Abstract
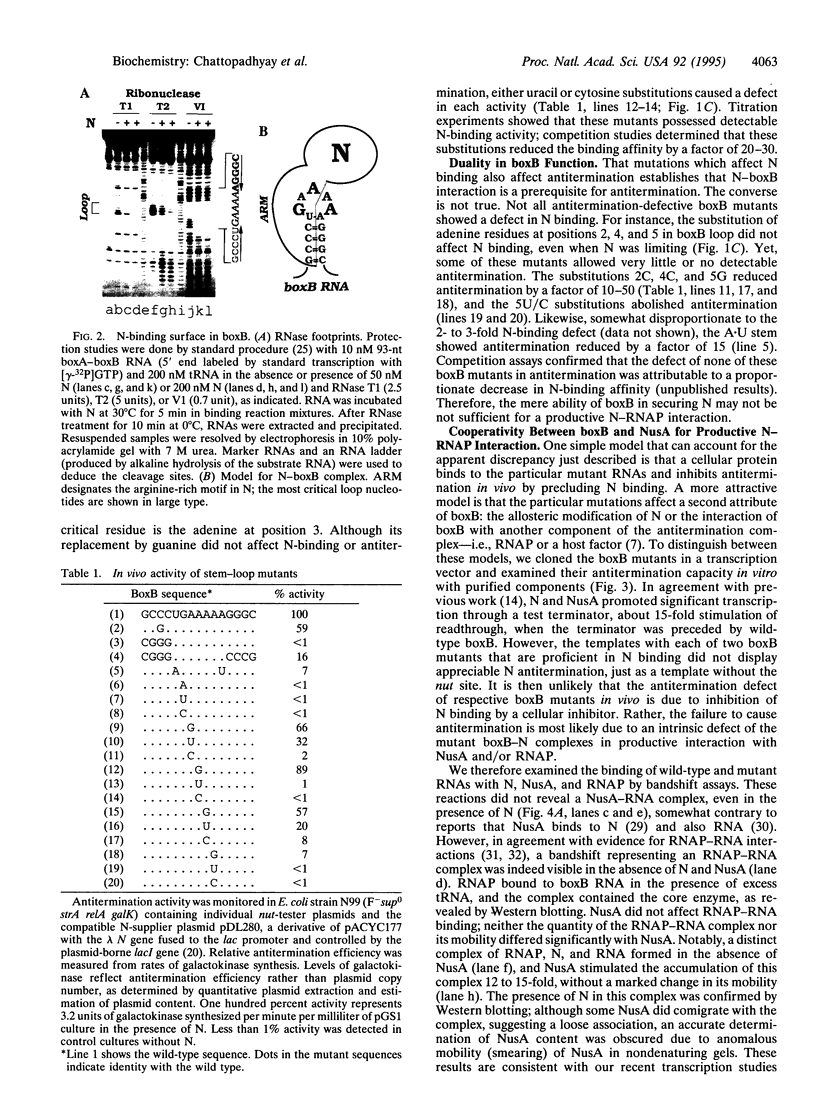
Transcription of downstream genes in the early operons of phage lambda requires a promoter-proximal element known as nut. This site acts in cis in the form of RNA to assemble a transcription antitermination complex which is composed of lambda N protein and at least four host factors. The nut-site RNA contains a small stem-loop structure called boxB. Here, we show that boxB RNA binds to N protein with high affinity and specificity. While N binding is confined to the 5' subdomain of the stem-loop, specific N recognition relies on both an intact stem-loop structure and two critical nucleotides in the pentamer loop. Substitutions of these nucleotides affect both N binding and antitermination. Remarkably, substitutions of other loop nucleotides also diminish antitermination in vivo, yet they have no detectable effect on N binding in vitro. These 3' loop mutants fail to support antitermination in a minimal system with RNA polymerase (RNAP), N, and the host factor NusA. Furthermore, the ability of NusA to stimulate the formation of the RNAP-boxB-N complex is diminished with these mutants. Hence, we suggest that boxB RNA performs two critical functions in antitermination. First, boxB binds to N and secures it near RNAP to enhance their interaction, presumably by increasing the local concentration of N. Second, boxB cooperates with NusA, most likely to bring N and RNAP in close contact and transform RNAP to the termination-resistant state.
