Abstract
Over the last decade, many varied resources have become available for genome studies in rice. These resources include over 4000 DNA markers, several bacterial artificial chromosome (BAC) libraries, P‐1 derived artificial chromosome (PAC) libraries and yeast artificial chromosome (YAC) libraries (genomic DNA clones, filters and end‐sequences), retrotransposon tagged lines, and many chemical and irradiated mutant lines. Based on these, high‐density genetic maps, cereal comparative maps, YAC and BAC physical maps, and quantitative trait loci (QTL) maps have been constructed, and 93 % of the genome has also been sequenced. These data have revealed key features of the genetic and physical structure of the rice genome and of the evolution of cereal chromosomes. This Botanical Briefing examines aspects of how the rice genome is organized structurally, functionally and evolutionarily. Emphasis is placed on the rice centromere, which is composed of long arrays of centromere‐specific repetitive sequences. Differences and similarities amongst various cereal centromeres are detailed. These indicate essential features of centromere function. Another view of various kinds of interactive relationships within and between genomes, which could play crucial roles in genome organization and evolution, is also introduced. Constructed genetic and physical maps indicate duplication of chromosomal segments and spatial association between specific chromosome regions. A genome‐wide survey of interactive genetic loci has identified various reproductive barriers that may drive speciation of the rice genome. The significance of these findings in genome organization and evolution is discussed.
Key words: Oryza sativa, rice, genomics, centromere, genome interaction, chromosome complement, spatial association, repetitive sequence, reproductive barrier
INTRODUCTION
Rice has the simplest of the monocotyledonous genomes analysed to date, which is a considerable advantage when assessing chromosome structure and behaviour. The draft genome sequences for japonica (Goff et al., 2002) and indica (Yu et al., 2002) rice have been published recently. These sequences cover 92–93 % of the genome and predict 40 000–50 000 genes, the largest number in any organism sequenced to date. The International Rice Genome Sequencing Project (http://rgp.dna.affrc.go.jp/cgi‐bin/ statusdb/status.pl) aims to complete full and accurate genome sequencing by the end of 2002. These projects generate the resources needed to assess basic genome structure.
A linkage map, based on a very high density of DNA markers, was constructed by Kurata et al. (1994b), Causse et al. (1994) and Harushima et al. (1998). The markers have since been used as anchor sites for genome sequencing (http://rgp.dna.affrc.go.jp). These markers, together with YAC (yeast artificial chromosome)‐, BAC (bacterial artificial chromosome)‐ and PAC (P‐1 derived artificial chromosome)‐based physical maps have also been utilized for much gene mapping and map‐based gene cloning, including genes dissected by QTL (quantitative trait loci) analysis (Yoshimura et al., 1998; Yano et al., 2000; Takahashi et al., 2001; Wang W et al., 2001). Mapping work with rice and other cereals has revealed highly syntenic genomes amongst cereals (Ahn and Tanksley, 1993; Kurata et al., 1994a; Moore et al., 1995, 1997b). [Synteny is genome similarity between organisms, seen at the level of gene order and sequences.] Large‐scale physical mapping using YAC (Kurata et al., 1997) and BAC clones (Mao et al., 2000; Tao et al., 2001) has also been used to probe several aspects of genome organization including centromere structure (Nonomura and Kurata, 2001) and duplicated regions (Wu et al., 1998). Here, a brief but comprehensive description of the progress on rice genome research is presented, with a focus on the structure of the centromere.
Several studies of genome organization have identified interactions and recognition processes at genetic, physical and functional levels. One such study indicated the possibility of chromosome interactions during mitosis. Other evidence pointed to genetic interactions between and within genomes. A number of reproductive barriers involved in hybrid sterility, hybrid weakness, hybrid breakdown and segregation distortion have also been reported (Nagato and Yoshimura, 1998; and the classified genes listed in ‘Oryzabase’ at http://www.shigen.nig.ac.jp/rice/oryzabase/servlet/rice.oryzabase.ClassListView?key=0). Although neither the causal genes nor the factors responsible for imposing reproductive barriers have been isolated in cereals, these studies present crucial aspects of genome interaction and evolution. In this article, we present an example of such work (Harushima et al., 2001, 2002) and propose mechanisms that may maintain rice genome organization and drive its evolution.
BASIC FEATURES OF THE RICE GENOME
Genome size and relationships between cereal genomes
The rice genome (Oryza sativa; AA genome) is composed of 12 chromosomes (2n = 24) and has a total length of 430 Mb (megabase, a nucleotide length of 1000 000 base pairs) corresponding to about 1500 cM (centiMorgan, a genetic unit of length measured by the crossing‐over frequency in genetic recombinations at meiosis) (Kurata et al., 1994b; Harushima et al., 1998). Thus, in rice, each cM corresponds to a mean of 290 kb and 15 meiotic recombinations. This statistic varies amongst species and with genome size; rice has a relatively small genome (430 Mb) compared with that of other common cereals [maize, 3000 Mb; barley, 3500 Mb, wheat (a hexaploid species with A, B and D genomes), 7000 Mb]. However, in contrast to genome size, the genetic distance of these plants is very similar and lies between 1200 and 1500 cM. Thus, recombination occurs at much the same frequency in meiosis in each cereal, even though the genome sizes differ by more than eight‐fold. One reason for the similar numbers of recombinations could be that they reflect a similar length of gene‐rich regions; the remaining stretches of the genome being gene‐rare regions, each with its unique size and representing heterochromatic blocks suppressed for recombination. Wheat, for example, has been reported to possess large blocks of centromeric/pericentromeric heterochromatin, with gene‐rich regions located on distal ends of chromosomes (Gill et al., 1996). This type of large genome, possessing long heterochromatic regions having a low recombination frequency, is consistent with wheat’s similar genetic distance to that of other cereals with smaller genomes. The rice genome can be expected to possess gene‐rich regions that are similar to those of other cereals but with a smaller proportion of repetitive sequences. Accordingly, a highly conserved order of genes has been identified on chromosomes of rice and other cereal species, i.e. genome synteny is evident among members of the Poaceae (Moore et al., 1995; Devos and Gale, 1997). This synteny was first recognized in rice and maize (Ahn and Tanksley, 1993) and between rice and wheat (Kurata et al., 1994a) in the genetic maps. Later, the evidence was expanded to the level of physical maps (Chen et al., 1997; Foote et al., 1997). These results revealed that even on the physical map, gene order and distances between genes are much the same among the cereal genomes. Rice has become the primary target for cereal genome analysis because of its compact genome and high synteny with other cereal genomes, together with its great economic value.
