Abstract
The role of STAT2 in mediating the antigrowth effects of type I interferon (IFN) is well-documented in vitro. Yet evidence of IFN-activated STAT2 as having tumor suppressor function in vivo and participation in antitumor immunity is lacking. Here we show in a syngeneic tumor transplantation model that STAT2 reduces tumor growth. Stat2−/− mice formed larger tumors compared to wild type (WT) mice. IFN-β treatment of Stat2−/− mice did not cause tumor regression. Gene expression analysis revealed a small subset of immunomodulatory genes to be downregulated in tumors established in Stat2−/− mice. Additionally, we found tumor antigen cross-presentation by Stat2−/− dendritic cells to T cells to be impaired. Adoptive transfer of tumor antigen specific CD8+ T cells primed by Stat2−/− dendritic cells into tumor-bearing Stat2−/− mice did not induce tumor regression with IFN-β intervention. We observed that an increase in the number of CD4+ and CD8+ T cells in the draining lymph nodes of IFN-β-treated tumor-bearing WT mice was absent in IFN-β treated Stat2−/− mice. Thus our study provides evidence for further evaluation of STAT2 function in cancer patients receiving type I IFN based immunotherapy.
Keywords: STAT2, STAT1, interferon, melanoma, dendritic cell, antitumor, cross-presentation
Introduction
STAT2 is an essential component of type I interferon (IFN-I) signaling.1-3 IFN-I induces the activation and assembly of STAT2/STAT1 heterodimers that, together with the DNA binding protein IRF9, form the transcriptional complex ISGF3. ISGF3 activates the expression IFN-I target genes with antiviral, antiproliferative, and immunomodulatory activities.3-5 In vitro studies conducted in multiple tumor cell lines have demonstrated that STAT2 directly mediates the growth inhibitory and the pro-apoptotic effects of IFN-I (IFN-α/β) 6-9 leading to the premise that STAT2 has tumor suppressor function. Yet limited experimental data exist to confirm it.
The biological significance of STAT2-mediated IFN-I signaling in the immune response comes from observations made in Stat2−/− mice.10 In these mice, the IFN-I autocrine/paracrine loop that is critical in the increased production of IFN-I and subsequent antiviral activity is impaired. IFN-I does not block the growth of proliferating T cells.10-12 The expression of IFN-I target genes is also severely affected. Loss of STAT2, however, does not cause spontaneous tumor formation, but can be promoted by carcinogen exposure.13
Evidence that STAT2 has tumor suppressor activity in vivo comes from one single study using Stat2−/− transgenic mice with constitutive IFN-α production (GIFN/Stat2−/−) in the central nervous system (CNS). These mice died prematurely due to spontaneous development of medulloblastoma.14 In contrast, Stat1−/− mice with transgenic CNS production of IFN-α did not develop tumors. These mice eventually died, but due to other causes. These findings indicated that in the absence of STAT2, IFN-α was tumorigenic independently of STAT1.
IFN-I is essential in tumor immunosurveillance. IFN receptor (IFNAR1) null mice show enhanced tumor growth 15. IFN-I can target the tumor directly16, 17 but also indirectly. The indirect antitumor effects of IFN are demonstrated in transplanted tumors lacking IFNAR1, which are rejected by relying on the host hematopoietic compartment.18, 19 Recent studies now show that IFN-I signaling in dendritic cells (DCs) is pivotal in the development of antitumor immunity. Antigen cross-presentation by DCs to T cells is defective in Ifnar1−/− DC.20, 21 Although STAT2 has not been studied for this function, there is some evidence that STAT2 positively regulates the development and maturation of DCs22.
We now report that in the Stat2−/− mice, tumor growth was accelerated. These tumors showed reduced expression of key immunomodulatory factors. Loss of STAT2 also impaired the anti-tumorigenic effects of IFN-I in part via STAT2 signaling in DCs. Together, our study highlights the importance of a host STAT2/IFN-I axis in antitumor immunity.
Materials and Methods
Mice
129SvJ/Stat2−/− mice provided by Dr. Christian Schindler (Columbia University) were backcrossed onto the C57/BL6 genetic background for 11 generations (>99.7% purity). C57/BL6 wild type mice were purchased from the Animal Production Area of the Frederick National Laboratory for Cancer Research (Frederick, MD). Mice were bred in our animal facility and housed under a pathogen-free environment. C57/BL6/Pmel-1 transgenic mice were purchased from The Jackson Laboratory (Bar Harbor, Maine). All animal protocols were approved by Temple University Animal Care and Use Committee guidelines.
Tumor cell lines
Murine B16-F1 melanoma and MC38 colon adenocarcinoma cell lines were maintained in DMEM medium (Mediatech, Inc; Herndon, VA) supplemented with 5% heat-inactivated FBS, 2 mM L-glutamine, 1mM sodium pyruvate, 100 U/mL penicillin and 100 μg/mL streptomycin (Invitrogen Corp; Carlsbad, CA) at 37°C and 5% CO2.
Antibodies and cytokines
Anti-STAT1 antibody (E-23) and anti-STAT2 (07-140) antibody were purchased from Santa Cruz Biotechnologies (Santa Cruz, CA) and Millipore (Billerica, MA), respectively. Anti-STAT2 antibody for immunofluorescence analysis was provided by Dr. Christian Schindler. Anti-STAT3 antibody was provided by Dr. Andrew Larner (Virginia Commonwealth University). Anti-pY701-STAT1, anti-mouse CD16/CD32 (clone 2.4G2), anti-CD3-FITC (Clone: 17A2), CD8α-APC (Clone: 53-6.7), CD4-PE-Cy7 (Clone: RM4-5), B220-PE (Clone: RA3-6B2), Vβ13-FITC (clone: MR12-3) and CD25-Percp-Cy5.5 (Clone: P61) were purchased from BD Biosciences (San Jose, CA). Anti-pY705-STAT3 antibody and anti-Actin antibody were purchased from Cell Signaling (Danvers, MA) and Abcam (Cambridge, MA), respectively. HRP-conjugated secondary antibodies were purchased from Invitrogen (Carlsbad, CA). FITC-anti-Rabbit IgG antibody and PE-anti-Mouse IgG (Biolegend; San Diego, CA) were used for immunocytochemistry and flow cytometry analysis. Foxp3-PE (Clone: FJk-16s) was purchased from eBioscience (San Diego, CA). Recombinant murine IFN-β was provided by Biogen-Idec.
