Abstract
Tourette syndrome is a neurodevelopmental disorder characterized by multiple motor and vocal tics, and the disorder is often accompanied by comorbidities such as attention-deficit hyperactivity-disorder and obsessive compulsive disorder. Tourette syndrome has a complex etiology, but the underlying environmental and genetic factors are largely unknown. IMMP2L (inner mitochondrial membrane peptidase, subunit 2) located on chromosome 7q31 is one of the genes suggested as a susceptibility factor in disease pathogenesis. Through screening of a Danish cohort comprising 188 unrelated Tourette syndrome patients for copy number variations, we identified seven patients with intragenic IMMP2L deletions (3.7%), and this frequency was significantly higher (P=0.0447) compared with a Danish control cohort (0.9%). Four of the seven deletions identified did not include any known exons of IMMP2L, but were within intron 3. These deletions were found to affect a shorter IMMP2L mRNA species with two alternative 5′-exons (one including the ATG start codon). We showed that both transcripts (long and short) were expressed in several brain regions, with a particularly high expression in cerebellum and hippocampus. The current findings give further evidence for the role of IMMP2L as a susceptibility factor in Tourette syndrome and suggest that intronic changes in disease susceptibility genes should be investigated further for presence of alternatively spliced exons.
Keywords: ADHD, IMMP2L, intronic deletions, OCD, susceptibility gene, Tourette syndrome
Introduction
Tourette syndrome (TS; MIM 137580) is a childhood onset chronic neurodevelopmental disorder characterized by multiple motor tics and at least one vocal/phonic tic.1 TS is a complex disorder where several environmental and genetic components are likely to be involved in the disease etiology. About 88% of TS patients are diagnosed with comorbidities such as attention-deficit hyperactivity-disorder (ADHD), obsessive compulsive disorder/behavior (OCD/OCB), rage attacks, sleep difficulties, autism spectrum disorders (ASD) and affective disorders2,3 suggesting a shared background between these disorders.4 This notion is strengthened by finding of genes, such as CHRNA7, CNTNAP2 and NLGN4, which are implicated in a wide variety of neurodevelopmental disorders, including TS and ASD.5
Although it is widely accepted that genetic factors have a significant role in TS etiology, only a few genes have been suggested to render susceptibility to TS and IMMP2L (inner mitochondrial membrane peptidase, subunit 2) located on chromosome 7q31 is one of these genes.5 The chromosome region 7q31 was first implicated as a candidate locus for TS through identification of a constitutional chromosome translocation t(7;18)(q22;q22.3) in a family with OCD, motor tics and/or vocal tics; where the 7q22 breakpoint was localized between markers D7S515 and D7S522.6 In 2001, a boy with a de novo inverted duplication, dup(7)(pter-q31.1::q31.1-q22.1::q31.1-qter), presenting with vocal and motor tics was reported.7, 8 The 7q31 breakpoint was within the region described by Boghosian-Sell and colleagues6 and disrupted a novel gene, IMMP2L.7 These two studies implicated 7q31 as a candidate TS region. Subsequently, a family-based association study was performed in a cohort of French–Canadian patients from Quebec, and a biased transmission of alleles from heterozygote parents to their TS offspring was found for markers D7S522, D7S523 and D7S1516 (located within IMMP2L).9 In a subsequent study, 39 TS patients were screened for intragenic sequence variations and 22 patients were further investigated for structural copy number variations (CNVs) involving IMMP2L, without finding any changes.10 Recent description of another patient with TS-like tics and a translocation, t(2;7)(p24.2;q31), where a 7.2-Mb microdeletion at the chromosome 7q31 breakpoint partly deleted IMMP2L, has again awakened the discussion about involvement of this gene in TS pathogenesis.11
The IMMP2L gene is the human homolog of Imp2 of Saccharomyces cerevisiae and the transcript identified and published by Petek and colleagues7 has 6 exons, where the ATG start codon is within the second exon (NM_032549.3). IMMP2L is expressed in a wide range of tissues including the fetal and adult brain, but not in the adult liver and lung.7 Mammalian IMMP1L and IMMP2L constitute, like the yeast Imp1 and Imp2, the subunits in the inner membrane peptidase complex, which catalyze the removal of the signal peptides required for the targeting of proteins from the mitochondrial matrix into the intermembrane space across the inner membrane.12 The two subunits share a conserved core and they are both catalytically active, but process different substrates.13, 14 Cytochrome c1 and mitochondrial glycerol-3-phosphate dehydrogenase 2 are substrates for IMMP2L, and IMMP2L mutations have been shown to impair signal peptide processing of both proteins.15, 16 Both the substrates have important roles in energy metabolism and oxidation in the mitochondria. In recent studies, mitochondria isolated from the brain tissue of mice homozygous for an Immp2l mutation manifested hyperpolarization of mitochondrial inner membrane potential, increased superoxide ion generation and higher levels of ATP.16 Incomplete processing of IMMP2L substrates seems to result in a hyperactive mitochondrion with an increased production of superoxide16 and to promote the formation of a mitochondrial permeability transition pore which leads to release of pro-apoptotic proteins, and thereby activating cell death pathways.17, 18
In this study, we describe seven Danish TS patients with intragenic IMMP2L deletions, affecting one of two alternative IMMP2L transcripts (the previously described long transcript and a short alternative transcript with the two first exons residing within intron 3 of the long transcript). The deletions are significantly more frequent in the patients compared with the controls giving further evidence for IMMP2L as a TS susceptibility gene.
