Abstract
We report a regulation system in Escherichia coli for independent regulation of two distinct reporter genes by application of Tet repressors with different specificities. One Tet repressor variant comprises wild-type tet operator (tetO) recognition and exclusive induction with the novel inducer 4-dedimethylamino-anhydrotetracycline. The other Tet repressor variant shows tetO-4C recognition and induction with tetracycline. We demonstrate that both variants are independently active in vivo and allow selective regulation of two genes in the same cell without any cross talk.
Inducible promoters are powerful tools for studying gene function in prokaryotes. The most widely used regulated expression systems in Escherichia coli have been developed from the lac, ara, and tet genes (2, 9, 14), while the tet, spac, and xyl systems have been used mostly with gram-positive bacteria (1, 6, 7, 16, 18). The advantage of tet regulation lies in the combination of tight control and sensitive induction by compounds that diffuse passively across biological membranes and do not require the presence of uptake proteins. Since complex cellular processes are often determined by more than one gene, multiple gene regulation systems that can be distinctly addressed are of general interest. We describe here modifications of the tet regulatory system that allow differential expression control of two genes in E. coli.
Independent tet regulation of two genes requires different tet operators and TetR variants with unique recognition specificity for them. Two such pairs have been described previously (4, 5). For differential induction, TetR variants with exclusive specificity for distinct inducers are needed. We have recently constructed a TetR variant that is inducible by 4-dedimethylamino-anhydrotetracycline (4-DDMA-ATC) (Fig. 1) but not inducible by tetracycline (TET). This would allow independent induction of two genes. Finally, since native TetR is a homodimer, cross talk between the TetR variants needs to be avoided, which can be accomplished by using tetR alleles from different naturally occurring sequence variants, as has been described previously (8, 11). Here, we ask whether the mutations leading to these functionally different TetR proteins can be combined to yield specifically addressable TetR variants and if their phenotypes are stringent enough for distinct regulation in E. coli. In addition to the E. coli lacZ, we used xynB from Bacillus subtilis encoding a β-xylosidase (β-Xyl) as a second reporter gene to quantitatively distinguish regulatory efficiencies.
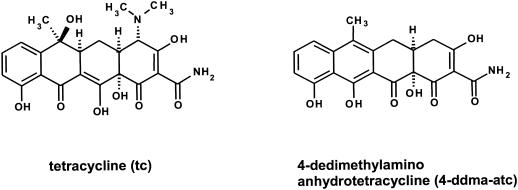
FIG. 1.
Chemical structures and designations of tetracyclines used in this study. The chemical structures of tetracycline and the antibiotically inactive 4-ddma-atc are shown.
Construction of the dual regulation system.
Strains and plasmids used in this study are presented in Table 1. Both reporter genes were expressed in the E. coli strain WH207λtet50, which carries a chromosomal tetA-lacZ transcriptional fusion (17) in which lacZ is under wild-type tetO control. This construct is regulated by the TetR(BD) mutant H64K L131I S135L, which is inducible with 4-DDMA-ATC but not with TET. The second reporter gene, xynB, is regulated by the tet operator 4C (tetO-4C) mutant, with two palindromic sequence changes from the wild type (4). xynB was amplified by PCR from chromosomal DNA of B. subtilis strain 168, introducing a downstream BsmBI site. tetO-4C was amplified from the plasmid pWH1012-4C (13) containing an upstream BsmBI site and then fused to the xynB-carrying fragment in a subsequent PCR without primers. The resulting product was cloned in two steps into a variant of pWH1411 with the tetR mutant. The resulting plasmid was termed pWH628 (Fig. 2). TetR-4C, the TetR(B) E37A P39Q Y42M mutant that specifically recognizes tetO-4C instead of wild-type tetO (4), was expressed from the plasmid pWH1925-tetR-4C. Due to different origins of replication (Fig. 2), pWH1925-tetR-4C and pWH628 can coexist in WH207λtet50. This plasmid compatibility allows the regulation of xynB with TetR-4C. In addition, TetR-4C is inducible with TET but not with 4-DDMA-ATC, and the class B and BD sequence variants of TetR do not heterodimerize (11).