Full text
PDF

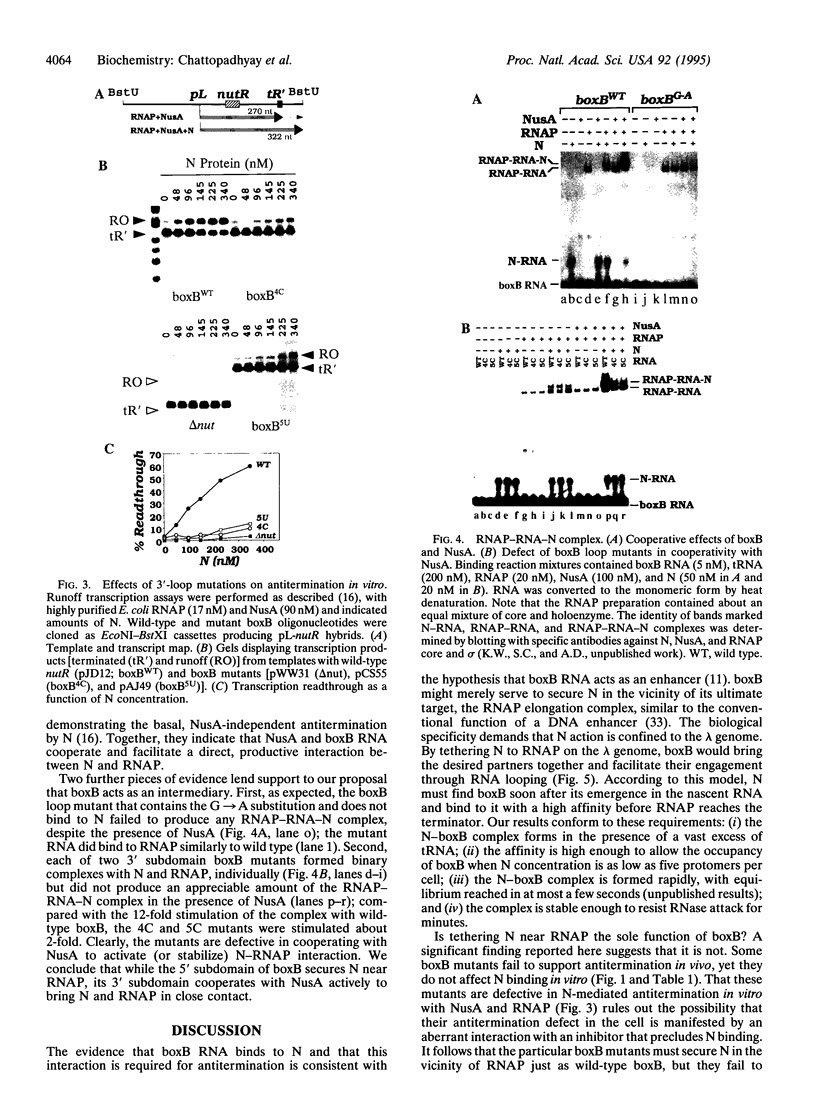

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barik S., Ghosh B., Whalen W., Lazinski D., Das A. An antitermination protein engages the elongating transcription apparatus at a promoter-proximal recognition site. Cell. 1987 Sep 11;50(6):885–899. doi: 10.1016/0092-8674(87)90515-0. [DOI] [PubMed] [Google Scholar]
- Baron J., Weisberg R. A. Mutations of the phage lambda nutL region that prevent the action of Nun, a site-specific transcription termination factor. J Bacteriol. 1992 Mar;174(6):1983–1989. doi: 10.1128/jb.174.6.1983-1989.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Busby S., Spassky A., Buc H. On the binding of tRNA to Escherichia coli RNA polymerase. Interactions between the core enzyme, DNA and tRNA. Eur J Biochem. 1981 Sep 1;118(3):443–451. doi: 10.1111/j.1432-1033.1981.tb05540.x. [DOI] [PubMed] [Google Scholar]
- Carey J. Gel retardation. Methods Enzymol. 1991;208:103–117. doi: 10.1016/0076-6879(91)08010-f. [DOI] [PubMed] [Google Scholar]
- Chamberlin M. J. New models for the mechanism of transcription elongation and its regulation. Harvey Lect. 1992 1993;88:1–21. [PubMed] [Google Scholar]
- Das A. Control of transcription termination by RNA-binding proteins. Annu Rev Biochem. 1993;62:893–930. doi: 10.1146/annurev.bi.62.070193.004333. [DOI] [PubMed] [Google Scholar]
- Das A. How the phage lambda N gene product suppresses transcription termination: communication of RNA polymerase with regulatory proteins mediated by signals in nascent RNA. J Bacteriol. 1992 Nov;174(21):6711–6716. doi: 10.1128/jb.174.21.6711-6716.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeVito J., Das A. Control of transcription processivity in phage lambda: Nus factors strengthen the termination-resistant state of RNA polymerase induced by N antiterminator. Proc Natl Acad Sci U S A. 1994 Aug 30;91(18):8660–8664. doi: 10.1073/pnas.91.18.8660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doelling J. H., Franklin N. C. Effects of all single base substitutions in the loop of boxB on antitermination of transcription by bacteriophage lambda's N protein. Nucleic Acids Res. 1989 Jul 25;17(14):5565–5577. doi: 10.1093/nar/17.14.5565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franklin N. C. Clustered arginine residues of bacteriophage lambda N protein are essential to antitermination of transcription, but their locale cannot compensate for boxB loop defects. J Mol Biol. 1993 May 20;231(2):343–360. doi: 10.1006/jmbi.1993.1287. [DOI] [PubMed] [Google Scholar]
- Franklin N. C. Conservation of genome form but not sequence in the transcription antitermination determinants of bacteriophages lambda, phi 21 and P22. J Mol Biol. 1985 Jan 5;181(1):75–84. doi: 10.1016/0022-2836(85)90325-0. [DOI] [PubMed] [Google Scholar]
