Abstract
Background
Many bacteria causing systemic invasive infections originate from the oral cavity by entering the bloodstream. Recently, a novel pathogenic bacterium, Streptococcus tigurinus, was identified as causative agent of infective endocarditis, spondylodiscitis and meningitis. In this study, we sought to determine the prevalence of S. tigurinus in the human oral microbial flora and analyzed its association with periodontal disease or health.
Results
We developed a diagnostic highly sensitive and specific real-time TaqMan PCR assay for detection of S. tigurinus in clinical samples, based on the 16S rRNA gene. We analyzed saliva samples and subgingival plaque samples of a periodontally healthy control group (n = 26) and a periodontitis group (n = 25). Overall, S. tigurinus was detected in 27 (53%) out of 51 patients. There is no significant difference of the frequency of S. tigurinus detection by RT-PCR in the saliva and dental plaque samples in the two groups: in the control group, 14 (54%) out of 26 individuals had S. tigurinus either in the saliva samples and/or in the plaque samples; and in the periodontitis group, 13 (52%) out of 25 patients had S. tigurinus in the mouth samples, respectively (P = 0.895). The consumption of nicotine was no determining factor.
Conclusion
Although S. tigurinus was a frequently detected species of the human oral microbial flora, it was not associated with periodontal disease. Further investigations are required to determine whether S. tigurinus is a commensal or an opportunistic oral pathogen with a potential for development of invasive infections.
Keywords: Streptococcus tigurinus, Specific RT TaqMan PCR, Periodontitis, Oral microbiome
Background
The human oral microbiome consists of a number of bacteria; most of them are non-pathogenic commensals or act as opportunistic pathogens [1]. Some oral bacteria are implicated in oral diseases such as dental caries and periodontitis, which are among the most common infections in humans. Periodontitis in particular represents an inflammatory disease that affects 15-47% of the world-wide population [2],[3] and contributes to the morbidity of other chronic diseases [4]. Although more than 700 species were shown to colonize the oral cavity [5], evidence suggests that only a few of them, such as Aggregatibacter actinomycetemcomitans or Porphyromonas gingivalis, are associated with the pathogenesis of periodontitis or systemic complications [6],[7].
In recent years, significant associations have been elucidated between periodontitis and other very common systemic diseases, including diabetes mellitus [8] and cardiovascular diseases [9]. This pathogenic association between the oral cavity and other parts of the human body is potentially triggered by oral bacteria entering the bloodstream, which increases the risk for invasive infections such as infective endocarditis [10]. Streptococcus tigurinus was recently identified as a novel pathogen associated with infective endocarditis, prosthetic joint infections or meningitis [11]-[13]. It has also been shown to be highly virulent in experimental animal models [14].
S. tigurinus belongs to the Streptococcus mitis group and is most closely related to Streptococcus mitis, Streptococcus oralis, Streptococcus pneumoniae, Streptococcus pseudopneumoniae and Streptococcus infantis. S. tigurinus forms α-hemolytic, smooth colonies with a diameter of 0.5 to 1 mm after incubation at 37°C for 24 h on sheep blood agar [11]. Because of the morphological resemblance to its most closely related species, accurate identification of S. tigurinus by conventional phenotypic methods is limited. Therefore, commercial test systems such as VITEK 2 (bioMérieux, Marcy L’Etoile, France) or matrix-assisted laser desorption ionization-time of flight mass spectrometry analyses are helpful for initial assignment to the S. mitis group, but genetic analyses are required for definitive assignment as S. tigurinus. Analysis of the 5′-end of the 16S rRNA gene allows accurate identification of S. tigurinus based on a significant sequence demarcation to the most closely related species [11].
To date, the oral cavity per se could not yet be identified as niche of S. tigurinus. In addition, no data exists, whether or not S. tigurinus is a frequent commensal of the human oral cavity. Therefore, a S. tigurinus specific real-time (RT) TaqMan PCR based on the 16S rRNA gene was developed to identify S. tigurinus directly in clinical oral samples. In this context, saliva and dental plaque samples from a non-periodontitis control group and periodontitis patients as a test group were investigated as we hypothesized that the prevalence of S. tigurinus may be influenced by periodontitis. In addition, the influence of smoking on the occurrence of S. tigurinus was assessed.