Tumor transplantation
One million tumor cells in 200 μL of endotoxin free saline solution were injected subcutaneously (s.c.) on the dorsal flank of 6-8 weeks old C57BL/6 WT or Stat2−/− mice. Three days later mice received IFN-β (2×104U) in and around the site of tumor transplantation twice weekly in 100 μL of PBS intratumorally. Tumor measurements were started at day 7 using a digital caliper. Tumor volume was determined with the formula: V= a2b, where a is the shorter diameter and b is the longer diameter of the tumor. For pulmonary metastasis, 5×105 B16-F1 tumor cells were injected into the tail vein of C57BL/6 WT or Stat2−/− mice. On day 20, animals were sacrificed and lungs and livers were harvested, preserved in Bouins fixative solution, and metastatic colonies counted. For antibody mediated T-cell depletion, each mice received an intraperitoneal injection of 150 μg of anti-Thy-1 antibody (M5/49; BioXcell, West Lebanon, NH) three days prior to tumor inoculation and every five days thereafter. T cell depletion was confirmed by flow cytometry analysis.
Generation of bone marrow-derived dendritic cells (BMDCs)
Bone marrow precursors were harvested from femurs and tibias and depleted of red blood cells with ACK lysis reagent. Bone marrow cells were differentiated into DCs with complete IMDM medium (Iscove’s DMEM supplemented with 10% FBS, penicillin/streptomycin, gentamicin, and β-ME) enriched with 3.3 ng/mL of GM-CSF (BD Biosciences)23. At day 6, resting DC cultures were stimulated with 100 ng/mL of LPS 3755 (Sigma-Aldrich), and pulsed with 5 μg/mL of human peptide gp10025–33 (GenScript, Piscataway, NJ) for 2h at 37°C.
Antigen cross-presentation assay
Cells from spleens and peripheral lymph nodes from Pmel-1 transgenic (Tg) mice were isolated by smashing onto a 100 μm cell strainer and depleted of red blood cells by ACK treatment. Cells were washed, resuspended in complete IMDM and plated on 10 cm petri dishes for 1h at 37°C. The non-adherent lymphocytes, enriched in Tg T cells, were re-plated in 96-well U-bottom plates at 100,000 cells in 100 μL volume and co-cultured with BMDCs (10,000 cells/100 μL). Culture supernatants were collected 5 days later and tested for cytokine release. The primed Pmel-1 CD8+ T lymphocytes were subsequently used for adoptive transfer.
Adoptive transfer of Pmel-1 Tg T lymphocytes
Five million Pmel-1 Tg T cells primed with gp100 peptide-loaded BMDCs, were resuspended in endotoxin-free DPBS and injected i.v. in a volume of 500 μL in mice three days after tumor cell injection. The next day, mice received IFN-β (2×104 U) as described earlier, twice weekly. At the end of the study, draining inguinal and contralateral lymph nodes were harvested for flow cytometry analysis.
Cytokine measurement
Supernatants were analyzed in triplicate for mouse IFN-γ and IL-2 release using ELISA (BD OptEIA ELISA kits). Tumor-conditioned media collected from individual tumors (WT and Stat2−/− mice) were analyzed using the CBA Mouse Inflammation Kit (BD Biosciences; San Diego, CA).
Flow cytometry
Pmel-1 T Tg lymphocytes obtained before and after in vitro activation with BMDCs were incubated with anti-CD16/CD32 to block Fcγ receptors. Cells were then stained for T and B cell markers. Lymphocytes were also stained separately with anti-FITC-Vβ13 antibody to quantify the percentage of transgenic lymphocytes. Lymph nodes were stained for CD4, CD8α, CD25, and Foxp3 to analyze regulatory T cells. FlowJo software (Tree star) was used for data analysis.
Western blot analysis
Tumor cells were disrupted in lysis buffer as described previously24. Protein extracts were resolved on precast NuPAGE 4-12% gradient gels (Invitrogen) and transferred onto polyvinylidene difluoride membranes. Membranes were probed with the indicated primary antibody followed by incubation with HRP-conjugated secondary antibody. Membranes were developed using enhanced chemiluminescence reagent (GE healthcare; Piscataway, NJ) and images captured with Alpha-Innotech HD2 image system (San Leandro, CA). Quantification was carried out using AlphaView software (San Leandro, CA).
Quantitative real-time PCR
Total RNA was extracted using RNA Bee Reagent (Ambion; Lake Forest, CA). RT-PCR was performed as a two-step process using High Capacity cDNA Reverse Transcription (Applied Biosystems; Foster City, CA) and ran using a 7300 Real Time PCR system with primer sets from Taqman Gene Expression Assays following the manufacturer’s instructions. Quantitation of gene expression was calculated using the following equation for determining fold changes: 2 –(CT target gene – CT β-Actin). Actin served as a housekeeping gene.
Immunocytochemistry
Tumors were embedded in tissue preserving compound (Sakura Finetek, Torrance, CA) and snapped frozen in liquid nitrogen. Tissue sections of 5 μm thickness placed on glass slides were fixed in 100% methanol. Slides were blocked in 10% normal goat serum and incubated with anti-STAT1 (1:100) and anti-STAT2 antibody (1:2500). Signals were detected after incubation with FITC-anti-Rabbit IgG and PE-anti-Mouse IgG. Nuclei were counterstained with Vectashield mounting medium (Vector laboratories). Images were taken with a Nikon Eclipse TE2000-U image system (Nikon; Melville, N.Y.)
Tumor-conditioned media
Tumors were individually minced, weighed, and 100 mg of tumor tissue was cultured in 1 mL of serum free DMEM. Supernatants free of cell debris were collected 24h later and stored at –80°C.
Isolation of tumor cells
Tumors were minced into small pieces with scalpels. The dissociated tumor material was passed through a 330 μm filter bag (Nasco). The tumor cell suspension was further subjected to serial filtration through 100 μm and 40 μm cell strainers (Fisher). Cell debris was removed by Ficoll gradient centrifugation. Viable tumor cells were resuspended in complete DMEM medium and incubated at 37°C.
Microarray analysis
Total RNA was isolated using TRIzol reagent (Invitrogen, Carlsbad, CA) and further purified by RNeasy kit (Qiagen, Valencia, CA). The quality of total RNA was assessed using an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA). RNA samples were labeled and hybridized to the Affymetrix mouse whole-genome Gene 2.0 ST Array (Affymetrix, Santa Clara, CA). For each condition, three individual tumor samples were used for microarray experiments. Scanned microarray images were analyzed using the Affymetrix Gene Expression Console with RMA (Robust Multi-array Average) normalization algorithm. Further statistical Analyses were performed using BRB-ArrayTools 25. Gene classification into ontology categories (OG) was performed using BLAST2GO® version 2.6.4 (Biobam Bioinformatics, Valencia, Spain). Gene expression was validated by qRT- PCR using three independent samples different from the samples analyzed in Affymetrix microarray.
