Abstract
Myeloid-derived suppressor cells (MDSC's), a heterogeneous population of cells capable of suppressing anti-tumor T cell function in the tumor microenvironment, represent an imposing obstacle in the development of cancer immunotherapeutics. Thus, identifying elements essential to the development and perpetuation of these cells will undoubtedly improve our ability to circumvent their suppressive impact. HDAC11 has emerged as a key regulator of IL-10 gene expression in myeloid cells, suggesting that this may represent an important targetable axis through which to dampen MDSC formation. Using a murine transgenic reporter model system where eGFP expression is controlled by the HDAC11 promoter (Tg-HDAC11-eGFP), we provide evidence that HDAC11 appears to function as a negative regulator of MDSC expansion/function in vivo. MDSCs isolated from EL4 tumor-bearing Tg-HDAC11-eGFP display high expression of eGFP, indicative of HDAC11 transcriptional activation at steady state. In striking contrast, immature myeloid cells in tumor-bearing mice display a diminished eGFP expression, implying that the transition of IMC to MDSC's require a decrease in the expression of HDAC11, where we postulate that it acts as a gate-keeper of myeloid differentiation. Indeed, tumor-bearing HDAC11-knockout mice (HDAC11-KO) demonstrate a more suppressive MDSC population as compared to wild-type (WT) tumor-bearing control. Notably, the HDAC11-KO tumor-bearing mice exhibit enhanced tumor growth kinetics when compare to the WT control mice. Thus, through a better understanding of this previously unknown role of HDAC11 in MDSC expansion and function, rational development of targeted epigenetic modifiers may allow us to thwart a powerful barrier to efficacious immunotherapies.
Keywords: HDAC11, MDSCs, immuno-suppression, myelopoiesis
1. Introduction
In treatment of cancer, successful immunotherapy hinges on the effective function of antigen-presenting cells (APCs) and T cells. In fact, the concept of immunity is based on the capacity of T cells to mount an effective immune response against malignant cells and harmful pathogens. One of the major hurdles in cancer immunotherapy is the failure of T cells to attain an effective response to malignant cells. Such problems arise when T cells become unresponsive to tumor specific antigens due to physiological changes in the tumor microenvironment(Wells 2003). In the past several years, mounting evidence has demonstrated that negative regulation of the host immune response is due to two groups of cells: 1-regulatory T cells (Tregs)(Wang 2006) and 2- myeloid-derived suppressor cells (MDSCs)(Youn and Gabrilovich 2010). MDSCs are a distinct population of cells with the ability to suppress various T cell functions. They are a heterogeneous population of cells generally composed of precursors to dendritic cells, granulocytes, macrophages, as well as myeloid cells at various stages of differentiation(Serafini, Borrello et al. 2006). In cancer patients, MDSCs are defined as cells that express the common myeloid marker CD33 but lack expression of mature myeloid and lymphoid cells(Almand, Clark et al. 2001). In mice, these cells are recognized by co-expression of CD11b and GR-1 and have been more precisely identified by their immature myeloid origin, and most significantly their strong suppressive ability in various facets of immune response, most importantly in T-cell activation, proliferation and cytokine production(Marigo, Dolcetti et al. 2008, Youn and Gabrilovich 2010). In recent years, these cells have been further sub-categorized into two subsequent subsets based on their expression of two molecules Ly-6C and Ly-6G(Hestdal, Ruscetti et al. 1991, Youn, Nagaraj et al. 2008). CD11b+ Ly-6G− Ly-6Chigh cells have been identified to have monocytic-like morphology and are subsequently termed monocytic-MDSCs (M-MDSCs) and CD11b+ Ly-6G+ Ly-6Clow cells have been identified to have granulocytic-like morphology and are termed granulocytic MDSCs (G-MDSCs)(Condamine and Gabrilovich 2011). MDSCs have a very fast proliferative capacity and rapidly accumulate in lymphoid organs of mice with infectious diseases(Gomez-Garcia, Lopez-Marin et al. 2005), inflammation(Ezernitchi, Vaknin et al. 2006), sepsis(Delano, Scumpia et al. 2007), and more importantly in mice bearing tumors(Sawanobori, Ueha et al. 2008). Since the identification of this suppressive subset, numerous studies have convincingly demonstrated possible molecules such as arginase, nitric oxide (NO), and reactive oxygen species (ROS) as major culprits responsible for the immunosuppressive ability of these cells(Bronte and Zanovello 2005, Rodriguez and Ochoa 2008, Gabrilovich and Nagaraj 2009). It has been described that MDSCs have the potential to promote de novo development of Tregs.(Foell, Wittkowski et al. 2007, Rodriguez and Ochoa 2008, Gabrilovich and Nagaraj 2009).
