Abstract
TGF-β family signaling through Smads is conceptually a simple and linear signaling pathway, driven by sequential phosphorylation, with type II receptors activating type I receptors, which in turn activate R-Smads. Nevertheless, TGF-β family proteins induce highly complex programs of gene expression responses that are extensively regulated, and depend on the physiological context of the cells. Regulation of TGF-β signaling occurs at multiple levels, including TGF-β activation, formation, activation and destruction of functional TGF-β receptor complexes, activation and degradation of Smads, and formation of Smad transcription complexes at regulatory gene sequences that cooperate with a diverse set of DNA binding transcription factors and coregulators. Here we discuss recent insights into the roles of post-translational modifications and molecular interaction networks in the functions of receptors and Smads in TGF-β signal responses. These layers of regulation demonstrate how a simple signaling system can be coopted to exert exquisitely regulated, complex responses.
Keywords: Transforming growth factor-β, signal transduction, Smad, receptor kinase, post-translational modification, phosphorylation, ubiquitylation, sumoylation, acetylation, ADP-ribosylation, ectodomain shedding, miRNA, nucleocytoplasmic shuttling, signaling crosstalk
Introduction
The TGF-β family of secreted proteins is encoded by 33 genes for structurally related polypeptides marked by a signal peptide, a large pro-domain and a C-terminal mature polypeptide that is characterized by its spacing of seven or nine cysteines (1). The mature polypeptides form disulfide-bonded dimers, which have primarily been studied as homodimers, yet also form physiological heterodimers with potent biological activities. Among the TGF-β family proteins, TGF-β1 has served as preferred model and prototype to study mechanisms of ligand-induced receptor activation and signaling mechanisms that are activated by the TGF-β family proteins.
TGF-β family proteins are involved in a large variety of processes in all metazoans, in both development and tissue physiology, and the deregulation of their activities is at the basis of many diseases. Most TGF-β proteins regulate cell proliferation and cell differentiation programs. Additionally, activins, bone morphogenetic proteins (BMPs) and growth and differentiation factors (GDFs) are perhaps best known for their key roles in cell and tissue differentiation, and morphogenesis. Activins also play major roles in hormonal control and homeostasis, and TGF-β controls differentiation and function of hematopoietic and immune cells. While deregulation of BMP signaling can lead to many developmental syndromes, increased TGF-β1 expression and activities are known to drive cancer progression and fibrosis in many tissues. These and other observations illustrate the pervasive involvement of TGF-β family proteins in a plethora of biological processes and physiological contexts (2).
TGF-β signaling through cell surface receptor complexes and Smads
TGF-β family proteins initiate intracellular signaling by binding to tetrameric cell surface complexes of two pairs of transmembrane kinases, the type II and the type I receptors (Fig. 1). Cytoplasmic receptor phosphorylation upon TGF-β ligand binding then provides the basis for signal transduction by TGF-β proteins. Viewing TGF-β as prototype, ligand binding stabilizes the interaction of TβRII dimers with two TβRI molecules, enabling the TβRII kinases to phosphorylate the short juxtamembrane, glycine-and serine-rich GS domains of the TβRI receptors on serines and threonines (3). Their phosphorylation then triggers conformational changes in TβRI that activate the TβRI receptor kinase (4), concomitantly with the release of the immunophilin inhibitor FKBP12 that associates with the unliganded TβRI receptor and helps maintain it in a silenced state (4, 5). The phosphorylated GS domain of the activated TβRI also provides the type I receptors with a surface to interact with the basic patch of the MH2 domains of Smad2 and Smad3 (4, 6), which will serve as signaling effectors of TGF-β. The Smad-TβRI interaction, presumably in a 2:2 stoichiometry, allows the TβRI kinase to phosphorylate the Smad on two serines within its conserved C-terminal -SSXS motif. Upon phosphorylation, the charged C-terminal sequence of the receptor-activated Smad (R-Smad) can then intramolecularly interact with its basic patch, thus weakening R-Smad binding to TβRI, and enabling the release of the activated R-Smad from TβRI (6, 7). A similar mechanism may allow BMP-induced activation of Smad1 and Smad5 by BMP type I receptors. Binding of TGF-β or TGF-β-related factors to their cognate receptor complexes also results in activation of non-Smad signaling pathways, such as MAP kinase pathways and the PI3K-Akt-TOR pathway (8). Activation of these non-Smad signaling pathways by TGF-β will not be discussed.
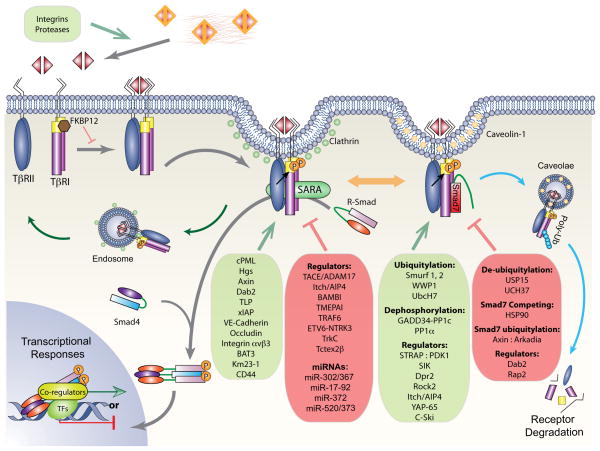
Figure 1. Regulation of the TGF-β receptors.
The TGF-β signaling pathway is outlined in thick gray lines, whereas the endosomal recycling pathway and lipid rafts-caveolae degradation pathway are indicated by green and blue lines, respectively. Binding of ligand stabilizes the heteromeric complex of TβRII and TβRI receptors, resulting in activation of TβRI through phosphorylation of its GS domain by TβRII. R-Smads are then activated by TβRI with SARA as scaffold, and form trimeric complexes with Smad4 that translocate into the nucleus, where they direct transcription responses of target genes. Activated receptor complexes are also internalized through lipid rafts and caveolae, where they are poly-ubiquitylated by E3 ubiquitin ligases recruited by Smad7 and destined for degradation. Various regulators of functional TGF-β receptor complexes are schematically shown. Inhibitory mechanisms are listed in a red box with blunt-headed lines, whereas those that enhance the complexes formation are listed in a green box with arrows.
Subsequently partnered with one Smad4 molecule, two activated R-Smads rapidly translocate into the nucleus as trimeric complexes, where they elicit transcription responses on target genes by associating with high affinity DNA binding transcription factors and conducive Smad binding DNA sequences at regulatory promoter sequences (Fig. 2). Activated Smad complexes have been shown to associate with, and regulate the activities of, a large variety of DNA binding transcription factors. Additional recruitment of coactivators and corepressors further defines the TGF-β-induced transcription activation or repression of target genes. The association of R-Smads with the coactivators CBP or p300 often allows for enhanced transcription activation, while also conferring histone acetylation. Conversely, R-Smads have also been shown to recruit histone deacetylases, leading to transcription repression and histone deacetylation. The interactions of the activated Smad complexes with divergent DNA-binding transcription factors, coactivators and corepressors allow for multiple types of functional crosstalk between TGF-β/Smad signaling and other signaling pathways, since many of the Smad interaction partners are usually also targeted by signaling pathways (9, 10).
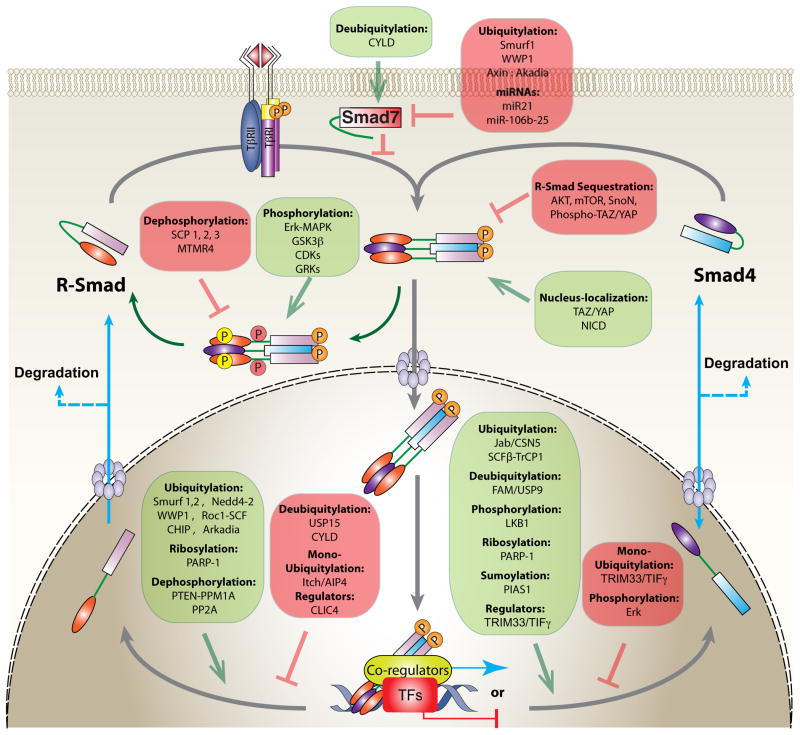
Figure 2. Regulation of Smad signaling.
Thick gray lines illustrate Smad signaling, whereas nucleocytoplasmic shuttling and degradation of Smads are indicated by solid and dashed light-blue lines, respectively. Activation of R-Smads by TβRI facilitates the formation of trimeric R-Smad/Smad4 complexes, which are imported into the nucleus to regulate transcription of target genes. R-Smad phosphorylation at various sites besides the C-terminus prevents nuclear import of the complexes, or favors degradation of the complexes, resulting in termination of signaling. Diverse regulators modulate the activated Smad complexes inside the nucleus, and control the balance between complex formation and dissociation, thus determining the overall signaling outcome in a context-dependent manner. Inhibitory mechanisms are listed in a red box with blunt-headed lines, and those that promote the indicated processes are listed in a green box with arrows.
We will summarize recent insights into the direct regulation of the TGF-β receptors (Fig. 1) and Smads (Fig. 2) by post-translational modifications, often resulting from signaling crosstalk. We will not elaborate on the signaling crosstalk that targets the many transcription factors that associate with Smads at the DNA, and thus regulates the transcriptional cooperativity.
Activation of latent TGF-β regulates TGF-β receptor signaling
The three TGF-βs, TGF-β1, -β2 and -β3 are secreted by most cell types as latent complexes, consisting of the mature TGF-β dimer in a non-covalent association with a dimer of the large pro-peptide, i.e. the sequence preceding the mature monomer in the TGF-β precursor, and, covalently attached to this pro-peptide, a latent TGF-β binding protein (LTBP). The latency of the TGF-β complex allows for storage of TGF-β in association with extracellular matrix proteins without inadvertent activation of TGF-β signaling, and requires an activation process to release biologically active TGF-β that only then can bind and activate the TGF-β receptors (11, 12). Various mechanisms lead to latent TGF-β activation, with the mode of TGF-β activation likely dependent on the cell type and physiological context. Generally, activation of TGF-β occurs through the actions of proteases, such as matrix metalloproteases (MMPs), plasmin, and thrombin that target LTBP, allowing release from the latent complexes in the extracellular matrix (11, 12).
Increasing evidence highlights key roles of integrins, specifically αvβ6 and αvβ8 dimers, in latent TGF-β activation and TGF-β signaling (13). The pro-region of TGF-β1 comprises an RGD sequence that is recognized by several integrin combinations, and this interaction is thought to be essential for integrin-mediated activation of TGF-β. Mice that express the TGF-β1 precursor with an inactivated RGD sequence show a phenotype that strongly resembles that of TGF-β1 null mice (14), highlighting the roles of integrins in TGF-β1 activation. Consistent with this notion, conditional loss of integrin αvβ8 expression in dendritic cells causes severe inflammatory bowel disease and age-related autoimmunity in mice, presumably resulting form the absence of TGF-β activity and failure of regulatory T cell induction (15, 16).
Integrin αvβ8-mediated activation of TGF-β appears to involve proteolytic cleavage, e.g. through the metalloproteases MT1-MMP, also called MMP14 (17). In contrast, αvβ6-mediated activation of TGF-β1 may occur without participation of a protease and involve a conformation change that results in TGF-β activation (18, 19). Furthermore, myofibroblast contraction leads to activation of TGF-β1 from extracellular matrix stores, through a process that requires integrins (19). Additionally, TGF-β released from platelets or fibroblasts is activated in response to shear stress (20), illustrating the importance of force-catalyzed release of TGF-β ligands.
Regulation of TGF-β receptor signaling by microRNAs
MicroRNAs (miRNAs) are small non-coding RNAs that specifically bind target mRNAs and inhibit mRNA translation or promote mRNA degradation (21). MicroRNAs are increasingly seen as dynamic regulators of signaling pathways, by controlling levels of key components of these pathways (22). Several miRNAs directly target the mRNAs encoding TβRII, or the type II Nodal receptor. These include the miR-302/367 cluster, miR-372, miR-520/373, miR-17–92 cluster, miR-15, and miR-16 (23–27). Since they are capable of inhibiting TGF-β receptor expression, and the receptor levels correlate with TGF-β responsiveness (28, 29), these miRNAs control the threshold for signaling initiation in response to TGF-β or TGF-β-related factors (23–27). Additional components for each step of the TGF-β signaling cascade may also be modulated by specific miRNAs, raising the possibility of a previously unanticipated, complex and context-dependent miRNA network controlling TGF-β signaling.
Control of TGF-β receptor availability through ectodomain shedding
The cell surface levels of TGF-β receptors, and, consequently, the sensitivity of cells to TGF-β, are controlled by ectodomain shedding. TACE, a transmembrane ADAM metalloproteinase also known as ADAM17, proteolytically removes the ectodomain of TβRI, but not TβRII (29), and this process is facilitated by TRAF6-mediated TβRI ubiquitylation (30). TACE-mediated ectodomain shedding is activated in response to ERK or p38 MAPK pathway signaling (29, 31). Activation of either pathway by growth factors or inflammatory signals consequently decreases cell surface levels of TβRI, but not TβRII, thus attenuating TGF-β-induced Smad3 activation without affecting TGF-β binding to the cell surface TβRII receptors (29). Conversely, inhibition of TACE increases the cell surface TβRI level, Smad3 activation, and TGF-β-induced cytostasis and epithelial-mesenchymal transition (29). Downregulation of TGF-β signaling through TACE activation in response to growth stimulatory or inflammatory cues provides a strategy for evasion of autocrine tumor suppression by TGF-β, and for modulation of epithelial-mesenchymal transition in cancer progression (29, 31). Upon release of the TβRI ectodomain, the TβRI cytoplasmic domain can be released from the plasma membrane, translocates into the nucleus, and then acts as a transcription factor in potentiating the expression of Snail and MMP2 (30).
