Abstract
This paper documents on-going work to facilitate ITK-based processing and 3D Slicer scene management in ParaView. We believe this will broaden the use of ParaView for high performance computing and visualization in the medical imaging research community. The effort is focused on developing ParaView plug-ins for managing VTK structures from 3D Slicer MRML scenes and encapsulating ITK filters for deployment in ParaView. In this paper, we present KWScene, an open source cross-platform library that is being developed to support implementation of these types of plugins. We describe the overall design of the library and provide implementation details and conclude by presenting a concrete example that demonstrates the use of the KWScene library in computational anatomy research at Johns Hopkins Center for Imaging Science.
1 Introduction
ParaView [1] and 3D Slicer [2] are both free, open source analysis and visualization applications but with different focuses. ParaView is designed to handle large-sized datasets with a focus on interactive scientific visualization. It can run on distributed and shared memory parallel as well as single processor systems. 3D Slicer provides an extensive suite of segmentation and registration algorithms for medical images. Slicer uses a Medical Reality Markup Language ( MRML) , a library for managing medical image data types (volumes, models, transforms, fiducials, Cameras, etc) and their visualization. Exchanging scene data between the two applications would provide medical imaging researchers a huge opportunity to take advantage of the unique functionalities available in each application. However, this is not possible currently because of a different scene representation. In addition, ITK-based processing is not easily and readily available within ParaView.
This technical report discusses an on-going work to facilitate ITK-based processing and 3D Slicer scene management in ParaView. The effort focuses on developing KWScene, a library that provides functionalities for managing VTK structures from 3D Slicer MRML scenes.
2 KWScene
KWScene is being developed based on the MRML standard to support scene graph-type manipulation of volume and surface primitives for VTK-based visualization and ITK-based processing.
2.1 Design
The overall design of KWScene consists of three modules
- Input/Output
- The purpose of this module is to read MRML XML scene files and extract objects and their parameters to generate an internal scene graph representation i.e kwScene.
- Scene Graph Internal Representation
- The purpose of this module is to encapsulate the scene graph. It contains a map of node names and pointers to the node. Access to the nodes is provided using iterators.
- Generators
- The purpose of this module is to generate the appropriate ITK, VTK and ParaView filters from the parsed MRML data.
The processing pipeline involves three steps ( shown in Figure 1 )
Step 1: Read and parse XML data elements from MRML Scene files.
Step 2: Parse the node data elements and generate internal representation of the Scene graph (KWScene)
Step 3: For each parsed node element, identify ITK, VTK or ParaView classes.
Figure 1.

Processing pipeline
2.2 Implementation
Scene: Encapsulating Scene Graph
kwsNode is the base class for all nodes. We define two types of nodes: ShapedNodes and ShapelessNodes. The ShapedNodes are all nodes that incorporate geometric information. We have implemented the following types of shaped nodes:
ImageNode
Landmark
LandmarkListNode
SurfaceNode
The following shapeless nodes are implemented:
CompositeNode
RepresentationNode
TransformNode
IO: MRML Reader, Parser, Writer
XMLReaderBase encapsulates the XML2 library and defines the methods needed in derived classes to receive the contents of an XML file in a structured manner. The reader outputs a KWScene object.
MRMLReader reads MRML scene files and instantiates and invokes node parsers. MRMLNodeParser is an abstract class used to parse MRML node read from the MRML scene file. Several classes are derived from this class such as MRMLImageNodeParser and MRMLSurfaceNodeParser.
MRMLNodeWriter is an abstract class used to write MRML node to a scene file. Several classes are derived from this class such MRMLImageNodeWriter and MRMLSurfaceNodeWriter.
Generators
Generators provide an interface for constructing appropriate VTK, ITK, and ParaView objects that are needed to manage data associated with a given KWScene node.
