DNA caretaker genes are defective in several human syndromes known as “chromosomal breakage” or “chromosomal fragility” disorders, which include autosomal recessive diseases such as Fanconi anemia (FA), ataxia telangiectasia (AT), and Nijmegen breakage syndrome (NBS), among others. Dominantly inherited high risk for breast and ovarian cancer is associated with mutations in the BRCA1/BRCA2 genes, whose protein products are also known to be crucial for chromosome stability.
Many of the insights that we have gained into the mechanisms involved in cellular DNA-damage response pathways have come from studies of human cancer susceptibility syndromes that are altered in DNA-damage responses. Basic research on these syndromes is therefore of interest to understand key biological processes that have evolved to maintain a stable genome and to prevent cancer. Research over the last three years has led to the conviction that the pathways involved in many chromosome fragility syndromes converge in a common tumor suppression nuclear network of interactions (Fig. 1). To further characterize how the various proteins involved in these syndromes are interconnected, an International Workshop was organized in the Juan March Centre for International Meetings on Biology (Madrid, Spain, February 2–4, 2004). In this article we summarize the main results presented in this workshop.
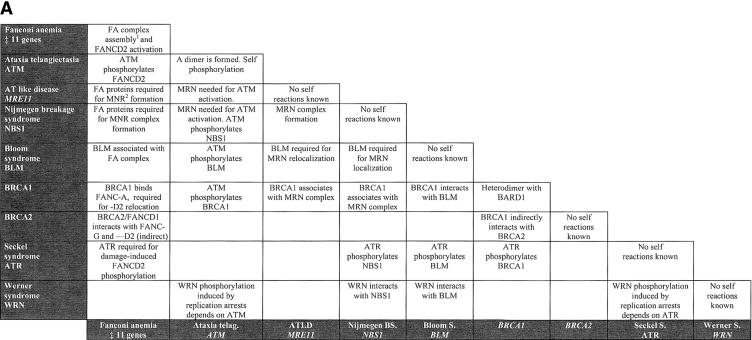
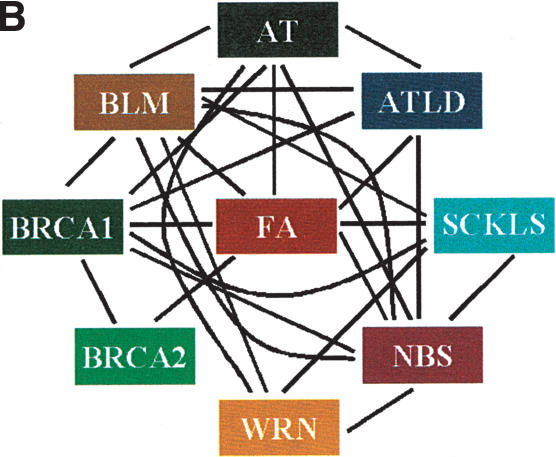
Figure 1.
Molecular interactions among chromosome fragility syndromes. (A) Each square indicates the molecular nature of the existing interaction between two given syndromes. (B) Diagram representation of the molecular cross-talk. Each line connects syndromes with known interaction at the molecular level as detailed in A.
The first session was fully dedicated to FA, which is characterized at the cellular level by chromatid-type breakage of metaphase chromosomes, in particular after exposure to cross-linking agents such as mitomycin C (MMC). Hans Joenje (Amsterdam, The Netherlands) presented a comprehensive up-to-date summary of the latest discoveries relating to genetics and molecular biology of this rare disease. Evidence from somatic cell hybridization, positional cloning, and protein association studies collectively indicate that there are at least 11 FA complementation groups (A, B, C, D1, D2, E, F, G, I, J, L), each connected with a distinct disease gene (Levitus et al. 2004). Protein coimmunoprecipitation studies have implicated four proteins to be associated with a nuclear multiprotein core complex that contains most of the other FA proteins. One of these (FANCL) was recently shown to be mutated in an unclassified FA patient (Meetei et al. 2003a). This protein has E3 ubiquitin ligase activity and is thought to represent the activity responsible for the monoubiquitination of FANCD2 at K561 (Meetei et al. 2004), a key posttranslational modification in the FA pathway (Garcia-Higuera et al. 2001). All FA proteins, except FANCD1/BRCA2 and FANCJ, which function downstream, are essential for this reaction to occur. Apart from FANCL, the domain structure of the other proteins in the nuclear complex is largely unknown and their precise molecular functions are obscure. FANCG/XRCC9 has recently been found to belong to the family of tetratricopeptide repeat proteins, suggesting that it may play an important role in the assembly and/or stabilization of the FA protein core complex (Blom et al. 2004). The putative product of the FANCI gene functions at the level of the core complex, or downstream, and seems to be required for binding of FANCD2 to chromatin. Activated FANCD2 is thought to function at the site of DNA damage, presumably in concert with BRCA1 and FANCD1/BRCA2. BRCA2 probably acts downstream or separately from the FA pathway, consistent with the distinct syndromic association observed in FA-D1 patients (see below). Joenje pointed out that BRCA2 has been considered the disease gene in both FA-D1 and FA-B patients (Howlett et al. 2002), but evidence on RAD51 foci formation suggests that the putative FANCB gene is distinct from BRCA2 (Godthelp et al. 2002).
Joenje also discussed the evidence that a disruption of the FA pathway may lead to some forms of sporadic cancer. An example is the reported silencing of FANCF by a promoter hypermethylation found in a proportion of sporadic tumors of different types (ovarian, oral, and lung tumors and acute myeloid leukaemia; AML), which may have important implications for the prognosis and treatment of patients having such tumors (Taniguchi et al. 2003; Tischkowitch et al. 2003; Marsit et al. 2004). Joenje commented on a recent report in which no pathogenic mutations could be found in six FA genes (FANCA, FANCC, FANCD2, FANCE, FANCF, and FANCG) in a population of familial breast cancer patients in whom no mutations in BRCA1 or BRCA2 had been found (Seal et al. 2003). This study suggests that, in spite of extensive molecular cross-talk between the FA pathway and BRCA1 and BRCA2, at least the FA genes tested are not likely to be tumor suppressor genes in familial breast cancer. However, the study would need to be extended to include the remaining FA genes as well. Moreover, silencing of FA genes (as found for FANCF) by promoter hypermethylation rather than nucleotide sequence alterations, will need to be addressed in the future.
Given the apparent implications of FANCF silencing in sporadic cancer, further functional studies of FANCF are of particular interest. In this context, Johan de Winter (Amsterdam, The Netherlands) presented an extensive study based on site-directed mutagenesis of those residues of FANCF that are highly conserved between humans and Xenopus laevis. He showed how a C-terminal truncated FANCF can still complement FANCF-deficient cells, although this part of the protein is important for binding FANCA and FANCG in a stable core complex. He then reported that the N terminus of FANCF stabilizes the interaction between FANCA and FANCG and is essential for binding FANCC and FANCE. De Winter also discussed that this part of the protein tolerates many amino acid changes, indicating that FANCF is a flexible protein. This is consistent with the lack of missense mutations in FA-F patients. The main conclusion from his work is that FANCF plays a pivotal role in the interaction of the other FA proteins to forms a stable and functional FA core complex.
The next speaker (K.J. Patel, Cambridge, UK) reported his recent studies on the molecular biology of the FA pathway using reverse genetics in chicken DT40 cell system. Patel first focused on the genetic consequences of FANCC ablation in the DT40 cells. He reported that FANCC-deficient cells have a defect in gene conversion and gene targeting and that these cells have decreased frequencies of DNA cross-link-induced sister chromatid exchanges (SCE), resembling RAD51-deficient cells. He also reported that FANCC and the RAD51 paralog XRCC2 function in the same epistatic group to resolve DNA replication blocking lesions, as cell lines disrupted for both proteins are equally sensitive to DNA cross-linking agents. Another key finding reported by Patel is the observation in this genetic model that FANCC and the translesion synthesis (TLS) polymerases Rev1 and Rev3 function in the same pathway. To reach this conclusion, he generated double KOs and demonstrated that these genes are epistatic. In addition, Rev1 and FANCD2 colocalize at hydroxyurea- and thymidine-induced stalled replication forks in HeLa cells. In conclusion, FANCC is required for homologous recombination (HR) in concert with XRCC2 and in an error-prone pathway together with Rev1 and Rev3, and the intersection between these two pathways takes place on replication forks to resolve DNA cross-links.