Materials and methods
Study subjects
The present cohort includes 188 unrelated TS patients (154 males and 34 females), who were recruited through the Herlev Tourette Clinic (Denmark) as part of a clinical study.3 Patients were examined using validated diagnostic instruments19 and clinical information about the parents of the patients with IMMP2L deletions was obtained through interviews. The study was approved by the Danish Institutional Review Board (H-KA-05118) and informed consent was obtained from each participant.
The control population includes material from 316 Danish individuals (203 males and 113 females) retrieved from the biobanks of the Kennedy Center and the Center for Neuropsychiatric Schizophrenia Research. Overt neurological disorders were ruled out for this cohort. Additional CNV calls from 1038 controls from the reference database genotyped on the Affymetrix CytoScan HD array platform were also included as control population (Affymetrix, Santa Clara, CA, USA).
Chromosome microarray
Chromosome microarray was initially carried out using Affymetrix Cytogenetics Whole-Genome 2.7 M array (82 patients), but the manufacturer (Affymetrix) replaced it by a similar and more robust platform (CytoScan HD array), which was then used for the remaining patients (106 patients and 1038 Affymetrix controls). Both array platforms have comparable probe localization and density (∼1 probe/kb) within the IMMP2L gene, and they are therefore not expected to introduce any bias in the pick-up rate of IMMP2L CNVs in the investigated individuals. Data were analyzed with ChAS software (Affymetrix) using the following filtering criteria: deletions >40 kb (a minimum of 30 markers) and duplications >40 kb (a minimum of 30 markers).
Quantitative PCR (qPCR)
Genomic DNA was extracted from peripheral blood using standard protocols and qPCR was carried out with Power SYBR Green PCR technology (Applied Biosystems, Foster City, CA, USA) using a 7500 Fast Real-time PCR system machine (Applied Biosystems). Each amplicon was quantified in triplicates and copy number estimates were derived by ΔΔCT calculations using a copy number neutral GAPDH amplicon and control individuals.
Verification of the IMMPL2 deletions was performed using qPCR amplicons targeting exon 3 and intron 3 of the long transcript (Figure 1). The same amplicons were used to investigate the parental origin of the deletions and to screen the control cohort of 316 Danish individuals. Validation and investigation of parental origin of all the other CNVs were also carried out using qPCR.
Figure 1.
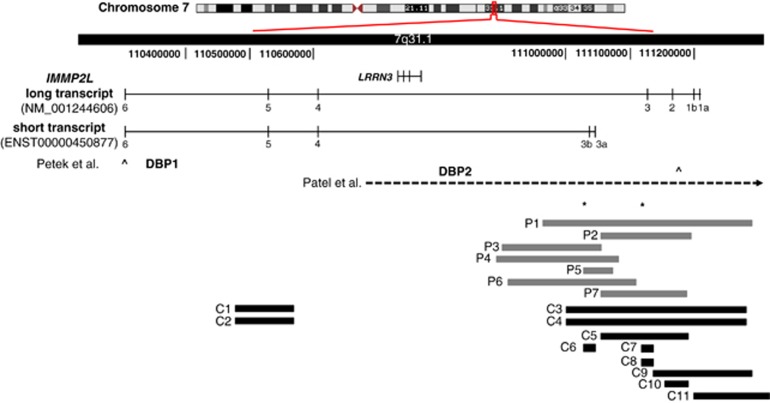
Structural variations within the IMMP2L transcripts. (Genomic region showing the long (NM_001244606.1) and the short (ENST00000450877) transcripts of IMMP2L, and the structural variations identified in this study or by others. DBP1 and DBP2 indicate the duplication breakpoints and the open arrows indicate the amplicons (corresponding to exon 6 and exon 1b/IVS1a) used to screen for IMMP2L deletions (Petek et al7 and Petek et al,10 respectively). Horizontal dashed-arrow shows the 7.2-Mb deletion described by Patel et al.11 Horizontal gray blocks indicate the seven intragenic deletions (P1–P7) identified in Tourette patients in this study. The three deletions identified in the Danish background population (C6–C8) and the eight deletions identified in the Affymetrix CytoScan HD reference database (C1–C5 and C9–C11) are shown as horizontal black blocks. Asterisks represent localization of the qPCR amplicons.
IMMP2L-deletion breakpoints were narrowed down with qPCR. The deletion junction fragments were amplified with long range PCR and sequenced in both directions.
Sequence analysis of IMMP2L
Exons and flanking intron sequences were investigated by Sanger sequencing using genomic DNA.
Transcription studies
Reverse transcription PCR was carried out using peripheral blood leukocytes from TS patients, total RNA from various brain regions of a normal human adult (Biochain, Newark, CA, USA), human universal reference total RNA (Clontech, Palo Alto, CA, USA) and human whole-brain total RNA (Stratagene, La Jolla, CA, USA). cDNA was synthesized from 0.5 μg total RNA using SuperScript III reverse transcriptase (Invitrogen, San Diego, CA, USA).
In situ hybridization
In situ hybridization was performed on eight-micron sections of snap-frozen or formalin-fixed, paraffin-embedded (FFPE) brains from adult, 7 days old (D7) and embryonic (E16.5 or E15.5) BALB/c mice as previously published20 with some modifications: Hybridizations were carried out overnight at 47 °C and the tyramide signal amplification was detected using tyramide signal amplification Plus Fluorescein system for frozen sections or tyramide signal amplification DNT kits for FFPE sections (Perkin Elmer, Skovlunde, Denmark). The oligonucleotide probes were 5′- and 3′-FITC labeled and targeted the long (5′-ACAATGTCACCACGGTGTACTTCAAAATTCCTCACTT-3′) (exon 3) or the short (5′-ATGCACTTCTTCAATCTTAGGCTAACTCTTCTCGGTTGGC-3′) (exon 3b) transcripts of human IMMP2L (TAG Copenhagen, Copenhagen, Denmark). The same probes were used against mouse and human sections. Animals were handled in accordance with the guidelines of the European Communities directive of 86/609/EEC. Paraffin-embedded human corpus callosum sections were purchased from Capital Biosciences (Rockville, MD, USA). Images were obtained using an Olympus Macroview Microscope (MVX10) equipped with an Olympus FIII View and ColorView II camera and Cell P AnalySIS software (Olympus Denmark, Ballerup, Denmark).
