Abstract
Sterol regulatory element binding protein1c (SREBP1c) is a key transcription factor for de novo lipogenesis during the postprandial state. During nutritional deprivation, hepatic SREBP1c is rapidly suppressed by fasting signals to prevent lipogenic pathways. However, the molecular mechanisms that control SREBP1c turnover in response to fasting status are not thoroughly understood. To elucidate which factors are involved in the inactivation of SREBP1c, we attempted to identify SREBP1c-interacting proteins by mass spectrometry analysis. Since we observed that ring finger protein20 (RNF20) ubiquitin ligase was identified as one of SREBP1c-interacting proteins, we hypothesized that fasting signaling would promote SREBP1c degradation in an RNF20-dependent manner. In this work, we demonstrate that RNF20 physically interacts with SREBP1c, leading to degradation of SREBP1c via ubiquitination. In accordance with these findings, RNF20 represses the transcriptional activity of SREBP1c and turns off the expression of lipogenic genes that are targets of SREBP1c. In contrast, knockdown of RNF20 stimulates the expression of SREBP1c and lipogenic genes and induces lipogenic activity in primary hepatocytes. Furthermore, activation of protein kinase A (PKA) with glucagon or forskolin enhances the expression of RNF20 and potentiates the ubiquitination of SREBP1c via RNF20. In wild-type and db/db mice, adenoviral overexpression of RNF20 markedly suppresses FASN promoter activity and reduces the level of hepatic triglycerides, accompanied by a decrease in the hepatic lipogenic program. Here, we reveal that RNF20-induced SREBP1c ubiquitination down-regulates hepatic lipogenic activity upon PKA activation. Conclusion: RNF20 acts as a negative regulator of hepatic fatty acid metabolism through degradation of SREBP1c upon PKA activation. Knowledge regarding this process enhances our understanding of how SREBP1c is able to turn off hepatic lipid metabolism during nutritional deprivation.
Sterol regulatory element binding proteins (SREBPs) play key roles in lipid homeostasis from yeast to humans.1,2 In mammals, three different SREBP isoforms, including SREBP1a, SREBP1c (also known as ADD1), and SREBP2, are encoded by two genes: SREBF1 and SREBF2. SREBP1 regulates fatty acid metabolism, whereas SREBP2 controls cholesterol metabolism.3 When the cellular sterol level is low, SREBP cleavage-activating protein (SCAP) escorts the SREBP precursors from the endoplasmic reticulum (ER) to the Golgi, where SREBPs are cleaved by Site-1 and Site-2 proteases.4 Subsequently, the mature forms of SREBPs are translocated into the nucleus and stimulate the expression of target genes.5,6 SREBP1a and SREBP1c are generated through transcription from alternative promoters and splicing from a single SREBF1 gene.
In metabolic tissues such as adipose tissue and liver, SREBP1c is the predominant isoform of SREBP1.1,7 SREBP1c governs de novo lipogenesis by stimulating its target genes, including fatty acid synthase (FASN), acetyl-CoA carboxylase1 (ACC1), steroyl-CoA desaturase1 (SCD1), and long-chain fatty acid elongase (ELOVL6).8–10 Furthermore, SREBP1c is sensitively regulated by nutritional and hormonal changes to achieve energy balance. For example, SREBP1c is suppressed by fasting, whereas SREBP1c is activated by feeding in adipose tissue and liver.11,12 In parallel, the expression of most lipogenic genes, including FASN, ACC1, and SCD1, is also modulated in a fashion analogous to that of nutritionally regulated SREBP1c.12–14 Accordingly, it has been reported that various hormones, such as insulin, glucagon, and adrenaline, participate in the regulation of SREBP1c and its target lipogenic genes.15,16 Insulin, a key postprandial hormone, stimulates the expression and activity of SREBP1c to accommodate anabolic processes, such as fatty acid synthesis, upon feeding.12,17 In contrast, glucagon, a key catabolic hormone, suppresses the activity of SREBP1c in fasting states, leading to a decrease in lipogenesis.18
In the nucleus, mature SREBPs are very unstable and are rapidly degraded by the proteasome.19,20 Previous reports have shown that SREBP1 is phosphorylated by glycogen synthase kinase-3 beta (GSK-3β), which leads to F-box and WD repeat domain-containing7 (FBXW7)-dependent ubiquitination of SREBP1.21,22 However, a recent in vivo study revealed that inhibition of FBXW7 does not alter the expression of SREBP1c or lipogenic genes in the liver.23
Although SREBP1c-mediated lipogenic program in liver is rapidly repressed by nutritional deprivations, the factors that are involved in the suppression of SREBP1c activity during fasting have not been thoroughly characterized. The finding that ring finger protein20 (RNF20) ubiquitin ligase was identified as one of the novel SREBP1c-interating proteins led us to test whether fasting signaling would promote SREBP1c degradation in an RNF20-dependent manner. In this study, we demonstrate that RNF20 promotes polyubiquitination and degradation of SREBP1c. Overexpression of RNF20 represses SREBP1c activity, leading to a decrease in the expression of lipogenic genes. In obese db/db mice, RNF20 overexpression alleviates hepatic steatosis by reducing the lipogenic program by way of SREBP1c down-regulation. Furthermore, activated PKA, a major signaling cascade that mediates the fasting state, induces degradation of SREBP1c by increasing RNF20 expression. Taken together, these data suggest that RNF20 plays a critical role in the regulation of hepatic lipid metabolism by modulating the protein stability and transcriptional activity of SREBP1c during hormonal changes.
Materials and Methods
Flag Affinity Purification of the SREBP1c Complex
Adenoviruses encoding green fluorescent protein (GFP) or Flag-SREBP1c, which contains the transcriptionally active fragment of rat SREBP1c (amino acids 1-403) fused with a Flag-tag, were used to infect primary hepatocytes. The infected hepatocytes were then gently washed with ice-cold phosphate-buffered saline (PBS) and lysed in hypotonic buffer (20 mM HEPES [pH 7.9], 1 mM EDTA, 1 mM EGTA, 1 mM DTT, 0.5 mM PMSF, 0.2% [v/v] Nonidet P-40 [NP-40]) and protease inhibitor cocktail (Roche, Rotkreuz, Switzerland). After incubation in hypotonic buffer for 10 minutes, the homogenates were centrifuged at 8,000 rpm for 1 minute at 4°C and the supernatants (cytosolic fraction) were transferred to a fresh tube. The pellet was homogenized in ice-cold high salt buffer (10 mM HEPES [pH 7.9], 420 mM NaCl, 20% [v/v] glycerol, 1 mM EDTA, 1 mM EGTA, 1 mM DTT, 0.5 mM PMSF, and protease inhibitor cocktail) on a rotating shaker for 30 minutes at 4°C and subsequently centrifuged at 12,000 rpm for 15 minutes at 4°C. Consequently, the supernatants (nuclear fraction) were incubated with anti-Flag M2-agarose affinity gel (Sigma-Aldrich, St. Louis, MO) for 2 hours at 4°C on a rotating shaker. The beads were then rinsed four times for 10 minutes each in washing buffer (20 mM HEPES [pH 7.9], 150 mM NaCl, 0.5 mM EDTA, 0.5 mM PMSF, 1% [v/v] Triton X-100, and protease inhibitor cocktail), followed by elution with sodium dodecyl sulfate (SDS) buffer (250 mM Tris-HCl [pH 6.6], 10% [w/v] SDS, 50% [v/v] glycerol, 500 mM DTT, and 0.5% [w/v] bromophenol blue). The eluates were then subjected to mass spectrometry.
Cell-Based Ubiquitination Assay
COS-1 cells were transfected with plasmids encoding Myc-SREBP1c, Flag-RNF20, and HA-ubiquitin in the presence or absence of forskolin (20 μM). After transfection for 36 hours, the cells were incubated with MG132 (10 μM) for 12 hours and lysed with cold RIPA buffer. Equal amounts of total cell lysates were incubated with the Myc antibodies for 2 hours at 4°C. Immunocomplexes were collected using protein-A sepharose beads (GE Healthcare, Buckinghamshire, UK) for 1 hour at 4°C. Further, the immunoprecipitates were washed with RIPA buffer and subjected to SDS-polyacrylamide gel electrophoresis (SDS-PAGE) followed by western blotting analyses with anti-HA antibodies.
Animals
All animal experiments were approved by the Seoul National University Animal Experiment Ethics Committee. Male C57BL/6 mice were obtained from Samtako (Osan, Korea) and db/db mice were obtained from Central Lab (Seoul, Korea). The animals were housed in colony cages in 12-hour light/dark cycles. Standard chow (Purina Mills) was given ad libitum.
