Abstract
Trees are used to produce a variety of wood‐based products including timber, pulp and paper. More recently, their use as a source of renewable energy has also been highlighted, as has their value for carbon mitigation within the Kyoto Protocol. Relative to food crops, the domestication of trees has only just begun; the long generation time and complex nature of juvenile and mature growth forms are contributory factors. To accelerate domestication, and to understand further some of the unique processes that occur in woody plants such as dormancy and secondary wood formation, a ‘model’ tree is needed. Here it is argued that Populus is rapidly becoming accepted as the ‘model’ woody plant and that such a ‘model’ tree is necessary to complement the genetic resource being developed in arabidopsis. The genus Populus (poplars, cottonwoods and aspens) contains approx. 30 species of woody plant, all found in the Northern hemisphere and exhibiting some of the fastest growth rates observed in temperate trees. Populus is fulfilling the ‘model’ role for a number of reasons. First, and most important, is the very recent commitment to sequence the Populus genome, a project initiated in February 2002. This will be the first woody plant to be sequenced. Other reasons include the relatively small genome size (450–550 Mbp) of Populus, the large number of molecular genetic maps and the ease of genetic transformation. Populus may also be propagated vegetatively, making mapping populations immortal and facilitating the production of large amounts of clonal material for experimentation. Hybridization occurs routinely and, in these respects, Populus has many similarities to arabidopsis. However, Populus also differs from arabidopsis in many respects, including being dioecious, which makes selfing and back‐cross manipulations impossible. The long time‐to‐flower is also a limitation, whilst physiological and biochemical experiments are more readily conducted in Populus compared with the small‐statured arabidopsis. Recent advances in the development of large expressed sequence tagged collections, microarray analysis and the free distribution of mapping pedigrees for quantitative trait loci analysis secure Populus as the ideal subject for further exploitation by a wide range of scientists including breeders, physiologists, biochemists and molecular biologists. In addition, and in contrast to other model plants, the genus Populus also has genuine commercial value as a tree for timber, plywood, pulp and paper.
Key words: Review, Populus, poplar, model plant, genomics, QTL, arabidopsis, genome sequence
INTRODUCTION
The need for sustainable products from plants has never been greater. In many respects, this is driven by the move away from oil‐based economies that is likely to develop during this century. Trees are unique amongst plants since they have extreme longevity and are characterized by the ability to generate woody biomass from secondary xylem tissue formed from the vascular cambium. The genus Populus makes an important contribution to meeting the global need for paper, timber and other wood‐based products. The role of fast‐growing trees, like Populus, in carbon mitigation aligned to the Kyoto Protocol is also being quantified and may be considerable. Smith et al. (2000) have shown that in Europe, of all options examined, bioenergy crops show the greatest potential for carbon mitigation. Bioenergy crop‐production also shows an indefinite mitigation potential compared with other options where the mitigation potential is finite (Tuskan and Walsh, 2001).
The necessity for model species of plants is well recognized and, in this role, Arabidopsis thaliana has gained a supreme acceptance amongst plant scientists. The reasons for this are clear. It is a small plant with a small genome, a rapid life cycle and some value in experimental studies at the physiological and biochemical levels (Goodman et al., 1995). It is easily transformed and has a wide natural distribution. This natural genetic variation may be exploited to produce crosses to follow segregation and, in the creation of genetic maps, recombinant inbred lines (RILs) and near isogeneic lines (NILs). The recent completion of sequencing of the entire physical genome (AGI, 2000) should be of major benefit to many researchers. Coupled with the large investment now committed to genomics using large collections of ESTs (expressed sequence tags) and knock‐out mutations (Walbot, 1999), the future of arabidopsis as a central resource for plant science is ensured. Recently, it was suggested that more than 7000 researchers are using arabidopsis as an experimental system (Huala et al., 2001).
It has often been considered, however, that arabidopsis cannot be a useful model for crop plants since the most important of these—rice, wheat and maize—are monocotyledonous, and because the physiology, biochemistry and development of these species may not be adequately studied in a dicotyledonous weed (Bennetzen et al., 1998). This suggests that a full understanding of the biology of monocotyledonous crops is unlikely to be achieved using arabidopsis. Thus, other systems for food plants are needed and are being developed, as evidenced recently by the sequencing of the genome of rice (Yu et al., 2002).
Whether arabidopsis can act as the model for a tree can also be questioned. Trees differ fundamentally from other plant species in that they are adapted to survive on a long time‐scale. The most obvious manifestation of this is the development of wood, or secondary xylem, from the vascular cambium. This secondary meristem is essential for tree growth and development and in providing support for a tall structure. It is difficult to imagine that the formation of secondary xylem can ever be studied in anything other than a true tree. Surprisingly, however, arabidopsis has proved useful in this role: many xylem‐forming genes from arabidopsis have been shown to be present in wood‐forming tissues of pine (Allona et al., 1998).
In addition to secondary xylem, trees also exhibit complex patterns of activity and dormancy and the control of, for example, bud‐burst must involve a complex of interactions between environmental signals (including day‐length and temperature) and plant signal transduction pathways. It is hard to imagine that arabidopsis could be adequate for the study of such processes in their entirety, since control may not be at the level of gene expression. Similarly, the age at which plants flower varies in woody species, occurring approx. 1 (Salix), 6 (Populus) and 60 (Quercus) years after germination. Despite this, many of the genes that control flowering in arabidopsis are probably present in trees, and recent reports suggest that some may be used to promote flowering in transgenic Populus (Weigel and Nilsson, 1995), a result of considerable value to breeding and improvement of long‐lived trees that are slow to flower. There is little doubt, then, that arabidopsis is an important model for trees, but it is also apparent that true woody systems are necessary to investigate some ‘unique’ tree processes.
Here it is proposed that the woody genus Populus (poplars, aspens and cottonwoods) has already proved valuable as a model and that recent developments in functional genomics and molecular genetics and biology will accelerate this position. The recent commitment in the USA to sequence the Populus genome can only further strengthen the role of Populus as the model tree.
THE NATURAL GENETIC VARIATION ACROSS THE POPULUS GENUS
Natural genetic variation is a vital tool for future understanding and exploitation in biological systems. Arabidopsis has a very wide global distribution, which has been used to great effect in developing recombinant inbred lines, such as those of Lister and Dean (1993) and Alonso‐Blanco and Koornneef (2000).