Another ADAM metalloproteinase, ADAM12, was shown to form a complex with the type II TGF-β receptor, and facilitate activation of TGF-β signaling through a mechanism independent of its protease activity. ADAM12 also induces TβRII accumulation in early endosomal vesicles and stabilizes TβRII, and was proposed to do so by suppressing the association of TβRII with Smad7 (32).
TGF-β receptor phosphorylation and dephosphorylation
Receptor phosphorylation provides the basis for signal transduction in response to TGF-β. TGF-β binding stabilizes the TβRII-TβRI interactions, enabling TβRII to phosphorylate the GS domain of TβRI on serines and threonines. The resulting conformational change in TβRI then activates the TβRI kinase, which then initiates Smad and non-Smad signaling (4). TGF-β binding to the receptor complex also induces TβRI phosphorylation outside its GS domain, presumably by TβRII and through autophosphorylation upon TβRI activation. The identities, functions, and mechanism of phosphorylation of these sites remain to be defined (33). The phosphorylation status of TβRII also remains largely uncharacterized, and TβRII phosphorylation may similarly combine autophosphorylation and phosphorylation by TβRI. Whether additional kinases can phosphorylate TβRI or TβRII on serine or threonine also remains to be determined.
The TβRI and TβRII receptors are additionally phosphorylated on tyrosines. TβRII and TβRI are both dual specificity kinases, which are characterized by strong serine and threonine but weaker tyrosine phosphorylation capacities (34). Their dual specificities are consistent with structural features in their sequences that phylogenetically place TβRII, TβRI and the other type I and type II TGF-β family receptors between tyrosine and serine-threonine kinases, and close to other dual specificity kinases (35). TβRII phosphorylation on tyrosine may result from autophosphorylation (36), although TβRII can also be phosphorylated by the tyrosine kinase Src (37). Phosphorylation by Src enables TβRII to recruit Grb2 and Shc, contributing to the TGF-β-induced activation of p38 MAPK (37). TGF-β also induces tyrosine phosphorylation of TβRI, resulting from autophosphorylation or mediated by TβRII (38). TβRI phosphorylation on Tyr may play a role in TGF-β-induced activation of Erk MAPK signaling, which is initiated by Shc association with TβRI through its phosphoTyr-binding SH2 and PTB domains residues, and subsequent Shc phosphorylation on Tyr by TβRI (38).
The activation state of the TGF-β receptors is also regulated by dephosphorylation (39). Smad7 was shown to interact with GADD34, a regulatory subunit of the protein phosphatase 1 (PP1) holoenzyme, and thus recruit the catalytic subunit of PP1 (PP1c) to TβRI, resulting in TβRI dephosphorylation and inhibition of TGF-β-induced cell cycle arrest (40). Additionally, SARA, an endosomal adaptor protein that associates with TβRI, may also target PP1c to Drosophila receptor complexes that are activated by the BMP ortholog Dpp. In this way, PP1c can attenuate Dpp signaling through dephosphorylation of the type I receptor, Tkv (41). Finally, regulatory subunits of protein phosphatase 2A (PP2A) also associate with TGF-β receptors and regulate their activity (42, 43). Surprisingly, the regulatory Bα and Bδ subunits modulate TGF-β/Activin/Nodal signaling in opposite ways. The Bα subunit of PP2A enhances the cytostasis effect of TGF-β (42, 43), whereas Bδ is thought to negatively modulate these responses by restricting the receptor activity (42).
Ubiquitylation and sumoylation of the TGF-β receptor
Ubiquitylation, i.e. the covalent attachment of ubiquitin polypeptides, and ubiquitin-mediated protein degradation define the levels and turnover of proteins in a wide range of processes. Ubiquitylation results from sequential actions of E1, E2 and E3 ubiquitin ligases that provide substrate specificity (44). The stability and levels of TGF-β receptor complexes are determined by ubiquitylation. Specifically, Smad7 recruits, through direct association, an E3 ubiquitin ligase to TβRI, resulting in TβRI ubiquitylation and degradation (45). Multiple E3 ligases were shown to be involved in TβRI ubiquitylation, including the HECT-type ligases Smurf1, Smurf2, WWP1, and NEDD4-2, which associates with the E2 ligase UbcH7 and an N-terminal sequence in Smad7 (45–48). The contributions of the individual E3 ligases to the regulation of the TGF-β receptors in vivo remain to be fully characterized.
Conversely, the deubiquitylating enzyme USP15 was shown to associate with the Smad7-Smurf2 complex, and to deubiquitylate and stabilize TβRI, leading to enhanced TGF-β signaling in glioblastoma, breast and ovarian cancer (49). Similarly to USP15, another deubiquitylating enzyme, UCH37, forms a complex with Smad7 by binding to a sequence that is distinct from the PY motif with which Smurf1 or Smurf2 interact, and also deubiquitylates and stabilizes TβRI (50). Finally, the 90-kD heat shock protein HSP90 was reported to associate with TβRI and TβRII, and thus prevent Smurf2-mediated receptor ubiquitylation and degradation (51).
Through a sequential process similar to ubiquitylation, sumoylation covalently attaches a SUMO polypeptide that resembles ubiquitin to its substrate. Sumoylation does not lead to degradation of the targeted protein, occurs primarily on perinuclear and nuclear proteins, including transcription factors, and often regulates subcellular localization and functions (44). The TβRI receptor is sumoylated at one defined lysine residue in response to TGF-β. TβRI sumoylation requires the kinase activities of TβRI and TβRII, indicating that phosphorylation and possibly a conformational change of TβRI are required for docking of the E3 ligase. Sumoylation of TβRI stabilizes the Smad2/3 binding to the TβRI receptor, leading to enhanced Smad activation. Accordingly, lack of TβRI sumoylation decreases TGF-β-induced Smad2 and Smad3 activation, and TGF-β-induced transcription of target genes (52).
Endocytic routing of receptors
The endocytic routing of TGF-β receptors relies on constitutive receptor internalization and recycling through either clathrin- or caveolin-1-mediated endocytosis (53–55). Through a di-leucine internalization motif in the TβRII cytoplasmic domain, TGF-β receptor complexes are internalized in clathrin-coated pits, and enter EEA-1- and Rab5-positive early endosomes, which are recycled to the cell surface in a manner that depends on the GTPase Rab11 (54, 56). In clathrin-coated pits, the association of the adaptor protein β2-adaptin with the cytoplasmic domain of TβRII couples the TGF-β receptor complexes to the AP-2 subunit of clathrin (57). TGF-β receptor internalization through clathrin-coated endosomes is thought to be essential for Smad activation, since inhibition of this endocytosis-recycling pathway prevents Smad activation (54, 56). TβRII, TβRI and the adaptor protein SARA, which stabilizes the TβRI-Smad interaction, are enriched in EEA-1-positive early endosomes, and this localization is required for Smad2/3 activation (58). RIN1, a guanine nucleotide exchange factor of Rab5, promotes TGF-β/Smad signaling by enhancing endocytosis (59).
In contrast to clathrin-mediated endocytosis, caveolin-1- and lipid-raft-dependent endocytosis is thought to direct the TGF-β receptors to degradation. Caveolin-1 interacts with TβRI as well as Smad7 and Smurf2, which ubiquitylates TβRI, and inhibits TGF-β-induced Smad signaling (54) (60). Silencing the expression of caveolin-1 enhances the half-life of the TGF-β receptors and the responsiveness of T-cells to TGF-β stimulation, whereas caveolin-1 overexpression suppresses TGF-β-induced Smad signaling (61). Other findings, however, argue for TGF-β-induced Smad activation in both lipid rafts and non-lipid rafts, and for receptor degradation in clathrin-mediated endocytosis (56). Therefore, the exclusive association of TGF-β-induced Smad activation and TGF-β receptor degradation with clathrin-mediated versus caveolin-1-mediated endocytosis, respectively, may not be unambiguous as sometimes believed. Interestingly, TGF-β-induced MAPK activation may require the presence of TGF-β receptors in lipid rafts (62). Distinct internalization routes may confer different signaling capabilities of TGF-β receptor.
In addition to Rab5 and Rab11, other GTPases have been implicated in directing TGF-β receptor endocytosis and recycling (63). For instance, Rap2 competes with Smad7 for binding to TβRI, and directs internalized receptor complexes to the recycling pathway to prevent their degradation, thus delaying receptor turnover, and contributing to upregulation of Smad activation (64).
Other TGF-β receptor interacting proteins
Since the identification of the TβRI and TβRII receptors, a steadily increasing number of proteins were shown to associate with TGF-β receptors and function as modulators of TGF-β signaling (Fig. 1). This diversity of interacting proteins reflects the dynamic changes in the life cycle of the receptors, ranging from maturation and cell surface transport of newly synthesized TβRI and TβRII, to defining their subcellular localization, cell surface presentation and ligand binding, differential endosomal and intracellular routing, Smad accessibility and differential activation of Smad and non-Smad pathways, and receptor stability and degradation. These proteins are listed in Tables 1 and 2 with a brief description of their actions. The extensive and complex regulation of the TGF-β receptors through protein associations and post-translational modifications allows for a highly regulated and finely tuned receptor signaling system that defines the TGF-β responsiveness of the cells and differential regulation of TGF-β responses (Fig. 1).
Table 1.
TGF-β receptor-interacting proteins that potentiate TGF-β signaling
| Interacting Protein | Regulatory Mechanism | References |
|---|---|---|
| SARA | Stabilizing R-Smads and TGF-β receptor association; Directing the activated receptor towards clathrin-mediated endocytic routing |
(58, 165, 166) |
| Hgs | Cooperating with SARA to stabilize TGF-β receptor:R-Smad association | (167) |
| cPML | Interacting with R-Smads and SARA to improve their association; Improving TGF-β receptor and SARA accumulation in early endosomes |
(168) |
| Axin | Adapter of Smad3 to facilitate its activation by TβRI; Forming a multimeric complex consisting of Smad7 and Arkadia to degrade Smad7 |
(74, 145, 153) |
| Dab2 | Associating with both TGF-β receptors and R-Smads to facilitate R-Smad activation; Promoting clathrin-mediated recycling of TβRII to protect it from degradation |
(169, 170) |
| TβRIII/betaglycan | Mediating Smad-independent activation of TRAF6-TAK1-p38 MAPK with TβRI | (171) |
| VE-cadherin | Enhancing TβRII:TβRI assembly into an active receptor complex to improve R-Smad phosphorylation. | (172) |
| Occludin | Regulating TβRI localization to enhance Smad activation | (173) |
| TRAF6 | Mediating the activation of p38 and JNK MAPK by ubiqitylation and phosphorylation of TAK1; Inhibiting TGF-β-induced R-Smad activation and IL-2 blockage in T cells |
(174–176) |
| Integrin αvβ3 | Interacting with TβRII to potentiate TGF-β1 activation | (177) |
| HSP90 | Binding to TβRI and TβRII to prevent Smurf2-mediated receptor ubiquitylation and degradation | (51) |
| Itch/AIP4 | Facilitating TGF-β receptor-Smad2 complex formation and enhancing TGF-β−induced transcription | (91) |
| CD44 | Associating with TβRI to promote Smad2/3 activation and PTH-rP expression | (178) |
| Mgat5 | Modifying N-glycans on TGF-β receptors to delay their removal at the cell surface by constitutive endocytosis | (179) |
| TLP | Associating with TβRI and TβRII to regulate the balance of Smad2 and Smad3 signaling by localizing Smad4 intracellularly | (180) |
| BAT3 | Interacting with TβRI and TβRII to potentiate Smad activation | (181) |
| km23-1 | Associating with TGF-β receptors and Smad2 in early endosomes to potentiate Smad activation | (182) |
| xIAP | Interacting with TβRI, mediating the activation of NF-κB by ubiquitiylation of TAK1, and coupling of Smad2/3 activation | (183) |
Table 2.
TGF-β receptor-interacting proteins that attenuate TGF-β signaling
| Interacting Protein | Regulatory Mechanism | References |
|---|---|---|
| Smad7 | Competing with recruitment of R-Smads to TβRI receptor, and preventing R-Smad activation; Recruiting ubiqitin E3 ligases such as Smurf2, WWP1 and SIK to TβRI receptor to degrade receptors; Recruiting subunit of phosphatase such as GADD34-PP1c to dephosphorylate TβRI |
(40, 45, 48, 133, 184) |
| FKBP12 | Binding to GS domain of TβRI to prevent leaky signaling; Interacting with Smad7 and TβRI to form a complex with Smurf1 to degrade TβRI |
(4, 185) |
| TMEPAI | Interacting with R-Smads to compete for association with the TβRI receptor | (186) |
| Dpr2 | Binding to TβRI receptor to promote its lysosomal degradation | (187, 188) |
| Rock2 | Binding to TβRI and accelerating its lysosomal degradation | (189) |
| STRAP | Interacting with TβRI and TβRII receptors and stabilizing Smad7:receptor complexes | (190) |
| SIK | Associating with Smad7, TβRI, and Smurf2 to form a complex to degrade TβRI | (184, 191) |
| PDK1 | Interacting with STRAP and enhancing STRAP-induced inhibition | (192) |
| BAMBI | Interfering with TβRI and TβRII complex formation, and forming a ternary complex with activated TβRI and Smad7 to prevent Smad association | (193, 194) |
| ALK1 | Associating with TβRI and TβRII to form a complex and antagonize Smad2/3 activation | (195) |
| Itch/AIP4 | Enhancing TβRI:Smad7 association and inhibiting Smad activation | (196) |
| UbcH7 | Serving as ubiquitin-conjugating enzyme (E2) to facilitate TβRI degradation by Smurf2 | (46) |
| YAP-65 | Augmenting the association of Smad7 to activated TβRI and inhibiting Smad activation | (197) |
| c-Ski | Associating with activated TβRI, resulting in constitutive association of TβRI with nonfunctional R-Smad/Smad4 complexes | (198) |
| TrkC | Binding to TβRII and preventing it from interacting with TβRI, and blocking Smad activation | (199) |
| Tctex2β | Interacting with TβRII and inhibiting TGF-β signaling | (200) |
| ETV6-NTRK3 | Directly binding to TβRII and preventing it from interacting with TβRI | (201) |
Regulation of receptor-activated Smad function by phosphorylation
Among the eight Smads in vertebrates, Smad2 and Smad3 are activated in response to TGF-β nodals, and activins, and Smad1, Smad5 and Smad8 respond to BMPs and GDFs (65). Once activated by type I receptors, R-Smads form complexes of two R-Smads and one Smad4, and translocate into the nucleus where they activate or repress transcription of target genes (Figs. 1 and 2). These Smads are characterized by a highly conserved N-terminal MH1 and C-terminal MH2 domain, separated by a variable serine and proline-rich linker region (2, 6).