Alan D'Andrea (Boston, MA) began by explaining his view on important questions yet to be resolved in the FA field: the sensor of the FA complex, the molecular function of monoubiquitinated FANCD2 (FANCD2-Ub) in chromatin, and the enzyme that deubiquitinates FANCD2-Ub. According to D'Andrea, ATR (AT-Rad3-related) could be the sensor of the FA/BRCA pathway, in agreement with the reported interactions between ATR and the FA/BRCA pathway (Tibbetts et al. 2000; Pichierri and Rosselli 2004). D'Andrea then reported that FANCD2-Ub recruits BRCA2 into a chromatin complex and that FANCD2-deficient cells are defective in DNA-damage-inducible BRCA2 foci and in chromatin loading of BRCA2 following DNA damage. Functional complementation with the FANCD2 cDNA restores BRCA2 foci and chromatin loading. BRCA2 mutant cells, expressing a C-terminal truncated BRCA2 protein, form ionizing radiation (IR)-inducible BRCA2 and FANCD2 foci that fail to colocalize, and the FANCD2 and truncated BRCA2 proteins fail to coimmunoprecipitate. Functional complementation of these cells with BRCA2 restores the interaction of BRCA2 and FANCD2. The C terminus of BRCA2 is therefore required for the functional interaction of BRCA2 and FANCD2 in chromatin. These results suggest that the FA/BRCA pathway and monoubiquitinated FANCD2 function to load BRCA2 onto chromatin complexes, a process required for normal homology-directed DNA repair.
Finally, D'Andrea described his recent attempts to find inhibitors and enhancers of the FA pathway, using a technology that allows the rapid screening of >200,000 molecules. Among the six molecules that he found to interact with the FA pathway, he described wortmannin, a drug known to inhibit the kinase activity of ATM, ATR, and DNA-PK, as an inhibitor of FANCD2 monoubiquitination.
Given the key role of FANCD2 in the FA pathway and its cross-talk with other chromosome fragility syndromes, Jordi Surrallés (Barcelona, Spain) focused his lecture on the molecular biology of this protein. He first noted that together with FANCL and FANCD1/BRCA2, FANCD2 is the only FA proteins conserved in nonvertebrates and, therefore, these proteins may represent the ancestor of the FA pathway. He then presented the results of the molecular cloning and characterization of FANCD2 in Drosophila melanogaster. Interestingly enough, both the monoubiquitination site and the ATM-dependent phosphorylation site are highly conserved from humans to flies, suggesting functional conservation of this pathway (Castillo et al. 2003). In support of this view, he then described the expression pattern of FANCD2 in Drosophila at the protein and RNA level. A high expression was detected in the abdominal part of adult flies, especially in males where the two FANCD2 isoforms are most evident. These results suggest a major role of dmFANCD2 in spermatogenesis, resembling FANCD2 function in mammals (Houghtaling et al. 2003).
Surrallés then focused on an unexpected role of FANCD2 in human cells. He reported a highly regulated activation of the FA/BRCA pathway in response to ultraviolet radiation in the G1 phase of the cell cycle. He showed that FANCD2 relocates to the site of damage in locally UV-irradiated nuclei in a FANCA, BRCA1, and FANCD2 K561-dependent manner, but independently of BRCA2, ATM, XPA, and RNA Pol II-mediated transcription. He suggested that FANCD2-Ub relocates to UV-induced DSBs (as detected by phosphorylated histone H2AX immunohistochemistry). This role for the FA/BRCA pathway in G1 is independent of BRCA2/RAD51-mediated HR, as these proteins do not relocate together with FANCD2 at the site of damage. One main conclusion from his work is that there are two types of FANCD2 aggregates: those foci connected to HR during S phase (D'Andrea 2003) and those induced in the G1 phase independent of HR. Surrallés also reported that oxidative damage (8-OxoG) is removed by base excision repair in FANCD2-deficient cells, suggesting that the well-known accumulation of 8-OxoG in FA patients (for review, see Pagano and Youssoufian 2003), is not caused by a deficiency in the removal of this lesion. In the last part of his talk, Surrallés presented a Spanish FA patient who died of Wilms tumor at the age of 1 yr and who had a family history of cancer in various tissues including a bilateral breast cancer. In this patient, all upstream FA genes were functional, based on FANCD2-Ub and complementation studies by retroviral transduction; BRCA2 and BRCA1 were not mutated; and the clinical phenotype was not consistent with a FANCJ mutation. Given the Wilms tumor (see below) and the lack of BRCA2 mutation, the gene mutated in this family could be an as-yet-unidentified tumor suppressor with a functional relationship to BRCA2.
The last speaker in this first session was Arleen Auerbach (New York, NY). She began by summarizing the main outcome of the International Fanconi Anemia Register (IFAR), with 906 FA patients currently included. Of these, 468 patients have been successfully subtyped and the pathogenic mutations have been found in many of them allowing extensive correlations between genotype and cancer and other clinical phenotypes (Kutler et al. 2003). Because of the extensive molecular and clinical overlap among chromosome fragility syndromes, Auerbach pointed out that after ruling out FA complementation groups FA-A, FA-C, FA-D2, FA-E, FA-F, and FA-G, patients are tested for mutations in NBS1 and BRCA2, and in some cases for Bloom syndrome (BLM) by the SCE assay. She then focused on these rare, latter cases with an FA-like phenotype but where most of the FA genes were ruled out. The range of congenital malformations in some BRCA2, NBS, and BLM patients was similar to those seen in the classical FA patients. Of special interest were three BLM patients with a clinical phenotype identical to FA but properly diagnosed as BLM only on the basis of high SCE. In addition, the 14 FA patients identified with biallelic BRCA2 mutations are characterized as having early onset of AML and solid tumors, especially brain and Wilms tumors (Offit et al. 2003). Interestingly, brain and Wilms tumors were not found in any other IFAR patients in which the complementation group was known, and leukemia occurred at a median age of 2.2 yr in BRCA2 patients in contrast to the 13.4 yr in all other IFAR patients. In addition, an increased incidence of early-onset breast cancer was noted in several BRCA2 patients but not in other IFAR patients. The main conclusion from Auerbach's work is that the cancer spectrum in FANCD1/BRCA2 patients is unique, suggesting that the pathogenesis of cancer in these patients may be via a different pathway from the other FA patients. Thus, FANCD1/BRCA2 probably acts separately from the main FA pathway, suggesting that FANCD1/BRCA2 should actually be classified as “FA-like patients.”
The phenotypic overlap between some FA, NBS, and BLM patients is consistent with a tight molecular cross-talk among the proteins involved in these syndromes (see Fig. 1). Thus, it is known that FANCD2-Ub interacts with NBS1 (Nakanishi et al. 2002) and that the BLM helicase is a component of the FA core complex (Meetei et al. 2003b). The same is true with FA, SCKLS, and ATM (see above and Fig. 1). In line with these observations, the next session was focused on AT, AT-Like disorder (ATLD), ATR, and on recent insights on DNA-damage-signaling pathways and the cellular and organismal phenotypes associated with various mutations in gene products that participate in these pathways.
AT is characterized by multiple physiological abnormalities, including neurodegeneration, immunological abnormalities, cancer predisposition, sterility, and metabolic abnormalities. The gene mutated in this disorder, ATM, is a protein kinase that is activated by the introduction of DNA DSBs in cells. Clinical disorders that have some similarities to AT include NBS, ATLD, and SCKLS, resulting from mutations in NBS1, MRE11, and ATR, respectively. Studies using cells from all of these syndromes have been invaluable in elucidating the molecular steps involved in stress-signaling pathways.
