Abstract
Background
Olive cultivation blends with the history of the Mediterranean countries since ancient times. Even today, activities around the olive tree constitute major engagements of several people in the countryside of both sides of the Mediterranean basin. The olive fly is, beyond doubt, the most destructive pest of cultivated olives. The female fly leaves its eggs in the olive fruit. Upon emergence, the larvae feed on the olive sap, thus destroying the fruit. If untreated, practically all olives get infected. The use of chemical insecticides constitutes the principal olive fly control approach. The Sterile Insect Technique (SIT), an environmentally friendly alternative control method, had been tried in pilot field applications in the 1970's, albeit with no practical success. This was mainly attributed to the low, non-antagonistic quality of the mixed-sex released insects. Many years of experience from successful SIT applications in related species, primarily the Mediterranean fruit fly, Ceratitis capitata, demonstrated that efficient SIT protocols require the availability of fundamental genetic and molecular information.
Results
Among the primary systems whose understanding can contribute towards novel SIT approaches (or its recently developed alternative RIDL: Release of Insects carrying a Dominant Lethal) is the reproductive, since the ability to manipulate the reproductive system would directly affect the insect's fertility. In addition, the analysis of early embryonic promoters and apoptotic genes would provide tools that confer dominant early-embryonic lethality during mass-rearing. Here we report the identification of several genes involved in these systems through whole transcriptome analysis of female accessory glands (FAGs) and spermathecae, as well as male testes. Indeed, analysis of differentially expressed genes in these tissues revealed higher metabolic activity in testes than in FAGs/spermathecae. Furthermore, at least five olfactory-related genes were shown to be differentially expressed in the female and male reproductive systems analyzed. Finally, the expression profile of the embryonic serendipity-α locus and the pre-apoptotic head involution defective gene were analyzed during embryonic developmental stages.
Conclusions
Several years of molecular studies on the olive fly can now be combined with new information from whole transcriptome analyses and lead to a deep understanding of the biology of this notorious insect pest. This is a prerequisite for the development of novel embryonic lethality female sexing strains for successful SIT efforts which, combined with improved mass-reared conditions, give new hope for efficient SIT applications for the olive fly.
Keywords: Tephritidae, Bactrocera oleae, sry-α, hid
Background
When Athena, the goddess of peace and wisdom, offered an olive tree to the people of Attica to sway them into choosing her name for their city - and not that of her brother's Poseidon - neither she nor the people of Attica were aware of the 'worm' that could destroy the precious fruit of that tree. That was described much later in the 3rd century AD, by the botanist Theophrastus who, in his works "Enquiry into Plants" and "Causes of Plants" [1], talked about the 'worm underneath the skin of the olive that destroys the fruit'. Indeed, the female olive fly (Bactrocera oleae, Rossi) lays her eggs in an olive fruit and the resulting larva feeds on the olive sap, opening channels inside it, thus destroying it. In this way, a female fly can damage more than 300 olives in her lifetime. Given the fact that during the summer and fall months about five generations of these flies are born, one can imagine the cumulative damage that can take place in an olive orchard. If untreated, practically every single olive will get infested. It is estimated that due to olive fly infestation olive oil production is reduced by more than 30% annually [2].
Control of these flies is traditionally based on cover or bait sprays with chemical insecticides. During the last 40-50 years, organophosphate insecticides have been extensively used against the olive fly, mainly dimethoate and fenthion. More recently, pyrethroids as well as the naturalyte spinosad have been added in the arsenal against the olive fly. The use of chemical pesticides, however, entails many known hazards. Among these are ecological disturbances, the development and spread of insecticide resistance, harmful toxicological effects on human health [3]. Many of these risks are apparent not only to scientists but also to growers and consumers who require a cleaner and safer environment as well as products of high quality. Alternative, environmentally friendly control methods against insect pests, such as the Sterile Insect Technique (SIT) have been experimented in the past with considerable success [4]. The SIT involves the mass production, sterilization and subsequent release of the sterilized insects [5]. The sterilized males will mate with wild females, whose unfertilized eggs will never hatch, thus reducing the numbers of the following generation. In theory, if continued releases are performed over several consecutive generations, the population will progressively be reduced and, eventually, a total eradication could occur.
Given the substantial economic burden of the olive fly in olive producing countries and the concerns raised about the heavy use of insecticides to control the flies, the SIT was proposed [6] and implemented in two pilot efforts. In the early 1970s, about 150,000 laboratory-reared male and female flies were sterilized by gamma-irradiation and subsequently released in the environment [7]. Although initially the releases seemed to contribute to low infestation levels, by the end of the season olives were as highly infested as in the two nearby control plantations. The sterilized flies were proven ineffective to reduce infestation. Similar results were obtained in a second pilot SIT effort that took place in the late '70s in a small Greek island. These unsuccessful pilot experiments led to funding suspension and the eventual abandonment of the program [8-10]. Apart from the high cost and labor-intensive rearing of the olive fly, extensive research that followed these first pilot efforts revealed several key issues of olive fly biology that should have been sorted out before a successful SIT could be implemented. The first issue regarded assortative mating of the released and wild populations. Laboratory-reared flies mated several hours before scotophase whereas wild flies mated at the end of the photophase [11]. Apparently, mass-laboratory rearing caused substantial alterations in the genetic makeup of the flies due to selective pressures in the artificial laboratory environment [12,13]. The second issue regarded the quality of the radiation-sterilized mass-reared flies. Radiation did not leave the vigor of the flies unaffected [14]. Another factor that probably exacerbated the low fitness of the laboratory reared flies was the use of antibiotics in the flies' diet that destroyed the endosymbiotic bacteria that are now known to play a very important role in the organism's fitness [15-19]. Finally, but equally importantly, extensive stinging of the olive fruits from the released females led to further fungal infestation [7].
Since those early years, several molecular and genetic studies have changed B. oleae's research landscape. First, the development of microsatellite markers [20] and the analysis of the mitochondrial genome [21] have offered tools for a fairly detailed analysis of population structure and dynamics in the Mediterranean basin [22-26]. Second, cytogenetic analysis, including in situ hybridization of several molecular markers, established the details of the chromosomal complement [27-31]. Third, isolation and characterization of various genes has shed light on important processes such as insecticide resistance [32-35], female germline differentiation and morphogenesis of epidermal cells [36], enzyme catalytic mechanisms [37], sex-determining cascades [38,39]. Fourth, an initial assessment of the genome of the olive fly was gained by an accurate estimate of its size [40] and the characterization and analysis of centromeric repeats [41] and several EST loci [42]. This was followed by a whole transcriptome analysis with 454 pyrosequencing [43]. Fifth, B. oleae was successfully transformed with the use of a Minos-based transposon [44]. Transformation efforts recently led to the development of piggyBac-based conditional female-lethal olive fly strains that provide highly penetrant female specific lethality, dominant fluorescent marking and genetic sterility [45]. Sixth, B. oleae was recently trans-infected with a cherry fly Wolbachia strain and shown to induce complete cytoplasmic incompatibility in the fly [46]. Finally, the experience gained during the first two pilot SIT efforts and the relevant research that followed, underlined a few key requirements for the maintenance of high quality and well-fit mass-reared olive flies (reviewed in [47]). Among them were changes in larval and adult diets (eg removal of antibiotics) that would preserve the endosymbiotic flora (that is now known to improve fitness) and occasional enrichments of the long-term laboratory colonies with wild individuals (that provide natural vigor). These achievements have renewed the interest in using SIT for olive fly control. In fact, there is a large international effort led by the Joint Division of the Food and Agricultural Organization and the International Atomic Energy Agency (FAO/IAEA) to develop a vigorous laboratory olive fly strain that could be used in such new SIT efforts.
Further scientific and technological developments, in addition to successful SIT applications in other insects, point to the direction olive fly research could go. Indeed, SIT has proven particularly effective in the medfly, the prototype Tephritid species where most genetic and molecular tools have been developed. One of the most active medfly research areas in recent years has been the development of the RIDL technology. RIDL (Release of Insects carrying a Dominant Lethal; [48,49]) is a variant of the conventional SIT, in which sterilization of the released insects is induced not by irradiation but by homozygocity for a dominant lethal gene. Mating with wild individuals results in offspring that are heterozygous for the lethal gene leading to the death of all progeny [50,51]. This dominant lethal gene can be placed under the control of an inducible early embryonic female promoter [51,52] that could achieve genetic sexing at a very early developmental stage. In this way, both genetic sexing and sterilization can be accomplished by the same construct. One other active research area regards the analysis of biological systems with relevance to SIT. Of particular interest are those that regard reproduction and olfaction. The first one is involved in successful mating and egg development, while the second in food and mate localization. A possible manipulation of either or both of these systems would severely affect the destructive ability of the flies. In that sense, transgenic flies could be developed in which genes regulating food and mate recognition or fertility are knocked-down, over-expressed or mis-expressed (depending on the case). Such flies would be safer and more efficient to be released in control programs in an SIT context.
The falling prices of next generation sequencing make it now possible to sequence the entire transcriptome of non-model organisms under different settings and identify differentially expressed genes relevant to the chosen conditions. Subsequently, these genes can be manipulated in vitro and re-introduced into the genome of the organism through well-established transgenic technologies. In a first attempt to explore the relevant-to-SIT transcriptome of the olive fly, we present differences observed in female and male reproductive systems and we examine the differential expression of olfactory genes in the same tissues. Finally, we assess the developmental expression of two of the most commonly used early embryonic genes.
Results and discussion
1. Sequencing and annotation
1.1. Solid ABI sequencing and reads assembly
In order to explore differentially expressed genes in the transcriptome of reproductive organs of the olive fly that could be useful in SIT development, the entire transcriptomes from female accessory glands and spermathecae were compared to male testes. For transcriptome assembly, the sequences from these two libraries (FEMALE and MALE) were combined with two more obtained from heads of spinosad-sensitive (LAB) and spinosad-resistant (SPIN) olive flies [53]. Paired-end sequencing with 35nt and 50nt read sizes was performed for each library and a total of 122,623,894 read pairs was obtained. All reads of the libraries were pooled to obtain a reference transcriptome assembly using SOAPdenovo assembler [54].
1.2. Sequence annotation
Annotation of the assembled sequences was obtained by aligning the 69,359 assembled B. oleae sequences against the NCBI non-redundant (Nr) protein database using blastx and collecting the annotations with the BLAST2GO tool [55]. Using an E-value threshold of ≤1e-6, 20207 (29.13 %) of the contigs were aligned. Of the 69,359 contigs, 23,042 (33.22%) have almost exact hits in the B. oleae transcriptome of Pavlidi et al [43] (E-value ≤1e-6).
2. Female vs male differential expression
The Cuffdiff [56] tool was used in order to reveal the differentially expressed genes between the reproductive systems of female and male flies, a stringent cutoff (p value adjusted for multiple testing, called q value <0.05) was used. This resulted in 1568 differentially expressed transcripts in the FEMALE vs. MALE comparison. Three hundred and thirty of these transcripts were up-regulated in FEMALE, while 1238 were up-regulated in MALE B. oleae flies. The top 40 up-regulated genes in each category are listed in Table S1. The entire lists of all significantly (q<0.05) up-regulated genes in FEMALE and MALE are given in Tables S3 and S4, respectively.