Genetic variation within the Populus genome is also considerable. This genus contains between 22 and 75 species, depending on which taxonomic classification is considered (Eckenwalder, 1996), although it is reasonable to assume approx. 30 species, with some broad agreement that clear morphological characteristics are present. It is accepted that the genus can be split into five sections and a summary of their distribution, main characteristics and species is given in Table 1.
Table 1.
Main species of Populus (poplars, cottonwoods and aspen) and their occurrence globally
| Section | Leaf characteristics | Distribution | Major species |
| Leuce (Populus) White and grey poplar and aspens | Large, three lobed or palmate leaves. Dense white hairs | North and Central Europe | P. alba (white poplar) |
| Europe, Western Asia, North Africa | P. tremula (aspen) | ||
| North America | P. tremuloides (quaking aspen) | ||
| Aigeiros Black poplars and cottonwoods | Large, deltoid, cordate | Eastern North America | P. deltoides (eastern cottonwood) |
| Western USA | P. fremontii | ||
| Europe to central Asia | P. nigra (black poplar) | ||
| North America (east of Rocky mountains from Saskatchewan to New Mexico; west to Texas). | P. sargentii | ||
| USA (west Texas, New Mexico) | P. wisilizeni | ||
| Tacamahaca Balsam poplars | Small–large ovate‐lanceolate. Underside silvery‐brownish. Without translucent margins | USA (Montana and south Dakota to New Mexico and Arizona) | P. acuminata |
| Western USA (east of Rocky mountains) | P. angustiolia | ||
| North‐eastern USA | P. balsamifera | ||
| North‐western Chile to Manchuria and Korea | P. cathayana | ||
| Himalayas | P. ciliata | ||
| Korea | P. koreana | ||
| Siberia | P. laurifolia | ||
| North‐eastern Asia, Japan | P. maximowiczii | ||
| North‐western China | P. purdomii | ||
| North and west central China | P. simonii | ||
| East Turkey, Siberia to Russia, Far East | P suaveolens | ||
| Western China | P. szechuanica | ||
| Alaska and British Columbia to California. Rocky mountain plains (Idaho to Montana) | P. trichocarpa | ||
| Central Asia | P. tristis | ||
| South‐west China | P. yunnanensis | ||
| Leucoides | Very large, cordate, underside violet when young | Eastern USA | P. heterophylla |
| Central and Western China | P. lasiocarpa (Chinese necklace poplar) | ||
| China | P. violascens | ||
| Central and Western China | P. wilsonii | ||
| Turanga | Lanceolate‐linear | Central Asia to North Africa | P. euphratica |
Approx. 30 species make up the Populus genus with a Northern hemisphere distribution. Modified from Eckenwalder (1996).
Genetic variation and genetic relatedness have been studied at the molecular level in only a limited number of Populus species, but an excellent example is an on‐going study of P. nigra, the native European poplar, where plants from across widely differing European climates have been collected and grown in several European countries. Molecular genetic analysis has been employed to determine genetic diversity and to assess the contribution of clonal and sexual reproduction to populations of P. nigra as well as the likely threats to native plants in the future. The utilization of genetic variation in arabidopsis has focused on developing RILS, NILs and in using ecotypes. In trees, there are no RILs, largely because of the time‐to‐flower limitation, but also because many breeding systems are difficult in woody plants and ‘back‐crossing’ and ‘near isogenic lines’ (lines that have been repeatedly back‐crossed to isolate a trait of interest) are impossible to generate. However, of the many woody systems of interest, Populus is perhaps most tractable for several reasons. First, it hybridizes readily (Stettler et al., 1996), and even across sections crosses are possible with the minimum of intervention; for example, using embryo rescue (Stanton and Villar, 1996). Once an F1 is produced, it can be immortal, as are other progeny, because Populus is readily propagated from both hardwood and softwood cuttings (Bradshaw, 1996). Nevertheless, this does not overcome the limitations imposed by the absence of true RILs. A very new approach to this problem utilizes the concept of linkage disequilibrium mapping, where divergent genotypes may be utilized to identify putative molecular markers for traits.
THE GENOME OF POPLAR
In many tree species the genome is large and complex, making any type of genetic characterization or manipulation more difficult than that in arabidopsis or even in monocotyledonous crop plants. Table 2 illustrates this point for eucalyptus and pines. The Pinus (pine) genome in particular is complex and unwieldy. Interestingly, however, the genome size of Populus is small, relative to that of other trees. Work on poplar genetic maps has confirmed the presence of 19 linkage groups (Bradshaw et al., 1994; Cervera et al., 2001) with similar genetic‐physical map distances to that of arabidopsis (Bradshaw et al., 2000).
Table 2.
The genome size of Populus relative to that of other tree species and the model plant Arabidopsis thaliana
| Species | Genome size (Mbp) |
| Arabidopsis | 100–150 |
| Pinus | 20 000 |
| Populus/Salix | 450–550 |
GENETIC MAPS IN POPLAR
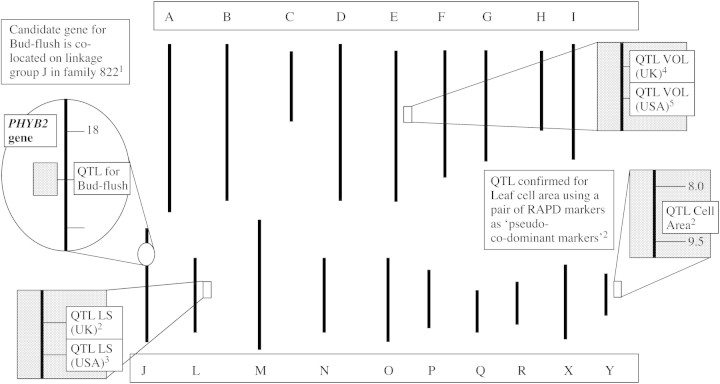
The advent of DNA‐based markers enabled rapid development of genetic maps in species such as arabidopsis (http://nasc.nott.ac.uk/RI_data/how_to_map.html), and these have provided a valuable resource for map‐based cloning of very many useful traits (Gibson and Somerville, 1993). With considerable foresight, the first molecular genetic linkage map of Populus was developed by Bradshaw (1996) from an original cross made in the 1970s at the University of Washington, USA. This cross led to the development of family 331, a reasonably large F2 progeny, and a molecular genetic map that now has several hundred molecular markers (Bradshaw et al., 1994). The maternal parent, P. trichocarpa, clone 93–968 of the native black cottonwood from western Washington state, was hybridized with the male clone ILL‐129 of the eastern cottonwood P. deltoides from Illinois. Two siblings of the resulting family 53 were crossed in 1988 to produce family 331. This genetic resource is similar to the RILs for arabidopsis developed by Lister and Dean (1993) and has provided a framework for numerous studies since its original development. It has been used to map quantitative trait loci (QTL) for a number of traits of importance to yield and disease resistance, some of which are shown in Fig. 1, whilst the diversity in leaf morphology in the parents is illustrated in Fig. 2. Mapped QTL include stem growth, leaf morphology and branch traits such as sylleptic production. Leaf cell traits have also been mapped recently as QTL in this pedigree (Taylor et al., 2001b), as have stomatal traits, including a QTL for stomatal index, a quantification of stomatal initiation (Ferris et al., 2002). The value of this original molecular genetic resource cannot be underestimated, since on publication, few such maps were available for woody species. In future, these parents will be utilized on microarrays, where identification of candidate genes and their map position should be rapid. Several other molecular genetic maps of Populus have been developed, mostly with F1 progeny using the pseudo test‐cross strategy described by Grattapaglia and Sederoff (1994). This includes a Belgian cross between P. trichocarpa, P. nigra and P. deltoides (Cervera et al., 2001) and a French cross using P. nigra and P. deltoides (reported by Cervera et al., 1997). A new map for P. deltoides was also reported recently by Wu et al. (2000b), based entirely on AFLP markers.