R-Smad activation results from ligand-induced phosphorylation of the C-terminal two serines in their –SSXS motif by type I receptors. However, the linker region can also be phosphorylated on Ser or Thr by other kinases, e.g. Erk MAPK and CDK kinases, which consequently control Smad activities (8, 10). Considering the flexible nature of the Smad linker domains, different patterns of linker phosphorylation are likely to impose conformation differences that then affect protein interactions and Smad functions and stability. Recent insights illustrate an extensive control of Smad function by linker phosphorylation. For less recent findings, we refer to an excellent review on regulation of Smad function by phosphorylation (39).
Besides the rapid activation of Smad3 through C-terminal phosphorylation, TGF-β induces linker phosphorylation at three sites in Smad2 and Smad3, albeit with slower kinetics. This phosphorylation is mediated by GSK3, also a downstream effector of Wnt signaling, or CDK8 and CDK9 (66–68), and marks activated Smad2 or Smad3 for proteasomal destruction. The E3 ubiquitin ligase Nedd4L then recognizes the phosphoT-P-Y motif in the linker, leading to Smad2/3 polyubiquitylation and degradation (69), limiting the half-life of activated Smads and, consequently, the amplitude and duration of TGF-β responses. CDK8 and CDK9 also phosphorylate the linker region of Smad1, resulting in recruitment of YAP, an effector of the Hippo pathway of cell size control. This interaction enhances Smad-mediated transcription, although the phosphorylated linker is ultimately recognized by the E3 ubiquitin ligase Smurf1, leading to Smad degradation (68, 70). The duration of Smad1 signaling is also regulated by sequential linker phosphorylation at conserved MAPK and GSK3 sites, leading to polyubiquitylation, and eventual transport to the centrosome for proteasomal degradation (71). Because Smad linker phosphorylation by CDK8 or CDK9, and/or GSK3, results in recruitment of YAP and the peptidyl-prolyl isomerase Pin1 for regulating transcription activity (70, 72), or Nedd4L or Smurf1 for R-Smad destruction (68), both processes are coordinately regulated. The functional switch from initial Smad activation to subsequent destruction appears to result from a switch in recognition of Smad phosphoserines by WW domains of both transcription factors and E3 ubiquitylating ligases (70). For example, in the TGF-β pathway, Smad3 phosphorylation by CDK8/9 creates binding sites for WW domains of Pin1, and subsequent phosphorylation by GSK3 switches off Pin1 binding, and adds binding sites for Smurf1 WW domains (70).
GRK2, a kinase involved in desensitizing G protein-coupled receptors, also associates with R-Smads (73). Phosphorylation of R-Smad linker regions by GRK2 somehow inhibits ligand-induced C-terminal phosphorylation, and prevents nuclear translocation of Smad complexes, thus inhibiting TGF-β-induced target gene expression. In this way, the induction of GRK2 expression in response to TGF-β provides negative feedback to control TGF-β responses. Non-activated Smad3 can also be phosphorylated in its MH1 domain by GSK3-β in association with the scaffolding protein Axin. This phosphorylation results in Smad3 ubiquitylation and degradation, and helps define the basal Smad3 level and activity (74).
Finally, it has been reported that the C-terminal -SSXS motif of R-Smads can be targeted by kinases that are distinct from the type I TGF-β family receptors. For example, Mps1, a kinase that regulates mitotic progression, can C-terminally phosphorylate and thus activate Smad2 and Smad3 (75). Additionally, the WNK family kinase WNK1 (76) and MPK38 kinase (77) also directly activate Smad2 and Smad3. The biological significance of Smad activation by these kinases needs to be further explored.
Dephosphorylation of receptor-activated Smads
Dephosphorylation of the C-terminal serines regulates the termination of Smad signaling. Among many phosphatases evaluated, PPM1A/PP2Cα acts as Smad phosphatase for Smad2 and Smad3. PPMA1 removes the receptor-mediated phosphorylation at the -SSXS motif, and promotes nuclear export of TGF-β-activated Smad2/3, thereby terminating TGF-β signaling (78). The phosphatase PTEN, a negative regulator of the PI3K/Akt pathway, remarkably serves as co-factor of PPM1A, and helps abrogate Smad2/3 phosphorylation by stabilizing PPM1A (79). The phosphatase MTMR4 was also found to C-terminally dephosphorylate the activated Smad2 and Smad3 in endosomes (80). Additionally, SCP phosphatases remove the C-terminal phosphorylation of Smad1 and thus attenuate BMP signaling (81). Under hypoxic conditions, the phosphatase PP2A was shown to C-terminally dephosphorylate Smad3, but not Smad2, suggesting a mechanism by which hypoxia regulates growth factor responses (82). Activated Smad2 or Smad3 dephosphorylation can be prevented by CLIC4, a multifunctional protein that shuttles between the cytoplasm and nucleus. Nuclear CLIC4 associates with activated Smad2 and Smad3, and protects them from dephosphorylation (83). Finally, the phosphatases SCP1, SCP2 and SCP3 remove linker phosphorylation at certain sites, without affecting C-terminal phosphorylation, of Smad2/3, thus enhancing TGF-β signaling (84, 85).
Ubiquitylation of R-Smads
TGF-β-induced Smad2 activation was shown to be followed by poly-ubiquitylation and proteasomal degradation, thus targeting nuclear Smad2 for degradation, and terminating its signaling (86). Since this initial observation, multiple ubiquitin ligases have been implicated in R-Smad degradation, including Smurf1, Smurf2, Nedd4-2, WWP1, ROC1-SCF, and CHIP (87). As mentioned, TGF-β-induced linker phosphorylation marks activated Smads for proteasomal destruction, and this involves the E3 ubiquitin ligase Nedd4L (69). Therefore, Nedd4L limits the half-life of TGF-β-activated Smads and restricts the intensity and duration of TGF-β signaling. The E3 ubiquitin ligase Arkadia, which was initially shown to ubiquitylate Smad7 and the Smad corepressor SnoN, and target them for degradation (88), also ubiquitylates activated R-Smads, causing their degradation (89). Additionally, the estrogen receptor was shown to form a complex with Smads, facilitating estrogen-dependent recruitment of Smurf1 for subsequent R-Smad degradation (90).
In contrast to poly-ubiquitylation, which leads to degradation, mono-ubiquitylation has different effects on Smad2 and Smad3 functions. The HECT-domain E3 ligase Itch/AIP4 can form a complex with Smad2 and activated TβRI, resulting in Smad2 mono-ubiquitylation, which in turn enhances the Smad2 interaction with the receptor, and Smad2 activation (91). Smad3 was shown to be mono-ubiquitylated in its MH1 domain at multiple lysines, and these modifications interfere with Smad3 binding to regulatory promoter sequences (92). The de-ubiquitylating enzyme USP15 opposes the MH1 domain monoubiquitylation, and thus enhances the occupancy of target promoters by Smad complexes (92), suggesting a dynamic balance of mono-ubiquitylation and de-ubiquitylation in the control of R-Smad activities. Consequently, silencing USP15 expression abolishes the recruitment of TGF-β-activated Smad complexes to regulatory DNA sequences. Smad3 is additionally mono-ubiquitylated at four lysine residues in the MH2 domain through the action of the E3 ligase Smurf2 (93). Rather than controlling Smad3 stability through poly-ubiquitylation, Smurf2-mediated mono-ubiquitylation interferes with the formation of functional Smad3 complexes. This mono-ubiquitylation requires prior TGF-β-induced phosphorylation of Thr179 and adjacent PY motif in the linker region by CDKs. Finally, the deubiquitylase CYLD inhibits TGF-β signaling by decreasing the stability of Smad3. This effect, however results from Akt de-ubiquitylation that relieves Akt-mediated inhibition of GSK3β-CHIP–induced degradation of Smad3 (94).
Acetylation and ADP-ribosylation of R-Smads
Resulting from their TGF-β-induced association in nucleoprotein complexes, Smad2 and Smad3 are acetylated by the coactivators CBP and p300 in response to TGF-β. Smad3 is acetylated in its MH2 domain, whereas Smad2 is primarily acetylated in its MH1 domain, yet both acetylations enhance Smad-mediated transcription (95–97). Thus, coactivator-mediated acetylation of receptor-activated Smad molecules in the nucleus may represent a novel way to modulate TGF-β signaling.
Smad3 can also be ADP-ribosylated in its MH1 domain through the action of poly(ADP-ribose) polymerase-1 (PARP-1), which associates with these Smads. This modification results in dissociation of Smad complexes from DNA, and, thus, attenuation of Smad-mediated TGF-β responses (98). ADP-ribosylation of Smad proteins may serve as an important step in controlling the intensity and duration of Smad-mediated transcription.
Regulation of nuclear shuttling of R-Smads
The disparate subcellular sites of R-Smad activation by cell surface TGF-β receptor complexes and their subsequent roles in transcription regulation of target genes requires efficient and highly regulated nucleo-cytoplasmic shuttling (99). This regulation involves nuclear localization sequences (NLS) in R-Smads, and the actions of importins β1, 7, 8 in nuclear import, and exportin 4 and Ran GTPases for nuclear export (100–102). RanBP3, an interacting partner of Ran, CRM1 and nucleopore components, is required for nuclear export of Smad2 and Smad3, as it directly recognizes dephosphorylated Smad2 and Smad3, and facilitates their nuclear export to terminate the TGF-β signaling (103). Phosphorylation of R-Smads and Smad complex formation promote their accumulation in the nucleus, while dephosphorylation by Smad phosphatases favors nuclear export (78, 81, 104).
The nuclear import of Smad complexes is also regulated by Akt. Attenuation of TGF-β-induced growth inhibition and apoptosis by PI3K/Akt signaling may result from direct association of Akt with unphosphorylated Smad3, thus preventing Smad3 activation, complex formation with Smad4, nuclear translocation and Smad-mediated transcription (105, 106). This function is thought to be independent of the Akt kinase activity, and creates a scenario that the Smad3 to Akt ratio correlates with the sensitivity of cells to TGF-β (106). Other evidence, however, suggests that the kinase activity of Akt is required and that mTOR plays a role in suppressing Smad3 activation and induction of apoptosis (107). SnoN, a co-repressor of Smad transcription complexes, which is predominant in the cytoplasm, can retain Smad proteins in the cytoplasm through its association with R-Smads. SnoN therefore represses TGF-β signaling using a dual mechanism, by preventing nuclear translocation of the Smad proteins, and as Smad corepressor at promoters of target genes (108). The Notch1 intracellular domain (NICD) also facilitates nuclear import of activated Smad3 and enhances its transcriptional activity, which may explain the enhanced TGF-β-induced functional activation of regulatory T-cells upon Notch signaling (109).
TAZ and YAP, transcription regulators in the Hippo pathway, also play a key role in nuclear import of TGF-β-activated Smad complexes and their recruitment to the Smad-binding DNA sequences. TAZ associates with Smad2/3-Smad4 complexes and is required for their nuclear import in embryonic stem cells, and is also co-recruited to Smad-mediated transcription complexes (110). At high cell density, and in response to the formation of polarity complexes, TAZ and YAP translocate into the cytoplasm and sequester Smad complexes, thereby preventing nuclear translocation and suppressing TGF-β signaling. During mouse embryogenesis, TAZ and/or YAP phosphorylation drives their nuclear accumulation, as well as that of Smad2/3 (111). Therefore, TAZ and YAP define a hierarchical system that regulates nuclear accumulation and transcription functions of activated Smad complexes.
Post-translational regulation of Smad4
Upon TGF-β-induced activation, complexes of Smad2 and/or Smad3 with Smad4 translocate into the nucleus. Smad4, however, is not required for nuclear import of R-Smads (112). Once incorporated in nucleoprotein complexes with DNA binding transcription factors, Smad4 functions as transcription coactivator, thus enhancing the effectiveness of R-Smad-mediated transcription (113). Although Smad4 is not essential for R-Smad mediated transcription, cells that lack Smad4 generally exhibit much weaker target gene responses (65). Like the R-Smads, the functions of Smad4 are modulated by post-translational modifications that define the interactions of Smad4 with other proteins, its subcellular localization, and its stability (Fig. 2).
Similarly to other Smads, Smad4 shuttles between the cytoplasm and nucleus, with nuclear import and export mediated by distinct nuclear localization and export sequences. Several importins and nuclear pore proteins, including CAN/Nup214, have been implicated in nuclear import of Smad4 or heteromeric Smad complexes (65, 100). Nuclear export of Smad4 requires exportin 1, also known as CRM1, and a nuclear export sequence near the Smad4 linker region (114, 115). The oncogene product v-ErbA can interact with Smad4 and sequester it in the cytoplasm, which results in attenuated TGF-β responsiveness and growth inhibition, and thus contributes to the oncogenicity of v-ErbA (116).
In addition to constitutive phosphorylation, Smad4 is also regulated by signal-dependent phosphorylation (39). The kinase LKB1 can phosphorylate Smad4 in its MH1 domain, thus interfering with Smad4 binding to DNA, and inhibiting Smad-mediated gene expression (117). Erk MAPK can phosphorylate Smad4 at a sequence in the linker region that stabilizes the binding of the co-activators p300 and CBP with the heteromeric Smad complex. This phosphorylation may enhance Smad-mediated transcription (118). The serine/threonine kinase MPK38 can also phosphorylate Smad4, and this phosphorylation is thought to contribute to enhanced TGF-β signaling induced by MPK38 (77).
Similarly to Smad2 and Smad3, ADP-ribosylation of the DNA-binding MH1 domain of Smad4 by PARP-1 interferes with the DNA binding of Smad4, resulting in attenuated Smad-mediated transcription (98). Smad4 can also be sumoylated in its MH1 domain by the E3 ligase PIAS1. Smad4 sumoylation may prevent Smad4 ubiquitylation, thus protecting Smad4 from degradation, and enhances TGF-β-induced Smad signaling (119–121). However, Smad4 sumoylation was also reported to repress Smad-mediated transcription by recruiting Daxx to the sumoylated form of Smad4 at regulatory promoter sequences (122).
Like R-Smads, Smad4 is targeted by poly-ubiquitylation, leading to degradation. While Smad4 degradation is mediated by the E3 ligases JAB1/CSN5 (123) and SCFβ-TrCP1 (124), some Smad4 mutants found in cancers are targeted for SCFskp2-mediated ubiquitylation (125).
Ectodermin, identified in Xenopus embryos, was shown to function as E3 ubiquitin ligase for Smad4 that drives Smad4 degradation and thus inhibits TGF-β signaling (126). Conflicting findings showed that this same protein, identified as TIF1γ and later renamed TRIM33, associates with TGF-β-activated Smad2/3 in competition with Smad4, and directs Smad4-independent transcription that regulates erythriod differentiation (127). Ectodermin/TIF1γ was also shown to mono-ubiquitylate Smad4 in its MH2 domain, thus interfering with Smad4 association with activated Smad2/3 (128). Furthermore, the PHD finger-bromodomain of TIF1γ constitutes a histone-binding module that specifically interacts with histone H3, and Smad4 ubiquitylation by TIF1γ requires this domain, and is induced by TIF1γ binding to histone (129). The PHD-Bromo domain of TIF1γ/TRIM33 also facilitates binding of TRIM33-Smad2/3 to H3K9me3 and H3K18ac on the promoters of mesendoderm regulators, thus displacing the chromatin-compacting factor HP1γ, and making Nodal response elements accessible to Smad4-Smad2/3 for later RNA polymerase II recruitment. Thus, Smad complexes use TIF1γ and the H3K9me3 mark as platform to trigger differentiation of mammalian embryonic stem cells (130). Despite their complexity, these findings all support the notion that, ectodermin/TIF1γ/TRIM33 antagonizes the transcription coactivator role of Smad4.