An M-A plot was constructed for comparison of the genes for FEMALE vs MALE flies with q value < 0.05. In Figure 1 the de-regulated genes are depicted in red.
Figure 1.
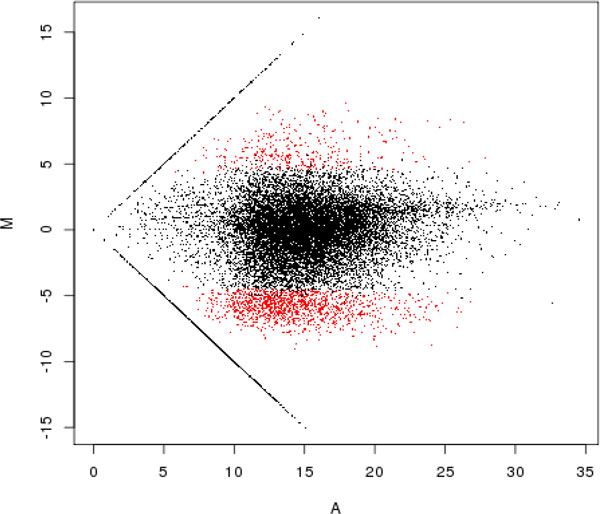
M-A plot of gene expression for female and male flies. Genes up-regulated in females have positive M values. Significantly differentially expressed genes (q-value < 0.05) are shown in red.
Functional annotation was made for the assembled sequences of the significantly differentially expressed female- and male- specific genes mentioned in Table S1, based on gene ontology (GO) categorization obtained using BLAST2GO. The FEMALE and MALE GO analysis performed for biological process of the top 40 female and male expressed genes is shown in Figure 2. In general, more GO terms appear in female tissues than in male (16 vs 12), a point that holds even in deeper GO-term analysis. This can be attributed to the fact that the FEMALE library was comprised of both FAGs and spermathecae, while the MALE from testes only. Furthermore, there were more male- than female-specific genes involved in metabolism and development, a fact that can be attributed to sperm activity in the MALE tissue. Finally, the presence of three immune system process genes in the female list should be noted. In fact, increased levels of immune response genes have been found in transcriptome analyses of insect female reproductive systems, particularly after mating [57,58]. Upregulation of these genes may assist females to combat pathogens introduced during copulation. Alternatively, it could be a result of female's perception of sperm as non-self molecules.
Figure 2.
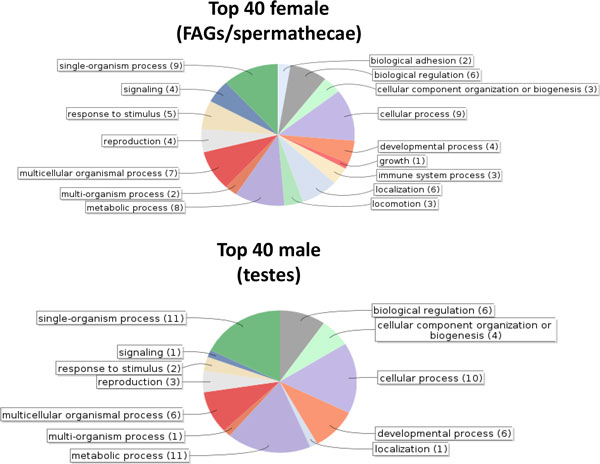
GO Term associations for the top 40 genes expressed in the female and male tissues. Associations were identified with BLAST2GO, using terms at the second level of the GO hierarchy.
A more direct comparison between FEMALE-only and MALE-only GO-term distribution is shown in Figure 3. Interestingly, numbers of GO-terms for biological process appear different in the two datasets, suggesting a different complexity of the studied female and male reproductive tissues. In most terms, there are more male- than female-specific transcripts that are differentially expressed. Many of these terms (cell cycle, intracellular organelle part, primary metabolic process, organic substance metabolic process, macromolecule metabolic process, cellular metabolic process, multicellular organismal development) refer to higher metabolic processes. This could be attributed to higher metabolic and cellular activity that takes place in the testes before mating.
Figure 3.
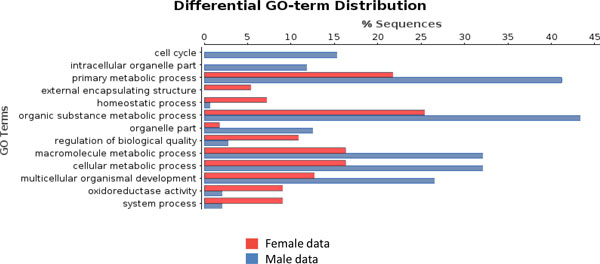
GO-terms (GO-Slim) differentially distributed between male and female transcriptomes. Contigs expressed only in female tissue are used as the test set (red bars) and contigs expressed only in male tissues as the reference set (blue bars).
3. Genes that might be implicated in sexual differentiation in B. oleae
In order to validate the differential expression of various genes observed after the RNAseq analysis of reproductive tissues of female and male olive flies, further functional analysis was performed for twelve genes that were differentially expressed in female accessory glands and spermathecae, on one hand, and male testes, on the other (Figure 4). These genes were selected on the basis of known involvement in sexual differentiation in other insects. Seven of them were selected from the 1238 significantly up-regulated in MALE (Table S4): kl2 (male fertility factor kl2), kl3 (male fertility factor kl3), kl5 (male fertility factor kl5), ory (occludin-related Y protein), fem-1 (sex-determining protein fem-1), gas8 (growth arrest specific protein 8) and lobo (lost boys). Three more genes that were up-regulated in MALE [ix (intersex), pbl (pebble) and hcf (host cell factor C1)] and two that were up-regulated in FEMALE [sox and pcp (pupal cuticle protein 78E)], albeit with lower statistical power (i.e., q>0.05) were also selected for further validation.
Figure 4.
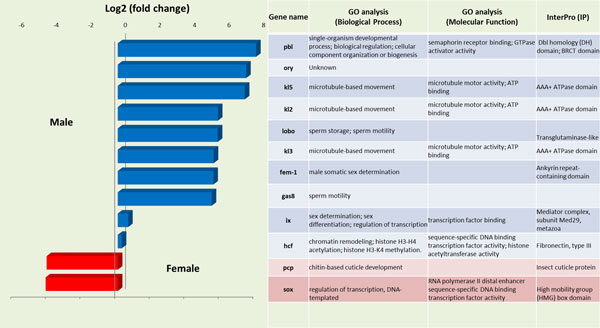
Functional annotation of differentially expressed sex-differentiation genes. In the left part of the figure, the gene expression levels of the differentially expressed sex-differentiation genes (Log2, fold change) are shown, as resulted from the RNA-seq analysis. The up-regulated genes in males are depicted in blue bars and the up-regulated genes in females in red bars. At the right part of the figure, the Gene Ontology (GO) classification of the same genes for the ontologies: Biological Process (BP), Molecular Function (MF), and Interpro (IP) protein domains is listed.
3.1. Drosophila Y-linked genes kl3, kl5 and ory
Quantitative RT-PCR confirmed the elevated expression of kl2, kl3, kl5 and ory in male testes of the olive fly (Figure 5). In Drosophila melanogaster, kl3 and kl5 (along with kl2) are known Y-linked fertility factors. The lack of kl3 or kl5 causes the loss of the outer arm of the sperm tail axoneme [59], a structure known to contain the molecular motor protein dynein in other organisms [60]. Indeed, Goldstein et al. showed in 1982 that sperm from mutant kl3- and kl5- males lack three discrete high molecular weight proteins with mobility similar to dynein heavy chains of Chlamydomonas reinhardtii and proposed that these fertility factors are the structural genes of three different dynein heavy chain proteins [61]. In 1993, Gepner and Hays sequenced part of kl5 and showed that it encodes an axonemal β-dynein heavy chain that is expressed in the testes [62].
Figure 5.
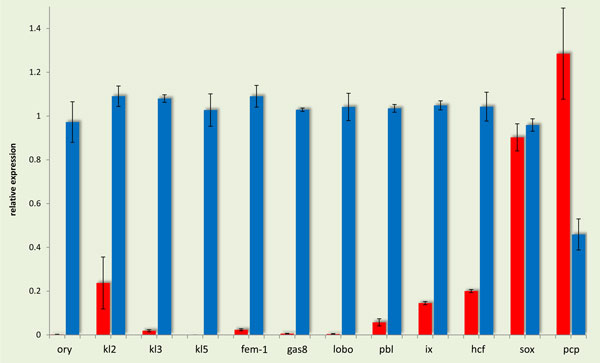
Validation profiles of differentially expressed sex-differentiation genes. Differentially expressed sex-differentiation genes of Figure 4 were further validated by qRT-PCR. Expression in male testes is depicted in blue color columns and expression in female accessory glands and spermathecae in red. Standard error of the mean of the two biological replicates is shown in bars. In all genes, except sox and pcp, expression in FEMALE and MALE was significantly different, as determined by t-test (p < 0.05).
ory is also Y-linked in D. melanogaster, although details on this gene are scarce. kl3, kl5 and ory are Y-linked in 12 different sequenced Drosophila genomes [63]. In Drosophila, the closest paralogs of kl2, kl3, and kl5 are autosomal and not X-linked, suggesting that the evolution of the Drosophila Y chromosome has been driven by an accumulation of male-related genes arising de novo from the autosomes [64]. While the most likely function of the three genes in the olive fly might be similar to that of Drosophila, we have no indication with regard to their chromosomal localization in the olive fly. Such information could shed some light to the evolutionary origin of the olive fly's Y chromosome.
3.2. Spermatogenesis and sperm motility genes
One spermatogenesis and two sperm motility genes were shown to be differentially over-expressed in male olive fly tissues both in the transcriptome analysis and after q-RT PCR (Figure 4 and 5). The first locus, sex-determining protein fem-1 (fem-1), encodes an essential spermatogenesis product in Caenorhabditis elegans. Three fem genes, fem-1, fem-2, and fem-3, have been shown to be essential for male development [65]. Loss-of-function mutations in any one of the fem genes prevent all aspects of male development and transform the animals that are genetically males into females [66,67]. The predicted product of the fem-1 gene is an intracellular protein that contains ankyrin repeats, which in many other proteins mediate specific protein-protein interaction [67]. In D. melanogaster, a fem-1 homolog with similar structure has been found [68]. The second locus, growth arrest-specific protein 8 (Gas8) is a microtubule-binding protein localized to regions of dynein regulation in mammalian cells. In mouse, Gas8 is predominantly a testicular protein, whose expression is developmentally regulated during puberty and spermatogenesis. In humans, it is absent in infertile males who lack the ability to generate gametes [69]. Gas8 has not been studied in insects. Finally, lost boys (lobo), has been shown to affect sperm entry movement into the female seminal receptacle and does not affect sperm exit movement from the seminal vesicle of D. melanogaster [70]. Given a similar function of these two loci in the olive fly, over-expression in male testes is expected.
3.3. Sex determination genes
In D. melanogaster, intersex (ix) controls somatic sexual differentiation only in females, acting near the end of the sex determination hierarchy. Its product does not have a known DNA-binding domain and, therefore, it is thought to act as a transcriptional co-factor for the female variant of Doublesex protein (DSXF), a key gene of the sexual determination cascade in D. melanogaster [71]. Minimal differences were observed in ix expression between the two sexes of the olive flies.