Methods
Study population
Human saliva samples and pooled plaque samples of two different groups, i.e., a non-periodontitis control group (n = 26; 18 females, mean age 27.7 years, range 16 to 58) and a periodontitis group (n = 25; 14 females, mean age 59.4 years, range 26 to 83) of patients of the Center of Dental Medicine, University of Zurich, Switzerland, were prospectively analyzed. This study was approved by the Ethics committee of the canton Zurich, Switzerland (reference number KEK-ZH-2012-0322) and was conducted according to the guidelines of the Declaration of Helsinki. Pregnant patients or patients under antibiotic therapy were excluded from the study. All patients gave their written informed consent for the study. Clinical data were retrieved from the patients’ medical and dental records. Smoking status was anamnestically registered.
Periodontal health status
In order to assess the periodontal health status of the patients, a periodontal examination was performed using a pressure-sensitive probe (Hawe Click Probe, Kerr Hawe, Bioggio, Switzerland), which included measurement of probing pocket depth (PPD) at six sites around each tooth. The dichotomous measurement of bleeding on probing (BOP) and presence of plaque/calculus or overhanging restorations were also recorded. All recordings were made by one calibrated investigator.
Based on this clinical data set, the periodontal health status was assessed by the periodontal screening index (PSI). This index provides an overall expression of the health status of the periodontium by assessing the PPD and BOP [15].
In brief, the staging is as follows: grade 0: no pockets >3 mm and no bleeding, grade 1: no pockets >3 mm, but presence of bleeding, grade 2: no pockets >3 mm, presence of bleeding plus the presence of calculus and/or overhanging restorations, grade 3: pockets of 4–5 mm, grade 4: pockets ≥6 mm. The highest score of a subject determined the clinical diagnosis according to the definition of Cutress and co-workers [16]: scores 0, 1, and 2: “no periodontitis”; scores 3 and 4: “periodontitis”.
Clinical sample collection
Saliva samples of each patient were obtained by paraffin stimulation for 5 min. In addition, one week after the periodontal charting, subgingival plaque samples were collected from the four deepest pockets in the periodontitis group and from the mesial sulcus of the first molars in the non-periodontitis control group by paper points and curette method as described earlier [17]. Four subgingival plaque samples were pooled together for each patient.
Primer design and TaqMan hydrolysis probes
To establish a S. tigurinus specific RT TaqMan PCR, 16S rRNA gene sequences of S. mitis group species available from GenBank database and of S. tigurinus type strain AZ_3aT (GenBank accession number JN004270) and S. tigurinus strain AZ_4a (JQ696861) were aligned using Clustal V program (Lasergene MegAlign software version 7, DNAstar, Madison, WI) (Figure 1). The sequence of S. tigurinus strain AZ_4a was included in the alignment as we observed a single nucleotide polymorphism at nucleotide position 150 at the 5′-end of the 16S rRNA gene. RT-PCR primers and TaqMan hydrolysis probes were chosen using PrimerExpress software version 3.0 (Life Technologies, Zug, Switzerland) following visual inspection of the aligned target sequences: forward primer StiF [5′-TGAAGAGAGGAGCTTGCTCTTCTTG-3′], reverse primer StiR [5′-GTTGCTCGGTCAGACTTCCGTC-3′], probe Sti3 [5′-6-FAM-AATGGATTATCGCATGATAA-MGB-3′, where FAM is 6-carboxyfluorescein and MGB is minor groove binder] and probe Sti4 [5′-NED-AATTGATTATCGCATGATAAT-MGB-3′, where NED is 2,7′,8′-benzo-5′-fluoro-2′,4,7-trichloro-5-carboxyfluorescein].
Figure 1.
