Abstract
A fundamental catalytic principle for protein enzymes in the use of binding interactions away from the site of chemical transformation for catalysis. We have compared the binding and reactivity of a series of oligonucleotide substrates and products of the Tetrahymena ribozyme, which catalyzes a site-specific phosphodiester cleavage reaction: CCCUCUpA+G<-->CCCUCU-OH+GpA. The results suggest that this RNA enzyme, like protein enzymes, can utilize binding interactions to achieve substantial catalysis via entropic fixation and substrate destabilization. The stronger binding of the all-ribose oligonucleotide product compared to an analog with a terminal 3' deoxyribose residue gives an effective concentration of 2200 M for the 3' hydroxyl group, a value approaching those obtained with protein enzymes and suggesting the presence of a structurally well defined active site capable of precise positioning. The stabilization from tertiary binding interactions is 40-fold less for the oligonucleotide substrate than the oligonucleotide product, despite the presence of the reactive phosphoryl group in the substrate. This destabilization is accounted for by a model in which tertiary interactions away from the site of bond cleavage position the electron-deficient 3' bridging phosphoryl oxygen of the oligonucleotide substrate next to an electropositive Mg ion. As the phosphodiester bond breaks and this 3' oxygen atom develops a negative charge in the transition state, the weak interaction of the substrate with Mg2+ becomes strong. These strategies of "substrate destabilization" and "transition state stabilization" provide estimated rate enhancements of approximately 280- and approximately 60-fold, respectively. Analogous substrate destabilization by a metal ion or hydrogen bond donor may be used more generally by RNA and protein enzymes catalyzing reactions of phosphate esters.
Full text
PDF
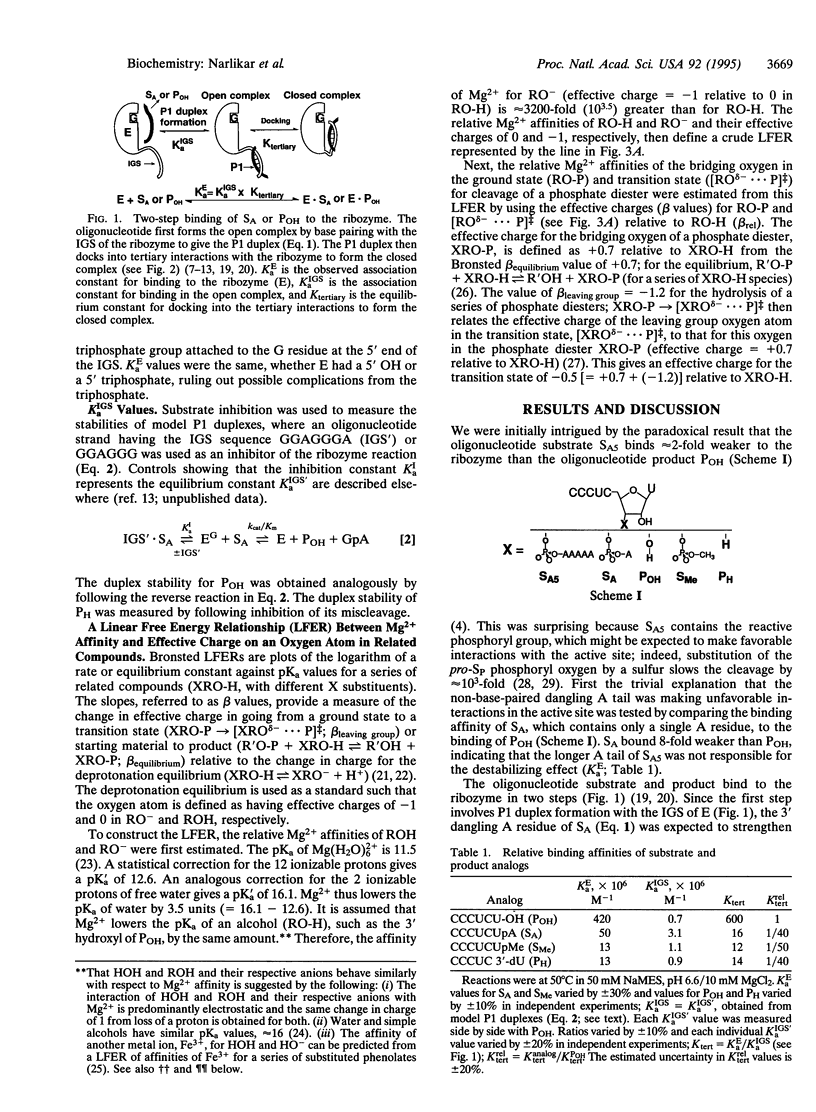
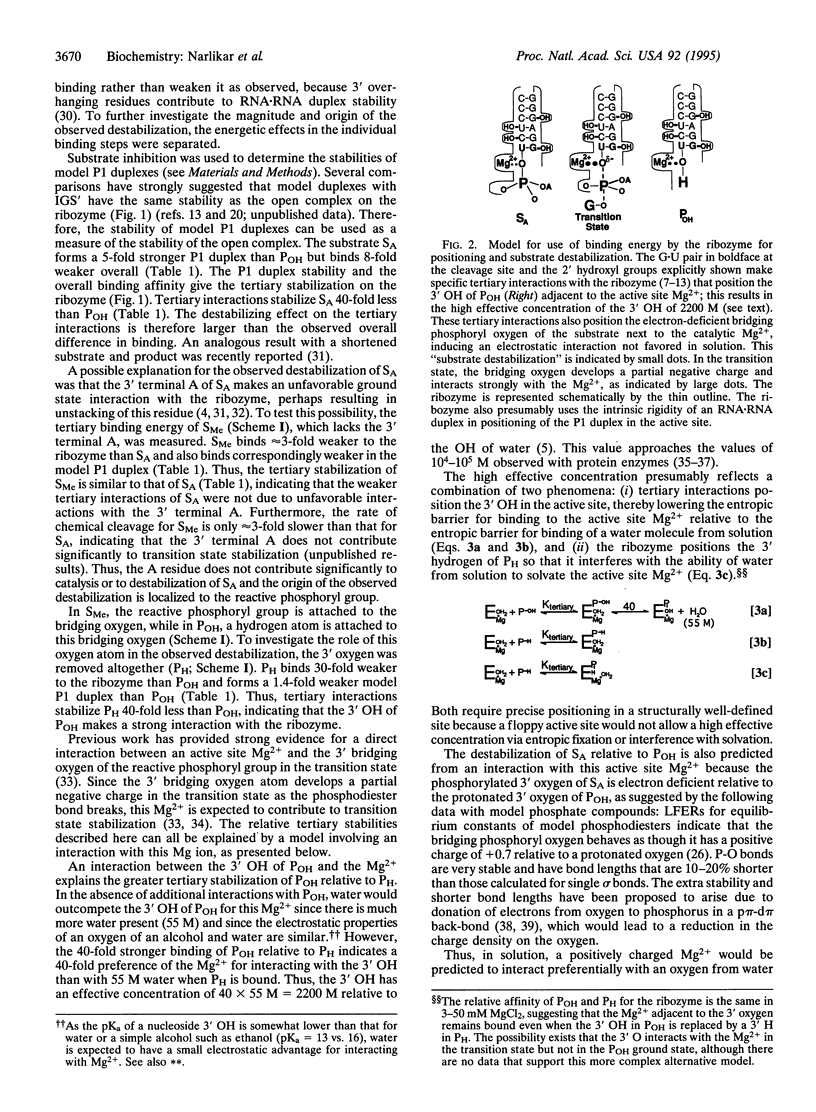



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altman S. Ribonuclease P: an enzyme with a catalytic RNA subunit. Adv Enzymol Relat Areas Mol Biol. 1989;62:1–36. doi: 10.1002/9780470123089.ch1. [DOI] [PubMed] [Google Scholar]
- Beebe J. A., Fierke C. A. A kinetic mechanism for cleavage of precursor tRNA(Asp) catalyzed by the RNA component of Bacillus subtilis ribonuclease P. Biochemistry. 1994 Aug 30;33(34):10294–10304. doi: 10.1021/bi00200a009. [DOI] [PubMed] [Google Scholar]