Results
Identification and characterization of IMMP2L deletions
Screening of 188 unrelated TS patients for CNVs resulted in identification of intragenic IMMP2L deletions in seven patients (P1–P7) (Figure 1, Table 1). In three patients, we found an additional CNV (duplication), which may contribute to the phenotype (Table 1). Other likely benign CNVs (>40 kb) are listed in Supplementary Table S1.
Table 1. Overview of the deletions encompassing IMMP2L, identified in 7 out of 188 unrelated TS patients.
| Patient | Sex | Symptom | Deletion | Deletion size (kb) | Exons deleted | Parental origin | Parental phenotype | Additional deletions/duplications (CNVs) |
|---|---|---|---|---|---|---|---|---|
| P1 TS101 6972466 | M | TS ADHD | chr7.hg19:g.110.959.798 _111.290.945delinsCATACTACAGGACTTACAAACTTACAA | 331 | 1a, 1b, 2, 3, 3a, 3b | Pat | Pat: dyslexia, temper. Mat: unaffected | |
| P2 TS71 6962812 | M | TS ADHD | chr7.hg19:g.111.051.907 _111.199.868del | 148 | 2, 3 | Mat | Pat: ADHD symptoms Mat: tics, OCB | |
| P3 TS131 6982155 | M | TS ADHD Asperger | chr7.hg19:g.(?_110.892.600) _(111.055.300_?)del | ∼163 | 3a, 3b | Mat | Pat: unaffected Mat: unaffected | chr1.hg19:g.(?_231711404)_(231814354_?) dup 102 kb paternal duplication which includes DISC1 partially |
| P4 TS133 6982192 | M | TS OCD | chr7.hg19:g.110.889.476 _111.080.538delinsACACCTACTGCTCA | 191 | 3a, 3b | Pat | Pat: OCB, stuttering Mat: unaffected | chr4.hg19:g.(?_188104036)_(189795272_?) dup 1.7 Mb paternal duplication which includes ZFP42, TRIML1, TRIML2, LOC339975, LOC401164 |
| P5 TS32 6952838 | M | TS | chr7.hg19:g.111.029.575 _111.079.376del | 49 | 3a, 3b | NA | Pat: unaffected Mat: unaffected | chr15.hg19:g.(?_32019187)_(32439298_?) dup 420 kb duplication which includes OTUD7A and CHRNA7 partially |
| P6 TS36 8942118 | M | TS ADHD OCD | chr7.hg19:g.110.904.173 _111.113.977delinsCA | 210 | 3a, 3b | Pat | Pat: stubbornness Mat: unaffected | |
| P7 TS126 8932009 | M | TS ADHD | chr7.hg19:g.(?_111.054.100) _(111.197.400_?)del | ∼143 | 2, 3 | Pat | Pat: unaffected Mat: unaffected |
Abbreviations: del, deletion; dup, duplication; M, male; mat, mater/mother; NA, not analyzed as parental DNA was not available; pat, pater/father.
Patient identities (P1–P7) are followed by family numbers and encrypted patient identification codes.
All the IMMP2L deletions were at the 5′-end of the gene and ranged in size from 49 to 331 kb. In four patients (P3–P6), the deletions did not include any coding exons of the long IMMP2L transcript (NM_001244606.1), but were within intron 3. We speculated whether this intron included alternatively spliced exons; and using Ensemble database, we identified a shorter IMMP2L transcript (ENST00000450877) with two unique exons (designated 3a and 3b). The translation start site was within exon 3a, and the short transcript included exons 3a, 3b and exons 4–6 (common with the long transcript).
In five patients (P1, P2 and P4–P6), we characterized the deletions at base-pair level. In three families (P1/TS101, P2/TS71 and P4/TS133), the deletion was inherited from a parent with a neuropsychiatric feature, whereas in three cases it was inherited from an unaffected parent (P3/TS131, P6/TS36 and P7/TS126) (Table 1). The genomic location of all IMMP2L deletions has been submitted to the Leiden Open Variation Database.
Frequency of IMMP2L deletions
Using two qPCR amplicons (in exon 3 and intron 3), we identified three deletions in a cohort of 316 individuals representing the Danish background population (0.9%). One deletion involved only the short transcript (C6) and two deletions involved only the long transcript (C7 and C8) (Figure 1). The frequency of IMMP2L deletions in our TS cohort is 3.7% (7/188) and this is significantly higher than that of the background population (P=0.0447, two-tailed Fisher's exact test).
To avoid a potential bias, we investigated the presence of CNVs along the entire gene among 1038 controls from the Affymetrix CytoScan HD reference database. IMMP2L deletions affecting the long transcript were present in eight controls (C1–C5 and C9–11) (0.8%) and two of these (C3 and C4) also included the short IMMP2L transcript (Figure 1). The frequency of IMMP2L deletions in the TS cohort is thus significantly higher than that observed in 1038 controls (P=0.0039; Fisher's exact test).