Genome structure and organization
Almost the whole genome of rice has been sequenced by the Syngenta group (Goff et al., 2002) and the Chinese sequencing group (Yu et al., 2002). Analysis of those draft sequences for the number and functional categories of predicted genes indicates that rice has 40 000–50 000 functional genes with about 15 000 distinct gene families. Surprisingly, the number of genes is larger than that known for any other organisms sequenced to date, and is double the number estimated for Arabidopsis thaliana. Even in A. thaliana, duplication has occurred randomly in most regions of the chromosomes, resulting in the chimeric tetraploid nature of the genome (Vision et al., 2000; Sankoff, 2001). As in arabidopsis, the large number of genes in rice might reflect a polyploid genome. Maize also has a tetraploid structure closely related to the rice genome (Ahn and Tanksley, 1993), suggesting a contribution of polyploidization to the genome evolution of this clade of species.
Analyses of approx. 10 000 expression sequence tags (ESTs) and over 15 000 full‐length cDNAs indicate the presence of many single copy genes and various gene families in rice (Yamamoto and Sasaki, 1997; Kawai et al., 2001; Otomo et al., 2001). Approximately 66 700 unique ESTs have been mapped recently: they cover almost 80 % of the genome but most were mapped to distal parts of chromosomes (Wu et al., 2002). Typical of the redundant sequence families are 18S‐25S and 5S ribosomal RNA genes, which are repeated hundreds of times and clustered in certain regions of particular chromosomes (Shishido et al., 2000). Several gene families, such as ribosomal protein genes (Wu et al., 1995) and histone genes (Sentoku et al., 1999), are scattered over all 12 chromosomes. Numerous protein kinase genes were identified in the whole genome and many disease resistance genes belonging to protein kinase families are clustered on several chromosomes (Wang Z et al., 2001). It has yet to be determined whether there is a mechanism that controls gene order and location on the chromosomes. If so, it would play a role not only in chromosome configuration but also in the functional organization of the genome and might well be involved in genome evolution.
About 50 % of the rice genome, including some parts of open reading frames (ORFs), are made up of repetitive sequences (Kurata et al., 1994b). Precise identification of many kinds of repetitive sequences has been carried out using the full genome draft sequences, revealing that the rice genome is composed of approx. 38 Mb of long and 150 Mb of short repetitive DNA (Goff et al., 2002). This showed more than 48 000 repetitions of di‐, tri‐ and tetra‐nucleotide simple sequence repeats (SSRs). The shortest repetitions are categorized into microsatellites of SSRs (less than 10 bp units), minisatellites (less than 40 bp units) and satellite DNAs (several hundreds bp units). Different sub‐units of various microsatellites have been calculated to repeat 5700–10 000 times in the genome and to be highly scattered (McCouch et al., 1997; Temnykh et al., 2000). One typical microsatellite (CCCTAAA), is a telomere repeat unit, which is tandemly arrayed in blocks ranging in size from several to dozens of kilobases on every chromosome end (Ashikawa et al., 1994; Ohmido et al., 2001). A large proportion of the moderately repeated sequences comprises transposon‐ and other mobile DNA‐related sequences (Mao et al., 2000). DNA transposons of the En/Spm family (Motohashi et al., 1996), several types of retrotransposons of LINE (Noma et al., 1999), SINE (Mochizuki et al., 1992; Bureau et al., 1996), RIRE (Kumekawa et al., 1999b; Kumekawa et al., 2001), TOS (Hirochika, 1997), MITE (Mao et al., 2000), and various other kinds of degenerate sequences including solo‐LTR have been found (see also Goff et al., 2002; Yu et al., 2002). One member of the TOS type of retrotransposon, Tos17, was recently revealed to transpose to various genome positions when cells were subjected to stress conditions, such as cell culture (Hirochika, 1997). Tos17 also proved to be very useful in enabling thousands of insertion mutant lines to be generated (Hirochika, 2001). Several genes isolated from the tagged mutants by PCR screening for pooled DNAs of Tos17 transposed lines have been reported (Sato et al., 1999; Takano et al., 2001). This is a powerful means by which to construct a functional genomic system to search for many genes by both phenotypic screening and DNA sequence analysis. Distribution and transposition of endogenous retrotransposon sequences may contribute to the organization of genome structure and evolution, but this remains purely speculative.
THE CENTROMERE
Centromeres mapped on the chromosomes
Centromeres are highly organized functional components necessary to maintain chromosome integrity during cell division. In rice, the first successful cytological identification of centromeres was by Kurata and Omura (1978) using a novel method of chromosome preparation. However, on the genetic map, the positions of centromeres have not been defined on all 12 rice chromosomes. Attempts have been made to locate centromeres on the genetic map by dosage analysis of RFLP markers using secondary trisomics and telotrisomics (Singh et al., 1996). More precise mapping on the high‐density genetic map could, theoretically, locate up to five centromeres on a single locus or within a 1 cM interval, but it is impossible for seven centromeres to be restricted to less than 1 cM (Harushima et al., 1998). One or two loci that carry multiple DNA markers are located on all candidate centromere regions of each chromosome. These clustered markers could not be segregated from each other in the population of 186 F2 plants used for analysis. The markers are known to be distributed on several YAC clones larger than 1 Mb. The minimum physical lengths at single centromere loci were calculated to be 1390, 2160, 1610 and 1220 kb for chromosomes 1, 7, 9 and 11, respectively, by YAC physical mapping (http://rgp.dna.affrc.go.jp/public data/physicalmap2001/YACall2001.html). These centromere regions should correspond well to the centromeric heterochromatic blocks detected cytologically and are probably suppressed for recombination along a 1 Mb length, as is already known for other organisms (Frary et al., 1996; Copenhaver et al., 1999).