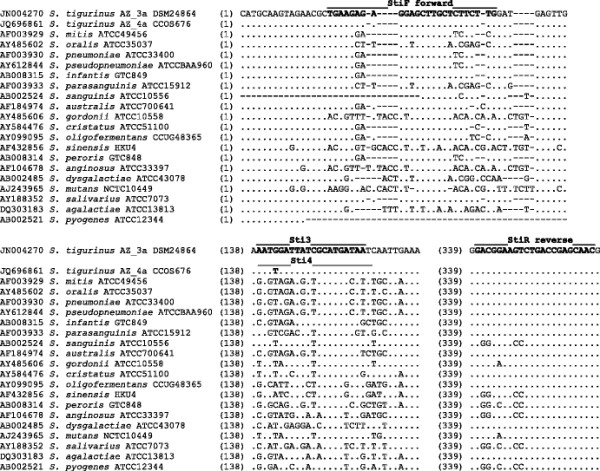
Homology analysis of partial 16S rRNA gene sequences ofS.tigurinusstrains,S.mitisgroup species and more distantly related streptococci shows hypervariable regions. Multiple alignment of the sequences was performed with the Clustal V program, sequence of the type strain S. tigurinus AZ_3aT (CCOS 600T; DSM 24864T), is the reference sequence. The lines above the reference sequence depict the positions of the forward and reverse primers and the S. tigurinus specific TaqMan probes Sti3 (specific for S. tigurinus AZ_3a) and Sti4 (specific for S. tigurinus AZ_4a).
DNA extraction and RT TaqMan PCR
DNA was extracted with an EZ1 DNA Tissue Kit (Qiagen, Hombrechtikon, Switzerland) following the manufacturer’s instructions. DNA extracts were eluted in 50 μl of PCR-grade water (Limulus amebocyte lysate [LAL] water; Lonza, Walkersville, MD). RT TaqMan PCR was performed on an Applied Biosystems 7500 fast instrument with 7500 System software (version 2.0.4). Each 25 μl mixture contained 12.5 μl of 2x PCR Mastermix (Roche Diagnostics, Rotkreuz, Switzerland), 2.5 μl of 10x exogenous internal positive-control primer and probe mix (VIC-labeled), 0.5 μl of 50x exogenous internal positive-control target DNA (both, Life Technologies), 0.25 μl of each primer (stock concentration, 30 μM), 0.5 μl of each probe (stock concentration, 5 μM), and 5.0 μl of DNA extract. The exogenous internal positive-control reagents were added to distinguish truly negative from falsely negative results due to PCR inhibition. PCR conditions were 2 min at 50°C and 10 min at 95°C, followed by 40 cycles of 15 s at 95°C and 60 s at 60°C. The positive-control plasmid pST3A containing a 435-bp segment of the 5′-end of the 16S rRNA gene (corresponding to positions 10 to 444 of the 16S rRNA gene of S. tigurinus AZ_3aT), containing the region as depicted in Figure 1, was constructed using in silico design and de novo synthesis and subcloning (Genscript, CA). The analytical sensitivity of the assay was determined by repeated testing of 10-fold dilutions of the plasmid positive control pST3A ranging from 5 × 105 to 5 × 10−1 copies. PCR-grade water (LAL water) was used as a negative control. Sensitivity was evaluated by testing DNA extracts of S. tigurinus strains AZ_1 (CCOS 683, Culture Collection of Switzerland), AZ_2 (CCOS 675), AZ_3aT (CCOS 600T; DSM 24864T, Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Braunschweig, Germany), AZ_4a (CCOS 676), AZ_6 (CCOS 681), AZ_7a (CCOS 677), AZ_8 (CCOS 678), AZ_10 (CCOS 679), AZ_11 (CCOS 682), AZ_12 (CCOS 680) and AZ_14 (CCOS 689); and of DNA extracts of 5 uncultured S. tigurinus (GenBank accession numbers JQ696868, JQ696870, JQ696871, JQ696872, JQ820471). Specificity was evaluated by testing DNA extracts of closely related streptococci, i.e., type strains of S. pneumoniae (DSM 20566T), S. mitis (DSM 12643T), S. oralis (DSM 20627T), S. pseudopneumoniae (CIP 108659T, Institut Pasteur, Paris, France) and S. infantis (CIP 105949T); and of clinical isolates of Streptococcus gordonii, Streptococcus sanguinis, Streptococcus parasanguinis, Streptococcus salivarius, Streptococcus anginosus, Streptococcus mutans and Streptococcus dysgalactiae. To further assess the assay specificity, amplification products from a sample tested positive with the S. tigurinus probes was sequenced and compared to known sequences using the NCBI BLAST tool and SmartGene software (SmartGene, Zug, Switzerland).