Michael Kastan (Memphis, TN) discussed the signals that go to and from ATM after DNA damage. ATM activity is required for cell cycle arrest induced by IR in G1, S, and G2 phases of the cell cycle. Several targets of the ATM kinase have been identified that participate in IR-induced cell cycle arrests (Fig. 1). For example, phosphorylation of p53, mdm2, and Chk2 participates in the G1 checkpoint; NBS1, BRCA1, FANCD2, and SMC1 participate in the transient IR-induced S-phase arrest; and BRCA1 and hRad17 have been implicated in the G2/M checkpoint. Although ATM is critical for cellular responses to IR, related kinases, like ATR, appear to be important for responses to other cellular stresses. Some substrates appear to be shared by the two kinases, with the major difference being which stimulus is present and which kinase is used to initiate the signaling pathway. Recent studies also elucidated the mechanism by which DNA damage activates the ATM kinase (Bakkenist and Kastan 2003). ATM normally exists in the cell as an inactive homodimer bound to nuclear chromatin in unperturbed cells, and the introduction of DNA damage induces intermolecular autophosphorylation on Ser 1981 in both ATM molecules. This phosphorylation causes a dissociation of the ATM molecules and frees them up to now circulate around the cell and phosphorylate the substrates that regulate cell cycle progression and DNA repair processes. This regulation of ATM activity in the cell represents a novel mechanism of protein kinase regulation and appears to result from alterations in higher-order chromatin structure rather than direct binding of ATM to DNA double-strand breaks (DSBs). SMC1 is the only ATM substrate identified whose phosphorylation has an impact on radiosensitivity (Kim et al. 2002). Phosphorylation of SMC1 by ATM was found to be dependent on the presence of both NBS1 and BRCA1 proteins because both proteins appear to recruit activated ATM to the DNA strand break where it can phosphorylate SMC1. Kastan suggested that the phosphorylation of SMC1 is a critical downstream target of this signaling pathway for determining cell survival and chromosomal repair.
Yosef Shiloh (Tel Aviv, Israel) discussed overlaps and distinctions among the genomic instability syndromes, AT, NBS, and ATLD. Interestingly, whereas cells from each of these disorders have very similar abnormalities in culture, there are distinctive clinical differences. One of the major differences is the fact that NBS patients are microcephalic, whereas AT and ATLD patients exhibit cerebellar neurodegeneration. Shiloh's lab recently suggested that the MRE11/RAD50/NBS1 (MRN) protein complex functions both “upstream” of and “downstream” from the ATM kinase (Uziel et al. 2003). It was already known that NBS1 was a substrate of the ATM kinase, but they also found abnormalities in the activation of the ATM kinase in cells lacking any of the three MRN proteins. He also suggested that the neurodegeneration seen in AT and ATLD resulted from DNA-damage response abnormalities.
Christian Carson, from Matthew Weitzman's laboratory (La Jolla, CA), discussed their recent results using an adenovirus system to probe cellular responses to DNA damage. Starting with the interesting question of whether viral genomes are recognized by the DNA-damage machinery, they found that adenoviruses lacking the E4 genetic region formed concatemers during infection in normal cells, but not in cells lacking DNA-PK and ligase IV, or in cells harboring hypomorphic mutations in NBS1 or MRE11. Furthermore, this concatemerization coincided with signaling from the ATM and ATR kinases. They also observed the recruitment of MRE11 and activated ATM to replication centers of E4-deleted adenovirus. Unexpectedly, they found that wild-type adenovirus did not induce a DNA-damage response and observed that MRE11 was actually degraded following the infection. This degradation appeared to depend on the adenoviral E1b55k and E4orf6 proteins. Interestingly, the degradation of MRE11 appeared to be associated with an inhibition of ATM activation by DNA damage. In response to infection with E4-deleted adenovirus, ATM activation was greatly reduced in A-TLD cells, but was normal in NBS cells. Thus, there appeared to be some distinctive functions of MRE11 and NBS1 (Stracker et al. 2002; Carson et al. 2003).
Studies of ATM activation in cells lacking normal MRE11 function were also discussed by Malcolm Taylor (Birmingham, UK). Careful characterization of AT patients led to the identification of a group of families carrying a leaky mutation such that they produced some normal ATM protein, with homozygotes having ∼11% of normal ATM levels and compound heterozygotes (carrying two different ATM mutations) with ∼4% of normal activity. The patients with 11% of normal activity had significantly slower progression of their cerebellar degeneration and no predisposition to tumors. Although the neurodegeneration was still severe in the patients with 4% of normal ATM activity, they also did not have significant predisposition to lymphomas. Thus, this low amount of ATM activity was sufficient to reduce cancer risk. Similarly, studies of ATLD patients with various amounts of MRE11 deficiencies were evaluated. Similar to the conclusions from the Kastan laboratory, but conflicting with an absolute requirement for MRN complex for ATM activation suggested by the Shiloh and Carson/Weitzman labs, Taylor's studies demonstrated that ATM is activated in both NBS and ATLD cells. Discussions of these differences by the audience led to general agreement that there are differences between measuring ATM activation and measuring ATM activity, the latter requiring the ability of the activated ATM to find and phosphorylate its substrates in the cell. This concept was suggested by Kastan's data showing that NBS1 and BRCA1 are required to recruit the activated ATM to the sites of DNA breaks. Similar to the suggestion made by the Carson/Weitzman lab, Taylor's data also suggested that there may be distinct functions of MRE11 and NBS1.
Penny Jeggo (Sussex, UK) showed data suggesting that ATM and DNA-PK both participate in the phosphorylation of histone H2AX after DNA damage and that the absence of just one of these kinases in a cell has only a minor quantitative effect on H2AX phosphorylation. She then went on to relate an intriguing story about SCKLS, a disorder characterized by growth retardation, microcephaly, and mental retardation among other abnormalities as discussed before. Cells from these patients have features that overlap with those seen in FA, NBS, and BLM, and Jeggo's lab found that some of these families carry germ-line splicing mutations in the ATR gene (O'Driscoll et al. 2003). In particular, these cells had abnormal initiation of DNA-damage-signaling pathways in response to UV irradiation, although responses to IR appeared relatively normal. Although all of the cell lines from Seckel patients exhibited these abnormal UV responses, most did not have mutations in ATR. This observation suggests that these other families may carry germ-line mutations in other genes in the ATR signaling pathway.
John Petrini (New York, NY) described mice generated in his lab that carry mutations in either MRE11 or RAD50 (Petrini and Theunissen 2004). Surprisingly, a mutation in RAD50 (RAD50S) suppressed lymphomagenesis in ATM-deficient mice, and Petrini suggested that this appears to reflect an RAD50S-dependent compensatory activity of ATR. Conversely, ATM heterozygosity rescued the decreased survival of RAD50S mice. Further elucidation of the mechanisms underlying these unexpected effects may provide significant insights into the roles of these damage-responsive proteins. Mutations in MRE11 that mimic the human ATLD mutations were also made, and, interestingly, there was no apparent cancer predisposition in these mice. Cells from these mice exhibited normal G1 checkpoints (suggesting normal ATM activation in these MRE11 mutants), defective intra-S-phase and G2/M checkpoints, and increased radiation-induced chromosomal aberrations. These results led Petrini to suggest that chromosome breakage in S phase may enhance the penetrance of an initiating lesion with respect to lymphomagenesis, but is insufficient as an initiator of lymphomas itself.
The next session was dedicated to the mechanisms of HR and how they may be controlled by BRCA2. The enzymes of HR can re-establish broken replication forks and promote the repair of DSB using a sister chromatid as the template for faithful repair (Cox et al. 2000; Johnson and Jasin 2000; West 2003). Our understanding of the mechanism of recombinational repair, and its importance for genome integrity, has recently been advanced by significant developments in several areas of research. Firstly, biochemical and structural studies are showing us how key recombination proteins manipulate DNA and catalyze the molecular gymnastics that are necessary for DNA pairing, strand exchange, and finally the resolution of repaired molecules. Secondly, molecular analysis of DSB repair in vivo has demonstrated the importance of HR as a DSB repair mechanism and the outcome of repair in several circumstances. Thirdly, we are beginning to understand that recombination proteins are tightly controlled and relocalize to sites of DNA damage as and when required. Finally, the discovery that cancer susceptibility genes such as BRCA1 and BRCA2 are required for normal levels of recombinational repair demonstrates the connection between repair efficiency, the ability to maintain genome stability, and the potential for tumorigenesis.
A key player in the recombination process is RAD51, which mediates the homologous pairing and DNA strand exchange reactions leading to recombination between interacting DNA molecules. RAD51 activity is controlled by BRCA2, a large (384 kDa) tumor-suppressor protein. Both proteins localize to distinct nuclear foci upon treatment with IR, and it is at these sites that the repair reactions essential for genome stability are thought to occur. However, our understanding of the molecular reactions that take place within these foci remains limited (West 2003).