Centromere constitution with repetitive DNAs
In many organisms, such as fruit fly, human, A. thaliana and maize, centromeres have been shown to be composed of long stretches of short tandem repeats and various other kinds of repetitive sequences (Karpen and Allshire, 1997; Ananiev et al., 1998; Tyler‐Smith and Floridia, 2000), as shown in Fig. 1. In rice, centromere‐specific repetitive sequences and several kinds of repeat units have been described. These are shown in Table 1 together with other plant centromeric repetitive sequences. Aragón‐Alcaide et al. (1996) first isolated a repetitive sequence CCS1, commonly located on the centromeres of most cereal plants (e.g. wheat, barley, maize and rice). Dong et al. (1998) identified a BAC clone that possessed seven kinds of repeat elements, all of which were localized on centromeres. Estimations of copy numbers in the genome showed six of them exist in 50–300 copy repeats. The RCS2 sequence of a 168 bp unit is repeated in over 5000 copies in the rice genome. RCS2 was also shown to be arranged as short tandem repeat blocks of various sizes of up to 150 kb in length.
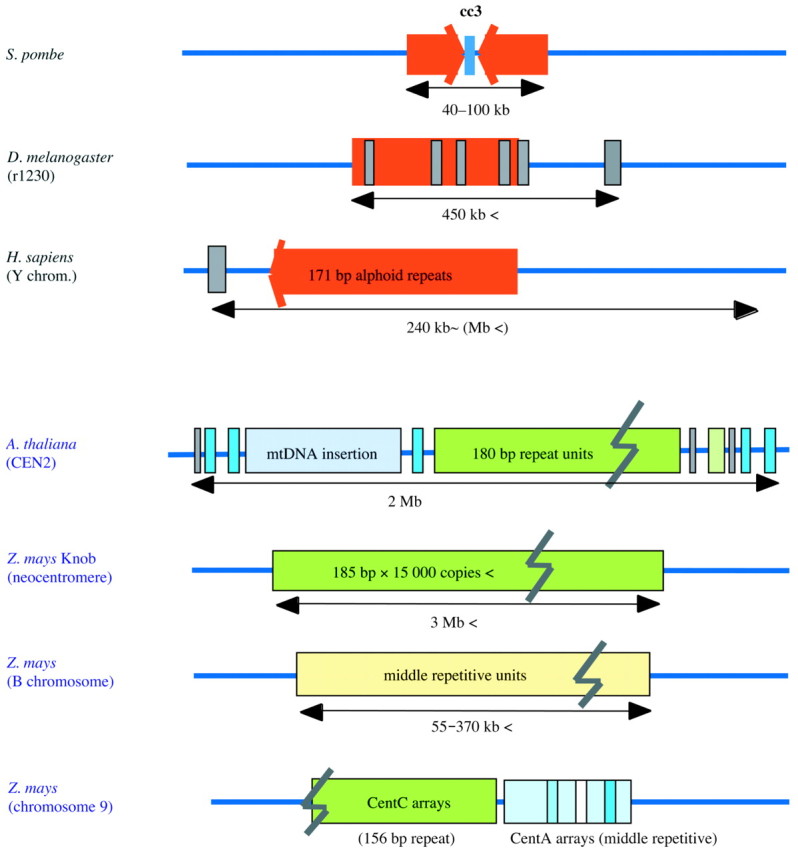
Fig. 1. Schematic illustration of centromere structure for animal and plant species. All centromeres have short tandem repeats (approx. 170 bp unit length) of different, but extensive, length, and transposon‐like moderately repeated sequences surrounding the tandem repeat sequences. Orange and green blocks represent short tandem repeat arrays and other colour blocks are moderately repetitive sequences. Diagrams are for Schizosaccharomyces pombe, Drosophila melanogaster, Homo sapiens, Arabidopsis thaliana and Zea mays.
Table 1.
Centromere repetitive sequences in plants
| Plant | Centromere repeat | Status | Total length or copy number (cp) | Species specificity | Reference |
| Arabidopsis | CEN unique (180 bp) | Short tandem | 400 kb? 1·4 Mb | Arabidopsis specific | Round et al. (1997) |
| Over 10 kinds | Middle dispersed | Common to family | see Copenhaver et al. (1999) | ||
| Cereal | |||||
| Maize | B repeat (?1·4k bp) | Short/middle | 55 kb? Mb/CEN | Maize specific | Alfenito and Birchler (1993) |
| neocent.rep.(185 bp) | Short tandem | ?9 Mb | Maize specific | Birchler (1997) | |
| CentC (156 bp) | Short tandem | 1000 cp</CEN | Maize specific | Ananiev et al. (1998) | |
| CCS1/CentA | Middle dispersed | RT‐LTR‐like | Aragón‐Alcaide et al. (1996) | ||
| False brome and cereals | CCS1 | Middle dispersed | Gramineae(T3/gp) | Aragón‐Alcaide et al. (1996) | |
| Sorghum | pSau3A9 (745 bp) | Middle dispersed | Gramineae(T3/gp) | Jiang et al. (1996) | |
| pSau3A10 (137 bp) | Short tandem | 81 kb </CEN | Sorghum specific | Miller et al. (1998) | |
| Rice | CCS1/RCS1 | Middle dispersed | Gramineae(T3/gp) | Aragón‐Alcaide et al. (1996) | |
| RCS1 (877 bp) | Middle dispersed | 130 cp | Gramineae (RT) | Dong et al. (1998) | |
| RCS2 (168 bp) | Short tandem | 6200 cp (1 Mb) | Rice specific | Dong et al. (1998) | |
| RCH1 (827 bp) | Middle dispersed | 53 cp | Gramineae (RT) | Dong et al. (1998) | |
| RCH2 (1·2 kbp) | Middle dispersed | 99 cp | Gramineae (RT) | Dong et al. (1998) | |
| RCH3 (1·3 kbp) | Middle dispersed | 67 cp | Gramineae (RT) | Dong et al. (1998) | |
| RCE1 701bp) | Middle dispersed | 287 cp | Bambusoideae (T3/gp) | Dong et al. (1998) | |
| RCE2 (2·7 kbp) | Middle dispersed | 305 cp | Gramineae (RT) | Dong et al. (1998) | |
| RCE1 (1·9 kbp) | Middle dispersed | 400–800 cp | Gypsy‐type RT | Nonomura and Kurata (1999) | |
| (Bambusoideae) | Nonomura and Kurata (2001) | ||||
| RIRE7 (7·6 kbp) | Middle dispersed | 200–1700 cp | Gypsy‐type RT | Kumekawa et al. (2001) | |
| RIRE3 (8·0 kbp) | Middle dispersed | Gypsy‐type RT (pl K) | Kumekawa et al. (1999) | ||
| CEN distribution | Nonomura and Kurata (2001) |
Another attempt to isolate centromere repeat sequences was made by targeting the centromere protein‐binding box (CENP‐B). This isolated an RCE1 sequence that was a 1·9 kb unit, tandemly arrayed with intervening sequences (Nonomura and Kurata, 1999). On the other hand, retrotransposon studies in rice presented evidence that most copies of gypsy‐type retrotransposons, such as RIRE3, RIRE7 and RIRE8, are clustered on centromeres though they are truncated and modified (Kumekawa et al., 1999a, 2001; Nonomura and Kurata, 2001). All centromeric repeat units, other than the RCS2 short tandem repeat, were shown to be a part of gypsy‐type retrotransposon sequences of RIRE3, RIRE7 and RIRE8. Using the information shown in Fig. 1 and Table 1, together with sequence and distribution information for many centromere repetitive sequences, it becomes possible to deduce the compositional feature of centromere structure. Accumulated results reveal that the centromere of each organism is composed of a region of genus‐specific highly repetitive short tandem repeats, each about 180 bp, and of long stretches with mixed moderate repetitive transposon‐related sequences common to family members.
