 10.1073/pnas.1219961110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hammond JW, et al. Tubulin modifications and their cellular functions. Curr Opin Cell Biol. 2008;20:71–76. doi: 10.1016/j.ceb.2007.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hammond JW, et al. Posttranslational modifications of tubulin and the polarized transport of kinesin-1 in neurons. Mol Biol Cell. 2010;21:572–583. doi: 10.1091/mbc.E09-01-0044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Janke C, Bulinski JC. Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat Rev Mol Cell Biol. 2011;12:773–786. doi: 10.1038/nrm3227. [DOI] [PubMed] [Google Scholar]
- 15.Nakata T, et al. Preferential binding of a kinesin-1 motor to GTP-tubulin-rich microtubules underlies polarized vesicle transport. J Cell Biol. 2011;194:245–255. doi: 10.1083/jcb.201104034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Vershinin M, et al. Multiple-motor based transport and its regulation by Tau. Proc Natl Acad Sci U S A. 2007;104:87–92. doi: 10.1073/pnas.0607919104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.McVicker DP, et al. The nucleotide-binding state of microtubules modulates kinesin processivity and the ability of Tau to inhibit kinesin-mediated transport. J Biol Chem. 2011;286:42873–42880. doi: 10.1074/jbc.M111.292987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Barlan K, et al. The microtubule-binding protein ensconsin is an essential cofactor of kinesin-1. Curr Biol. 2013;23:317–322. doi: 10.1016/j.cub.2013.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Balint S, et al. Correlative live-cell and superresolution microscopy reveals cargo transport dynamics at microtubule intersections. Proc Natl Acad Sci U S A. 2013;110:3375–3380. doi: 10.1073/pnas.1219206110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ross JL, et al. Cargo transport: molecular motors navigate a complex cytoskeleton. Curr Opin Cell Biol. 2008;20:41–47. doi: 10.1016/j.ceb.2007.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zajac AL, et al. Local cytoskeletal and organelle interactions impact molecular-motor- driven early endosomal trafficking. Curr Biol. 2013;23:1173–1180. doi: 10.1016/j.cub.2013.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Friedman JR, et al. Endoplasmic reticulum-endosome contact increases as endosomes traffic and mature. Mol Biol Cell. 2013;24:1030–1040. doi: 10.1091/mbc.E12-10-0733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Jordens I, et al. The Rab7 effector protein RILP controls lysosomal transport by inducing the recruitment of dynein-dynactin motors. Curr Biol. 2001;11:1680–1685. doi: 10.1016/s0960-9822(01)00531-0. [DOI] [PubMed] [Google Scholar]
- 24.Johansson M, et al. Activation of endosomal dynein motors by stepwise assembly of Rab7-RILP-p150Glued, ORP1L, and the receptor betalll spectrin. J Cell Biol. 2007;176:459–471. doi: 10.1083/jcb.200606077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Holleran EA, et al. beta III spectrin binds to the Arp1 subunit of dynactin. J Biol Chem. 2001;276:36598–36605. doi: 10.1074/jbc.M104838200. [DOI] [PubMed] [Google Scholar]
- 26.Rai AK, et al. Molecular adaptations allow dynein to generate large collective forces inside cells. Cell. 2013;152:172–182. doi: 10.1016/j.cell.2012.11.044. [DOI] [PubMed] [Google Scholar]
- 27.Rocha N, et al. Cholesterol sensor ORP1L contacts the ER protein VAP to control Rab7-RILP-p150 Glued and late endosome positioning. J Cell Biol. 2009;185:1209–1225. doi: 10.1083/jcb.200811005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hong Z, et al. The retromer component SNX6 interacts with dynactin p150(Glued) and mediates endosome-to-TGN transport. Cell Res. 2009;19:1334–1349. doi: 10.1038/cr.2009.130. [DOI] [PubMed] [Google Scholar]