In Vivo Imaging System
Ten-week-old C57BL/6 mice and 9-week-old db/db mice were injected through the tail vein with adenoviruses encoding GFP (Ad-Mock), RNF20 (Ad-RNF20), and FASN luciferase (Ad-FASN-luc). After 7 days, adenovirus-infected mice were injected intraperitoneally with 100 mg/kg sterile D-luciferin. The mice were then anesthetized with Zoletil and imaged using the IVIS 100 Imaging System (Xenogen) as described previously.24
Results
SREBP1c Is Decreased by Ubiquitination Upon PKA Activation
SREBP1c is tightly regulated by hormonal and nutritional changes to reflect the energy status.15,16 When we examined the protein stability of SREBP1c in the presence of cycloheximide, an inhibitor of protein synthesis, the degradation rate of nuclear SREBP1c was reduced in the presence of MG132, an inhibitor of the 26S proteasome (Fig. 1A; Supporting Fig. S1A), indicating that the rapid turnover of SREBP1c protein might be mediated by proteasomal degradation. Although it has been demonstrated that the cyclic adenosine monophosphate (cAMP)/PKA pathway is involved in the suppression of SREBP1c,18,25,26 it is largely unknown how PKA affects SREBP1c protein stability. To address this issue, primary hepatocytes were treated with forskolin, an activator of the PKA cascade. The level of SREBP1 protein was reduced by forskolin (Fig. 1B; Supporting Fig. S1B), while that of SREBP1 protein was restored by MG132 in a dose-dependent manner (Fig. 1C; Supporting Fig. S1C), implying that PKA would be involved in the regulation of SREBP1c protein stability through the ubiquitin-proteasome system. Next, to investigate whether SREBP1c is indeed ubiquitinated by PKA activation, cell-based ubiquitination assays were carried out. As shown in Fig. 1D, forskolin efficiently polyubiquitinated the SREBP1c protein, while H-89, the inhibitor of PKA, greatly attenuated forskolin-induced SREBP1c ubiquitination. Therefore, these data suggest that SREBP1c is degraded by ubiquitination upon PKA activation.
Fig 1.
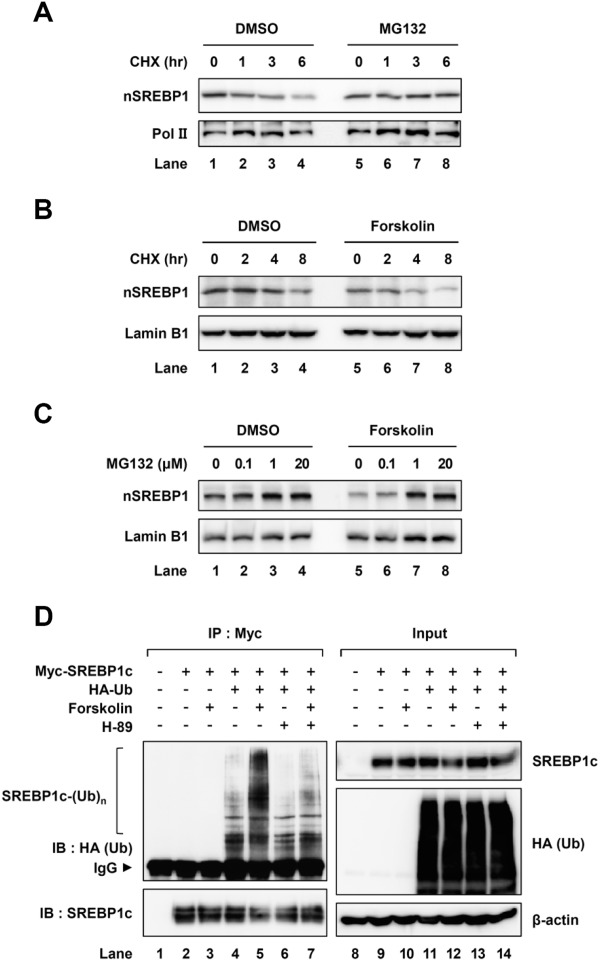
PKA activation decreases SREBP1c protein stability. (A) Mouse primary hepatocytes were treated with cycloheximide (20 μM) for the indicated periods with or without MG132 (20 μM) treatment for 3 hours. After the preparation of nuclear extracts, western blotting analyses were performed with the indicated antibodies. DMSO, dimethyl sulfoxide; CHX, cycloheximide; nSREBP1, nuclear SREBP1. (B) Mouse primary hepatocytes were treated with forskolin (50 μM) for 6 hours and harvested at the indicated timepoints after cycloheximide (50 μM) treatment. After isolating nuclear extracts, western blotting analyses were performed with the indicated antibodies. (C) Mouse primary hepatocytes were treated with or without forskolin (50 μM) in the presence of MG132 with the indicated dose for 4 hours. Nuclear extracts were isolated and analyzed by western blotting analyses with the indicated antibodies. (D) COS-1 cells were cotransfected with plasmids encoding Myc-SREBP1c and HA-ubiquitin. After transfection, the cells were treated with MG132 (10 μM) for 12 hours and subsequently pretreated with H-89 (20 μM) for 1 hour. Then the cells were treated with or without forskolin (20 μM) for another 3 hours. Total cell lysates were subjected to coimmunoprecipitation with anti-Myc antibody, followed by western blotting analyses with the indicated antibodies. IP, immunoprecipitation; IB, immunoblotting; IgG, immunoglobulin G. All experiments were repeated independently at least three times and representative results are shown.
RNF20 Physically Interacts With SREBP1c
To investigate which factors are involved in the degradation and ubiquitination of nuclear SREBP1c protein, we attempted to identify SREBP1c-interacting proteins; particularly, we were eager to identify an E3 ubiquitin ligase. Mouse primary hepatocytes were infected with adenovirus expressing nuclear SREBP1c, and affinity purifications were conducted. Mass spectrometry analyses indicated that RNF20 (also known as BRE1A) was a potential SREBP1c-interacting protein (Supporting Fig. S2A,B). To validate the physical interaction between RNF20 and SREBP1c, glutathione S-transferase (GST) pull-down assays were performed. Because SREBP1c forms homodimers, 35S-Met labeled SREBP1c was used as a positive control (Fig. 2A, lane 5). RNF20 protein was detected in the fractions eluted from the GST-SREBP1c protein complex (Fig. 2A, lane 6). Further, when we biochemically tested the physical interaction between RNF20 and SREBP1c, RNF20 was coimmunoprecipitated with SREBP1c (Fig. 2B), indicating that RNF20 could physically interact with SREBP1c. Moreover, we observed that endogenous RNF20 formed immunocomplex with endogenous SREBP1 in liver nuclear extracts (Fig. 2C). In parallel immunocytochemistry experiments, exogenous Flag-SREBP1c and Myc-RNF20 proteins colocalized in the nucleus (Fig. 2D), implying that RNF20 can interact with SREBP1c in the nucleus.
Fig 2.
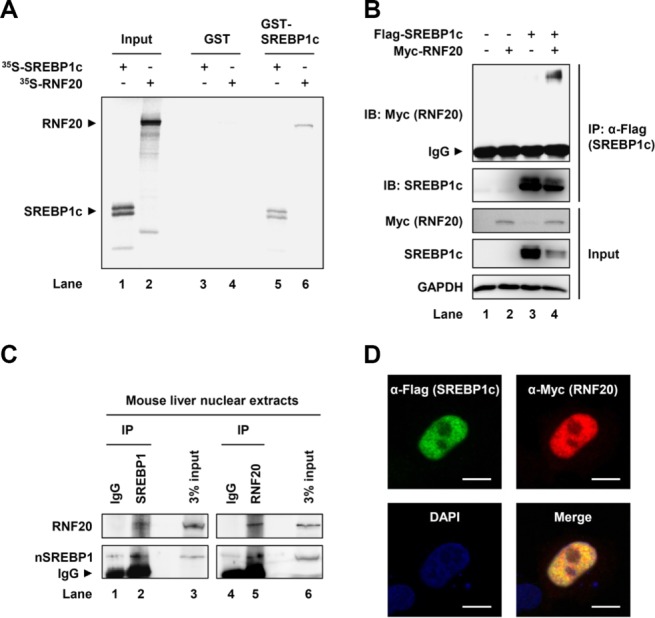
RNF20 physically interacts with SREBP1c. (A) GST pull-down assays were performed as described in the Supporting Material. Inputs of in vitro-translated 35S-Met-labeled SREBP1c (nuclear form) and RNF20 are shown in lanes 1 and 2, respectively. Radioisotope-labeled proteins were mixed with GST or GST-SREBP1c recombinant proteins and GST pull-down analyses were performed. GST, glutathione S-transferase. (B) HEK293T cells were transiently cotransfected with Flag-SREBP1c and/or Myc-RNF20 expression vectors. After preparing total cell lysates, coimmunoprecipitation with anti-Flag antibody and western blotting analyses were performed with the indicated antibodies. IP, immunoprecipitation; IB, immunoblotting; IgG, immunoglobulin G. (C) Nuclear extracts were isolated from mouse livers and subsequently coimmunoprecipitated with IgG or the indicated antibody. Immuno-protein complexes were detected by western blotting analyses with the indicated antibodies. IgG, immunoglobulin G; nSREBP1, nuclear SREBP1. (D) Immunocytochemistry analyses of coexpressed Flag-SREBP1c and Myc-RNF20 in HeLa cells. DAPI, 4′,6-diamidino-2-phenylindole. Scale bars = 10 μm.