Transcriptome analysis also showed a four-fold over-expression of sox in female tissues, a result that was not confirmed after validation. The sox gene family is a group of related transcription factors that play critical roles in embryonic development. This family was originally identified in mammals based on sequence similarity to SRY, the sex-determining region Y chromosome [72]. In the honeybee, as SOX proteins play key roles in gonad differentiation, the SoxE group orthologues were up-regulated in the drone testes [73]. In Drosophila SoxN is a new group B Sox gene expressed in the developing CNS and is one of the earliest transcription factors to be expressed in a pan-neuroectodermal manner [74].
3.4. Other genes
The Pebble (pbl) gene belongs to a family of GTP exchange factors that are essential for the construction of a contractile ring and the initiation of cytokinesis during the embryonic division cycles of the somatic cells in D. melanogaster [75,76]. Its role in spermatogenesis has not been elucidated yet. Expression of pbl in D. melanogaster testes is low [68]. On the other hand, expression in olive fly testes was found elevated in comparison to its expression in female accessory glands/spermathecae (Figure 4 and 5).
Host cell factor C1 (Hcf) is involved in a wide variety of cellular functions, including regulation of transcription, cytokinesis, cell cycle progression and chromatin remodeling [77]. The protein is essential for cellular viability and demonstrates similar activity among a broad range of species. A single hcf homolog is also present in Drosophila (called dHCF) and is expressed in all tissues, although at relatively low levels [68]. The transcriptome analysis in the olive fly tissues showed a ~0,2-fold higher expression in the male tissues. This result was confirmed after qRT-PCR in the same tissues, where higher levels of expression in testes were observed in comparison with female accessory glands/spermathecae (Figure 4 and 5).
Quantitation by RT-PCR confirmed the over-expression of pupal cuticle protein (pcp) in female accessory glands/spermathecae as compared to male testes. Cuticle proteins, along with chitin, are the two components of insect cuticle. The cuticular proteins seem to be specific to the type of cuticle that occurs at stages of the insect development. Flexible proteins are found in the flexible cuticle of larva and pupa, but can also be found in the soft endocuticle of adult insects [78].
Female insects require the steroid hormone 20-hydroxyecdysone (20E) in order to activate vitellogenesis, a process required for egg development. In Anopheles gambiae mosquitoes, large amounts of 20E are produced and stored in male accessory glands and subsequently delivered to female mosquitoes during mating [79]. Pupal cuticle proteins, on the other hand, are known to accumulate in response to a pulse of 20E [80]. However, given that FAGs/spermathecae collected were from unmated females, we cannot offer a plausible explanation for the over-expression of pcps.
4. Validation of olfactory gene differential expression
Insects possess very sensitive chemosensory systems that can detect and discriminate among a diverse array of odors. These systems play a crucial role in insect survival and reproductive success, mediating responses to food detection, mating and oviposition. Odor recognition is a coordinated process requiring the combined specificities contributed by odorant-binding proteins (OBPs) and chemosensory proteins (CSPs) as well as odorant receptors (ORs) (Reviewed in [81]). Insect odorant-binding proteins (OBPs) are soluble proteins surrounding the extracellular lymph of olfactory neurons [82]. OBPs are capable of binding and solubilizing small hydrophobic molecules from the environment and therefore transport them to the underlying ORs, which are expressed on peripheral olfactory receptor neurons. Insect ORs are either ionotropic receptors (IRs) or seven-transmembrane proteins (ORs) with an inverse topology compared to GPCRs, that form heterodimers of a ligand-binding OR and an ubiquitous highly conserved co-receptor named Orco [83]. These complexes are suggested to constitute ligand-gated nonselective cation channels triggering the olfactory signaling [81].
While OR expression in olfactory tissues is obvious and well-established, the distribution of ORs beyond the olfactory system has also been documented in different mammalian species [84-86], suggesting that ORs may play an important role in the ectopic expression of non-chemosensory tissues. Interestingly, OR expression has been documented in human and mouse germ cells [87-91] and recently in mosquitoes [92]. Similarly, other non-olfactory functions have been reported for OBP-like proteins including the B proteins of Tenebrio molitor accessory glands [93], the male specific serum proteins of Ceratitis capitata [94], and the heme-binding protein of Rhodnius prolixus [95]. These demonstrate that OBPs are not restricted to olfaction and are likely to be involved in broader physiological functions, suggesting that their roles may be restricted to general carrier capabilities with broad specificity for lipophilic compounds [96].
With that in mind, we opted to explore the expression of various olfactory-related genes in the reproductive systems under investigation. Twelve olfactory-related genes were present in the annotated list that resulted from the transcriptome assembly of the FEMALE and MALE olive fly tissues (Figure 6), nine of which presented various levels of over-expression in MALE, whereas the remaining three in FEMALE. In order to get a deeper insight, the relative expression of five of these genes was further analyzed in female FAGs/spermathecae, male testes and male accessory glands (MAGs), before and after mating.
Figure 6.
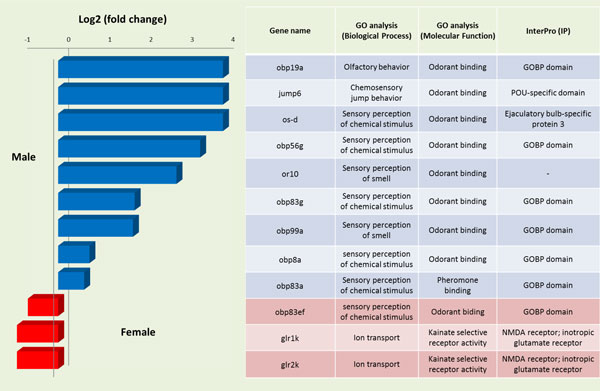
Functional annotation of differentially expressed olfactory genes. At the left part of the figure, the expression levels of the differentially expressed olfactory genes (Log2, fold change) are shown, as resulted from the RNA-seq analysis. The up-regulated genes in males are depicted in blue bars and the up-regulated genes in females in red bars. At the right part of the figure, the Gene Ontology (GO) classification of the same genes for the ontologies: Biological Process (BP), Molecular Function (MF) and Interpro (IP) protein domains is listed. Gene names are based on the nomenclature of the Drosophila melanogaster homologues [68].
obp83a, obp8a and obp19a genes are over-expressed in MALE tissue (Figure 6). qRT-PCR revealed that these genes share the same expression pattern in MAGs. obp83a and obp8a are over-expressed before mating in testes while obp83a and obp19a are over-expressed after mating in FAGs/spermathecae (Figure 7). All three genes are characterized by a GOBP (general odorant binding protein) domain that is also found in their orthologues in Drosophila melanogaster. This structural domain is found in pheromone binding proteins, which exist in extracellular fluid surrounding odorant receptors [97]. The presence of these OBPs in the reproductive tissues implicates their interaction with other substrates except the olfactory system as transporters in the post-mating events in the male reproductive system. In fact, D. melanogaster's obp8a shows the highest levels of expression in male accessory glands [98,99] and has been associated with non-olfactory functions such as RNA transcription [100].
Figure 7.
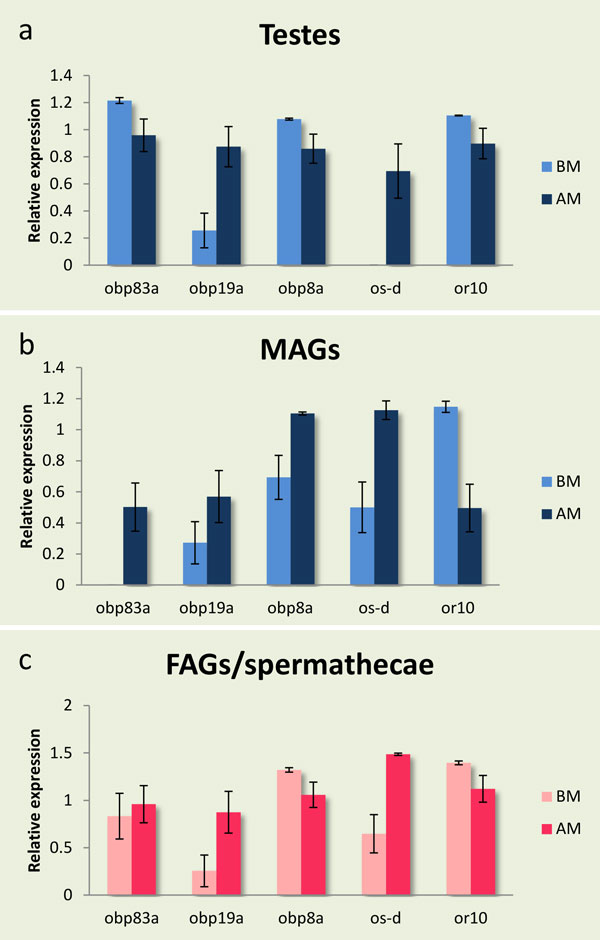
Relative expression profiles of differentially expressed olfactory genes. Expression profiles of five olfactory genes [odorant binding proteins obp83a, obp19a, obp8a, chemosensory protein, os-d, and odorant receptor 10, or10] as determined by qRT-PCR in three different tissues: Testes (a), MAGs (b) and FAGs/spermatheca (c) before (BM) and after (AM) mating. Standard error of the mean of five biological replicates is depicted in bars. No significant difference (for P < 0.05) was detected.
os-d is over-expressed in MALE tissue (Figure 6) while qRT-PCR showed similar expression patterns in mature FAGs/spermathecae, MAGs and testes, but no expression in MAGs before mating (Figure 7). Os-D is a chemosensory protein (CSP) that encodes the antennal protein 10 in D. melanogaster. CSPs are secreted in the sensillum lymph of insect chemosensory sensilla and some OS-D-like proteins bind short to medium chain length fatty acid derivatives with low specificity [101,102]. Their specific function remains uncertain [103], suggesting a more general physiological function relating to the transport/solubility of hydrophobic ligands in various tissues.
or10 showed expression in male tissues (Figure 6) while qRT-PCR detected same transcriptional profiles in all three tissues before and after mating (Figure 7). or10 encodes an olfactory receptor protein and has a G-protein coupled receptor activity. The expression of ORs in testes has been reported for a number of species [90,104]. ORs' function in mammalian sperm is thought to regulate motility in response to exogenous signals derived from the existence of sperm-egg chemotaxis in invertebrates. The small peptides, speract and resact, are secreted by sea urchin eggs and attract spermatozoa in a species-specific manner by stimulating sperm motility and respiration [105,106]. The presence of a similar chemoreceptor may be essential in female spermatheca in order to establish a concentration gradient of a putative chemo-attractant. Since female accessory glands and spermatheca were dissected together, we are not able at this point to establish which exact tissue is the source of the observed expression of or10.
5. Early embryonic gene expression in the olive fly
As mentioned in the Background, promoters of early embryonic genes in combination with pro-apoptotic cell death genes are very important tools in inducing dominant early-embryonic lethality during insect transgenesis [107]. In that regard, the serendipity-α (sry-α) and head involution defective (hid) genes were selected for expression evaluation during embryonic development in the olive fly.