- Bevilacqua P. C., Kierzek R., Johnson K. A., Turner D. H. Dynamics of ribozyme binding of substrate revealed by fluorescence-detected stopped-flow methods. Science. 1992 Nov 20;258(5086):1355–1358. doi: 10.1126/science.1455230. [DOI] [PubMed] [Google Scholar]
- Bevilacqua P. C., Li Y., Turner D. H. Fluorescence-detected stopped flow with a pyrene labeled substrate reveals that guanosine facilitates docking of the 5' cleavage site into a high free energy binding mode in the Tetrahymena ribozyme. Biochemistry. 1994 Sep 20;33(37):11340–11348. doi: 10.1021/bi00203a032. [DOI] [PubMed] [Google Scholar]
- Bevilacqua P. C., Turner D. H. Comparison of binding of mixed ribose-deoxyribose analogues of CUCU to a ribozyme and to GGAGAA by equilibrium dialysis: evidence for ribozyme specific interactions with 2' OH groups. Biochemistry. 1991 Nov 5;30(44):10632–10640. doi: 10.1021/bi00108a005. [DOI] [PubMed] [Google Scholar]
- Bode W. The transition of bovine trypsinogen to a trypsin-like state upon strong ligand binding. II. The binding of the pancreatic trypsin inhibitor and of isoleucine-valine and of sequentially related peptides to trypsinogen and to p-guanidinobenzoate-trypsinogen. J Mol Biol. 1979 Feb 5;127(4):357–374. doi: 10.1016/0022-2836(79)90227-4. [DOI] [PubMed] [Google Scholar]
- Cech T. R. Self-splicing of group I introns. Annu Rev Biochem. 1990;59:543–568. doi: 10.1146/annurev.bi.59.070190.002551. [DOI] [PubMed] [Google Scholar]
- Cotton F. A., Hazen E. E., Jr, Legg M. J. Staphylococcal nuclease: proposed mechanism of action based on structure of enzyme-thymidine 3',5'-bisphosphate-calcium ion complex at 1.5-A resolution. Proc Natl Acad Sci U S A. 1979 Jun;76(6):2551–2555. doi: 10.1073/pnas.76.6.2551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freemont P. S., Friedman J. M., Beese L. S., Sanderson M. R., Steitz T. A. Cocrystal structure of an editing complex of Klenow fragment with DNA. Proc Natl Acad Sci U S A. 1988 Dec;85(23):8924–8928. doi: 10.1073/pnas.85.23.8924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freier S. M., Kierzek R., Jaeger J. A., Sugimoto N., Caruthers M. H., Neilson T., Turner D. H. Improved free-energy parameters for predictions of RNA duplex stability. Proc Natl Acad Sci U S A. 1986 Dec;83(24):9373–9377. doi: 10.1073/pnas.83.24.9373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herschlag D., Cech T. R. Catalysis of RNA cleavage by the Tetrahymena thermophila ribozyme. 