Histone deacetylases (HDACs) are enzymes that are frequently recruited by transcriptional factors or co-repressors to the gene promoters, where they regulate transcription through chromatin modification without directly binding response elements on DNA. It has also been suggested that some HDACs have a broad range of protein substrates, in addition to factors involved directly in transcription, and have the potential to deacetylase non-histone proteins(Glozak, Sengupta et al. 2005). Despite the rapidly increasing knowledge about the role of HDACs in cancer biology, as well as other pathological conditions such as autoimmunity, it is imperative to delineate specific mechanisms induced by these molecules which govern the physiological outcome of such diseases. Recently, it has been shown that HDAC inhibition enhances MDSC generation and expansion(Condamine and Gabrilovich 2011). Also, important to mention are the new roles assign to specific HDACs which are particularly involved in controlling the immune response(Villagra, Sotomayor et al. 2010). We recently unveiled the role of HDAC11 in the regulation of antigen presenting cells and T cell response(Villagra, Cheng et al. 2009). This deacetylase is the newest member of the histone deacetylase family and has previously been identified as tissue-restricted and exclusively expressed in the brain, kidney and testis(Gao, Cueto et al. 2002). Several studies have also highlighted the role of this HDAC in regulating the differentiation and development of neural cells(Liu, Hu et al. 2008, Liu, Hu et al. 2009). Beyond these studies, little was known regarding the role of HDAC11 in other cell types, until demonstrated by our group that HDAC11, by interacting at the chromatin level with the IL-10 promoter, down-regulates IL-10 transcription in murine and human APCs(Villagra, Cheng et al. 2009). Unpublished data from our lab also suggests that HDAC11 is involved in hematopoietic lineage differentiation, as well as graft vs host disease (GVHD) (both manuscripts in preparation); however the mechanistic role of HDAC11 in myeloid differentiation and MDSC expansion/function still remains to be elucidated. Here we demonstrate that HDAC11 appears to be involved in the regulation of MDSCs in vivo. A better understanding of this previously unknown role of HDAC11 in MDSC biology may lead to the development of targeted epigenetic therapies, in order to modulate the suppressive ability of these cells and augment the efficacy of immunotherapy against autoimmunity, GVHD and malignancies.
2. Materials and Methods
2.1 Flow Cytometry Immunophenotyping
Peripheral blood mononuclear cells (PBMCs), bone marrow aspirates (BM), and splenocytes were harvested under sterile condition. Single-cell suspensions were prepared, and red blood cells were eliminated using ACK lysis buffer (Gibco). Flow cytometric analysis was performed using fluorochrome-labeled monoclonal antibodies (mAbs; anti-CD3, -CD11b, -Ly6C, -Ly6G, Becton Dickinson, San Jose, CA and eBiosciences, San Diego, CA) and the vitality dye 4’,6-diamidino-2-phenylindole (DAPI, Sigma). Data was acquired on an LSRII cytometer (Beckman Coulter), and analyzed with FlowJo software v9.52 (Tree Star, Ashland, OR). Flow cytometric sorting was performed using the same fluorochrome panel and utilizing the FacsARIA (Beckman Coulter) device.
2.2 IFN-gamma Suppression/Functional Assay
Whole spleens and tumors were isolated from tumor burdened or naive mice, cells were isolated and sorted into MDSC population (CD11b+/GR-1+) and their subsets (M-MDSC & GMDSC, Ly6Chigh/Ly6G− & Ly6Clow/Ly6G+ respectively) using FACSAria cell sorter (BD Bioscience) The purity of cell population was 99%. Anti-OVA CD8+ T-cells (OT-I) in the presence -or absence- of cognate peptide (OVA peptide323-339 for CD4+ T-cells and OVA peptide257-264 for CD8+ T-cells) were incubated for 48 hours with MDSCs. An MDSC:OT-I Splenocyte ratio of 1:3, (5 × 104/well MDSC to 15 × 104/well OT-I splenocytes), was used and cells were stimulated with OVA peptide (10ug/mL) and incubated at 37C in a 96 well plate for 48hrs in RPMI/10%FBS. Cytokine production by T-cells was determined using Enzyme-linked immunosorbent assay (ELISA) and Enzyme-linked immunosorbent spot (ELISPOT). For CD8+ T-cells IFN-γ production was measured using an IFN-γ ELISA kit (DY485 R&D Systems) following manufacturer provided protocols.
2.3 Quantitative Reverse Transcriptase-Polymerase Chain Reaction (qRT-PCR)
Total RNA was prepared from centrifugally pelleted and pre-sorted cells (RNeasy mini columns and RNAse free DNAse, Qiagen, Valencia, CA). cDNA was prepared using iScript cDNA Synthesis Kit (Bio-Rad) and qRT-PCR reactions were conducted using the SYBR green two-step qRT-PCR (Bio-Rad) with transcript-specific primers (Supplied upon request) and cDNA from MDSCs as templates. qRT-PCR amplification reactions were resolved on CFX iCycler (Bio-Rad) and fold changes were quantified (2 − ΔΔC t ).
2.4 Mice and Cell Lines
OT-I mice were purchased from Jackson laboratories, Tg-HDAC11-eGFP(Gong, Zheng et al. 2003) reporter mice were provided by Nathaniel Heintz through the Mutant Mouse Regional Centers, and HDAC11-KO kindly supplied by Merck and obtained from Dr. Seto's lab respectively. Mice were kept in pathogen-free condition and handled in accordance with the requirements of the Guideline for Animal Experiments. EL4 thymoma was purchased from American Type Culture Collection (ATCC) and cultured and maintained in DMEM with 10% FBS, at 5%CO2 and 37°C.
2.5 Statistical Analysis
The statistical significance between values was determined by student's t test. Data were expressed as the mean ± SD. Probability values of p ≤ 0.05 were considered significant.