The embryonic developmental progress begins with the egg maturation and formation of the zygote, then enters the stage of blastoderm formation and gastrulation and ultimately ends with the organogenesis. Accordingly, three stages of embryogenesis have been also designated in B. oleae, whose average duration is 65-70h at 25 ± 1°C under standard laboratory conditions [108]. Microscopy morphological observations in living embryos report that cellularization of the blastoderm begins 6h after oviposition and lasts until 10h. During the third stage of organogenesis, the ventral furrow formation starts by 22h and the head and abdominal lobe masses become visible by 46h. Gut and mouth hook formation can be identified by 52h, whereas the development of other systems are distinct by 60h.
In Drosophila melanogaster, sry-α gene is specifically transcribed at the blastoderm stage in all somatic nuclei, from nuclear cycle 11 to the onset of gastrulation [109]. The gene product is required for the complete reorganization of the microfilaments at the onset of membrane invagination [110]. sry-α is fast evolving even within the Drosophilidae [111] and extensive divergence of many developmental genes within dipterans has also been reported [112-114]. This was most likely the reason for the unsuccessful efforts in C. capitata to obtain sry-α by degenerate PCR on the basis of sequence similarity with the homologous D. melanogaster [115]. Given the availability of both D. melanogaster and C. capitata sry-α sequences in the NCBI database, a homology search in the B. oleae transcriptome identified the relevant B. oleae sry-α gene homologue.
Based on this sequence, B. oleae-specific primers were designed and the expression profiles of sry-α mRNA were studied by qRT-PCR analysis at different stages of B. oleae embryonic development. Eggs were collected throughout embryogenesis from the time of egg laying to larval hatching. The selected time points represented embryos at 0h, 4h, 8h, 9h, 10h, 11h, 12h, 15h and 18h after oviposition (Figure 8, panel A). This analysis revealed that sry-α mRNA is developmentally regulated during the second major event in the first stage of embryogenesis. It is initially present in large amounts just after oviposition (0h embryos), following a reduction in 4h embryos. The larger amounts of the transcripts among all time points examined were detected in 8h embryos. This suggests the presence of maternal mature transcripts which in turn are eliminated probably in the first event of maternal-to-zygotic transition (MZT). The subsequent wave of 'zygotic' activity requires zygotically synthesized transcripts [116]. In D. melanogaster as well as in C. capitata, sry-α is expressed only in the zygote [117]. However the retrieved B. oleae transcript shared greater amino acid similarity to the D. melanogaster CG8247 gene than to sry-α, as was also reported for the Ccsry-α like gene [118]. The orthologous CG8247 in D. melanogaster is characterized as a sry-α-like gene being also involved in cellular blastoderm formation. However, it is maternally inherited in contrast to sry-α, demonstrating a different mechanism of molecular control of transcription. In our case Bosry-α like gene seems to be maternally supplied in the embryos as mature transcripts. Previous studies have designated that the cellular blastoderm formation in C. capitata occurs within 9 h and 11 h after oviposition [115]. In accordance with C. capitata, a relative Tephritid species, we suggest that the cellurarization process in B. oleae during embryogenesis also occurs at 8h, since the sry-α transcripts were detected at higher levels during this time.
Figure 8.
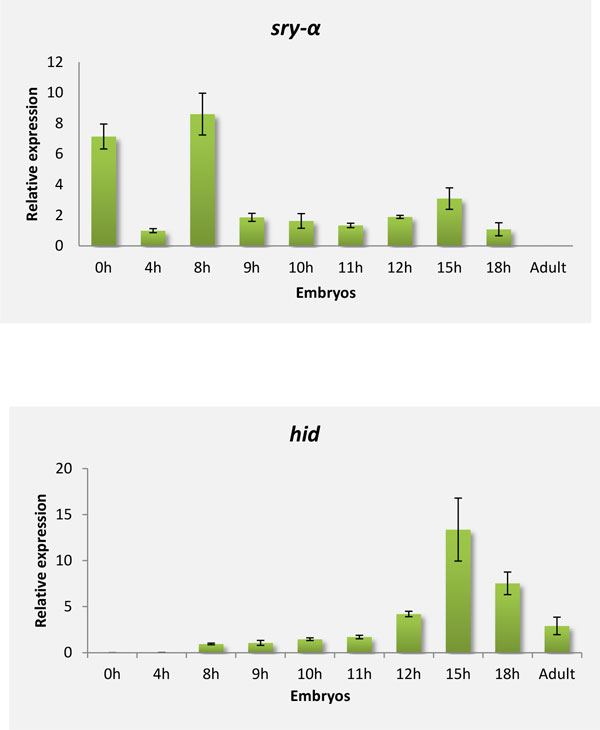
Expression profile analysis during the early stages of embryogenesis. Expression levels of Α) Bosry-α and B) Bohid in individual eggs collected at different time points during embryonic development, as determined by qRT-PCR. Standard error of the mean of two biological replicates per time point is depicted in bars.
6. Apoptotic gene expression
At the same time, head involution defective (hid), known to have a central role in apoptosis pathway, was also selected for further study. Apoptosis is a genetically controlled mechanism of cytological events that results in programmed cell death. During development, programmed cell death plays a key role by eliminating unwanted cells from a variety of tissues, such as, for example, larval tissues during insect metamorphosis (Reviewed in [119]). A series of caspases, a family of cysteine proteases, play a central role during apoptosis. Once activated, caspases can cleave more than 100 different cell target proteins, bringing about ultimately the cell death [120]. Regulators of caspase activation may either promote apoptosis (pro-apoptotic) or inhibit apoptosis (anti-apoptotic). Drosophila Hid belongs to a family of pro-apoptotic proteins which act as antagonists of IAPs (Inhibitor of Apoptosis Proteins), thus resulting in caspase activation and apoptosis [119,121,122]. Such pro-apoptotic genes have been used in transgenic control systems for pest insects. In tetracycline-suppressible systems for female-specific lethality and conditional embryonic expression of a Drosophila hid-containing transgene, for example, 100% lethality was observed in Drosophila [123], as well as in the Tephritid flies Ceratitis capitata [117] and Anastrepha suspensa [124].
The developmental regulation of Bohid was explored by determining the transcript levels during embryogenesis. A qRT-PCR approach with species-specific primers was used to evaluate the expression pattern of hid in embryos at 0 h, 4 h, 8 h, 9 h, 10 h, 11 h, 12 h, 15 h, 18 h after oviposition. Based on D. melanogaster hid expression pattern, no expression was expected in embryos prior to formation of the syncytial blastoderm [125]. Indeed, until 8h no transcripts were detected. hid expression was first detected at 12h and peaked during 15h (Figure 8, panel B).
It is noteworthy that most developmental programmed cell death occurs during the gastrulation process of D. melanogaster embryonic development [126], suggesting that the onset of this period in B. oleae could be defined approximately at 12h, occurring mainly within 15-18h.
However, further examination of the pro-apoptotic function of hid gene is required in order to explore its ability of inducing apoptosis in B. oleae cells. Specific lethal embryonic phenotypes need to be obtained to characterize its role in the cell-death pathway. Ongoing analysis for the isolation of the complete gene will provide the essential tools for the generation of an endogenous effective lethal effector system.
Conclusions
In serious agricultural pests (like the olive fly) which are not model experimental organisms (unlike the medfly), the major focus of most scientific research is, in the end, directed towards control of the pest. Old and new environmental concerns and sensibilities, that regard mostly insecticide use, drive science to the quest of alternative, environmentally friendlier methods of pest control. Time and again it has been shown that such methods go through thorough understanding of the biology and ecology of the target organism. Since the initial unsuccessful SIT efforts, molecular and genetic studies in the olive fly have focused on genetic analyses of natural populations, cytogenetics, isolation and characterization of genes that control important biological processes, as well as the identification and mapping of several microsatellite loci. Just a few years ago, B. oleae was successfully transformed, an achievement that gave new perspective towards the efficient use of the SIT. Lately, this is being coupled with genomics studies and transcriptomics analyses of various important systems, as well as efforts in advancing olive fly mass-rearing, that are setting the ground for the application of modern control approaches through the genetic manipulation of the insect.
Methods
Ethics statement
The study was carried out on laboratory reared olive flies. No specific permissions are required for these experiments, since these studies did not involve endangered or protected species.
Fly culture and stocks
Laboratory strain
The laboratory strain of the olive fly (LAB) is part from the original stock from the Department of Biology, 'Demokritos' Nuclear Research Centre, Athens, Greece, and has been reared in our laboratory for over 15 years. The flies are reared at 25°C with a 12h light/12h dark photoperiod in 30x30x30cm3 cages, as described by [127-129].
Egg collection
For embryo analysis, eggs were collected from 10-day old mated females maintained in our laboratory, which were fed with artificial adult diet to ensure high oviposition rates and embryo viability. Adults were exposed to paraffin oviposition domes for 10 minutes and the eggs were obtained with a 0.3% propionic acid solution, assigning this as the start time point. Eggs were maintained in an incubator according to the standard rearing conditions.
RNA isolation for library preparation and functional analysis
Total RNA was isolated from female accessory glands (FAGs) and spermathecae of ~300 female flies and from testes of ~150 male flies. Four-day old sexually immature unmated insects were used. For RNA isolation, the TRIzol® Reagent (Ambion-Invitrogen) was used, following the instructions of the manufacturer with minor modifications. RNA extraction was followed by an additional DNA removal using the TURBO DNA-free Kit (Ambion-Invitrogen), according to manufacturer's instructions. The integrity of RNA was assessed by 1% agarose gel electrophoresis and the purity of all RNA samples was evaluated at Fleming Institute (Greece) with the use of (Agilent 2100 Bioanalyzer) and NanoDrop (2000).
Whole transcriptome library preparation for next-generation sequencing with the SOLiD 4 Sequencing System
RNA transcripts from olive fly FAGs/spermathecae (FEMALE) and testes (MALE) were used to construct two cDNA libraries for sequencing analysis on the SOLiD 4 Sequencing System. More specifically, polyadenylated RNA (polyA-RNA) was isolated from 5 μg of total RNA using the Dynabeads Oligo(dT) kit (Ambion, Life Technologies Corporation). The isolated polyA-RNA was randomly fragmented by chemical hydrolysis at 94°C for 5 minutes and was then treated with antarctic phosphatase to remove phosphate groups from the fragments' ends, followed by treatment with T4 polynucleotide kinase to add a Pi at the 5' end of each fragment. The resulting RNA fragments were hybridized and ligated to the P1 and P2 adaptor sequences specifically designed for sequencing with the SOLiD system (SOLiD Total RNA-Seq Kit, Life Technologies Corporation). The RNA produced was reverse transcribed to cDNA which was then amplified in a 15-cycle PCR. At this step, the use of different barcoded 3' PCR primers from the selection included in the SOLiD barcoding kit allowed the preparation of cDNA libraries for multiplex sequencing. From the cDNA produced, only fragments of average size 200-300 bp were selected with two rounds of magnetic bead purification (Agencourt AMPure XP Reagent, Beckman Coulter).