- 29.Wassmer T, et al. The retromer coat complex coordinates endosomal sorting and dynein-mediated transport, with carrier recognition by the trans-Golgi network. Dev Cell. 2009;17:110–122. doi: 10.1016/j.devcel.2009.04.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Niu Y, et al. PtdIns(4)P regulates retromer-motor interaction to facilitate dynein-cargo dissociation at the trans-Golgi network. Nat Cell Biol. 2013;15:417–429. doi: 10.1038/ncb2710. [DOI] [PubMed] [Google Scholar]
- 31.MacAskill AF, et al. Mitochondrial trafficking and the provision of energy and calcium buffering at excitatory synapses. Eur J Neurosci. 2010;32:231–240. doi: 10.1111/j.1460-9568.2010.07345.x. [DOI] [PubMed] [Google Scholar]
- 32.Misgeld T, et al. Imaging axonal transport of mitochondria in vivo. Nat Methods. 2007;4:559–561. doi: 10.1038/nmeth1055. [DOI] [PubMed] [Google Scholar]
- 33.Bilsland LG, et al. Deficits in axonal transport precede ALS symptoms in vivo. Proc Natl Acad Sci U S A. 2010;107:20523–20528. doi: 10.1073/pnas.1006869107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Glater EE, et al. Axonal transport of mitochondria requires milton to recruit kinesin heavy chain and is light chain independent. J Cell Biol. 2006;173:545–557. doi: 10.1083/jcb.200601067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wang X, Schwarz TL. The mechanism of Ca2+ -dependent regulation of kinesin-mediated mitochondrial motility. Cell. 2009;136:163–174. doi: 10.1016/j.cell.2008.11.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Macaskill AF, et al. Miro1 is a calcium sensor for glutamate receptor-dependent localization of mitochondria at synapses. Neuron. 2009;61:541–555. doi: 10.1016/j.neuron.2009.01.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang X, et al. PINK1 and Parkin target Miro for phosphorylation and degradation to arrest mitochondrial motility. Cell. 2011;147:893–906. doi: 10.1016/j.cell.2011.10.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.van Spronsen M, et al. TRAK/Milton motor-adaptor proteins steer mitochondrial trafficking to axons and dendrites. Neuron. 2013;77:485–502. doi: 10.1016/j.neuron.2012.11.027. [DOI] [PubMed] [Google Scholar]
- 39.Kang JS, et al. Docking of axonal mitochondria by syntaphilin controls their mobility and affects short-term facilitation. Cell. 2008;132:137–148. doi: 10.1016/j.cell.2007.11.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Courchet J, et al. Terminal Axon Branching Is Regulated by the LKB1-NUAK1 Kinase Pathway via Presynaptic Mitochondrial Capture. Cell. 2013;153:1510–1525. doi: 10.1016/j.cell.2013.05.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Chen Y, Sheng ZH. Kinesin-1-syntaphilin coupling mediates activity-dependent regulation of axonal mitochondrial transport. J Cell Biol. 2013;202:351–364. doi: 10.1083/jcb.201302040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Caviston JP, et al. Huntingtin facilitates dynein/dynactin-mediated vesicle transport. Proc Natl Acad Sci U S A. 2007;104:10045–10050. doi: 10.1073/pnas.0610628104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Twelvetrees AE, et al. Delivery of GABAARs to synapses is mediated by HAP1-KIF5 and disrupted by mutant huntingtin. Neuron. 2010;65:53–65. doi: 10.1016/j.neuron.2009.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.McGuire JR, et al. Interaction of Huntingtin-associated protein-1 with kinesin light chain: implications in intracellular trafficking in neurons. J Biol Chem. 2006;281:3552–3559. doi: 10.1074/jbc.M509806200. [DOI] [PubMed] [Google Scholar]
- 45.Engelender S, et al. Huntingtin-associated protein 1 (HAP1) interacts with the p150Glued subunit of dynactin. Hum Mol Genet. 1997;6:2205–2212. doi: 10.1093/hmg/6.13.2205. [DOI] [PubMed] [Google Scholar]
- 46.Li SH, et al. Interaction of huntingtin-associated protein with dynactin P150Glued. J Neurosci. 1998;18:1261–1269. doi: 10.1523/JNEUROSCI.18-04-01261.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Caviston JP, et al. Huntingtin coordinates the dynein-mediated dynamic positioning of endosomes and lysosomes. Mol Biol Cell. 2011;22:478–492. doi: 10.1091/mbc.E10-03-0233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gunawardena S, et al. Disruption of axonal transport by loss of huntingtin or expression of pathogenic polyQ proteins in Drosophila. Neuron. 2003;40:25–40. doi: 10.1016/s0896-6273(03)00594-4. [DOI] [PubMed] [Google Scholar]