3. Results
3.1 HDAC11 is differentially expressed in immature myeloid cells
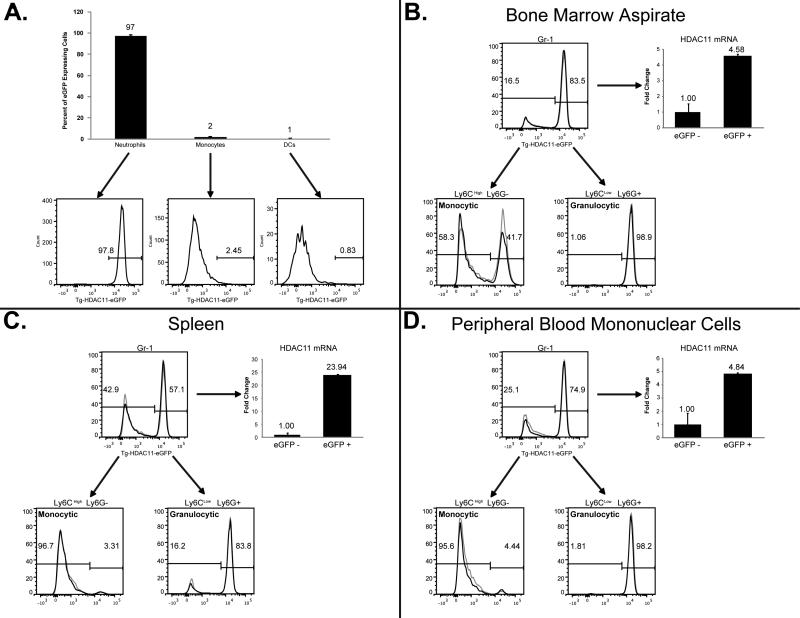
To investigate the endogenous expression of HDAC11 in various hematopoietic compartments, we first examined the expression of HDAC11 in terminally differentiated myeloid cells. Experiments using the Tg-HDAC11-eGFP reporter mice revealed that at steady state, percent of eGFP expressing cells in neutrophils were at the highest (97%) and conversely eGFP expressing cells in the monocytes and dendritic cells (DCs) had the lowest percentage of eGFP at 2% and 1% respectively (Fig.1A). Within the myeloid compartment, precursors of MDSCs are immature myeloid cells (IMCs) which are identified by the expression of CD11b+GR-1+. Next we ventured to examine the expression pattern of HDAC11 in these cells within in the bone marrow (BM), spleen, and peripheral blood mononuclear cell (PBMC) compartments. To accomplish this task, HDAC11 promoter-driven eGFP reporter transgenic mice (Tg-HDAC11-eGFP) were used to evaluate dynamic changes in HDAC11 gene expression (transcriptional activation) activity by evaluating the eGFP expressing cells in each compartment. Results demonstrated that about 90% of all Gr-1+CD11b+ IMCs present in the BM were positive for HDAC11 transcription (Figure 1B). This percentage changes in the spleen to 57% and (Figure 1C) significantly decreases to 27% in the peripheral blood (Figure 1D). Once we looked further and analyzed the expression of eGFP in the granulocytic and monocytic compartments of IMCs in these tissues, we noticed that almost the entire granulocytic populations in all three compartments were active for HDAC11 transcript. The monocytic compartment however painted a different image where monocytic IMCs, even though largely negative for eGFP expression, gradually became entirely devoid of eGFP expression, when transitioning from BM to the spleen, and the PBMCs respectively (Fig. 1). Overall, results suggest that HDAC11 is differentially expressed in various myeloid cells and appears be associated in the lineage differentiation and the fate of monocytic and granulocytic differentiation/maturation.
Figure 1. The expression of HDAC11 in different compartments of IMCs at steady state (without tumor challenge).
Bone marrow aspirate (A & B), splenocytes (C), and PBMCs (D) were isolated from two naive Tg-HDAC11-eGFP mice. Using flow cytometric analysis, first expression of HDAC11 transcript (by examining the expression of eGFP protein) in the neutrophils, monocytes, and DCs (A) were assessed. Next, expression of eGFP in CD11b+/GR-1+ as well as CD11b+Ly6G−/Ly6Chigh and CD11b+/Ly6G+/Ly6Clow compartments of IMCs were determined. The percentages depicted in these histograms are indicative of HDAC11 promoter-driven eGFP reporter gene expression. q-RT-PCR analysis further demonstrates that in this transgenic model, eGFP expression is consistent with HDAC11 mRNA expression. Data presented here is a representative of three individual experiments.
3.2 Expression of HDAC11 changes CD11b+/GR-1+ compartment concomitantly with the expansion of MDSCs in tumor challenged Tg-HDAC11-eGFP mice
In these series of experiments, we subcutaneously inoculated Tg-HDAC11-eGFP mice with either 2.5 × 105 EL4 cells or Hank's Balanced Salt Solution (HBSS) (vehicle control for cell suspension). After 25 days, these mice were euthanized and the spleens as well as the tumors (from tumor bearers) were removed. MDSCs from spleens and tumors were isolated, analyzed and quantified for the expression of eGFP. Markedly, the CD11b+/GR-1+ cells isolated from the spleen of tumor bearing mice appeared less positive for the expression of eGFP reporter gene when compared to HBSS control naïve mice demonstrated by poly-chromatic flow analysis (Figure 2A). In this figure, the expression of eGFP is represented as green dots on the flow plot. As the MDSCs expand in the spleen of tumor bearing mice, the dots become black which signifies loss of eGFP expression. Interestingly, when the percentage of each population from our flow analysis was graphed, MDSCs isolated from the tumor itself displayed the most significant reduction in the expression of eGFP (Figure 2B). This observation highlights the conceivable role of HDAC11 in the expansion of MDSCs. To verify our model, in Figure 2C and D we also analyzed and quantified the expression of eGFP as well as HDAC11 message. Together the data proposes that HDAC11 plays a role in the expansion of MDSCs.