The quality and size of the purified cDNA library was assessed on the Agilent Bioanalyzer 2100 (Agilent Technologies Inc.) and with quantitative PCR using the Library Quant Kit ABI Solid (KAPA Biosystems). A multiplex library mix (500pM) was used to prepare a full-slide for analysis on the SOliD 4 Sequencing System (Applied Biosystems) with 35+50 bp PE-chemistry.
RNA isolation and expression analysis of selected genes
RNA extraction for expression analysis of sexually differentially expressed genes. For the validation of the differential expression of sexually differentially expressed genes, RNA was extracted from two pools of 40 pairs of spermathecae/FAGs and 40 pairs of testes (two biological pool replicates), dissected from an equivalent number of female and male adult laboratory flies, respectively.
RNA extraction for expression analysis of olfactory and early embryonic developmental genes. For the validation of the olfactory genes expression, RNA was extracted from five female and five male individual insects (five biological replicates, respectively) before and after mating of the aforementioned laboratory strain. Two groups of insects were considered. Firstly, unmated insects, i.e., sexually mature 7-day old unmated insects (before mating, BM). Secondly, mated insects, i.e., sexually mature 7-day old insects that were allowed to mate on the seventh day and were dissected 12 hours after mating (after mating, AM). For the validation of the sexually differentially expressed genes, the RNA isolated for the construction of the two libraries was used. RNA was extracted using TriZol reagent according to manufacturer's protocol.
For the validation of the early embryonic genes, eggs were removed from the incubator at different time intervals throughout embryonic development and total RNA was extracted from each egg using TriZol reagent according to the manufacturer's protocol. Two individual eggs (two biological replicates) from the various time points during the embryonic developmental stages were used for the extractions.
Following extraction, the RNA was treated with 1.0 unit of DNase I (Invitrogen) according to manufacturer's instructions. In all of the above cases, the total amount of DNA-free RNA obtained from each tissue (between 400 to 700 ng) was converted into cDNA using 300ng Random hexamer primers (equimolar mix of N5A, N5G, N5C and N5T), 200 units MMLV Reverse Transcriptase (Geneon), 5X reaction buffer, 40mM dNTP mix and 40 units RNase Inhibitor (GeneOn) according to the manufacturer's instructions. Reverse transcription was conducted at 42°C for 50 min and 70°C for 15 min. The resulting cDNA was used in the subsequent qPCR reactions.
Specific primers for the amplification of selected differentially expressed genes revealed by the transcriptome analysis were designed by Primer-BLAST (http://www.ncbi.nlm.nih.gov/tools/primer-blast) (Table S2). To identify sequences with homology to the genes sry-α and hid, the orthologous genes of C. capitata and An. suspensa were used as queries to search for B. oleae transcripts using tBLASTX in the TSA Database. Species-specific Blast hits for each of the query sequences were retrieved (Genbank: GAKB01005111.1, GAKB01003654.1) and used to design primers (Table S2) for the subsequent amplification of gene-specific sequences by quantitative real-time PCR (qRT-PCR).
Relative quantitation was used to analyze changes in expression levels of the selected genes using a Real-time PCR approach. Expression values were calculated relatively to the housekeeping rpl19 gene. Rpl19 and 14-3-3z genes were used as reference in MAGs and testes while actin3 and a-tubulin in FAGs/spermathecae. The qRT-PCR conditions were: polymerase activation and DNA denaturation step at 95 °C for 4 min, followed by 40 cycles of denaturation at 95 °C for 30 s, annealing/extension and plate read at 56 °C for 30 s and finally, a step of melting curve analysis at a gradual increase of temperature over the range 55 °C → 95 °C. In this step, the detection of one gene specific peak and the absence of primer dimer peaks was assured. Each reaction was performed in a total volume of 15 μl, containing 5 μl from a dilution 1:10 of the cDNA template, 1X iTaq Universal SYBR Green Supermix (Biorad, Gaithesburg, MD) and 400nM of each primer. The reactions were carried out on Bio-Rad Real-Time thermal cycler CFX96 (Bio-Rad, Hercules, CA, USA) and data analysed using the CFX Manager™ software. All qRT-PCRs were performed in triplicate (i.e., three technical replicates).
Bioinformatics analysis
All paired and unpaired reads of the libraries were assembled to construct the reference transcriptome using the SOAPdenovo assembler [54] with a word size of 25 nt. Annotation of the assembled sequences was obtained by comparing to the NCBI non-redundant (Nr) protein database (May 7th, 2014 version) using blastx [130] and collecting the annotations with the BLAST2GO tool [55]. TopHat [131] was used to generate a spliced alignment to the reference transcriptome. Transcripts were assembled using Cufflinks and differentially expressed genes were identified using Cuffdiff [56]. GO-term enrichment between male and female transcriptomes was analyzed using the using the GOSSIP [132] application embedded in BLAST2GO.
Availability of supporting data
The data sets supporting the results of this article are included within the article and its additional files. Additional File 1, Additional File 2, Additional File 3 and Additional File 4
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
ES was involved in the transcriptome library construction and performed the analysis of the sex-determination genes; MR performed the bioinformatics analysis of the transcriptome; VH constructed the transcriptome libraries and analysed the sequencing data; KTT and AMM analyzed the embryonic and apoptotic genes; MEG, ST and KA analysed the olfactory genes; JR directed the bioinformatics analysis; KDM designed and coordinated the study. All authors participated in drafting the manuscript and read and approved the final document.
Supplementary Material
Contributor Information
Efthimia Sagri, Email: esagr@bio.uth.gr.
Martin Reczko, Email: reczko@fleming.gr.
Konstantina T Tsoumani, Email: kotsouma@bio.uth.gr.
Maria-Eleni Gregoriou, Email: magrigoriou@bio.uth.gr.
Vaggelis Harokopos, Email: harokopos@fleming.gr.
Anna-Maria Mavridou, Email: anmavrid@med.uth.gr.
Spyros Tastsoglou, Email: tastsogl@bio.uth.gr.
Konstantinos Athanasiadis, Email: konathas@bio.uth.gr.
Jiannis Ragoussis, Email: ioannis.ragoussis@mcgill.ca.
Kostas D Mathiopoulos, Email: kmathiop@bio.uth.gr.
Acknowledgements
This research has been co-financed by: the Actions Heracleitus II and "ARISTEIA" (“OLFLY SMELL & SEX”) of the "Operational programme Education and Life Long Learning", co-funded by the European Social Fund and Greek National Resources”; and the two postgraduate programs of the Department of Biochemistry and Biotechnology of the University of Thessaly ("Biotechnology - Nutrition and Environment" and "Molecular Biology and Genetics applications"). Special acknowledgements should also go the Joint FAO/IAEA Division of Nuclear Techniques in Food and Agriculture for their support in the organization of a Coordinated Research Project on "Development and evaluation of imported strains of insect pests for SIT.
This article has been published as part of BMC Genetics Volume 15 Supplement 2, 2014: Development and evaluation of improved strains of insect pests for SIT. The full contents of the supplement are available online at http://www.biomedcentral.com/bmcgenet/supplements/15/S2.
Publication of this supplement was funded by the International Atomic Energy Agency. The peer review process for articles published in this supplement was overseen by the Supplement Editors in accordance with BioMed Central's peer review guidelines for supplements. The Supplement Editors declare that they have no competing interests.
References
- Theophrastus. Enquiry into plants (History of plants HP), I & II (HORT, A. F., translator) London, Cambridge & Massachusetts; 1916. [in ancient Greek with English translation] [Google Scholar]
- Daane KM, Johnson MW. Olive fruit fly: managing an ancient pest in modern times. Annu Rev Entomol. 2010;55:151–69. doi: 10.1146/annurev.ento.54.110807.090553. [DOI] [PubMed] [Google Scholar]
- Pimentel D. Ecological Effects of Pesticides on Non-target Species. Washington, D.C.: Executive Office of the President, Office of Science and Technology; 1971. p. 220. [Google Scholar]
- Baumhover A, Graham A, Bitter B, Hopkins D, New W, Dudleyandr F, Bushland C. Screwworm control through release of sterilized flies. J Econ Entomol. 1955. pp. 462–466.
- Knipling E. Possibilities of insect control or eradication through the use of sexually sterile males. J Econ Entomol. 1955. pp. 459–462.
- Greek Ministry of Agriculture. Description of research organization for the control of the olive fruit fly. 1961. p. 33. (in Greek)
- Economopoulos A, Avtzis N, Zervas G, Tsitsipis J, Haniotakis G, Tsiropoulos G, Manoukas A. Control of the olive fly, Dacus oleae (Gmelin), by the combined effects of insecticides and release of gamma sterilized insects. J Appl Entomol. 1977. pp. 201–215.
- Economopoulos AP, Haniotakis GE, Mathioudis J, Missis N, Kinigakis P. Long-distance flight of wild and artificially-reared Dacus oleae (Gmelin) (Diptera, Tephritidae) Z Angew Entomol. 1978. pp. 101–108.
- Economopoulos A, Zervas G. The quality problem in olive flies produced for SIT experiments. IAEA STI/PUB. 1982.
- Economopoulos A. The olive fruit fly, Bactrocera (Dacus) oleae (Gmelin) (Diptera: Tephritidae): its importance and control; previous SIT research and pilot testing. Int At Energy Agency, Vienna, Austria. 2002.
- Zervas GA, Economopoulos AP. Mating frequency in caged populations of wild and artificially reared (normal or γ-sterilized) olive fruit flies. Environ Entomol. 1982. pp. 17–20.
- Loukas M, Economopoulos AP, Zouros E, Vergini Y. Genetic changes in artificially reared colonies of the olive fruit fly. Ann Ent Soc Amer. 1985. pp. 159–165.
- Economopoulos A, Loukas M. ADH allele frequency changes in olive fruit flies shift from olives to artificial larval food and vice versa, effect of temperature. Entomol Exp Appl. 1986. pp. 215–221.
- Economopoulos A. Sexual competitiveness of gamma-ray sterilized males of Dacus oleae. Mating frequency of artificially reared and wild females. Env Entomol. 1972. pp. 490–497.
