Abstract
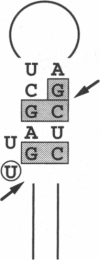
A 17-amino acid arginine-rich peptide from the bovine immunodeficiency virus Tat protein has been shown to bind with high affinity and specificity to bovine immunodeficiency virus transactivation response element (TAR) RNA, making contacts in the RNA major groove near a bulge. We show that, as in other peptide-RNA complexes, arginine and threonine side chains make important contributions to binding but, unexpectedly, that one isoleucine and three glycine residues also are critical. The isoleucine side chain may intercalate into a hydrophobic pocket in the RNA. Glycine residues may allow the peptide to bind deeply within the RNA major groove and may help determine the conformation of the peptide. Similar features have been observed in protein-DNA and drug-DNA complexes in the DNA minor groove, including hydrophobic interactions and binding deep within the groove, suggesting that the major groove of RNA and minor groove of DNA may share some common recognition features.
Full text
PDF

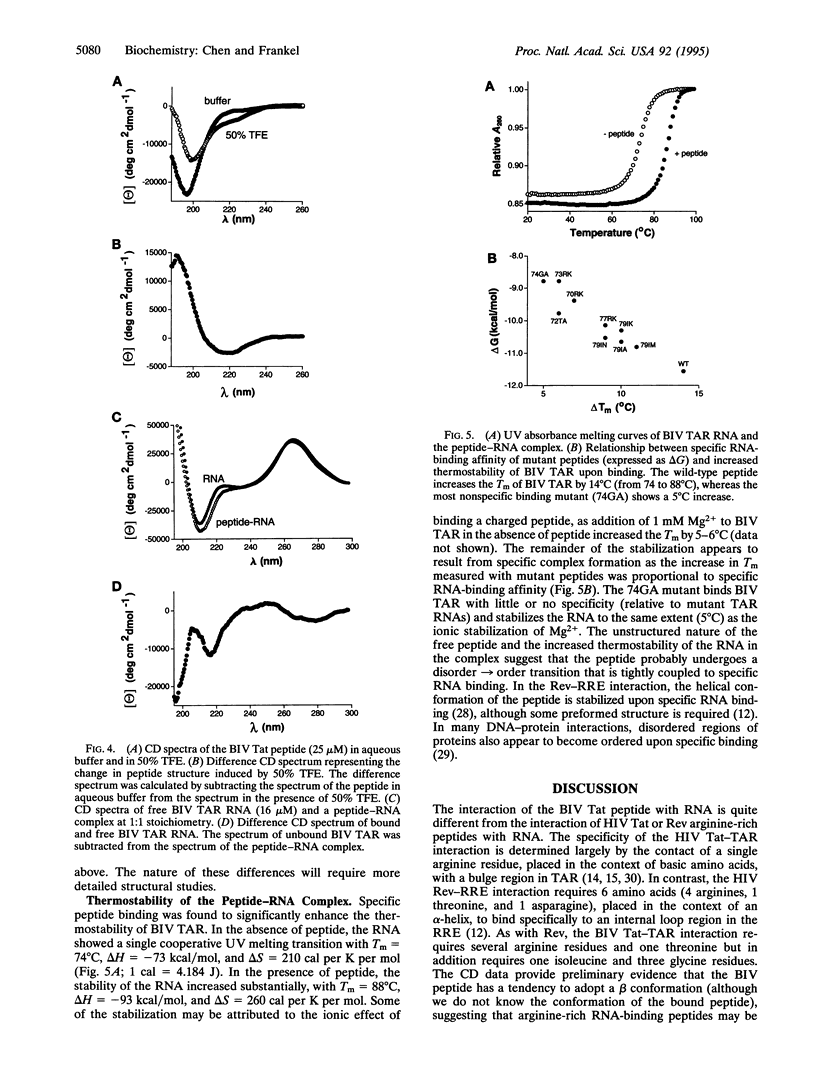

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Battiste J. L., Tan R., Frankel A. D., Williamson J. R. Binding of an HIV Rev peptide to Rev responsive element RNA induces formation of purine-purine base pairs. Biochemistry. 1994 Mar 15;33(10):2741–2747. doi: 10.1021/bi00176a001. [DOI] [PubMed] [Google Scholar]
- Biou V., Yaremchuk A., Tukalo M., Cusack S. The 2.9 A crystal structure of T. thermophilus seryl-tRNA synthetase complexed with tRNA(Ser). Science. 1994 Mar 11;263(5152):1404–1410. doi: 10.1126/science.8128220. [DOI] [PubMed] [Google Scholar]
- Blanco F. J., Jiménez M. A., Pineda A., Rico M., Santoro J., Nieto J. L. NMR solution structure of the isolated N-terminal fragment of protein-G B1 domain. Evidence of trifluoroethanol induced native-like beta-hairpin formation. Biochemistry. 1994 May 17;33(19):6004–6014. doi: 10.1021/bi00185a041. [DOI] [PubMed] [Google Scholar]
- Burd C. G., Dreyfuss G. Conserved structures and diversity of functions of RNA-binding proteins. Science. 1994 Jul 29;265(5172):615–621. doi: 10.1126/science.8036511. [DOI] [PubMed] [Google Scholar]
- Calnan B. J., Biancalana S., Hudson D., Frankel A. D. Analysis of arginine-rich peptides from the HIV Tat protein reveals unusual features of RNA-protein recognition. Genes Dev. 1991 Feb;5(2):201–210. doi: 10.1101/gad.5.2.201. [DOI] [PubMed] [Google Scholar]
- Calnan B. J., Tidor B., Biancalana S., Hudson D., Frankel A. D. Arginine-mediated RNA recognition: the arginine fork. Science. 1991 May 24;252(5009):1167–1171. doi: 10.1126/science.252.5009.1167. [DOI] [PubMed] [Google Scholar]