Characterization of rice CEN5
A recent study that characterized centromere composition and isolated a large centromere clone made it possible to unravel centromere structure in chromosome 5 (Nonomura and Kurata, 2001). Contig (a contiguous clone stretch constructed by connecting overlapping clones) formation with YAC and BAC clones and analysis of the distribution of major repetitive sequences on the contigs coupled with genetic analysis showed that the centromere of chromosome 5 (CEN5) occupies more than 2 Mb. CEN5 is composed of multiple arrays of various degenerate, moderately repetitive sequences and large blocks of RCS2, short, highly repetitive tandem units at the centre (Fig. 2). The distribution of major moderately repetitive sequences has shown that the copy number of each repetitive member was highest close to RCS2 blocks and decreased towards distal ends of the centromere and away from the central core of the RCS2 blocks. Several functional genes were also scattered between repetitive sequences and were not found to recombine with each other in meiosis when 480 F2 plants were analysed. These genetic and physical characters of CEN5 in rice are similar to those of centromeres of A. thaliana (Copenhaver et al., 1999) and other organisms (Frary et al., 1996; Mahtani and Willard, 1998). The maize centromere was also shown to have a similar constitution, centred on a high copy number block of short tandem repeats being adjacent to moderately repeated transposon‐like sequences (Ananiev et al., 1998). Centromere‐specific short tandem repeat units consisting of 180 bp (approx.) were genus‐ or species‐specific. On the other hand, various retrotransposon‐like sequences located next to the tandem repeat blocks seemed to be common for all species in any one family (Table 1). This common feature might play some functional and evolutionary role in the organization of centromeres.
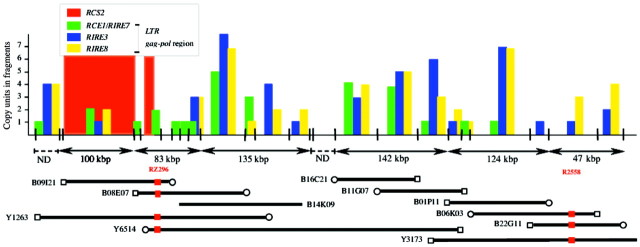
Fig. 2. Distribution of centromere‐specific repetitive sequences of rice on one of the three YAC/BAC contigs of CEN5 that spans more than 630 kb. Numbers of copies were calculated from both strands and expressed in arbitrary units. RCS2 short tandem repeat blocks were observed only on this contig. Other retrotransposon‐related sequences of RIRE3, RIRE8 and RIRE7 included the RCE1 sequence clustered around the RCS2 blocks but were scattered on two other contigs (modified from Nonomura and Kurata, 2001).
Thus, rice CEN5 has been partly analysed, and several YAC clones possessing repeat blocks and multiple copies of retrotransposons in its 380 kb DNA should be promising candidates for examining centromere function. YAC arms with rice telomeres and plant selection markers have been constructed and used for retrofitting (replacement of a YAC vector with other necessary sequences by using homologous recombination in yeast cells) the candidate YAC clone (Adam et al., 1997; Kurata et al., 2001). The retrofitted YAC clone, which possesses both centromere and telomere sequences, should function as an artificial chromosome in rice cells. If the introduced artificial chromosome could be transmitted to the next generation, it would provide a valuable opportunity to study genomic organization and interactions between original chromosomes and artificial chromosomes. These might also provide essential knowledge for generating novel plants by the use of artificial chromosomes.
GENOME INTERACTIONS
Physical interaction in the genome
In addition to sequence‐based characterizations, some findings relate to genome organization and intragenomic interactions. Intensive genetic mapping has detected very few duplicated regions in the rice genome, but occasionally very small regions have shown similarities with each other. However, two segments positioned more than 2·5 Mb from the distal ends of the short arm of chromosomes 11 and 12 show high levels of similarity in their gene order and sequences, suggesting that one segment was generated by duplication of the other. These duplicated segments on chromosomes 11 and 12 may hold promise for detecting structural and functional differences between regions of similarity. Particularly interesting was the finding that the genetic distance differs between these two regions of similar physical length. Only about half the length of the duplicated segments corresponds closely with their genetic length. Of the other half, the distal region of chromosome 11, lying within 300 kb of the tip, corresponded to only 0·3 cM, but to 5·8 cM of genetic distance on chromosome 12. The central region of a duplicated segment of about 1 Mb length also showed this type of difference: in chromosome 11 it extended to 5·1 cM while in chromosome 12 it corresponded to only 0·3 cM (Wu et al., 1998). These observations could reflect a different functional status in the duplicated segments, such as a transcriptionally active or inactive state and/or a conformationally tight or loose state of the chromatin during recombination. If true, structural and functional differences that influence the recombination rate would reveal useful information about genome recognition events.
