Figure 1.
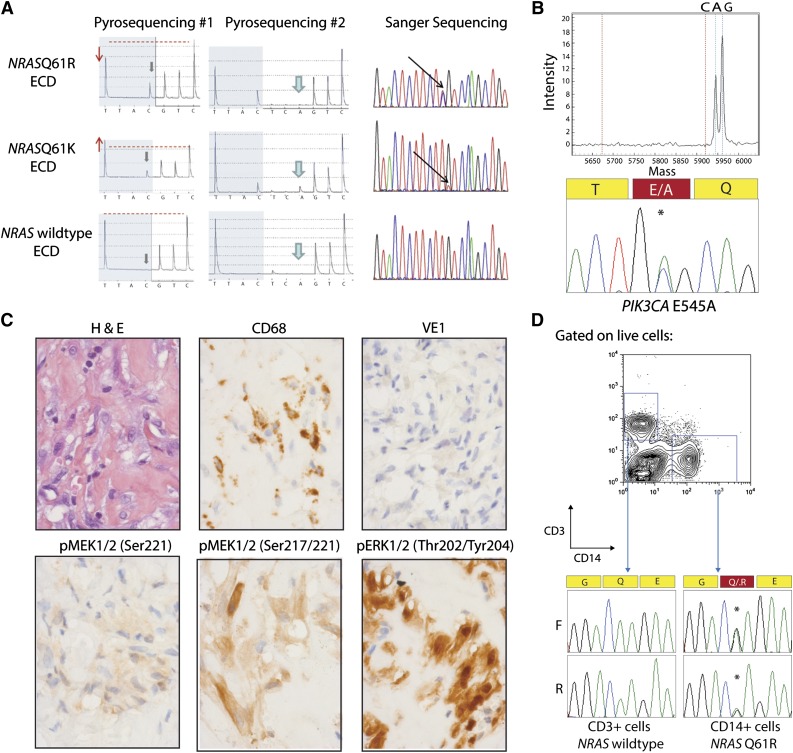
NRAS and PIK3CA mutations in ECD histiocytes and CD14+ cells from peripheral blood. (A) Detection of NRAS p.Q61 mutation by pyrosequencing (first 2 columns) and confirmation by Sanger sequencing (third column). The lower pyrogram corresponds to an NRAS–wild-type ECD case, and the 2 others correspond to patients with NRAS mutations. Mutants are detectable with the appearance of a new peak at the first C injection (gray arrows), and size variation of the peak at the first T injection compared with the second C injection (red arrows and dashed line). Pyrosequencing with another sequence of injection (ESGTTACTCAGTCAGCT) was used to further identify the mutations (middle column) as c.181C>A mutations. (B) Detection of PIK3CA E545A mutation by Sequenom and Sanger sequencing in an NRAS/KRAS/BRAF–wild-type ECD patient. (C) Phosphorylated MEK (pMEK) and ERK (pERK) detected by immunohistochemistry in BRAFV600E–wild-type, NRAS-mutant ECD. Histiocytes (noted by hematoxylin and eosin and CD68 stain) failed to stain for BRAFV600E VE1 monoclonal antibody but had high cytoplasmic expression of pMEK1/2 with 2 different antibodies as well as cytoplasmic and nuclear expression of pERK1/2 (original magnification ×600). (D) Genotyping of CD14+ monocytes and CD3+ T cells purified from the peripheral blood of an NRASQ61R-mutant ECD patient with double-FACS sorting reveals the presence of NRAS mutation in CD14+ cells but not in T cells.