Fig. 1. Summary schematic of molecular genetic map of Populus and the occurrence of some QTL for leaf morphological and wood‐yield traits. The genome is diploid and consists of 19 linkage groups, here labelled A–Y. The inset shows the candidate gene approach used to map candidate gene PhyB2 on the Populus genetic map and co‐locating for the QTL ‘date of bud‐flush’. Other traits are for QTL located either in contrasting growing environments of the UK and USA (LS, leaf size; VOL, stem volume) or for traits confirmed through other analyses (Cell Area, leaf epidermal cell area). Data modified from Bradshaw et al. (1994), 1 Frewen et al. (2000), 2 Taylor et al. (2001b), 3 Wu (1998), 5 Bradshaw (1996) and 4 G.Taylor et al. (pers. comm.).
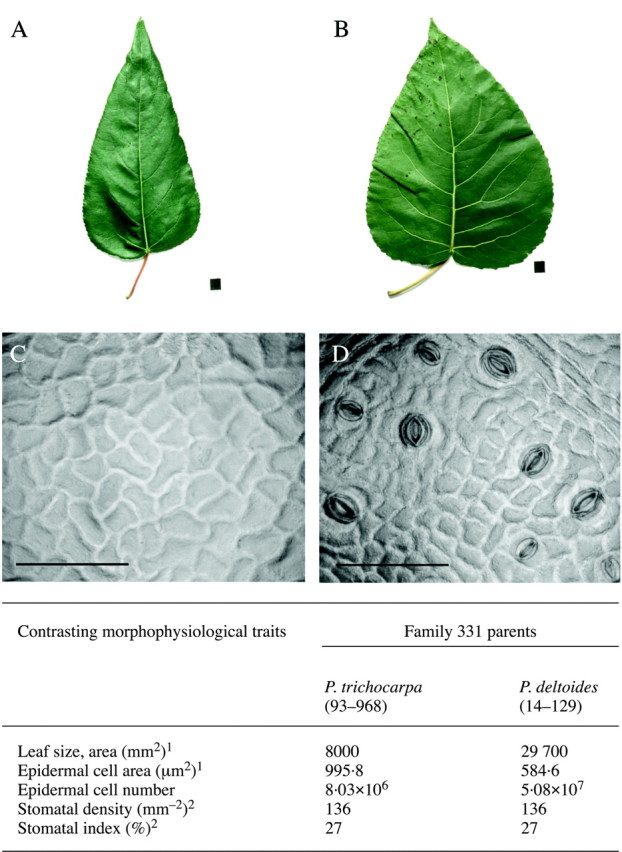
Fig. 2. Contrasting parents of the mapping pedigree ‘family 331’, where a cross between a female P. trichocarpa (A) and male P. deltoides (B) was used to generate an F1 and then F2 pedigree. Black square = 1 cm2. Images of adaxial epidermal cells are also shown for both parents (C, P. trichocarpa; D, P. deltoides). Bar = 100 µm. The table illustrates the contrasting morphophysiological characteristics that have been used to detect a wide range of QTL in this pedigree. 1 Taylor et al. (2001a), 2 Ferris et al. (2002). Photographic image reproduced from Ferris et al. (2002), with permission.
Together these maps form a powerful resource for two reasons. First, they will be ‘anchored’ in future, enriching the markers available for mapping traits in a generic Populus map. Markers may then be linked to traits for yield and disease resistance and used in a programme of advanced tree selection, improvement and breeding. In this instance the gene behind a QTL may be of little significance since the value of the study is in developing a marker that can predict yield quality or quantity. This has yet to become a reality in forest tree breeding, although if any system offered great potential it is that of forest trees. Populus is an excellent model in which to test the idea of molecular breeding of trees through marker assisted selection, since a number of QTL for yield‐related traits (Wu, 1998) appear to be fairly robust in that they have co‐located on the molecular genetic map for plants grown in more than one geographical location (Bradshaw, 1996). This is true for stem height (see Fig. 1), which mapped to similar chromosomal locations for material grown in the USA and UK (Taylor et al., 2001b). Wu et al (2000a) believe that molecular tree breeding will become a reality in the future.