1. Kinetic description of the reaction of an RNA substrate complementary to the active site. Biochemistry. 1990 Nov 6;29(44):10159–10171. doi: 10.1021/bi00496a003. [DOI] [PubMed] [Google Scholar]
- Herschlag D., Cech T. R. Catalysis of RNA cleavage by the Tetrahymena thermophila ribozyme. 2. Kinetic description of the reaction of an RNA substrate that forms a mismatch at the active site. Biochemistry. 1990 Nov 6;29(44):10172–10180. doi: 10.1021/bi00496a004. [DOI] [PubMed] [Google Scholar]
- Herschlag D., Eckstein F., Cech T. R. Contributions of 2'-hydroxyl groups of the RNA substrate to binding and catalysis by the Tetrahymena ribozyme. An energetic picture of an active site composed of RNA. Biochemistry. 1993 Aug 17;32(32):8299–8311. doi: 10.1021/bi00083a034. [DOI] [PubMed] [Google Scholar]
- Herschlag D., Eckstein F., Cech T. R. The importance of being ribose at the cleavage site in the Tetrahymena ribozyme reaction. Biochemistry. 1993 Aug 17;32(32):8312–8321. doi: 10.1021/bi00083a035. [DOI] [PubMed] [Google Scholar]
- Herschlag D. Evidence for processivity and two-step binding of the RNA substrate from studies of J1/2 mutants of the Tetrahymena ribozyme. Biochemistry. 1992 Feb 11;31(5):1386–1399. doi: 10.1021/bi00120a015. [DOI] [PubMed] [Google Scholar]
- Herschlag D., Piccirilli J. A., Cech T. R. Ribozyme-catalyzed and nonenzymatic reactions of phosphate diesters: rate effects upon substitution of sulfur for a nonbridging phosphoryl oxygen atom. Biochemistry. 1991 May 21;30(20):4844–4854. doi: 10.1021/bi00234a003. [DOI] [PubMed] [Google Scholar]
- Jencks W. P. Binding energy, specificity, and enzymic catalysis: the circe effect. Adv Enzymol Relat Areas Mol Biol. 1975;43:219–410. doi: 10.1002/9780470122884.ch4. [DOI] [PubMed] [Google Scholar]
- Kim E. E., Wyckoff H. W. Reaction mechanism of alkaline phosphatase based on crystal structures. Two-metal ion catalysis. J Mol Biol. 1991 Mar 20;218(2):449–464. doi: 10.1016/0022-2836(91)90724-k. [DOI] [PubMed] [Google Scholar]
- Knitt D. S., Narlikar G. J., Herschlag D. Dissection of the role of the conserved G.U pair in group I RNA self-splicing. Biochemistry. 1994 Nov 22;33(46):13864–13879. doi: 10.1021/bi00250a041. [DOI] [PubMed] [Google Scholar]