Statistical analyses
The crosstab chi-square test of independence was performed by the IBMS PSS statistic software version 20. P < 0.05 was considered statistically significant.
Results
Development of a RT-PCR for the detection of S. tigurinus
A TaqMan-based RT-PCR for highly sensitive and specific detection of S. tigurinus in clinical samples was developed. A 288-bp fragment at the 5′-end of the 16S rRNA gene was selected, which allowed discrimination between S. tigurinus and the most closely related species within the S. mitis group (Figure 1). All S. tigurinus samples including S. tigurinus strain AZ_4a were detected due to the incorporation of two probes Sti3 and Sti4, respectively. Closely related species such as S. pneumoniae, S. mitis, S. oralis, S. pseudopneumoniae and S. infantis were not detected by the S. tigurinus specific RT-PCR, as well as other more distantly related species, i.e., S. gordonii, S. sanguinis, S. parasanguinis, S. salivarius, S. anginosus, S. mutans and S. dysgalactiae, showing the specificity of the assay.
Repeated testing of 10-fold serial dilutions of purified pST3A DNA consistently showed that the limit of detection for S. tigurinus was around 5 copies of the 16S rRNA gene using the Sti3 probe. In addition, specificity of the assay was supported by the lack of reactivity of the Sti4 probe with pST3A, which contains the 16S rRNA gene of S. tigurinus strain AZ_3aT. No amplification was detected for a template dilution of less than 5 copies and the negative control.
Detection of S. tigurinus in the human oral cavity
In total, 51 saliva samples and 51 subgingival plaque samples obtained of 51 individuals were analyzed. In 22 (43%) out of 51 saliva samples and in 18 (35%) out of 51 plaque samples, S. tigurinus was detected by the specific RT-PCR, respectively. Overall, in 27 (53%) out of 51 individuals, S. tigurinus was detected in the saliva samples and/or in the plaque samples. In 13 (26%) individuals, S. tigurinus was detected both in the saliva and in the plaque samples. When comparing age groups <39 yr (n = 25), 40–65 yr (n = 16) and >65 yr (n = 10), no significant difference was observed for detection of S. tigurinus in the oral samples (P = 0.756).
Systemic comorbidities of patients were as follows: diabetes mellitus (n = 5), coronary heart disease (n = 3), rheumatoid arthritis (n = 1) and juvenile polyarthritis (n = 1); no immunosuppression was observed.
Influence of periodontitis in the occurrence of S. tigurinus
Clinical diagnosis of periodontitis was based on the PSI. Individuals of the non-periodontitis control group (n = 26) had PSI grades <3 whereas patients of the periodontitis group (n = 25) had PSI grades 3 (n = 2) and 4 (n = 23). There is no significant difference of the frequency of S. tigurinus detection by RT-PCR in the saliva and dental plaque samples in the two groups: in the control group, 14 (54%) out of 26 individuals had S. tigurinus either in the saliva samples and/or in the plaque samples, and in the periodontitis group, 13 (52%) out of 25 patients had S. tigurinus in the mouth samples, respectively (P = 0.895) (Tables 1 and 2). Four (15%) out of 26 individuals of the non-periodontitis group and 9 (36%) out of 25 patients of the periodontitis group had S. tigurinus in both the saliva and the plaque samples, respectively (P = 0.091).
Table 1.