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Relevant characteristic(s) | Reference or source |
|---|---|---|
| Strains | ||
| E. coli DH5α | recA1 endA1 gyrA96 thi relA1 hsdR17(rK− mK+) supE44 φ80dlacZΔM15 ΔlacU169 | 3 |
| B. subtilis 168 | trpC2 | Laboratory stock |
| E. coli WH207 | galK rpsL thi ΔlacX74 recA13 | 17 |
| E. coli WH207λtet50 | Tn10 tetA-lacZ transcriptional fusion, bet+gam+cIII+cI+lacZ+ | 15 |
| Plasmids | ||
| pWH1925-tetR-4C | Apr, pMB1 tetR-4C | This study |
| pWH1925-Δ | pWH1925-tetR-4C without tetR gene | 12a |
| pWH1411-i1.9 | CmrtetR-H64K L131I S135L | 12 |
| pWH628 | Cmr, pWH1411-i1.9 with tetO-4C-xynB region | This study |
| pWH627 | pWH628 without tetR gene | This study |
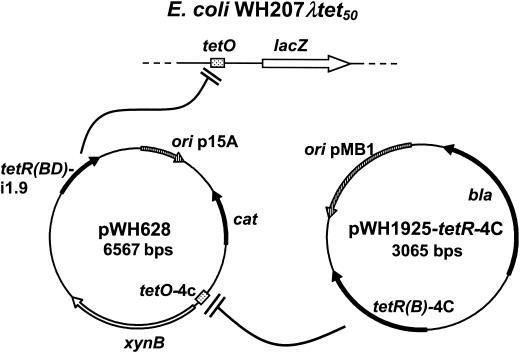
FIG. 2.
Components of the dual regulation system. The host strain is WH207λtet50. tetR(BD)-i1.9 [encoding the 4-DDMA-ATC-specific mutant TetR(BD) H64K L131I S135L] is constitutively expressed from plasmid pWH628 and binds tetO located upstream of lacZ on the chromosome. XynB, expressed from pWH628, is regulated via tetO-4C that is bound by TetR(B)-4C. TetR(B)-4C is constitutively expressed from pWH1925-tetR-4C. ori codes for origin of replication, cat codes for chloramphenicol acetyltransferase, bla codes for β-lactamase, xynB codes for β-Xyl, and lacZ codes for β-Gal.
In vivo evaluation of enzyme activities.
β-Galactosidase (β-Gal) activity was determined as described previously (10) by using ortho-nitrophenyl-β-d-galactopyranoside as a substrate. β-Xyl activity was determined in the same way except with para-nitrophenyl-xylanopyranoside as a substrate. We tested for potential cross-reaction of the enzymes by adding ortho-nitrophenyl-β-d-galactopyranoside to a culture expressing only β-Xyl. No absorption was detected at 420 nm, and the same result was obtained with β-Gal and para-nitrophenyl-xylanopyranoside (data not shown). Measurements of dual regulation were performed in parallel, where one part of the culture was assayed for β-Gal and the other for β-Xyl activity. Both effector concentrations were adjusted to achieve optimal induction within the tolerance range of E. coli (0.2 μM for TET and 1.6 μM for 4-DDMA-ATC) (data not shown). The regulation of β-Gal expression is shown in Fig. 3A. Independent of the presence or absence of any inducer, TetR-4C does not repress β-Gal expression and therefore does not recognize tetO, as is shown on the left side of Fig. 3A. The right side of Fig. 3A shows that in the absence of inducer, the 4-DDMA-ATC-specific TetR binds to tetO, as indicated by the tight repression of lacZ, while TET does not induce this TetR variant at all. In contrast, lacZ is highly expressed when 4-DDMA-ATC is added, indicating that this TetR variant is inducible only with 4-DDMA-ATC and not with TET. β-Gal activity remains high in the presence of both inducers, suggesting that TET does not interfere with the induction of TetR H64K L131I S135L by 4-DDMA-ATC. Figure 3B shows the results of β-Xyl activity determinations of the same cultures. TetR H64K L131I S135L does not bind to tetO-4C (Fig. 3B, left). No XynB expression was detectable in the absence of an inducer, indicating that TetR-4C binds tightly to its cognate sequence tetO-4C. Upon addition of TET, β-Xyl was induced to 65% activity. 4-DDMA-ATC does not induce TetR-4C, as shown by the repression of xynB, and it does not interfere with induction by TET (Fig. 3B, right).