Proliferation assay
Tumor cells were incubated with medium or 1:2 diluted tumor-conditioned medium. Cell proliferation was assessed every 24h using CellTiter 96-Aqueous One Solution Reagent (Promega, Madison, WI). Absorbance was measured at 490 nm using a VictorX5 multilabel plate reader (Perkin Elmer, Waltham MA). Background values were subtracted prior to data analysis.
Statistical analysis
Prism software (GraphPad, San Diego) was used for statistical analysis. In vitro results were analyzed using the Student’s t-test to assess significance. In vivo data were analyzed using the Mann-Whitney U test. Values of p< 0.05 were considered statistically significant.
Results
Enhanced tumor growth and lung metastasis in Stat2−/− mice
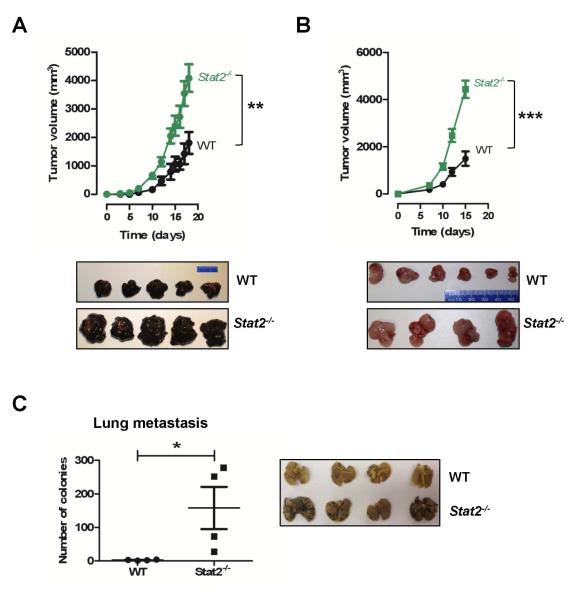
To assess differences in tumor formation, wild type (WT) and Stat2−/− mice received a single s.c. injection of B16-F1 melanoma cells. Stat2−/− mice developed progressively larger tumors (4100 mm3 ± 480 mm3) compared to WT mice (1800 mm3 ± 400 mm3; Fig. 1A and lower panel). Similarly, s.c. injection of MC38 colon adenocarcinoma cells in Stat2−/− mice caused the formation of larger tumors (4434 mm3 ± 807 mm3) compared to WT mice (1493 mm3 ± 272 mm3; Fig. 1B and lower panel). We next examined whether Stat2−/− mice would be more susceptible to metastasis. B16-F1 cells are poorly metastatic, and yet we observed a significant increase in the number of metastatic colonies in the lungs of Stat2−/− mice (157 ± 62) as opposed to WT mice (2.5 ± 0.6; Fig. 1C and right panel). No metastatic colonies in the livers were detected in both strains. Our findings indicate that STAT2 exhibits tumor suppressor function as it restricts tumor growth and metastasis.
Figure 1. Targeted deletion of host STAT2 enhances tumor growth and lung metastasis.
WT or Stat2−/− mice were injected with 1 × 106 (A) B16-F1 or (B) MC38 tumor cells. Data represent mean tumor volume ± SEM of one representative study conducted with wild type (WT) and Stat2−/− mice. Representative images of individual B16-F1 melanoma tumors or MC38 tumors harvested from WT and Stat2−/− mice at day 20 of the study. (C) WT and Stat2−/− mice received a tail vein injection of 5×105 B16-F1 tumor cells. Mice were sacrificed at day 20 and tumor colonies in the lungs were counted, (n=5). Representative image of individual lungs from wild type (WT) and Stat2−/− mice are shown. *, p< 0.05; **, p< 0.01; and ***, p< 0.001. n=8-10 animals/group.
Requirement of STAT2 in the antitumor effects of IFN-I
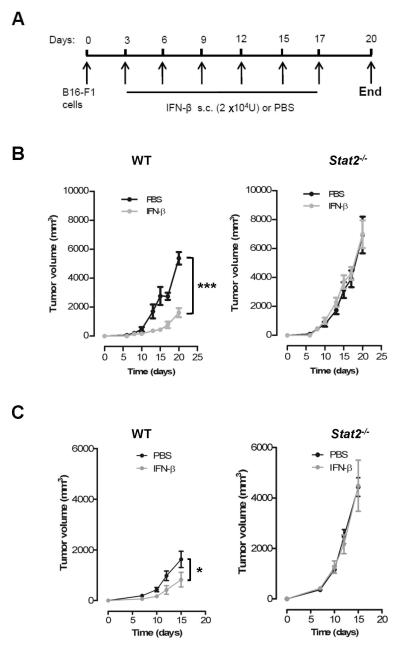
STAT2 mediates the antigrowth effects of IFN-I. Hence we evaluated the antitumor capacity of IFN-β in B16-F1 tumor-bearing WT and Stat2−/− mice. IFN-β treatment of WT mice caused a significant reduction in tumor size compared to the vehicle control group (Fig. 2B left panel; 1620 ± 325 mm3 vs. 5376 ± 470 mm3). In contrast, IFN-β treatment of Stat2−/− mice caused no tumor regression (Fig. 2B right panel; 6936±1240 mm3 vs. 6990±930 mm3 in IFN-β). Similar effects were observed in MC38 tumor-bearing WT and Stat2−/− mice (Fig. 2C). Thus our data show that STAT2 is critical in eliciting the antitumor effects of IFN-I.
Figure 2. The antitumor effects of IFN-I are abrogated in Stat2−/− mice.
(A) Animal study design and scheduling of IFN-β administration. WT and Stat2−/− mice received 1 × 106 (B) B16-F1 tumor cells or (C) MC38 tumor cells. Three days later mice received subcutaneous (s.c.) injections of PBS or IFN-β (2×104 U) around the site of tumor implantation. Tumor size was measured three times weekly. Values are shown as mean tumor volume ± SEM. *, p< 0.05; ***, p< 0.001. Data are representative of three independent experiments. n=8/group.