DNA sequence analyses
None of the seven patients had small base-pair changes of the non-deleted IMMP2L allele. Furthermore, we sequenced the remaining 174 patients for mutations within exons 3a and 3b without detecting any pathogenic variations. We did not include the other exons as these were previously screened for mutations in 39 TS patients by Petek and colleagues,10 without detecting any mutations.
Other alternative transcripts of IMMP2L
In this study, we could not amplify the entire transcript (NM_032549.3), which was predicted to have a 440-bp long exon 1.7 On the other hand, we could amplify the entire NM_001244606.1 transcript and we showed that this transcript had two 5′ non-coding exons of 54 and 74 bp (designated exon 1a and 1b, respectively). Throughout the text we use NM_001244606.1 as the long IMMP2L transcript.
Further studies using different combinations of primers showed that both the long (NM_001244606.1) and the short (ENST00000450877) transcripts had two isoforms, with and without exon 5, respectively. All these transcripts had open-reading frames. Several other smaller transcripts were also identified, but none of these had a reasonable open-reading frame that would be translated to a functional protein.
Predicted protein encoded by the short IMMP2L transcript
The 157 amino-acid protein product of the short transcript does not contain an N-terminal transmembrane domain (TMpred) and is likely to be a soluble protein with an average hydropathicity of 0.71 (SOSUI). The structurally important substrate-binding cleft NX5S-motif of the long IMMP2L transcript and the highly conserved C-terminal12 are present in the short isoform.
Tissue-expression profiles of the long and the short IMMP2L transcripts
We investigated tissue-expression profiles of the long (primers against exons 2 and 6) and the short transcript (primers against exons 3b and 6) to gather further evidence of their involvment in the central nervous system pathology implicated in TS. Both transcripts were present in human total RNA and total brain RNA, whereas the short transcript was not expressed in blood leukocytes. Studies with 19 different human brain regions showed that the long transcript was expressed in all the regions whereas the short transcript was expressed in the cerebellum, meninges, corpus callosum, hippocampus, medulla oblongata, occipital lobe, pituitary, pons, precentral gyrus, temporal lobe and thalamus (Figure 2a).
Figure 2.
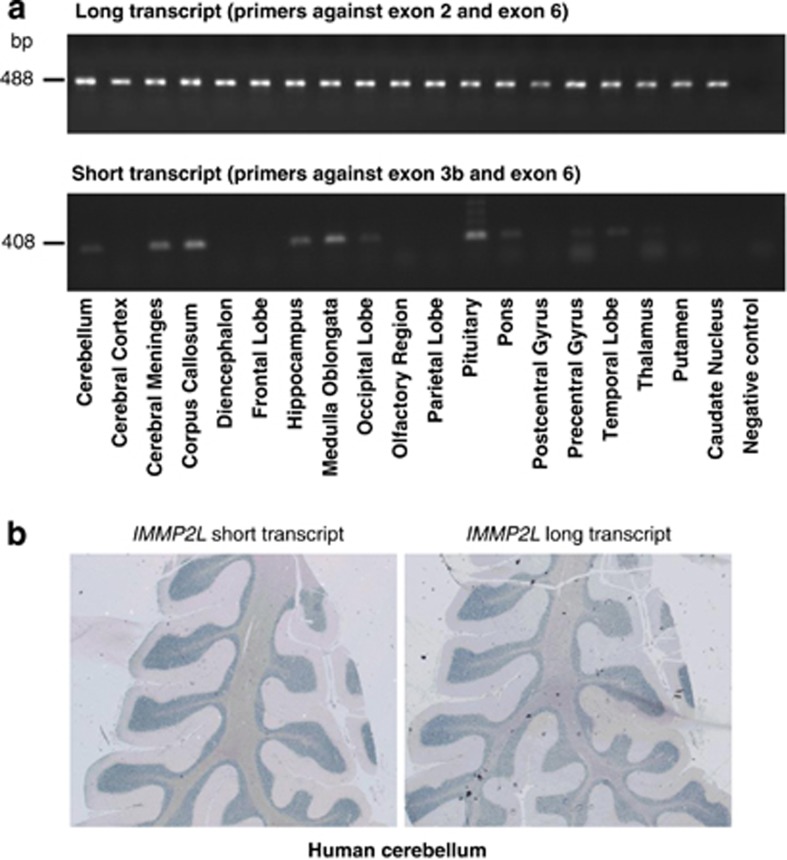
Expression studies of the IMMP2L transcripts in human brain. (a) The expression profile of long (above) and short (below) IMMP2L transcripts in 19 human brain regions using reverse transcription PCR. The long transcript is expressed in all regions, whereas the short transcript is expressed in the cerebellum, meninges, corpus callosum, hippocampus, medulla oblongata, occipital lobe, pituitary, pons, precentral gyrus, temporal lobe and thalamus. (b) Expression of the short and long transcripts of IMMP2L in adult human cerebellum showing expression especially in the granular layer and the medullar region.
In situ hybridization studies
In the whole-mouse embryo sections (E16.5), expression of the short transcript was more specific to the brain compared with the long transcript (RNA-In situ hybridization studies, data not shown). In the brain sections of the mouse embryos (E15.5), both transcripts were widely expressed at a moderate level (Figure 3a). In the 7 days old mice, both transcripts gave strong signals in the cerebellum and hippocampus (Figure 3b). On the sagittal sections of the adult mice brain both transcripts gave strong signals in the cerebellum and the hippocampus/dental gyrus, and the short transcript was also expressed in the olfactory bulb (Figure 3c). On the coronal sections, expression was detected in the hippocampus/dental gyrus, together with hypothalamic nuclei, fornix, cingulum, superior colliculus, posterior commissure, substantia nigra, amygdala, as well as putamen and thalamus (Figure 3d).