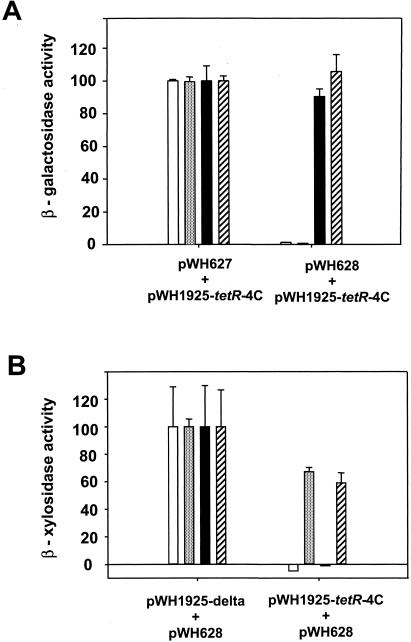
FIG. 3.
Induction analysis of the dual regulation system. (A) Induction of β-Gal expression is shown in the presence of the respective inducers. β-Gal activity in the absence of tetR(BD)-i1.9 (pWH627) was set to 100% and represents about 2,500 Miller units. Open columns, no inducer; gray columns, 0.2 μM TET; black columns, 1.2 μM 4-DDMA-ATC; striped columns, 0.2 μM TET plus 2 μM 4-DDMA-ATC. (B) Induction of XynB expression in the presence of the inducers. XynB activity obtained in the absence of tetR(B)-4C (pWH1925-Δ) was set to 100% and represents about 10 Miller units.
Conclusions.
We present for the first time a dual regulation system that combines different TetR inducer specificities with different TetR operator recognition mutations for application in prokaryotes, allowing independent and reversible in vivo regulation of two different genes by two distinct effectors. Apparently, 4-DDMA-ATC resembles TET as a well-permeating effector showing fast and efficient induction in E. coli. In addition, this effector exhibits no antibiotic activity and may thus overcome limitations in efficiency of expression that are sometimes observed with induction by TET in sensitive strains. The use of these two independently functioning repressors should be a powerful tool for the analysis of prokaryotic phenotypes that depend on the expression of two genes. It should also have merits in target validation or inhibitor screening assays for heterooligomeric proteins.
Acknowledgments
This work was supported by the Deutsche Forschungsgemeinschaft through SFB 473 and Graduiertenkolleg 805 and the Fonds der Chemischen Industrie.
We thank Peter Gmeiner and Susanne Lochner for kindly providing 4-DDMA-ATC. We thank Ralph Bertram and Christian Berens for helpful discussions and for critically reading the manuscript.