IFN-I targets tumor cells indirectly in a STAT2 dependent manner
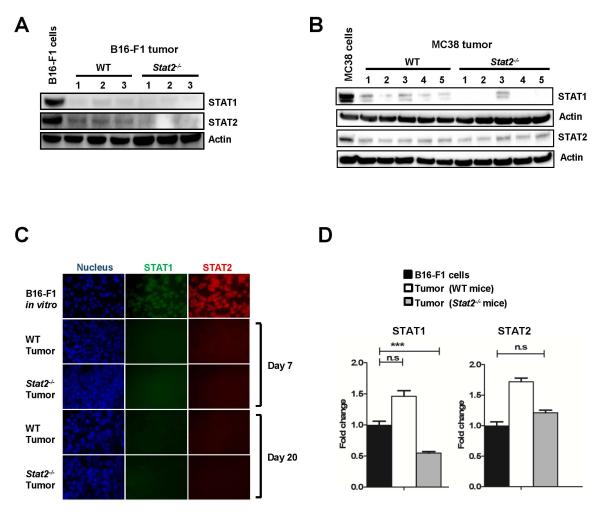
IFN-I promotes antitumor activity directly by engaging surface IFN receptors present on tumor cells and also indirectly by relying on the immune system.18, 26 Consequently, we compared by Western blot analysis STAT2 and STAT1 protein levels in individual B16-F1 and MC38 tumors harvested from WT and Stat2−/− mice. Both STAT2 and STAT1 within the tumors were similarly reduced in both strains of mice when compared against tumor cells expanded in vitro (Fig. 3A-B). Immunofluorescence staining of B16-F1 tumor sections indicated that at day 7 (when tumors are palpable) and until the end of the study at day 20, STAT2 and STAT1 proteins were barely detectable. (Fig. 3C). Images of the same tumor sections showed that STAT2 and STAT1 proteins were expressed at the peripheral edges of WT tumors indicative of host STAT1/2 expression (Fig. S1). Moreover, we found that within the tumors established in Stat2−/− mice, mRNA levels of STAT1 were decreased by 50% whereas STAT2 mRNA levels remained unchanged (Fig. 3D). In contrast, no significant changes were detected in tumors established in WT mice. IFN-β treatment, however, restored STAT1 and STAT2 expression, but only in tumors established in WT mice (Fig. S2). Thus our data show that loss of tumor STAT2 and STAT1 proteins in the WT and Stat2−/− host are regulated differently.
Figure 3. Dramatic decrease of STAT2/STAT1 within tumors.
Western blot analysis of STAT1 and STAT2 in individual (A) B16-F1 and (B) MC38 tumors harvested from WT or Stat2−/− mice on day 20. Tumor cells expanded in vitro represent the basal level of STAT proteins. (C) Tumors harvested on the indicated days were evaluated for STAT1 (green) and STAT2 (red) by immunofluorescence staining. Nuclei are blue. Images are representative of at least three sections from three individual tumors. (D) mRNA levels of STAT1 and STAT2 measured in six individual B16-F1 tumors harvested from WT and Stat2−/− mice on day 20 by qRT-PCR. Data are shown as the mean fold change determined by normalization to the housekeeping gene Actin. ***, p< 0.001 and n.s.; not significant.
Tumor-conditioned medium does not suppress STAT1 and STAT2 expression but contains soluble factors that enhance STAT3 activation
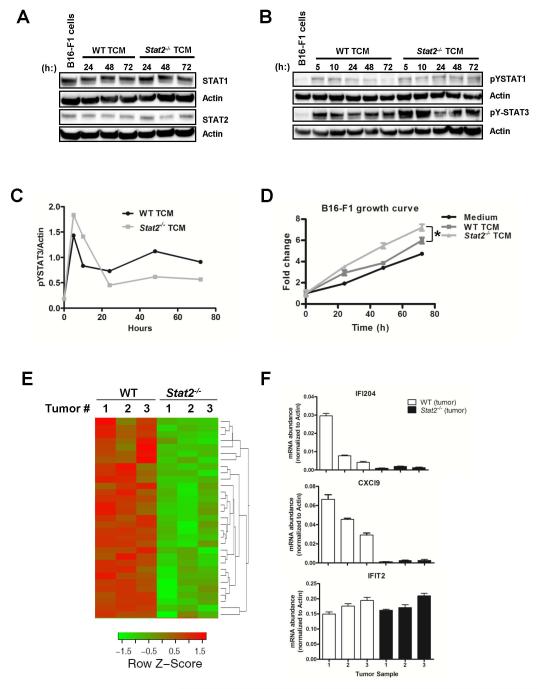
To determine whether soluble factors released by the tumor suppressed STAT1 and STAT2 protein expression within the tumors, we generated tumor-conditioned medium (TCM) from tumors established in WT and Stat2−/− mice. We cultured B16-F1 tumors cells expanded in vitro in medium alone or in 50% diluted TCM for different times. First, Western blot analysis revealed no reduction in the protein levels of STAT2 and STAT1 after incubation with either WT or Stat2−/− TCM (Fig. 4A). Next, we observed STAT3 activation with WT TCM, but the level of activation was not as pronounced as with Stat2−/− TCM in which enhanced STAT3 activation was detected between 5h and 10h (Fig. 4B and 4C). While the identity of the soluble factors present in Stat2−/− TCM that augmented STAT3 activation remains to be determined, no differences in the secretion of inflammatory mediators IL-6 and MCP-1 were observed in WT and Stat2−/− TCM (Fig. S3).
Figure 4. B16-F1 tumors established in Stat2−/− mice show reduced expression of immunomodulatory genes.
Tumor-conditioned medium (TCM) prepared from tumors harvested from WT and Stat2−/− mice was incubated with B16-F1 cells. Western blot analysis of (A) STAT1 and STAT2 protein levels and (B) Activated forms of STAT1 (pY701) and STAT3 (pY705) are shown. (C) The ratio of pY705-STAT3/Actin. (D) Cell proliferation measured by MTS assay every 24h; *, p< 0.05. (E) Heatmap of relative gene expression levels on a log2 scale comparing transcriptional signatures of tumors explants from WT and Stat2−/− mice. (F) Validation of select genes by qRT-PCR.
Tumors established in Stat2−/− mice show downregulation of immunomodulatory genes
We next compared the mitogenic activity of Stat2−/− TCM and WT TCM given that STAT3 activation was enhanced with Stat2−/− TCM. We observed that B16-F1 cells proliferated more rapidly with Stat2−/− TCM (Fig. 4D). In an effort to identify genes implicated in the enhanced tumor growth observed in Stat2−/− mice, we performed comparative microarray analysis on individual tumors established in WT and Stat2−/− mice. A small subset of genes was downregulated in tumors formed in Stat2−/− mice. Many of these genes are involved in immune cell differentiation and function (Fig. 4E and Table S1). The mRNA expression level of the top two candidate genes, Ifi204 and Cxcl927, 28 were validated by qRT-PCR. In agreement with our microarray data, we found the mRNA expression levels of IFIT2, an IFN-I target gene, to be the same in the two tumor groups. These findings indicate that tumors proliferate more efficiently in the Stat2−/− host due to impaired expression of key genes involved in the development of antitumor immunity.