In addition, there might well be spatial interactions between and within chromosomes in interphase nuclei. Recent evidence suggests that a highly probable close association exists between chromosome segments within the nucleus. For example, chimeric clone formation has often been reported in the course of cloning particularly long segments of genomic DNA into YAC vectors. In rice, we detected that about 40 % of a constructed YAC library comprised chimeric clones (Umehara et al., 1995). The majority of chimeric clones landed on two different and random chromosome positions (Saji et al., 1996; Wang et al., 1996; Koike et al., 1997; Umehara et al., 1997). However, a unique phenomenon was that fragments of the resulting chimeric clones were derived from chromosome 5 at the 100 cM position and the centromere position of chromosome 7. No other such specified association was detected for any other chimeric clones (N. Kurata, unpubl. obs.). Spatial proximity or physical interaction between two genomic regions explains the chimeric clones made up of specific combinations of fragments. If such an association really does occur in the nucleus, further evidence should be forthcoming.
Further evidence of intragenomic chromatin interaction may be sought in events of transposition of DNA elements. DNA‐type transposons, such as the Ac element of maize, can be transposed to closer sites on the same chromosome when they are introduced into A. thaliana (Machida et al., 1997). There is also evidence from rice that a Ds element was transposed to more closely linked positions (Nakagawa et al., 2000). These data suggest that the transposition of DNA elements is apt to occur between chromatin that is in close juxtaposition in the nucleus. If transposition of a DNA‐type transposon from the original integrated site could be detected in its own adjacent position of both the same chromosome and a part of another chromosome, spatial interaction between chromosomes would be demonstrated. Accumulation of such data could be expected to reveal intragenomic interactions and aspects of the spatial and functional organization of the nuclear genome in vivo.
Genetic and functional interaction between genomes
Direct studies to clarify functional and organizational differences between different genomes are difficult because of hybrid barriers. An alternative approach is to detect interference between genomes with the same polyploid species. Closely related chromosome complements with appropriate differences in gene function and in genome organization can provide fertile hybrids that show segregation distortions in their progeny. These offspring permit the identification and analysis of many interactive factors operating between two complementary genomes. In the fruit fly, there is a well‐studied system that prevents transmission of one particular genotype to the next generation in appropriate genetic backgrounds (Temin et al., 1991). One causal factor (the responder) was a repeated sequence (Wu et al., 1988) and sperm with the repeated array failed at the chromatin condensation stage (Tokuyasu et al., 1977). Another ‘reproductive barrier’ on the same chromosome was identified as a truncated Ran‐GAP activator functional in combination with the repeated sequence (Merrill et al., 1999). This is an interesting example of genome interactions at the level of genome organization which strongly influences chromosome function. Research to find reproductive barriers based on genome interactions in rice has recently been initiated in our laboratory.
(1) Reproductive isolation by interaction between homologous chromosome sets.
When crosses are made between different rice strains or species belonging to the same genus, various phenomena can be observed. These include hybrid vigour, hybrid weakness, hybrid incompatibility, hybrid inviability, hybrid sterility and hybrid breakdown. These phenomena are due to complicated interactions between homologous chromosome complements of the same genome. Mapping causal loci and characterization of the genes is the first step to discovering how homologous chromosome complements interact with each other. The viability or fertility of a hybrid is usually controlled by multiple loci. However, the use of QTL analysis is not valid for inviability and sterility because genotypes with serious effects on viability cannot survive, and sterility is determined by the genotypes of both parents and their F2 progeny. However, by using regression analysis, Harushima et al. (2001) succeeded in mapping all the reproductive barriers causing deviations from Mendelian segregation ratios over the whole rice genome. This is the first time all reproductive barriers on a genome have been successfully located on a linkage map. The accurate and intensive mapping of many barriers makes it possible to clone them as genes or DNA elements by means of positional cloning. This would identify the important gene products or DNA structure that interfere in the reproductive processes.
(2) Rapid evolution of reproductive barriers.
Oryza sativa can be classified into two major ecotypes (sometimes called sub‐species, japonica and indica). However, the fertility of japnica–indica hybrids ranges from several per cent to completely fertile. To analyse possible reasons for this variability, all reproductive barriers were mapped and compared between three different japnica–indica crosses (Harushima et al., 2002). More than 30 reproductive barriers were detected, including both gametophytic and zygotic ones in all three crosses. Although genetic polymorphism detected by RFLP analysis was conserved at a 75 % probability from one cross to the others (Antonio et al., 1996), the mapped reproductive barrier loci were largely different among the three crosses. Only 33 % of barriers detected in each cross were conserved in their position and category, even when the maximal possibility was counted among three crosses. This finding suggests that reproductive barriers have evolved more rapidly than other regions of the genome in rice. One of the forces responsible for such rapid evolution of the barriers is probably the domestication of rice.
NEXT STEPS FOR THE STUDY OF GENOME ORGANIZATION
Many features of the structure and function of the rice genome have now been identified. Several tools and experimental systems have also been established that facilitate structural and functional genome analyses; several have been introduced in this Botanical Briefing. The genus Oryza has 22 wild rice species with nine genomes (AA, BB, CC, BBCC, CCDD, EE, FF, GG and HHJJ). This is in addition to the two widely cultivated species Oryza sativa (AA genome) and O. glaberrima (AA genome) (Vaughan, 1994). Following the completion of genome sequencing of O. sativa, genome diversity within the genus will be one of the next targets for genomic analysis. These studies will necessarily include problems of genome organization and comparisons at a chromosome level, sequence level, functional level and evolutionary level. There will be many mechanisms involved in the organization of genome functions and in maintaining complicated programmes of genome organization. Accordingly, studies for resolving them will need novel breakthroughs in molecular biological, cytogenetical, biochemical and genetic methods.
Detailed analyses of interactions between homologous, non‐homologous and/or homoeologous chromosomes in meiosis and comparisons of centromere organization among them can be expected to reveal principles for genome functioning at the chromosome level. Experimental insertion of artificial chromosomes into transgenic plant cells should be a powerful tool by which to clarify essential and supportive elements for centromere function and to identify active factors involved in chromosome organization.