In contrast to the inhibitory effects of K519 mono-ubiquitylation by ectodermin (128), Smad4 mono-ubiquitylation at K507 enhanced the capacity of Smad4 to form complexes with activated R-Smads (131). Ubiquitylation-mediated degradation of Smad4 can be antagonized by de-ubiquitylases. Thus, the de-ubiquitylase FAM/USP9 can remove K519 mono-ubiquitylation of Smad4 imposed by TIF1γ, and thus restore Smad4 function (128).
In summary, like Smad2 and Smad3, the function and stability of Smad4 are extensively post-translationally regulated, thus providing high versatility to TGF-β/Smad signaling (Fig. 2).
Regulation of TGF-β-induced Smad signaling by inhibitory Smad7
In vertebrates, two inhibitory Smads, Smad6 and Smad7, inhibit Smad activation by binding to the activated type I receptor, thus interfering with R-Smad binding and activation. Smad6 preferentially inhibits BMP signaling through Smad1/5, and Smad7 inhibits both TGF-β/activin and BMP-induced R-Smad activation (132). Focusing on TGF-β signaling, Smad7 binds to activated TβRI, interfering with TGF-β-induced Smad2/3 activation (133), yet is also instrumental in attenuating the TGF-β receptors. Indeed, Smad7 binding to TβRI can recruit E3 ubiquitin ligases such as Smurf2 (45) or WWP1 (48) to the receptor complex, leading to TβRI ubiquitylation and degradation (45, 48), or the GADD34-PP1c phosphatase complex, which may play a role in terminating TGF-β signaling through dephosphorylation of the activated TβRI (40). Remarkably, Smad7 also acts as adaptor required for TGF-β-induced activation of p38 MAPK signaling (134). In the nucleus, Smad7 may antagonize TGF-β/Smad-mediated transcription by interfering with functional Smad-DNA complex formation (135), and was shown to associate with MyoD and promote myogenic differentiation (136). Continuously shuttling between nucleus and cytoplasm enables Smad7 to provide negative feedback in the control of TGF-β signaling in multiple compartments (Fig. 2) (65).
Many signaling pathways activate Smad7 expression and thus lead to attenuation of TGF-β signaling. Consistent with its role as feedback mechanism, TGF-β directly induces Smad7 expression though Smad3/4 binding to its promoter (8). Additionally, activation of ERK, JNK or p38 MAPK all result in transcriptional activation of Smad7 expression, thus dampening TGF-β signaling (137). Similarly, cytokines, such as IFN-γ or IL-7, can induce Smad7 expression through activation of Jak/STAT signaling (138, 139). Likewise, TNF-α induces Smad7 expression through NF-κB, thus inhibiting TGF-β/Smad-mediated responses (140). Hypoxia also activates Smad7 expression, which is mediated by the transcription factor HIF1α and contributes to malignant cell invasiveness (141).
Smad7 expression is additionally regulated by miRNAs, specifically miR-106b-25 and miR-21, resulting in decreased Smad7 expression and increased TGF-β signaling (142, 143). However, TGF-β can directly induce miR-21 expression through R-Smad activation. Thus, miR-21-mediated decrease of Smad7 expression could counteract Smad7-mediated negative feedback mechanisms of TGF-β signaling, and allow prolonged TGF-β effects, as proposed in the induction of fibrosis (143).
Smad7 protein levels are also controlled by ubiquitylation, e.g. by the E3 ligases Smurf1 and WWP1, and degradation (87). Furthermore, through its association with Axin, Smad7 is ubiquitylated by Arkadia (144, 145), and the resulting Smad7 degradation decreases the feedback inhibition of TGF-β signaling, resulting in increased TGF-β signaling, which may contribute to atrial fibrillation-induced atrial fibrosis (146). The role of Arkadia in Smad7 degradation and enhanced TGF-β signaling may provide a basis for its tumor suppressor properties in colorectal cancer (147). The deubiquitylating enzyme CYLD additionally inhibits TGF-β signaling by forming a complex with Smad7 and facilitates its deubiquitylation at two sites in its MH2 domain (148).
Finally, the function of Smad7 is regulated by acetylation. The interaction of Smad7 with the transcription coactivator p300 leads to direct Smad7 acetylation on two lysines close to the N-terminus (149). As these residues are also targeted by ubiquitylation, their acetylation prevents Smad7 ubiquitylation by Smurf1 and subsequent degradation (149). Conversely, the histone deacetylases HDAC1 and SIRT1 are able to interact with the N-terminus of Smad7 and reverse acetylation, thus enhancing Smurf1-mediated ubiquitylation and proteasomal degradation of Smad7 (150, 151). The competition between ubiquitylation and acetylation likely regulate Smad7 stability inside nucleus.
Regulation through Smad transcriptional crosstalk
In the nucleus, the activated Smad complexes orchestrate changes in transcription of target genes through cooperation with sequence-specific, DNA-binding transcription factors at conducive Smad binding sites. These transcription complexes coopt transcription coactivators and/or corepressors to define the amplitude of the transcription activation or repression, modify the local chromatin structure and engage the basal transcription machinery (9, 10). In this way, TGF-β directly induces Smad-mediated transcription activation or repression of several hundreds of genes, dependent on the physiological context (152). Intrinsic in this mechanism of Smad-mediated transcription is the cooperation with, and dependence on, a large and diverse set of DNA binding transcription factors that not only define the target regulatory DNA sequences of Smad cooperation, but are themselves extensively regulated by signaling pathways. These transcription complexes therefore intrinsically serve as excellent platforms for functional crosstalk between TGF-β/Smad signaling and other pathways. The complexity and versatility of transcription regulation of Smads, and the many opportunities for signaling crosstalk at the level of nucleoprotein transcription complexes have been reviewed (2, 9, 10, 137), but will be illustrated with a few examples of signaling crosstalk.
TGF-β/Smad signaling extensively cooperates with Wnt signaling, and such crosstalk occurs at multiple levels, including induction of TGF-β ligand expression (137), and interactions of TGF-β receptors or Smads with Axin or GSK3 (74, 145, 153), as mentioned. In the nucleoprotein transcription complexes, activated Smads were found to directly associate and cooperate with TCF and LEF, two DNA-binding transcription factors that serve as effectors of Wnt-induced transcription responses, and to transcriptionally regulate diverse Wnt target genes (154, 155). Smad also form transcription complexes with β-catenin and their association is thought to be stabilized by p300/CBP coactivator (155). For example, in mesenchymal stem cells, TGF-β induces a fast co-translocation of Smad3 and β-catenin into the nucleus, to then cooperatively regulate a set of genes that cannot be recognized by either Smads or β-catenin alone (156).
Cooperating with Notch signaling, TGF-β-activated Smad3 complexes associate directly with the Notch intracellular domain (NICD) and the DNA-binding transcription factor CSL to regulate transcription of the gene encoding Hes-1, a Notch signaling target gene (157). Conceptually similarly, activated Smad3 associated with NICD to cooperatively activate the Foxp3 expression in murine regulatory T cells (158). However, NICD also promotes cell growth and cancer development by suppressing the growth inhibitory effects of TGF-β by sequestering p300 from activated Smad3 (159).
Smads also crosstalk with MAPK pathways at multiple levels. In addition to activating the expression of TGF-β ligands and the inhibitory Smad7, MAPKs phosphorylate R-Smads and Smad4 at multiple sites, further defining their activities and stabilities (137). At the level of transcription regulation by Smads, MAPKs target various transcription factors that associate with Smads and enable Smads to cooperatively orchestrate gene expression responses. Among these, Smads associate and cooperate with c-Jun, Jun-B, c-Fos and other AP-1 transcription factors. Indeed, TGF-β-induced transcription activation of many genes is thought to result from crosstalk with MAPK signaling, and often occurs through cooperation of Smads with AP-1 complexes (160–163). As another example, TGF-β directly induces, through Smad3/4, expression of the transcription factor ATF3, which then cooperates with Smad3 in ATF3-mediated secondary gene expression responses. As target of both TGF-β signaling and p38 MAPK stress signaling, ATF3 integrates both pathways in the response of epithelial cells to stress and injury (164).
In summary, Smad-mediated transcription through interactions with many types of transcription factors allow extensive versatility, dependent upon activation of other signaling pathways.
Concluding remarks
TGF-β family signaling through Smads is conceptually a simple and linear signaling pathway, driven by sequential phosphorylation, with the type II receptors activating the type I receptors, which in turn activate R-Smads. However, as we dissect the complexities of the TGF-β responses and their context-dependence, we have come to appreciate its amazing versatility and regulation. Heteromeric combinations of type II and type I receptors, and of activated Smads provide levels of versatility that accommodate the many ligands and complex signaling patterns and cellular responses (9, 10, 65). As illustrated in this review, many levels of post-translational regulation of the receptors and Smads define receptor and Smad stability, provide negative feedback mechanisms, and help define their functions, thus providing insight into the exquisite regulation of the complex responses (Figs. 1 and 2). Protein interactions with the receptors and Smads further help define their functions and are required for the many phases in their life cycles, and the elaboration of appropriately tuned TGF-β responses. Finally, illustrated with some examples, functional crosstalk of TGF-β/Smad signaling with other pathways explains the context-dependence of the TGF-β response, depending on the physiological state of the cells. In fact, Smad-mediated transcription complexes intrinsically depend on cooperation with DNA-binding transcription factors and coregulators that are themselves also targets of many other pathways.
Going forward, further characterization of the regulation and roles of post-translational modifications and protein interactions in TGF-β/Smad signaling will provide exciting insights into how such conceptually simple signaling system can give rise to exquisite versatility and complexity. More richness to our understanding will be unraveled from the use of efficient and sensitive mass spectrometry methods, systems approaches using CHIP-seq, deep RNA sequencing techniques, and bioinformatics, in combination with detailed mechanistic studies.
Acknowledgments
In reviewing recent advances, we had to limit ourselves to only some observations to illustrate the roles of post-translational modifications and functional crosstalk. Inevitably, many highly relevant findings could not be included. We therefore apologize to those researchers whose work was not included in this review. Research in the lab of R.D. was supported by NIH grants RO1 CA63101 and CA136690 to R.D. J. L. is supported by a scientist development award from the Muscular Dystrophy Association.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Derynck R, Miyazono K. TGF-β and the TGF-β Family. In: Derynck R, Miyazono K, editors. The TGF-β Family. Cold Spring Harbor Laboratory Press; 2008. pp. 29–44. [Google Scholar]
- 2.Derynck R, Miyazono K, editors. The TGF-β Family. Cold Spring Harbor Laboratory Press; 2008. [Google Scholar]
- 3.Souchelnytskyi S, ten Dijke P, Miyazono K, Heldin CH. Phosphorylation of Ser165 in TGF-β type I receptor modulates TGF-β1-induced cellular responses. Embo J. 1996;15:6231–6240. [PMC free article] [PubMed] [Google Scholar]
- 4.Huse M, Muir TW, Xu L, Chen YG, Kuriyan J, Massague J. The TGF β receptor activation process: an inhibitor- to substrate-binding switch. Mol Cell. 2001;8:671–682. doi: 10.1016/s1097-2765(01)00332-x. [DOI] [PubMed] [Google Scholar]
- 5.Chen YG, Liu F, Massague J. Mechanism of TGFβ receptor inhibition by FKBP12. Embo J. 1997;16:3866–3876. doi: 10.1093/emboj/16.13.3866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shi Y, Massague J. Mechanisms of TGF-β signaling from cell membrane to the nucleus. Cell. 2003;113:685–700. doi: 10.1016/s0092-8674(03)00432-x. [DOI] [PubMed] [Google Scholar]
- 7.Chacko BM, Qin BY, Tiwari A, Shi G, Lam S, Hayward LJ, De Caestecker M, Lin K. Structural basis of heteromeric smad protein assembly in TGF-β signaling. Mol Cell. 2004;15:813–823. doi: 10.1016/j.molcel.2004.07.016. [DOI] [PubMed] [Google Scholar]
- 8.Derynck R, Zhang YE. Smad-dependent and Smad-independent pathways in TGF-β family signalling. Nature. 2003;425:577–584. doi: 10.1038/nature02006. [DOI] [PubMed] [Google Scholar]
