Abstract
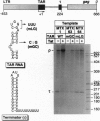
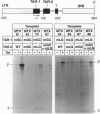
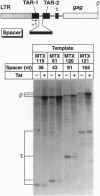
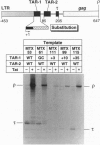
The human immunodeficiency virus type 1 (HIV-1) trans-activator protein, Tat, specifically stimulates transcription from the viral long terminal repeat. Tat binds to an RNA stem-loop structure encoded by the trans-activation response region (TAR). To test whether TAR is functional when displaced downstream of the start of transcription, we assayed a series of templates carrying duplicated TAR elements in cell-free transcription systems. When the normally positioned TAR element (TAR-1) is inactivated by mutations in either the Tat binding site or the apical loop sequence, which acts as the binding site for a cellular factor, transactivation can be rescued by a wild-type TAR element placed downstream (TAR-2). The TAR-2 element is functional even when placed > 200 nt downstream of TAR-1. TAR complementation experiments have also shown that a functional TAR element requires both an intact Tat binding site and an intact apical loop sequence. For example, if TAR-1 carries a mutation in the loop element it cannot be rescued by a TAR-2 element carrying a mutation in the Tat binding site. Substitution mutations in TAR-1 show that the 5' half of TAR also encodes an essential DNA element which is required for efficient transcription initiation. These results strongly suggest that Tat and cellular cofactors which bind TAR RNA associate with the transcription complex during its transit through TAR.
Full text
PDF
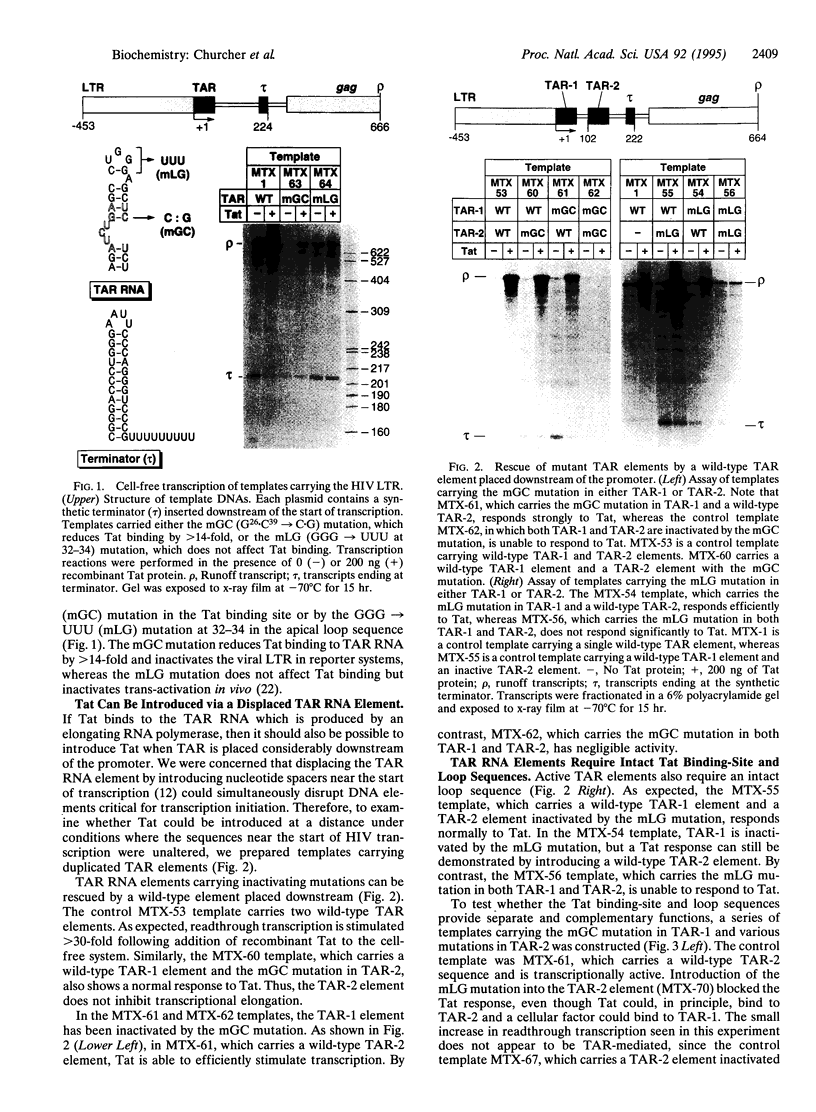
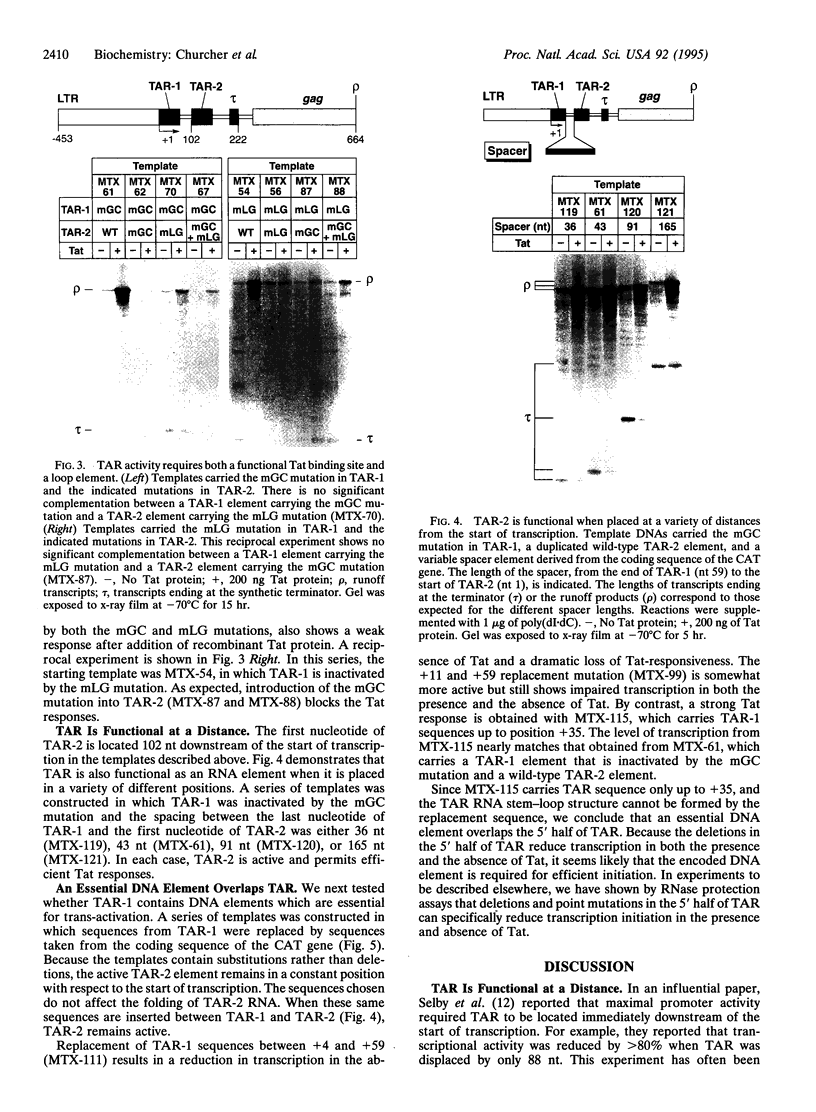


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bengal E., Aloni Y. Transcriptional elongation by purified RNA polymerase II is blocked at the trans-activation-responsive region of human immunodeficiency virus type 1 in vitro. J Virol. 1991 Sep;65(9):4910–4918. doi: 10.1128/jvi.65.9.4910-4918.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berkhout B., Gatignol A., Rabson A. B., Jeang K. T. TAR-independent activation of the HIV-1 LTR: evidence that tat requires specific regions of the promoter. Cell. 1990 Aug 24;62(4):757–767. doi: 10.1016/0092-8674(90)90120-4. [DOI] [PubMed] [Google Scholar]
- Berkhout B., Gatignol A., Silver J., Jeang K. T. Efficient trans-activation by the HIV-2 Tat protein requires a duplicated TAR RNA structure. Nucleic Acids Res. 1990 Apr 11;18(7):1839–1846. doi: 10.1093/nar/18.7.1839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berkhout B., Jeang K. T. Detailed mutational analysis of TAR RNA: critical spacing between the bulge and loop recognition domains. Nucleic Acids Res. 1991 Nov 25;19(22):6169–6176. doi: 10.1093/nar/19.22.6169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berkhout B., Silverman R. H., Jeang K. T. Tat trans-activates the human immunodeficiency virus through a nascent RNA target. Cell. 1989 Oct 20;59(2):273–282. doi: 10.1016/0092-8674(89)90289-4. [DOI] [PubMed] [Google Scholar]