The second use of molecular genetic maps is in determining the link between QTL and the underlying gene or genes. Map‐based cloning may be relatively straightforward in arabidopsis and even in other plants, for example in determining a gene controlling sugar content in tomato (Fridman et al., 2000). However, in the absence of recombinant inbred lines and near isogenic lines, and with the large map‐distances and small progeny numbers that exist in trees, this can be extremely difficult and more akin to some of the problems faced in mammals (Nadeau and Frankel, 2000). Despite this, considerable progress has been made in some areas. One such area is in understanding the genetic basis of resistance to the rust Melampsora spp. that is critical for the commercial development of Populus plantations. Here, a quantitative genetic study of resistance to the US species M. medusae f. sp. deltoidae has identified a locus, Mmd1, that plays a major role in resistance to this rust in the Pacific Northwest. Mmd1 was mapped to linkage group Q, approx. 5 centimorgan (cM) from a restriction fragment length polymorphism (RFLP) marker, P222 (Newcombe et al., 1996). Advanced research is also in progress to clone the QTL of major importance in determining resistance to the European M. larici‐populina. A genomic region involved in qualitative resistance has also been tagged, with a random amplified polymorphic DNA (RAPD) marker OPM03/04_480, approx. 1 cM from the R locus (Villar et al., 1996). Recently, further fine mapping for the qualitative resistance locus (MER) was reported by Zhang et al. (2001). Not only does Populus have probably the most extensive range of molecular genetic maps, which themselves will act as an important model in which to study a variety of biological and tree‐specific phenomena, but the genus is also at the forefront for the development of tree genomics. In bringing these two technologies together, there is real hope that candidate genes can be identified and mapped more rapidly than was previously possible, making ‘map‐based’ cloning in a tree species a reality. A candidate gene approach in Populus was recently described by Frewen et al. (2001), where the genes ABI3 and PhyB were mapped in family 822 and found to co‐locate with a QTL for bud‐burst and bud set (Fig. 1), suggesting that in the future such candidate genes can be mapped and investigated.
THE ABILITY TO TRANSFORM POPLAR
Any model plant must be readily transformable, preferably using a number of techniques, and to a high efficiency, not overly sensitive to the specifics of protocol (Birch, 1997). For arabidopsis, transformation has become a routine tool to study the expression of a vast array of gene constructs that may be identified through both forward and reverse genetic approaches. This is linked to the use of several reporter genes including β‐glucoronidase (GUS; Jefferson et al., 1987) and the green fluorescent protein (GFP) from Aquorea victoria (Haseloff et al., 1997). More recently, in planta techniques of transformation that do not require tissue regeneration, for example exposure to Agrobacterium using vacuum filtration, have been developed for arabidopsis (Feldmann and Marks, 1987). The latest technique involves ‘dipping’ plants or shoot pieces into an appropriate solution (Clough and Bent, 1998).
Populus was one of the first woody systems to undergo genetic transformation; this was noted as early as 1970 and has been reviewed by Han et al. (1996). The key to this early success was the ease with which in vitro cultures could be generated. All of the routine methods of plant transformation (Agrobacterium mediated, direct DNA transfer, electroporation) have been tried and have proved successful with Populus. The current emphasis, however, is on Agrobacterium‐mediated transformation and on improving regeneration efficiency in commercially relevant genotypes. Currently, there are at least three areas of interest that dominate molecular transformation research in the genus Populus. These are the control of flowering (Rottmann et al., 2000), engineering of Bt and glyphosate resistance (Meilan et al., 2000), and the manipulation in lignin quality and quantity during wood formation. Using wood formation as an example, it is easy to demonstrate the value of Populus as a model (Mellerowicz et al., 2001). Understanding wood formation and manipulating lignin content have a high commercial value. Lignin formation is an essential part of the development of xylem tissue in vascular plants. Although poplar wood (which is lignin‐rich) has many uses, including timber, plywood and pallets, one major use is in the pulp and paper industries, where lignin must be removed in energy‐demanding processes that currently require considerable chemical input. Lignin bioengineering would now seem a real possibility (Chen et al., 2001) following the initial discovery that enzymes in the complex lignin biosynthetic pathway, such as cinnamyl alcohol dehydrogenase (CAD), could be down‐regulated using transformed Populus (Baucher et al., 1996). Recent work has shown that down‐regulation of the lignin biosynthetic pathway gene Pt4CL1 encoding 4‐coumarate‐coenzyme A ligase using antisense technology in P. tremuloides resulted in a 45 % reduction in lignin content, whilst the amount of cellulase was increased by 15 %. Gene silencing has also been used for lignin modification in P. tremula × P. alba, with the gene encoding caffeic acid O‐methyltransferase (COMT) silenced. This resulted in the production of trees with a 17 % decrease in lignin and an associated improvement in pulp yield (Jouanin et al., 2000). These examples not only illustrate the value of Populus as a model for genetic transformation, but also suggest that such trees could provide an improved source of pulp which is less dependent on chemical inputs.
EXPERIMENTAL MANIPULATIONS: PHYSIOLOGY AND BIOCHEMISTRY
If a model plant is one in which experimental manipulation and measurements can be easily performed, then Populus is an ideal choice and, in many respects, is even preferable to arabidopsis, which may be difficult to use in physiological and biochemical studies given its small stature. For Populus, considerable progress has been made in determining the physiological basis of yield and relating yield characteristics to specific genotypes. Yield in a tree crop does not involve reproductive structures and so is often developmentally less complex than that in many food‐crops. Indeed, the following analysis shows that many of the traits ‘selected’ for high yield in F1 hybrids of Populus are actually the opposite [particularly for leaf size, branching and leaf are index (LAI)], to those that have been bred into monocotyledonous crops such as maize over the last hundred years or so (Duvick, 2001). In Populus, there is particular emphasis on the rapid growth of young trees for pulp and paper and also for biomass. Yield, along with disease resistance, is a top priority for future improvement and planting of this forest tree. Ceulemans (1990) has quantified yield traits for several European and US genotypes at the cell, leaf and canopy level. This information has been used to effect in developing models to describe canopy development in Populus (Chen et al., 1994). Complementary to this, models using a photosynthesis and gas‐exchange approach have also been developed (Rauscher et al., 1990).
Following on from these and other studies, an ‘ideotype’ for high yield in Populus includes: (a) large leaves, produced with fast leaf expansion rates (Ridge et al., 1986); (b) rapid development and long duration of high LAI (Ceulemans, 1990); (c) increased ‘branchiness’ and, in particular, the development of sylleptic branches (those formed from current‐year meristems in the axils of current‐year leaves; Ceulemans et al., 1992; Wu and Stettler, 1996); (d) high leaf photosynthetic and respiration rates (Isebrands et al., 1988); (e) highly branched rooting systems; (f) stomata responsive to prevailing environmental conditions (Braatne et al., 1992); (g) erectophile leaves and branches (Dickmann et al, 1990; Wu, 1994).