- Capuzzo C, Firrao G, Mazzon L, Squartini A, Girolami V. "Candidatus Erwinia dacicola", a coevolved symbiotic bacterium of the olive fly Bactrocera oleae (Gmelin) Int J Syst Evol Microbiol. 2005;55(Pt 4):1641–7. doi: 10.1099/ijs.0.63653-0. [DOI] [PubMed] [Google Scholar]
- Sacchetti P, Granchietti A, Landini S, Viti C, Giovannetti L, Belcari A. Relationships between the olive fly and bacteria. J Appl Entomol. 2008;132:682–689. doi: 10.1111/j.1439-0418.2008.01334.x. [DOI] [Google Scholar]
- Estes AM, Hearn DJ, Bronstein JL, Pierson EA. The olive fly endosymbiont, "Candidatus Erwinia dacicola," switches from an intracellular existence to an extracellular existence during host insect development. Appl Environ Microbiol. 2009;75:7097–106. doi: 10.1128/AEM.00778-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Yosef M, Aharon Y, Jurkevitch E, Yuval B. Give us the tools and we will do the job: symbiotic bacteria affect olive fly fitness in a diet-dependent fashion. Proc Biol Sci. 2010;277:1545–52. doi: 10.1098/rspb.2009.2102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kounatidis I, Crotti E, Sapountzis P, Sacchi L, Rizzi A, Chouaia B, Bandi C, Alma A, Daffonchio D, Mavragani-Tsipidou P, Bourtzis K. Acetobacter tropicalis is a major symbiont of the olive fruit fly (Bactrocera oleae) Appl Environ Microbiol. 2009;75:3281–8. doi: 10.1128/AEM.02933-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Augustinos AA, Stratikopoulos EE, Zacharopoulou A, Mathiopoulos KD. Polymorphic microsatellite markers in the olive fly, Bactrocera oleae. Mol Ecol Notes. 2002;2:278–280. doi: 10.1046/j.1471-8286.2002.00222.x. [DOI] [Google Scholar]
- Nardi F, Carapelli A, Dallai R, Frati F. The mitochondrial genome of the olive fly Bactrocera oleae: two haplotypes from distant geographical locations. Insect Mol Biol. 2003;12:605–611. doi: 10.1046/j.1365-2583.2003.00445.x. [DOI] [PubMed] [Google Scholar]
- Augustinos AA, Mamuris Z, Stratikopoulos EE, D'Amelio S, Zacharopoulou A, Mathiopoulos KD. Microsatellite analysis of olive fly populations in the Mediterranean indicates a westward expansion of the species. Genetica. 2005;125:231–41. doi: 10.1007/s10709-005-8692-y. [DOI] [PubMed] [Google Scholar]
- Nardi F, Carapelli A, Dallai R, Roderick GK, Frati F. Population structure and colonization history of the olive fly, Bactrocera oleae (Diptera, Tephritidae) Mol Ecol. 2005;14:2729–38. doi: 10.1111/j.1365-294X.2005.02610.x. [DOI] [PubMed] [Google Scholar]
- Nardi F, Carapelli A, Boore JL, Roderick GK, Dallai R, Frati F. Domestication of olive fly through a multi-regional host shift to cultivated olives: comparative dating using complete mitochondrial genomes. Mol Phylogenet Evol. 2010;57:678–86. doi: 10.1016/j.ympev.2010.08.008. [DOI] [PubMed] [Google Scholar]
- Zygouridis NE, Augustinos AA, Zalom FG, Mathiopoulos KD. Analysis of olive fly invasion in California based on microsatellite markers. Heredity (Edinb) 2009;102:402–12. doi: 10.1038/hdy.2008.125. [DOI] [PubMed] [Google Scholar]
- Dogaç E, Kandemir İ, Taskin V. The genetic polymorphisms and colonization process of olive fly populations in Turkey. PLoS One. 2013;8:e56067. doi: 10.1371/journal.pone.0056067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mavragani-Tsipidou P. Genetic and cytogenetic analysis of the olive fruit fly Bactrocera oleae (Diptera: Tephritidae) Genetica. 2002;116:45–57. doi: 10.1023/A:1020907624816. [DOI] [PubMed] [Google Scholar]
- Mavragani-Tsipidou P, Karamanlidou G, Zacharopoulou A, Koliais S, Kastritisis C. Mitotic and polytene chromosome analysis in Dacus oleae (Diptera: Tephritidae) Genome. 1992;35:373–8. doi: 10.1139/g92-056. [DOI] [PubMed] [Google Scholar]
- Zambetaki A, Kleanthous K, Mavragani-Tsipidou P. Cytogenetic analysis of Malpighian tubule and salivary gland polytene chromosomes of Bactrocera oleae (Dacus oleae) (Diptera: Tephritidae) Genome. 1995;38:1070–81. doi: 10.1139/g95-143. [DOI] [PubMed] [Google Scholar]
- Drosopoulou E, Chrysopoulou A, Nikita V, Mavragani-Tsipidou P. The heat shock 70 genes of the olive pest Bactrocera oleae: genomic organization and molecular characterization of a transcription unit and its proximal promoter region. Genome. 2009;52:210–4. doi: 10.1139/G08-110. [DOI] [PubMed] [Google Scholar]
- Drosopoulou E, Nakou I, Síchová J, Kubíčková S, Marec F, Mavragani-Tsipidou P. Sex chromosomes and associated rDNA form a heterochromatic network in the polytene nuclei of Bactrocera oleae (Diptera: Tephritidae) Genetica. 2012;140:169–80. doi: 10.1007/s10709-012-9668-3. [DOI] [PubMed] [Google Scholar]
- Vontas JG, Hejazi MJ, Hawkes NJ, Cosmidis N, Loukas M, Hemingway J, Janes RW. Resistance-associated point mutations of organophosphate insensitive acetylcholinesterase, in the olive fruit fly Bactrocera oleae. Insect Mol Biol. 2002;11:329–336. doi: 10.1046/j.1365-2583.2002.00343.x. April. [DOI] [PubMed] [Google Scholar]
- Vontas J, Blass C, Koutsos AC, David JP, Kafatos FC, Louis C, Hemingway J, Christophides GK, Ranson H. Gene expression in insecticide resistant and susceptible Anopheles gambiae strains constitutively or after insecticide exposure. Insect Mol Biol. 2005;14:509–21. doi: 10.1111/j.1365-2583.2005.00582.x. [DOI] [PubMed] [Google Scholar]
- Kakani EG, Mathiopoulos KD. Organophosphosphate resistance-related mutations in the acetylcholinesterase gene of Tephritidae. J Appl Entomol. 2008;132:762–771. doi: 10.1111/j.1439-0418.2008.01373.x. [DOI] [Google Scholar]
- Kakani EG, Bon S, Massoulié J, Mathiopoulos KD. Altered GPI modification of insect AChE improves tolerance to organophosphate insecticides. Insect Biochem Mol Biol. 2011;41:150–8. doi: 10.1016/j.ibmb.2010.11.005. [DOI] [PubMed] [Google Scholar]
- Khila A, El Haidani A, Vincent A, Payre F, Souda SI. The dual function of ovo/shavenbaby in germline and epidermis differentiation is conserved between Drosophila melanogaster and the olive fruit fly Bactrocera oleae. Insect Biochem Mol Biol. 2003;33:691–9. doi: 10.1016/S0965-1748(03)00063-8. [DOI] [PubMed] [Google Scholar]
- Benos P, Tavernarakis N, Brogna S, Thireos G, Savakis C. Acquisition of a potential marker for insect transformation: isolation of a novel alcohol dehydrogenase gene from Bactrocera oleae by functional complementation in yeast. Mol Gen Genet. 2000;263:90–5. doi: 10.1007/PL00008679. [DOI] [PubMed] [Google Scholar]
- Lagos D, Ruiz MF, Sánchez L, Komitopoulou K. Isolation and characterization of the Bactrocera oleae genes orthologous to the sex determining Sex-lethal and doublesex genes of Drosophila melanogaster. Gene. 2005;348:111–21. doi: 10.1016/j.gene.2004.12.053. [DOI] [PubMed] [Google Scholar]
- Lagos D, Koukidou M, Savakis C, Komitopoulou K. The transformer gene in Bactrocera oleae: the genetic switch that determines its sex fate. Insect Mol Biol. 2007;16:221–30. doi: 10.1111/j.1365-2583.2006.00717.x. [DOI] [PubMed] [Google Scholar]
- Tsoumani KT, Mathiopoulos KD. Genome size estimation with quantitative real-time PCR in two Tephritidae species: Ceratitis capitata and Bactrocera oleae. J Appl Entomol. 2012;136:626–631. doi: 10.1111/j.1439-0418.2011.01684.x. [DOI] [Google Scholar]
- Tsoumani KT, Drosopoulou E, Mavragani-Tsipidou P, Mathiopoulos KD. Molecular characterization and chromosomal distribution of a species-specific transcribed centromeric satellite repeat from the olive fruit fly, Bactrocera oleae. PLoS One. 2013;8:e79393. doi: 10.1371/journal.pone.0079393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsoumani KT, Augustinos AA, Kakani EG, Drosopoulou E, Mavragani-Tsipidou P, Mathiopoulos KD. Isolation, annotation and applications of expressed sequence tags from the olive fly, Bactrocera oleae. Mol Genet Genomics. 2011;285:33–45. doi: 10.1007/s00438-010-0583-y. [DOI] [PubMed] [Google Scholar]
- Pavlidi N, Dermauw W, Rombauts S, Chrisargiris A, Van Leeuwen T, Vontas J. Analysis of the Olive Fruit Fly Bactrocera oleae Transcriptome and Phylogenetic Classification of the Major Detoxification Gene Families. PLoS One. 2013;8:e66533. doi: 10.1371/journal.pone.0066533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koukidou M, Klinakis A, Reboulakis C, Zagoraiou L, Tavernarakis N, Livadaras I, Economopoulos A, Savakis C. Germ line transformation of the olive fly Bactrocera oleae using a versatile transgenesis marker. Insect Mol Biol. 2006;15:95–103. doi: 10.1111/j.1365-2583.2006.00613.x. [DOI] [PubMed] [Google Scholar]
- Ant T, Koukidou M, Rempoulakis P, Gong HF, Economopoulos A, Vontas J, Alphey L. Control of the olive fruit fly using genetics-enhanced sterile insect technique. BMC Biol. 2012;10:51. doi: 10.1186/1741-7007-10-51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apostolaki A, Livadaras I, Saridaki A, Chrysargyris A, Savakis C, Bourtzis K. Transinfection of the olive fruit fly Bactrocera oleae with Wolbachia: towards a symbiont-based population control strategy. J Appl Entomol. 2011;135:546–553. doi: 10.1111/j.1439-0418.2011.01614.x. [DOI] [Google Scholar]
- Estes AM, Hearn DJ, Burrack HJ, Rempoulakis P, Pierson EA. Prevalence of Candidatus Erwinia dacicola in wild and laboratory olive fruit fly populations and across developmental stages. Environ Entomol. 2012;41:265–74. doi: 10.1603/EN11245. [DOI] [PubMed] [Google Scholar]
- Alphey L, Andreasen M. Dominant lethality and insect population control. Mol Biochem Parasitol. 2002;121:173–8. doi: 10.1016/S0166-6851(02)00040-3. [DOI] [PubMed] [Google Scholar]
- Alphey L, Beard C Ben, Billingsley P, Coetzee M, Crisanti A, Curtis C, Eggleston P, Godfray C, Hemingway J, Jacobs-Lorena M, James AA, Kafatos FC, Mukwaya LG, Paton M, Powell JR, Schneider W, Scott TW, Sina B, Sinden R, Sinkins S, Spielman A, Touré Y, Collins FH. Malaria control with genetically manipulated insect vectors. Science. 2002;298:119–21. doi: 10.1126/science.1078278. [DOI] [PubMed] [Google Scholar]
- Heinrich JC, Scott MJ. A repressible female-specific lethal genetic system for making transgenic insect strains suitable for a sterile-release program. Proc Natl Acad Sci USA. 2000;97:8229–32. doi: 10.1073/pnas.140142697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas DD, Donnelly CA, Wood RJ, Alphey LS. Insect population control using a dominant, repressible, lethal genetic system. Science. 2000;287:2474–6. doi: 10.1126/science.287.5462.2474. [DOI] [PubMed] [Google Scholar]
- Gong P, Epton MJ, Fu G, Scaife S, Hiscox A, Condon KC, Condon GC, Morrison NI, Kelly DW, Dafa'alla T, Coleman PG, Alphey L. A dominant lethal genetic system for autocidal control of the Mediterranean fruitfly. Nat Biotechnol. 2005;23:453–6. doi: 10.1038/nbt1071. [DOI] [PubMed] [Google Scholar]
- Sagri E, Reczko M, Gregoriou M-E, Tsoumani KT, Zygouridis NE, Salpea KD, Zalom FG, Ragoussis J, Mathiopoulos KD. Olive fly transcriptomics analysis implicates energy metabolism genes in spinosad resistance. BMC Genomics. 2014;15:714. doi: 10.1186/1471-2164-15-714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li R, Zhu H, Ruan J, Qian W, Fang X, Shi Z, Li Y, Li S, Shan G, Kristiansen K, Li S, Yang H, Wang J, Wang J. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 2010;20:265–72. doi: 10.1101/gr.097261.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008;36:3420–35. doi: 10.1093/nar/gkn176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–5. doi: 10.1038/nbt.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Domanitskaya E V, Liu H, Chen S, Kubli E. The hydroxyproline motif of male sex peptide elicits the innate immune response in Drosophila females. FEBS J. 2007;274:5659–68. doi: 10.1111/j.1742-4658.2007.06088.x. [DOI] [PubMed] [Google Scholar]