- ITK ( ITKGenerator ):
- ■ ITK surface
- ■ ITK image
- ■ Landmark
- VTK (VTKGenerator)
- ■ Image
- ■ Surface
- ■ Landmark
ParaView
2.3 KWScene Code Management and Build Instructions
KWScene is an open-source project and is hosted in a public SVN repository https://www.kitware.com/svn/KWPublic/trunk/KWScene
ITK, VTK and external ParaView builds are required. The following version are recommended.
- ParaView
- Version 3.10.1
- Build ParaView with the SHARED_LIBRARY option turned on
- ITK
- Use ITK version 3.20
- Turn on the ITK_USE_REVIEW option
- VTK
- Use VTK version 5.6.1
A project-dedicated Wiki is available for collaborative documentation http://public.kitware.com/Wiki/KWScene
3 KWScene for Computational Anatomy Research
Computational anatomy [4] involves the analysis of the shapes of structures in the human body. Researchers at the Center for Imaging Science (CIS) at John Hopkins University try to correlate the shape of subcortical structures in brains with disease. Quantifying and visualizing structural change can provide insight on the effects and progression of diseases, and possibly help with patient diagnosis and prognosis.
CIS and Washington University researchers used typical computationl anatomy techniques to group patients with Dementia of the Alzheimer's Type (DAT) from control subjects [5]. As a control subject was first selected to act as an “Atlas” or “Template”, and the template hippocampus was manually segmented from the MR image. Large Deformation Diffeomorphic Metric Mapping (LDDMM) [6] is used to create segmented images of each subject hippocampus. A triangulated surface is created from each subject to create the comparison data sets, and each data set is transformed to the template coordinate space by LDDMM surface matching via currents [7]. These transformations are encoded in the initial momentum data at each vertex of the template for each subject. All the data necessary to make the statistical comparisons of the population is in the initial momentum data. In this study, the principal components of the momentum data are computed and used to determine the direction of greatest variability of the deformation. The researchers were able to use the left hippocampus results to distinguish between control and mildly cognitive impaired subjects in the population to a high degree of statistical significance.
In a study such as this, data originally captured in MR scans goes through a pipeline of processing where intermediate results are produced and stored, typically in free format directory structures of the user's own design. Several data types are used in computational anatomy and they include:
Volumes
Point Data (for landmarks)
Mesh Data for surfaces
Each of these types of data could have vector or scalar values associated with each voxel or point (as in the case of LDDMM output momentum data). Using KWScene, we are able to capture the entire pipeline, in an organized way that is compatible with ParaView (with the KWScene plugin) and Slicer3D.
As part of a multiple-year NIH project, CIS is performing analysis similar to [5], while additionally adding parts of our Computational Anatomy pipeline to ITK. As the pipeline components continue to come on line, their functionality can be added to ParaView under the KWScene plugin, and KWScene will be available to store all results.
In the following example, a small population of subject caudates are compared to a template caudate created by automated template estimation. For this use case, intermediate results from a part of the pipeline being created for are loaded and stored in a KWScene. After starting ParaView, click Tools->Manage Plugins, and select the KWScene plugin to load, as shown in Figure 2. After each subject surface has been transformed via LDDMM into the coordinate space of the template surface, each initial momentum set (a surface with vector values at each vertex) is loaded into ParaView, as seen in Figure 2. The vectors that define the transform can be visualized on the surfaces by coloring the Momentum Point Data set.
Figure 2.
Loading KWScene plugin to ParaView
As in [5], to determine the areas of greatest variability in the set of transforms, a calculation of the Principal Components is used on the Momentum vectors. The results are loaded into ParaView, as shown in Figure 4.
Figure 4.
Visualizing the principal components
This intermediate step performed in this small study can be saved as a MRML based scene for future processing and/or Slicer visualization. Click File->Save into a KWScene, and select a file to save the scene into.