Frequency of S . tigurinus detected in the oral microbial flora of the periodontally healthy subjects (n = 26) by specific RT TaqMan PCR
| Individual | Age, sex | Nicotine consumption | Detection of S . tigurinus in saliva sample by RT-PCR | Detection of S . tigurinus in subgingival plaque sample by RT-PCR |
|---|---|---|---|---|
| 1 |
23, f |
Yes |
Negative |
Positive |
| 2 |
23, f |
Yes |
Negative |
Negative |
| 3 |
18, f |
No |
Negative |
Negative |
| 4 |
18, f |
No |
Positive |
Negative |
| 5 |
22, f |
Yes |
Positive |
Positive |
| 6 |
16, f |
No |
Positive |
Negative |
| 7 |
23, f |
No |
Positive |
Negative |
| 8 |
18, f |
Yes |
Negative |
Negative |
| 9 |
39, f |
Yes |
Positive |
Positive |
| 10 |
16, f |
Yes |
Negative |
Negative |
| 11 |
26, f |
No |
Negative |
Negative |
| 12 |
26, m |
No |
Negative |
Negative |
| 13 |
24, f |
No |
Negative |
Negative |
| 14 |
48, m |
No |
Positive |
Negative |
| 15 |
31, m |
Yes |
Negative |
Negative |
| 16 |
53, m |
No |
Negative |
Negative |
| 17 |
24, f |
No |
Positive |
Positive |
| 18 |
26, f |
No |
Positive |
Negative |
| 19 |
33, m |
No |
Negative |
Positive |
| 20 |
58, m |
No |
Negative |
Negative |
| 21 |
25, m |
No |
Positive |
Positive |
| 22 |
23, m |
Yes |
Positive |
Negative |
| 23 |
34, f |
No |
Negative |
Negative |
| 24 |
25, f |
No |
Negative |
Negative |
| 25 |
24, f |
No |
Negative |
Positive |
| 26 | 25, f | No | Positive | Negative |
Table 2.
Frequency of S . tigurinus detected in the oral microbial flora of the periodontitis group (n = 25) by specific RT TaqMan PCR
| Patient | Age, sex | Nicotine consumption | Detection of S . tigurinus in saliva sample by RT-PCR | Detection of S . tigurinus in subgingival plaque sample by RT-PCR |
|---|---|---|---|---|
| 1 |
50, f |
Yes |
Positive |
Negative |
| 2 |
69, m |
Yes |
Negative |
Negative |
| 3 |
63, f |
No |
Negative |
Negative |
| 4 |
46, f |
Yes |
Negative |
Positive |
| 5 |
43, f |
No |
Negative |
Negative |
| 6 |
71, m |
Yes |
Negative |
Positive |
| 7 |
66, m |
No |
Negative |
Negative |
| 8 |
52, f |
Yes |
Positive |
Positive |
| 9 |
74, m |
Yes |
Positive |
Positive |
| 10 |
50, f |
No |
Positive |
Positive |
| 11 |
54, m |
No |
Negative |
Negative |
| 12 |
26, f |
No |
Positive |
Positive |
| 13 |
55, f |
No |
Negative |
Negative |
| 14 |
51, f |
Yes |
Negative |
Negative |
| 15 |
83, m |
No |
Positive |
Positive |
| 16 |
65, f |
No |
Positive |
Negative |
| 17 |
82, f |
No |
Positive |
Positive |
| 18 |
57, f |
Yes |
Positive |
Positive |
| 19 |
38, f |
No |
Positive |
Positive |
| 20 |
76, m |
No |
Negative |
Negative |
| 21 |
65, m |
Yes |
Negative |
Negative |
| 22 |
63, m |
Yes |
Negative |
Negative |
| 23 |
45, m |
Yes |
Positive |
Positive |
| 24 |
74, m |
Yes |
Negative |
Negative |
| 25 | 68, f | No | Negative | Negative |
Influence of nicotine consumption in the occurrence of S. tigurinus
In total, 20 out of 51 individuals had nicotine consumption, of which 11 had S. tigurinus detected in at least the saliva and/or plaque samples. This was not significant compared to individuals without nicotine consumption (31 out of 51, 16 with S. tigurinus detected in the oral samples), P = 0.813. In the periodontitis group, the number of patients with nicotine consumption and S. tigurinus detected in the oral samples (n = 7) did not differ significantly from the patients without nicotine consumption and S. tigurinus in the mouth (n = 6), P = 0.543, respectively. Similar results were observed in the non-periodontitis control group, 4 individuals with nicotine consumption and S. tigurinus detected in the oral samples were identified compared to 10 individuals without nicotine consumption but S. tigurinus detected in the mouth, P = 0.793.