- Kruger K., Grabowski P. J., Zaug A. J., Sands J., Gottschling D. E., Cech T. R. Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell. 1982 Nov;31(1):147–157. doi: 10.1016/0092-8674(82)90414-7. [DOI] [PubMed] [Google Scholar]
- McConnell T. S., Cech T. R., Herschlag D. Guanosine binding to the Tetrahymena ribozyme: thermodynamic coupling with oligonucleotide binding. Proc Natl Acad Sci U S A. 1993 Sep 15;90(18):8362–8366. doi: 10.1073/pnas.90.18.8362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michel F., Hanna M., Green R., Bartel D. P., Szostak J. W. The guanosine binding site of the Tetrahymena ribozyme. Nature. 1989 Nov 23;342(6248):391–395. doi: 10.1038/342391a0. [DOI] [PubMed] [Google Scholar]
- Moran S., Kierzek R., Turner D. H. Binding of guanosine and 3' splice site analogues to a group I ribozyme: interactions with functional groups of guanosine and with additional nucleotides. Biochemistry. 1993 May 18;32(19):5247–5256. doi: 10.1021/bi00070a037. [DOI] [PubMed] [Google Scholar]
- Nakamura C. E., Abeles R. H. Mode of interaction of beta-hydroxy-beta-methylglutaryl coenzyme A reductase with strong binding inhibitors: compactin and related compounds. Biochemistry. 1985 Mar 12;24(6):1364–1376. doi: 10.1021/bi00327a014. [DOI] [PubMed] [Google Scholar]
- Piccirilli J. A., Vyle J. S., Caruthers M. H., Cech T. R. Metal ion catalysis in the Tetrahymena ribozyme reaction. Nature. 1993 Jan 7;361(6407):85–88. doi: 10.1038/361085a0. [DOI] [PubMed] [Google Scholar]
- Pon R. T., Usman N., Ogilvie K. K. Derivatization of controlled pore glass beads for solid phase oligonucleotide synthesis. Biotechniques. 1988 Sep;6(8):768–775. [PubMed] [Google Scholar]
- Pyle A. M., Cech T. R. Ribozyme recognition of RNA by tertiary interactions with specific ribose 2'-OH groups. Nature. 1991 Apr 18;350(6319):628–631. doi: 10.1038/350628a0. [DOI] [PubMed] [Google Scholar]
- Pyle A. M., Moran S., Strobel S. A., Chapman T., Turner D. H., Cech T. R. Replacement of the conserved G.U with a G-C pair at the cleavage site of the Tetrahymena ribozyme decreases binding, reactivity, and fidelity. Biochemistry. 1994 Nov 22;33(46):13856–13863. doi: 10.1021/bi00250a040. [DOI] [PubMed] [Google Scholar]
- Pyle A. M., Murphy F. L., Cech T. R. RNA substrate binding site in the catalytic core of the Tetrahymena ribozyme. Nature. 1992 Jul 9;358(6382):123–128. doi: 10.1038/358123a0. [DOI] [PubMed] [Google Scholar]