- Cavarelli J., Rees B., Ruff M., Thierry J. C., Moras D. Yeast tRNA(Asp) recognition by its cognate class II aminoacyl-tRNA synthetase. Nature. 1993 Mar 11;362(6416):181–184. doi: 10.1038/362181a0. [DOI] [PubMed] [Google Scholar]
- Chen L., Frankel A. D. An RNA-binding peptide from bovine immunodeficiency virus Tat protein recognizes an unusual RNA structure. Biochemistry. 1994 Mar 8;33(9):2708–2715. doi: 10.1021/bi00175a046. [DOI] [PubMed] [Google Scholar]
- Churcher M. J., Lamont C., Hamy F., Dingwall C., Green S. M., Lowe A. D., Butler J. G., Gait M. J., Karn J. High affinity binding of TAR RNA by the human immunodeficiency virus type-1 tat protein requires base-pairs in the RNA stem and amino acid residues flanking the basic region. J Mol Biol. 1993 Mar 5;230(1):90–110. doi: 10.1006/jmbi.1993.1128. [DOI] [PubMed] [Google Scholar]
- Fackelmayer F. O., Richter A. Purification of two isoforms of hnRNP-U and characterization of their nucleic acid binding activity. Biochemistry. 1994 Aug 30;33(34):10416–10422. doi: 10.1021/bi00200a024. [DOI] [PubMed] [Google Scholar]
- Geierstanger B. H., Volkman B. F., Kremer W., Wemmer D. E. Short peptide fragments derived from HMG-I/Y proteins bind specifically to the minor groove of DNA. Biochemistry. 1994 May 3;33(17):5347–5355. doi: 10.1021/bi00183a043. [DOI] [PubMed] [Google Scholar]
- Jasanoff A., Fersht A. R. Quantitative determination of helical propensities from trifluoroethanol titration curves. Biochemistry. 1994 Mar 1;33(8):2129–2135. doi: 10.1021/bi00174a020. [DOI] [PubMed] [Google Scholar]
- Kim J. L., Nikolov D. B., Burley S. K. Co-crystal structure of TBP recognizing the minor groove of a TATA element. Nature. 1993 Oct 7;365(6446):520–527. doi: 10.1038/365520a0. [DOI] [PubMed] [Google Scholar]
- Kim Y., Geiger J. H., Hahn S., Sigler P. B. Crystal structure of a yeast TBP/TATA-box complex. Nature. 1993 Oct 7;365(6446):512–520. doi: 10.1038/365512a0. [DOI] [PubMed] [Google Scholar]
- King C. Y., Weiss M. A. The SRY high-mobility-group box recognizes DNA by partial intercalation in the minor groove: a topological mechanism of sequence specificity. Proc Natl Acad Sci U S A. 1993 Dec 15;90(24):11990–11994. doi: 10.1073/pnas.90.24.11990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lazinski D., Grzadzielska E., Das A. Sequence-specific recognition of RNA hairpins by bacteriophage antiterminators requires a conserved arginine-rich motif. Cell. 1989 Oct 6;59(1):207–218. doi: 10.1016/0092-8674(89)90882-9. [DOI] [PubMed] [Google Scholar]
- Majerfeld I., Yarus M. An RNA pocket for an aliphatic hydrophobe. Nat Struct Biol. 1994 May;1(5):287–292. doi: 10.1038/nsb0594-287. [DOI] [PubMed] [Google Scholar]