RNF20 Promotes Ubiquitination and Degradation of SREBP1c
Given that the E3 ubiquitin ligase RNF20 associates with SREBP1c, the effect of RNF20 on SREBP1c stability was examined. As shown in Fig. 3A, the level of SREBP1c protein was decreased by RNF20 in a dose-dependent manner. Moreover, the RNF20-induced reduction of SREBP1c protein was alleviated by MG132 (Fig. 3A, lane 6), indicating that RNF20 would stimulate SREBP1c degradation by way of ubiquitin-proteasomal degradation. Consistent with these data, cycloheximide-chase assays revealed that the half-life of SREBP1c was shortened by ectopic expression of RNF20 (Fig. 3B). Next, as RNF20 exhibits E3 ubiquitin ligase activity, we tested whether SREBP1c is ubiquitinated by RNF20. As shown in Fig. 3C, RNF20 greatly promoted the level of SREBP1c polyubiquitination. To confirm whether endogenous RNF20 plays a role in SREBP1c ubiquitination, we investigated the effect of RNF20 knockdown by way of small interfering RNA (siRNA) on the ubiquitination of SREBP1c. We found that suppression of RNF20 relieved the level of SREBP1c polyubiquitination (Fig. 3D). These data indicate that RNF20 can act as an E3 ubiquitin ligase for SREBP1c and can accelerate polyubiquitination and degradation of SREBP1c protein.
Fig 3.
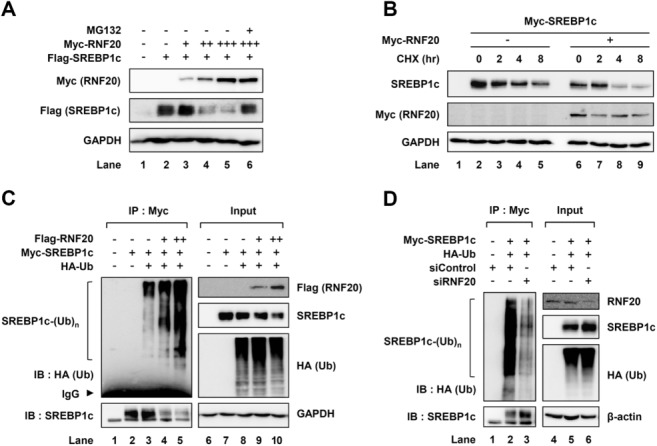
RNF20 mediates ubiquitination and degradation of SREBP1c. (A) HEK293T cells transfected with Flag-SREBP1c and/or Myc-RNF20 expression vectors were incubated with or without MG132 (20 μM) for 4 hours and total cell lysates were subjected to SDS-PAGE followed by western blotting analyses with the indicated antibodies. (B) COS-1 cells transfected with Myc-SREBP1c and/or Myc-RNF20 expression vectors were harvested at 0, 2, 4, and 8 hours after cycloheximide treatment (30 μM). After isolating total cell lysates, western blotting analyses were performed with the indicated antibodies. CHX, cycloheximide. (C) COS-1 cells were cotransfected with Myc-SREBP1c, HA-tagged ubiquitin, and/or Flag-RNF20 expression vectors. After incubation for 36 hours, the cells were treated with MG132 (10 μM) for 12 hours. Total cell lysates were isolated and subjected to cell-based ubiquitination assays. IP, immunoprecipitation; IB, immunoblotting; IgG, immunoglobulin G. (D) COS-1 cells were transfected with nonspecific control siRNA (siControl) or RNF20-specific siRNA (siRNF20). After incubation for 36 hours, the cells were incubated with MG132 (10 μM) for 12 hours. Total cell lysates were isolated and subjected to cell-based ubiquitination assays. The data shown are representative results of at least three independent experiments.
RNF20 Suppresses the Transcriptional Activity of SREBP1c
To understand the biological significance of SREBP1c degradation by RNF20, we investigated the transcriptional activity of SREBP1c with or without RNF20 overexpression. Consistent with previous reports,12,27 SREBP1c transactivated the promoters of FASN and adiponectin (Acrp30) genes, whereas coexpression with RNF20 suppressed the transcriptional activity of SREBP1c (Fig. 4A). When the level of SREBP1 protein was investigated with or without RNF20 overexpression in primary hepatocytes, RNF20 overexpression clearly decreased in endogenous and ectopic nuclear SREBP1 protein (Fig. 4B). Moreover, RNF20 overexpression indeed decreased elevated endogenous SREBP1 protein in the presence of insulin and/or T0901317, a LXR agonist (Supporting Fig. S3). Next, to address the question whether RNF20 could affect the expression of SREBP1c target genes, we analyzed the messenger RNA (mRNA) levels of its target genes in RNF20 and/or SREBP1c-overexpressing hepatocytes. In primary hepatocytes, RNF20 overexpression alone significantly decreased many lipogenic genes including SREBP1c, FASN, SCD1, ACC1, and ELOVL6 (Fig. 4C, lane 2). Consistent with previous reports,3,12 SREBP1c elevated the mRNA expression of lipogenic genes such as FASN, SCD1, and ELOVL6 in primary hepatocytes (Fig. 4C, lane 3). However, the effects of RNF20 overexpression on lipogenic suppression were marginal in SREBP1c-overexpressing hepatocytes (Fig. 4C, lane 4). It is feasible to speculate that high levels of SREBP1c protein may partly stimulate lipogenic gene expression even in the presence of RNF20 overexpression. On the other hand, overexpression of RNF20 did not significantly alter the mRNA level of other SREBP isoforms, including SREBP1a and 2 (Fig. 4C). Moreover, in primary hepatocytes, RNF20 overexpression decreased intracellular lipid accumulation as determined by Oil Red O staining (Fig. 4D). These data indicate that RNF20 can indeed down-regulate lipogenic gene expression by suppressing SREBP1c.
Fig 4.
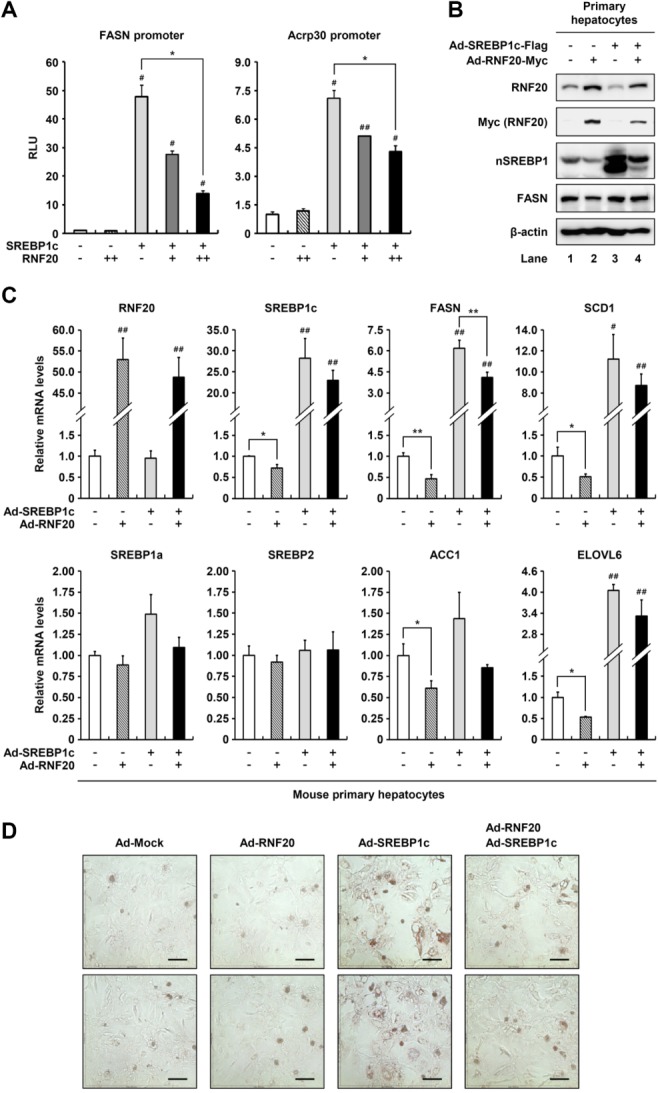
RNF20 down-regulates SREBP1c transcriptional activity and the expression of SREBP1c target genes. (A) HEK293T cells were cotransfected with luciferase reporter plasmids containing the FASN or Acrp30 promoter along with pRSV-β-gal, Myc-SREBP1c, and/or Myc-RNF20 expression vectors. Total cell lysates were subjected to luciferase and β-galactosidase assays. The values are representative data from three independent experiments carried out in triplicate. Each bar represents the mean ± SD of three individual samples. #P < 0.05 versus negative control. ##P < 0.01 versus negative control. *P < 0.05. RLU, relative luminescence units. (B) Mouse primary hepatocytes were infected with adenoviruses containing GFP alone (Ad-Mock), Flag-SREBP1c (Ad-SREBP1c), and/or Myc-RNF20 (Ad-RNF20) as indicated. After infection for 12 hours, total cell lysates were subjected to SDS-PAGE followed by western blotting analyses with the indicated antibodies. nSREBP1, nuclear SREBP1. (C) Mouse primary hepatocytes were infected with Ad-SREBP1c and/or Ad-RNF20. After infection for 12 hours, the culture media were replaced with fresh media and subsequently incubated for 36 hours. The relative mRNA levels were measured using quantitative reverse-transcription polymerase chain reaction (qRT-PCR). The relative values were normalized to the level of GAPDH mRNA. Each bar represents the mean ± SD of three individual samples. #P < 0.05 versus negative control. ##P < 0.01 versus negative control. *P < 0.05. **P < 0.01. (D) Mouse primary hepatocytes were transduced with Ad-SREBP1c and/or Ad-RNF20 adenoviruses. After incubation for 48 hours, intracellular lipid droplets were visualized by Oil Red O staining and the cells were photographed. Microscopic views of cells at a magnification of ×200 are shown. Scale bars = 100 μm.