Ashok Venkitaraman (Cambridge, UK) described how BRCA2 interacts with RAD51. Previously, it was shown that interactions between BRCA2 and RAD51 take place at a series of BRC repeats present within BRCA2 exon 11 (Bignell et al. 1997; Wong et al. 1997). Although these sites are unlikely to be equivalent to each other, BRCA2 appears to be capable of binding multiple RAD51 molecules in an inactive state, and yet is also required for the accumulation of RAD51 to repair foci where it actively promotes repair. Venkitaraman discussed how BRCA2 might provide both positive and negative control over the actions of RAD51. He showed the results of structural studies in which the interactions between RAD51 and a BRC repeat were analyzed using a fusion protein containing the nucleotide-binding core of RAD51 linked to BRC4 (Pellegrini et al. 2002). Venkitaraman described how the BRC4 region remained in continuous contact with RAD51 over a stretch of 28 amino acids. If the same interaction occurs in the cell, the binding of RAD51 to BRC4 will impair the ability of one RAD51 monomer to interact with another, so that RAD51 nucleoprotein filament formation will be blocked. Consistent with this observation, the inhibition of RAD51 filament formation by peptides corresponding to BRC3 and BRC4 has been observed previously (Davies et al. 2001). But what this structural study provides is a remarkable demonstration that the BRC4 polypeptide effectively mimics the structure of the interaction domain of two adjacent RAD51 monomers. Although it is likely that studies with a short isolated BRC repeat may be oversimplistic in terms of how BRCA2 functions as a whole, they provide us with some insight into the control mechanism that BRCA2 exerts over RAD51 and will one day give us a clear indication of how multiple units of RAD51 associate with BRCA2. However, our understanding of the control of RAD51 is not helped by the puzzling observation that only a fraction (∼20%) of the RAD51 within the nucleus appears to be bound by BRCA2. New studies of the dynamics of wild-type and mutant GFP–RAD51 fusion proteins in the nucleus of living cells indicates that it is this BRCA2-bound fraction of RAD51 that becomes selectively mobilized after DNA damage (Yu et al. 2003). The nature of the bound and unbound fractions, and potential interacting partners, will clearly be a topic for future study.
Venkitaraman also pointed out that DNA replication intermediates formed in BRCA2tr/tr mouse embryonic fibroblasts (MEFs) treated with hydroxyurea to stall fork progression were processed into DSBs. In cells lacking wild-type BRCA2, he suggested that it is the repair of these breaks by HR that is compromised, and that the broken forks may well be the cause of the gross chromosomal rearrangements associated with BRCA2 defects (Lomonosov et al. 2003). However, if one considers BRCA2-related cancers to be a disease associated with DNA replication, then it is reasonable to think that some breast and ovarian tumors might therefore be good therapeutic targets.
It is known that BRCA2-defective cells, like those of FA complementation group D1, are sensitive to DNA cross-linking agents such as cisplatin and MMC (see above), but the difference in sensitivity compared with normal or BRCA2 heterozygous cells may not be enough to be of therapeutic value. It was therefore very interesting to hear Alan Ashworth (London, UK), who presented new data that his laboratory has generated in collaboration with KuDOS Pharmaceuticals showing that BRCA2-deficient cells are exquisitely sensitive (∼1000-fold compared with control cells) to the inhibition of a second DNA repair pathway involving poly(ADP-ribose) polymerase (PARP). The further development of chemical inhibitors of PARP may therefore have great therapeutic potential against BRCA-associated cancers.
Because cells defective in BRCA1/BRCA2 and those derived from individuals with FA are sensitive to DNA cross-linking agents and are defective in HR, it is important that the recombination repair process is understood fully at the molecular level. Recently, significant progress was made toward understanding how recombination intermediates are processed in mammalian cells, as Steve West (London, UK) described his group's recent results showing that extracts prepared from cell lines carrying mutations in the recombination/repair genes RAD51C and XRCC3 exhibit reduced levels of Holliday junction (HJ) resolvase activity (Liu et al. 2004). Moreover, RAD51C protein was found to be an essential component of a highly purified fraction from human cells that was capable of promoting branch migration and Holliday junction resolution in vitro. It is hoped that the identification of RAD51C and XRCC3 as key components of the HJ resolvasome will now open the door toward understanding the mechanisms of Holliday junction processing in eukaryotic cells.
Details of the mechanism by which one chromatid uses its sister as a template for recombinational repair were the topic of Ralph Scully (Boston, MA). It is well known that HR is used to repair DSBs that arise at stalled replication forks, but the molecular details of these reactions are unclear because the sequences of the two sister chromatids are identical and SCE is generally mutationally silent and error free. Sister chromatid recombination (SCR) in mammalian cells has traditionally been studied by cytological techniques that allow the microscopic visualization of crossover events between sister chromatids. But Scully pointed out that information provided by this technique is limited because it fails to provide a molecular picture of the repair event, nor does it detect recombination events that do not result in crossovers. An alternative has been to study SCR using I-SceI endonuclease-generated DSBs, which has allowed a molecular analysis of repair (Johnson and Jasin 2000). To refine our understanding of the mechanism of SCR events, Scully detailed how he is developing novel recombination reporter systems that will allow the selection of SCR events that involve long gene conversion tracts. These systems are likely to be very useful in the analysis of mutant cell lines such as BRCA1, BRCA2, FA, and BLM, all of which are known to be defective in some aspect of recombinational repair.
The fact that sister chromatids are used for error-free exchange during DSB repair by HR links DSB repair to chromatid and chromosome positioning in the nucleus. Roland Kanaar (Rotterdam, The Netherlands) described how repair foci formed at defined locations are able to maintain their global nuclear position upon cell division. His data showed that global chromosome domain position is heritable, consistent with observations from another laboratory (Gerlich et al. 2003). Using time-lapse imaging of cells undergoing division such that global chromosome location could be monitored directly, Kanaar described how, rather than being directly inherited, the global chromosome neighborhoods were re-established in the G1 phase of the cell cycle. Furthermore, by exposing cells to a source of α-particles that track in a linear path through the nucleus, it was shown that mechanisms exist in G1 that facilitate the movement of DSBs (visualized by immunodetection of phosphorylated histone H2AX) such that they gather at sites where the repair proteins accumulate. The observation that chromosome domains containing DSBs exhibit a mobility that allows them to interact may have important implications in terms of the generation of potentially tumorigenic translocations between broken chromosomes (Aten et al. 2004).
Having well-characterized cell mutants is key for the delineating the role of genes mutated in the chromosome fragility syndromes. BRCA2 mutant cells are difficult to come by. Either their growth is severely compromised, like the CAPAN-1 tumor cell line, or the cells have weak hypomorphic alleles, like some of the targeted mouse cell lines. Margaret Zdzienicka (Leiden, The Netherlands) previously described a Brca2 mutant hamster cell line (Kraakman-van der Zwet et al. 2002) that grows surprisingly well and will certainly be useful for many studies. She has now characterized the mutations in the two Brca2 alleles in the cell line, and as well has created revertant lines in which either one of the alleles becomes functional. Notably, this Brca2 cell line recapitulates the phenotypes of other strong HR mutants, including a high level of spontaneous chromosome aberrations. The revertants are not fully wild type, implying a heterozygous phenotype. A higher spontaneous mutation rate is also found in the Brca2 mutant (Kraakman-van der Zwet et al. 2003). The additional mutations are deletions (14-fold increased), rather than point mutations, which is consistent with the cells being defective in HR.
The inability to repair spontaneously arising DNA damage or the misrepair of such damage is what causes chromosome fragility in these syndromes. Replication inevitably stalls during every cell cycle at sites of DNA damage, but can be resumed with the intervention of appropriate DNA repair mechanisms. HR factors play a key role in this regard, as is well described in Escherichia coli (Cox et al. 2000). Many of the proteins deficient in the chromosome fragility syndromes are HR factors or interact with factors involved in the HR. One of the clearest examples is BRCA2 (Moynahan et al. 2001). An alternative mechanism to re-establish replication is to replicate past DNA lesions as they are encountered, that is, by translesion synthesis, using one of a number of specialized polymerases that have been recently discovered (see above; Friedberg et al. 2002). The fidelity of these polymerases is variable, but in some cases can be quite high. Mutation of one of these polymerases is associated with a variant of Xeroderma pigmentosum, a syndrome involving DNA repair defects but that is not considered to cause chromosomal fragility. Taking advantage of the DT40 chicken cell line, Shunichi Takeda (Kyoto, Japan) reported his group's efforts to characterize the genetic interactions between these two mechanisms for dealing with replication problems in vertebrates. Mutants for genes involved in translesion synthesis, that is, Rev3 or Rad18, exhibit increased SCE, indicating that HR is elevated (Yamashita et al. 2002; Sonoda et al. 2003). When either of these mutations is combined with a mutation in Rad54, a gene involved in HR, cells are not viable, even though both single mutants proliferate with nearly normal kinetics. These results suggest that when either HR or translesion synthesis is compromised, the other pathway can compensate, but that when both pathways are compromised, cells are irreparably damaged.