- Mattaj I. W. RNA recognition: a family matter? Cell. 1993 Jun 4;73(5):837–840. doi: 10.1016/0092-8674(93)90265-r. [DOI] [PubMed] [Google Scholar]
- Mujeeb A., Bishop K., Peterlin B. M., Turck C., Parslow T. G., James T. L. NMR structure of a biologically active peptide containing the RNA-binding domain of human immunodeficiency virus type 1 Tat. Proc Natl Acad Sci U S A. 1994 Aug 16;91(17):8248–8252. doi: 10.1073/pnas.91.17.8248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oubridge C., Ito N., Evans P. R., Teo C. H., Nagai K. Crystal structure at 1.92 A resolution of the RNA-binding domain of the U1A spliceosomal protein complexed with an RNA hairpin. Nature. 1994 Dec 1;372(6505):432–438. doi: 10.1038/372432a0. [DOI] [PubMed] [Google Scholar]
- Peterson R. D., Bartel D. P., Szostak J. W., Horvath S. J., Feigon J. 1H NMR studies of the high-affinity Rev binding site of the Rev responsive element of HIV-1 mRNA: base pairing in the core binding element. Biochemistry. 1994 May 10;33(18):5357–5366. doi: 10.1021/bi00184a001. [DOI] [PubMed] [Google Scholar]
- Puglisi J. D., Tan R., Calnan B. J., Frankel A. D., Williamson J. R. Conformation of the TAR RNA-arginine complex by NMR spectroscopy. Science. 1992 Jul 3;257(5066):76–80. doi: 10.1126/science.1621097. [DOI] [PubMed] [Google Scholar]
- Puglisi J. D., Tinoco I., Jr Absorbance melting curves of RNA. Methods Enzymol. 1989;180:304–325. doi: 10.1016/0076-6879(89)80108-9. [DOI] [PubMed] [Google Scholar]
- Reardon B. J., Winters R. S., Gordon D., Winter E. A peptide motif that recognizes A.T tracts in DNA. Proc Natl Acad Sci U S A. 1993 Dec 1;90(23):11327–11331. doi: 10.1073/pnas.90.23.11327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rould M. A., Perona J. J., Steitz T. A. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature. 1991 Jul 18;352(6332):213–218. doi: 10.1038/352213a0. [DOI] [PubMed] [Google Scholar]
- Rould M. A., Perona J. J., Söll D., Steitz T. A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA(Gln) and ATP at 2.8 A resolution. Science. 1989 Dec 1;246(4934):1135–1142. doi: 10.1126/science.2479982. [DOI] [PubMed] [Google Scholar]
- Rubin J., Sundaralingam M. An unexpected major groove binding of netropsin and distamycin A to tRNA(phe). J Biomol Struct Dyn. 1984 Aug;2(1):165–174. doi: 10.1080/07391102.1984.10507555. [DOI] [PubMed] [Google Scholar]
- Ruff M., Krishnaswamy S., Boeglin M., Poterszman A., Mitschler A., Podjarny A., Rees B., Thierry J. C., Moras D. Class II aminoacyl transfer RNA synthetases: crystal structure of yeast aspartyl-tRNA synthetase complexed with tRNA(Asp). Science. 1991 Jun 21;252(5013):1682–1689. doi: 10.1126/science.2047877. [DOI] [PubMed] [Google Scholar]