- Friedman D. I., Olson E. R., Johnson L. L., Alessi D., Craven M. G. Transcription-dependent competition for a host factor: the function and optimal sequence of the phage lambda boxA transcription antitermination signal. Genes Dev. 1990 Dec;4(12A):2210–2222. doi: 10.1101/gad.4.12a.2210. [DOI] [PubMed] [Google Scholar]
- Grayhack E. J., Yang X. J., Lau L. F., Roberts J. W. Phage lambda gene Q antiterminator recognizes RNA polymerase near the promoter and accelerates it through a pause site. Cell. 1985 Aug;42(1):259–269. doi: 10.1016/s0092-8674(85)80121-5. [DOI] [PubMed] [Google Scholar]
- Greenblatt J., Li J. Interaction of the sigma factor and the nusA gene protein of E. coli with RNA polymerase in the initiation-termination cycle of transcription. Cell. 1981 May;24(2):421–428. doi: 10.1016/0092-8674(81)90332-9. [DOI] [PubMed] [Google Scholar]
- Greenblatt J., Li J. Interaction of the sigma factor and the nusA gene protein of E. coli with RNA polymerase in the initiation-termination cycle of transcription. Cell. 1981 May;24(2):421–428. doi: 10.1016/0092-8674(81)90332-9. [DOI] [PubMed] [Google Scholar]
- Greenblatt J., Li J. The nusA gene protein of Escherichia coli. Its identification and a demonstration that it interacts with the gene N transcription anti-termination protein of bacteriophage lambda. J Mol Biol. 1981 Mar 25;147(1):11–23. doi: 10.1016/0022-2836(81)90076-0. [DOI] [PubMed] [Google Scholar]
- Greenblatt J., Nodwell J. R., Mason S. W. Transcriptional antitermination. Nature. 1993 Jul 29;364(6436):401–406. doi: 10.1038/364401a0. [DOI] [PubMed] [Google Scholar]
- Horwitz R. J., Li J., Greenblatt J. An elongation control particle containing the N gene transcriptional antitermination protein of bacteriophage lambda. Cell. 1987 Nov 20;51(4):631–641. doi: 10.1016/0092-8674(87)90132-2. [DOI] [PubMed] [Google Scholar]
- Knapp G. Enzymatic approaches to probing of RNA secondary and tertiary structure. Methods Enzymol. 1989;180:192–212. doi: 10.1016/0076-6879(89)80102-8. [DOI] [PubMed] [Google Scholar]
- Landick R., Yanofsky C. Isolation and structural analysis of the Escherichia coli trp leader paused transcription complex. J Mol Biol. 1987 Jul 20;196(2):363–377. doi: 10.1016/0022-2836(87)90697-8. [DOI] [PubMed] [Google Scholar]
- Landick R., Yanofsky C. Isolation and structural analysis of the Escherichia coli trp leader paused transcription complex. J Mol Biol. 1987 Jul 20;196(2):363–377. doi: 10.1016/0022-2836(87)90697-8. [DOI] [PubMed] [Google Scholar]
- Lazinski D., Grzadzielska E., Das A. Sequence-specific recognition of RNA hairpins by bacteriophage antiterminators requires a conserved arginine-rich motif. Cell. 1989 Oct 6;59(1):207–218. doi: 10.1016/0092-8674(89)90882-9. [DOI] [PubMed] [Google Scholar]