Figure 3.
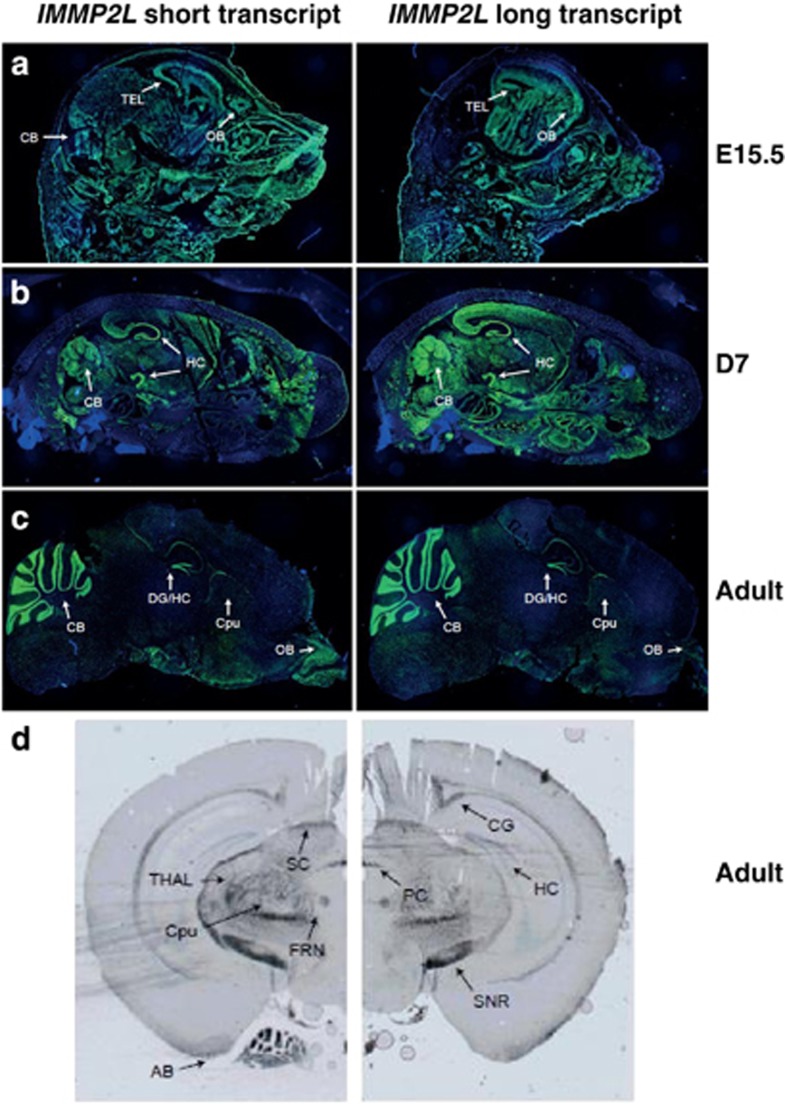
Expression of the short and long transcripts of IMMP2L in mouse brain sections. Sagittal sections from (a) mouse embryo (E15.5); (b) mouse at age of seven days (D7); and (c) adult mouse are hybridized with long (exon 3) and short (exon 3b) IMMP2L transcripts using fluorescence in situ hybridization. In the embryo, a general expression of both the transcripts were observed, whereas in the D7 mouse, and especially the adult mouse, the signals were mainly localized to the hippocampus/dental gyrus and the cerebellum. (d) The coronal sections (Bregma −2.92) of adult mouse were hybridized with both transcripts using DNP chromogenic detection. SC, superior commissure; CG, cingulum; PC, posterior commissure; TH, thalamus; CP, caudate putamen; FRN, fornix; HC, hippocampus; SNR, substantia nigra; AM, amygdala; DG, dentate gyrus, CB, cerebellum; OB, olfactory bulb; TEL, telencephalon.
In the human cerebellum, both transcripts were highly expressed in the granular layer of the cerebellar cortex and some staining was also obtained in the medulla composed of the white matter (Figure 2b).
Discussion
In this study, we investigated a cohort of 188 unrelated TS patients and identified seven IMMP2L deletions (Figure 1). Four deletions included only two exons (3a and 3b), which were not part of the long IMMP2L transcript (NM_001244606.1). This resulted in identification of an alternative transcript (ENST00000450877), where the ATG start codon was within exon 3a. Both transcripts were expressed in several brain regions. This study underlines that intronic deletions should be investigated further for presence of alternative transcription start sites, alternative exons and regulatory sequences.
Partial deletions within IMMP2L have not previously been associated with TS and deletions of this gene have been observed in control populations. However, the deletion frequency was significantly higher in the present TS cohort (3.7%) compared with the background population (0.9%) and the Affymetrix reference cohort (0.8%). The use of the Danish background population as a control cohort enabled us to minimize the population stratification bias between patients and controls, whereas the Affymetrix reference cohort ensured to eliminate a potential bias produced by different detection methods (chromosome microarray and qPCR). The present results, therefore, suggest that intragenic deletions of IMMP2L may be genetic risk factors in TS pathogenesis. In a previous study, IMMP2L deletions were not detected among 22 TS patients, but this may be due to the small size of this cohort or the localization of the amplicons used.10 The type of IMMP2L mutations found in TS patients is chromosome aberrations or deletions, but not small base-pair changes. This suggests that haploinsufficiency of one or more IMMP2L transcripts, rather than structural changes in the protein structure (eg, due to base-pair substitutions), is associated with TS susceptibility. However, screening of larger TS cohorts for small base-pair changes in IMMP2L is warranted to support this hypothesis.