- Schumacher M. A., Choi K. Y., Zalkin H., Brennan R. G. Crystal structure of LacI member, PurR, bound to DNA: minor groove binding by alpha helices. Science. 1994 Nov 4;266(5186):763–770. doi: 10.1126/science.7973627. [DOI] [PubMed] [Google Scholar]
- Siomi H., Choi M., Siomi M. C., Nussbaum R. L., Dreyfuss G. Essential role for KH domains in RNA binding: impaired RNA binding by a mutation in the KH domain of FMR1 that causes fragile X syndrome. Cell. 1994 Apr 8;77(1):33–39. doi: 10.1016/0092-8674(94)90232-1. [DOI] [PubMed] [Google Scholar]
- Spolar R. S., Record M. T., Jr Coupling of local folding to site-specific binding of proteins to DNA. Science. 1994 Feb 11;263(5148):777–784. doi: 10.1126/science.8303294. [DOI] [PubMed] [Google Scholar]
- Tan R., Chen L., Buettner J. A., Hudson D., Frankel A. D. RNA recognition by an isolated alpha helix. Cell. 1993 Jun 4;73(5):1031–1040. doi: 10.1016/0092-8674(93)90280-4. [DOI] [PubMed] [Google Scholar]
- Tan R., Frankel A. D. Circular dichroism studies suggest that TAR RNA changes conformation upon specific binding of arginine or guanidine. Biochemistry. 1992 Oct 27;31(42):10288–10294. doi: 10.1021/bi00157a016. [DOI] [PubMed] [Google Scholar]
- Tan R., Frankel A. D. Costabilization of peptide and RNA structure in an HIV Rev peptide-RRE complex. Biochemistry. 1994 Dec 6;33(48):14579–14585. doi: 10.1021/bi00252a025. [DOI] [PubMed] [Google Scholar]
- Tao J., Frankel A. D. Electrostatic interactions modulate the RNA-binding and transactivation specificities of the human immunodeficiency virus and simian immunodeficiency virus Tat proteins. Proc Natl Acad Sci U S A. 1993 Feb 15;90(4):1571–1575. doi: 10.1073/pnas.90.4.1571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tao J., Frankel A. D. Specific binding of arginine to TAR RNA. Proc Natl Acad Sci U S A. 1992 Apr 1;89(7):2723–2726. doi: 10.1073/pnas.89.7.2723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valegård K., Murray J. B., Stockley P. G., Stonehouse N. J., Liljas L. Crystal structure of an RNA bacteriophage coat protein-operator complex. Nature. 1994 Oct 13;371(6498):623–626. doi: 10.1038/371623a0. [DOI] [PubMed] [Google Scholar]
- Weeks K. M., Crothers D. M. Major groove accessibility of RNA. Science. 1993 Sep 17;261(5128):1574–1577. doi: 10.1126/science.7690496. [DOI] [PubMed] [Google Scholar]
- Weeks K. M., Crothers D. M. RNA recognition by Tat-derived peptides: interaction in the major groove? Cell. 1991 Aug 9;66(3):577–588. doi: 10.1016/0092-8674(81)90020-9. [DOI] [PubMed] [Google Scholar]
- Zhong L., Johnson W. C., Jr Environment affects amino acid preference for secondary structure. Proc Natl Acad Sci U S A. 1992 May 15;89(10):4462–4465. doi: 10.1073/pnas.89.10.4462. [DOI] [PMC free article] [PubMed] [Google Scholar]