- McGraw LA, Clark AG, Wolfner MF. Post-mating gene expression profiles of female Drosophila melanogaster in response to time and to four male accessory gland proteins. Genetics. 2008;179:1395–408. doi: 10.1534/genetics.108.086934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hardy RW, Tokuyasu KT, Lindsley DL. Analysis of spermatogenesis in Drosophila melanogaster bearing deletions for Y-chromosome fertility genes. Chromosoma. 1981;83:593–617. doi: 10.1007/BF00328522. [DOI] [PubMed] [Google Scholar]
- Gibbons IR. Dynein family of motor proteins: present status and future questions. Cell Motil Cytoskeleton. 1995;32:136–44. doi: 10.1002/cm.970320214. [DOI] [PubMed] [Google Scholar]
- Goldstein LS, Hardy RW, Lindsley DL. Structural genes on the Y chromosome of Drosophila melanogaster. Proc Natl Acad Sci USA. 1982;79:7405–9. doi: 10.1073/pnas.79.23.7405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gepner J, Hays TS. A fertility region on the Y chromosome of Drosophila melanogaster encodes a dynein microtubule motor. Proc Natl Acad Sci USA. 1993;90:11132–6. doi: 10.1073/pnas.90.23.11132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koerich LB, Wang X, Clark AG, Carvalho AB. Low conservation of gene content in the Drosophila Y chromosome. Nature. 2008;456:949–51. doi: 10.1038/nature07463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carvalho AB, Lazzaro BP, Clark AG. Y chromosomal fertility factors kl-2 and kl-3 of Drosophila melanogaster encode dynein heavy chain polypeptides. Proc Natl Acad Sci USA. 2000;97:13239–44. doi: 10.1073/pnas.230438397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimble J, Edgar L, Hirsh D. Specification of male development in Caenorhabditis elegans: the fem genes. Dev Biol. 1984;105:234–9. doi: 10.1016/0012-1606(84)90279-3. [DOI] [PubMed] [Google Scholar]
- Doniach T, Hodgkin J. A sex-determining gene, fem-1, required for both male and hermaphrodite development in Caenorhabditis elegans. Dev Biol. 1984;106:223–35. doi: 10.1016/0012-1606(84)90077-0. [DOI] [PubMed] [Google Scholar]
- Spence AM, Coulson A, Hodgkin J. The product of fem-1, a nematode sex-determining gene, contains a motif found in cell cycle control proteins and receptors for cell-cell interactions. Cell. 1990;60:981–90. doi: 10.1016/0092-8674(90)90346-G. [DOI] [PubMed] [Google Scholar]
- flybase. http://www.flybase.org
- Yeh SD, Chen YJ, Chang AC, Ray R, She BR, Lee WS, Chiang HS, Cohen SN, Lin-Chao S. Isolation and properties of Gas8, a growth arrest-specific gene regulated during male gametogenesis to produce a protein associated with the sperm motility apparatus. J Biol Chem. 2002;277:6311–7. doi: 10.1074/jbc.M106941200. [DOI] [PubMed] [Google Scholar]
- Yang Y, Cochran DA, Gargano MD, King I, Samhat NK, Burger BP, Sabourin KR, Hou Y, Awata J, Parry DAD, Marshall WF, Witman GB, Lu X. Regulation of flagellar motility by the conserved flagellar protein CG34110/Ccdc135/FAP50. Mol Biol Cell. 2011;22:976–87. doi: 10.1091/mbc.E10-04-0331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garrett-Engele CM, Siegal ML, Manoli DS, Williams BC, Li H, Baker BS. intersex, a gene required for female sexual development in Drosophila, is expressed in both sexes and functions together with doublesex to regulate terminal differentiation. Development. 2002;129:4661–75. doi: 10.1242/dev.129.20.4661. [DOI] [PubMed] [Google Scholar]
- Gubbay J, Collignon J, Koopman P, Capel B, Economou A, Münsterberg A, Vivian N, Goodfellow P, Lovell-Badge R. A gene mapping to the sex-determining region of the mouse Y chromosome is a member of a novel family of embryonically expressed genes. Nature. 1990;346:245–50. doi: 10.1038/346245a0. [DOI] [PubMed] [Google Scholar]
- Wilson MJ, Dearden PK. Evolution of the insect Sox genes. BMC Evol Biol. 2008;8:120. doi: 10.1186/1471-2148-8-120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crémazy F, Berta P, Girard F. Sox neuro, a new Drosophila Sox gene expressed in the developing central nervous system. Mech Dev. 2000;93:215–9. doi: 10.1016/S0925-4773(00)00268-9. [DOI] [PubMed] [Google Scholar]
- Prokopenko SN, Brumby A, O'Keefe L, Prior L, He Y, Saint R, Bellen HJ. A putative exchange factor for Rho1 GTPase is required for initiation of cytokinesis in Drosophila. Genes Dev. 1999;13:2301–14. doi: 10.1101/gad.13.17.2301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Keefe L, Somers WG, Harley A, Saint R. The pebble GTP exchange factor and the control of cytokinesis. Cell Struct Funct. 2001;26:619–26. doi: 10.1247/csf.26.619. [DOI] [PubMed] [Google Scholar]
- Khurana B, Kristie TM. A protein sequestering system reveals control of cellular programs by the transcriptional coactivator HCF-1. J Biol Chem. 2004;279:33673–83. doi: 10.1074/jbc.M401255200. [DOI] [PubMed] [Google Scholar]
- Talbo G, Højrup P, Rahbek-Nielsen H, Andersen SO, Roepstorff P. Determination of the covalent structure of an N- and C-terminally blocked glycoprotein from endocuticle of Locusta migratoria. Combined use of plasma desorption mass spectrometry and Edman degradation to study post-translationally modified proteins. Eur J Biochem. 1991;195:495–504. doi: 10.1111/j.1432-1033.1991.tb15730.x. [DOI] [PubMed] [Google Scholar]
- Pondeville E, Maria A, Jacques J-C, Bourgouin C, Dauphin-Villemant C. Anopheles gambiae males produce and transfer the vitellogenic steroid hormone 20-hydroxyecdysone to females during mating. Proc Natl Acad Sci USA. 2008;105:19631–6. doi: 10.1073/pnas.0809264105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doctor J, Fristrom D, Fristrom JW. The pupal cuticle of Drosophila: biphasic synthesis of pupal cuticle proteins in vivo and in vitro in response to 20-hydroxyecdysone. J Cell Biol. 1985;101:189–200. doi: 10.1083/jcb.101.1.189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leal WS. Odorant reception in insects: roles of receptors, binding proteins, and degrading enzymes. Annu Rev Entomol. 2013;58:373–91. doi: 10.1146/annurev-ento-120811-153635. [DOI] [PubMed] [Google Scholar]
- Pelosi P, Maida R. Odorant-binding proteins in insects. Comp Biochem Physiol B Biochem Mol Biol. 1995;111:503–14. doi: 10.1016/0305-0491(95)00019-5. [DOI] [PubMed] [Google Scholar]
- Vosshall LB, Hansson BS. A unified nomenclature system for the insect olfactory coreceptor. Chem Senses. 2011;36:497–8. doi: 10.1093/chemse/bjr022. [DOI] [PubMed] [Google Scholar]
- Vanderhaeghen P, Schurmans S, Vassart G, Parmentier M. Olfactory receptors are displayed on dog mature sperm cells. J Cell Biol. 1993;123(6 Pt 1):1441–52. doi: 10.1083/jcb.123.6.1441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanderhaeghen P, Schurmans S, Vassart G, Parmentier M. Specific repertoire of olfactory receptor genes in the male germ cells of several mammalian species. Genomics. 1997;39:239–46. doi: 10.1006/geno.1996.4490. [DOI] [PubMed] [Google Scholar]
- Kang N, Koo J. Olfactory receptors in non-chemosensory tissues. BMB Rep. 2012;45:612–22. doi: 10.5483/BMBRep.2012.45.11.232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spehr M, Gisselmann G, Poplawski A, Riffell JA, Wetzel CH, Zimmer RK, Hatt H. Identification of a testicular odorant receptor mediating human sperm chemotaxis. Science. 2003;299:2054–8. doi: 10.1126/science.1080376. [DOI] [PubMed] [Google Scholar]
- Spehr M, Schwane K, Heilmann S, Gisselmann G, Hummel T, Hatt H. Dual capacity of a human olfactory receptor. Curr Biol. 2004;14:R832–3. doi: 10.1016/j.cub.2004.09.034. [DOI] [PubMed] [Google Scholar]
- Spehr M, Schwane K, Riffell JA, Zimmer RK, Hatt H. Odorant receptors and olfactory-like signaling mechanisms in mammalian sperm. Mol Cell Endocrinol. 2006;250:128–36. doi: 10.1016/j.mce.2005.12.035. [DOI] [PubMed] [Google Scholar]
- Fukuda N, Yomogida K, Okabe M, Touhara K. Functional characterization of a mouse testicular olfactory receptor and its role in chemosensing and in regulation of sperm motility. J Cell Sci. 2004;117(Pt 24):5835–45. doi: 10.1242/jcs.01507. [DOI] [PubMed] [Google Scholar]
- Veitinger T, Riffell JR, Veitinger S, Nascimento JM, Triller A, Chandsawangbhuwana C, Schwane K, Geerts A, Wunder F, Berns MW, Neuhaus EM, Zimmer RK, Spehr M, Hatt H. Chemosensory Ca2+ dynamics correlate with diverse behavioral phenotypes in human sperm. J Biol Chem. 2011;286:17311–25. doi: 10.1074/jbc.M110.211524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pitts RJ, Liu C, Zhou X, Malpartida JC, Zwiebel LJ. Odorant receptor-mediated sperm activation in disease vector mosquitoes. Proc Natl Acad Sci USA. 2014;111:2566–71. doi: 10.1073/pnas.1322923111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paesen GC, Happ GM. The B proteins secreted by the tubular accessory sex glands of the male mealworm beetle, Tenebrio molitor, have sequence similarity to moth pheromone-binding proteins. Insect Biochem Mol Biol. 1995;25:401–8. doi: 10.1016/0965-1748(94)00085-V. [DOI] [PubMed] [Google Scholar]
- Thymianou S, Mavroidis M, Kokolakis G, Komitopoulou K, Zacharopoulou A, Mintzas AC. Cloning and characterization of a cDNA encoding a male-specific serum protein of the Mediterranean fruit fly, Ceratitis capitata, with sequence similarity to odorant-binding proteins. Insect Mol Biol. 1998;7:345–53. doi: 10.1046/j.1365-2583.1998.740345.x. [DOI] [PubMed] [Google Scholar]
- Paiva-Silva GO, Sorgine MHF, Benedetti CE, Meneghini R, Almeida IC, Machado EA, Dansa-Petretski M, Yepiz-Plascencia G, Law JH, Oliveira PL, Masuda H. On the biosynthesis of Rhodnius prolixus heme-binding protein. Insect Biochem Mol Biol. 2002;32:1533–41. doi: 10.1016/S0965-1748(02)00074-7. [DOI] [PubMed] [Google Scholar]
- Forêt S, Maleszka R. Function and evolution of a gene family encoding odorant binding-like proteins in a social insect, the honey bee (Apis mellifera) Genome Res. 2006;16:1404–13. doi: 10.1101/gr.5075706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogt RG, Prestwich GD, Lerner MR. Odorant-binding-protein subfamilies associate with distinct classes of olfactory receptor neurons in insects. J Neurobiol. 1991;22:74–84. doi: 10.1002/neu.480220108. [DOI] [PubMed] [Google Scholar]
- Arya GH, Weber AL, Wang P, Magwire MM, Negron YL, Mackay TF, Anholt RR. Natural variation, functional pleiotropy and transcriptional contexts of odorant binding protein genes in Drosophila melanogaster. Genetics. 2010;186:1475–85. doi: 10.1534/genetics.110.123166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou S, Stone EA, Mackay TF, Anholt RR. Plasticity of the chemoreceptor repertoire in Drosophila melanogaster. PLoS Genet. 2009;5:e1000681. doi: 10.1371/journal.pgen.1000681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kodrík D, Filippov VA, Sehnal F, Filippova MA. Sericotropin: an insect neurohormonal factor affecting RNA transcription. Netherlands J Zool. 1995.