STAT2/IFN-I signaling in DCs is essential in tumor antigen cross-presentation and restricting tumor growth
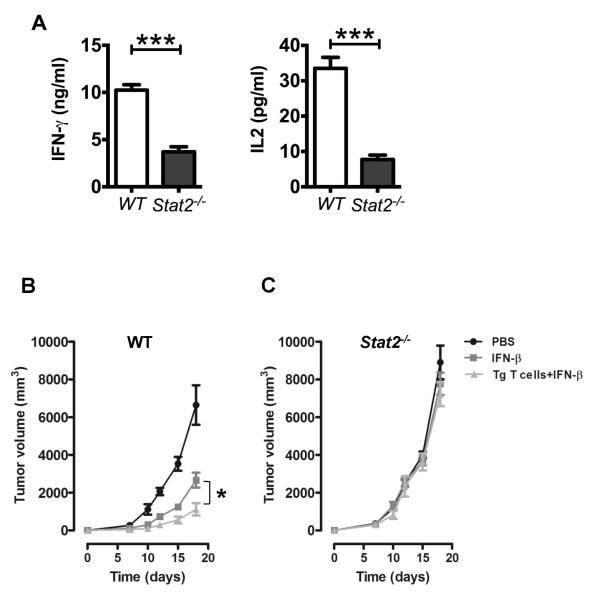
STAT2 was reported to participate in the development and differentiation of dendritic cells (DCs)22. Most recently, IFN-I signaling in DCs was identified as crucial for antigen cross-presentation.20, 21 Therefore, we evaluated potential defects in tumor antigen-cross presentation by Stat2−/− DCs. WT and Stat2−/− bone marrow-derived DCs were loaded with gp100 peptide and incubated with Pmel-1 TCR transgenic (Tg) T cells, which recognize the endogenous gp100 tumor antigen expressed by B16-F1 cells29. First, no differences in the percentages of CD8α+ Pmel-1 Tg T cells before or after priming by WT or Stat2−/− DCs were noted (Fig. S4A-B). Second, we found a significant reduction (~60-70%) in the levels of IFN-γ and IL-2 produced by Pmel-1 Tg T cells stimulated with Stat2−/− DCs compared to WT DCs (Fig. 5A).
Figure 5. Impaired tumor antigen cross-presentation by STAT2−/− DC.
LPS-stimulated WT or Stat2−/− BMDCs were pulsed with gp100 peptide and co-cultured with Pmel-1 T cells. (A) Supernatants were analyzed for IFN-γ and IL-2. (B-C) Antigen-primed Pmel-1 T cells were adoptively transferred into recipient mice injected three days before with B16-F1 tumor cells (T cells primed by WT-DCs → WT mice and T cells primed by Stat2−/−DCs → Stat2−/− mice). The next day mice began IFN-β treatment. Mice treated with vehicle or IFN-β alone served as controls; n=5/group. Values are shown as mean tumor volume ± SEM. ***, p< 0.001 and *, p>0.05. Data are representative of two independent experiments.
We next determined in vivo the cytotoxic activity of Pmel-1 CD8+ Tg T cells primed by WT or Stat2−/− DCs in vitro by measuring tumor regression with IFN-β administration. We must emphasize that for Pmel-1 Tg T cells to cause tumor rejection, accessory help is required in the form of immunomodulatory cytokines that include IL-2, IL-15, and IFN-I30-32. We adoptively transferred the corresponding primed Pmel-1 T cells into WT and Stat2−/− recipient mice that were injected with B16-F1 melanoma cells three days earlier. The next day, mice received IFN-β twice weekly (Fig. 5B and 5C). Adoptive transfer of Pmel-1 T cells primed by WT DCs with IFN-β treatment significantly reduced tumor growth in WT mice when compared to WT mice treated with IFN-β alone (Fig. 5B). In contrast, Stat2−/− mice with adoptive transfer of Pmel-1 T cells primed by Stat2−/− DCs did not cause tumor shrinkage (Fig. 5C). These results suggest that tumor antigen cross-presentation to CD8+ T cells by Stat2−/− DCs in vivo leads to impaired differentiation of CD8+ T cells.
IFN-I enhances the recruitment of T cells in a STAT2-dependent manner
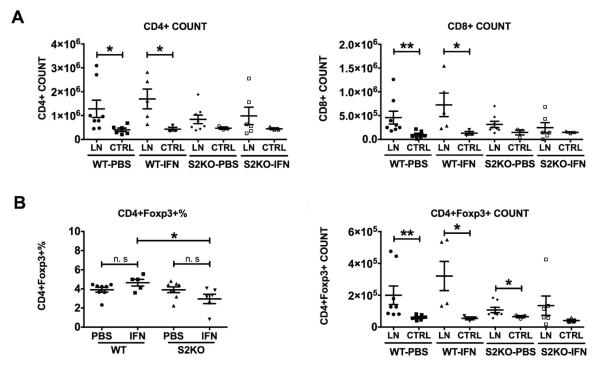
We next examined the composition of CD4+ and CD8+ T cells in WT and Stat2−/− tumor-bearing mice. The draining inguinal lymph nodes (LN, proximal to the tumor site) as opposed to the contralateral LN (CTRL, distal from the tumor site) in both strains showed an increase in the absolute number of both CD4+ and CD8+ T cell populations that was statistically significant in WT mice. A further increase in T cells was only observed in IFN-β treated WT mice (Fig. 6A). We also analyzed the same LNs for changes in the population of regulatory T cells (Tregs). We found that the percentage and absolute number of the classic regulatory CD4+Foxp3+ T cells were not significantly altered before or after IFN-β treatment in WT and Stat2−/− tumor bearing mice.
Figure 6. Increase in T cells in IFN-β treated tumor-bearing mice is STAT2 dependent.
Draining inguinal lymph nodes (LN) and contralateral LN (CTRL) from individual tumor-bearing WT or Stat2−/− (S2KO) mice injected with PBS or IFN-β were analyzed by flow cytometry. (A) Absolute numbers of CD4+ and CD8+ T cells. (B) Percentage and absolute numbers of CD4+Foxp3+ T cells. Data of one representative experiment of two are shown. Draining LNs; 5-8 mice/group. CTRL; 3-6 mice/group. *, p< 0.05; **, p< 0.01; and n.s., not significant.