Figure 2. The expression of HDAC11 message in tumor challenged TgHDAC11-eGFP mice.
(A) (Top panel) Flow cytometric data analysis demonstrating the destitution of CD11b+/GR-1+ cells in a naïve TgHDAC11-eGFP mice (Top-Left) and the expansion of MDSCs in EL4 tumor challenged TgHDAC11-eGFP mice (Day 25 after inoculation of tumors sub cutaneous 2.5 × 105 cells/injection). The expansions of these cells were compared to the IMCs percentage in naïve mice. Concurrently, polychromatic representation of data was utilized to highlight the changes in expression of HDAC11 transcript. (Black dots represent HDAC11- while green dots show HDAC11+ cells) (Top-right). (B) A graphic demonstration of eGFP negative percent of CD11b+/GR-1+ cell population in naïve splenocytes compared to EL4 tumor challenged (Day 25) mouse splenocytes as well as tumor cells. (C) q-RT-PCR data analysis for eGFP message expression in CD11b+/GR-1+ cell population isolated from naïve and EL4 tumor challenged Tg-HDAC11-eGFP mice. (D) q-RT-PCR data analysis for HDAC11 message expression in CD11b+/GR-1+ cell population isolated from naïve and EL4 tumor challenged Tg-HDAC11-eGFP mice. The flow data represented in this figure was analyzed by collecting 50,000 events and is a representative figure out of three individual experiments.
3.3 CD11b+/GR-1+/eGFP− population of MDSCs are more suppressive when compared to CD11b+/GR-1+/eGFP+
To understand the physiological consequence of HDAC11 absence in MDSC function, we designed an experiment to study the suppressive capacity of MDSCs lacking HDAC11 (CD11b+/GR-1+/eGFP−) when compared to the MDSCs expressing HDAC11 message (CD11b+/GR-1+/eGFP+). Briefly, Tg-HDAC11-eGFP mice were subcutaneously inoculated with 2.5 × 105 EL4 cells, and on day 24 the mice were euthanized and eGFP+ and eGFP− MDSCs were sorted. Using our OT-I transgenic mouse model, in an ELISA assay we compared the suppressive ability of CD11b+/GR-1+/eGFP+ MDSCs with CD11b+/GR-1+/eGFP−. Although both samples reduced the induction of IFN-γ by OT-I responder cells after stimulation with SIINFEKL OVA peptide, the eGFP negative MDSCs showed a more suppression when compared to the eGFP positive cells (Figure 3). This observation adds to our previously described finding that indeed loss of HDAC11 is associated with the changes seen not only in the expansion of MDSCs but also in the acquisition of their function.
Figure 3. Suppressive capacity of GR-1+ eGFP negative MDSCs vs GR-1+ eGFP positive MDSCs.
(A) eGFP+ and or eGFP- tumor MDSCs were sorted (FACSAria Sorting BD) by either GR-1+/eGFP− (HDAC11-) and or GR-1+/eGFP+ (HDAC11+) populations from three EL4 tumor challenged mice (24 days). Functional assay analysis in this experiment was performed using the OT-I transgenic mouse/OVA-peptide CD8 T cells stimulation model. Functionality of T cells from OT-I mice in the presence -or absence of cognate peptide (OVA peptide257-264 for CD8+ T-cells) were measured by their capacity to produce IFN-γ upon peptide stimulation and in the presence or absence of MDSCs. Probability values of p ≤ 0.05 were considered significant in eGFP negative MDSCs vs eGFP Positive MDSCs. The bar graph is a representative functional assay ELISA analysis for IFN-γ production from three independent experiments.
3.4 MDSCs devoid of HDAC11 are more suppressive
In this section of our studies, HDAC11-KO mice along with C57BL/6 wild type mice were inoculated with EL4 tumor cells as described above. The MDSCs were isolated from the spleen of these mice and using OT-I responder cells and OVA peptide stimulation, IFN-γ production was measured by an ELISA assay. As seen in Figure 4, MDSCs isolated from the HDAC11-KO mice appear to be more suppressive than the control mouse MDSCs. Interestingly, numerous laboratories have demonstrated that per cell basis MMDSCs are more suppressive when compared to GMDSCs(Nausch, Galani et al. 2008, Dolcetti, Peranzoni et al. 2010, Gabrilovich, Ostrand-Rosenberg et al. 2012, Youn, Kumar et al. 2013) which may in part shed some light on our observation in our HDAC11-KO mouse studies. Observations from this part of our studies reiterate the associative role HDAC11 plays in suppressive capacity of MDSC function. Also, demonstrated in Figure 5, we observed that deficiency of HDAC11 induces the up-regulation of suppressive cytokine IL-10, which in part may be involved in the overall suppressive phenotype observed in HDAC11-KO mice. Notably, tumor growth kinetics is enhanced in mice lacking HDAC11, as observed in two different tumor models (Figure 6), indicating a plausible negative regulatory association by HDAC11.
Figure 4. Suppressive capacity of isolated MDSCs from tumor challenged C57BL/6 WT vs HDAC11-KO mice.