HR is a major mechanism for repairing DSBs and other lesions that arise from replication and other cellular processes. A second major pathway of DSB repair is NHEJ. Similar to the approach taken by Takeda and colleagues in chicken cells, mutations in HR and NHEJ genes have been combined in the mouse by Maria Jasin (New York, NY) and colleagues. Rad54 mutant mice show no overt phenotype (Essers et al. 1997), whereas mice mutated for the NHEJ gene Ku80 have several phenotypes, including small size and immunodeficiency (Nussenzweig et al. 1996). Rad54/Ku80 double-mutant mice are severely compromised for viability, much more than Ku80 mice, and have synergistic cellular defects (Couëdel et al. 2004). Thus, although usually considered separable pathways, HR and NHEJ may act in some cases on the same lesion.
Thus far, an unambiguous relationship between NHEJ defects and chromosome fragility syndromes has not been determined. However, individuals have been identified by the laboratories of Pat Concannon (Seattle, WA) and Penny Jeggo together with colleague Mark O'Driscoll, who have hypomorphic mutations in the gene for DNA ligase IV, an NHEJ factor (O'Driscoll et al. 2001). These patients present with several of the same phenotypes as NBS patients: microcephaly, developmental delays, radiosensitivity, and immunodeficiency. Although lymphoid cancers have been observed in several of these patients, others have remained cancer free, possibly because, unlike in NBS, cell cycle checkpoints are intact.
At the heart of chromosome fragility syndromes is the molecular events that go awry to give rise to chromosome aberrations. Unrepaired or misrepaired DNA breaks, single-strand but especially DSBs, are the starting point for these aberrations. Maria Jasin (New York, NY) described a system for examining the molecular events giving rise to reciprocal chromosomal translocations in wild-type cells (Richardson and Jasin 2000). Although one DSB will not give rise to a translocation, two DSBs give rise to translocations at a readily detected frequency. Importantly, the molecular analysis of these events demonstrates that HR is not involved. That is, when HR occurs between two different chromosomes during the repair of DSBs, the events are completed precisely, without giving rise to genomic rearrangements (Richardson and Jasin 2000). The results indicate a strong preference for noncrossover HR events in mammalian cells. The other DSB repair pathways, NHEJ and single-strand annealing, are in contrast much more prone to give rise to translocations. This system, therefore, lends itself to examining in molecular detail the effect of genes mutated in chromosomal fragility syndromes.
The next two speakers, Steve Jackson (Cambridge, UK) and María Blasco (Madrid, Spain), focused on overlaps and cross-talk between pathways controlling DNA-damage responses and those involved in maintaining telomeres, the ends of linear chromosomes. Telomeres contain long stretches of short DNA tandem repeats (TTAGGG in mammals) that are added by the specialized reverse transcriptase, telomerase, which is composed of a catalytic subunit (Tert in humans, Terc in mice) and an associated RNA component. Despite terminating the DNA double-helix, telomeres do not normally trigger DNA-damage responses, and this presumably reflects them being sequestered by specialized telomeric proteins. Strikingly, however, in the past few years a range of studies has shown that many proteins associated with DNA-damage responses localize to telomeres and actually play key roles in controlling telomeric integrity (Blasco 2003).
Blasco began by explaining how mice inactivated for Terc exhibit gradual telomere shortening over several generations and that this is associated with a progressive loss of mouse vitality and, in the later generations, infertility and a range of other organ and tissue pathologies. Furthermore, cells derived from late-generation Terc-/- mice exhibit frequent telomere-to-telomere fusions, lose their proliferative potential, and tend to enter p53-dependent apoptosis. Blasco noted, however, that generation 5 (G5) Terc-/- mice are more resistant to induced skin tumorigenesis than wild-type mice, which is consistent with a model in which proliferative defects associated with telomere shortening can provide an important tumor-suppressor function. She then discussed how the DNA-dependent protein kinase catalytic subunit (DNA-PKcs) and its DNA-targeting subunit Ku86— two proteins involved in the DNA-repair pathway of NHEJ—play important roles at the telomere. One main conclusion from this work was that DNA-PKcs and Ku provide a capping function that prevents telomeres entering into end-fusions. Another key finding was that inactivation of DNA-PKcs dramatically accelerates the rate of telomere loss in Terc-/- backgrounds, such that G1 of the doubly mutant mouse has a telomeric length phenotype as severe as that of a G2 Terc-/- animal. Furthermore, Blasco reported that aged DNA-PKcs-deficient mice display evidence of telomere shortening, suggesting that this may be associated with the reduced life span and certain features of premature aging seen in this animal. Finally, Blasco reported that no telomere deficiencies are readily apparent in mice lacking the gene for Fancg (XRCC9). This suggests that the more rapid rates of telomere shortening observed in human FA cells as compared with controls (Callén et al. 2002), may be a secondary consequence of loss of the FA pathway, rather than reflecting a direct function at the telomere.
Jackson began by explaining that the standard lagging-strand DNA replication machinery is unable to copy the most distal telomere sequences and that without specialized systems to add new telomeric repeats, telomeres shorten progressively with every cell division. Notably, most human somatic cells do not express enough telomerase to maintain their telomeres in this manner, and, as a consequence, their telomeres gradually shorten as cells are serially passaged in culture, ultimately becoming so short that their normal functions are lost. This leads to apoptosis in some cell types whereas in others, such as human fibroblasts, it elicits senescence—a state in which the cell has permanently withdrawn from the cell cycle but remains metabolically active. Jackson described recent work revealing that cells undergoing telomere-initiated senescence seem to bear all the hallmarks of cells experiencing DSBs (d'Adda di Fagagna et al. 2003). These features include the activation of upstream and downstream components of the DNA-damage checkpoint pathway and the recruitment of checkpoint and DNA DSB repair proteins to subnuclear foci that are readily detected by indirect immunofluorescence. Jackson thus drew attention to the diagnostic potential of DNA-damage response markers in cancer and perhaps also other age-related diseases.
Jackson then showed that ablation of DNA-damage checkpoint signaling by using siRNA or dominant-negative constructs for the checkpoint kinases ATM, ATR, CHK1, and CHK2 resulted in a significant proportion of senescent cells resuming cell cycle progression into S phase. Although these results did not exclude other mechanisms for initiating and/or maintaining senescence, they revealed that DNA-damage checkpoint activation is causally associated with the senescent state. Jackson then described the whole-genome approach of ChIP-on-ChIP and how it had been used to reveal that telomeric ends directly participate in the DNA-damage response in senescent human cells. Furthermore, this approach showed that phosphorylated histone H2AX extends for large distances inward from the telomeric end in senescent cells. He explained how this type of analysis might not only better explain telomere-initiated senescence but might also provide important insights into DNA-repair and DNA-damage signaling in other contexts. Finally, Jackson pointed out that defects in many DNA-damage response pathways result in telomere defects and/or premature cell senescence in culture, raising the possibility that these might cause various phenotypes associated with human chromosomal fragility syndromes.
A good example of cross-talk between chromosome fragility syndromes was described by Wen-Hsing Cheng (Baltimore, MD), who discussed functional interactions between the proteins WRN (deficient in Werner syndrome) and NBS1. Previous work has implicated WRN, which contains helicase and DNA exonuclease domains, in DNA repair, and has established that NBS1 functions in a complex with MRE11 and RAD50 in DNA repair and checkpoint signaling following the generation of DSBs (see above). Cheng showed that when human cells are treated with agents that produce DSBs, WRN leaves its normal nucleolar location and over the course of several hours becomes concentrated in foci that also contain NBS1 and therefore correspond to sites of DNA damage. Consistent with this colocalization, Cheng showed that NBS1 and WRN can be coimmunoprecipitated and this interaction was enhanced following DNA damage. He also explained that the relocalization of WRN was hampered in NBS cells, whereas NBS1 relocalization still ensued in WRN cells. These data therefore implied that NBS1 facilitates the recruitment of WRN to sites of DNA damage. Consistent with these data, Cheng showed that siRNA-mediated depletion of WRN enhanced the sensitivity of wild-type cells to IR or MMC but did not further sensitize NBS cells to such agents. Thus, WRN appears to operate in the same pathway as NBS1 in mediating resistance to these DNA-damaging drugs. Finally, Cheng provided biochemical data indicating that the MRE11–RAD50–NBS1 complex, but not the MRE11–RAD50 complex, stimulates the DNA helicase activity of WRN. Taken together then, these results suggest a model in which NBS1 facilitates the recruitment of WRN to sites of DNA damage in vivo and that the combined activities of WRN and NBS1 may be required to bring about the repair of recalcitrant DNA lesions.