Finally, we determined if the enhanced tumor growth observed in Stat2−/− mice could be achieved by depleting T cells in WT mice. T-cell depleted WT mice formed larger tumors when compared to control mice. These tumors, however, were smaller than those formed in Stat2−/− mice (Fig. S5A-B). Together, these data demonstrate that a constitutive and IFN-I-induced T cell- mediated antitumor response is impaired Stat2−/− mice. In addition, immune cells other than T cells participate in tumor rejection via STAT2.
Discussion
Based on a tumor transplantation model, we conclude that IFN-I-activated STAT2 is important in restricting tumor growth. Earlier studies showed a requirement of transcriptionally-active STAT2 to directly mediate the antigrowth effects of IFN-I.6, 9 During the course of our study, we found unexpectedly that the protein expression of both STAT2 and STAT1 within the tumors was markedly reduced. This is the first time, to our knowledge, that this effect has been reported in syngeneic tumor transplantation models. This surprising finding explains, in part, why in several spontaneous and tumor transplantation animal models, host IFN-I responsiveness and to a lesser degree a direct effect on the tumor, has been reported to be critical in tumor rejection.18, 26, 33 We speculate that in tumors established in Stat2−/− mice, microRNA-146a may be induced to suppress STAT1 expression.34 In contrast, suppression of STAT1 and STAT2 in tumors established in WT mice can be overcome with IFN-β treatment suggesting different mechanisms to reduce IFN-I signaling.
Our results also support an earlier study in which tumor biopsies of patients with squamous cell carcinoma showed a gradual reduction in the protein levels IFN-I signaling molecules STAT1 and STAT2 during tumor progression35. In a different study, however, melanoma biopsies taken from patients before and after IFN-α immunotherapy showed variable levels of STAT1 and STAT2 protein. This report concluded a lack of correlation between the levels of STAT proteins with outcome of IFN response26. Although STAT2 was found localized in the nucleus or in the cytoplasm or both compartments, it remains to be determined whether STAT2 was functional.
Another interesting observation we report is that Stat2−/− tumor-conditioned media enhanced tumor cell proliferation and activation of STAT3. While the composition of factors secreted by B16-F1 tumor cells is quite extensive36, we initially postulated that loss of host STAT2 could lead to an increase in the secretion of mitogenic factors by the tumors. However, whole genome microarray analysis indicated quite the opposite outcome. We identified a small subset of genes with immunomodulatory function to be downregulated in tumors established in Stat2−/− mice. The two top candidate genes were Ifi204 and Cxcl9. Ifi204 is an IFN-I target gene important in cell growth inhibition and macrophage cell differentiation. 27, 37 Cxcl9 is another IFN target gene induced by IFN-α/β and IFN-γ and functions as a T-cell chemoattractant.38, 39 Recently, Cxcl9−/− mice were reported to form large tumors due to immunosurveillance escape.28 Thus our findings indicate that STAT2 orchestrates the expression of genes that are important in the differentiation and recruitment of immune effector cells to the tumor site.
Most tumors carry activated STAT3 including B16-F1 and MC38 tumor cell lines. Consequently, one can speculate that loss of STAT2 and STAT1 within tumors may confer tumor cells a survival advantage by relying on STAT3 to activate oncogenic survival pathways in the presence of IFN-I. In fact, IFN-I has been shown to be mitogenic and promote cell proliferation of Stat1−/− or Stat2−/− T cells11. Furthermore, transgenic expression of IFN-α in the brain of Stat2−/− mice has been reported to be tumorigenic. Our studies, however, did not show enhanced tumor growth in Stat2−/−mice treated with IFN-β compared with untreated mice.
Our study also revealed that Stat2−/− DCs were defective in tumor antigen cross-presentation in vivo, a feature for which Stat2−/− mice may have been more permissive to form tumors due to its inherent IFN signaling defect10. The role of IFN signaling in maturation and differentiation of DCs has been intensively studied.20, 21, 40-42 IFN-I signaling in DCs is important for cross-presenting antigen to CD8+ T cells; the driving force behind tumor elimination. Similarly, STAT2 mutant mice with close functional likeness to Stat2−/− mice were also shown to exhibit a deficiency in maturation and differentiation of DCs22. These observations agree with the transcriptional signature of tumors established in Stat2−/− mice.
IFN-Is are known to enhance the proliferation of Pmel-1 Tg CD8+ T cells in a tumor vaccination model and suppress B16 melanoma growth.32 In our study, adoptive transfer of Pmel-1 Tg T cells primed by Stat2−/− DCs into Stat2−/− mice treated with IFN-β did not cause tumor regression. This confirms that T cell-mediated antitumor immunity is important for host IFN-I responsiveness33, and that IFN-I adjuvant immunotherapy of melanoma may prove to be ineffective if DCs have reduced expression of STAT2 or impaired STAT2 signaling.
In summary, our study highlights a critical interplay between host STAT2 and the tumor. A deficiency in STAT2 disables IFN-I-mediated antitumor immune response and reduces the expression of key immunomodulatory factors thus allowing tumors to thrive. Consequently, the efficacy of cytokine-based cancer immunotherapy may be compromised if STAT2 function is impaired.
Supplementary Material
Novelty and Impact.
We provide evidence of the in vivo role of STAT2 in the antitumor activity of type I interferons (IFN-I). We found STAT2 to be critical in controlling tumor growth and in tumor antigen cross-presentation. Microarray analysis identified a small subset of STAT2-dependent genes that actively participates in antitumor immunity. T cell recruitment is also enhanced by IFN-I via STAT2. Our findings indicate that impairment in host STAT2 function can diminish the efficacy of IFN-I immunotherapy.
Acknowledgements
We thank Dr. Håkan Steen and Dr. Scott Durum for their critical review of the manuscript and Mr. Michael Slifker for conducting the microarray data analysis. This work was supported by NIH grant RO1 CA140499 to AMG.
Footnotes
The authors have no conflicting financial interests, with the exception of Darren P. Baker who is an employee of BiogenIdec and who owns BiogenIdec stock.