Suppression of RNF20 Enhances Hepatic Lipid Metabolism via SREBP1c
To determine whether endogenous RNF20 influences lipogenic activity, we investigated the effect of RNF20 suppression by way of siRNA on lipogenic gene expression in primary hepatocytes. When the level of RNF20 was decreased (by ∼25%), the mRNA levels of SREBP1c and its target genes, including FASN and ELOVL6, were significantly elevated (Fig. 5A). On the contrary, the mRNA levels of other SREBP isoforms, including SREBP1a and 2, were not altered by RNF20 suppression (Fig. 5A), whereas the level of SREBP1c mRNA was up-regulated, probably through an autoregulatory mechanism.28,29 Consistently, suppression of RNF20 with siRNA in primary hepatocytes increased the level of endogenous SREBP1 protein (both precursor and nuclear forms) (Fig. 5B) and promoted intracellular neutral lipid accumulation as determined by Oil Red O staining (Fig. 5C). Additionally, RNF20-suppressed hepatocytes stored more intracellular triglycerides than control primary hepatocytes (Fig. 5D). On the contrary, the mRNA expression levels of other lipogenic transcription factors, such as peroxisome proliferator-activated receptor-gamma (PPARγ), liver X receptor-alpha (LXRα), and carbohydrate responsive element-binding protein (ChREBP) were not altered by RNF20 suppression (Fig. 5A; Supporting Fig. S4). Besides, the mRNA levels of fatty acid oxidation pathway genes, including peroxisome proliferator-activated receptor-alpha (PPARα), carnitine palmitoyltransferase 1A (CPT1A), medium-chain acyl-CoA dehydrogenase (MCAD), and aconitase 1 (ACO1) were not changed by RNF20 suppression in primary hepatocytes (Supporting Fig. S4), implying that RNF20 can selectively down-regulate lipogenic gene expression via SREBP1c in hepatocytes.
Fig 5.
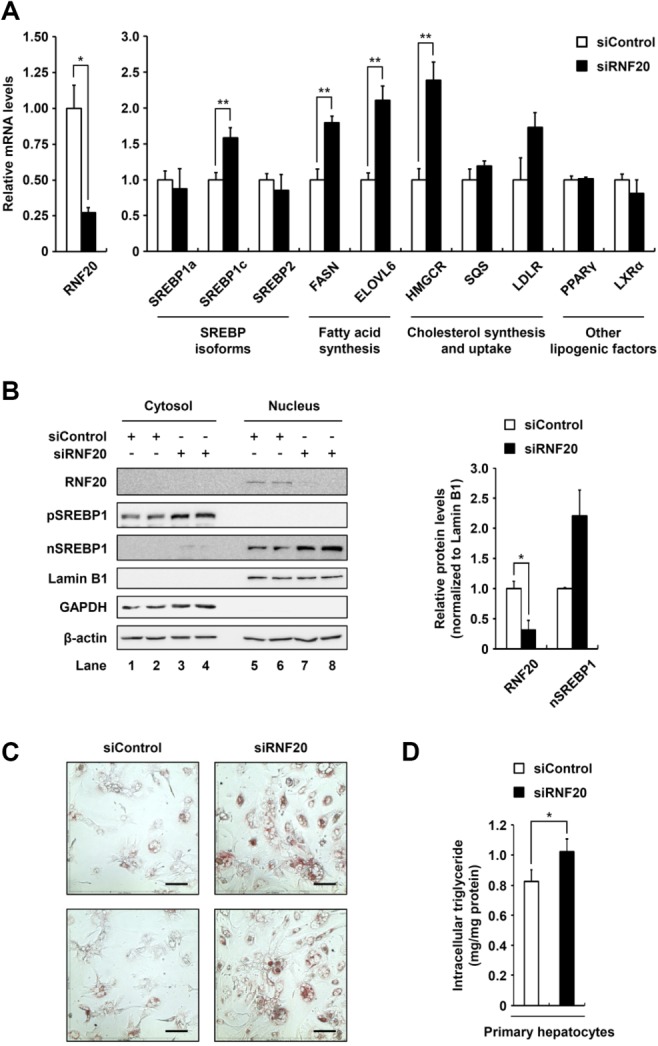
RNF20 negatively regulates hepatic lipogenesis. (A) Mouse primary hepatocytes were transfected with siControl or siRNF20 and relative mRNA levels were determined using qRT-PCR. The level of each mRNA was normalized to the mRNA level of the TATA-binding protein (TBP) gene. Each bar represents the mean ± SD of three individual samples. *P < 0.05 and **P < 0.01 were considered significant. SQS, squalene synthetase; LDLR, low-density lipoprotein receptor. (B) Mouse primary hepatocytes were transfected with siControl or siRNF20. After isolating nuclear extracts, western blotting analyses were performed with the indicated antibodies. Relative amounts of RNF20 and nuclear SREBP1 proteins were calculated using Labworks software (UVP Bioimaging System) and normalized to Lamin B1. The western blots shown are representative of three independent experiments. pSREBP1, precursor SREBP1; nSREBP1, nuclear SREBP1. (C) Mouse primary hepatocytes were transfected with siControl or siRNF20. After incubation for 48 hours, intracellular lipid droplets were visualized by Oil Red O staining and the cells were photographed. Microscopic views of cells at a magnification of ×200 are shown. Scale bars = 100 μm. (D) Intracellular triglyceride contents were measured biochemically.
Expression of Hepatic RNF20 Is Nutritionally Regulated
Nutritional and hormonal changes coordinate energy homeostasis, including lipid and glucose metabolism. Thus, we examined whether the nutritional status might modulate hepatic RNF20 expression. In the liver of fasted mice, the expression of RNF20 mRNA was significantly increased (Fig. 6A), whereas that of SREBP1c and FASN mRNA was decreased. These observations indicate that the change in RNF20 expression might be associated with the regulation of SREBP1c during nutritional changes. In contrast, the mRNA level of FBXW7, an E3 ligase of SREBP proteins, was unchanged by either feeding or fasting conditions (Fig. 6A). Upon fasting, glucagon stimulates the PKA cascade to regulate lipid and glucose metabolism to accomplish catabolic responses.14,30 Thus, to address the question of whether RNF20 expression might be regulated by PKA activation, we examined the level of RNF20 mRNA in hepatocytes with or without forskolin. Notably, forskolin increased the level of RNF20 mRNA, whereas it decreased SREBP1c and SCD1 mRNA levels (Fig. 6B). Furthermore, in primary hepatocytes, both glucagon and forskolin increased the RNF20 protein level and decreased the nuclear SREBP1 protein level (Fig. 6C). Next, to determine whether cytosolic or nuclear RNF20 is regulated upon PKA activation, cytosolic and nuclear extracts were isolated from hepatoma cells with or without forskolin treatment. As shown in Fig. 6D, forskolin enhanced the nuclear RNF20 protein level and reduced the nuclear SREBP1 protein level. The results from a cell-based ubiquitination assay revealed that RNF20 increased the level of SREBP1c polyubiquitination in the presence of forskolin (data not shown). However, unlike RNF20, FBXW7 expression was not altered by forskolin (Fig. 6B). In addition, we investigated the mRNA levels of SREBP1c target genes in RNF20-suppressed hepatocytes treated with forskolin. As shown in Fig. 6E, the mRNA levels of lipogenic genes such as SREBP1c, FASN, and SCD1 were decreased by forskolin, whereas RNF20 suppression by way of siRNA restored the expression of lipogenic genes, even in the presence of forskolin (Fig. 6E). To understand whether RNF20 is involved in the PKA-dependent SREBP1c ubiquitination, we examined the level of SREBP1c ubiquitination upon PKA activation with RNF20 suppression. As shown in Fig. 6F, knockdown of RNF20 with siRNA successfully decreased the level of PKA-mediated SREBP1c ubiquitination, implying that PKA-dependent SREBP1c degradation would require RNF20. These data strongly indicate that RNF20 is a negative regulator of SREBP1c and the lipogenic program upon PKA activation in hepatocytes.