(A) HDAC11-KO mice and their control counter parts C57BL/6 WT mice were inoculated with EL4 tumors (sub cutaneous 2.5 × 105 cells/injection for 24 day as previously described. Splenocytes for each animal group (3 mice/group) were harvested, isolated and sorted (FACSAria Sorting BD) for CD11b+/GR-1+. Functional assay analysis in this experiment was performed using the OT-I transgenic mouse/OVA-peptide CD8 T cell stimulation model. Functionality of T cells from OT-I mice in the presence -or absence of cognate peptide (OVA peptide257-264 for CD8+ T-cells) were measured by their capacity to produce IFN-γ upon peptide stimulation and in the presence or absence of MDSCs. Probability values of p ≤ 0.05 were considered significant in C57BL/6 WT MDSCs vs HDAC11-KO MDSCs. The bar graph is a representative functional assay ELISA analysis for IFN-γ production from three independent experiments.
Figure 5. Expression of suppressive cytokine IL-10 is increased in HDAC11 null Gr-1+ population at steady state as well as under tumor burden.
(A) HDAC11-KO and their control counter parts C57BL/6 WT mice (no tumor inoculation—steady-state) were euthanized and splenocytes for each animal group (3 mice/group) were harvested, isolated and sorted (FACSAria Sorting BD) for CD11b+/GR-1+. These cells were next treated with or without LPS (1ug/mL) for 6hrs. IL-10 expression was assessed using qRT-PCR analysis. (B) eGFP+ and or eGFP- tumor MDSCs (CD11b+/GR-1+) were sorted by either GR-1+/eGFP- (HDAC11-) and or GR-1+/eGFP+ (HDAC11+) populations from three EL4 tumor challenged mice (24 days). Probability values of p ≤ 0.05 were considered significant in comparing populations within groups. The bar graph illustrations for the expression IL-10 is a representative figure from two independent experiments.
Figure 6. HDAC11 deficient mice demonstrate a more enhanced tumor growth when compared to C57BL/6 wild-type counterparts.
HDAC11-KO and their control counterparts C57BL/6 WT mice (3 mice per group) were inoculated with sub cutaneous injection of EL4 cells at 2.5 × 105 cells/injection for 21 days (A) or Panco cells at 5 × 104 for 23 days (B). Tumors were measured at 3 day intervals once palpable. Graphs presented here are linear representation of tumor growth in each tumor model. Probability values of p ≤ 0.05 were considered significant in comparing populations within groups. The graph representation in figure 6A is pooled data from two independent EL4 tumor growth experiments and figure 6B is a representative graph of 2 independent Panco tumor experiments.
Hematopoiesis in general is typically a well-regulated scheme intended to replenish blood cells at a perpetual and steady rate, sustaining equilibrium in the myeloid and lymphoid compartments. Nonetheless, the frequency of yield in certain blood cell types can be changed by conditions such as acute inflammation/infection, leukemia, and radiation or chemical damage(Fuchs, Hausen et al. 1991, Najman, Kobari et al. 1991, Mauch, Constine et al. 1995, Chandra, Villanueva et al. 2008). The condition termed “emergency hematopoiesis”(Zhan, Lieschke et al. 1998, Basu, Hodgson et al. 2000), is characterized by increased and rapid production of myeloid cells, occurring during acute physiological stress such as infection. In order to further evaluate the role of HDAC11 in myelopoiesis, we conducted a preliminary emergency hematopoiesis experiment where we induced myelopoiesis using Freund's complete adjuvant. Strikingly, HDAC11-KO mice had 3 fold higher expansions of GR-1+ population when compared to the wild type control C57BL/6 mice (Data not shown). These findings again highlight the possible regulatory role HDAC11 plays in myelopoiesis.
4 Discussion
Myeloid cells have vast and diverse functions and signify the most abundant hematopoietic cells. In this study we demonstrated that HDAC11 is differentially expressed in various lymphoid compartments and its expression appears to be significant in granulocytic vs monocytic differentiation in myeloid cells. We further demonstrated the role of HDAC11 in the expansion of MDSC as Tg-HDAC11-eGFP mice inoculated with EL4 cells, revealed a decrease in the expression of HDAC11 message in the CD11b/GR-1+ population. These interesting findings are correlated with a more suppressive phenotype of MDSCs in the OT-I transgenic mouse model. In this study we hypothesized that these findings are perhaps due to additional regulation of MDSCs by factors produced/regulated by HDAC11. To support our hypothesis and using the HDAC11-KO tumor-bearing mice, we demonstrated that MDSCs isolated from the HDAC11 deficient mice were more suppressive on per-cell basis when compared to the wild-type counterparts. These observations become more evident as we demonstrate that HDAC11 deficient mice demonstrate a more enhanced tumor growth kinetics when compared to the wild-type controls. We do acknowledge that a germ-line knock-out of HDAC11 may consequently have other immunological aberrations in various compartments of the immune system (As of yet, no disease or developmental aberrations have been reported in these mice), and that these possible changes may affect the overall extrapolative outcome in the total knock-out murine model; therefore we are further investigating the exact mechanistic role of HDAC11 in the process of MDSC function and expansion. Using the Cre-transgenic mouse technology, we are in the final stages of developing targeted HDAC11-KO in the myeloid compartment of mice, which will allow us to observe the direct consequence of HDAC11 deficiency in the myeloid compartment—while evading other possible physiological influences other immune cell compartments.
To this day, there has been no study in the MDSC field investigating the role of HDAC11. However, there has been new evidence linking HDACs with factors that are ultimately responsible for the induction of these cells. Recently, a study from our lab has revealed a novel regulatory role of HDAC6 in STAT3 activation. The additional demonstration that HDAC6 is required for STAT3 phosphorylation and recruitment to the nucleus highlights the important role HDACs play in the activation of factors such as STAT3 and reiterates their integral role in MDSC biology (Cheng et al accepted for publication JI July, 2014).