- 49.Her LS, Goldstein LS. Enhanced sensitivity of striatal neurons to axonal transport defects induced by mutant huntingtin. J Neurosci. 2008;28:13662–13672. doi: 10.1523/JNEUROSCI.4144-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Colin E, et al. Huntingtin phosphorylation acts as a molecular switch for anterograde/retrograde transport in neurons. EMBO J. 2008;27:2124–2134. doi: 10.1038/emboj.2008.133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wong YC, Holzbaur EL. The Regulation of Autophagosome Dynamics by Huntingtin and HAP1 Is Disrupted by Expression of Mutant Huntingtin, Leading to Defective Cargo Degradation. J Neurosci. 2014;34:1293–1305. doi: 10.1523/JNEUROSCI.1870-13.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ma B, et al. Huntingtin mediates dendritic transport of beta-actin mRNA in rat neurons. Scientific reports. 2011;1:140. doi: 10.1038/srep00140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Liot G, et al. Mutant Huntingtin alters retrograde transport of TrkB receptors in striatal dendrites. J Neurosci. 2013;33:6298–6309. doi: 10.1523/JNEUROSCI.2033-12.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Sahlender DA, et al. Optineurin links myosin VI to the Golgi complex and is involved in Golgi organization and exocytosis. J Cell Biol. 2005;169:285–295. doi: 10.1083/jcb.200501162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Caviston JP, Holzbaur EL. Huntingtin as an essential integrator of intracellular vesicular trafficking. Trends Cell Biol. 2009;19:147–155. doi: 10.1016/j.tcb.2009.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zala D, et al. Phosphorylation of mutant huntingtin at S421 restores anterograde and retrograde transport in neurons. Hum Mol Genet. 2008;17:3837–3846. doi: 10.1093/hmg/ddn281. [DOI] [PubMed] [Google Scholar]
- 57.Morfini G, et al. JNK mediates pathogenic effects of polyglutamine-expanded androgen receptor on fast axonal transport. Nat Neurosci. 2006;9:907–916. doi: 10.1038/nn1717. [DOI] [PubMed] [Google Scholar]
- 58.Morfini GA, et al. Pathogenic huntingtin inhibits fast axonal transport by activating JNK3 and phosphorylating kinesin. Nat Neurosci. 2009;12:864–871. doi: 10.1038/nn.2346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Deberg HA, et al. Motor domain phosphorylation modulates Kinesin-1 transport. J Biol Chem. 2013;288:32612–32621. doi: 10.1074/jbc.M113.515510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.van Niekerk EA, et al. Sumoylation in axons triggers retrograde transport of the RNA-binding protein La. Proc Natl Acad Sci U S A. 2007;104:12913–12918. doi: 10.1073/pnas.0611562104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Whitmarsh AJ. The JIP family of MAPK scaffold proteins. Biochemical Society transactions. 2006;34:828–832. doi: 10.1042/BST0340828. [DOI] [PubMed] [Google Scholar]
- 62.Cavalli V, et al. Sunday Driver links axonal transport to damage signaling. J Cell Biol. 2005;168:775–787. doi: 10.1083/jcb.200410136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Abe N, et al. Sunday driver interacts with two distinct classes of axonal organelles. J Biol Chem. 2009;284:34628–34639. doi: 10.1074/jbc.M109.035022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Sun F, et al. Sunday Driver/JIP3 binds kinesin heavy chain directly and enhances its motility. EMBO J. 2011;30:3416–3429. doi: 10.1038/emboj.2011.229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Arimoto M, et al. The Caenorhabditis elegans JIP3 Protein UNC-16 Functions As an Adaptor to Link Kinesin-1 with Cytoplasmic Dynein. J Neurosci. 2011;31:2216–2224. doi: 10.1523/JNEUROSCI.2653-10.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Deinhardt K, et al. Rab5 and Rab7 control endocytic sorting along the axonal retrograde transport pathway. Neuron. 2006;52:293–305. doi: 10.1016/j.neuron.2006.08.018. [DOI] [PubMed] [Google Scholar]