Fig 6.
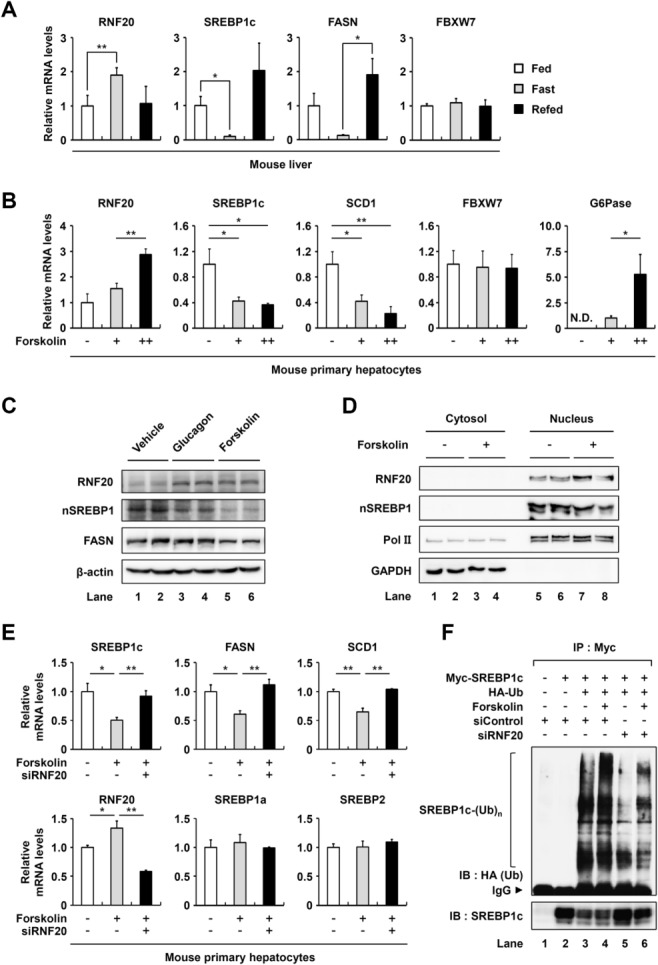
Hepatic RNF20 is regulated by nutritional changes. (A) The level of each mRNA in the livers of fed and fasted mice were determined by qRT-PCR analysis. The TBP mRNA level was used for normalization. The fed control (Fed) was allowed free access to food and the fasted groups (Fast) were denied access to food for 24 hours. The refed group (Refed) was allowed free access to food for 7 hours after 24 hours of fasting. n = 4 for each group. *P < 0.05 and **P < 0.01 were considered significant. (B) Mouse primary hepatocytes were incubated with or without forskolin (+ indicates 10 μM forskolin and ++ indicates 100 μM forskolin) for 6 hours. Each relative mRNA level was determined using qRT-PCR and normalized to the TBP mRNA level. As a positive control, the expression of G6Pase mRNA was greatly enhanced by forskolin. Each bar represents the mean ± SD of three individual samples. *P < 0.05 and **P < 0.01 were considered significant. N.D., not detected. (C) Mouse primary hepatocytes were incubated with glucagon (100 nM) or forskolin (50 μM) for 4 hours. Total cell lysates were subjected to western blotting analyses with the indicated antibodies. nSREBP1, nuclear SREBP1. (D) H4IIE rat hepatoma cells were incubated with forskolin (50 μM) for 5 hours. Cytosolic and nuclear extracts were subjected to western blotting analyses with the indicated antibodies. The results are representative of three independent experiments. (E) Mouse primary hepatocytes transfected with siControl or siRNF20 were incubated with forskolin (10 μM) for 6 hours. The level of each mRNA was determined using qRT-PCR and normalized to the TBP mRNA level. Each bar represents the mean ± SD of experiments carried out in triplicate. *P < 0.05 and **P < 0.01 were considered significant. (F) COS-1 cells were cotransfected with DNA plasmids and siRNAs as indicated. After incubation for 36 hours, the cells were incubated with MG132 (10 μM) for 12 hours and subsequently treated with forskolin (20 μM) for 4 hours. Then the cells were treated with or without forskolin (20 μM) for another 3 hours. Total cell lysates were isolated and subjected to cell-based ubiquitination assays. IP, immunoprecipitation; IB, immunoblotting; IgG, immunoglobulin G.
RNF20 Represses Hepatic Lipid Metabolism In Vivo
To further examine whether RNF20 confers hepatic lipid metabolism via SREBP1c in vivo, adenovirus-expressing RNF20 was intravenously injected into wild-type mice. In agreement with the above data, in vivo RNF20 overexpression decreased the level of hepatic SREBP1 protein (Fig. 7A). Consistently, the hepatic triglyceride level was decreased by RNF20 overexpression (Fig. 7B). To confirm the possibility that the decrease in hepatic lipid metabolism is mediated by RNF20-dependent SREBP1c suppression, we tested FASN promoter activity in vivo. As shown in Fig. 7C, optical in vivo imaging analyses revealed that adenoviral RNF20 overexpression repressed FASN promoter activity compared with control mice. In addition, we found that the expression of lipogenic genes such as SREBP1c, FASN, SCD1, and ELOVL6 was significantly attenuated in the liver of RNF20-overexpressing mice (Fig. 7D). However, hepatic RNF20 overexpression did not significantly alter the mRNA level of other SREBPs such as SREBP1a and 2 (Fig. 7D). Moreover, adenovirally overexpressed RNF20 in vivo did not change the mRNA levels of other lipogenic transcription factors, including PPARγ, LXRα, and ChREBP, and fatty acid oxidation pathway genes in liver (Fig. 7D; Supporting Fig. S5). Thus, these in vivo data confirm the notion that RNF20 plays an important role in the regulation of hepatic lipogenesis via SREBP1c.
Fig 7.
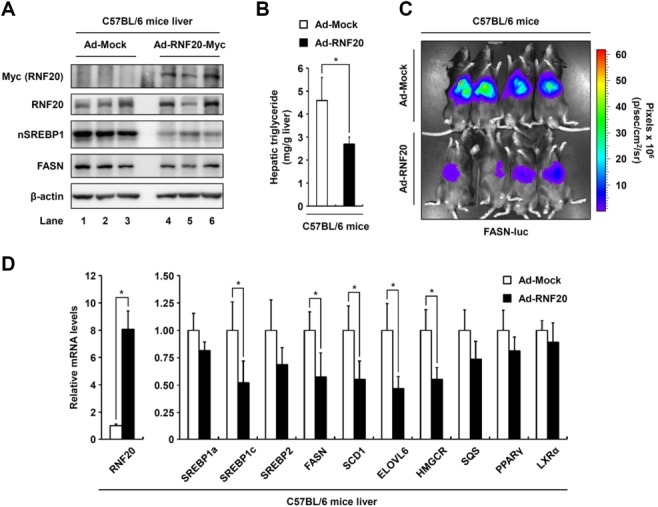
RNF20 overexpression inhibits the hepatic lipogenic program in vivo. (A) Ten-week-old male C57BL/6 mice were infected through the tail vein with adenovirus encoding GFP (Ad-Mock) as the control or mouse RNF20 (Ad-RNF20) (adenoviral dose of 1 × 1010 viral particles per mouse). After 7 days of adenoviral injection, the mice were sacrificed in fed states. The expression levels of RNF20, nSREBP1, and FASN protein in the livers of mice infected with Ad-Mock or Ad-RNF20 were monitored by western blotting analyses. nSREBP1, nuclear SREBP1. (B) The hepatic triglyceride level was measured from 100 mg liver tissue as described in the Supporting Material. *P < 0.05 was considered significant (versus the Ad-Mock control group). (C) Live imaging of in vivo FASN-luciferase (FASN-luc) activity in response to RNF20 overexpression in C57BL/6 mice. In vivo luminescence was measured 7 days postadenoviral infection as described in the Materials and Methods. (D) In C57BL/6 mouse liver injected Ad-Mock or Ad-RNF20, the effects of adenoviral RNF20 overexpression on lipogenic gene expression were determined by qRT-PCR analyses. The level of TBP mRNA was used for normalization. Each mRNA level is shown as a ratio relative to the Ad-Mock control group. *P < 0.05 was considered significant (versus the Ad-Mock control group). n = 3 for each group in panel A, and n = 4 for each group in all other panels.