New findings propose that STAT3 also regulates MDSC expansion by inducing the expression of S100A8 and S100A9(Yang, Cai et al. 2006). STAT3-dependent induction of these proteins by myeloid progenitors halts the differentiation of immature myeloid cells resulting in the expansion of MDSCs ultimately(Huang, Pan et al. 2006). Preliminary data from our lab has also revealed that S100A8 and S100A9 appear to be unregulated in the splenocytes of HDAC11-KO mice when compared to wild-type counterpart (data not shown). We are currently investigation the role of these proteins in this phenomenon. More recently, Youn et al have revealed yet another HDAC—HDAC2 that is involved in silencing of the Rb1 MDSCs(Youn, Kumar et al. 2013) which reiterates the importance of investigating these epigenetic factors in the context of MDSC biology. There is also new evidence that HDAC6 and HDAC11 proteins physically interact and are being recruited to the IL-10 gene promoter dictating dynamic transcriptional responses(Cheng, Lienlaf et al. 2014).
In the past several years, the role of tumor-induced MDSCs in cancer immune suppression has been widely recognized(Kusmartsev and Gabrilovich 2006, Talmadge 2007, Nagaraj and Gabrilovich 2010, Condamine and Gabrilovich 2011). These findings have emphasized the heterogeneity of these cells(Serafini, Borrello et al. 2006) where immature myeloid cells are present at various stages of differentiation(Youn, Collazo et al. 2012). This heterogeneity along with the fact that MDSCs are induced in response to variety of tumor-derived factors presents a challenge in depicting a distinct and clear mechanism. Hence, the characterization of specific pathways responsible for induction of these cells is yet to be elucidated. Here in this study, we have identified HDAC11 as one possible culprit which may in part elucidate a plausible pathway responsible for induction and function of MDSCs.
Observation and findings in this manuscript provide important insights into the regulatory role HDAC11 plays in MDSC expansion and function. These studies also provide a platform to further interrogate the role of HDAC11 in myeloid differentiation. Furthermore, these findings raise implications for the therapeutic modulation of this HDAC, with the potential to control the expansion and function of MDSCs in cancer immunotherapy.
Acknowledgments
We acknowledge the flow cytometry core facilities at H. Lee Moffitt Cancer Center and their extended technical support for our project.
This work was supported by National Genomic Center US Army W81XWH-08-2-0101 & National Institutes of Health NIH-RCA184612A
Abbreviation
- MDSC
myeloid derived suppressor cells
- IMC
immature myeloid cells
- HDAC
histone deacetylase
- eGFP
enhanced green fluorescent protein
- APCs
antigen presenting cells
- ROS
reactive oxygen species
- NO
nitric oxide
- GVHD
graft vs host disease
- PBMC
peripheral blood mononuclear cells
- BM
bone marrow
- DAPI
diamidino-2-phenylindole
- DCs
dendritic cells
- EL4
murine lymphoma cell line
- Panco
murine pancreatic adenocarcinoma cell line
- LPS
lipopolysaccharide
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Disclosure of Potential Conflicts of Interest:
No potential conflicts of interest were disclosed.
References
- Almand B, Clark JI, Nikitina E, van Beynen J, English NR, Knight SC, Carbone DP, Gabrilovich DI. Increased production of immature myeloid cells in cancer patients: a mechanism of immunosuppression in cancer. J Immunol. 2001;166(1):678–689. doi: 10.4049/jimmunol.166.1.678. [DOI] [PubMed] [Google Scholar]
- Basu S, Hodgson G, Zhang HH, Katz M, Quilici C, Dunn AR. Emergency” granulopoiesis in G-CSF-deficient mice in response to Candida albicans infection. Blood. 2000;95(12):3725–3733. [PubMed] [Google Scholar]
- Bronte V, Zanovello P. Regulation of immune responses by L-arginine metabolism. Nat Rev Immunol. 2005;5(8):641–654. doi: 10.1038/nri1668. [DOI] [PubMed] [Google Scholar]
- Chandra R, Villanueva E, Feketova E, Machiedo GW, Hasko G, Deitch EA, Spolarics Z. Endotoxemia down-regulates bone marrow lymphopoiesis but stimulates myelopoiesis: the effect of G6PD deficiency. J Leukoc Biol. 2008;83(6):1541–1550. doi: 10.1189/jlb.1207838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng F, Lienlaf M, Perez-Villarroel P, Wang HW, Lee C, Woan K, Woods D, Knox T, Bergman J, Pinilla-Ibarz J, Kozikowski A, Seto E, Sotomayor EM, Villagra A. Divergent roles of histone deacetylase 6 (HDAC6) and histone deacetylase 11 (HDAC11) on the transcriptional regulation of IL10 in antigen presenting cells. Mol Immunol. 2014;60(1):44–53. doi: 10.1016/j.molimm.2014.02.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Condamine T, Gabrilovich DI. Molecular mechanisms regulating myeloid-derived suppressor cell differentiation and function. Trends Immunol. 