IMMP2L has previously been associated with autism21, 22 and ADHD.23 Furthermore, in the Decipher database, deletions and duplications spanning IMMP2L are reported in patients with developmental delay and ASD. Notably, among the seven families investigated in this study, five index patients have ADHD and one has Asperger syndrome. These findings, together with the previous studies, suggest that IMMP2L may be one of the shared genetic factors between neurodevelopmental disorders. However, it is likely that IMMP2L deletions are not fully penetrant as underlined by the finding of deletions in unaffected parents (Table 1) and in control populations. Investigation of further and larger TS cohorts is necessary to understand the role of this gene in the pathogenesis of TS and other neurodevelopmental disorders.
In addition to the IMMP2L deletions, three patients have CNVs, which have previously been associated with neuropsychiatric disorders (Table 1). One patient with TS only (P5) has a duplication including the CHRNA7 gene (15q13.3) and duplications of this region have been observed across a range of neuropsychiatric phenotypes including intellectual disability, ASD and ADHD.24, 25, 26, 27, 28 It is therefore possible that both the IMMP2L deletion and the CHRNA7 duplication contribute to the disease pathogenesis in this patient. In another patient (P4) with TS and OCD both the IMMP2L deletion and the 4q35.2 duplication are inherited from the father with OCB and stuttering. This duplication includes five genes, and CNVs of TRIML1, TRIML2 and LOC401164 have been observed among ASD patients.29 It is thus likely that both CNVs contribute to the disease pathogenesis. In the third patient (P3) with TS, ADHD and Asperger syndrome, the IMMP2L deletion and the partial DISC1 duplication are inherited from phenotypically normal parents (mother and father, respectively). DISC1 has been implicated in several psychiatric disorders,30 including ADHD,31 and may contribute to the patients symptoms. Finding of these additional CNVs in TS patients with IMMP2L deletions supports the notion of a complex pattern of inheritance in TS, such as oligogenic heterozygosity or epistatic interactions.
Mitochondrial dysfunction has been associated with a range of human disorders, including neuropsychiatric disorders.32 A defective IMMP2L may therefore be related to the pathogenesis of neuropsychiatric disorders, including TS, due to a hyperactive mitochondrion with increased production of superoxide through a defective processing of IMMP2L substrates, such as cytochrome c1 and glycerol-3-phosphate dehydrogenase 2, leading to apoptosis. It is also possible that a defective IMMP2L leading to an unbalance in mitochondria function may be a risk factor affecting myelination. In support of this hypothesis, it has recently been reported that mitochondria dysfunction through oxidative stress, perturbed calcium homeostasis and release of pro-apoptotic factors could compromise myelinogenesis.33 Both IMMP2L transcripts show high expression in the cerebellum, which is involved in TS pathogenesis.34, 35 Mitochondria dysfunction at the cellular level and cerebellum involvement at the anatomic level are also related to other movement disorders such as dystonia and ataxia.36, 37 It is therefore plausible that an unbalanced mitochondria function through a defective cerebellar IMMP2L may contribute to tic formation. However, further studies are necessary to understand the involvement of mitochondria dysfunction due to a deficient IMMP2L in TS.
Different isoforms of IMMP2L may have differential roles in development and in different brain regions. In mouse embryo (E16.5), the short transcript is mainly expressed in the brain whereas the long transcript shows an ubiquitous expression (data not shown). In the E15.5 and D7 mouse both transcripts are expressed throughout the brain, whereas in the adult mouse both transcripts are mainly expressed in the hippocampus/dental gyrus and cerebellum (Figures 3a and b). In human adult brain (RNA panel), the long transcript shows an overall expression, whereas the short transcript shows a higher expression in the cerebellum, hippocampus and corpus callosum, regions which are shown to be involved in TS pathogenesis.38 It is thus possible that differences in the expression of the alternative IMMP2L transcripts in specific brain regions due to the deletions, may lead to TS susceptibility.39
Noradrenergic drugs, such as clonidine, are used in the treatment of patients with tics and ADHD, but evidence for the effect of this medicine is limited.40 Notably, clonidine has recently been associated with improvement in tic symptoms and overall well-being of a patient with a chromosome aberration involving IMMP2L.41 Further studies are warranted to investigate whether clonidine supplement is treatment of choice in patients with a nonfunctional IMMP2L gene.
The present findings together with previous studies support that genomic rearrangements affecting IMMP2L may be one of the predisposing factors involved in TS and overlapping neurodevelopmental disorders. Investigation of other larger cohorts for IMMP2L deletions and functional studies of both transcripts are necessary to gain a better insight into the role of this gene in disease pathogenesis.
Web Resources
UCSC Genome Browser, Human Genome Database Build hg19, February 2009 (http://www.genome.ucsc.edu).
Database of Genomic Variants (DGV), http://projects.tcag.ca/cgi-bin/variations/gbrowse/hg19.
Decipher database, Database of Chromosomal Imbalance and Phenotype in Humans; http://decipher.sanger.ac.uk.
Ensembl database, http://www.ensembl.org/index.html.
TMpred, http://www.ch.embnet.org/software/TMPRED_form.html.
SOSUI, http://bp.nuap.nagoya-u.ac.jp/sosui/sosui_submit.html.
Acknowledgments
We thank all the individuals and families for their participation and contribution to this work. We acknowledge the assistance of Judy Grejsen. This study was approved by the Danish Institutional Review Board (H-KA-05118). The study is supported by Lundbeck Foundation (R24-A2419). B B is supported with a fellowship from the University of Copenhagen.