RNF20 Alleviates Hepatic Steatosis in db/db Mice
Several obese rodent animals, including db/db mice, exhibit hepatic steatosis with enhanced SREBP1c and lipogenic activity.31–33 To investigate whether RNF20 overexpression might alleviate fatty liver through the regulation of SREBP1c, RNF20 was overexpressed by adenovirus in the liver of db/db mice. In db/db mice, adenoviral overexpression of RNF20 did not cause major differences in body weight and fasting blood glucose (Supporting Fig. S6A,B). Similar to lean mice, hepatic RNF20 overexpression decreased both precursor and nuclear forms of SREBP1 protein in db/db mice (Fig. 8A). In accordance with reduced SREBP1 expression, hepatic steatosis was alleviated by RNF20 overexpression in db/db mice (Fig. 8B,D; Supporting Fig. S6D). Furthermore, hepatic RNF20 overexpression in db/db mice significantly lowered FASN promoter activity (Fig. 8C) and reduced the expression of lipogenic genes (Fig. 8E). However, similar to lean mice, hepatic RNF20 overexpression in db/db mice did not alter the mRNA levels of fatty acid oxidation pathway genes (Supporting Fig. S6C), indicating that hepatic RNF20 overexpression would alleviate hepatic steatosis by suppression of SREBP1c and lipogenic gene expression. Next, to investigate whether hepatic RNF20 might be involved in glucose homeostasis in diabetic db/db mice, oral glucose tolerance tests were examined. As shown in Fig. 8F, RNF20 overexpression improved glucose intolerance in db/db mice. Together, these data propose the idea that hepatic RNF20 might also affect the regulation of whole body energy metabolisms such as lipid and glucose.
Fig 8.
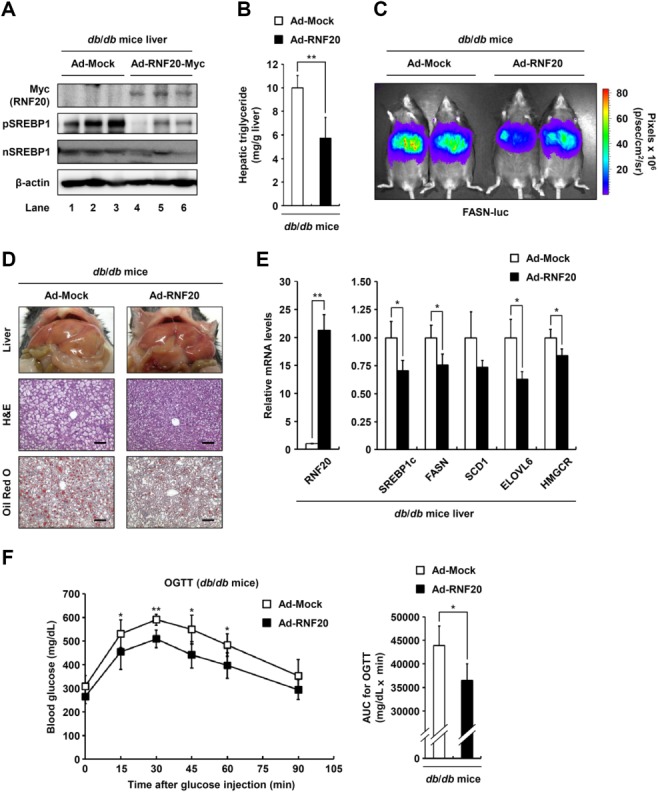
RNF20 overexpression alleviates hepatic steatosis in db/db mice. (A) Nine-week-old male db/db mice were infected through the tail vein with adenovirus encoding GFP or RNF20 (adenoviral dose of 2 × 1010 viral particles per mouse). After 7 days of adenoviral injection, the mice were sacrificed in fed states. The liver tissue was then subjected to SDS-PAGE followed by western blotting analyses with the indicated antibodies. n = 3 for each group. (B) The level of hepatic triglycerides was measured from 100 mg liver tissue in db/db mice infected with each adenovirus. n = 5 for each group. *P < 0.05 was considered significant (versus the Ad-Mock control group). (C) Live imaging of in vivo FASN-luciferase (FASN-luc) activity in response to RNF20 overexpression in db/db mice. n = 2 for each group. (D) Representative hematoxylin and eosin (H&E) and Oil Red O staining of liver sections of db/db mice infected with Ad-Mock or Ad-RNF20. Scale bars = 100 μm. See also Supporting Fig. S6D. (E) The relative mRNA levels of various lipogenic genes in the livers of db/db mice infected with Ad-Mock or Ad-RNF20. Each mRNA level was determined by qRT-PCR analyses and normalized to the TBP mRNA level. Each relative mRNA level is presented as a ratio relative to the Ad-Mock-infected db/db control group. n = 3 for each group. *P < 0.05 and **P < 0.01 were considered significant (versus the Ad-Mock control group). (F) db/db mice were infected with Ad-Mock or Ad-RNF20 and subjected to oral glucose tolerance test. n = 5 at each timepoint. *P < 0.05 and **P < 0.01 were considered significant (versus the Ad-Mock control group). This result was confirmed by area-under-the-curve (AUC) analysis.
Discussion
Due to its key roles in lipid metabolism, several metabolic diseases are associated with SREBP1c dysregulation. Both animal models and human subjects with obesity and diabetes mellitus often suffer from hepatic steatosis and hyperlipidemia, accompanied by elevated SREBP1c.31–33 Thus, elucidation of the molecular mechanisms of SREBP1c function and its lipogenic activity under physiological conditions is important. SREBP1c is controlled by several different mechanisms, including transcriptional regulation, proteolytic maturation, and posttranslational modifications.19–21,34–37 It has been reported that PKA, one of the fasting-induced kinases, phosphorylates SREBP1c and consequently suppresses lipogenic activity.18 However, the factors that are involved in SREBP1c degradation under fasting conditions are poorly understood. Here, we demonstrated that SREBP1c protein is ubiquitinated and degraded by an E3 ubiquitin ligase concurrent with PKA activation during fasting.
RNF20 was first identified as yeast Bre1 and possesses a RING finger domain that primarily functions as an E3 ligase for histone H2B monoubiquitination, which regulates transcription of certain genes.38–40 In addition, it has been reported that knockdown of RNF20 leads to abrogation of H2B monoubiquitination and elevated expression of several proto-oncogenes for tumorigenesis, indicating that RNF20 could act as a tumor suppressor protein.41,42 However, to date, there is no report that RNF20 could selectively regulate hepatic lipid metabolism. In this work, we identified that RNF20 would act as a negative regulator of hepatic lipid metabolism in an SREBP1c-dependent pathway. Several lines of evidence from our in vitro and in vivo data support the above idea. First, ectopic expression of RNF20 repressed SREBP1c and lipogenic gene expression in primary hepatocytes (Fig. 4) and mouse liver (Figs. 7, 8). Second, the level of nuclear SREBP1 protein and the expression of its target genes were increased by RNF20 knockdown in primary hepatocytes, accompanied by augmentation of intracellular lipid accumulation (Fig. 5). Moreover, the mRNA levels of other lipogenic factors, such as PPARγ, LXRα, and ChREBP, and fatty acid oxidation pathway genes were not significantly altered by RNF20-overexpressing or suppressing conditions (Figs. 5A, 7D, 8E; Supporting Figs. S4, S5, S6C), indicating that RNF20 would control hepatic lipid metabolism through SREBP1c modulation.
In order to examine the effect of RNF20 on other SREBP isoforms such as SREBP1a and 2, we tested the effect of RNF20 on degradation of SREBP1a and 2. As shown in Supporting Fig. S7A, RNF20 was also able to induce the degradation of nuclear SREBP1a and 2, implying that RNF20 may regulate the stability of all isoforms of SREBP proteins, at least in an in vitro cell culture system. Although RNF20 may influence hepatic lipid metabolism by way of SREBP1a, 1c, and/or 2, several current in vitro and in vivo data supported the idea that hepatic lipid metabolism would be primarily regulated by SREBP1c rather than 1a or 2 (Figs. 4C, 5A, 6E, 7D, 8E). Nevertheless, future studies are necessary to clarify whether RNF20 might be involved in the regulation of other SREBP isoforms in vivo.
Next, to test the possibility that the expression of hepatic RNF20 might be altered in pathophysiological conditions, we examined the mRNA level of hepatic RNF20 by comparison with db/+ versus db/db and normal-chow diet versus high-fat diet-fed mice. Although the levels of lipogenic genes such as SREBP1c, FASN, and SCD1 were increased in the liver of insulin-resistant mouse models, the level of hepatic RNF20 was not significantly altered (Supporting Fig. S8A,B). Thus, it is likely that RNF20 might regulate lipogenic activity upon hormonal changes in normal conditions rather than pathophysiological conditions. However, since overexpression of hepatic RNF20 markedly improved glucose intolerance in diabetic db/db mice (Fig. 8F), we cannot exclude the possibility that hepatic RNF20 might affect glucose metabolism in vivo. Further studies are required to understand the detailed mechanisms for the role of RNF20 in whole body energy homeostasis.