Andre Nussenzweig (Bethesda, MD) addressed the topical issues of how chromatin modulates cellular responses to DNA damage and, conversely, how DNA damage can lead to alterations in chromatin structure. After briefly reviewing the field, he explained how the DNA-damage activated kinases ATM, ATR, and DNA-PK bring about extensive phosphorylation of the C terminus of mammalian histone H2AX flanking sites of DNA damage—particularly DSBs (Rogakou et al. 1998). This phosphorylated species, often termed γ-H2AX, plays important roles in DNA-damage responses as indicated by the radiosensitivity and chromosomal instability phenotypes of H2AX knockout mice (Celeste et al. 2003a). Nussenzweig described how fibroblasts derived from such animals grow very poorly in culture and display defective G2/M checkpoint responses at low radiation doses. He also showed that, although such cells display no overt defects in telomere maintenance during mitotic cell cycles, meiotic cells of H2AX knockout mice display prolonged telomeric associations, suggesting a role for γ-H2AX in controlling meiotic recombination. Consistent with such an idea, H2AX-/- mice are also defective in forming the meiotic X-Y body that contains the sex chromosomes in an apparently condensed and inactive state. Notably, Nussenzweig explained that, although H2AX-/- mice are not strongly tumorprone, H2AX inactivation in p53-/- backgrounds significantly enhances rates of tumor development. Strikingly, this is even the case in H2AX heterozygotes, and analysis of the resulting tumors revealed that they retained the wild-type H2AX allele (Bassing et al. 2003; Celeste et al. 2003b). Nussenzweig noted that histone H2AX might also act as a tumor suppressor in humans and pointed out that its gene maps to a chromosome region frequently lost or rearranged in lymphomas as well as solid cancers.
In the last part of his talk, Nussenzweig explored how γ-H2AX promotes genome stability. Describing elegant studies using “laser scissors” to rapidly generate DNA damage in defined subnuclear volumes, he established that γ-H2AX is required for the retention of DNA-repair and checkpoint factors within foci at sites of DNA damage but not the initial recruitment of such factors to these sites.
Finally, Nussenzweig showed that γ-H2AX foci in irradiated cells colocalize with foci recognized by antibodies directed against Ser 14-phosphorylated histone H2B (P-Ser 14 H2B). Moreover, P-Ser 14 H2B foci formation was shown to depend on γ-H2AX. As γ-H2AX was not required for P-Ser H2B phosphorylation as ascertained by Western blot analysis, however, Nussenzweig speculated that P-Ser 14 H2B foci might reflect the condensation of chromatin at sites of DNA damage. In the discussion, it was speculated that such condensation might facilitate the coalescence of DNA-repair foci, as described earlier by Kanaar.
Hein te Riele (Amsterdam, The Netherlands) finally addressed the functions of DNA mismatch repair (MMR) proteins, deficiencies which are associated with hereditary nonpolyposis colorectal cancer (HNPCC) and a subset of sporadic cancers in humans. He began by explaining that DNA mismatches can arise through mistakes during DNA replication and when HR takes place between related but nonidentical sequences, and that such mismatches are recognized by two heterodimeric protein complexes, MSH2/MSH6 and MSH2/MSH3. To understand the functions of these proteins in more detail, te Riele described the generation and analysis of mice and of mouse cell lines deficient in these components. One key conclusion from these studies was that, as in humans, inactivation of Msh2 in the mouse leads to highly penetrant cancer predisposition and, at the cellular level, increased rates of spontaneous mutagenesis yet higher tolerance toward DNA methylating agents. te Riele then described the generation of an Msh2 allele, Msh2low, that causes Msh2 protein levels to be only ∼10% of that of wild type. Notably, Msh2-/low embryonic stem (ES) cells were found to be as resistant toward DNA-methylating agents as Msh2-/- cells but had spontaneous mutation rates similar to those of wild-type cells. Moreover, te Riele explained that Msh2-/low mice displayed only low levels of tumor incidence. Taken together, these results lead to the striking conclusion that the tumor-prone phenotype of Msh2-deficient mice primarily reflects increased spontaneous mutagenesis rather than aberrant responses to exogenous DNA-damaging agents.
Next, te Riele described a series of elegant studies that explored the role of MMR proteins in suppressing recombination between related but nonidentical DNA sequences. These studies measured recombination rates in wild-type or MMR mutant embryonic stem cells that contained stably integrated constructs of the neomycin-resistance gene (neo) bearing a 2-bp insertion or a base-pair substitution. Targeting vectors containing wild-type neo sequences were then transfected into these cells, and they were screened for neomycin resistance. The main conclusion from this work was that MMR-deficient cells are 10- to 50-fold more efficient than wild-type cells at mediating HR within DNA regions containing even subtle 1-bp or 2-bp mismatches. These findings therefore reveal that the MMR pathway is very potent at inhibiting recombination in such situations and raises the possibility that aberrant HR between related but nonidentical sequences plays an important role in the pathology of HNPCC. Conversely, whereas MMR can inhibit recombination, Pablo Huertas from Andrés Aguilera's laboratory (Seville, Spain) explained that transcription elongation impairment can cause hyperrecombination. He discussed recent data indicating that DNA:RNA hybrids can be formed cotranscriptionally, diminishing transcription elongation efficiency and promoting recombination (Huertas and Aguilera 2003). These results further enforce the complex interplay between transcription, recombination, replication, DNA repair, and chromatin and nuclear organization in eukaryotic genomes (Aguilera 2002; Surrallés et al. 2002).
As we learn about the function and molecular biology of the proteins involved in genome and chromosome stability, the final picture is more and more complicated. The list of interactions between factors is rapidly increasing to make an integrated network of genome stability pathways (Fig. 1). FA is the syndrome with the highest number of possible interactions and, therefore, the FA pathway would act as the spider in this macrotumor suppressor spider's web. Although the number of yet unresolved questions is increasing, we are now closer to understand the basis for the differential site-specificity in cancer among syndromes, their heterogeneous mutagen sensitivity spectra and clinical phenotypes, and the role of these tumor-suppressor proteins in DNA-damage responses, telomere function, cell senescence, and aging. This will undoubtedly lead us to a better understanding of the origin of cancer and chromosome fragility syndromes and, consequently, to a knowledge base for the development of novel therapies. These important challenges will keep us busy for years to come.
Acknowledgments
This International Workshop was held in Madrid, Spain, a few weeks before the tragic terrorist attack of March 11, 2004. This paper is dedicated to the memory of all innocent people who died in this sad and nonsense event. We are grateful to all speakers for helpful comments and for sharing unpublished data and to Elsa Callén for assistance with the figure. Finally, we thank Lucía Franco and the rest of the staff of the Juan March Foundation Center for International Meetings on Biology for creating such an excellent scientific and social atmosphere that strongly promoted exiting discussions and interpersonal contacts.
Article and publication are at http://www.genesdev.org/cgi/doi/10.1101/gad.1216304.