- Churcher M. J., Lamont C., Hamy F., Dingwall C., Green S. M., Lowe A. D., Butler J. G., Gait M. J., Karn J. High affinity binding of TAR RNA by the human immunodeficiency virus type-1 tat protein requires base-pairs in the RNA stem and amino acid residues flanking the basic region. J Mol Biol. 1993 Mar 5;230(1):90–110. doi: 10.1006/jmbi.1993.1128. [DOI] [PubMed] [Google Scholar]
- Cullen B. R. Does HIV-1 Tat induce a change in viral initiation rights? Cell. 1993 May 7;73(3):417–420. doi: 10.1016/0092-8674(93)90126-b. [DOI] [PubMed] [Google Scholar]
- Dingwall C., Ernberg I., Gait M. J., Green S. M., Heaphy S., Karn J., Lowe A. D., Singh M., Skinner M. A. HIV-1 tat protein stimulates transcription by binding to a U-rich bulge in the stem of the TAR RNA structure. EMBO J. 1990 Dec;9(12):4145–4153. doi: 10.1002/j.1460-2075.1990.tb07637.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingwall C., Ernberg I., Gait M. J., Green S. M., Heaphy S., Karn J., Lowe A. D., Singh M., Skinner M. A., Valerio R. Human immunodeficiency virus 1 tat protein binds trans-activation-responsive region (TAR) RNA in vitro. Proc Natl Acad Sci U S A. 1989 Sep;86(18):6925–6929. doi: 10.1073/pnas.86.18.6925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg M. B., Baltimore D., Frankel A. D. The role of Tat in the human immunodeficiency virus life cycle indicates a primary effect on transcriptional elongation. Proc Natl Acad Sci U S A. 1991 May 1;88(9):4045–4049. doi: 10.1073/pnas.88.9.4045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng S., Holland E. C. HIV-1 tat trans-activation requires the loop sequence within tar. Nature. 1988 Jul 14;334(6178):165–167. doi: 10.1038/334165a0. [DOI] [PubMed] [Google Scholar]
- Fenrick R., Malim M. H., Hauber J., Le S. Y., Maizel J., Cullen B. R. Functional analysis of the Tat trans activator of human immunodeficiency virus type 2. J Virol. 1989 Dec;63(12):5006–5012. doi: 10.1128/jvi.63.12.5006-5012.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gait M. J., Karn J. RNA recognition by the human immunodeficiency virus Tat and Rev proteins. Trends Biochem Sci. 1993 Jul;18(7):255–259. doi: 10.1016/0968-0004(93)90176-n. [DOI] [PubMed] [Google Scholar]