- 9.Massague J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19:2783–2810. doi: 10.1101/gad.1350705. [DOI] [PubMed] [Google Scholar]
- 10.Feng XH, Derynck R. Specificity and versatility in TGF-β signaling through Smads. Annu Rev Cell Dev Biol. 2005;21:659–693. doi: 10.1146/annurev.cellbio.21.022404.142018. [DOI] [PubMed] [Google Scholar]
- 11.Dabovic B, Rifkin DB. TGF-β Bioavailability: Latency, Targeting, and Activation. In: Derynck R, Miyazono K, editors. The TGF-β Family. Cold Spring Harbor Laboratory Press; 2008. pp. 179–202. [Google Scholar]
- 12.Annes JP, Munger JS, Rifkin DB. Making sense of latent TGFβ activation. J Cell Sci. 2003;116:217–224. doi: 10.1242/jcs.00229. [DOI] [PubMed] [Google Scholar]
- 13.Wipff PJ, Hinz B. Integrins and the activation of latent transforming growth factor β1 - an intimate relationship. Eur J Cell Biol. 2008;87:601–615. doi: 10.1016/j.ejcb.2008.01.012. [DOI] [PubMed] [Google Scholar]
- 14.Yang Z, Mu Z, Dabovic B, Jurukovski V, Yu D, Sung J, Xiong X, Munger JS. Absence of integrin-mediated TGFβ1 activation in vivo recapitulates the phenotype of TGFβ1-null mice. J Cell Biol. 2007;176:787–793. doi: 10.1083/jcb.200611044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kudo M, Melton AC, Chen C, Engler MB, Huang KE, Ren X, Wang Y, Bernstein X, Li JT, Atabai K, Huang X, Sheppard D. IL-17A produced by αβ T cells drives airway hyper-responsiveness in mice and enhances mouse and human airway smooth muscle contraction. Nat Med. 2012;18:547–554. doi: 10.1038/nm.2684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Melton AC, Bailey-Bucktrout SL, Travis MA, Fife BT, Bluestone JA, Sheppard D. Expression of αvβ8 integrin on dendritic cells regulates Th17 cell development and experimental autoimmune encephalomyelitis in mice. J Clin Invest. 2010;120:4436–4444. doi: 10.1172/JCI43786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Mu D, Cambier S, Fjellbirkeland L, Baron JL, Munger JS, Kawakatsu H, Sheppard D, Broaddus VC, Nishimura SL. The integrin α(v)β8 mediates epithelial homeostasis through MT1-MMP-dependent activation of TGF-β1. J Cell Biol. 2002;157:493–507. doi: 10.1083/jcb.200109100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Annes JP, Chen Y, Munger JS, Rifkin DB. Integrin αVβ6-mediated activation of latent TGF-β requires the latent TGF-β binding protein-1. J Cell Biol. 2004;165:723–734. doi: 10.1083/jcb.200312172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wipff PJ, Rifkin DB, Meister JJ, Hinz B. Myofibroblast contraction activates latent TGF-β1 from the extracellular matrix. J Cell Biol. 2007;179:1311–1323. doi: 10.1083/jcb.200704042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ahamed J, Burg N, Yoshinaga K, Janczak CA, Rifkin DB, Coller BS. In vitro and in vivo evidence for shear-induced activation of latent transforming growth factor-β1. Blood. 2008;112:3650–3660. doi: 10.1182/blood-2008-04-151753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. doi: 10.1016/s0092-8674(04)00045-5. [DOI] [PubMed] [Google Scholar]
- 22.Inui M, Martello G, Piccolo S. MicroRNA control of signal transduction. Nat Rev Mol Cell Biol. 2010;11:252–263. doi: 10.1038/nrm2868. [DOI] [PubMed] [Google Scholar]
- 23.Martello G, Zacchigna L, Inui M, Montagner M, Adorno M, Mamidi A, Morsut L, Soligo S, Tran U, Dupont S, Cordenonsi M, Wessely O, Piccolo S. MicroRNA control of Nodal signalling. Nature. 2007;449:183–188. doi: 10.1038/nature06100. [DOI] [PubMed] [Google Scholar]
- 24.Keklikoglou I, Koerner C, Schmidt C, Zhang JD, Heckmann D, Shavinskaya A, Allgayer H, Guckel B, Fehm T, Schneeweiss A, Sahin O, Wiemann S, Tschulena U. MicroRNA-520/373 family functions as a tumor suppressor in estrogen receptor negative breast cancer by targeting NF-kappaB and TGF-β signaling pathways. Oncogene. 2011 doi: 10.1038/onc.2011.571. [DOI] [PubMed] [Google Scholar]
- 25.Subramanyam D, Lamouille S, Judson RL, Liu JY, Bucay N, Derynck R, Blelloch R. Multiple targets of miR-302 and miR-372 promote reprogramming of human fibroblasts to induced pluripotent stem cells. Nat Biotechnol. 2011;29:443–448. doi: 10.1038/nbt.1862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lipchina I, Elkabetz Y, Hafner M, Sheridan R, Mihailovic A, Tuschl T, Sander C, Studer L, Betel D. Genome-wide identification of microRNA targets in human ES cells reveals a role for miR-302 in modulating BMP response. Genes Dev. 2011;25:2173–2186. doi: 10.1101/gad.17221311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mestdagh P, Bostrom AK, Impens F, Fredlund E, Van Peer G, De Antonellis P, von Stedingk K, Ghesquiere B, Schulte S, Dews M, Thomas-Tikhonenko A, Schulte JH, Zollo M, Schramm A, Gevaert K, Axelson H, Speleman F, Vandesompele J. The miR-17–92 microRNA cluster regulates multiple components of the TGF-β pathway in neuroblastoma. Mol Cell. 2010;40:762–773. doi: 10.1016/j.molcel.2010.11.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wu L, Derynck R. Essential role of TGF-β signaling in glucose-induced cell hypertrophy. Dev Cell. 2009;17:35–48. doi: 10.1016/j.devcel.2009.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liu C, Xu P, Lamouille S, Xu J, Derynck R. TACE-mediated ectodomain shedding of the type I TGF-β receptor downregulates TGF-β signaling. Mol Cell. 2009;35:26–36. doi: 10.1016/j.molcel.2009.06.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Mu Y, Sundar R, Thakur N, Ekman M, Gudey SK, Yakymovych M, Hermansson A, Dimitriou H, Bengoechea-Alonso MT, Ericsson J, Heldin CH, Landstrom M. TRAF6 ubiquitinates TGFβ type I receptor to promote its cleavage and nuclear translocation in cancer. Nat Commun. 2011;2:330. doi: 10.1038/ncomms1332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Xu P, Derynck R. Direct activation of TACE-mediated ectodomain shedding by p38 MAP kinase regulates EGF receptor-dependent cell proliferation. Mol Cell. 2010;37:551–566. doi: 10.1016/j.molcel.2010.01.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Atfi A, Dumont E, Colland F, Bonnier D, L'Helgoualc'h A, Prunier C, Ferrand N, Clement B, Wewer UM, Theret N. The disintegrin and metalloproteinase ADAM12 contributes to TGF-β signaling through interaction with the type II receptor. J Cell Biol. 2007;178:201–208. doi: 10.1083/jcb.200612046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kang JS, Liu C, Derynck R. New regulatory mechanisms of TGF-β receptor function. Trends Cell Biol. 2009;19:385–394. doi: 10.1016/j.tcb.2009.05.008. [DOI] [PubMed] [Google Scholar]
- 34.Hanks SK, Hunter T. Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. Faseb J. 1995;9:576–596. [PubMed] [Google Scholar]
- 35.Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science. 2002;298:1912–1934. doi: 10.1126/science.1075762. [DOI] [PubMed] [Google Scholar]
- 36.Lawler S, Feng XH, Chen RH, Maruoka EM, Turck CW, Griswold-Prenner I, Derynck R. The type II transforming growth factor-β receptor autophosphorylates not only on serine and threonine but also on tyrosine residues. J Biol Chem. 1997;272:14850–14859. doi: 10.1074/jbc.272.23.14850. [DOI] [PubMed] [Google Scholar]
- 37.Galliher AJ, Schiemann WP. Src phosphorylates Tyr284 in TGF-β type II receptor and regulates TGF-β stimulation of p38 MAPK during breast cancer cell proliferation and invasion. Cancer Res. 2007;67:3752–3758. doi: 10.1158/0008-5472.CAN-06-3851. [DOI] [PubMed] [Google Scholar]
- 38.Lee MK, Pardoux C, Hall MC, Lee PS, Warburton D, Qing J, Smith SM, Derynck R. TGF-β activates Erk MAP kinase signalling through direct phosphorylation of ShcA. Embo J. 2007;26:3957–3967. doi: 10.1038/sj.emboj.7601818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wrighton KH, Lin X, Feng XH. Phospho-control of TGF-β superfamily signaling. Cell Res. 2009;19:8–20. doi: 10.1038/cr.2008.327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Shi W, Sun C, He B, Xiong W, Shi X, Yao D, Cao X. GADD34-PP1c recruited by Smad7 dephosphorylates TGFβ type I receptor. J Cell Biol. 2004;164:291–300. doi: 10.1083/jcb.200307151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bennett D, Alphey L. PP1 binds Sara and negatively regulates Dpp signaling in Drosophila melanogaster. Nat Genet. 2002;31:419–423. doi: 10.1038/ng938. [DOI] [PubMed] [Google Scholar]
- 42.Batut J, Schmierer B, Cao J, Raftery LA, Hill CS, Howell M. Two highly related regulatory subunits of PP2A exert opposite effects on TGF-β/Activin/Nodal signalling. Development. 2008;135:2927–2937. doi: 10.1242/dev.020842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Griswold-Prenner I, Kamibayashi C, Maruoka EM, Mumby MC, Derynck R. Physical and functional interactions between type I transforming growth factor β receptors and Bα, a WD-40 repeat subunit of phosphatase 2A. Mol Cell Biol. 1998;18:6595–6604. doi: 10.1128/mcb.18.11.6595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kerscher O, Felberbaum R, Hochstrasser M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol. 2006;22:159–180. doi: 10.1146/annurev.cellbio.22.010605.093503. [DOI] [PubMed] [Google Scholar]
- 45.Kavsak P, Rasmussen RK, Causing CG, Bonni S, Zhu H, Thomsen GH, Wrana JL. Smad7 binds to Smurf2 to form an E3 ubiquitin ligase that targets the TGF β receptor for degradation. Mol Cell. 2000;6:1365–1375. doi: 10.1016/s1097-2765(00)00134-9. [DOI] [PubMed] [Google Scholar]
- 46.Ogunjimi AA, Briant DJ, Pece-Barbara N, Le Roy C, Di Guglielmo GM, Kavsak P, Rasmussen RK, Seet BT, Sicheri F, Wrana JL. Regulation of Smurf2 ubiquitin ligase activity by anchoring the E2 to the HECT domain. Mol Cell. 2005;19:297–308. doi: 10.1016/j.molcel.2005.06.028. [DOI] [PubMed] [Google Scholar]
- 47.Kuratomi G, Komuro A, Goto K, Shinozaki M, Miyazawa K, Miyazono K, Imamura T. NEDD4–2 (neural precursor cell expressed, developmentally down-regulated 4–2) negatively regulates TGF-β (transforming growth factor-β) signalling by inducing ubiquitin-mediated degradation of Smad2 and TGF-β type I receptor. Biochem J. 2005;386:461–470. doi: 10.1042/BJ20040738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Komuro A, Imamura T, Saitoh M, Yoshida Y, Yamori T, Miyazono K, Miyazawa K. Negative regulation of transforming growth factor-β (TGF-β) signaling by WW domain-containing protein 1 (WWP1) Oncogene. 2004;23:6914–6923. doi: 10.1038/sj.onc.1207885. [DOI] [PubMed] [Google Scholar]
- 49.Eichhorn PJ, Rodon L, Gonzalez-Junca A, Dirac A, Gili M, Martinez-Saez E, Aura C, Barba I, Peg V, Prat A, Cuartas I, Jimenez J, Garcia-Dorado D, Sahuquillo J, Bernards R, Baselga J, Seoane J. USP15 stabilizes TGF-β receptor I and promotes oncogenesis through the activation of TGF-β signaling in glioblastoma. Nat Med. 2012;18:429–435. doi: 10.1038/nm.2619. [DOI] [PubMed] [Google Scholar]
- 50.Wicks SJ, Haros K, Maillard M, Song L, Cohen RE, Dijke PT, Chantry A. The deubiquitinating enzyme UCH37 interacts with Smads and regulates TGF-β signalling. Oncogene. 2005;24:8080–8084. doi: 10.1038/sj.onc.1208944. [DOI] [PubMed] [Google Scholar]
- 51.Wrighton KH, Lin X, Feng XH. Critical regulation of TGFβ signaling by Hsp90. Proc Natl Acad Sci U S A. 2008;105:9244–9249. doi: 10.1073/pnas.0800163105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kang JS, Saunier EF, Akhurst RJ, Derynck R. The type I TGF-β receptor is covalently modified and regulated by sumoylation. Nat Cell Biol. 2008;10:654–664. doi: 10.1038/ncb1728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Le Roy C, Wrana JL. Clathrin- and non-clathrin-mediated endocytic regulation of cell signalling. Nat Rev Mol Cell Biol. 2005;6:112–126. doi: 10.1038/nrm1571. [DOI] [PubMed] [Google Scholar]
- 54.Di Guglielmo GM, Le Roy C, Goodfellow AF, Wrana JL. Distinct endocytic pathways regulate TGF-β receptor signalling and turnover. Nat Cell Biol. 2003;5:410–421. doi: 10.1038/ncb975. [DOI] [PubMed] [Google Scholar]
- 55.Chen YG. Endocytic regulation of TGF-β signaling. Cell Res. 2009;19:58–70. doi: 10.1038/cr.2008.315. [DOI] [PubMed] [Google Scholar]
- 56.Mitchell H, Choudhury A, Pagano RE, Leof EB. Ligand-dependent and -independent transforming growth factor-β receptor recycling regulated by clathrin-mediated endocytosis and Rab11. Mol Biol Cell. 2004;15:4166–4178. doi: 10.1091/mbc.E04-03-0245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Yao D, Ehrlich M, Henis YI, Leof EB. Transforming growth factor-β receptors interact with AP2 by direct binding to β2 subunit. Mol Biol Cell. 2002;13:4001–4012. doi: 10.1091/mbc.02-07-0104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hayes S, Chawla A, Corvera S. TGF β receptor internalization into EEA1-enriched early endosomes: role in signaling to Smad2. J Cell Biol. 2002;158:1239–1249. doi: 10.1083/jcb.200204088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hu H, Milstein M, Bliss JM, Thai M, Malhotra G, Huynh LC, Colicelli J. Integration of transforming growth factor β and RAS signaling silences a RAB5 guanine nucleotide exchange factor and enhances growth factor-directed cell migration. Mol Cell Biol. 2008;28:1573–1583. doi: 10.1128/MCB.01087-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Razani B, Zhang XL, Bitzer M, von Gersdorff G, Bottinger EP, Lisanti MP. Caveolin-1 regulates transforming growth factor (TGF)-β/SMAD signaling through an interaction with the TGF-β type I receptor. J Biol Chem. 2001;276:6727–6738. doi: 10.1074/jbc.M008340200. [DOI] [PubMed] [Google Scholar]