Discussion
Members of the microbial flora originating from the oral cavity may be involved in the pathogenesis of systemic infections [18]. Biofilm formation, complex mechanisms with other bacteria or underlying diseases might play a crucial role in the development of invasive infections. Regarding the pathogenesis of chronic periodontal diseases, complex host-bacterial interactions are responsible for the initiation of tissue destruction [19],[20]. Earlier studies have demonstrated that S. mitis, which is the closest related species to S. tigurinus, is a predominant early colonizing species of dental biofilms [21]. Although S. mitis is not a potent inducer of immune responses, it can antagonize the capacity of A. actinomycetemcomitans to stimulate IL-8 [22]. Interaction of S. tigurinus with A. actinomycetemcomitans (a key pathogen associated with aggressive form of periodontitis in younger individuals) might be of interest [23]. Since its recent identification [11],[12], it is not clear whether modifying factors are associated with the presence of S. tigurinus in the human oral microbiome and if its detection in the oral cavity has direct clinical implications in systemic diseases.
Our data shows that S. tigurinus is a frequent bacterium colonizing the human oral cavity in periodontal health and disease. In more than half of the individuals (53%) investigated S. tigurinus was detected by the S. tigurinus specific RT-PCR. Overall, the frequency of S. tigurinus in the saliva and plaque samples in patients with periodontitis did not differ significantly from individuals in the non-periodontitis control group. Both, individuals with or without nicotine consumption, had S. tigurinus in the saliva/plaque samples, independent of the individual’s age. However, it remains to be investigated how S. tigurinus interacts with other oral bacteria and if there might be a similar inhibitory effect.
Whole-genome analyses of S. tigurinus revealed the presence of several virulence factors such as fibronectin-binding protein or exfoliative toxin [24], which might differentiate this bacterium from other oral commensal organisms of the normal microbial flora. However, little is understood how exactly S. tigurinus causes various invasive diseases. An enhanced resistance to phagocytosis by macrophages of S. tigurinus was shown without induction of platelet aggregation [14].
Previous studies have shown that S. tigurinus is a frequent and aggressive pathogen causing infective endocarditis [11],[12],[14]. For patient management and guidance of appropriate therapy, accurate identification of the causative agent is of major importance. The S. tigurinus specific RT-PCR allows accurate discrimination between S. tigurinus and the most closely related species within the S. mitis group. In future, the S. tigurinus specific RT-PCR might be useful for direct application on clinical samples, e.g., heart valves, for timely identifications of the pathogen in a routine diagnostic laboratory.
The human oral microbiome is comprised of a bacterial diversity including different phyla, e.g., Firmicutes, Actinobacteria, Proteobacteria, Bacteroidetes and Proteobacteria[5],[25]. Viridans streptococci, e.g., S. mitis, are known to be the predominant bacterial species in the human oral cavity and were detected in various dental sites [5]. The present study is the first to show comparatively that S. tigurinus can be detected both in saliva and in subgingival plaque samples, however, it remains to be determined if the occurrence of S. tigurinus is site specific. It is not surprising that S. tigurinus can be found in saliva in higher frequency than individually selected subgingival sites, since saliva has representatively bacteria from different oral sites including the subgingival area. Saliva has been shown to be a suitable biological fluid, alternative to pooled subgingival plaque samples for detection of oral bacteria such as newly identified Synergistetes[26].
Conclusions
We developed a diagnostic, highly sensitive and specific RT TaqMan PCR for direct detection of S. tigurinus in clinical samples. The data of the present study suggests for the first time that S. tigurinus represents a prevalent organism of the oral microbiome and that its occurrence is not increased by the presence of periodontal disease or smoking. However, prospective studies with larger populations are required to determine whether S. tigurinus is a commensal or an opportunistic oral pathogen with a potential for development of invasive infections.
Competing interests
The authors declare that they have no competing interests.