- 67.Mitchell DJ, et al. Trk activation of the ERK1/2 kinase pathway stimulates intermediate chain phosphorylation and recruits cytoplasmic dynein to signaling endosomes for retrograde axonal transport. J Neurosci. 2012;32:15495–15510. doi: 10.1523/JNEUROSCI.5599-11.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Huang SH, et al. JIP3 mediates TrkB axonal anterograde transport and enhances BDNF signaling by directly bridging TrkB with kinesin-1. J Neurosci. 2011;31:10602–10614. doi: 10.1523/JNEUROSCI.0436-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Montagnac G, et al. ARF6 Interacts with JIP4 to control a motor switch mechanism regulating endosome traffic in cytokinesis. Curr Biol. 2009;19:184–195. doi: 10.1016/j.cub.2008.12.043. [DOI] [PubMed] [Google Scholar]
- 70.Horiuchi D, et al. APLIP1, a kinesin binding JIP-1/JNK scaffold protein, influences the axonal transport of both vesicles and mitochondria in Drosophila. Curr Biol. 2005;15:2137–2141. doi: 10.1016/j.cub.2005.10.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Muresan Z, Muresan V. c-Jun NH2-terminal kinase-interacting protein-3 facilitates phosphorylation and controls localization of amyloid-beta precursor protein. J Neurosci. 2005;25:3741–3751. doi: 10.1523/JNEUROSCI.0152-05.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Fu MM, Holzbaur EL. JIP1 regulates the directionality of APP axonal transport by coordinating kinesin and dynein motors. J Cell Biol. 2013;202:495–508. doi: 10.1083/jcb.201302078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Verhey KJ, et al. Cargo of kinesin identified as JIP scaffolding proteins and associated signaling molecules. J Cell Biol. 2001;152:959–970. doi: 10.1083/jcb.152.5.959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Matsuda S, et al. c-Jun N-terminal kinase (JNK)-interacting protein-1b/islet-brain-1 scaffolds Alzheimer’s amyloid precursor protein with JNK. J Neurosci. 2001;21:6597–6607. doi: 10.1523/JNEUROSCI.21-17-06597.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Scheinfeld MH, et al. Jun NH2-terminal kinase (JNK) interacting protein 1 (JIP1) binds the cytoplasmic domain of the Alzheimer’s beta-amyloid precursor protein (APP) J Biol Chem. 2002;277:3767–3775. doi: 10.1074/jbc.M108357200. [DOI] [PubMed] [Google Scholar]
- 76.Nihalani D, et al. Recruitment of JNK to JIP1 and JNK-dependent JIP1 phosphorylation regulates JNK module dynamics and activation. J Biol Chem. 2003;278:28694–28702. doi: 10.1074/jbc.M304212200. [DOI] [PubMed] [Google Scholar]
- 77.Willoughby EA, et al. The JNK-interacting protein-1 scaffold protein targets MAPK phosphatase-7 to dephosphorylate JNK. J Biol Chem. 2003;278:10731–10736. doi: 10.1074/jbc.M207324200. [DOI] [PubMed] [Google Scholar]
- 78.Blasius TL, et al. Two binding partners cooperate to activate the molecular motor Kinesin-1. J Cell Biol. 2007;176:11–17. doi: 10.1083/jcb.200605099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Kristensen O, et al. A unique set of SH3-SH3 interactions controls IB1 homodimerization. EMBO J. 2006;25:785–797. doi: 10.1038/sj.emboj.7600982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Hammond JW, et al. Co-operative versus independent transport of different cargoes by Kinesin-1. Traffic. 2008;9:725–741. doi: 10.1111/j.1600-0854.2008.00722.x. [DOI] [PubMed] [Google Scholar]
- 81.Muresan Z, Muresan V. Coordinated transport of phosphorylated amyloid-beta precursor protein and c-Jun NH2-terminal kinase-interacting protein-1. J Cell Biol. 2005;171:615–625. doi: 10.1083/jcb.200502043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Hammond JW, et al. Autoinhibition of the kinesin-2 motor KIF17 via dual intramolecular mechanisms. J Cell Biol. 2010;189:1013–1025. doi: 10.1083/jcb.201001057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Hammond JW, et al. Mammalian Kinesin-3 motors are dimeric in vivo and move by processive motility upon release of autoinhibition. PLoS Biol. 2009;7:e72. doi: 10.1371/journal.pbio.1000072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Setou M, et al. Kinesin superfamily motor protein KIF17 and mLin-10 in NMDA receptor-containing vesicle transport. Science. 2000;288:1796–1802. doi: 10.1126/science.288.5472.1796. [DOI] [PubMed] [Google Scholar]