- Garcia J. A., Wu F. K., Mitsuyasu R., Gaynor R. B. Interactions of cellular proteins involved in the transcriptional regulation of the human immunodeficiency virus. EMBO J. 1987 Dec 1;6(12):3761–3770. doi: 10.1002/j.1460-2075.1987.tb02711.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graeble M. A., Churcher M. J., Lowe A. D., Gait M. J., Karn J. Human immunodeficiency virus type 1 transactivator protein, tat, stimulates transcriptional read-through of distal terminator sequences in vitro. Proc Natl Acad Sci U S A. 1993 Jul 1;90(13):6184–6188. doi: 10.1073/pnas.90.13.6184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones K. A., Luciw P. A., Duchange N. Structural arrangements of transcription control domains within the 5'-untranslated leader regions of the HIV-1 and HIV-2 promoters. Genes Dev. 1988 Sep;2(9):1101–1114. doi: 10.1101/gad.2.9.1101. [DOI] [PubMed] [Google Scholar]
- Kamine J., Subramanian T., Chinnadurai G. Sp1-dependent activation of a synthetic promoter by human immunodeficiency virus type 1 Tat protein. Proc Natl Acad Sci U S A. 1991 Oct 1;88(19):8510–8514. doi: 10.1073/pnas.88.19.8510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kao S. Y., Calman A. F., Luciw P. A., Peterlin B. M. Anti-termination of transcription within the long terminal repeat of HIV-1 by tat gene product. Nature. 1987 Dec 3;330(6147):489–493. doi: 10.1038/330489a0. [DOI] [PubMed] [Google Scholar]
- Karn J., Graeble M. A. New insights into the mechanism of HIV-1 trans-activation. Trends Genet. 1992 Nov;8(11):365–368. doi: 10.1016/0168-9525(92)90284-b. [DOI] [PubMed] [Google Scholar]
- Kato H., Sumimoto H., Pognonec P., Chen C. H., Rosen C. A., Roeder R. G. HIV-1 Tat acts as a processivity factor in vitro in conjunction with cellular elongation factors. Genes Dev. 1992 Apr;6(4):655–666. doi: 10.1101/gad.6.4.655. [DOI] [PubMed] [Google Scholar]
- Kessler M., Mathews M. B. Premature termination and processing of human immunodeficiency virus type 1-promoted transcripts. J Virol. 1992 Jul;66(7):4488–4496. doi: 10.1128/jvi.66.7.4488-4496.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laspia M. F., Rice A. P., Mathews M. B. HIV-1 Tat protein increases transcriptional initiation and stabilizes elongation. Cell. 1989 Oct 20;59(2):283–292. doi: 10.1016/0092-8674(89)90290-0. [DOI] [PubMed] [Google Scholar]
- Laspia M. F., Rice A. P., Mathews M. B. Synergy between HIV-1 Tat and adenovirus E1A is principally due to stabilization of transcriptional elongation. Genes Dev. 1990 Dec;4(12B):2397–2408. doi: 10.1101/gad.4.12b.2397. [DOI] [PubMed] [Google Scholar]
- Laspia M. F., Wendel P., Mathews M. B. HIV-1 Tat overcomes inefficient transcriptional elongation in vitro. J Mol Biol. 1993 Aug 5;232(3):732–746. doi: 10.1006/jmbi.1993.1427. [DOI] [PubMed] [Google Scholar]
- Marciniak R. A., Calnan B. J., Frankel A. D., Sharp P. A. HIV-1 Tat protein trans-activates transcription in vitro. Cell. 1990 Nov 16;63(4):791–802. doi: 10.1016/0092-8674(90)90145-5. [DOI] [PubMed] [Google Scholar]