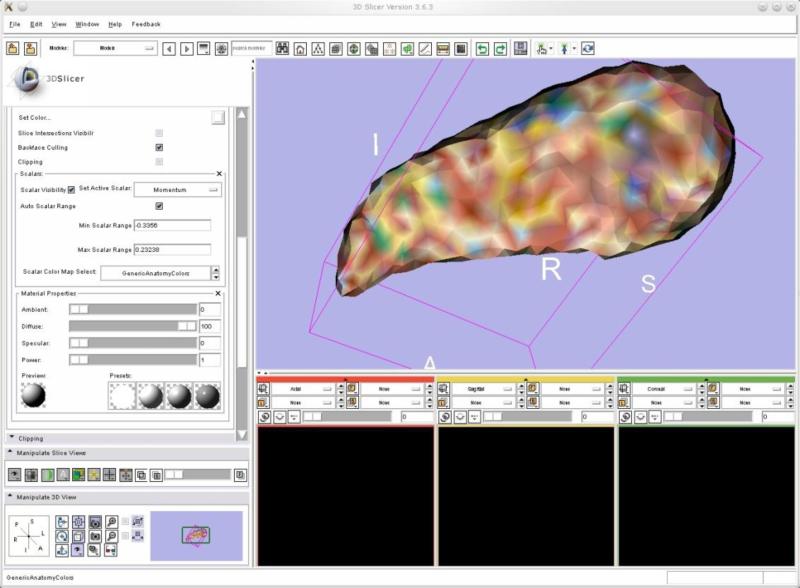
After the file is saved, its contents can be viewed and worked on by loading it into Slicer3D. To color the data, go to the Models module and to the Hierarchy and Display tab. In the Model Display Section, select the Mesh you would like to view and check the Visibility box. In the Scalars section, check the Scalar Visibility box and Set the Active Scalar (Figure 5 shows the deformed template with its momentums).
Figure 5.
Visualizing the deformed template created within ParaView in Slicer 3.6
4 Summary and Future Work
This paper presents KWScene, an open source library, that is useful to for managing VTK structures from 3D Slicer MRML scenes and encapsulating ITK filters for deployment in ParaView. We believe this will broaden the use of ParaView for high performance computing and visualization in the medical imaging research community. The future work in this project focuses on adding more generators for different types of VTK and ITK data structures to the library, applying the library in other computational anatomy applications and improving the overall software quality.
Figure 3.
Visualizing the deformation templates in ParaView
Acknowledgment
This research is supported in part by NIH grants R01-EB008171-01A1 and P41-RR015241. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Reference
- 1. http://www.paraview.org/
- 2.Pieper S, Lorensen B, Schroeder W, Kikinis R. The NA-MIC Kit: ITK, VTK, Pipelines, Grids and 3D Slicer as An Open Platform for the Medical Image Computing Community. Proceedings of IEEE International Symposium on Biomedical Imaging: From Nano to Macro 2006. 2006:698–701. [Google Scholar]
- 3.Ibanez Luis, Schroeder Will, Ng Lydia, Cates Josh. The ITK Software Guide: The Insight Segmentation and Registration Toolkit. Kitware Inc.; 2006. [Google Scholar]
- 4.Grenander U, Miller MI. Computational anatomy: an emerging discipline. Q. Appl. Math. LVI. 1998;4:617–694. (Dec. 1998) [Google Scholar]
- 5.Beg Mirza Faisal, Miller Michael I., Trouvé Alain, Younes Laurent. Computing Large Deformation Metric Mappings via Geodesic Flows of Diffeomorphisms. International Journal of Computer Vision. 2005;61(2):139–157. [Google Scholar]
- 6.Wang Lei, Beg Faisal, Ratnanather Tilak, Ceritoglu Can, Younes Laurent, Morris John C., Csernansky John G., Miller Michael I. Large Deformation Diffeomorphism and Momentum Based Hippocampal Shape Discrimination in Dementia of the Alzheimer Type. IEEE Transactions on Medical Imaging. 2007 Apr;264 doi: 10.1109/TMI.2005.853923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Vaillant Marc, Glaunes Joan. Surface Matching via Currents. Information Processing for Medical Imaging. 2005 doi: 10.1007/11505730_32. [DOI] [PubMed] [Google Scholar]