Abstract
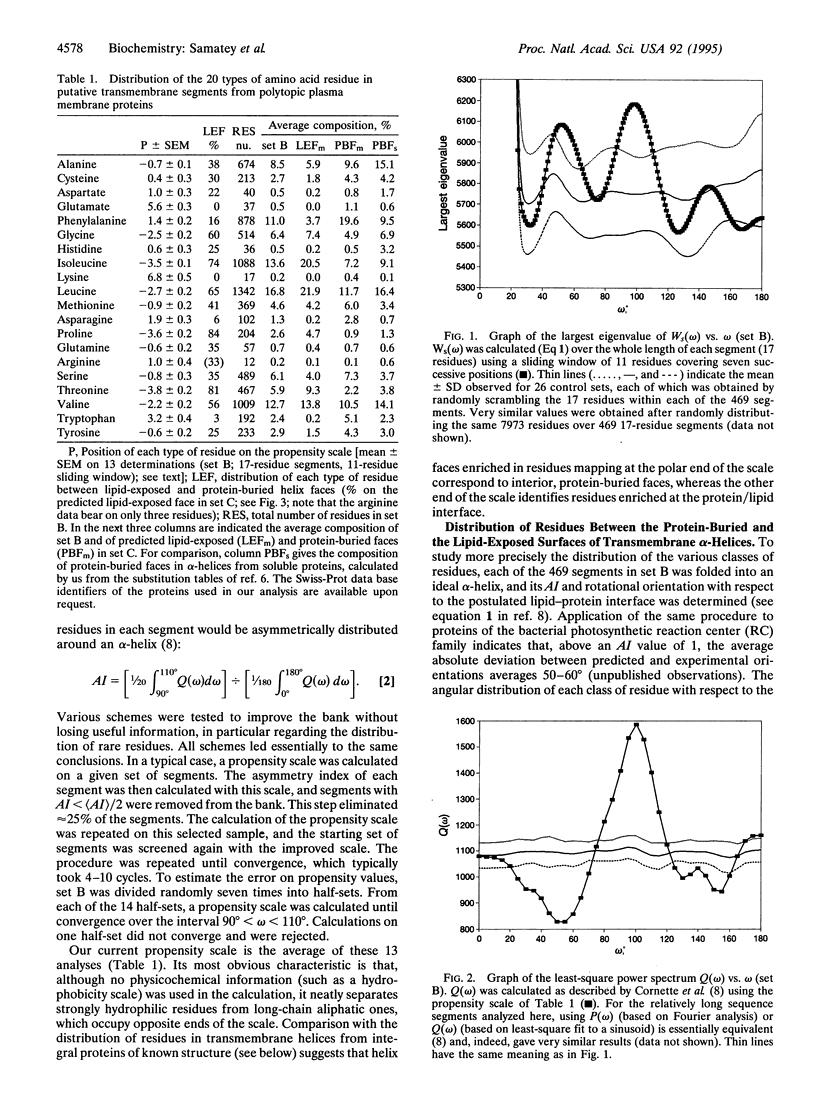
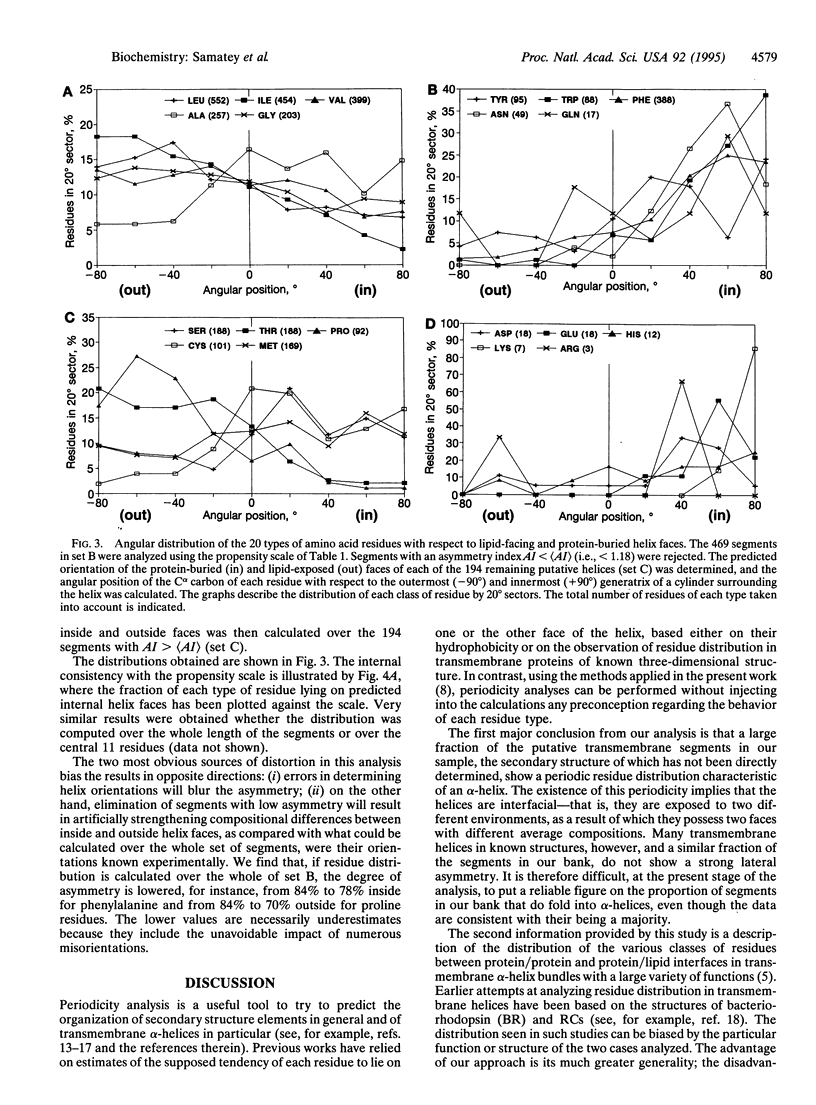
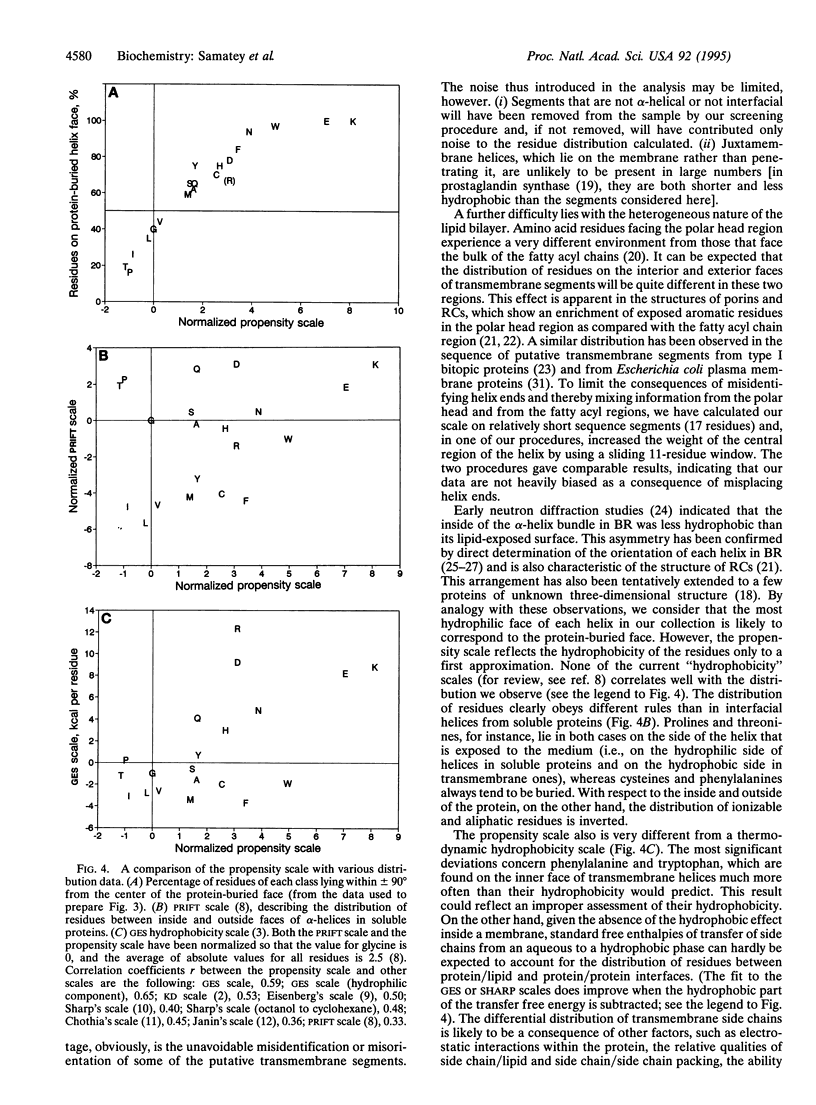
The periodic distribution of residues in the sequence of 469 putative transmembrane alpha-helices from eukaryotic plasma membrane polytopic proteins has been analyzed with correlation matrices. The method does not involve any a priori assumption about the secondary structure of the segments or about the physicochemical properties of individual amino acid residues. Maximal correlation is observed at 3.6 residues per period, characteristic of alpha-helices. A scale extracted from the data describes the propensity of the various residues to lie on the same or on opposite helix faces. The most polar face of transmembrane helices, presumably that buried in the protein core, shows a strong enrichment in aromatic residues, while residues likely to face the fatty acyl chains of lipids are largely aliphatic.
Full text
PDF

Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Chothia C. The nature of the accessible and buried surfaces in proteins. J Mol Biol. 1976 Jul 25;105(1):1–12. doi: 10.1016/0022-2836(76)90191-1. [DOI] [PubMed] [Google Scholar]
- Cornette J. L., Cease K. B., Margalit H., Spouge J. L., Berzofsky J. A., DeLisi C. Hydrophobicity scales and computational techniques for detecting amphipathic structures in proteins. J Mol Biol. 1987 Jun 5;195(3):659–685. doi: 10.1016/0022-2836(87)90189-6. [DOI] [PubMed] [Google Scholar]