References
- Aguilera A. 2002. The connection between transcription and genomic instability. EMBO J. 21: 195-201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aten J.A., Stap, J., Krawczyk, P.M., van Oven, C.H., Hoebe, R.A., Essers, J., and Kanaar, R. 2004. Dynamics of DNA double-strand breaks revealed by clustering of damaged chromosome domains. Science 303: 92-95. [DOI] [PubMed] [Google Scholar]
- Bakkenist C.J. and Kastan, M.B. 2003. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421: 499-506. [DOI] [PubMed] [Google Scholar]
- Bassing C.H., Suh, H., Ferguson, D.O., Chua, K.F., Manis, J., Eckersdorff, M., Gleason, M., Bronson, R., Lee, C., and Alt, F.W. 2003. Histone H2AX: A dosage-dependent suppressor of oncogenic translocations and tumors. Cell 114: 359-370. [DOI] [PubMed] [Google Scholar]
- Bignell G., Micklem, G., Stratton, M.R., Ashworth, A., and Wooster, R. 1997. The BRC repeats are conserved in mammalian BRCA2 proteins. Hum. Mol. Genet. 6: 53-58. [DOI] [PubMed] [Google Scholar]
- Blasco M. 2003. Telomeres and cancer: A tale with many endings. Curr. Opin. Genet. Dev. 13: 70-76. [DOI] [PubMed] [Google Scholar]
- Blom E., Van de Vrugt, H.J., De Vries, Y., De Winter, J.P., Arwert, F., and Joenje, H. 2004. Multiple TPR motifs characterize the Fanconi anemia FANCG protein. DNA Repair 5: 77-84. [DOI] [PubMed] [Google Scholar]
- Callén E., Samper, E., Ramírez, M.J., Creus, A., Marcos, R., Ortega, J.J., Olive, T., Badell, I., Blasco, M.A., and Surrallés, J. 2002. Breaks at telomeres and TRF2-independent end fusions in Fanconi anemia. Hum. Mol. Genet. 11: 439-444. [DOI] [PubMed] [Google Scholar]
- Carson C.T., Schwartz, R.A., Stracker, T.H., Lilley, C.E., Lee, D.V., and Weitzman, M.D. 2003. The Mre11 complex is required for ATM activation and the G2/M checkpoint. EMBO J. 22: 6610-6620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castillo V., Cabre, O., Marcos, R., and Surrallés, J. 2003. Molecular cloning of the Drosophila Fanconi anaemia gene FANCD2 cDNA. DNA Repair 2: 751-758. [DOI] [PubMed] [Google Scholar]
- Celeste A., Petersen, S., Romanienko, P.J., Fernandez-Capetillo, O., Chen, H.T., Sedelnikova, O.A., Reina-San-Martin, B., Coppola, V., Meffre, E., Difilippantonio, M.J., et al. 2003a. Genomic instability in mice lacking histone H2AX. Science 296: 922-927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Celeste A., Difilippantonio, S., Difilippantonio, M.J., Fernandez-Capetillo, O., Pilch, D.R., Sedelnikova, O.A., Eckhaus, M., Ried, T., Bonner, W.M., and Nussenzweig, A. 2003b. H2AX haploinsufficiency modifies genomic stability and tumor susceptibility. Cell 114: 371-383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Couëdel C., Mills, K.D., Barchi, M., Shen, L., Olshen, A., Johnson, R.D., Nussenzweig, A., Essers, J., Kanaar, R., Li, G.C., et al. 2004. Collaboration of homologous recombina-tion and nonhomologous end-joining factors to the survival and integrity of mice and cells. Genes & Dev. 18: 1293-1304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cox M.M., Goodman, M.F., Kreuzer, K.N., Sherratt, D.J., Sandler, S.J., and Marians, K.J. 2000. The importance of repairing stalled replication forks. Nature 404: 37-41. [DOI] [PubMed] [Google Scholar]
- d'Adda di Fagagna F., Reaper, P.M., Clay-Farrace, L., Fiegler, H., Carr, P., Von Zglinicki, T., Saretzki, G., Carter, N.P., and Jackson, S.P. 2003. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426: 194-198. [DOI] [PubMed] [Google Scholar]
- D'Andrea A.D. 2003. The Fanconi road to cancer. Genes & Dev. 17: 1933-1936. [DOI] [PubMed] [Google Scholar]
- Davies A.A., Masson, J.-Y., McIlwraith, M.J., Stasiak, A.Z., Stasiak, A., Venkitaraman, A.R., and West, S.C. 2001. Role of BRCA2 in control of the RAD51 recombination and DNA repair protein. Mol. Cell 7: 273-282. [DOI] [PubMed] [Google Scholar]
- Essers J., Hendriks, R.W., Swagemakers, S.M., Troelstra, C., de Wit, J., Bootsma, D., Hoeijmakers, J.H., and Kanaar, R. 1997. Disruption of mouse RAD54 reduces ionizing radiation resistance and homologous recombination. Cell 89: 195-204. [DOI] [PubMed] [Google Scholar]
- Friedberg E.C., Wagner, R., and Radman, M. 2002. Specialized DNA polymerases, cellular survival, and the genesis of mutations. Science 296: 1627-1630. [DOI] [PubMed] [Google Scholar]
- Garcia-Higuera I., Taniguchi, T., Ganesan, S., Meyn, M.S., Timmers, C., Hejna, J., Grompe, M., and D'Andrea, A.D. 2001. Interaction of the Fanconi anemia proteins and BRCA1 in a common pathway. Mol. Cell 7: 249-262. [DOI] [PubMed] [Google Scholar]
- Gerlich D., Beaudouin, J., Kalbfuss, B., Daigle, N., Eils, R., and Ellenberg, J. 2003. Global chromosome positions are transmitted through mitosis in mammalian cells. Cell 112: 751-764. [DOI] [PubMed] [Google Scholar]
- Godthelp B.C., Artwert, F., Joenje, H., and Zdzienicka, M.A. 2002. Impaired DNA damage-induced nuclear Rad51 foci formation uniquely characterizes Fanconi anemia group D1. Oncogene 21: 5002-5005. [DOI] [PubMed] [Google Scholar]
- Houghtaling S., Timmers, C., Noll, M., Finegold, M.J., Jones, S.N., Meyn, M.S., and Grompe, M. 2003. Epithelial cancer in Fanconi anemia complementation group D2 (FancD2) knockout mice. Genes & Dev. 17: 2021-2035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howlett N.G., Taniguchi, T., Olson, S., Cox, B., Waisfisz, Q., De Die-Smulders, C., Persky, N., Grompe, M., Joenje, H., Pals, G., et al. 2002. Biallelic inactivation of BRCA2 in Fanconi anemia. Science 297: 606-609. [DOI] [PubMed] [Google Scholar]
- Huertas P. and Aguilera, A. 2003. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12: 711-721. [DOI] [PubMed] [Google Scholar]
- Johnson R.D. and Jasin, M. 2000. Sister-chromatid recombination is a prominent DNA repair pathway in mammalian cells. EMBO J. 19: 3398-3407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim S.T., Xu, B., and Kastan, M.B. 2002. Involvement of the cohesin protein, Smc1, in ATM-dependent and independent responses to DNA damage. Genes & Dev. 16: 560-570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kraakman-van der Zwet M., Overkamp, W.J., van Lange, R.E., Essers, J., van Duijn-Goedhart, A., Wiggers, I., Swaminathan, S., van Buul, P.P., Errami, A., Tan, R.T., et al. 2002. Brca2 (XRCC11) deficiency results in radioresistant DNA synthesis and a higher frequency of spontaneous deletions. Mol. Cell. Biol. 22: 669-679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kraakman-van der Zwet M., Wiegant, W., and Zdzienicka, M.Z. 2003. Brca2 (XRCC11) deficiency results in enhanced mutagenesis. Mutagenesis 18: 521-525. [DOI] [PubMed] [Google Scholar]
- Kutler D.I., Singh, B., Satagopan, J., Batish, S.D., Berwick, M., Giampietro, P.F., Hanenberg, H., and Auerbach, A.D. 2003. A 20-year perspective on the International Fanconi Anemia Registry (IFAR). Blood 101: 1249-1256. [DOI] [PubMed] [Google Scholar]
- Levitus M., Rooimans, M.A., Steltenpool, J., Cool, N.F.C., Oostra, A.B., Mathew, A.G., Hoatlin, M.E., Waisfisz, Q., Arwert, F., De Winter, J.P., et al. 2004. Genetic heterogeneity in Fanconi anemia: Evidence for two new genetic subtypes. Blood 103: 2498-2503. [DOI] [PubMed] [Google Scholar]