- 85.Guillaud L, et al. Disruption of KIF17-Mint1 interaction by CaMKII-dependent phosphorylation: a molecular model of kinesin-cargo release. Nat Cell Biol. 2008;10:19–29. doi: 10.1038/ncb1665. [DOI] [PubMed] [Google Scholar]
- 86.Yin X, et al. Regulation of NMDA receptor transport: a KIF17-cargo binding/releasing underlies synaptic plasticity and memory in vivo. J Neurosci. 2012;32:5486–5499. doi: 10.1523/JNEUROSCI.0718-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Deacon SW, et al. Dynactin is required for bidirectional organelle transport. J Cell Biol. 2003;160:297–301. doi: 10.1083/jcb.200210066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Berezuk MA, Schroer TA. Dynactin enhances the processivity of kinesin-2. Traffic. 2007;8:124–129. doi: 10.1111/j.1600-0854.2006.00517.x. [DOI] [PubMed] [Google Scholar]
- 89.Niwa S, et al. KIF1Bbeta- and KIF1A–mediated axonal transport of presynaptic regulator Rab3 occurs in a GTP-dependent manner through DENN/MADD. Nat Cell Biol. 2008;10:1269–1279. doi: 10.1038/ncb1785. [DOI] [PubMed] [Google Scholar]
- 90.Zhang J, et al. HookA is a novel dynein-early endosome linker critical for cargo movement in vivo. J Cell Biol. 2014;204:1009–1026. doi: 10.1083/jcb.201308009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Bielska E, et al. Hook is an adapter that coordinates kinesin-3 and dynein cargo attachment on early endosomes. J Cell Biol. 2014;204:989–1007. doi: 10.1083/jcb.201309022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Walenta JH, et al. The Golgi-associated hook3 protein is a member of a novel family of microtubule-binding proteins. J Cell Biol. 2001;152:923–934. doi: 10.1083/jcb.152.5.923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Maldonado-Baez L, et al. Microtubule-dependent endosomal sorting of clathrin-independent cargo by Hook1. J Cell Biol. 2013;201:233–247. doi: 10.1083/jcb.201208172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Xu L, et al. An FTS/Hook/p107(FHIP) complex interacts with and promotes endosomal clustering by the homotypic vacuolar protein sorting complex. Mol Biol Cell. 2008;19:5059–5071. doi: 10.1091/mbc.E08-05-0473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Hirokawa N, et al. Kinesin superfamily motor proteins and intracellular transport. Nat Rev Mol Cell Biol. 2009;10:682–696. doi: 10.1038/nrm2774. [DOI] [PubMed] [Google Scholar]
- 96.Kanai Y, et al. KIF5C, a novel neuronal kinesin enriched in motor neurons. J Neurosci. 2000;20:6374–6384. doi: 10.1523/JNEUROSCI.20-17-06374.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Nakajima K, et al. Molecular motor KIF5A is essential for GABA(A) receptor transport, and KIF5A deletion causes epilepsy. Neuron. 2012;76:945–961. doi: 10.1016/j.neuron.2012.10.012. [DOI] [PubMed] [Google Scholar]
- 98.Verhey KJ, Hammond JW. Traffic control: regulation of kinesin motors. Nat Rev Mol Cell Biol. 2009;10:765–777. doi: 10.1038/nrm2782. [DOI] [PubMed] [Google Scholar]
- 99.Stock MF, et al. Formation of the compact confomer of kinesin requires a COOH-terminal heavy chain domain and inhibits microtubule-stimulated ATPase activity. J Biol Chem. 1999;274:14617–14623. doi: 10.1074/jbc.274.21.14617. [DOI] [PubMed] [Google Scholar]
- 100.Friedman DS, Vale RD. Single-molecule analysis of kinesin motility reveals regulation by the cargo-binding tail domain. Nat Cell Biol. 1999;1:293–297. doi: 10.1038/13008. [DOI] [PubMed] [Google Scholar]
- 101.Hirokawa N, et al. Submolecular domains of bovine brain kinesin identified by electron microscopy and monoclonal antibody decoration. Cell. 1989;56:867–878. doi: 10.1016/0092-8674(89)90691-0. [DOI] [PubMed] [Google Scholar]