Populus makes an excellent subject for any such study since it is fast‐growing (one of the fastest growing temperate trees) and because the use of clonal material ensures that experiments are repeatable and with small error. Many physiological studies of Populus confirm this, in contrast to virtually all other tree species, where little selection has taken place and where the use of seedlings can result in large experimental errors and difficulty in designing valid experiments. Other physiological parameters that have been characterized in Populus in relation to the environment include response to nutrient and water stress (Chen et al., 1997; Tschaplinski et al., 1998) and the responses of trees to increased atmospheric CO2 (Ceulemans et al., 1997; Ferris et al., 2001; Taylor et al., 2001a). Perhaps some of the best progress has been made in understanding plant response to ozone pollution, where Populus has easily provided a superior tree‐model for developing and testing hypotheses on both short‐term acute and long‐term chronic effects of this stress. Using natural variation in response to ozone as a tool to understand physiological mechanisms, Coleman et al. (1995) were able to show that ozone tolerance was related to the maintenance of high photosynthetic rates in recently mature leaves and retention of lower leaves. Rapid photosynthetic decline and accelerated senescence are also important variables in the response of Populus to ozone (Bredley and Pell, 1998). However, few effects on ozone tolerance were found between wild‐type and transformed plants over‐expressing glutathione synthetase, suggesting a secondary role for glutathione in the ozone response (Strohm et al., 1999). In contrast, more recent research on Populus suggests a role for salicylic and jasmonic acid signalling pathways, analogous to the hypersensitive response observed following pathogen attack (Koch et al., 2000).
MICROARRAY AND DEVELOPING GENOMIC RESOURCES: A PHYSICAL MAP OF POPLAR?
Populus is currently the best placed of all woody genera with which to exploit the new technologies offered by genomics and proteomics. This is largely the result of a timely programme funded through collaboration between Swedish scientists centred at the Umeå Plant Science Centre, University of Umeå. In the initial phase of this project, approx. 5000 ESTs were developed from a wood cDNA library. This large‐scale production of ESTs from woody angiosperm tissue is unique and utilized an aspen hybrid, P. tremula × P. tremuloides, to generate ESTs for both cambial and developing xylem (Sterky et al., 1998). These findings are summarized in Fig. 3. This large‐scale gene discovery programme sets poplar apart from other trees, and will be extended further in the near future (http://www.pappel.fysbot.umu.se). A new ‘poplar chip’ with 14 000 ESTs from both wood and leaf will offer the opportunity to consider a range of biological questions in trees. This will be complemented with poplar root EST programmes in France and a poplar genomics initiative in Canada. Although genomics is also being developed in Pinus, at the moment the resource offered by Populus is superior. The arabidopsis EST database (http//:www.ncbi.nih.gov) has formed the core of plant genomics in the USA and more recently in the UK (http://www.york.ac.uk/res/garnet/garnet.htm). Widely diverging topics of interest will now be studied in essentially the same way as that outlined by Pesaresi et al. (2001) for photosynthetic genes, where a genomic, microarray approach is coupled to the generation of arabidopsis knock‐out mutant screens and then the identification of proteins through mass spectrometry and bioinformatics. All this should be possible with Populus within the next few years, linked to the discovery of ESTs identified as having a putative role in the trait of interest and where several QTL are already available for that trait in well‐defined mapping progeny. This could provide an efficient route for gene discovery in these difficult plants, as demonstrated recently for rice ESTs.
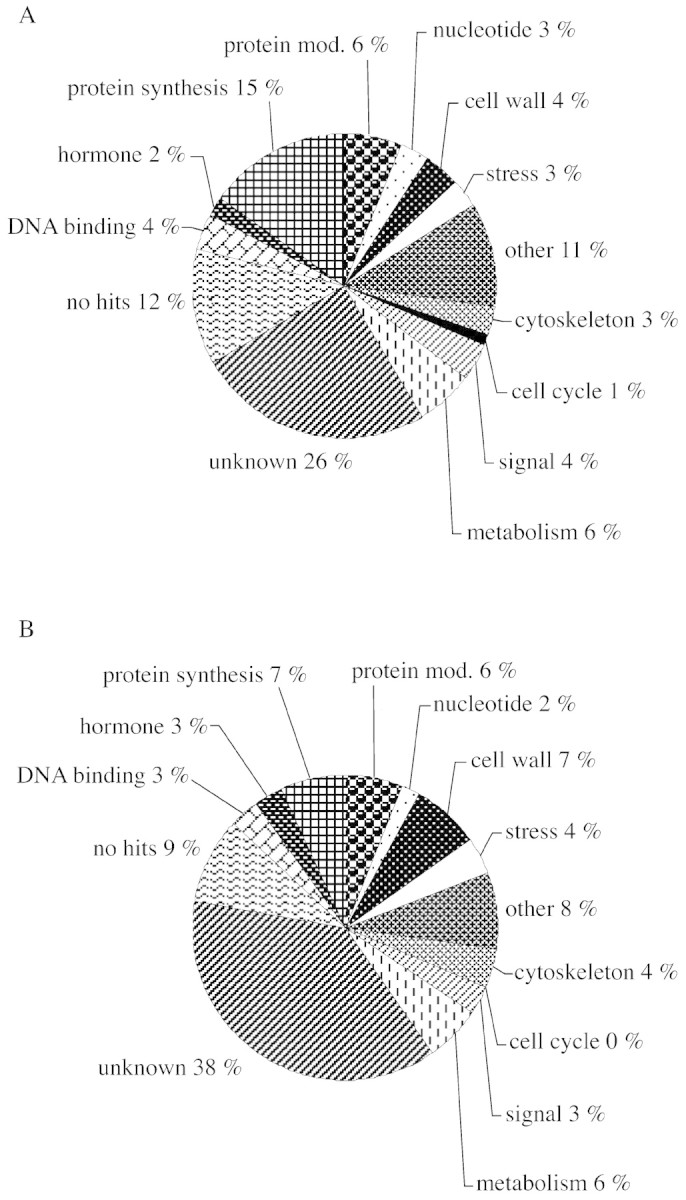
Fig. 3. Classification of ESTs from the cambial region (A) and developing xylem (B) libraries. Modified from the BLASTX scores > 100, modified from Sterky et al. (1998).