- 61.Sawada S, Ishikawa C, Tanji H, Nakachi S, Senba M, Okudaira T, Uchihara JN, Taira N, Ohshiro K, Yamada Y, Tanaka Y, Uezato H, Ohshima K, Sasai K, Burgering BM, Duc Dodon M, Fujii M, Sunakawa H, Mori N. Overexpression of caveolin-1 in adult T-cell leukemia. Blood. 2010;115:2220–2230. doi: 10.1182/blood-2009-08-240044. [DOI] [PubMed] [Google Scholar]
- 62.Zuo W, Chen YG. Specific activation of mitogen-activated protein kinase by transforming growth factor-β receptors in lipid rafts is required for epithelial cell plasticity. Mol Biol Cell. 2009;20:1020–1029. doi: 10.1091/mbc.E08-09-0898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Kardassis D, Murphy C, Fotsis T, Moustakas A, Stournaras C. Control of transforming growth factor β signal transduction by small GTPases. Febs J. 2009;276:2947–2965. doi: 10.1111/j.1742-4658.2009.07031.x. [DOI] [PubMed] [Google Scholar]
- 64.Choi SC, Kim GH, Lee SJ, Park E, Yeo CY, Han JK. Regulation of activin/nodal signaling by Rap2-directed receptor trafficking. Dev Cell. 2008;15:49–61. doi: 10.1016/j.devcel.2008.05.004. [DOI] [PubMed] [Google Scholar]
- 65.Heldin CH, Moustakas A. Role of Smads in TGFβ signaling. Cell Tissue Res. 2012;347:21–36. doi: 10.1007/s00441-011-1190-x. [DOI] [PubMed] [Google Scholar]
- 66.Wang G, Matsuura I, He D, Liu F. Transforming growth factor-{β}-inducible phosphorylation of Smad3. J Biol Chem. 2009;284:9663–9673. doi: 10.1074/jbc.M809281200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Millet C, Yamashita M, Heller M, Yu LR, Veenstra TD, Zhang YE. A negative feedback control of transforming growth factor-β signaling by glycogen synthase kinase 3-mediated Smad3 linker phosphorylation at Ser-204. J Biol Chem. 2009;284:19808–19816. doi: 10.1074/jbc.M109.016667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Alarcon C, Zaromytidou AI, Xi Q, Gao S, Yu J, Fujisawa S, Barlas A, Miller AN, Manova-Todorova K, Macias MJ, Sapkota G, Pan D, Massague J. Nuclear CDKs drive Smad transcriptional activation and turnover in BMP and TGF-β pathways. Cell. 2009;139:757–769. doi: 10.1016/j.cell.2009.09.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Gao S, Alarcon C, Sapkota G, Rahman S, Chen PY, Goerner N, Macias MJ, Erdjument-Bromage H, Tempst P, Massague J. Ubiquitin ligase Nedd4L targets activated Smad2/3 to limit TGF-β signaling. Mol Cell. 2009;36:457–468. doi: 10.1016/j.molcel.2009.09.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Aragon E, Goerner N, Zaromytidou AI, Xi Q, Escobedo A, Massague J, Macias MJ. A Smad action turnover switch operated by WW domain readers of a phosphoserine code. Genes Dev. 2011;25:1275–1288. doi: 10.1101/gad.2060811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Fuentealba LC, Eivers E, Ikeda A, Hurtado C, Kuroda H, Pera EM, De Robertis EM. Integrating patterning signals: Wnt/GSK3 regulates the duration of the BMP/Smad1 signal. Cell. 2007;131:980–993. doi: 10.1016/j.cell.2007.09.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Matsuura I, Chiang KN, Lai CY, He D, Wang G, Ramkumar R, Uchida T, Ryo A, Lu K, Liu F. Pin1 promotes transforming growth factor-β-induced migration and invasion. J Biol Chem. 2010;285:1754–1764. doi: 10.1074/jbc.M109.063826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Ho J, Cocolakis E, Dumas VM, Posner BI, Laporte SA, Lebrun JJ. The G protein-coupled receptor kinase-2 is a TGFβ-inducible antagonist of TGFβ signal transduction. Embo J. 2005;24:3247–3258. doi: 10.1038/sj.emboj.7600794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Guo X, Ramirez A, Waddell DS, Li Z, Liu X, Wang XF. Axin and GSK3- control Smad3 protein stability and modulate TGF- signaling. Genes Dev. 2008;22:106–120. doi: 10.1101/gad.1590908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zhu S, Wang W, Clarke DC, Liu X. Activation of Mps1 promotes transforming growth factor-β-independent Smad signaling. J Biol Chem. 2007;282:18327–18338. doi: 10.1074/jbc.M700636200. [DOI] [PubMed] [Google Scholar]
- 76.Lee BH, Chen W, Stippec S, Cobb MH. Biological cross-talk between WNK1 and the transforming growth factor β-Smad signaling pathway. J Biol Chem. 2007;282:17985–17996. doi: 10.1074/jbc.M702664200. [DOI] [PubMed] [Google Scholar]
- 77.Seong HA, Jung H, Ha H. Murine protein serine/threonine kinase 38 stimulates TGF-β signaling in a kinase-dependent manner via direct phosphorylation of Smad proteins. J Biol Chem. 2010;285:30959–30970. doi: 10.1074/jbc.M110.138370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Lin X, Duan X, Liang YY, Su Y, Wrighton KH, Long J, Hu M, Davis CM, Wang J, Brunicardi FC, Shi Y, Chen YG, Meng A, Feng XH. PPM1A functions as a Smad phosphatase to terminate TGFβ signaling. Cell. 2006;125:915–928. doi: 10.1016/j.cell.2006.03.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Bu S, Kapanadze B, Hsu T, Trojanowska M. Opposite effects of dihydrosphingosine 1-phosphate and sphingosine 1-phosphate on transforming growth factor-β/Smad signaling are mediated through the PTEN/PPM1A-dependent pathway. J Biol Chem. 2008;283:19593–19602. doi: 10.1074/jbc.M802417200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Yu J, Pan L, Qin X, Chen H, Xu Y, Chen Y, Tang H. MTMR4 attenuates transforming growth factor β (TGFβ) signaling by dephosphorylating R-Smads in endosomes. J Biol Chem. 2010;285:8454–8462. doi: 10.1074/jbc.M109.075036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Knockaert M, Sapkota G, Alarcon C, Massague J, Brivanlou AH. Unique players in the BMP pathway: small C-terminal domain phosphatases dephosphorylate Smad1 to attenuate BMP signaling. Proc Natl Acad Sci U S A. 2006;103:11940–11945. doi: 10.1073/pnas.0605133103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Heikkinen PT, Nummela M, Leivonen SK, Westermarck J, Hill CS, Kahari VM, Jaakkola PM. Hypoxia-activated Smad3-specific dephosphorylation by PP2A. J Biol Chem. 2010;285:3740–3749. doi: 10.1074/jbc.M109.042978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Shukla A, Malik M, Cataisson C, Ho Y, Friesen T, Suh KS, Yuspa SH. TGF-β signalling is regulated by Schnurri-2-dependent nuclear translocation of CLIC4 and consequent stabilization of phospho-Smad2 and 3. Nat Cell Biol. 2009;11:777–784. doi: 10.1038/ncb1885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Wrighton KH, Willis D, Long J, Liu F, Lin X, Feng XH. Small C-terminal domain phosphatases dephosphorylate the regulatory linker regions of Smad2 and Smad3 to enhance transforming growth factor-β signaling. J Biol Chem. 2006;281:38365–38375. doi: 10.1074/jbc.M607246200. [DOI] [PubMed] [Google Scholar]
- 85.Sapkota G, Knockaert M, Alarcon C, Montalvo E, Brivanlou AH, Massague J. Dephosphorylation of the linker regions of Smad1 and Smad2/3 by small C-terminal domain phosphatases has distinct outcomes for bone morphogenetic protein and transforming growth factor-β pathways. J Biol Chem. 2006;281:40412–40419. doi: 10.1074/jbc.M610172200. [DOI] [PubMed] [Google Scholar]
- 86.Lo RS, Massague J. Ubiquitin-dependent degradation of TGF-β-activated smad2. Nat Cell Biol. 1999;1:472–478. doi: 10.1038/70258. [DOI] [PubMed] [Google Scholar]
- 87.Inoue Y, Imamura T. Regulation of TGF-β family signaling by E3 ubiquitin ligases. Cancer Sci. 2008;99:2107–2112. doi: 10.1111/j.1349-7006.2008.00925.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Levy L, Howell M, Das D, Harkin S, Episkopou V, Hill CS. Arkadia activates Smad3/Smad4-dependent transcription by triggering signal-induced SnoN degradation. Mol Cell Biol. 2007;27:6068–6083. doi: 10.1128/MCB.00664-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Mavrakis KJ, Andrew RL, Lee KL, Petropoulou C, Dixon JE, Navaratnam N, Norris DP, Episkopou V. Arkadia enhances Nodal/TGF-β signaling by coupling phospho-Smad2/3 activity and turnover. PLoS Biol. 2007;5:e67. doi: 10.1371/journal.pbio.0050067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Ito I, Hanyu A, Wayama M, Goto N, Katsuno Y, Kawasaki S, Nakajima Y, Kajiro M, Komatsu Y, Fujimura A, Hirota R, Murayama A, Kimura K, Imamura T, Yanagisawa J. Estrogen inhibits transforming growth factor β signaling by promoting Smad2/3 degradation. J Biol Chem. 2010;285:14747–14755. doi: 10.1074/jbc.M109.093039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Bai Y, Yang C, Hu K, Elly C, Liu YC. Itch E3 ligase-mediated regulation of TGF-β signaling by modulating smad2 phosphorylation. Mol Cell. 2004;15:825–831. doi: 10.1016/j.molcel.2004.07.021. [DOI] [PubMed] [Google Scholar]
- 92.Inui M, Manfrin A, Mamidi A, Martello G, Morsut L, Soligo S, Enzo E, Moro S, Polo S, Dupont S, Cordenonsi M, Piccolo S. USP15 is a deubiquitylating enzyme for receptor-activated SMADs. Nat Cell Biol. 2011;13:1368–1375. doi: 10.1038/ncb2346. [DOI] [PubMed] [Google Scholar]
- 93.Tang LY, Yamashita M, Coussens NP, Tang Y, Wang X, Li C, Deng CX, Cheng SY, Zhang YE. Ablation of Smurf2 reveals an inhibition in TGF-β signalling through multiple mono-ubiquitination of Smad3. Embo J. 2011;30:4777–4789. doi: 10.1038/emboj.2011.393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Lim JH, Jono H, Komatsu K, Woo CH, Lee J, Miyata M, Matsuno T, Xu X, Huang Y, Zhang W, Park SH, Kim YI, Choi YD, Shen H, Heo KS, Xu H, Bourne P, Koga T, Xu H, Yan C, Wang B, Chen LF, Feng XH, Li JD. CYLD negatively regulates transforming growth factor-β-signalling via deubiquitinating Akt. Nat Commun. 2012;3:771. doi: 10.1038/ncomms1776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Inoue Y, Itoh Y, Abe K, Okamoto T, Daitoku H, Fukamizu A, Onozaki K, Hayashi H. Smad3 is acetylated by p300/CBP to regulate its transactivation activity. Oncogene. 2007;26:500–508. doi: 10.1038/sj.onc.1209826. [DOI] [PubMed] [Google Scholar]
- 96.Simonsson M, Kanduri M, Gronroos E, Heldin CH, Ericsson J. The DNA binding activities of Smad2 and Smad3 are regulated by coactivator-mediated acetylation. J Biol Chem. 2006;281:39870–39880. doi: 10.1074/jbc.M607868200. [DOI] [PubMed] [Google Scholar]
- 97.Tu AW, Luo K. Acetylation of Smad2 by the co-activator p300 regulates activin and transforming growth factor β response. J Biol Chem. 2007;282:21187–21196. doi: 10.1074/jbc.M700085200. [DOI] [PubMed] [Google Scholar]
- 98.Lonn P, van der Heide LP, Dahl M, Hellman U, Heldin CH, Moustakas A. PARP-1 attenuates Smad-mediated transcription. Mol Cell. 2010;40:521–532. doi: 10.1016/j.molcel.2010.10.029. [DOI] [PubMed] [Google Scholar]
- 99.Schmierer B, Tournier AL, Bates PA, Hill CS. Mathematical modeling identifies Smad nucleocytoplasmic shuttling as a dynamic signal-interpreting system. Proc Natl Acad Sci U S A. 2008;105:6608–6613. doi: 10.1073/pnas.0710134105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hill CS. Nucleocytoplasmic shuttling of Smad proteins. Cell Res. 2009;19:36–46. doi: 10.1038/cr.2008.325. [DOI] [PubMed] [Google Scholar]
- 101.Xu L, Yao X, Chen X, Lu P, Zhang B, Ip YT. Msk is required for nuclear import of TGF-{β}/BMP-activated Smads. J Cell Biol. 2007;178:981–994. doi: 10.1083/jcb.200703106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Kurisaki A, Kurisaki K, Kowanetz M, Sugino H, Yoneda Y, Heldin CH, Moustakas A. The mechanism of nuclear export of Smad3 involves exportin 4 and Ran. Mol Cell Biol. 2006;26:1318–1332. doi: 10.1128/MCB.26.4.1318-1332.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Dai F, Lin X, Chang C, Feng XH. Nuclear export of Smad2 and Smad3 by RanBP3 facilitates termination of TGF-β signaling. Dev Cell. 2009;16:345–357. doi: 10.1016/j.devcel.2009.01.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Chen HB, Shen J, Ip YT, Xu L. Identification of phosphatases for Smad in the BMP/DPP pathway. Genes Dev. 2006;20:648–653. doi: 10.1101/gad.1384706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Remy I, Montmarquette A, Michnick SW. PKB/Akt modulates TGF-β signalling through a direct interaction with Smad3. Nat Cell Biol. 2004;6:358–365. doi: 10.1038/ncb1113. [DOI] [PubMed] [Google Scholar]
- 106.Conery AR, Cao Y, Thompson EA, Townsend CM, Jr, Ko TC, Luo K. Akt interacts directly with Smad3 to regulate the sensitivity to TGF-β induced apoptosis. Nat Cell Biol. 2004;6:366–372. doi: 10.1038/ncb1117. [DOI] [PubMed] [Google Scholar]