The authors declare no conflict of interest.
Footnotes
Supplementary Information accompanies this paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary Material
References
- Leckman JF, Bloch MH, Scahill L, King RA. Tourette syndrome: the self under siege. J Child Neurol. 2006;21:642–649. doi: 10.1177/08830738060210081001. [DOI] [PubMed] [Google Scholar]
- Freeman RD, Fast DK, Burd L, Kerbeshian J, Robertson MM, Sandor P. An international perspective on Tourette syndrome: selected findings from 3,500 individuals in 22 countries. Dev Med Child Neurol. 2000;42:436–447. doi: 10.1017/s0012162200000839. [DOI] [PubMed] [Google Scholar]
- Debes NMM, Hjalgrim H, Skov L. Validation of the presence of comorbidities in a Danish cohort of children with Tourette Syndrome. J Child Neurol. 2008;23:1017–1027. doi: 10.1177/0883073808316370. [DOI] [PubMed] [Google Scholar]
- Clarke RA, Lee S, Eapen V. Pathogenetic model for Tourette syndrome delineates overlap with related neurodevelopmental disorders including Autism. Transl Psychiatry. 2012;2:e158. doi: 10.1038/tp.2012.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paschou P. The genetic basis of Gilles de la Tourette Syndrome. Neurosci Biobehav Rev. 2013;37:1026–1039. doi: 10.1016/j.neubiorev.2013.01.016. [DOI] [PubMed] [Google Scholar]
- Boghosian-Sell L, Comings DE, Overhauser J. Tourette syndrome in a pedigree with a 7;18 translocation: identification of a YAC spanning the translocation breakpoint at 18q22.3. Am J Hum Genet. 1996;59:999–1005. [PMC free article] [PubMed] [Google Scholar]
- Petek E, Windpassinger C, Vincent JB, et al. Disruption of a novel gene (IMMP2L) by a breakpoint in 7q31 associated with Tourette syndrome. Am J Hum Genet. 2001;68:848–858. doi: 10.1086/319523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kroisel PM, Petek E, Emberger W, Windpassinger C, Wladika W, Wagner K. Candidate region for Gilles de la Tourette syndrome at 7q31. Am J Med Genet. 2001;101:259–261. doi: 10.1002/1096-8628(20010701)101:3<259::aid-ajmg1374>3.0.co;2-#. [DOI] [PubMed] [Google Scholar]
- Díaz-Anzaldúa A, Joober R, Rivière JB, et al. Association between 7q31 markers and Tourette syndrome. Am J Med Genet A. 2004;127:17–20. doi: 10.1002/ajmg.a.20631. [DOI] [PubMed] [Google Scholar]
- Petek E, Schwarzbraun T, Noor A, et al. Molecular and genomic studies of IMMP2L and mutation screening in autism and Tourette syndrome. Mol Genet Genomics. 2007;277:71–81. doi: 10.1007/s00438-006-0173-1. [DOI] [PubMed] [Google Scholar]
- Patel C, Cooper-Charles L, McMullan DJ, Walker JM, Davison V, Morton J. Translocation breakpoint at 7q31 associated with tics: further evidence for IMMP2L as a candidate gene for Tourette syndrome. Eur J Hum Genet. 2011;19:634–639. doi: 10.1038/ejhg.2010.238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burri L, Strahm Y, Hawkins CJ, et al. Mature DIABLO/Smac is produced by the IMP protease complex on the mitochondrial inner membrane. Mol Biol Cell. 2005;16:2926–2933. doi: 10.1091/mbc.E04-12-1086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunnari J, Fox TD, Walter P. A mitochondrial protease with two catalytic subunits of nonoverlapping specificities. Science. 1993;262:1997–2004. doi: 10.1126/science.8266095. [DOI] [PubMed] [Google Scholar]
- Gakh O, Cavadini P, Isaya G. Mitochondrial processing peptides. Biochim Biophys Acta. 2002;1592:63–77. doi: 10.1016/s0167-4889(02)00265-3. [DOI] [PubMed] [Google Scholar]
- Luo W, Fang H, Green N. Substrate specificity of inner membrane peptidase in yeast mitochondria. Mol Genet Genomics. 2006;275:431–436. doi: 10.1007/s00438-006-0099-7. [DOI] [PubMed] [Google Scholar]
- Lu B, Poirier C, Gaspar T, et al. A mutation in the inner mitochondrial membrane peptidase 2-like gene (Immp2l) affects mitochondrial function and impairs fertility in mice. Biol Reprod. 2008;78:601–610. doi: 10.1095/biolreprod.107.065987. [DOI] [PubMed] [Google Scholar]
- Mattson MP, Gleichmann M, Cheng A. Mitochondria in neuroplasticity and neurological disorders. Neuron. 2008;60:748–766. doi: 10.1016/j.neuron.2008.10.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma Y, Metha SL, Lu B, Li PA. Deficiency in the inner mitochondrial membrane peptidase 2-like (Immp2l) gene increases ischemic brain damage and impairs mitochondrial function. Neurobiol Dis. 2011;44:270–276. doi: 10.1016/j.nbd.2011.06.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- American Psychiatric Association Diagnostic and Statistical Manual Text Revision (DSM-IV-TR)4th edn.Washington, DC: American Psychiatric Press; 2000 [Google Scholar]
- Silahtaroglu AN, Nolting D, Dyrskjøt L, et al. Detection of microRNAs in frozen tissue sections by fluorescence in situ hybridization using locked nucleic acid probes and tyramide signal amplification. Nat Protoc. 2007;2:2520–2528. doi: 10.1038/nprot.2007.313. [DOI] [PubMed] [Google Scholar]
- International Molecular Genetic Study of Autism Consortium A full genome screen for autism with evidence for linkage to a region on chromosome 7q. Hum Mol Genet. 1998;7:571–578. doi: 10.1093/hmg/7.3.571. [DOI] [PubMed] [Google Scholar]