- Deisenhofer J., Michel H. Nobel lecture. The photosynthetic reaction centre from the purple bacterium Rhodopseudomonas viridis. EMBO J. 1989 Aug;8(8):2149–2170. doi: 10.1002/j.1460-2075.1989.tb08338.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donnelly D., Overington J. P., Ruffle S. V., Nugent J. H., Blundell T. L. Modeling alpha-helical transmembrane domains: the calculation and use of substitution tables for lipid-facing residues. Protein Sci. 1993 Jan;2(1):55–70. doi: 10.1002/pro.5560020106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisenberg D. Three-dimensional structure of membrane and surface proteins. Annu Rev Biochem. 1984;53:595–623. doi: 10.1146/annurev.bi.53.070184.003115. [DOI] [PubMed] [Google Scholar]
- Engelman D. M., Steitz T. A., Goldman A. Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Annu Rev Biophys Biophys Chem. 1986;15:321–353. doi: 10.1146/annurev.bb.15.060186.001541. [DOI] [PubMed] [Google Scholar]
- Engelman D. M., Zaccai G. Bacteriorhodopsin is an inside-out protein. Proc Natl Acad Sci U S A. 1980 Oct;77(10):5894–5898. doi: 10.1073/pnas.77.10.5894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finer-Moore J., Stroud R. M. Amphipathic analysis and possible formation of the ion channel in an acetylcholine receptor. Proc Natl Acad Sci U S A. 1984 Jan;81(1):155–159. doi: 10.1073/pnas.81.1.155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson R., Baldwin J. M., Ceska T. A., Zemlin F., Beckmann E., Downing K. H. Model for the structure of bacteriorhodopsin based on high-resolution electron cryo-microscopy. J Mol Biol. 1990 Jun 20;213(4):899–929. doi: 10.1016/S0022-2836(05)80271-2. [DOI] [PubMed] [Google Scholar]
- Janin J. Surface and inside volumes in globular proteins. Nature. 1979 Feb 8;277(5696):491–492. doi: 10.1038/277491a0. [DOI] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Landolt-Marticorena C., Williams K. A., Deber C. M., Reithmeier R. A. Non-random distribution of amino acids in the transmembrane segments of human type I single span membrane proteins. J Mol Biol. 1993 Feb 5;229(3):602–608. doi: 10.1006/jmbi.1993.1066. [DOI] [PubMed] [Google Scholar]