2011;32(1):19–25. doi: 10.1016/j.it.2010.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delano MJ, Scumpia PO, Weinstein JS, Coco D, Nagaraj S, Kelly-Scumpia KM, O'Malley KA, Wynn JL, Antonenko S, Al-Quran SZ, Swan R, Chung CS, Atkinson MA, Ramphal R, Gabrilovich DI, Reeves WH, Ayala A, Phillips J, Laface D, Heyworth PG, Clare-Salzler M, Moldawer LL. MyD88-dependent expansion of an immature GR-1(+)CD11b(+) population induces T cell suppression and Th2 polarization in sepsis. J Exp Med. 2007;204(6):1463–1474. doi: 10.1084/jem.20062602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dolcetti L, Peranzoni E, Ugel S, Marigo I, Fernandez Gomez A, Mesa C, Geilich M, Winkels G, Traggiai E, Casati A, Grassi F, Bronte V. Hierarchy of immunosuppressive strength among myeloid-derived suppressor cell subsets is determined by GM-CSF. Eur J Immunol. 2010;40(1):22–35. doi: 10.1002/eji.200939903. [DOI] [PubMed] [Google Scholar]
- Ezernitchi AV, Vaknin I, Cohen-Daniel L, Levy O, Manaster E, Halabi A, Pikarsky E, Shapira L, Baniyash M. TCR zeta down-regulation under chronic inflammation is mediated by myeloid suppressor cells differentially distributed between various lymphatic organs. J Immunol. 2006;177(7):4763–4772. doi: 10.4049/jimmunol.177.7.4763. [DOI] [PubMed] [Google Scholar]
- Foell D, Wittkowski H, Vogl T, Roth J. S100 proteins expressed in phagocytes: a novel group of damage-associated molecular pattern molecules. J Leukoc Biol. 2007;81(1):28–37. doi: 10.1189/jlb.0306170. [DOI] [PubMed] [Google Scholar]
- Fuchs D, Hausen A, Reibnegger G, Werner ER, Werner-Felmayer G, Dierich MP, Wachter H. Immune activation and the anaemia associated with chronic inflammatory disorders. Eur J Haematol. 1991;46(2):65–70. doi: 10.1111/j.1600-0609.1991.tb00524.x. [DOI] [PubMed] [Google Scholar]
- Gabrilovich DI, Nagaraj S. Myeloid-derived suppressor cells as regulators of the immune system. Nat Rev Immunol. 2009;9(3):162–174. doi: 10.1038/nri2506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gabrilovich DI, Ostrand-Rosenberg S, Bronte V. Coordinated regulation of myeloid cells by tumours. Nat Rev Immunol. 2012;12(4):253–268. doi: 10.1038/nri3175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao L, Cueto MA, Asselbergs F, Atadja P. Cloning and functional characterization of HDAC11, a novel member of the human histone deacetylase family. J Biol Chem. 2002;277(28):25748–25755. doi: 10.1074/jbc.M111871200. [DOI] [PubMed] [Google Scholar]
- Glozak MA, Sengupta N, Zhang X, Seto E. Acetylation and deacetylation of non histone proteins. Gene. 2005;363:15–23. doi: 10.1016/j.gene.2005.09.010. [DOI] [PubMed] [Google Scholar]
- Gomez-Garcia L, Lopez-Marin LM, Saavedra R, Reyes JL, Rodriguez-Sosa M, Terrazas LI. Intact glycans from cestode antigens are involved in innate activation of myeloid suppressor cells. Parasite Immunol. 2005;27(10-11):395–405. doi: 10.1111/j.1365-3024.2005.00790.x. [DOI] [PubMed] [Google Scholar]
- Gong S, Zheng C, Doughty ML, Losos K, Didkovsky N, Schambra UB, Nowak NJ, Joyner A, Leblanc G, Hatten ME, Heintz N. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature. 2003;425(6961):917–925. doi: 10.1038/nature02033. [DOI] [PubMed] [Google Scholar]
- Hestdal K, Ruscetti FW, Ihle JN, Jacobsen SE, Dubois CM, Kopp WC, Longo DL, Keller JR. Characterization and regulation of RB6-8C5 antigen expression on murine bone marrow cells. J Immunol. 1991;147(1):22–28. [PubMed] [Google Scholar]
- Huang B, Pan PY, Li Q, Sato AI, Levy DE, Bromberg J, Divino CM, Chen SH. Gr-1+CD115+ immature myeloid suppressor cells mediate the development of tumor-induced T regulatory cells and T-cell anergy in tumor-bearing host. Cancer Res. 2006;66(2):1123–1131. doi: 10.1158/0008-5472.CAN-05-1299. [DOI] [PubMed] [Google Scholar]
- Kusmartsev S, Gabrilovich DI. Role of immature myeloid cells in mechanisms of immune evasion in cancer. Cancer Immunol Immunother. 2006;55(3):237–245. doi: 10.1007/s00262-005-0048-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H, Hu Q, D'Ercole A J, Ye P. Histone deacetylase 11 regulates oligodendrocyte-specific gene expression and cell development in OL-1 oligodendroglia cells. Glia. 2009;57(1):1–12. doi: 10.1002/glia.20729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H, Hu Q, Kaufman A, D'Ercole AJ, Ye P. Developmental expression of histone deacetylase 11 in the murine brain. J Neurosci Res. 2008;86(3):537–543. doi: 10.1002/jnr.21521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marigo I, Dolcetti L, Serafini P, Zanovello P, Bronte V. Tumor-induced tolerance and immune suppression by myeloid derived suppressor cells. Immunol Rev. 2008;222:162–179. doi: 10.1111/j.1600-065X.2008.00602.x. [DOI] [PubMed] [Google Scholar]