- Maestrini E, Pagnamenta AT, Lamb JA, et al. High-density SNP association study and copy number variation analysis of the AUTS1 and AUTS5 loci implicate the IMMP2L-DOCK4 gene region in autism susceptibility. Mol Psychiatry. 2010;15:954–968. doi: 10.1038/mp.2009.34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elia J, Gai X, Xie HM, et al. Rare structural variants found in attention-deficit hyperactivity disorder are preferentially associated with neurodevelopmental genes. Mol Psychiatry. 2010;15:637–646. doi: 10.1038/mp.2009.57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Bon BW, Mefford HC, Menten B, et al. Further delineation of the 15q31 microdeletion and duplication syndromes: a clinical spectrum varying from non-pathogenic to a severe outcome. J Med Genet. 2009;46:511–523. doi: 10.1136/jmg.2008.063412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leblond CS, Heinrich J, Delorme R, et al. Genetic and functional analyses of SHANK2 mutations suggest a multiple hit model of autism spectrum disorders. PLoS Genet. 2012;8:e1002521. doi: 10.1371/journal.pgen.1002521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams NM, Franke B, Mick E, et al. Genome-wide analysis of copy number variants in attention deficit hyperactivity disorder: the role of rare variants and duplications at 15q13.3. Am J Psychiatry. 2012;169:195–204. doi: 10.1176/appi.ajp.2011.11060822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chilian B, Abdollahpour H, Bierhals T, et al. Dysfunction of SHANK2 and CHRNA7 in a patient with intellectual disability and language impairment supports genetic epistasis of the two loci. Clin Genet. 2013;84:560–565. doi: 10.1111/cge.12105. [DOI] [PubMed] [Google Scholar]
- Melchior L, Bertelsen B, Debes NM, et al. Microduplication of 15q13.3 and Xq21.31 in a family with tourette syndrome and comorbidities. Am J Med Genet B Neuropsychiatr Genet. 2013;162:825–831. doi: 10.1002/ajmg.b.32186. [DOI] [PubMed] [Google Scholar]
- Menashe I, Larsen EC, Banerjee-Basu S. Prioritization of copy number variation loci associated with autism from AutDB–An integrative multi-study genetics database. PLoS One. 2013;8:e66707. doi: 10.1371/journal.pone.0066707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomson PA, Malavasi EL, Grünewald E, Soares DC, Borkowska M, Millar JK. DISC1 genetics, biology and psychiatric illness. Front Biol. 2013;8:1–31. doi: 10.1007/s11515-012-1254-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobsen KK, Halmøy A, Sánchez-Mora C, et al. DISC1 in adult ADHD patients: an association study in two European samples. Am J Med Genet B. 2013;162:227–234. doi: 10.1002/ajmg.b.32136. [DOI] [PubMed] [Google Scholar]
- Rezin GT, Amboni G, Zugno AI, Quevedo J, Streck EL. Mitochondrial dysfunction and psychiatric disorders. Neurochem Res. 2009;34:1021–1029. doi: 10.1007/s11064-008-9865-8. [DOI] [PubMed] [Google Scholar]
- Barateiro A, Vaz AR, Silva SL, Fernandes A, Brites D. ER stress, mitochondrial dysfunction and calpain/JNK activation are involved in oligodendrocyte precursor cell death by unconjugated bilirubin. Neuromolecular Med. 2012;14:285–302. doi: 10.1007/s12017-012-8187-9. [DOI] [PubMed] [Google Scholar]
- Bohlhalter S, Goldfine A, Matteson S, et al. Neural correlates of tic generation in Tourette syndrome: an event-related functional MRI study. Brain. 2006;129:2029–2037. doi: 10.1093/brain/awl050. [DOI] [PubMed] [Google Scholar]
- Tobe RH, Bansal R, Xu D, et al. Cerebellar morphology in Tourette syndrome and obsessive-compulsive disorder. Ann Neurol. 2010;67:479–487. doi: 10.1002/ana.21918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson VB, Jinnah HA, Hess EJ. Convergent mechanisms in etiologically-diverse dystonia. Brain. 2006;129:1357–1370. doi: 10.1517/14728222.2011.641533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dueñas AM, Goold R, Giunti P. Molecular pathogenesis of spinocerebellar ataxias. Brain. 2006;129:1357–1370. doi: 10.1093/brain/awl081. [DOI] [PubMed] [Google Scholar]
- Robertson MM. The Gilles de la Tourette syndrome: the current status. Arch Dis Child Educ Pract Ed. 2012;97:166–175. doi: 10.1136/archdischild-2011-300585. [DOI] [PubMed] [Google Scholar]
- Tian Y, Liao IH, Zhan X, et al. Exon expression and alternatively spliced genes in Tourette Syndrome. Am J Med Genet B. 2011;156:72–78. doi: 10.1002/ajmg.b.31140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilbert D. Treatment of children and adolescents with tics and Tourette syndrome. Neurology. 2006;21:690–700. doi: 10.1177/08830738060210080401. [DOI] [PubMed] [Google Scholar]
- Katuwawela I, Cavanna AE. Good response to clonidine in tourette syndrome associated with chromosomal translocation involving the IMMP2L gene. J Neuropsychiatry Clin Neurosci. 2012;24:E17. doi: 10.1176/appi.neuropsych.11010032. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.