Authors’ contribution
AZ contributed to the overall study design, analysis of molecular data and drafting the manuscript. FA contributed to the acquisition of the clinical samples and parameters. RZ contributed to the overall study design and laboratory data. FM contributed to the statistical analysis. PRS contributed to the overall study design, acquisition of clinical samples and critical revision of the draft. GVB contributed to overall study design, development of molecular methods and critical revision of the draft. NB contributed to the overall study design, acquisition of clinical samples and data and drafting the manuscript. All authors read and approved the final manuscript.
Contributor Information
Andrea Zbinden, Email: zbinden.andrea@virology.uzh.ch.
Fatma Aras, Email: fatma.aras@uzh.ch.
Reinhard Zbinden, Email: rzbinden@imm.uzh.ch.
Forouhar Mouttet, Email: f.mouttet@gmail.com.
Patrick R Schmidlin, Email: patrick.schmidlin@zzm.uzh.ch.
Guido V Bloemberg, Email: bloemberg@imm.uzh.ch.
Nagihan Bostanci, Email: nagihan.bostanci@zzm.uzh.ch.
Acknowledgment
The study was supported by the University of Zurich. We thank the laboratory technicians for their dedicated help.
References
- Marsh PD. Are dental diseases examples of ecological catastrophes? Microbiology. 2003;149(Pt 2):279–294. doi: 10.1099/mic.0.26082-0. [DOI] [PubMed] [Google Scholar]
- Albandar JM. Underestimation of periodontitis in NHANES surveys. J Periodontol. 2011;82(3):337–341. doi: 10.1902/jop.2011.100638. [DOI] [PubMed] [Google Scholar]
- Konig J, Holtfreter B, Kocher T. Periodontal health in Europe: future trends based on treatment needs and the provision of periodontal services–position paper 1. Eur J Dent Educ. 2010;14(Suppl 1):4–24. doi: 10.1111/j.1600-0579.2010.00620.x. [DOI] [PubMed] [Google Scholar]
- Marcenes W, Kassebaum NJ, Bernabe E, Flaxman A, Naghavi M, Lopez A, Murray CJ. Global burden of oral conditions in 1990–2010: a systematic analysis. J Dent Res. 2013;92(7):592–597. doi: 10.1177/0022034513490168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43(11):5721–5732. doi: 10.1128/JCM.43.11.5721-5732.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papapanou PN, Behle JH, Kebschull M, Celenti R, Wolf DL, Handfield M, Pavlidis P, Demmer RT. Subgingival bacterial colonization profiles correlate with gingival tissue gene expression. BMC Microbiol. 2009;9:221. doi: 10.1186/1471-2180-9-221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Socransky SS, Haffajee AD. Implications of periodontal microbiology for the treatment of periodontal infections. Compt Rendus Geosci. 1994;18:S684–S685. 688–693; quiz S714-687. [PubMed] [Google Scholar]
- Lalla E, Papapanou PN. Diabetes mellitus and periodontitis: a tale of two common interrelated diseases. Nat Rev Endocrinol. 2011;7(12):738–748. doi: 10.1038/nrendo.2011.106. [DOI] [PubMed] [Google Scholar]
- Beck JD, Offenbacher S. Systemic effects of periodontitis: epidemiology of periodontal disease and cardiovascular disease. J Periodontol. 2005;76(11 Suppl):2089–2100. doi: 10.1902/jop.2005.76.11-S.2089. [DOI] [PubMed] [Google Scholar]
- Spellerberg B, Brandt C: Streptococcus. In Manual of clinical microbiology. Volume 1. 10th edition. Edited by Versalovic J, Carroll KC, Funke G, Jorgensen JH, Landry ML, Warnock DW. Washington, DC: ASM Press; 2011:331–349.