- Nagnan-Le Meillour P, Cain AH, Jacquin-Joly E, François MC, Ramachandran S, Maida R, Steinbrecht RA. Chemosensory proteins from the proboscis of mamestra brassicae. Chem Senses. 2000;25:541–53. doi: 10.1093/chemse/25.5.541. [DOI] [PubMed] [Google Scholar]
- Jacquin-Joly E, Vogt RG, François MC, Nagnan-Le Meillour P. Functional and expression pattern analysis of chemosensory proteins expressed in antennae and pheromonal gland of Mamestra brassicae. Chem Senses. 2001;26:833–44. doi: 10.1093/chemse/26.7.833. [DOI] [PubMed] [Google Scholar]
- Wanner KW, Willis LG, Theilmann DA, Isman MB, Feng Q, Plettner E. Analysis of the insect os-d-like gene family. J Chem Ecol. 2004;30:889–911. doi: 10.1023/b:joec.0000028457.51147.d4. [DOI] [PubMed] [Google Scholar]
- Walensky LD, Ruat M, Bakin RE, Blackshaw S, Ronnett G V, Snyder SH. Two novel odorant receptor families expressed in spermatids undergo 5'-splicing. J Biol Chem. 1998;273:9378–87. doi: 10.1074/jbc.273.16.9378. [DOI] [PubMed] [Google Scholar]
- Suzuki N, Garbers DL. Stimulation of sperm respiration rates by speract and resact at alkaline extracellular pH. Biol Reprod. 1984;30:1167–74. doi: 10.1095/biolreprod30.5.1167. [DOI] [PubMed] [Google Scholar]
- Parmentier M, Libert F, Schurmans S, Schiffmann S, Lefort A, Eggerickx D, Ledent C, Mollereau C, Gérard C, Perret J. et al. Expression of members of the putative olfactory receptor gene family in mammalian germ cells. Nature. 1992;355:453–5. doi: 10.1038/355453a0. [DOI] [PubMed] [Google Scholar]
- Ogaugwu CE, Schetelig MF, Wimmer EA. Transgenic sexing system for Ceratitis capitata (Diptera: Tephritidae) based on female-specific embryonic lethality. Insect Biochem Mol Biol. 2013;43:1–8. doi: 10.1016/j.ibmb.2012.10.010. [DOI] [PubMed] [Google Scholar]
- Hanife G. Embryonic development of the olive fruit fly, Bactrocera oleae Rossi ( Diptera: Tephritidae ), in vivo. Turkish J Zool. 2014.
- Schweisguth F, Lepesant JA, Vincent A. The serendipity alpha gene encodes a membrane-associated protein required for the cellularization of the Drosophila embryo. Genes Dev. 1990;4:922–31. doi: 10.1101/gad.4.6.922. [DOI] [PubMed] [Google Scholar]
- Ibnsouda S, Schweisguth F, de Billy G, Vincent A. Relationship between expression of serendipity alpha and cellularisation of the Drosophila embryo as revealed by interspecific transformation. Development. 1993;119:471–83. doi: 10.1242/dev.119.2.471. [DOI] [PubMed] [Google Scholar]
- Schmid KJ, Tautz D. A screen for fast evolving genes from Drosophila. Proc Natl Acad Sci USA. 1997;94:9746–50. doi: 10.1073/pnas.94.18.9746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holt RA, Subramanian GM, Halpern A, Sutton GG, Charlab R, Nusskern DR, Wincker P, Clark AG, Ribeiro JM, Wides R, Salzberg SL, Loftus B, Yandell M, Majoros WH, Rusch DB, Lai Z, Kraft CL, Abril JF, Anthouard V, Arensburger P, Atkinson PW, Baden H, de Berardinis V, Baldwin D, Benes V, Biedler J, Blass C, Bolanos R, Boscus D, Barnstead M, et al. The genome sequence of the malaria mosquito Anopheles gambiae. Science. 2002;298:129–49. doi: 10.1126/science.1076181. [DOI] [PubMed] [Google Scholar]
- Zou Z, Lopez DL, Kanost MR, Evans JD, Jiang H. Comparative analysis of serine protease-related genes in the honey bee genome: possible involvement in embryonic development and innate immunity. Insect Mol Biol. 2006;15:603–14. doi: 10.1111/j.1365-2583.2006.00684.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haugen M, Flannery E, Tomchaney M, Mori A, Behura SK, Severson DW, Duman-Scheel M. Semaphorin-1a is required for Aedes aegypti embryonic nerve cord development. PLoS One. 2011;6:e21694. doi: 10.1371/journal.pone.0021694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schetelig MF, Horn C, Handler AM, Wimmer EA. In: Area-Wide Control Insect Pests. MJB Vreysen, AS Robinson J Hendrichs, editor. 2007. Development of an Embryonic Lethality System in Mediterranean Fruit Fly Ceratitis capitata; pp. 85–93. [Google Scholar]
- Tadros W, Lipshitz HD. The maternal-to-zygotic transition: a play in two acts. Development. 2009;136:3033–42. doi: 10.1242/dev.033183. [DOI] [PubMed] [Google Scholar]
- Schetelig MF, Caceres C, Zacharopoulou A, Franz G, Wimmer EA. Conditional embryonic lethality to improve the sterile insect technique in Ceratitis capitata (Diptera: Tephritidae) BMC Biol. 2009;7:4. doi: 10.1186/1741-7007-7-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gabrieli P, Gomulski LM, Bonomi A, Siciliano P, Scolari F, Franz G, Jessup A, Malacrida AR, Gasperi G. Interchromosomal duplications on the Bactrocera oleae Y chromosome imply a distinct evolutionary origin of the sex chromosomes compared to Drosophila. PLoS One. 2011;6:e17747. doi: 10.1371/journal.pone.0017747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bilak A, Su TT. Regulation of Drosophila melanogaster pro-apoptotic gene hid. Apoptosis. 2009;14:943–9. doi: 10.1007/s10495-009-0374-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kornbluth S, White K. Apoptosis in Drosophila: neither fish nor fowl (nor man, nor worm) J Cell Sci. 2005;118(Pt 9):1779–87. doi: 10.1242/jcs.02377. [DOI] [PubMed] [Google Scholar]
- Hay BA, Guo M. Caspase-dependent cell death in Drosophila. Annu Rev Cell Dev Biol. 2006;22:623–50. doi: 10.1146/annurev.cellbio.21.012804.093845. [DOI] [PubMed] [Google Scholar]
- Steller H. Regulation of apoptosis in Drosophila. Cell Death Differ. 2008;15:1132–8. doi: 10.1038/cdd.2008.50. [DOI] [PubMed] [Google Scholar]
- Horn C, Wimmer EA. A transgene-based, embryo-specific lethality system for insect pest management. Nat Biotechnol. 2003;21:64–70. doi: 10.1038/nbt769. [DOI] [PubMed] [Google Scholar]
- Schetelig MF, Nirmala X, Handler AM. Pro-apoptotic cell death genes, hid and reaper, from the tephritid pest species, Anastrepha suspensa. Apoptosis. 2011;16:759–68. doi: 10.1007/s10495-011-0610-4. [DOI] [PubMed] [Google Scholar]
- Grether ME, Abrams JM, Agapite J, White K, Steller H. The head involution defective gene of Drosophila melanogaster functions in programmed cell death. Genes Dev. 1995;9:1694–708. doi: 10.1101/gad.9.14.1694. [DOI] [PubMed] [Google Scholar]
- Abrams JM, White K, Fessler LI, Steller H. Programmed cell death during Drosophila embryogenesis. Development. 1993;117:29–43. doi: 10.1242/dev.117.1.29. [DOI] [PubMed] [Google Scholar]
- M.E Economopoulos A, Tsitsipis J. The importance of conditions during the adult stage in evaluating an artificial food for larvae of Dacus oleae (Gmel.) (Diptera, Tephritidae) Z Angew Entomol. 1967;59:127–130. [Google Scholar]
- Tsitsipis J. Development of a caging and egging system for mass rearing the olive fruit fly, Dacus oleae (Gmel.) (Diptera, Tephritidae) Ann Zool Ecol Anim. 1977;9:133–139. [Google Scholar]
- Tsitsipis JA, Kontos A. Improved solid adult diet for the olive fruit fly Dacus oleae. Entomol Hell. 1983;1:24–29. [Google Scholar]
- Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL. BLAST+: architecture and applications. BMC Bioinformatics. 2009;10:421. doi: 10.1186/1471-2105-10-421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25:1105–11. doi: 10.1093/bioinformatics/btp120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blüthgen N, Brand K, Cajavec B, Swat M, Herzel H, Beule D. Biological profiling of gene groups utilizing Gene Ontology. Genome Inform. 2005;16:106–15. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