Supplementary Material
Received: 11 September 2001; Returned for revision: 24 October 2001; Accepted: 1 July 2002 Published electronically: 4 September 2002
References
- AdamG, Muller JA, Kindle KL.1997. Retrofitting YACs for direct DNA transfer into plant cells. Plant Journal 11: 1349–1358. [DOI] [PubMed] [Google Scholar]
- AhnS, Tanksley SD.1993. Comparative linkage map of the rice and maize genomes. Proceedings of the National Academy of Sciences of the USA 90: 7980–7984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- AlfenitoMR, Birchler JA.1993. Molecular characterization of a maize B chromosome centric sequence. Genetics 135: 589–597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- AnanievEV, Phillips RL, Rines HW.1998. Chromosome‐specific molecular organization of maize (Zea mays L.) centromeric regions. Proceedings of the National Academy of Sciences of the USA 95: 13073–13078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- AntonioBA, Inoue T, Kajiya H, Nagamura Y, Kurata N, Minobe Y, Yano M, Nakagahra M, Sasaki T.1996. Comparison of genetic distance and order of DNA markers in five populations of rice. Genome 39: 946–956. [DOI] [PubMed] [Google Scholar]
- Aragón‐AlcaideL, Miller T, Schwarzacher T, Reader S, Moore G.1996. A cereal centromeric sequence. Chromosoma 105: 261–268. [DOI] [PubMed] [Google Scholar]
- AshikawaI, Kurata N, Nagamura Y, Minobe Y.1994. Cloning and mapping of telomere‐associated sequences from rice. DNA Research 1: 67–76. [DOI] [PubMed] [Google Scholar]
- BirchlerJA.1997. Do these sequences make CENs yet? Genome Research 7: 1035–1037. [DOI] [PubMed] [Google Scholar]
- BureauTE, Ronald PC, Wessler SR.1996. A computer‐based systematic survey reveals the predominance of small inverted‐repeat elements in wild‐type rice genes. Proceedings of the National Academy of Sciences of the USA 93: 8524–8529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CausseMAet al.1994. Saturated molecular map of the rice genome based on an interspecific backcross population. Genetics 138: 1251–1274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ChenMP, Sanmiguel AC, De Oliveira, Woo S‐S, Zhang H, Wing RA, Bennetzen J.1997. Microcolinearity in sh2‐homologous regions of the maize, rice, and sorghum genomes. Proceedings of the National Academy of Sciences of the USA 94: 3431–3435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CopenhaverGPet al.1999. Genetic definition and sequence analysis of Arabidopsis centromeres. Science 286: 2468–2474. [DOI] [PubMed] [Google Scholar]
- DevosKM, Gale MD.1997. Comparative genetics in the grasses. Plant Molecular Biology 35: 3–15. [PubMed] [Google Scholar]
- DongF, Miller JT, Jackson SA, Wang GL, Ronald PC, Jiang J.1998. Rice (Oryza sativa) centromeric regions consist of complex DNA. Proceedings of the National Academy of Sciences of the USA 95: 8135–8140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- FooteT, Roberts M, Kurata N, Sasaki T, Moore G.1997. Detailed comparative mapping of cereal chromosome regions corresponding to the Ph1 locus in wheat. Genetics 147: 801–807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- FraryA, Presting GG, Tanksley SD.1996. Molecular mapping of the centromeres of tomato chromosomes 7 and 9. Molecular and General Genetics 25: 295–304. [DOI] [PubMed] [Google Scholar]
- GillKS, Gill BS, Endo TR, Taylor T.1996. Identification and high‐density mapping of gene‐rich regions in chromosome group 1 of wheat. Genetics 144: 1883–1891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- GoffSAet al.2002. A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296: 92–100. [DOI] [PubMed] [Google Scholar]
- HarushimaYet al.1998. A high‐density rice genetic linkage map with 2275 markers using a single F2 population. Genetics 148: 479–494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HarushimaY, Nakagahra M, Yano M, Sasaki T, Kurata N.2001. A genome‐wide survey of reproductive barriers in an intraspecific hybrid. Genetics 159: 883–892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HarushimaY, Nakagahra M, Yano M, Sasaki T, Kurata N.2002. Diverse variation of reproductive barriers in three intraspecific rice crosses. Genetics 160: 313–322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HirochikaH.1997. Retrotransposons of rice: their regulation and use for genome analysis. Plant Molecular Biology 35: 231–240. [PubMed] [Google Scholar]
- HirochikaH.2001. Contributin of the Tos17 retrotransposon to rice functional genomics. Current Opinion in Plant Biology 4: 118–122. [DOI] [PubMed] [Google Scholar]
- JiangJ, Nasuda S, Dong F, Scherrer CW, Woo SS, Wing RA, Gill BS, Ward DC.1996. A conserved repetitive DNA element located in the centromere of cereal chromosomes. Proceedings of the National Academy of Sciences of the USA 93: 14210–14213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- KarpenGH, Allshire RC.1997. The case for epigenetic effects on centromere identity and function. Trends in Genetics 13: 489–496. [DOI] [PubMed] [Google Scholar]
- KawaiJet al.2001. Rice full‐length cDNA project: collection and sequencing in RIKEN.The 24th annual meeting of the Japanese Society of Molecular Biology. Proceedings 3P‐009, 608. Osaka: A Secretariat for Japanese Society Meetings. [Google Scholar]
- KoikeK, Yoshino K, Sue N, Umehara Y, Ashikawa I, Kurata N, Sasaki T.1997. Physical mapping of rice chromosomes 4 and 7 using YAC clones. DNA Research 4: 27–33. [DOI] [PubMed] [Google Scholar]
- KumekawaN, Ohtsubo E, Ohtsubo H.1999a Identification and phylogenetic analysis of gypsy‐type retrotransposons in the plant kingdom. Genes and Genetics Systems 74: 299–307. [DOI] [PubMed] [Google Scholar]