- Zbinden A, Mueller NJ, Tarr PE, Sproer C, Keller PM, Bloemberg G: Streptococcus tigurinussp. nov., isolated from blood of patients with endocarditis, meningitis and spondylodiscitis.Int J Syst Evol Microbiol 2012, 62(Pt 12):2941–2945. [DOI] [PubMed]
- Zbinden A, Mueller NJ, Tarr PE, Eich G, Schulthess B, Bahlmann AS, Keller PM, Bloemberg GV: Streptococcus tigurinus, a novel member of theStreptococcus mitisgroup, causes invasive infections.J Clin Microbiol 2012, 50(9):2969–2973. [DOI] [PMC free article] [PubMed]
- Zbinden A, Quiblier C, Hernandez D, Herzog K, Bodler P, Senn MM, Gizard Y, Schrenzel J, François P: Characterization ofStreptococcus tigurinussmall-colony variants causing prosthetic joint infection by comparative whole-genome analyses.J Clin Microbiol 2014, 52(2):467–474. [DOI] [PMC free article] [PubMed]
- Veloso TR, Zbinden A, Andreoni F, Giddey M, Vouillamoz J, Moreillon P, Zinkernagel AS, Entenza JM: Streptococcus tigurinusis highly virulent in a rat model of experimental endocarditis.Int J Med Microbiol 2013, 303(8):498–504. [DOI] [PubMed]
- Lo Frisco C, Cutler R, Bramson JB. Periodontal screening and recording: perceptions and effects on practice. J Am Dent Assoc. 1993;124(7):226–229. doi: 10.14219/jada.archive.1993.0256. 231–232. [DOI] [PubMed] [Google Scholar]
- Cutress TW, Ainamo J, Sardo-Infirri J. The community periodontal index of treatment needs (CPITN) procedure for population groups and individuals. Int Dent J. 1987;37(4):222–233. [PubMed] [Google Scholar]
- Belibasakis GN, Schmidlin PR, Sahrmann P. Molecular microbiological evaluation of subgingival biofilm sampling by paper point and curette. APMIS. 2014;122(4):347–352. doi: 10.1111/apm.12151. [DOI] [PubMed] [Google Scholar]
- Deshpande RG, Khan M, Genco CA: Invasion strategies of the oral pathogenPorphyromonas gingivalis: implications for cardiovascular disease.Invasion Metastasis 1998, 18(2):57–69. [DOI] [PubMed]
- Kinane DF, Bartold PM. Clinical relevance of the host responses of periodontitis. Periodontol 2000. 2007;43:278–293. doi: 10.1111/j.1600-0757.2006.00169.x. [DOI] [PubMed] [Google Scholar]
- Hamedi M, Belibasakis GN, Cruchley AT, Rangarajan M, Curtis MA, Bostanci N: Porphyromonas gingivalisculture supernatants differentially regulate interleukin-1beta and interleukin-18 in human monocytic cells.Cytokine 2009, 45(2):99–104. [DOI] [PubMed]
- Li J, Helmerhorst EJ, Leone CW, Troxler RF, Yaskell T, Haffajee AD, Socransky SS, Oppenheim FG. Identification of early microbial colonizers in human dental biofilm. J Appl Microbiol. 2004;97(6):1311–1318. doi: 10.1111/j.1365-2672.2004.02420.x. [DOI] [PubMed] [Google Scholar]
- Sliepen I, Van Damme J, Van Essche M, Loozen G, Quirynen M, Teughels W. Microbial interactions influence inflammatory host cell responses. J Dent Res. 2009;88(11):1026–1030. doi: 10.1177/0022034509347296. [DOI] [PubMed] [Google Scholar]
- Ennibi OK, Benrachadi L, Bouziane A, Haubek D, Poulsen K: The highly leukotoxic JP2 clone ofAggregatibacter actinomycetemcomitansin localized and generalized forms of aggressive periodontitis.Acta Odontol Scand 2009, 70(4):318–322. [DOI] [PubMed]
- Gizard Y, Zbinden A, Schrenzel J, François P: Whole-genome sequences ofStreptococcus tigurinustype strain AZ_3a andS. tigurinus1366, a strain causing prosthetic joint infection.Genome Announc 2013, 1(2):e00210–e00212. [DOI] [PMC free article] [PubMed]
- Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner AC, Yu WH, Lakshmanan A, Wade WG. The human oral microbiome. J Bacteriol. 2010;192(19):5002–5017. doi: 10.1128/JB.00542-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belibasakis GN, Ozturk VO, Emingil G, Bostanci N: Synergistetescluster A in saliva is associated with periodontitis.J Periodontal Res 2013, 48(6):727–732. [DOI] [PubMed]