Recently, it has been demonstrated that SREBP1c is dynamically modified by various posttranslational modifications. For example, SIRT1 promotes the deacetylation-mediated ubiquitination of SREBP1c and represses lipogenic activity during fasting.36,43 Additionally, another study showed that fasting-induced cyclin-dependent kinase 8 (CDK8) phosphorylates and sequentially degrades SREBP1c.44 Although it is unknown whether RNF20-mediated ubiquitination of SREBP1c might be required for prerequisite posttranslational modifications under catabolic conditions, we cannot exclude the possibility that any modification of SREBP1c might change the association between RNF20 and SREBP1c and/or E3 ligase activity of RNF20 during fasting.
In addition to RNF20, FBXW7 is another E3 ubiquitin ligase for SREBP1.21 FBXW7-dependent SREBP1 degradation requires for GSK-3β-mediated phosphorylation of SREBP1.21,35 It is of interest to note that RNF20 would induce the ubiquitination and degradation of SREBP1c upon PKA activation. In contrast, the expression level of FBXW7 was not altered by nutritional states such as feeding and fasting (Fig. 6A). Furthermore, PKA activation did not change the level of FBXW7. Given that PKA plays a crucial role in immediate catabolic responses, PKA activation with forskolin significantly decreased lipogenic gene expression with an increase in RNF20 in primary hepatocytes, whereas suppression of RNF20 reversed the effect of forskolin on lipogenic gene expression (Fig. 6E), indicating that RNF20 might mediate the PKA signaling cascade to down-regulate hepatic lipid metabolism via SREBP1c degradation. Additionally, it has been reported that suppression of FBXW7 in vivo causes fatty liver through the induction of PPARγ rather than SREBP1c.23 On the contrary, RNF20 overexpression did not affect the protein levels of PPARγ and LXRα (Supporting Fig. S7B), implying that RNF20 functions as a “turn-off” switch in hepatic lipogenesis through the regulation of SREBP1c, but not PPARγ, protein stability. Therefore, our data clearly support the hypothesis that RNF20 acts as a negative regulator of SREBP1c and hepatic lipogenesis under catabolic conditions.
Here, we elucidated a novel mechanism of ubiquitination and degradation of SREBP1c by RNF20 during nutritional deprivation. It is plausible to speculate that RNF20 is a suppressor of hepatic lipogenesis through the down-regulation of SREBP1c upon PKA activation. Furthermore, our data provide a clue to understand how SREBP1c is rapidly regulated by fasting signals to prevent excess lipid metabolism. Because there is a positive correlation between lipogenic activity and metabolic complications such as obesity, nonalcoholic fatty liver disease (NAFLD), and certain cancers, it is likely that treatments that activate RNF20 might be useful tools for ameliorating metabolic disorders associated with increased lipid metabolism.
Acknowledgments
We thank Dr. Jae Hong Seol (Seoul National University) for helpful discussions and Yoon Jeong Park for proofreading the article.
Glossary
- ACC1
acetyl-CoA carboxylase1
- db/db
leptin receptor-deficient
- ELOVL6
long-chain fatty acid elongase
- FASN
fatty acid synthase
- FBXW7
F-box and WD repeat domain-containing7
- HMGCR
HMG-CoA reductase
- PKA
protein kinase A
- RNF20
ring finger protein20
- SCD1
steroyl-CoA desaturase1
- SREBP1c
sterol regulatory element binding protein1c
- Ub
ubiquitin
- WT
wild-type
Supporting Information
Additional Supporting Information may be found in the online version of this article at the publisher's website.
References
- 1.Tontonoz P, Kim JB, Graves RA, Spiegelman BM. ADD1: a novel helix-loop-helix transcription factor associated with adipocyte determination and differentiation. Mol Cell Biol. 1993;13:4753–4759. doi: 10.1128/mcb.13.8.4753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Osborne TF. Sterol regulatory element-binding proteins (SREBPs): key regulators of nutritional homeostasis and insulin action. J Biol Chem. 2000;275:32379–32382. doi: 10.1074/jbc.R000017200. [DOI] [PubMed] [Google Scholar]
- 3.Horton JD, Goldstein JL, Brown MS. SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Invest. 2002;109:1125–1131. doi: 10.1172/JCI15593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yang T, Espenshade PJ, Wright ME, Yabe D, Gong Y, Aebersold R, et al. Crucial step in cholesterol homeostasis: sterols promote binding of SCAP to INSIG-1, a membrane protein that facilitates retention of SREBPs in ER. Cell. 2002;110:489–500. doi: 10.1016/s0092-8674(02)00872-3. [DOI] [PubMed] [Google Scholar]
- 5.Briggs MR, Yokoyama C, Wang X, Brown MS, Goldstein JL. Nuclear protein that binds sterol regulatory element of low density lipoprotein receptor promoter. I. Identification of the protein and delineation of its target nucleotide sequence. J Biol Chem. 1993;268:14490–14496. [PubMed] [Google Scholar]
- 6.Wang X, Briggs MR, Hua X, Yokoyama C, Goldstein JL, Brown MS. Nuclear protein that binds sterol regulatory element of low density lipoprotein receptor promoter. II. Purification and characterization. J Biol Chem. 1993;268:14497–14504. [PubMed] [Google Scholar]
- 7.Shimomura I, Shimano H, Horton JD, Goldstein JL, Brown MS. Differential expression of exons 1a and 1c in mRNAs for sterol regulatory element binding protein-1 in human and mouse organs and cultured cells. J Clin Invest. 1997;99:838–845. doi: 10.1172/JCI119247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Kim JB, Spiegelman BM. ADD1/SREBP1 promotes adipocyte differentiation and gene expression linked to fatty acid metabolism. Genes Dev. 1996;10:1096–1107. doi: 10.1101/gad.10.9.1096. [DOI] [PubMed] [Google Scholar]
- 9.Eberle D, Hegarty B, Bossard P, Ferre P, Foufelle F. SREBP transcription factors: master regulators of lipid homeostasis. Biochimie. 2004;86:839–848. doi: 10.1016/j.biochi.2004.09.018. [DOI] [PubMed] [Google Scholar]
- 10.Kumadaki S, Matsuzaka T, Kato T, Yahagi N, Yamamoto T, Okada S, et al. Mouse Elovl-6 promoter is an SREBP target. Biochem Biophys Res Commun. 2008;368:261–266. doi: 10.1016/j.bbrc.2008.01.075. [DOI] [PubMed] [Google Scholar]
- 11.Horton JD, Bashmakov Y, Shimomura I, Shimano H. Regulation of sterol regulatory element binding proteins in livers of fasted and refed mice. Proc Natl Acad Sci U S A. 1998;95:5987–5992. doi: 10.1073/pnas.95.11.5987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kim JB, Sarraf P, Wright M, Yao KM, Mueller E, Solanes G, et al. Nutritional and insulin regulation of fatty acid synthetase and leptin gene expression through ADD1/SREBP1. J Clin Invest. 1998;101:1–9. doi: 10.1172/JCI1411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Foufelle F, Gouhot B, Perdereau D, Girard J, Ferre P. Regulation of lipogenic enzyme and phosphoenolpyruvate carboxykinase gene expression in cultured white adipose tissue. Glucose and insulin effects are antagonized by cAMP. Eur J Biochem. 1994;223:893–900. doi: 10.1111/j.1432-1033.1994.tb19066.x. [DOI] [PubMed] [Google Scholar]
- 14.Girard J, Perdereau D, Foufelle F, Prip-Buus C, Ferre P. Regulation of lipogenic enzyme gene expression by nutrients and hormones. FASEB J. 1994;8:36–42. doi: 10.1096/fasebj.8.1.7905448. [DOI] [PubMed] [Google Scholar]
- 15.Foretz M, Pacot C, Dugail I, Lemarchand P, Guichard C, Le Liepvre X, et al. ADD1/SREBP-1c is required in the activation of hepatic lipogenic gene expression by glucose. Mol Cell Biol. 1999;19:3760–3768. doi: 10.1128/mcb.19.5.3760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang Y, Yin L, Hillgartner FB. SREBP-1 integrates the actions of thyroid hormone, insulin, cAMP, and medium-chain fatty acids on ACCalpha transcription in hepatocytes. J Lipid Res. 2003;44:356–368. doi: 10.1194/jlr.M200283-JLR200. [DOI] [PubMed] [Google Scholar]