Perhaps by far the most significant initiative, though, is that announced in February 2002, in the USA, where a sequencing project has been initiated. The entire 550 Mbp Populus genome is being sequenced by the Joint Genome Institute, meaning that Populus will be the first woody plant whose genome will be known in detail. This sequence information will allow a complete understanding of genetic architecture in a tree relative to other crop plants being sequenced. It should also revolutionize the speed with which genes can be discovered and their role in regulating processes unique to trees, such as wood formation and dormancy, can be elucidated. It is further evidence that Populus has fast become the model tree.
CONCLUSIONS
This brief report has demonstrated that there is a need for a model tree to study the ‘unique’ aspects of tree biology and to elucidate detailed understanding of trees. In this role, the value of Populus as a model is clear: it is fast growing, easy to propagate, has considerable genetic variation, and transformation and tree genomics programmes are up and running. It is likely that rapid progress will be made in the forthcoming years in linking the output from studies of Populus genomics to that of arabidopsis, and in our understanding of some of the difficult developmental and stress responses of trees, in particular, wood formation, signal transduction and the control of dormancy, seasonality, juvenile vs. mature growth and flowering. Populus scores well in a comparison of important features for model plants (Table 3). Populus is unique in that it will not only act as a model for all woody species, but is in itself a forest tree of considerable commercial importance.
Table 3.
Arabidopsis vs. Populus: a summary of major criteria on which to judge a ‘model’ plant system
| Criterion | Arabidopsis | Populus |
| Wide genetic diversity | Yes | Yes |
| Fast growth | Yes | Yes |
| Short life cycle | Yes | No |
| Ease of breeding system | Yes | No |
| Ability to transform | Yes | Yes |
| Molecular genetic maps | Yes | Yes |
| Recombinant inbred lines | Yes | No |
| Genomics | Yes | Yes |
| Immortal mapping populations | No | Yes |
| Sequencing data | Yes | On‐going |
ACKNOWLEDGEMENTS
The author gratefully acknowledges funding for research on Populus in her laboratory, provided by DEFRA (NFO410), NERC, BBSRC and by the EU Fourth (POPFACE, ENV4‐CT97‐0657) and Fifth (POPYOMICS, QLRT‐2001‐00953) frameworks.
Supplementary Material
Received: 11 August 2001; Returned for revision: 24 October 2001; Accepted: 5 September 2002 Published electronically: 24 October 2002
References
- AGI.2000. Arabidopsis thaliana genome. Nature 408: 791–826. [DOI] [PubMed] [Google Scholar]
- AllonaIet al.1998. Analysis of xylem formation in pine by cDNA sequencing. Proceedings of the National Academy of Sciences of the USA 95: 9693–9698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alonso‐BlancoC, Koornneef M.2000. Naturally occurring variation in Arabidopsis: an underexploited resource for plant genetics. Trends in Plant Science 5: 22–29. [DOI] [PubMed] [Google Scholar]
- BaucherMet al. Red xylem and higher lignin extractability by down‐regulating a cinnanyl alcohol dehydrogenase in poplar. Plant Physiology 112: 1479–1490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- BaucherM, Monties B, Van Montagu M, Boerjan W.1998. Biosynthesis and genetic engineering of lignin. Critical Reviews in Plant Sciences 17: 125–197. [Google Scholar]
- BennetzenJL, Kellogg EA, Lee M, Messing J.1998. A plant genome initiative. Plant Cell 10: 488–493. [Google Scholar]
- BirchRG.1997. Plant transformation: problems and strategies for practical application. Annual Review of Plant Physiology and Plant Molecular Biology 48: 297–326. [DOI] [PubMed] [Google Scholar]
- BraatneJH, Hinckley TM, Stettler RF.1992. Influence of soil‐water on the physiological and morphological components of plant water‐balance in Populus‐trichocarpa, Populus‐deltoides and their F1 hybrids. Tree Physiology 11: 325–339. [DOI] [PubMed] [Google Scholar]
- BradshawHD.1996. Molecular genetics of Populus In: Stettler RF, Bradshaw HD, Heilman PE, Hinckley TM, eds. Biology of Populus and its implications for management and conservation. Ottawa: NRC Research Press, 183–199. [Google Scholar]
- BradshawHD, Ceulemans R, Davis J, Stettler R.2000. Emerging model systems in plant biology: poplar (Populus) as a model forest tree. Journal of Plant Growth Regulation 19: 306–313. [Google Scholar]
- BradshawHD, Villar M, Watson BD, Otto KG, Stewart S, Stettler RF.1994. Molecular‐genetics of growth and development in Populus 3. A genetic‐linkage map of a hybrid poplar composed of RFLP, STS, and RAPD markers. Theoretical and Applied Genetics 89: 167–178. [DOI] [PubMed] [Google Scholar]
- BrendleyBW, Pell EJ.1998. Ozone‐induced changes in biosynthesis of Rubisco and associated compensation to stress in foliage of hybrid poplar. Tree Physiology 18: 81–90. [DOI] [PubMed] [Google Scholar]
- CerveraMT, Villar M, Faivre‐Rampant P, Goue M‐C, Van Montagu M, Boerjan W.1997. Applications of molecular marker technologies in Populus breeding. In: Klopfenstein NB, Chun YW, Kim M.‐S, Ahuja MR, Dillon MC, Carman RC, Eskew LG, eds. Micropropagation, genetic engineering, and molecular biology of Populus. Gen. Tech. Rep. RM‐GTR‐297. Fort Collins, CO, USA: US Department of Agriculture, 101–115. [Google Scholar]
- CerveraMT, Storme V, Ivens B, Gusmao J, Liu BH, Hostyn V, Van Slycken J, Van Montagu M, Boerjan W.2001. Dense genetic linkage maps of three Populus species (Populus deltoides, P. nigra and P. trichocarpa) based on AFLP and microsatellite markers. Genetics 158: 787–809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CeulemansR.1990. Genetic variation in functional and structural productivity determinants in poplar. Amsterdam: Amsterdam Thesis Publishers. [Google Scholar]