- Overington J., Donnelly D., Johnson M. S., Sali A., Blundell T. L. Environment-specific amino acid substitution tables: tertiary templates and prediction of protein folds. Protein Sci. 1992 Feb;1(2):216–226. doi: 10.1002/pro.5560010203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Picot D., Loll P. J., Garavito R. M. The X-ray crystal structure of the membrane protein prostaglandin H2 synthase-1. Nature. 1994 Jan 20;367(6460):243–249. doi: 10.1038/367243a0. [DOI] [PubMed] [Google Scholar]
- Popot J. L., Engelman D. M., Gurel O., Zaccaï G. Tertiary structure of bacteriorhodopsin. Positions and orientations of helices A and B in the structural map determined by neutron diffraction. J Mol Biol. 1989 Dec 20;210(4):829–847. doi: 10.1016/0022-2836(89)90111-3. [DOI] [PubMed] [Google Scholar]
- Popot J. L., Engelman D. M. Membrane protein folding and oligomerization: the two-stage model. Biochemistry. 1990 May 1;29(17):4031–4037. doi: 10.1021/bi00469a001. [DOI] [PubMed] [Google Scholar]
- Popot J. L., de Vitry C. On the microassembly of integral membrane proteins. Annu Rev Biophys Biophys Chem. 1990;19:369–403. doi: 10.1146/annurev.bb.19.060190.002101. [DOI] [PubMed] [Google Scholar]
- Rees D. C., DeAntonio L., Eisenberg D. Hydrophobic organization of membrane proteins. Science. 1989 Aug 4;245(4917):510–513. doi: 10.1126/science.2667138. [DOI] [PubMed] [Google Scholar]
- Rees D. C., Komiya H., Yeates T. O., Allen J. P., Feher G. The bacterial photosynthetic reaction center as a model for membrane proteins. Annu Rev Biochem. 1989;58:607–633. doi: 10.1146/annurev.bi.58.070189.003135. [DOI] [PubMed] [Google Scholar]