- Rajagopal J., Doudna J. A., Szostak J. W. Stereochemical course of catalysis by the Tetrahymena ribozyme. Science. 1989 May 12;244(4905):692–694. doi: 10.1126/science.2470151. [DOI] [PubMed] [Google Scholar]
- Scaringe S. A., Francklyn C., Usman N. Chemical synthesis of biologically active oligoribonucleotides using beta-cyanoethyl protected ribonucleoside phosphoramidites. Nucleic Acids Res. 1990 Sep 25;18(18):5433–5441. doi: 10.1093/nar/18.18.5433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlichting I., Almo S. C., Rapp G., Wilson K., Petratos K., Lentfer A., Wittinghofer A., Kabsch W., Pai E. F., Petsko G. A. Time-resolved X-ray crystallographic study of the conformational change in Ha-Ras p21 protein on GTP hydrolysis. Nature. 1990 May 24;345(6273):309–315. doi: 10.1038/345309a0. [DOI] [PubMed] [Google Scholar]
- Strobel S. A., Cech T. R. Tertiary interactions with the internal guide sequence mediate docking of the P1 helix into the catalytic core of the Tetrahymena ribozyme. Biochemistry. 1993 Dec 14;32(49):13593–13604. doi: 10.1021/bi00212a027. [DOI] [PubMed] [Google Scholar]
- Strobel S. A., Cech T. R. Translocation of an RNA duplex on a ribozyme. Nat Struct Biol. 1994 Jan;1(1):13–17. doi: 10.1038/nsb0194-13. [DOI] [PubMed] [Google Scholar]
- Suck D., Oefner C. Structure of DNase I at 2.0 A resolution suggests a mechanism for binding to and cutting DNA. Nature. 1986 Jun 5;321(6070):620–625. doi: 10.1038/321620a0. [DOI] [PubMed] [Google Scholar]
- Thompson R. C. Binding of peptides to elastase: implications for the mechanism of substrate hydrolysis. Biochemistry. 1974 Dec 31;13(27):5495–5501. doi: 10.1021/bi00724a007. [DOI] [PubMed] [Google Scholar]
- Zaug A. J., Grosshans C. A., Cech T. R. Sequence-specific endoribonuclease activity of the Tetrahymena ribozyme: enhanced cleavage of certain oligonucleotide substrates that form mismatched ribozyme-substrate complexes. Biochemistry. 1988 Dec 13;27(25):8924–8931. doi: 10.1021/bi00425a008. [DOI] [PubMed] [Google Scholar]
- Zhang Y., Liang J. Y., Huang S., Ke H., Lipscomb W. N. Crystallographic studies of the catalytic mechanism of the neutral form of fructose-1,6-bisphosphatase. Biochemistry. 1993 Feb 23;32(7):1844–1857. doi: 10.1021/bi00058a019. [DOI] [PubMed] [Google Scholar]