References
- 1.Fu XY, Schindler C, Improta T, Aebersold R, Darnell JE., Jr. The proteins of ISGF-3, the interferon alpha-induced transcriptional activator, define a gene family involved in signal transduction. Proc Natl Acad Sci U S A. 1992;89:7840–3. doi: 10.1073/pnas.89.16.7840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schindler C, Shuai K, Prezioso VR, Darnell JE., Jr. Interferon-dependent tyrosine phosphorylation of a latent cytoplasmic transcription factor. Science. 1992;257:809–13. doi: 10.1126/science.1496401. [DOI] [PubMed] [Google Scholar]
- 3.Darnell JE, Jr., Kerr IM, Stark GR. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science. 1994;264:1415–21. doi: 10.1126/science.8197455. [DOI] [PubMed] [Google Scholar]
- 4.Stark GR, Kerr IM, Williams BR, Silverman RH, Schreiber RD. How cells respond to interferons. Annu Rev Biochem. 1998;67:227–64. doi: 10.1146/annurev.biochem.67.1.227. [DOI] [PubMed] [Google Scholar]
- 5.Levy DE, Darnell JE., Jr. Stats: transcriptional control and biological impact. Nat Rev Mol Cell Biol. 2002;3:651–62. doi: 10.1038/nrm909. [DOI] [PubMed] [Google Scholar]
- 6.Clifford JL, Yang X, Walch E, Wang M, Lippman SM. Dominant negative signal transducer and activator of transcription 2 (STAT2) protein: stable expression blocks interferon alpha action in skin squamous cell carcinoma cells. Mol Cancer Ther. 2003;2:453–9. [PubMed] [Google Scholar]
- 7.Du Z, Fan M, Kim JG, Eckerle D, Lothstein L, Wei L, Pfeffer LM. Interferon-resistant Daudi cell line with a Stat2 defect is resistant to apoptosis induced by chemotherapeutic agents. J Biol Chem. 2009;284:27808–15. doi: 10.1074/jbc.M109.028324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gamero AM, Larner AC. Vanadate facilitates interferon alpha-mediated apoptosis that is dependent on the Jak/Stat pathway. J Biol Chem. 2001;276:13547–53. doi: 10.1074/jbc.M007948200. [DOI] [PubMed] [Google Scholar]
- 9.Romero-Weaver AL, Wang HW, Steen HC, Scarzello AJ, Hall VL, Sheikh F, Donnelly RP, Gamero AM. Resistance to IFN-alpha-induced apoptosis is linked to a loss of STAT2. Mol Cancer Res. 2010;8:80–92. doi: 10.1158/1541-7786.MCR-08-0344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Park C, Li S, Cha E, Schindler C. Immune response in Stat2 knockout mice. Immunity. 2000;13:795–804. doi: 10.1016/s1074-7613(00)00077-7. [DOI] [PubMed] [Google Scholar]
- 11.Gimeno R, Lee CK, Schindler C, Levy DE. Stat1 and Stat2 but not Stat3 arbitrate contradictory growth signals elicited by alpha/beta interferon in T lymphocytes. Mol Cell Biol. 2005;25:5456–65. doi: 10.1128/MCB.25.13.5456-5465.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Plumlee CR, Lee C, Beg AA, Decker T, Shuman HA, Schindler C. Interferons direct an effective innate response to Legionella pneumophila infection. J Biol Chem. 2009;284:30058–66. doi: 10.1074/jbc.M109.018283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gamero AM, Young MR, Mentor-Marcel R, Bobe G, Scarzello AJ, Wise J, Colburn NH. STAT2 contributes to promotion of colorectal and skin carcinogenesis. Cancer Prev Res (Phila) 2010;3:495–504. doi: 10.1158/1940-6207.CAPR-09-0105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wang J, Pham-Mitchell N, Schindler C, Campbell IL. Dysregulated Sonic hedgehog signaling and medulloblastoma consequent to IFN-alpha-stimulated STAT2-independent production of IFN-gamma in the brain. J Clin Invest. 2003;112:535–43. doi: 10.1172/JCI18637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Picaud S, Bardot B, De Maeyer E, Seif I. Enhanced tumor development in mice lacking a functional type I interferon receptor. J Interferon Cytokine Res. 2002;22:457–62. doi: 10.1089/10799900252952244. [DOI] [PubMed] [Google Scholar]
- 16.Takemoto Y, Yano H, Momosaki S, Ogasawara S, Nishida N, Kojiro S, Kamura T, Kojiro M. Antiproliferative effects of interferon-alphaCon1 on ovarian clear cell adenocarcinoma in vitro and in vivo. Clin Cancer Res. 2004;10:7418–26. doi: 10.1158/1078-0432.CCR-04-0279. [DOI] [PubMed] [Google Scholar]
- 17.Hisaka T, Yano H, Ogasawara S, Momosaki S, Nishida N, Takemoto Y, Kojiro S, Katafuchi Y, Kojiro M. Interferon-alphaCon1 suppresses proliferation of liver cancer cell lines in vitro and in vivo. J Hepatol. 2004;41:782–9. doi: 10.1016/j.jhep.2004.07.012. [DOI] [PubMed] [Google Scholar]
- 18.Dunn GP, Bruce AT, Sheehan KC, Shankaran V, Uppaluri R, Bui JD, Diamond MS, Koebel CM, Arthur C, White JM, Schreiber RD. A critical function for type I interferons in cancer immunoediting. Nat Immunol. 2005;6:722–9. doi: 10.1038/ni1213. [DOI] [PubMed] [Google Scholar]
- 19.Swann JB, Hayakawa Y, Zerafa N, Sheehan KC, Scott B, Schreiber RD, Hertzog P, Smyth MJ. Type I IFN contributes to NK cell homeostasis, activation, and antitumor function. J Immunol. 2007;178:7540–9. doi: 10.4049/jimmunol.178.12.7540. [DOI] [PubMed] [Google Scholar]
- 20.Diamond MS, Kinder M, Matsushita H, Mashayekhi M, Dunn GP, Archambault JM, Lee H, Arthur CD, White JM, Kalinke U, Murphy KM, Schreiber RD. Type I interferon is selectively required by dendritic cells for immune rejection of tumors. J Exp Med. 2011;208:1989–2003. doi: 10.1084/jem.20101158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fuertes MB, Kacha AK, Kline J, Woo SR, Kranz DM, Murphy KM, Gajewski TF. Host type I IFN signals are required for antitumor CD8+ T cell responses through CD8{alpha}+ dendritic cells. J Exp Med. 2011;208:2005–16. doi: 10.1084/jem.20101159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen LS, Wei PC, Liu T, Kao CH, Pai LM, Lee CK. STAT2 hypomorphic mutant mice display impaired dendritic cell development and antiviral response. J Biomed Sci. 2009;16:22. doi: 10.1186/1423-0127-16-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Sriram U, Biswas C, Behrens EM, Dinnall JA, Shivers DK, Monestier M, Argon Y, Gallucci S. IL-4 suppresses dendritic cell response to type I interferons. J Immunol. 2007;179:6446–55. doi: 10.4049/jimmunol.179.10.6446. [DOI] [PubMed] [Google Scholar]