- Samatey F. A., Zaccaï G., Engelman D. M., Etchebest C., Popot J. L. Rotational orientation of transmembrane alpha-helices in bacteriorhodopsin. A neutron diffraction study. J Mol Biol. 1994 Mar 4;236(4):1093–1104. doi: 10.1016/0022-2836(94)90014-0. [DOI] [PubMed] [Google Scholar]
- Sharp K. A., Nicholls A., Friedman R., Honig B. Extracting hydrophobic free energies from experimental data: relationship to protein folding and theoretical models. Biochemistry. 1991 Oct 8;30(40):9686–9697. doi: 10.1021/bi00104a017. [DOI] [PubMed] [Google Scholar]
- Sun S., Parthasarathy R. Protein sequence and structure relationship ARMA spectral analysis: application to membrane proteins. Biophys J. 1994 Jun;66(6):2092–2106. doi: 10.1016/S0006-3495(94)81004-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss M. S., Abele U., Weckesser J., Welte W., Schiltz E., Schulz G. E. Molecular architecture and electrostatic properties of a bacterial porin. Science. 1991 Dec 13;254(5038):1627–1630. doi: 10.1126/science.1721242. [DOI] [PubMed] [Google Scholar]
- von Heijne G., Gavel Y. Topogenic signals in integral membrane proteins. Eur J Biochem. 1988 Jul 1;174(4):671–678. doi: 10.1111/j.1432-1033.1988.tb14150.x. [DOI] [PubMed] [Google Scholar]
- von Heijne G. Membrane proteins: from sequence to structure. Annu Rev Biophys Biomol Struct. 1994;23:167–192. doi: 10.1146/annurev.bb.23.060194.001123. [DOI] [PubMed] [Google Scholar]
- von Heijne G. Proline kinks in transmembrane alpha-helices. J Mol Biol. 1991 Apr 5;218(3):499–503. doi: 10.1016/0022-2836(91)90695-3. [DOI] [PubMed] [Google Scholar]