REFERENCES
- 1.Geissendörfer, M., and W. Hillen. 1990. Regulated expression of heterologous genes in Bacillus subtilis using the Tn10 encoded tet regulatory elements. Appl. Microbiol. Biotechnol. 33:657-663. [DOI] [PubMed] [Google Scholar]
- 2.Guzman, L. M., D. Belin, M. J. Carson, and J. Beckwith. 1995. Tight regulation, modulation, and high-level expression by vectors containing the arabinose PBAD promoter. J. Bacteriol. 177:4121-4130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hanahan, D. 1983. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166:557-580. [DOI] [PubMed] [Google Scholar]
- 4.Helbl, V., and W. Hillen. 1998. Stepwise selection of TetR variants recognizing tet operator 4C with high affinity and specificity. J. Mol. Biol. 276:313-318. [DOI] [PubMed] [Google Scholar]
- 5.Helbl, V., B. Tiebel, and W. Hillen. 1998. Stepwise selection of TetR variants recognizing tet operator 6C with high affinity and specificity. J. Mol. Biol. 276:319-324. [DOI] [PubMed] [Google Scholar]
- 6.Kallio, P. T., and J. E. Bailey. 1996. Intracellular expression of Vitreoscilla hemoglobin (VHb) enhances total protein secretion and improves the production of alpha-amylase and neutral protease in Bacillus subtilis. Biotechnol. Prog. 12:31-39. [DOI] [PubMed] [Google Scholar]
- 7.Kim, L., A. Mogk, and W. Schumann. 1996. A xylose-inducible Bacillus subtilis integration vector and its application. Gene 181:71-76. [DOI] [PubMed] [Google Scholar]
- 8.Levy, S. B., L. M. McMurry, T. M. Barbosa, V. Burdett, P. Courvalin, W. Hillen, M. C. Roberts, J. I. Rood, and D. E. Taylor. 1999. Nomenclature for new tetracycline resistance determinants. Antimicrob. Agents Chemother. 43:1523-1524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lutz, R., and H. Bujard. 1997. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25:1203-1210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Miller, J. 1972. Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 11.Schnappinger, D., P. Schubert, K. Pfleiderer, and W. Hillen. 1998. Determinants of protein-protein recognition by four helix bundles: changing the dimerization specificity of Tet repressor. EMBO J. 17:535-543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Scholz, O., M. Köstner, M. Reich, S. Gastiger, and W. Hillen. 2003. Teaching TetR to recognize a new inducer. J. Mol. Biol. 329:217-227. [DOI] [PubMed] [Google Scholar]
- 12a.Scholz, O., et al. Activity reversal of Tet repressor caused by single amino acid changes. Mol. Microbiol., in press. [DOI] [PubMed]
- 13.Sizemore, C., A. Wissmann, U. Gülland, and W. Hillen. 1990. Quantitative analysis of Tn10 Tet repressor binding to a complete set of tet operator mutants. Nucleic Acids Res. 18:2875-2880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Skerra, A. 1994. Use of the tetracycline promoter for the tightly regulated production of a murine antibody fragment in Escherichia coli. Gene 151:131-135. [DOI] [PubMed] [Google Scholar]
- 15.Smith, L. D., and K. P. Bertrand. 1988. Mutations in the Tn10 tet repressor that interfere with induction. Location of the tetracycline-binding domain. J. Mol. Biol. 203:949-959. [DOI] [PubMed] [Google Scholar]
- 16.Stieger, M., B. Wohlgensinger, M. Kamber, R. Lutz, and W. Keck. 1999. Integrational plasmids for the tetracycline-regulated expression of genes in Streptococcus pneumoniae. Gene 226:243-251. [DOI] [PubMed] [Google Scholar]
- 17.Wissmann, A., L. V. Wray, Jr., U. Somaggio, R. Baumeister, M. Geissendörfer, and W. Hillen. 1991. Selection for Tn10 tet repressor binding to tet operator in Escherichia coli: isolation of temperature-sensitive mutants and combinatorial mutagenesis in the DNA binding motif. Genetics 128:225-232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhang, L., F. Fan, L. M. Palmer, M. A. Lonetto, C. Petit, L. L. Voelker, A. St. John, B. Bankosky, M. Rosenberg, and D. McDevitt. 2000. Regulated gene expression in Staphylococcus aureus for identifying conditional lethal phenotypes and antibiotic mode of action. Gene 255:297-305. [DOI] [PubMed] [Google Scholar]