- 24.Yue C, Soboloff J, Gamero AM. Control of type I interferon-induced cell death by Orai1-mediated calcium entry in T cells. J Biol Chem. 2012;287:3207–16. doi: 10.1074/jbc.M111.269068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Simon R, Lam A, Li MC, Ngan M, Menenzes S, Zhao Y. Analysis of gene expression data using BRB-ArrayTools. Cancer Inform. 2007;3:11–7. [PMC free article] [PubMed] [Google Scholar]
- 26.Lesinski GB, Anghelina M, Zimmerer J, Bakalakos T, Badgwell B, Parihar R, Hu Y, Becknell B, Abood G, Chaudhury AR, Magro C, Durbin J, et al. The antitumor effects of IFN-alpha are abrogated in a STAT1-deficient mouse. J Clin Invest. 2003;112:170–80. doi: 10.1172/JCI16603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Dauffy J, Mouchiroud G, Bourette RP. The interferon-inducible gene, Ifi204, is transcriptionally activated in response to M-CSF, and its expression favors macrophage differentiation in myeloid progenitor cells. J Leukoc Biol. 2006;79:173–83. doi: 10.1189/jlb.0205083. [DOI] [PubMed] [Google Scholar]
- 28.Petro M, Kish D, Guryanova OA, Ilyinskaya G, Kondratova A, Fairchild RL, Gorbachev AV. Cutaneous tumors cease CXCL9/Mig production as a result of IFN-gamma-mediated immunoediting. J Immunol. 2013;190:832–41. doi: 10.4049/jimmunol.1201906. [DOI] [PubMed] [Google Scholar]
- 29.Overwijk WW, Theoret MR, Finkelstein SE, Surman DR, de Jong LA, Vyth-Dreese FA, Dellemijn TA, Antony PA, Spiess PJ, Palmer DC, Heimann DM, Klebanoff CA, et al. Tumor regression and autoimmunity after reversal of a functionally tolerant state of self-reactive CD8+ T cells. J Exp Med. 2003;198:569–80. doi: 10.1084/jem.20030590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Klebanoff CA, Gattinoni L, Palmer DC, Muranski P, Ji Y, Hinrichs CS, Borman ZA, Kerkar SP, Scott CD, Finkelstein SE, Rosenberg SA, Restifo NP. Determinants of successful CD8+ T-cell adoptive immunotherapy for large established tumors in mice. Clin Cancer Res. 2011;17:5343–52. doi: 10.1158/1078-0432.CCR-11-0503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Roychowdhury S, May KF, Jr., Tzou KS, Lin T, Bhatt D, Freud AG, Guimond M, Ferketich AK, Liu Y, Caligiuri MA. Failed adoptive immunotherapy with tumor-specific T cells: reversal with low-dose interleukin 15 but not low-dose interleukin 2. Cancer Res. 2004;64:8062–7. doi: 10.1158/0008-5472.CAN-04-1860. [DOI] [PubMed] [Google Scholar]
- 32.Sikora AG, Jaffarzad N, Hailemichael Y, Gelbard A, Stonier SW, Schluns KS, Frasca L, Lou Y, Liu C, Andersson HA, Hwu P, Overwijk WW. IFN-alpha enhances peptide vaccine-induced CD8+ T cell numbers, effector function, and antitumor activity. J Immunol. 2009;182:7398–407. doi: 10.4049/jimmunol.0802982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Burnette BC, Liang H, Lee Y, Chlewicki L, Khodarev NN, Weichselbaum RR, Fu YX, Auh SL. The efficacy of radiotherapy relies upon induction of type i interferon-dependent innate and adaptive immunity. Cancer Res. 2011;71:2488–96. doi: 10.1158/0008-5472.CAN-10-2820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wang S, Zhang X, Ju Y, Zhao B, Yan X, Hu J, Shi L, Yang L, Ma Z, Chen L, Liu Y, Duan Z, et al. MicroRNA-146a feedback suppresses T cell immune function by targeting Stat1 in patients with chronic hepatitis B. J Immunol. 2013;191:293–301. doi: 10.4049/jimmunol.1202100. [DOI] [PubMed] [Google Scholar]
- 35.Clifford JL, Walch E, Yang X, Xu X, Alberts DS, Clayman GL, El-Naggar AK, Lotan R, Lippman SM. Suppression of type I interferon signaling proteins is an early event in squamous skin carcinogenesis. Clin Cancer Res. 2002;8:2067–72. [PubMed] [Google Scholar]
- 36.Obrador E, Benlloch M, Pellicer JA, Asensi M, Estrela JM. Intertissue flow of glutathione (GSH) as a tumor growth-promoting mechanism: interleukin 6 induces GSH release from hepatocytes in metastatic B16 melanoma-bearing mice. J Biol Chem. 2011;286:15716–27. doi: 10.1074/jbc.M110.196261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lembo M, Sacchi C, Zappador C, Bellomo G, Gaboli M, Pandolfi PP, Gariglio M, Landolfo S. Inhibition of cell proliferation by the interferon-inducible 204 gene, a member of the Ifi 200 cluster. Oncogene. 1998;16:1543–51. doi: 10.1038/sj.onc.1201677. [DOI] [PubMed] [Google Scholar]
- 38.Guirnalda P, Wood L, Goenka R, Crespo J, Paterson Y. Interferon gamma-induced intratumoral expression of CXCL9 alters the local distribution of T cells following immunotherapy with. Oncoimmunology. 2013;2:e25752. doi: 10.4161/onci.25752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Padovan E, Spagnoli GC, Ferrantini M, Heberer M. IFN-alpha2a induces IP-10/CXCL10 and MIG/CXCL9 production in monocyte-derived dendritic cells and enhances their capacity to attract and stimulate CD8+ effector T cells. J Leukoc Biol. 2002;71:669–76. [PubMed] [Google Scholar]
- 40.Montoya M, Schiavoni G, Mattei F, Gresser I, Belardelli F, Borrow P, Tough DF. Type I interferons produced by dendritic cells promote their phenotypic and functional activation. Blood. 2002;99:3263–71. doi: 10.1182/blood.v99.9.3263. [DOI] [PubMed] [Google Scholar]
- 41.Asselin-Paturel C, Brizard G, Chemin K, Boonstra A, O’Garra A, Vicari A, Trinchieri G. Type I interferon dependence of plasmacytoid dendritic cell activation and migration. J Exp Med. 2005;201:1157–67. doi: 10.1084/jem.20041930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Swiecki M, Wang Y, Vermi W, Gilfillan S, Schreiber RD, Colonna M. Type I interferon negatively controls plasmacytoid dendritic cell numbers in vivo. J Exp Med. 2011;208:2367–74. doi: 10.1084/jem.20110654. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.