- 107.Song K, Wang H, Krebs TL, Danielpour D. Novel roles of Akt and mTOR in suppressing TGF-β/ALK5-mediated Smad3 activation. Embo J. 2006;25:58–69. doi: 10.1038/sj.emboj.7600917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Krakowski AR, Laboureau J, Mauviel A, Bissell MJ, Luo K. Cytoplasmic SnoN in normal tissues and nonmalignant cells antagonizes TGF-β signaling by sequestration of the Smad proteins. Proc Natl Acad Sci U S A. 2005;102:12437–12442. doi: 10.1073/pnas.0504107102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Asano N, Watanabe T, Kitani A, Fuss IJ, Strober W. Notch1 signaling and regulatory T cell function. J Immunol. 2008;180:2796–2804. doi: 10.4049/jimmunol.180.5.2796. [DOI] [PubMed] [Google Scholar]
- 110.Varelas X, Sakuma R, Samavarchi-Tehrani P, Peerani R, Rao BM, Dembowy J, Yaffe MB, Zandstra PW, Wrana JL. TAZ controls Smad nucleocytoplasmic shuttling and regulates human embryonic stem-cell self-renewal. Nat Cell Biol. 2008;10:837–848. doi: 10.1038/ncb1748. [DOI] [PubMed] [Google Scholar]
- 111.Varelas X, Samavarchi-Tehrani P, Narimatsu M, Weiss A, Cockburn K, Larsen BG, Rossant J, Wrana JL. The Crumbs complex couples cell density sensing to Hippo-dependent control of the TGF-β-SMAD pathway. Dev Cell. 2010;19:831–844. doi: 10.1016/j.devcel.2010.11.012. [DOI] [PubMed] [Google Scholar]
- 112.Fink SP, Mikkola D, Willson JK, Markowitz S. TGF-β-induced nuclear localization of Smad2 and Smad3 in Smad4 null cancer cell lines. Oncogene. 2003;22:1317–1323. doi: 10.1038/sj.onc.1206128. [DOI] [PubMed] [Google Scholar]
- 113.Feng XH, Zhang Y, Wu RY, Derynck R. The tumor suppressor Smad4/DPC4 and transcriptional adaptor CBP/p300 are coactivators for smad3 in TGF-β-induced transcriptional activation. Genes Dev. 1998;12:2153–2163. doi: 10.1101/gad.12.14.2153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Pierreux CE, Nicolas FJ, Hill CS. Transforming growth factor β-independent shuttling of Smad4 between the cytoplasm and nucleus. Mol Cell Biol. 2000;20:9041–9054. doi: 10.1128/mcb.20.23.9041-9054.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Watanabe M, Masuyama N, Fukuda M, Nishida E. Regulation of intracellular dynamics of Smad4 by its leucine-rich nuclear export signal. EMBO Rep. 2000;1:176–182. doi: 10.1093/embo-reports/kvd029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Erickson RA, Liu X. Association of v-ErbA with Smad4 disrupts TGF-β signaling. Mol Biol Cell. 2009;20:1509–1519. doi: 10.1091/mbc.E08-08-0836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Moren A, Raja E, Heldin CH, Moustakas A. Negative regulation of TGFβ signaling by the kinase LKB1 and the scaffolding protein LIP1. J Biol Chem. 2011;286:341–353. doi: 10.1074/jbc.M110.190660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Roelen BA, Cohen OS, Raychowdhury MK, Chadee DN, Zhang Y, Kyriakis JM, Alessandrini AA, Lin HY. Phosphorylation of threonine 276 in Smad4 is involved in transforming growth factor-β-induced nuclear accumulation. Am J Physiol Cell Physiol. 2003;285:C823–830. doi: 10.1152/ajpcell.00053.2003. [DOI] [PubMed] [Google Scholar]
- 119.Ohshima T, Shimotohno K. Transforming growth factor-β-mediated signaling via the p38 MAP kinase pathway activates Smad-dependent transcription through SUMO-1 modification of Smad4. J Biol Chem. 2003;278:50833–50842. doi: 10.1074/jbc.M307533200. [DOI] [PubMed] [Google Scholar]
- 120.Lin X, Liang M, Liang YY, Brunicardi FC, Feng XH. SUMO-1/Ubc9 promotes nuclear accumulation and metabolic stability of tumor suppressor Smad4. J Biol Chem. 2003;278:31043–31048. doi: 10.1074/jbc.C300112200. [DOI] [PubMed] [Google Scholar]
- 121.Lee PS, Chang C, Liu D, Derynck R. Sumoylation of Smad4, the common Smad mediator of transforming growth factor-β family signaling. J Biol Chem. 2003;278:27853–27863. doi: 10.1074/jbc.M301755200. [DOI] [PubMed] [Google Scholar]
- 122.Chang CC, Lin DY, Fang HI, Chen RH, Shih HM. Daxx mediates the small ubiquitin-like modifier-dependent transcriptional repression of Smad4. J Biol Chem. 2005;280:10164–10173. doi: 10.1074/jbc.M409161200. [DOI] [PubMed] [Google Scholar]
- 123.Wan M, Cao X, Wu Y, Bai S, Wu L, Shi X, Wang N, Cao X. Jab1 antagonizes TGF-β signaling by inducing Smad4 degradation. EMBO Rep. 2002;3:171–176. doi: 10.1093/embo-reports/kvf024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Wan M, Tang Y, Tytler EM, Lu C, Jin B, Vickers SM, Yang L, Shi X, Cao X. Smad4 protein stability is regulated by ubiquitin ligase SCF β-TrCP1. J Biol Chem. 2004;279:14484–14487. doi: 10.1074/jbc.C400005200. [DOI] [PubMed] [Google Scholar]
- 125.Wan M, Huang J, Jhala NC, Tytler EM, Yang L, Vickers SM, Tang Y, Lu C, Wang N, Cao X. SCF(β-TrCP1) controls Smad4 protein stability in pancreatic cancer cells. Am J Pathol. 2005;166:1379–1392. doi: 10.1016/s0002-9440(10)62356-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Dupont S, Zacchigna L, Cordenonsi M, Soligo S, Adorno M, Rugge M, Piccolo S. Germ-layer specification and control of cell growth by Ectodermin, a Smad4 ubiquitin ligase. Cell. 2005;121:87–99. doi: 10.1016/j.cell.2005.01.033. [DOI] [PubMed] [Google Scholar]
- 127.He W, Dorn DC, Erdjument-Bromage H, Tempst P, Moore MA, Massague J. Hematopoiesis controlled by distinct TIF1γ and Smad4 branches of the TGFβ pathway. Cell. 2006;125:929–941. doi: 10.1016/j.cell.2006.03.045. [DOI] [PubMed] [Google Scholar]
- 128.Dupont S, Mamidi A, Cordenonsi M, Montagner M, Zacchigna L, Adorno M, Martello G, Stinchfield MJ, Soligo S, Morsut L, Inui M, Moro S, Modena N, Argenton F, Newfeld SJ, Piccolo S. FAM/USP9x, a deubiquitinating enzyme essential for TGFβ signaling, controls Smad4 monoubiquitination. Cell. 2009;136:123–135. doi: 10.1016/j.cell.2008.10.051. [DOI] [PubMed] [Google Scholar]
- 129.Agricola E, Randall RA, Gaarenstroom T, Dupont S, Hill CS. Recruitment of TIF1γ to chromatin via its PHD finger-bromodomain activates its ubiquitin ligase and transcriptional repressor activities. Mol Cell. 2011;43:85–96. doi: 10.1016/j.molcel.2011.05.020. [DOI] [PubMed] [Google Scholar]
- 130.Xi Q, Wang Z, Zaromytidou AI, Zhang XH, Chow-Tsang LF, Liu JX, Kim H, Barlas A, Manova-Todorova K, Kaartinen V, Studer L, Mark W, Patel DJ, Massague J. A poised chromatin platform for TGF-β access to master regulators. Cell. 2011;147:1511–1524. doi: 10.1016/j.cell.2011.11.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Moren A, Hellman U, Inada Y, Imamura T, Heldin CH, Moustakas A. Differential ubiquitination defines the functional status of the tumor suppressor Smad4. J Biol Chem. 2003;278:33571–33582. doi: 10.1074/jbc.M300159200. [DOI] [PubMed] [Google Scholar]
- 132.Goto K, Kamiya Y, Imamura T, Miyazono K, Miyazawa K. Selective inhibitory effects of Smad6 on bone morphogenetic protein type I receptors. J Biol Chem. 2007;282:20603–20611. doi: 10.1074/jbc.M702100200. [DOI] [PubMed] [Google Scholar]
- 133.Hayashi H, Abdollah S, Qiu Y, Cai J, Xu YY, Grinnell BW, Richardson MA, Topper JN, Gimbrone MA, Jr, Wrana JL, Falb D. The MAD-related protein Smad7 associates with the TGFβ receptor and functions as an antagonist of TGFβ signaling. Cell. 1997;89:1165–1173. doi: 10.1016/s0092-8674(00)80303-7. [DOI] [PubMed] [Google Scholar]
- 134.Ekman M, Mu Y, Lee SY, Edlund S, Kozakai T, Thakur N, Tran H, Qian J, Groeden J, Heldin CH, Landstrom M. APC and Smad7 link TGFβ type I receptors to the microtubule system to promote cell migration. Mol Biol Cell. 2012 doi: 10.1091/mbc.E10-12-1000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Zhang S, Fei T, Zhang L, Zhang R, Chen F, Ning Y, Han Y, Feng XH, Meng A, Chen YG. Smad7 antagonizes transforming growth factor β signaling in the nucleus by interfering with functional Smad-DNA complex formation. Mol Cell Biol. 2007;27:4488–4499. doi: 10.1128/MCB.01636-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Miyake T, Alli NS, McDermott JC. Nuclear function of Smad7 promotes myogenesis. Mol Cell Biol. 2010;30:722–735. doi: 10.1128/MCB.01005-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Guo X, Wang XF. Signaling cross-talk between TGF-β/BMP and other pathways. Cell Res. 2009;19:71–88. doi: 10.1038/cr.2008.302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Huang M, Sharma S, Zhu LX, Keane MP, Luo J, Zhang L, Burdick MD, Lin YQ, Dohadwala M, Gardner B, Batra RK, Strieter RM, Dubinett SM. IL-7 inhibits fibroblast TGF-β production and signaling in pulmonary fibrosis. J Clin Invest. 2002;109:931–937. doi: 10.1172/JCI14685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Ulloa L, Doody J, Massague J. Inhibition of transforming growth factor-β/SMAD signalling by the interferon-gamma/STAT pathway. Nature. 1999;397:710–713. doi: 10.1038/17826. [DOI] [PubMed] [Google Scholar]
- 140.Bitzer M, von Gersdorff G, Liang D, Dominguez-Rosales A, Beg AA, Rojkind M, Bottinger EP. A mechanism of suppression of TGF-β/SMAD signaling by NF-kappa B/RelA. Genes Dev. 2000;14:187–197. [PMC free article] [PubMed] [Google Scholar]
- 141.Heikkinen PT, Nummela M, Jokilehto T, Grenman R, Kahari VM, Jaakkola PM. Hypoxic conversion of SMAD7 function from an inhibitor into a promoter of cell invasion. Cancer Res. 2010;70:5984–5993. doi: 10.1158/0008-5472.CAN-09-3777. [DOI] [PubMed] [Google Scholar]
- 142.Smith AL, Iwanaga R, Drasin DJ, Micalizzi DS, Vartuli RL, Tan AC, Ford HL. The miR-106b-25 cluster targets Smad7, activates TGF-β signaling, and induces EMT and tumor initiating cell characteristics downstream of Six1 in human breast cancer. Oncogene. 2012 doi: 10.1038/onc.2012.11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Liu G, Friggeri A, Yang Y, Milosevic J, Ding Q, Thannickal VJ, Kaminski N, Abraham E. miR-21 mediates fibrogenic activation of pulmonary fibroblasts and lung fibrosis. J Exp Med. 2010;207:1589–1597. doi: 10.1084/jem.20100035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Koinuma D, Shinozaki M, Komuro A, Goto K, Saitoh M, Hanyu A, Ebina M, Nukiwa T, Miyazawa K, Imamura T, Miyazono K. Arkadia amplifies TGF-β superfamily signalling through degradation of Smad7. Embo J. 2003;22:6458–6470. doi: 10.1093/emboj/cdg632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Liu W, Rui H, Wang J, Lin S, He Y, Chen M, Li Q, Ye Z, Zhang S, Chan SC, Chen YG, Han J, Lin SC. Axin is a scaffold protein in TGF-β signaling that promotes degradation of Smad7 by Arkadia. Embo J. 2006;25:1646–1658. doi: 10.1038/sj.emboj.7601057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Bonni S, Wang HR, Causing CG, Kavsak P, Stroschein SL, Luo K, Wrana JL. TGF-β induces assembly of a Smad2-Smurf2 ubiquitin ligase complex that targets SnoN for degradation. Nat Cell Biol. 2001;3:587–595. doi: 10.1038/35078562. [DOI] [PubMed] [Google Scholar]
- 147.Sharma V, Antonacopoulou AG, Tanaka S, Panoutsopoulos AA, Bravou V, Kalofonos HP, Episkopou V. Enhancement of TGF-β signaling responses by the E3 ubiquitin ligase Arkadia provides tumor suppression in colorectal cancer. Cancer Res. 2011;71:6438–6449. doi: 10.1158/0008-5472.CAN-11-1645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Zhao Y, Thornton AM, Kinney MC, Ma CA, Spinner JJ, Fuss IJ, Shevach EM, Jain A. The deubiquitinase CYLD targets Smad7 protein to regulate transforming growth factor β (TGF-β) signaling and the development of regulatory T cells. J Biol Chem. 2011;286:40520–40530. doi: 10.1074/jbc.M111.292961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Gronroos E, Hellman U, Heldin CH, Ericsson J. Control of Smad7 stability by competition between acetylation and ubiquitination. Mol Cell. 2002;10:483–493. doi: 10.1016/s1097-2765(02)00639-1. [DOI] [PubMed] [Google Scholar]
- 150.Simonsson M, Heldin CH, Ericsson J, Gronroos E. The balance between acetylation and deacetylation controls Smad7 stability. J Biol Chem. 2005;280:21797–21803. doi: 10.1074/jbc.M503134200. [DOI] [PubMed] [Google Scholar]
- 151.Kume S, Haneda M, Kanasaki K, Sugimoto T, Araki S, Isshiki K, Isono M, Uzu T, Guarente L, Kashiwagi A, Koya D. SIRT1 inhibits transforming growth factor β-induced apoptosis in glomerular mesangial cells via Smad7 deacetylation. J Biol Chem. 2007;282:151–158. doi: 10.1074/jbc.M605904200. [DOI] [PubMed] [Google Scholar]
- 152.Koinuma D, Tsutsumi S, Kamimura N, Taniguchi H, Miyazawa K, Sunamura M, Imamura T, Miyazono K, Aburatani H. Chromatin immunoprecipitation on microarray analysis of Smad2/3 binding sites reveals roles of ETS1 and TFAP2A in transforming growth factor β signaling. Mol Cell Biol. 2009;29:172–186. doi: 10.1128/MCB.01038-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Furuhashi M, Yagi K, Yamamoto H, Furukawa Y, Shimada S, Nakamura Y, Kikuchi A, Miyazono K, Kato M. Axin facilitates Smad3 activation in the transforming growth factor β signaling pathway. Mol Cell Biol. 2001;21:5132–5141. doi: 10.1128/MCB.21.15.5132-5141.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Labbe E, Letamendia A, Attisano L. Association of Smads with lymphoid enhancer binding factor 1/T cell-specific factor mediates cooperative signaling by the transforming growth factor-β and wnt pathways. Proc Natl Acad Sci U S A. 2000;97:8358–8363. doi: 10.1073/pnas.150152697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Lei S, Dubeykovskiy A, Chakladar A, Wojtukiewicz L, Wang TC. The murine gastrin promoter is synergistically activated by transforming growth factor-β/Smad and Wnt signaling pathways. J Biol Chem. 2004;279:42492–42502. doi: 10.1074/jbc.M404025200. [DOI] [PubMed] [Google Scholar]