- KumekawaN, Ohtsubo H, Horiuchi T, Ohtsubo E.1999b Identification and characterization of novel retrotransposons of the gypsy type in rice. Molecular and General Genetics 260: 593–602. [DOI] [PubMed] [Google Scholar]
- KumekawaN, Ohmido N, Fukui K, Ohtsubo E, Ohtsubo H.2001. A new gypsy type retrotransposon RIRE7: preferential insertion into the tandem repeat sequence TrsD in pericentromeric heterochromatin regions of rice chromosomes. Molecular Genetics and Genomics 265: 480–488. [DOI] [PubMed] [Google Scholar]
- KurataN, Omura T.1978. Karyotype analysis in rice. I. A new method for identifying all chromosome pairs. Japanese Journal of Genetics 53: 251–255. [Google Scholar]
- KurataN, Moore G, Nagamura Y, Foote T, Yano M, Minobe Y, Gale M.1994a Conservation of genome structure between rice and wheat. Bio/technology 12: 276–278. [Google Scholar]
- KurataNet al.1994b A 300 kilobase interval genetic map of rice including 880 expressed sequences. Nature Genetics 8: 365–372. [DOI] [PubMed] [Google Scholar]
- KurataN, Umehara Y, Tanoue H. Sasaki T.1997. Physical mapping of the rice genome with yeast artificial chromosome clones. Special issue: Oryza: from molecular to plant. Plant Molecular Biology 35: 101–113. [PubMed] [Google Scholar]
- KurataN, Suzuki T, Nonomura K‐I.2001. Nuclear and chromosome organization in rice – functional and genomic approaches to find out organizing principles. In: Iida S, ed. Proceedings of the 46th International NIBB conference. Genetics and epigenetics – the first 100 years. Okazaki: NIBB, 23–24. [Google Scholar]
- McCouchSR, Chen X, Panaud O, Temnykh S, Xu Y, Cho YG, Huang N, Ishii T, Blair M.1997. Microsatellite marker development, mapping and applications in rice genetics and breeding. Plant Molecular Biology 35: 89–99. [PubMed] [Google Scholar]
- MachidaC, Onouchi H, Koizumi J, Hamada S, Semiarti E, Torikai S, Machida Y.1997. Characterization of the transposition pattern of the Ac element in Arabidopsis thaliana using endonuclease I‐SceI. Proceedings of the National Academy of Sciences of the USA 94: 8675–8680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MahtaniMM, Willard HF.1998. Physical and genetic mapping of the human X chromosome centromere repression of recombination. Genome Research 8: 100–110. [DOI] [PubMed] [Google Scholar]
- MaoLet al.2000. Rice transposable elements: a survey of 73 000 sequence‐tagged‐connectors. Genome Research 10: 982–990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MerrillC, Bayraktaroglu L, Kusano A, Ganetzky B.1999. Truncated RanGAP encoded by the segregation distorter locus of Drosophila Science 283: 1742–1745. [DOI] [PubMed] [Google Scholar]
- MillerJT, Jackson SA, Nasuda S, Gill BS, Wing RA, Dong F, Ward DC, Jiang J.1998. Cloning and characterization of a centromere‐specific repetitive DNA element from Sorghum bicolor Theoretical and Applied Genetics 96: 832–839. [Google Scholar]
- MochizukiK, Umeda M, Ohtsubo H, Ohtsubo E.1992. Characterization of a plant SINE, p‐SINE1, in rice genomes. Japanese Journal of Genetics (Genes and Genetic Systems) 67: 155–166. [DOI] [PubMed] [Google Scholar]
- MooreG, Foote T, Helentjaris T, Devos K, Kurata N, Gale M.1995. Was there a single ancestral cereal chromosome? Trends in Genetics 11: 81–82. [DOI] [PubMed] [Google Scholar]
- MooreG, Roberts M, Alcaide L, Foote T.1997a Centromere sites and cereal genome evolution. Chromosoma 105: 321–323. [DOI] [PubMed] [Google Scholar]
- MooreG, Aragon‐Alcaide L, Roberts M, Reader S, Miller T, Foote T.1997b Are rice chromosomes components of a holocentric chromosome ancestor? Plant Molecular Biology 35: 17–23. [PubMed] [Google Scholar]
- MotohashiR, Ohtsubo E, Ohtsubo H.1996. Identification of Tnr3, a suppressor‐mutator/enhancer‐like transposable element from rice. Molecular and General Genetics 250: 148–152. [DOI] [PubMed] [Google Scholar]
- NagatoY, Yoshimura A.1998. Report of the committee on gene symbolization, nomenclature and linkage groups. Rice Genetics Newsletter 15: 13–74. [Google Scholar]
- NakagawaY, Machida C, Machida Y, Toriyama K.2000. Frequency and pattern of transposition of the maize transposable element Ds in transgenic rice plants. Plant Cell Physiology 41: 733–742. [DOI] [PubMed] [Google Scholar]
- NomaK, Ohtsubo E, Ohtsubo H.1999. Non‐LTR retrotransposons (LINEs) as ubiquitous components of plant genomes. Molecular and General Genetics 261: 71–79. [DOI] [PubMed] [Google Scholar]
- NonomuraK‐I, Kurata N.1999. Organization of the 1·9‐kb repeat unit RCE1 in the centromeric region of rice chromosomes. Molecular and General Genetics 261: 1–10. [DOI] [PubMed] [Google Scholar]
- NonomuraK‐I, Kurata N.2001. The centromere composition of multiple repetitive sequences on rice chromosome 5. Chromosoma 110: 284–291. [DOI] [PubMed] [Google Scholar]
- OhmidoN, Kijima K, Ashikawa I, de Jong JH, Fukui K.2001. Visualization of the terminal structure of rice chromosomes 6 and 12 with multicolor FISH to chromosomes and extended DNA fibers. Plant Molecular Biology 47: 413–421. [DOI] [PubMed] [Google Scholar]
- OtomoY, Xie Q, Narikawa T, Kusunegi T, Miura J, Sugano S, Kikuchi S, Higo K, Murakami K, Matsubara K.2001. Large‐scale analysis of a rice full length cDNA. The 24th annual meeting of the Japanese Society of Molecular Biology. Proceedings 3p‐011, 608. Osaka: A Secretariat for Japanese Society Meetings. [Google Scholar]
- SajiS, Umehara Y, Kurata N, Ashikawa I, Sasaki T.1996. Construction of YAC contigs rice chromosome 5. DNA Research 3: 297–302. [DOI] [PubMed] [Google Scholar]
- SankoffD.2001. Gene and genome duplication. Current Opinion in Genetics and Development 11: 681–684. [DOI] [PubMed] [Google Scholar]