- CeulemansR, Taylor G, Bosac C, Wilkins D, Besford RB.1997. Photosynthetic acclimation of poplars to elevated CO2 in glasshouse cabinets or in open top chambers depends on duration of exposure. Journal of Experimental Botany 48: 1681–1689. [Google Scholar]
- CeulemansR, Scarascia‐Mugnozza G, Wiard BM, Braatne JH, Hinckley TM, Stettler RF, Isebrands JG, Heilman PE.1992. Production physiology and morphology of Populus species and their hybrids grown under short rotation. 1. Clonal comparisons of 4‐year growth and phenology. Canadian Journal of Forest Research–Revue Canadienne De Recherche Forestiere 22: 1937–1948. [Google Scholar]
- ChenCY, Baucher M, Holst Christensen J, Boerjan W.2001. Biotechnology in trees: Towards improved paper pulping by lignin engineering. Euphytica 118: 185–195. [Google Scholar]
- ChenSG, Ceulemans R, Impens I.1994. A fractal‐based Populus canopy structure model for the calculation of light interception. Forest Ecology and Management 69: 97–110. [Google Scholar]
- ChenSL, Wang SS, Altman A, Huttermann A.1997. Genotypic variation in drought tolerance of poplar in relation to abscisic acid. Tree Physiology 17: 797–803. [DOI] [PubMed] [Google Scholar]
- CloughSJ, Bent AF.1998. Floral dip: a simplified method for Agrobacterium‐mediated transformation of Arabidopsis thaliana Plant Journal 16: 735–743. [DOI] [PubMed] [Google Scholar]
- ColemanMD, Dickson RE, Isebrands JG, Karnosky DF.1995. Carbon allocation and partitioning in aspen clones varying in sensitivity to tropospheric ozone. Tree Physiology 15: 593–604. [DOI] [PubMed] [Google Scholar]
- DickmannDI, Michael DA, Isebrands JG, Westin S.1990. Effects of leaf display on light interception and apparent photosynthesis in 2 contrasting Populus cultivars during their 2nd growing‐season. Tree Physiology 7: 7–20. [DOI] [PubMed] [Google Scholar]
- DuvickDN.2001. Biotechnology in the 1930s: the development of hybrid maize. Nature Reviews Genetics 2: 69–74. [DOI] [PubMed] [Google Scholar]
- EckenwalderJE.1996. Systematics and evolution of Populus In: Stettler RF, Bradshaw HD, Heilman PE, Hinckley TM, eds. Biology of Populus and its implications for management and conservation. Ottawa, Ontario, Canada: NRC Research Press, 7–32. [Google Scholar]
- FeldmannKA, Marks MD.1987. Agrobacterium‐mediated trans formation of germinating seeds of Arabidopsis thaliana – a non‐tissue culture approach. Molecular and General Genetics 208: 1–9. [Google Scholar]
- FerrisR, Sabatti M, Mills RF, Miglietta F, Taylor G.2001. Leaf area is stimulated in Populus by free air CO2 enrichment (POPFACE), through increased cell expansion and production. Plant Cell and Environment 24: 305–315. [Google Scholar]
- FerrisR, Long L, Bunn SM, Bradshaw HD, Rae AM, Taylor G.2002. Leaf stomatal and epidermal cell development: the identification of putative quantitative trait loci in relation to elevated carbon dioxide concentration in poplar. Tree Physiology 22: 633–640. [DOI] [PubMed] [Google Scholar]
- FrewenBE,Chen THH, Howe GT,Davis J, Rohde A, Boerjan W,Bradshaw HD.2000. Quantitative trait loci and candidate gene mapping of bud set and bud flush in Populus Genetics 154: 837–845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- FridmanE, Pleban T, Zamir D.2000. A recombination hotspot delimits a wild‐species quantitative trait locus for tomato sugar content to 484 bp within an invertase gene. Proceedings of the National Academy of Sciences of the USA 97: 4718–4723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- GibsonS, Somerville C.1993. Isolating plant genes. Trends in Biotechnology 11: 306–313. [DOI] [PubMed] [Google Scholar]
- GoodmanHM, Ecker JR, Dean C.1995. The genome of Arabidopsis thaliana Proceedings of the National Academy of Sciences of the USA 92: 10831–10835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- GrattapagliaD, Sederoff R.1994. Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo‐testcross mapping strategy and RAPD markers. Genetics 137: 1121–1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HanKH, Gordon MP, Strauss SH.1996. Cellular and molecular biology of Agrobacterium‐mediated transformation of plants and its application to genetic transformation of Populus In: Stettler RF, Bradshaw HD, Heilman PE, Hinckley TM, eds. Biology of Populus and its implications for management and conservation. Ottawa, Ontario, Canada: NRC Research Press, 201–222. [Google Scholar]
- HaseloffJ, Siemering KR, Prasher DC, Hodge S.1997. Removal of a cryptic intron and subcellular localization of green fluorescent protein are required to mark transgenic Arabidopsis plants brightly. Proceedings of the National Academy of Sciences of the USA 94: 2122–2127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HualaEet al.2001. The Arabidopsis Information Resource (TAIR): a comprehensive database and web‐based information retrieval, analysis, and visualization system for a model plant. Nucleic Acids Research 29: 102–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- IsebrandsJG, Ceulemans R, Wiard B.1988. Genetic variation in photosynthetic traits among Populus clones in relation to yield. Plant Physiology and Biochemistry 26: 427–437. [Google Scholar]
- JeffersonRA, Kavanagh TA, Bevan MW.1987. Gus fusions beta‐glucuronidase as a sensitive and versatile gene fusion marker in higher‐plants. EMBO Journal 6: 3901–3907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- JouaninLet al.2000. Lignification in transgenic poplars with extremely reduced caffeic acid O‐methyltransferase activity. Plant Physiology 123: 1363–1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- KochJR, Creelman RA, Eshita SM, Seskar M, Mullet JE, Davis KR.2000. Ozone sensitivity in hybrid poplar correlates with insensitivity to both salicylic acid and jasmonic acid. The role of programmed cell death in lesion formation. Plant Physiology 123: 487–496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ListerC, Dean C.1993. Recombinant Inbred Lines for Mapping RFLP and phenotypic markers in Arabidopsis thaliana Plant Journal 4: 745–750. [DOI] [PubMed] [Google Scholar]