- 156.Jian H, Shen X, Liu I, Semenov M, He X, Wang XF. Smad3-dependent nuclear translocation of β-catenin is required for TGF-β1-induced proliferation of bone marrow-derived adult human mesenchymal stem cells. Genes Dev. 2006;20:666–674. doi: 10.1101/gad.1388806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Blokzijl A, Dahlqvist C, Reissmann E, Falk A, Moliner A, Lendahl U, Ibanez CF. Cross-talk between the Notch and TGF-β signaling pathways mediated by interaction of the Notch intracellular domain with Smad3. J Cell Biol. 2003;163:723–728. doi: 10.1083/jcb.200305112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Samon JB, Champhekar A, Minter LM, Telfer JC, Miele L, Fauq A, Das P, Golde TE, Osborne BA. Notch1 and TGFβ1 cooperatively regulate Foxp3 expression and the maintenance of peripheral regulatory T cells. Blood. 2008;112:1813–1821. doi: 10.1182/blood-2008-03-144980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Masuda S, Kumano K, Shimizu K, Imai Y, Kurokawa M, Ogawa S, Miyagishi M, Taira K, Hirai H, Chiba S. Notch1 oncoprotein antagonizes TGF-β/Smad-mediated cell growth suppression via sequestration of coactivator p300. Cancer Sci. 2005;96:274–282. doi: 10.1111/j.1349-7006.2005.00048.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Zhang Y. Non-Smad TGF-β Signaling Pathways. In: Derynck R, Miyazono K, editors. The TGF-β Family. Cold Spring Harbor Laboratory Press; 2008. pp. 419–438. [Google Scholar]
- 161.Zhang Y, Feng XH, Derynck R. Smad3 and Smad4 cooperate with c-Jun/c-Fos to mediate TGF-β-induced transcription. Nature. 1998;394:909–913. doi: 10.1038/29814. [DOI] [PubMed] [Google Scholar]
- 162.Shaulian E, Karin M. AP-1 as a regulator of cell life and death. Nat Cell Biol. 2002;4:E131–136. doi: 10.1038/ncb0502-e131. [DOI] [PubMed] [Google Scholar]
- 163.Javelaud D, Mauviel A. Crosstalk mechanisms between the mitogen-activated protein kinase pathways and Smad signaling downstream of TGF-β: implications for carcinogenesis. Oncogene. 2005;24:5742–5750. doi: 10.1038/sj.onc.1208928. [DOI] [PubMed] [Google Scholar]
- 164.Kang Y, Chen CR, Massague J. A self-enabling TGFβ response coupled to stress signaling: Smad engages stress response factor ATF3 for Id1 repression in epithelial cells. Mol Cell. 2003;11:915–926. doi: 10.1016/s1097-2765(03)00109-6. [DOI] [PubMed] [Google Scholar]
- 165.Wu G, Chen YG, Ozdamar B, Gyuricza CA, Chong PA, Wrana JL, Massague J, Shi Y. Structural basis of Smad2 recognition by the Smad anchor for receptor activation. Science. 2000;287:92–97. doi: 10.1126/science.287.5450.92. [DOI] [PubMed] [Google Scholar]
- 166.Tsukazaki T, Chiang TA, Davison AF, Attisano L, Wrana JL. SARA, a FYVE domain protein that recruits Smad2 to the TGFβ receptor. Cell. 1998;95:779–791. doi: 10.1016/s0092-8674(00)81701-8. [DOI] [PubMed] [Google Scholar]
- 167.Miura S, Takeshita T, Asao H, Kimura Y, Murata K, Sasaki Y, Hanai JI, Beppu H, Tsukazaki T, Wrana JL, Miyazono K, Sugamura K. Hgs (Hrs), a FYVE domain protein, is involved in Smad signaling through cooperation with SARA. Mol Cell Biol. 2000;20:9346–9355. doi: 10.1128/mcb.20.24.9346-9355.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Lin HK, Bergmann S, Pandolfi PP. Cytoplasmic PML function in TGF-β signalling. Nature. 2004;431:205–211. doi: 10.1038/nature02783. [DOI] [PubMed] [Google Scholar]
- 169.Hocevar BA, Smine A, Xu XX, Howe PH. The adaptor molecule Disabled-2 links the transforming growth factor β receptors to the Smad pathway. Embo J. 2001;20:2789–2801. doi: 10.1093/emboj/20.11.2789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Penheiter SG, Singh RD, Repellin CE, Wilkes MC, Edens M, Howe PH, Pagano RE, Leof EB. Type II transforming growth factor-β receptor recycling is dependent upon the clathrin adaptor protein Dab2. Mol Biol Cell. 2010;21:4009–4019. doi: 10.1091/mbc.E09-12-1019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171.Iwata J, Hacia JG, Suzuki A, Sanchez-Lara PA, Urata M, Chai Y. Modulation of noncanonical TGF-β signaling prevents cleft palate in TGFbr2 mutant mice. J Clin Invest. 2012;122:873–885. doi: 10.1172/JCI61498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172.Rudini N, Felici A, Giampietro C, Lampugnani M, Corada M, Swirsding K, Garre M, Liebner S, Letarte M, ten Dijke P, Dejana E. VE-cadherin is a critical endothelial regulator of TGF-β signalling. Embo J. 2008;27:993–1004. doi: 10.1038/emboj.2008.46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173.Barrios-Rodiles M, Brown KR, Ozdamar B, Bose R, Liu Z, Donovan RS, Shinjo F, Liu Y, Dembowy J, Taylor IW, Luga V, Przulj N, Robinson M, Suzuki H, Hayashizaki Y, Jurisica I, Wrana JL. High-throughput mapping of a dynamic signaling network in mammalian cells. Science. 2005;307:1621–1625. doi: 10.1126/science.1105776. [DOI] [PubMed] [Google Scholar]
- 174.Yamashita M, Fatyol K, Jin C, Wang X, Liu Z, Zhang YE. TRAF6 mediates Smad-independent activation of JNK and p38 by TGF-β. Mol Cell. 2008;31:918–924. doi: 10.1016/j.molcel.2008.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175.Sorrentino A, Thakur N, Grimsby S, Marcusson A, von Bulow V, Schuster N, Zhang S, Heldin CH, Landstrom M. The type I TGF-β receptor engages TRAF6 to activate TAK1 in a receptor kinase-independent manner. Nat Cell Biol. 2008;10:1199–1207. doi: 10.1038/ncb1780. [DOI] [PubMed] [Google Scholar]
- 176.Cejas PJ, Walsh MC, Pearce EL, Han D, Harms GM, Artis D, Turka LA, Choi Y. TRAF6 inhibits Th17 differentiation and TGF-β-mediated suppression of IL-2. Blood. 2010;115:4750–4757. doi: 10.1182/blood-2009-09-242768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Scaffidi AK, Petrovic N, Moodley YP, Fogel-Petrovic M, Kroeger KM, Seeber RM, Eidne KA, Thompson PJ, Knight DA. α(v)β(3) Integrin interacts with the transforming growth factor β (TGFβ) type II receptor to potentiate the proliferative effects of TGFβ1 in living human lung fibroblasts. J Biol Chem. 2004;279:37726–37733. doi: 10.1074/jbc.M403010200. [DOI] [PubMed] [Google Scholar]
- 178.Bourguignon LY, Singleton PA, Zhu H, Zhou B. Hyaluronan promotes signaling interaction between CD44 and the transforming growth factor β receptor I in metastatic breast tumor cells. J Biol Chem. 2002;277:39703–39712. doi: 10.1074/jbc.M204320200. [DOI] [PubMed] [Google Scholar]
- 179.Partridge EA, Le Roy C, Di Guglielmo GM, Pawling J, Cheung P, Granovsky M, Nabi IR, Wrana JL, Dennis JW. Regulation of cytokine receptors by Golgi N-glycan processing and endocytosis. Science. 2004;306:120–124. doi: 10.1126/science.1102109. [DOI] [PubMed] [Google Scholar]
- 180.Felici A, Wurthner JU, Parks WT, Giam LR, Reiss M, Karpova TS, McNally JG, Roberts AB. TLP, a novel modulator of TGF-β signaling, has opposite effects on Smad2- and Smad3-dependent signaling. Embo J. 2003;22:4465–4477. doi: 10.1093/emboj/cdg428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Kwak JH, Kim SI, Kim JK, Choi ME. BAT3 interacts with transforming growth factor-β (TGF-β) receptors and enhances TGF-β1-induced type I collagen expression in mesangial cells. J Biol Chem. 2008;283:19816–19825. doi: 10.1074/jbc.M802285200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Jin Q, Ding W, Mulder KM. Requirement for the dynein light chain km23–1 in a Smad2-dependent transforming growth factor-β signaling pathway. J Biol Chem. 2007;282:19122–19132. doi: 10.1074/jbc.M609915200. [DOI] [PubMed] [Google Scholar]
- 183.Neil JR, Tian M, Schiemann WP. X-linked inhibitor of apoptosis protein and its E3 ligase activity promote transforming growth factor-{β}-mediated nuclear factor-{kappa}B activation during breast cancer progression. J Biol Chem. 2009;284:21209–21217. doi: 10.1074/jbc.M109.018374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Kowanetz M, Lonn P, Vanlandewijck M, Kowanetz K, Heldin CH, Moustakas A. TGFβ induces SIK to negatively regulate type I receptor kinase signaling. J Cell Biol. 2008;182:655–662. doi: 10.1083/jcb.200804107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Yamaguchi T, Kurisaki A, Yamakawa N, Minakuchi K, Sugino H. FKBP12 functions as an adaptor of the Smad7-Smurf1 complex on activin type I receptor. J Mol Endocrinol. 2006;36:569–579. doi: 10.1677/jme.1.01966. [DOI] [PubMed] [Google Scholar]
- 186.Watanabe Y, Itoh S, Goto T, Ohnishi E, Inamitsu M, Itoh F, Satoh K, Wiercinska E, Yang W, Shi L, Tanaka A, Nakano N, Mommaas AM, Shibuya H, Ten Dijke P, Kato M. TMEPAI, a transmembrane TGF-β-inducible protein, sequesters Smad proteins from active participation in TGF-β signaling. Mol Cell. 2010;37:123–134. doi: 10.1016/j.molcel.2009.10.028. [DOI] [PubMed] [Google Scholar]
- 187.Su Y, Zhang L, Gao X, Meng F, Wen J, Zhou H, Meng A, Chen YG. The evolutionally conserved activity of Dapper2 in antagonizing TGF-β signaling. Faseb J. 2007;21:682–690. doi: 10.1096/fj.06-6246com. [DOI] [PubMed] [Google Scholar]
- 188.Zhang L, Zhou H, Su Y, Sun Z, Zhang H, Zhang L, Zhang Y, Ning Y, Chen YG, Meng A. Zebrafish Dpr2 inhibits mesoderm induction by promoting degradation of nodal receptors. Science. 2004;306:114–117. doi: 10.1126/science.1100569. [DOI] [PubMed] [Google Scholar]
- 189.Zhang Y, Li X, Qi J, Wang J, Liu X, Zhang H, Lin SC, Meng A. Rock2 controls TGFβ signaling and inhibits mesoderm induction in zebrafish embryos. J Cell Sci. 2009;122:2197–2207. doi: 10.1242/jcs.040659. [DOI] [PubMed] [Google Scholar]
- 190.Datta PK, Moses HL. STRAP and Smad7 synergize in the inhibition of transforming growth factor β signaling. Mol Cell Biol. 2000;20:3157–3167. doi: 10.1128/mcb.20.9.3157-3167.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Lonn P, Vanlandewijck M, Raja E, Kowanetz M, Watanabe Y, Kowanetz K, Vasilaki E, Heldin CH, Moustakas A. Transcriptional induction of salt-inducible kinase 1 by transforming growth factor β leads to negative regulation of type I receptor signaling in cooperation with the Smurf2 ubiquitin ligase. J Biol Chem. 2012 doi: 10.1074/jbc.M111.307249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Seong HA, Jung H, Choi HS, Kim KT, Ha H. Regulation of transforming growth factor-β signaling and PDK1 kinase activity by physical interaction between PDK1 and serine-threonine kinase receptor-associated protein. J Biol Chem. 2005;280:42897–42908. doi: 10.1074/jbc.M507539200. [DOI] [PubMed] [Google Scholar]
- 193.Yan X, Lin Z, Chen F, Zhao X, Chen H, Ning Y, Chen YG. Human BAMBI cooperates with Smad7 to inhibit transforming growth factor-β signaling. J Biol Chem. 2009;284:30097–30104. doi: 10.1074/jbc.M109.049304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194.Onichtchouk D, Chen YG, Dosch R, Gawantka V, Delius H, Massague J, Niehrs C. Silencing of TGF-β signalling by the pseudoreceptor BAMBI. Nature. 1999;401:480–485. doi: 10.1038/46794. [DOI] [PubMed] [Google Scholar]
- 195.Goumans MJ, Valdimarsdottir G, Itoh S, Lebrin F, Larsson J, Mummery C, Karlsson S, ten Dijke P. Activin receptor-like kinase (ALK)1 is an antagonistic mediator of lateral TGFβ/ALK5 signaling. Mol Cell. 2003;12:817–828. doi: 10.1016/s1097-2765(03)00386-1. [DOI] [PubMed] [Google Scholar]
- 196.Lallemand F, Seo SR, Ferrand N, Pessah M, L'Hoste S, Rawadi G, Roman-Roman S, Camonis J, Atfi A. AIP4 restricts transforming growth factor-β signaling through a ubiquitination-independent mechanism. J Biol Chem. 2005;280:27645–27653. doi: 10.1074/jbc.M500188200. [DOI] [PubMed] [Google Scholar]
- 197.Ferrigno O, Lallemand F, Verrecchia F, L'Hoste S, Camonis J, Atfi A, Mauviel A. Yes-associated protein (YAP65) interacts with Smad7 and potentiates its inhibitory activity against TGF-β/Smad signaling. Oncogene. 2002;21:4879–4884. doi: 10.1038/sj.onc.1205623. [DOI] [PubMed] [Google Scholar]
- 198.Ferrand N, Atfi A, Prunier C. The oncoprotein c-ski functions as a direct antagonist of the transforming growth factor-{β} type I receptor. Cancer Res. 2010;70:8457–8466. doi: 10.1158/0008-5472.CAN-09-4088. [DOI] [PubMed] [Google Scholar]
- 199.Jin W, Yun C, Kwak MK, Kim TA, Kim SJ. TrkC binds to the type II TGF-β receptor to suppress TGF-β signaling. Oncogene. 2007;26:7684–7691. doi: 10.1038/sj.onc.1210571. [DOI] [PubMed] [Google Scholar]
- 200.Meng Q, Lux A, Holloschi A, Li J, Hughes JM, Foerg T, McCarthy JE, Heagerty AM, Kioschis P, Hafner M, Garland JM. Identification of Tctex2β, a novel dynein light chain family member that interacts with different transforming growth factor-β receptors. J Biol Chem. 2006;281:37069–37080. doi: 10.1074/jbc.M608614200. [DOI] [PubMed] [Google Scholar]
- 201.Jin W, Kim BC, Tognon C, Lee HJ, Patel S, Lannon CL, Maris JM, Triche TJ, Sorensen PH, Kim SJ. The ETV6-NTRK3 chimeric tyrosine kinase suppresses TGF-β signaling by inactivating the TGF-β type II receptor. Proc Natl Acad Sci U S A. 2005;102:16239–16244. doi: 10.1073/pnas.0503137102. [DOI] [PMC free article] [PubMed] [Google Scholar]