- 102.Hackney DD, et al. Half-site inhibition of dimeric kinesin head domains by monomeric tail domains. Biochemistry. 2009;48:3448–3456. doi: 10.1021/bi8022575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Kaan HY, et al. The structure of the kinesin-1 motor-tail complex reveals the mechanism of autoinhibition. Science. 2011;333:883–885. doi: 10.1126/science.1204824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Roberts AJ, et al. Functions and mechanics of dynein motor proteins. Nat Rev Mol Cell Biol. 2013;14:713–726. doi: 10.1038/nrm3667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Ross JL, et al. Processive bidirectional motion of dynein-dynactin complexes in vitro. Nat Cell Biol. 2006;8:562–570. doi: 10.1038/ncb1421. [DOI] [PubMed] [Google Scholar]
- 106.Kardon JR, Vale RD. Regulators of the cytoplasmic dynein motor. Nat Rev Mol Cell Biol. 2009;10:854–865. doi: 10.1038/nrm2804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Holzbaur EL, et al. Homology of a 150K cytoplasmic dynein-associated polypeptide with the Drosophila gene Glued. Nature. 1991;351:579–583. doi: 10.1038/351579a0. [DOI] [PubMed] [Google Scholar]
- 108.Waterman-Storer CM, et al. The p150Glued component of the dynactin complex binds to both microtubules and the actin-related protein centractin (Arp-1) Proc Natl Acad Sci U S A. 1995;92:1634–1638. doi: 10.1073/pnas.92.5.1634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Culver-Hanlon TL, et al. A microtubule-binding domain in dynactin increases dynein processivity by skating along microtubules. Nat Cell Biol. 2006;8:264–270. doi: 10.1038/ncb1370. [DOI] [PubMed] [Google Scholar]
- 110.Karki S, Holzbaur EL. Affinity chromatography demonstrates a direct binding between cytoplasmic dynein and the dynactin complex. J Biol Chem. 1995;270:28806–28811. doi: 10.1074/jbc.270.48.28806. [DOI] [PubMed] [Google Scholar]
- 111.Vaughan KT, Vallee RB. Cytoplasmic dynein binds dynactin through a direct interaction between the intermediate chains and p150Glued. J Cell Biol. 1995;131:1507–1516. doi: 10.1083/jcb.131.6.1507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Siglin AE, et al. Dynein and dynactin leverage their bivalent character to form a high-affinity interaction. PLoS One. 2013;8:e59453. doi: 10.1371/journal.pone.0059453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Schroer TA. Dynactin. Annu Rev Cell Dev Biol. 2004;20:759–779. doi: 10.1146/annurev.cellbio.20.012103.094623. [DOI] [PubMed] [Google Scholar]
- 114.Verhey KJ, et al. Light chain-dependent regulation of Kinesin’s interaction with microtubules. J Cell Biol. 1998;143:1053–1066. doi: 10.1083/jcb.143.4.1053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Diefenbach RJ, et al. The C-terminal region of the stalk domain of ubiquitous human kinesin heavy chain contains the binding site for kinesin light chain. Biochemistry. 1998;37:16663–16670. doi: 10.1021/bi981163r. [DOI] [PubMed] [Google Scholar]
- 116.Hackney DD, et al. Kinesin undergoes a 9 S to 6 S conformational transition. J Biol Chem. 1992;267:8696–8701. [PubMed] [Google Scholar]
- 117.Palacios IM, St Johnston D. Kinesin light chain-independent function of the Kinesin heavy chain in cytoplasmic streaming and posterior localisation in the Drosophila oocyte. Development. 2002;129:5473–5485. doi: 10.1242/dev.00119. [DOI] [PubMed] [Google Scholar]
- 118.Pernigo S, et al. Structural basis for kinesin-1:cargo recognition. Science. 2013;340:356–359. doi: 10.1126/science.1234264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Williams LS, et al. The auto-inhibitory domain and ATP-independent microtubule-binding region of Kinesin heavy chain are major functional domains for transport in the Drosophila germline. Development. 2014;141:176–186. doi: 10.1242/dev.097592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Vagnoni A, et al. Phosphorylation of kinesin light chain 1 at serine 460 modulates binding and trafficking of calsyntenin-1. J Cell Sci. 2011;124:1032–1042. doi: 10.1242/jcs.075168. [DOI] [PMC free article] [PubMed] [Google Scholar]