- MeilanR, Han KH, Ma CP, James RR, Eaton JA, Stanton BJ, Hoien E, Crockett RP, Strauss SH.2000. Development of glyphosate‐tolerant hybrid cottonwoods. Tappi Journal 83: 164–166. [Google Scholar]
- MellerowiczEJ, Baucher M, Sundberg B, Boerjan W.2001. Unravelling cell wall formation in the woody dicot stem. Plant Molecular Biology 47: 239–274. [PubMed] [Google Scholar]
- NadeauJH, Frankel WN.2000. The roads from phenotypic variation to gene discovery: mutagenesis versus QTLs. Nature Genetics 25: 381–384. [DOI] [PubMed] [Google Scholar]
- NewcombeG, Bradshaw HD, Chastagner GA, Stettler RF.1996. A major gene for resistance to Melampsora medusae f. sp. deltoidae in a hybrid poplar pedigree. Phytopathology 86: 87–94. [DOI] [PubMed] [Google Scholar]
- PesaresiP, Varotto C, Richly E, Kurth J, Salamini F, Leister D.2001. Functional genomics of Arabidopsis photosynthesis. Plant Physio logy and Biochemistry 39: 285–294. [Google Scholar]
- RauscherHM, Isebrands JG, Host GE, Dickson RE, Dickmann DI, Crow TR, Michael DA.1990. Ecophys – an ecophysiological growth‐process model for juvenile poplar. Tree Physiology 7: 255–281. [DOI] [PubMed] [Google Scholar]
- RidgeCR, Hinckley TM, Stettler RF, Van Volkenburgh E.1986. Leaf growth characteristics of fast‐growing poplar hybrid, Populus trichocarpa × P. deltoides Tree Physiology 1: 209–216. [DOI] [PubMed] [Google Scholar]
- RottmannWH, Meilan R, Sheppard LA, Brunner AM, Skinner JS, Ma CP, Cheng SP, Jouanin L, Pilate G, Strauss SH.2000. Diverse effects of overexpression of LEAFY and PTLF, a poplar (Populus) homolog of LEAFY/FLORICAULA, in transgenic poplar and Arabidopsis. Plant Journal 22: 235–245. [DOI] [PubMed] [Google Scholar]
- SmithP, Powlson DS, Smith JU, Falloon P, Coleman K.2000. Meeting Europe’s climate change commitments: quantitative estimates of the potential for carbon mitigation by agriculture. Global Change Biology 6: 525–539. [Google Scholar]
- StantonBJ, Villar M.1996. Controlled reproduction of Populus In: Stettler RF, Bradshaw HD, Heilman PE, Hinckley TM, eds. Biology of Populus and its implications for management and conservation. Ottawa, Ontario, Canada: NRC Research Press, 113–138. [Google Scholar]
- SterkyFet al.1998. Gene discovery in the wood‐forming tissues of poplar: analysis of 5692 expressed sequence tags. Proceedings of the National Academy of Sciences of the USA 95: 13330–13335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- StettlerRF, Zsuffa L, Wu R.1996. The role of hybridisation in the genetic manipulation of Populus. In: Stettler FR, Bradshaw HD, Heilman PE, Hinckley TM, eds. Biology of Populus and its implications for management and conservation. Ottawa: NRC Research Press, 87–112. [Google Scholar]
- StrohmM, Eiblmeier M, Langebartels C, Jouanin L, Polle A, Sandermann H, Rennenberg H.1999. Responses of transgenic poplar (Populus tremula × P. alba) overexpressing glutathione synthetase or glutathione reductase to acute ozone stress: visible injury and leaf gas exchange. Journal of Experimental Botany 50: 365–374. [Google Scholar]
- TaylorG, Ceulemans R, Ferris R, Gardner SDL.2001a Increased leaf area expansion of hybrid poplar in elevated CO2 From controlled environment to open‐top chambers and to FACE. Environmental Pollution 115: 463–472. [DOI] [PubMed] [Google Scholar]
- TaylorG, Beckett KP, Robinson KM, Stiles K, Rae AM.2001b Identifying QTL for yield in UK biomass poplar. Aspects of Applied Biology 65: 173–182. [Google Scholar]
- TschaplinskiTJ, Tuskan GA, Gebre GM, Todd DE.1998. Drought resistance of two hybrid Populus clones grown in a large‐scale plantation. Tree Physiology 18: 653–658. [DOI] [PubMed] [Google Scholar]
- TuskanGA, Walsh ME.2001. Short‐rotation woody crop systems, atmospheric carbon dioxide and carbon management: a US case study. Forestry Chronicle 77: 259–264. [Google Scholar]
- VillarM, Lefevre F, Bradshaw HD, Teissier du Cros E.1996. Molecular genetics of rust resistance in poplars (Melampsora larici‐populina Kleb/Populus sp.i) by bulked segregant analysis in a 2 × 2 factorial mating design. Genetics 143: 531–536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- WalbotV.1999. Genes, genomes, genomics. What can plant biologists expect from the 1998 National Science Foundation Plant Genome Research Program? Plant Physiology 119: 1151–1155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- WeigelD, Nilsson O.1995. A developmental switch sufficient for flower initiation in diverse plants. Nature 377: 495–500. [DOI] [PubMed] [Google Scholar]
- WuR, Stettler RF.1996. The genetic resolution of juvenile canopy structure and function in a three‐generation pedigree of Populus Trees—Structure and Function 11: 99–108. [Google Scholar]
- WuRL.1994. Quantitative genetics of yield breeding for Populus short‐rotation culture. 2. Genetic determination and expected selection response of tree geometry. Canadian Journal of Forest Research–Revue Canadienne De Recherche Forestiere 24: 155–165. [Google Scholar]
- WuRL.1998. Genetic mapping of QTLs affecting tree growth and architecture in Populus: implication for ideotype breeding. Theoretical and Applied Genetics 96: 447–457. [DOI] [PubMed] [Google Scholar]
- WuRL, Hu XS, Han YF.2000a Molecular genetics and developmental physiology: Implications for designing better forest crops. Critical Reviews in Plant Sciences 19: 377–393. [Google Scholar]
- WuRL, Han YF, Hu JJ, Fang JJ, Li L, Li ML, Zeng ZB.2000b An integrated genetic map of Populus deltoides based on amplified fragment length polymorphisms. Theoretical and Applied Genetics 100: 1249–1256. [Google Scholar]
- YuJet al.2002. A draft sequence of the rice genome. Science 296: 79–92. [DOI] [PubMed] [Google Scholar]
- ZhangJ, Steenackers M, Storme V, Neyrinck S, Van Montagu M, Gerats T, Boerjan W.2001. Fine mapping and identification of the nucleotide binding site/leucine‐rich repeat sequences at the MER locus in Populus deltoides ‘S9‐2’. Phytopathology 91: 1069–1073. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.