- Mason S. W., Li J., Greenblatt J. Host factor requirements for processive antitermination of transcription and suppression of pausing by the N protein of bacteriophage lambda. J Biol Chem. 1992 Sep 25;267(27):19418–19426. [PubMed] [Google Scholar]
- Nodwell J. R., Greenblatt J. Recognition of boxA antiterminator RNA by the E. coli antitermination factors NusB and ribosomal protein S10. Cell. 1993 Jan 29;72(2):261–268. doi: 10.1016/0092-8674(93)90665-d. [DOI] [PubMed] [Google Scholar]
- Nodwell J. R., Greenblatt J. The nut site of bacteriophage lambda is made of RNA and is bound by transcription antitermination factors on the surface of RNA polymerase. Genes Dev. 1991 Nov;5(11):2141–2151. doi: 10.1101/gad.5.11.2141. [DOI] [PubMed] [Google Scholar]
- Patterson T. A., Zhang Z., Baker T., Johnson L. L., Friedman D. I., Court D. L. Bacteriophage lambda N-dependent transcription antitermination. Competition for an RNA site may regulate antitermination. J Mol Biol. 1994 Feb 11;236(1):217–228. doi: 10.1006/jmbi.1994.1131. [DOI] [PubMed] [Google Scholar]
- Ring B. Z., Roberts J. W. Function of a nontranscribed DNA strand site in transcription elongation. Cell. 1994 Jul 29;78(2):317–324. doi: 10.1016/0092-8674(94)90300-x. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Telesnitsky A. P., Chamberlin M. J. Sequences linked to prokaryotic promoters can affect the efficiency of downstream termination sites. J Mol Biol. 1989 Jan 20;205(2):315–330. doi: 10.1016/0022-2836(89)90343-4. [DOI] [PubMed] [Google Scholar]
- Telesnitsky A. P., Chamberlin M. J. Sequences linked to prokaryotic promoters can affect the efficiency of downstream termination sites. J Mol Biol. 1989 Jan 20;205(2):315–330. doi: 10.1016/0022-2836(89)90343-4. [DOI] [PubMed] [Google Scholar]
- Tsugawa A., Kurihara T., Zuber M., Court D. L., Nakamura Y. E. coli NusA protein binds in vitro to an RNA sequence immediately upstream of the boxA signal of bacteriophage lambda. EMBO J. 1985 Sep;4(9):2337–2342. doi: 10.1002/j.1460-2075.1985.tb03935.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whalen W. A., Das A. Action of an RNA site at a distance: role of the nut genetic signal in transcription antitermination by phage-lambda N gene product. New Biol. 1990 Nov;2(11):975–991. [PubMed] [Google Scholar]
- Whalen W., Ghosh B., Das A. NusA protein is necessary and sufficient in vitro for phage lambda N gene product to suppress a rho-independent terminator placed downstream of nutL. Proc Natl Acad Sci U S A. 1988 Apr;85(8):2494–2498. doi: 10.1073/pnas.85.8.2494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yarnell W. S., Roberts J. W. The phage lambda gene Q transcription antiterminator binds DNA in the late gene promoter as it modifies RNA polymerase. Cell. 1992 Jun 26;69(7):1181–1189. doi: 10.1016/0092-8674(92)90639-t. [DOI] [PubMed] [Google Scholar]