- SatoY,Sentoku N, Miura Y, Hirochika H, Kitano H, Matsuoka M.1999. Loss‐of‐function mutations in the rice homeobox gene OSH15 affect the architecture of internodes resulting in dwarf plants. EMBO Journal 18: 992–1002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SentokuN, Sato Y, Kurata N, Ito Y, Kitano H, Matsuoka M.1999. Regional expression of the rice KN1‐type homeobox gene family during embryo, shoot and flower development. Plant Cell 11: 1651–1664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ShishidoR, Sano Y, Fukui K.2000. Ribosomal DNAs: an exception to the conservation of gene order in rice genomes. Molecular and General Genetics 263: 586–591. [DOI] [PubMed] [Google Scholar]
- SinghK, Ishii T, Parco A, Huang N, Brar NH, Khush GS.1996. Centromere mapping and orientation of the molecular linkage map of rice (Oryza sativa L.). Proceedings of the National Academy of Sciences of the USA 93: 6163–6168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- TakahashiY, Shomura A, Sasaki T, Yano M.2001. Hd6, a rice quantitative trait locus involved in photoperiod sensitivity, encodes the a subunit of protein kinase CK2. Proceedings of the National Academy of Sciences of the USA 98: 7922–7927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- TakanoM, Kanegae H, Shinomura T, Miyao A, Hirochika H, Furuya M.2001. Isolation and characterization of rice phytochome A mutants. Plant Cell 13: 521–534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- TaoQ, Chang Y‐L, Wang J, Chen H, Islam‐Faridi MN, Scheuring C, Wang B, Stelly DM, Zhang H‐B.2001. Bacterial artificial chromosome‐based physical map of the rice genome constructed by restriction fingerprint analysis. Genetics 158: 1711–1724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- TeminRG, Ganetzky G, Powers PA, Lyttle TW, Pimpinelli S, Dimitri P, Wu C‐I, Hiraizumi Y.1991. Segregation distortion in Drosophila melanogaster: genetic and molecular analyses. American Naturalist 137: 287–331. [Google Scholar]
- TemnykhS, Park WD, Ayres N, Cartinhour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR.2000. Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theoretical and Applied Genetics 100: 697–712. [Google Scholar]
- TokuyasuKT, Peacock WJ, Hardy RW.1977. Dynamics of spermiogenesis in Drosophila melanogaster VII. Effects of segregation distorter (SD) chromosome. Journal of Ultrastructure Research 58: 96–107. [DOI] [PubMed] [Google Scholar]
- Tyler‐SmithC, Floridia G.2000. Many paths to the top of the mountain: diverse evolutionary solutions to centromere structure. Cell 102: 5–8. [DOI] [PubMed] [Google Scholar]
- UmeharaY, Miyazaki A, Tanoue H, Yasukouchi Y, Nagamura Y, Saji S, Fujimura T, Kurata N, Minobe Y.1995. Construction and characterization of rice YAC libraries for physical mapping. Molecular Breeding 1: 79–89. [Google Scholar]
- UmeharaY, Kurata N, Ashikawa I. Sasaki T.1997. Yeast artificial chromosome clones of rice chromosome 2 ordered using DNA markers. DNA Research 4: 127–131. [DOI] [PubMed] [Google Scholar]
- VaughanDA.1994. The wild relatives of rice. Manila: International Rice Research Institute Press. [Google Scholar]
- VisionTJ, Brown DG, Tanksley SD.2000. The origins of genomic duplications in Arabidopsis. Science 290: 2114–2117. [DOI] [PubMed] [Google Scholar]
- WangW, Zhai W, Luo M, Jiang G, Chen X, Li X, Wing RA, Zhu L.2001. Chromosome landing at the bacterial blight resistance gene Xa4 locus using a deep coverage rice BAC library. Molecular Genetics and Genomics 265: 118–125. [DOI] [PubMed] [Google Scholar]
- WangZ, Taramino G, Yang D, Liu G, Tingey SV, Miao GH, Wang GL.2001. Rice ESTs with disease‐resistance gene‐ or defense‐response gene‐like sequences mapped to regions containing major resistance genes or QTLs. Molecular Genetics and Genomics 265: 302–310. [DOI] [PubMed] [Google Scholar]
- WangZX, Idonuma A, Umehara Y, Van Houten W, Ashikawa I, Minobe Y, Kurata N, Sasaki T.1996. Physical mapping of rice chromosome 1 with yeast artificial chromosomes (YACs). DNA Research 3: 291–296. [DOI] [PubMed] [Google Scholar]
- WuC‐I, Lyttle TW, Wu M‐L, Lin G‐F.1988. Association between a satellite DNA sequence and the responder of segregation distorter in D. melanogaster Cell 54: 179–189. [DOI] [PubMed] [Google Scholar]
- WuJ, Matsui E, Yamamoto K, Nagamura Y, Kurata N, Sasaki T, Minobe Y.1995. Genomic organization of 57 ribosomal protein genes in rice (Oryza sativa L.) through RFLP mapping. Genome 38: 1189–1200 . [DOI] [PubMed] [Google Scholar]
- WuJ, Kurata N, Tanoue H, Shimokawa T, Umehara Y, Yano M. Sasaki T.1998. Physical mapping of duplicated genomic regions of two chromosome ends in rice. Genetics 150: 1595–1603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- WuJet al.2002. A comprehensive rice transcript map containing 6 591 EST sites. Plant Cell 14: 525–535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- YamamotoK, Sasaki T.1997. Large scale EST sequencing in rice. Special issue: Oryza: from molecular to plant. Plant Molecular Biology 35: 135–144. [PubMed] [Google Scholar]
- YanoM et al.2000. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 12: 2473–2484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- YoshimuraS, Yamanouchi U, Katayose Y, Wang Z‐X, Toki S, Kono K, Kurata N, Yano M, Iwata N, Sasaki T.1998. Expression of Xa‐1, a novel bacterial blight resistance gene in rice, is induced by bacterial inoculation. Proceedings of the National Academy of Sciences of the USA 95: 1663–1668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- YuJet al.2002. A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296: 79–92. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