- Marciniak R. A., Sharp P. A. HIV-1 Tat protein promotes formation of more-processive elongation complexes. EMBO J. 1991 Dec;10(13):4189–4196. doi: 10.1002/j.1460-2075.1991.tb04997.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muesing M. A., Smith D. H., Capon D. J. Regulation of mRNA accumulation by a human immunodeficiency virus trans-activator protein. Cell. 1987 Feb 27;48(4):691–701. doi: 10.1016/0092-8674(87)90247-9. [DOI] [PubMed] [Google Scholar]
- Ratnasabapathy R., Sheldon M., Johal L., Hernandez N. The HIV-1 long terminal repeat contains an unusual element that induces the synthesis of short RNAs from various mRNA and snRNA promoters. Genes Dev. 1990 Dec;4(12A):2061–2074. doi: 10.1101/gad.4.12a.2061. [DOI] [PubMed] [Google Scholar]
- Selby M. J., Bain E. S., Luciw P. A., Peterlin B. M. Structure, sequence, and position of the stem-loop in tar determine transcriptional elongation by tat through the HIV-1 long terminal repeat. Genes Dev. 1989 Apr;3(4):547–558. doi: 10.1101/gad.3.4.547. [DOI] [PubMed] [Google Scholar]
- Sheldon M., Ratnasabapathy R., Hernandez N. Characterization of the inducer of short transcripts, a human immunodeficiency virus type 1 transcriptional element that activates the synthesis of short RNAs. Mol Cell Biol. 1993 Feb;13(2):1251–1263. doi: 10.1128/mcb.13.2.1251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheline C. T., Milocco L. H., Jones K. A. Two distinct nuclear transcription factors recognize loop and bulge residues of the HIV-1 TAR RNA hairpin. Genes Dev. 1991 Dec;5(12B):2508–2520. doi: 10.1101/gad.5.12b.2508. [DOI] [PubMed] [Google Scholar]
- Sodroski J., Rosen C., Wong-Staal F., Salahuddin S. Z., Popovic M., Arya S., Gallo R. C., Haseltine W. A. Trans-acting transcriptional regulation of human T-cell leukemia virus type III long terminal repeat. Science. 1985 Jan 11;227(4683):171–173. doi: 10.1126/science.2981427. [DOI] [PubMed] [Google Scholar]
- Southgate C. D., Green M. R. The HIV-1 Tat protein activates transcription from an upstream DNA-binding site: implications for Tat function. Genes Dev. 1991 Dec;5(12B):2496–2507. doi: 10.1101/gad.5.12b.2496. [DOI] [PubMed] [Google Scholar]
- Toohey M. G., Jones K. A. In vitro formation of short RNA polymerase II transcripts that terminate within the HIV-1 and HIV-2 promoter-proximal downstream regions. Genes Dev. 1989 Mar;3(3):265–282. doi: 10.1101/gad.3.3.265. [DOI] [PubMed] [Google Scholar]
- Wu F., Garcia J., Sigman D., Gaynor R. tat regulates binding of the human immunodeficiency virus trans-activating region RNA loop-binding protein TRP-185. Genes Dev. 1991 Nov;5(11):2128–2140. doi: 10.1101/gad.5.11.2128. [DOI] [PubMed] [Google Scholar]
- Zenzie-Gregory B., Sheridan P., Jones K. A., Smale S. T. HIV-1 core promoter lacks a simple initiator element but contains a bipartite activator at the transcription start site. J Biol Chem. 1993 Jul 25;268(21):15823–15832. [PubMed] [Google Scholar]