- Liu Y., Masson, J.Y., Shah, R., O'Regan, P., and West, S.C. 2004. RAD51C is required for Holliday junction processing in mammalian cells. Science 303: 243-246. [DOI] [PubMed] [Google Scholar]
- Lomonosov M., Anand, S., Sangrithi, M., Davies, R., and Venkitaraman, A.R. 2003. Stabilization of stalled DNA replication forks by the BRCA2 breast cancer susceptibility protein. Genes & Dev. 17: 3017-3022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marsit C.J., Liu, M., Nelson, H., Posner, M., Suzuki, M., and Kelsey, K.T. 2004. Inactivation of the Fanconi anemia/BRCA pathway in lung and oral cancers: Implications for treatment and survival. Oncogene 23: 1000-1004. [DOI] [PubMed] [Google Scholar]
- Meetei A.R., De Winter, J.P., Medhurst, A.L., Wallisch, M., Waisfisz, Q., Van de Vrugt, H.J., Oostra, A.B., Yan, Z., Ling, C., Bishop, C.E., et al. 2003a. A novel ubiquitin ligase is deficient in Fanconi anaemia. Nat. Genet. 35: 165-170. [DOI] [PubMed] [Google Scholar]
- Meetei A.R., Sechi, S., Wallisch, M., Yang, D., Young, M.K., Joenje, H., Hoatlin, M.E., and Wang, W. 2003b. A multiprotein nuclear complex connects Fanconi anemia and Bloom syndrome. Mol. Cell. Biol. 23: 3417-3426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meetei A.R., Yan, Z., and Wang, W. 2004. FANCL Replaces BRCA1 as the likely ubiquitin ligase responsible for FANCD2 monoubiquitination. Cell Cycle 3: 179-181. [PubMed] [Google Scholar]
- Moynahan M.E., Pierce, A.J., and Jasin, M. 2001. BRCA2 is required for homology-directed repair of chromosomal breaks. Mol. Cell 7: 263-272. [DOI] [PubMed] [Google Scholar]
- Nakanishi K., Taniguchi, T., Ranganathan, V., New, H.V., Moreau, L.A., Stotsky, M., Mathew, C.G., Kastan, M.B., Weaver, D.T., and D'Andrea, A.D. 2002. Interaction of FANCD2 and NBS1 in the DNA damage response. Nat. Cell Biol. 4: 913-920. [DOI] [PubMed] [Google Scholar]
- Nussenzweig A., Chen, C., da Costa Soares, V., Sanchez, M., Sokol, K., Nussenzweig, M.C., and Li, G.C. 1996. Requirement for Ku80 in growth and immunoglobulin V(D)J recombination. Nature 382: 551-555. [DOI] [PubMed] [Google Scholar]
- O'Driscoll M., Cerosaletti, K.M., Girard, P.M., Dai, Y., Stumm, M., Kysela, B., Hirsch, B., Gennery, A., Palmer, S.E., Seidel, J., et al. 2001. DNA ligase IV mutations identified in patients exhibiting developmental delay and immunodeficiency. Mol. Cell 8: 1175-1185. [DOI] [PubMed] [Google Scholar]
- O'Driscoll M., Ruiz-Perez, V.L., Woods, C.G., Jeggo, P.A., and Goodship, J.A. 2003. A splicing mutation affecting expression of ataxia-telangiectasia and Rad3-related protein (ATR) results in Seckel syndrome. Nat. Genet. 33: 497-501. [DOI] [PubMed] [Google Scholar]
- Offit K., Levran, O., Mullaney, B., Mah, K., Nafa, K., Batish, S.D., Diotti, R., Schneider, R.H., Deffenbaugh, A., Scholl, T., et al. 2003. Shared genetic susceptibility to breast cancer, brain tumors, and Fanconi anemia. J. Nat. Cancer Inst. 95: 1548-1551. [DOI] [PubMed] [Google Scholar]
- Pagano G. and Youssoufian, H. 2003. Fanconi anaemia proteins: Major roles in cell protection against oxidative damage. Bioessays 25: 589-595. [DOI] [PubMed] [Google Scholar]
- Pellegrini L., Yu, D.S., Lo, T., Anand, S., Lee, M.Y., Blundell, T.L., and Venkitaraman, A.R. 2002. Insights into DNA recombination from the structure of a RAD51–BRCA2 complex. Nature 420: 287-293. [DOI] [PubMed] [Google Scholar]
- Petrini J.H. and Theunissen, J.W. 2004. Double strand break metabolism and cancer susceptibility: Lessons from the Mre11 complex. Cell Cycle May [Epub ahead of print]. [DOI] [PubMed]
- Pichierri P. and Rosselli, F. 2004. The DNA crosslink-induced S-phase checkpoint depends on ATR–CHK1 and ATR–NBS1–FANCD2 pathways. EMBO J. 23: 1178-1187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richardson C. and Jasin, M. 2000. Frequent chromosomal translocations induced by DNA double-strand breaks. Nature 405: 697-700. [DOI] [PubMed] [Google Scholar]
- Rogakou E.P., Pilch, D.R., Orr, A.H., Ivanova, V.S., and Bonner, W.M. 1998. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 273: 5858-5868. [DOI] [PubMed] [Google Scholar]
- Seal S., Barfoot, R., Jayatilake, H., Smith, P., Renwick, A., Bascombe, L., McGuffog, L., Evans, D.G., Eccles, D., Easton, D.F., et al. 2003. Breast cancer susceptibility collaboration. Evaluation of Fanconi anemia genes in familial breast cancer predisposition. Cancer Res. 63: 8596-8599. [PubMed] [Google Scholar]
- Sonoda E., Okada, T., Zhao, G.Y., Tateishi, S., Araki, K., Yamaizumi, M., Yagi, T., Verkaik, N.S., van Gent, D.C., Takata, M., et al. 2003. Multiple roles of Rev3, the catalytic subunit of polzeta in maintaining genome stability in vertebrates. EMBO J. 22: 3188-3197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stracker T.H., Carson, C.T., and Weitzman, M.D. 2002. Adenovirus oncoproteins inactivate the Mre11–Rad50–NBS1 DNA repair complex. Nature 418: 348-352. [DOI] [PubMed] [Google Scholar]
- Surrallés J., Ramirez, M.J., Marcos, R., Natarajan, A.T., and Mullenders, L.H. 2002. Clusters of transcription-coupled repair in the human genome. Proc. Natl. Acad. Sci. 99: 10571-10574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taniguchi T., Tischkowitz, M., Ameziane, N., Hodgson, S.V., Mathew, C.G., Joenje, H., Mok, S.C., and D'Andrea, A.D. 2003. Disruption of the Fanconi anemia–BRCA pathway in cisplatin-sensitive ovarian tumors. Nat. Med. 9: 568-574. [DOI] [PubMed] [Google Scholar]
- Tibbetts R.S., Cortez, D., Brumbaugh, K.M., Scully, R., Livingston, D., Elledge, S.J., and Abraham, R.T. 2000. Functional interactions between BRCA1 and the checkpoint kinase ATR during genotoxic stress. Genes & Dev. 14: 2989-3002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tischkowitch M., Ameziane, N., Waisfisz, Q., De Winter, J.P., Harris, R., Taniguchi, T., D'Andrea, A.D., Hodgson, S.V., Mathew, C.G., and Joenje, H. 2003. Bi-allelic silencing of the Fanconi anaemia gene FANCF in acute myeloid leukaemia. Br. J. Haematol. 123: 469-471. [DOI] [PubMed] [Google Scholar]
- Uziel T., Lerenthal, Y., Moyal, L., Andegeko, Y., Mittelman, L., and Shiloh, Y. 2003. Requirement of the MRN complex for ATM activation by DNA damage. EMBO J. 22: 5612-5621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West S.C. 2003. Molecular views of recombination proteins and their control. Nat. Rev. Mol. Cell Biol. 4: 435-445. [DOI] [PubMed] [Google Scholar]
- Wong A.K.C., Pero, R., Ormonde, P.A., Tavtigian, S.V., and Bartel, P.L. 1997. RAD51 interacts with the evolutionarily conserved BRC motifs in the human breast cancer susceptibility gene BRCA2. J. Biol. Chem. 272: 31941-31944. [DOI] [PubMed] [Google Scholar]
- Yamashita Y.M., Okada, T., Matsusaka, T., Sonoda, E., Zhao, G.Y., Araki, K., Tateishi, S., Yamaizumi, M., and Takeda, S. 2002. RAD18 and RAD54 cooperatively contribute to maintenance of genomic stability in vertebrate cells. EMBO J. 21: 5558-5566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu D.S., Sonoda, E., Takeda, S., Huang, C.L., Pellegrini, L., Blundell, T.L., and Venkitaraman, A.R. 2003. Dynamic control of RAD51 recombinase by self-association and interaction with BRCA2. Mol. Cell 12: 1029-1041. [DOI] [PubMed] [Google Scholar]