- Mauch P, Constine L, Greenberger J, Knospe W, Sullivan J, Liesveld JL, Deeg HJ. Hematopoietic stem cell compartment: acute and late effects of radiation therapy and chemotherapy. Int J Radiat Oncol Biol Phys. 1995;31(5):1319–1339. doi: 10.1016/0360-3016(94)00430-S. [DOI] [PubMed] [Google Scholar]
- Nagaraj S, Gabrilovich DI. Myeloid-derived suppressor cells in human cancer. Cancer J. 2010;16(4):348–353. doi: 10.1097/PPO.0b013e3181eb3358. [DOI] [PubMed] [Google Scholar]
- Najman A, Kobari L, Khoury E, Baillou CL, Lemoine F, Guigon M. Suppression of normal hematopoiesis during acute leukemias. Ann N Y Acad Sci. 1991;628:140–147. doi: 10.1111/j.1749-6632.1991.tb17231.x. [DOI] [PubMed] [Google Scholar]
- Nausch N, Galani IE, Schlecker E, Cerwenka A. Mononuclear myeloid-derived “suppressor“ cells express RAE-1 and activate natural killer cells. Blood. 2008;112(10):4080–4089. doi: 10.1182/blood-2008-03-143776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodriguez PC, Ochoa AC. Arginine regulation by myeloid derived suppressor cells and tolerance in cancer: mechanisms and therapeutic perspectives. Immunol Rev. 2008;222:180–191. doi: 10.1111/j.1600-065X.2008.00608.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawanobori Y, Ueha S, Kurachi M, Shimaoka T, Talmadge JE, Abe J, Shono Y, Kitabatake M, Kakimi K, Mukaida N, Matsushima K. Chemokine-mediated rapid turnover of myeloid-derived suppressor cells in tumor-bearing mice. Blood. 2008;111(12):5457–5466. doi: 10.1182/blood-2008-01-136895. [DOI] [PubMed] [Google Scholar]
- Serafini P, Borrello I, Bronte V. Myeloid suppressor cells in cancer: recruitment, phenotype, properties, and mechanisms of immune suppression. Semin Cancer Biol. 2006;16(1):53–65. doi: 10.1016/j.semcancer.2005.07.005. [DOI] [PubMed] [Google Scholar]
- Talmadge JE. Pathways mediating the expansion and immunosuppressive activity of myeloid-derived suppressor cells and their relevance to cancer therapy. Clin Cancer Res. 2007;13(18 Pt 1):5243–5248. doi: 10.1158/1078-0432.CCR-07-0182. [DOI] [PubMed] [Google Scholar]
- Villagra A, Cheng F, Wang HW, Suarez I, Glozak M, Maurin M, Nguyen D, Wright KL, Atadja PW, Bhalla K, Pinilla-Ibarz J, Seto E, Sotomayor EM. The histone deacetylase HDAC11 regulates the expression of interleukin 10 and immune tolerance. Nat Immunol. 2009;10(1):92–100. doi: 10.1038/ni.1673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villagra A, Sotomayor EM, Seto E. Histone deacetylases and the immunological network: implications in cancer and inflammation. Oncogene. 2010;29(2):157–173. doi: 10.1038/onc.2009.334. [DOI] [PubMed] [Google Scholar]
- Wang RF. Regulatory T cells and innate immune regulation in tumor immunity. Springer Semin Immunopathol. 2006;28(1):17–23. doi: 10.1007/s00281-006-0022-7. [DOI] [PubMed] [Google Scholar]
- Wells AD. Cell-cycle regulation of T-cell responses--novel approaches to the control of alloimmunity. Immunol Rev. 2003;196:25–36. doi: 10.1046/j.1600-065x.2003.00080.x. [DOI] [PubMed] [Google Scholar]
- Yang R, Cai Z, Zhang Y, Yutzy W. H. t., Roby KF, Roden RB. CD80 in immune suppression by mouse ovarian carcinoma-associated Gr-1+CD11b+ myeloid cells. Cancer Res. 2006;66(13):6807–6815. doi: 10.1158/0008-5472.CAN-05-3755. [DOI] [PubMed] [Google Scholar]
- Youn JI, Collazo M, Shalova IN, Biswas SK, Gabrilovich DI. Characterization of the nature of granulocytic myeloid-derived suppressor cells in tumor-bearing mice. J Leukoc Biol. 2012;91(1):167–181. doi: 10.1189/jlb.0311177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youn JI, Gabrilovich DI. Eur J Immunol. 2010;The biology of myeloid-derived suppressor cells: the blessing and the curse of morphological and functional heterogeneity.40(11):2969–2975. doi: 10.1002/eji.201040895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youn JI, Kumar V, Collazo M, Nefedova Y, Condamine T, Cheng P, Villagra A, Antonia S, McCaffrey JC, Fishman M, Sarnaik A, Horna P, Sotomayor E, Gabrilovich DI. Epigenetic silencing of retinoblastoma gene regulates pathologic differentiation of myeloid cells in cancer. Nat Immunol. 2013;14(3):211–220. doi: 10.1038/ni.2526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youn JI, Nagaraj S, Collazo M, Gabrilovich DI. Subsets of myeloid-derived suppressor cells in tumor-bearing mice. J Immunol. 2008;181(8):5791–5802. doi: 10.4049/jimmunol.181.8.5791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhan Y, Lieschke GJ, Grail D, Dunn AR, Cheers C. Essential roles for granulocyte-macrophage colony-stimulating factor (GM-CSF) and G-CSF in the sustained hematopoietic response of Listeria monocytogenes-infected mice. Blood. 1998;91(3):863–869. [PubMed] [Google Scholar]