- 17.Flier JS, Hollenberg AN. ADD-1 provides major new insight into the mechanism of insulin action. Proc Natl Acad Sci U S A. 1999;96:14191–14192. doi: 10.1073/pnas.96.25.14191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lu M, Shyy JY. Sterol regulatory element-binding protein 1 is negatively modulated by PKA phosphorylation. Am J Physiol Cell Physiol. 2006;290:C1477–1486. doi: 10.1152/ajpcell.00374.2005. [DOI] [PubMed] [Google Scholar]
- 19.Hirano Y, Yoshida M, Shimizu M, Sato R. Direct demonstration of rapid degradation of nuclear sterol regulatory element-binding proteins by the ubiquitin-proteasome pathway. J Biol Chem. 2001;276:36431–36437. doi: 10.1074/jbc.M105200200. [DOI] [PubMed] [Google Scholar]
- 20.Hirano Y, Murata S, Tanaka K, Shimizu M, Sato R. Sterol regulatory element-binding proteins are negatively regulated through SUMO-1 modification independent of the ubiquitin/26 S proteasome pathway. J Biol Chem. 2003;278:16809–16819. doi: 10.1074/jbc.M212448200. [DOI] [PubMed] [Google Scholar]
- 21.Sundqvist A, Bengoechea-Alonso MT, Ye X, Lukiyanchuk V, Jin J, Harper JW, et al. Control of lipid metabolism by phosphorylation-dependent degradation of the SREBP family of transcription factors by SCF(Fbw7) Cell Metab. 2005;1:379–391. doi: 10.1016/j.cmet.2005.04.010. [DOI] [PubMed] [Google Scholar]
- 22.Punga T, Bengoechea-Alonso MT, Ericsson J. Phosphorylation and ubiquitination of the transcription factor sterol regulatory element-binding protein-1 in response to DNA binding. J Biol Chem. 2006;281:25278–25286. doi: 10.1074/jbc.M604983200. [DOI] [PubMed] [Google Scholar]
- 23.Kumadaki S, Karasawa T, Matsuzaka T, Ema M, Nakagawa Y, Nakakuki M, et al. Inhibition of ubiquitin ligase F-box and WD repeat domain-containing 7alpha (Fbw7alpha) causes hepatosteatosis through Kruppel-like factor 5 (KLF5)/peroxisome proliferator-activated receptor gamma2 (PPARgamma2) pathway but not SREBP-1c protein in mice. J Biol Chem. 2011;286:40835–40846. doi: 10.1074/jbc.M111.235283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dentin R, Liu Y, Koo SH, Hedrick S, Vargas T, Heredia J, et al. Insulin modulates gluconeogenesis by inhibition of the coactivator TORC2. Nature. 2007;449:366–369. doi: 10.1038/nature06128. [DOI] [PubMed] [Google Scholar]
- 25.Yamamoto T, Shimano H, Inoue N, Nakagawa Y, Matsuzaka T, Takahashi A, et al. Protein kinase A suppresses sterol regulatory element-binding protein-1C expression via phosphorylation of liver X receptor in the liver. J Biol Chem. 2007;282:11687–11695. doi: 10.1074/jbc.M611911200. [DOI] [PubMed] [Google Scholar]
- 26.Yellaturu CR, Deng X, Cagen LM, Wilcox HG, Park EA, Raghow R, et al. Posttranslational processing of SREBP-1 in rat hepatocytes is regulated by insulin and cAMP. Biochem Biophys Res Commun. 2005;332:174–180. doi: 10.1016/j.bbrc.2005.04.112. [DOI] [PubMed] [Google Scholar]
- 27.Seo JB, Moon HM, Noh MJ, Lee YS, Jeong HW, Yoo EJ, et al. Adipocyte determination- and differentiation-dependent factor 1/sterol regulatory element-binding protein 1c regulates mouse adiponectin expression. J Biol Chem. 2004;279:22108–22117. doi: 10.1074/jbc.M400238200. [DOI] [PubMed] [Google Scholar]
- 28.Amemiya-Kudo M, Shimano H, Yoshikawa T, Yahagi N, Hasty AH, Okazaki H, et al. Promoter analysis of the mouse sterol regulatory element-binding protein-1c gene. J Biol Chem. 2000;275:31078–31085. doi: 10.1074/jbc.M005353200. [DOI] [PubMed] [Google Scholar]
- 29.Dif N, Euthine V, Gonnet E, Laville M, Vidal H, Lefai E. Insulin activates human sterol-regulatory-element-binding protein-1c (SREBP-1c) promoter through SRE motifs. Biochem J. 2006;400:179–188. doi: 10.1042/BJ20060499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jitrapakdee S. Transcription factors and coactivators controlling nutrient and hormonal regulation of hepatic gluconeogenesis. Int J Biochem Cell Biol. 2012;44:33–45. doi: 10.1016/j.biocel.2011.10.001. [DOI] [PubMed] [Google Scholar]
- 31.Li Z, Bowerman S, Heber D. Health ramifications of the obesity epidemic. Surg Clin North Am. 2005;85:681–701, v. doi: 10.1016/j.suc.2005.04.006. [DOI] [PubMed] [Google Scholar]
- 32.Muoio DM, Newgard CB. Obesity-related derangements in metabolic regulation. Annu Rev Biochem. 2006;75:367–401. doi: 10.1146/annurev.biochem.75.103004.142512. [DOI] [PubMed] [Google Scholar]
- 33.Marchesini G, Brizi M, Bianchi G, Tomassetti S, Bugianesi E, Lenzi M, et al. Nonalcoholic fatty liver disease: a feature of the metabolic syndrome. Diabetes. 2001;50:1844–1850. doi: 10.2337/diabetes.50.8.1844. [DOI] [PubMed] [Google Scholar]
- 34.Giandomenico V, Simonsson M, Gronroos E, Ericsson J. Coactivator-dependent acetylation stabilizes members of the SREBP family of transcription factors. Mol Cell Biol. 2003;23:2587–2599. doi: 10.1128/MCB.23.7.2587-2599.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Kim KH, Song MJ, Yoo EJ, Choe SS, Park SD, Kim JB. Regulatory role of glycogen synthase kinase 3 for transcriptional activity of ADD1/SREBP1c. J Biol Chem. 2004;279:51999–52006. doi: 10.1074/jbc.M405522200. [DOI] [PubMed] [Google Scholar]
- 36.Ponugoti B, Kim DH, Xiao Z, Smith Z, Miao J, Zang M, et al. SIRT1 deacetylates and inhibits SREBP-1C activity in regulation of hepatic lipid metabolism. J Biol Chem. 2010;285:33959–33970. doi: 10.1074/jbc.M110.122978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Li Y, Xu S, Mihaylova MM, Zheng B, Hou X, Jiang B, et al. AMPK phosphorylates and inhibits SREBP activity to attenuate hepatic steatosis and atherosclerosis in diet-induced insulin-resistant mice. Cell Metab. 2011;13:376–388. doi: 10.1016/j.cmet.2011.03.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hwang WW, Venkatasubrahmanyam S, Ianculescu AG, Tong A, Boone C, Madhani HD. A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol Cell. 2003;11:261–266. doi: 10.1016/s1097-2765(02)00826-2. [DOI] [PubMed] [Google Scholar]
- 39.Wood A, Krogan NJ, Dover J, Schneider J, Heidt J, Boateng MA, et al. Bre1, an E3 ubiquitin ligase required for recruitment and substrate selection of Rad6 at a promoter. Mol Cell. 2003;11:267–274. doi: 10.1016/s1097-2765(02)00802-x. [DOI] [PubMed] [Google Scholar]
- 40.Kim J, Hake SB, Roeder RG. The human homolog of yeast BRE1 functions as a transcriptional coactivator through direct activator interactions. Mol Cell. 2005;20:759–770. doi: 10.1016/j.molcel.2005.11.012. [DOI] [PubMed] [Google Scholar]
- 41.Shema E, Tirosh I, Aylon Y, Huang J, Ye C, Moskovits N, et al. The histone H2B-specific ubiquitin ligase RNF20/hBRE1 acts as a putative tumor suppressor through selective regulation of gene expression. Genes Dev. 2008;22:2664–2676. doi: 10.1101/gad.1703008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Shema E, Kim J, Roeder RG, Oren M. RNF20 inhibits TFIIS-facilitated transcriptional elongation to suppress pro-oncogenic gene expression. Mol Cell. 2011;42:477–488. doi: 10.1016/j.molcel.2011.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Walker AK, Yang F, Jiang K, Ji JY, Watts JL, Purushotham A, et al. Conserved role of SIRT1 orthologs in fasting-dependent inhibition of the lipid/cholesterol regulator SREBP. Genes Dev. 2010;24:1403–1417. doi: 10.1101/gad.1901210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhao X, Feng D, Wang Q, Abdulla A, Xie XJ, Zhou J, et al. Regulation of lipogenesis by cyclin-dependent kinase 8-mediated control of SREBP-1. J Clin Invest. 2012;122:2417–2427. doi: 10.1172/JCI61462. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional Supporting Information may be found in the online version of this article at the publisher's website.


