The expression, purification, crystallization and preliminary X-ray diffraction analysis of KstR2, a TetR-family transcriptional regulator of cholesterol metabolism from M. tuberculosis, is described.
Keywords: cholesterol, TetR family, Mycobacterium tuberculosis, steroid
Abstract
KstR2 (Rv3557c) is one of two TetR-family transcriptional repressors of cholesterol metabolism in Mycobacterium tuberculosis. The ability to degrade cholesterol fully is important for pathogenesis, and therefore this repressor was expressed, purified and crystallized. Crystals of KstR2 diffracted to better than 1.9 Å resolution and belonged to space group C2, with unit-cell parameters a = 72.3, b = 90.3, c = 49.7 Å, α = γ = 90, β = 128.2°.
1. Introduction
The degradation of host cholesterol is a hallmark of pathogenesis in Mycobacterium tuberculosis (Lee et al., 2013 ▶; Pandey & Sassetti, 2008 ▶). Two TetR-family repressors, KstR and KstR2, control the expression of genes involved in cholesterol transport and degradation (Kendall et al., 2007 ▶, 2010 ▶). In particular, KstR2 regulates 14 genes expected to be involved in the metabolism of the C and D rings of cholesterol (Kendall et al., 2010 ▶; Casabon et al., 2013 ▶). TetR-family repressors typically bind to compounds metabolized by the pathways that they control (Ramos et al., 2005 ▶): KstR2 releases DNA when bound to 3aα-H-4α(3′-propanoyl-CoA)-7aβ-methylhexahydro-1,5-indanedione, a breakdown product of cholesterol (Casabon et al., 2013 ▶).
Determination of the crystal structure of M. tuberculosis KstR2 would allow visualization of its active site and a better understanding of how C and D ring breakdown is regulated in M. tuberculosis. Here, we describe the expression, purification and crystallization of M. tuberculosis KstR2 together with preliminary crystallographic data.
2. Materials and methods
2.1. Expression and purification of KstR2
The M. tuberculosis KstR2 expression construct has been described previously (Kendall et al., 2010 ▶). Escherichia coli BL21(DE3)pLysS cells (Promega) were electroporated with this expression construct, selected on LB–agar containing 20 µg ml−1 kanamycin, pooled and resuspended in auto-induction Terrific Broth (Studier, 2005 ▶) without antibiotic (Agilent Technologies manual). After 24 h of expression at 291 K, the cells were collected by centrifugation and resuspended in lysis buffer (50 mM potassium phosphate pH 8.0, 300 mM NaCl, 30 mM imidazole pH 7.6). DNase (100 µg) and protease inhibitors [one cOmplete Protease Inhibitor Cocktail Mini Tablet (EDTA-free; Roche) per 20 g of cells] were added and the cells were lysed by two freeze–thaw cycles and sonication on ice (Misonix 3000; 12 × 5 s pulses with amplitude control set to 5). The soluble fraction was clarified by centrifugation and diluted sixfold with lysis buffer. KstR2 was immobilized on Ni2+-charged nitrilotriacetic acid resin (Bio-Rad) and washed with lysis buffer containing 60 mM imidazole. Half of the protein eluted with lysis buffer containing 250 mM imidazole; the remainder was released with 500 mM imidazole, possibly reflecting fractions of dimer immobilized by either one or both His6 tags (Lee et al., 2008 ▶). The latter eluate was incubated at 277 K overnight with recombinant TEV protease S219V (Kapust et al., 2001 ▶). KstR2 was further purified by size-exclusion chromatography (SEC) using a Superdex 75 16/60 column (GE Healthcare) into 50 mM HEPES pH 8, 150 mM NaCl, concentrated to 40 mg ml−1 in this buffer using a Vivaspin centrifugal concentrator (10 000 molecular-weight cutoff; GE Healthcare) and stored at 277 K. Size-exclusion chromatography with multi-angle light scattering (SEC-MALS) was performed on a Superdex 75 10/300 GL column (GE Healthcare) in 10 mM Tris pH 8, 150 mM NaCl using a Dionex HPLC with an SLD7000 MALS detector and a Shodex RI-101 differential refractive-index detector. Data were analyzed using the PSS WinGPC UniChrom software.
2.2. Crystallization
Initial crystallization experiments were carried out by sitting-drop vapour diffusion at 291 K using a 480-condition in-house screen (Moreland et al., 2005 ▶) and a Cartesian Honeybee dispensing robot (Genomic Solutions). Diamond-shaped plate crystals grew over three weeks in 14% MPEG 5000, 0.2 M citric acid pH 4.9, and these were reproduced in larger drops (1 µl protein:1 µl precipitant).
2.3. Data collection and processing
Crystals were cryoprotected by the addition of 20% glycerol to the precipitant condition and flash-cooled to 110 K in liquid nitrogen. X-ray diffraction data were collected using a Rigaku MicroMax-007 HF rotating-anode generator fitted with a MAR345 image-plate detector. The crystal-to-detector distance was 300 mm, the oscillation range was 1.0° and 360 images were collected. Data-collection statistics are shown in Table 1 ▶. The data were processed using iMosflm (Battye et al., 2011 ▶), SCALA (Evans, 2006 ▶) and TRUNCATE (French & Wilson, 1978 ▶) in the CCP4 suite (Winn et al., 2011 ▶). The space group was confirmed with POINTLESS (Evans, 2011 ▶).
Table 1. Data-collection statistics for KstR2.
Values in parentheses are for the highest resolution shell.
| Space group | C2 |
| a, b, c () | 72.3, 90.3, 49.7 |
| , , () | 90, 128.2, 90 |
| Mosaicity () | 0.82 |
| Resolution range () | 26.61.86 (1.941.86) |
| Total No. of reflections | 124168 (14882) |
| No. of unique reflections | 19815 (2441) |
| Completeness (%) | 94.3 (80) |
| Multiplicity | 6.3 (6.1) |
| I/(I) | 25.1 (4.5) |
| R merge | 0.046 (0.38) |
| Overall B factor from Wilson plot (2) | 24.7 |
| CC1/2 | 0.999 (0.897) |
3. Results and discussion
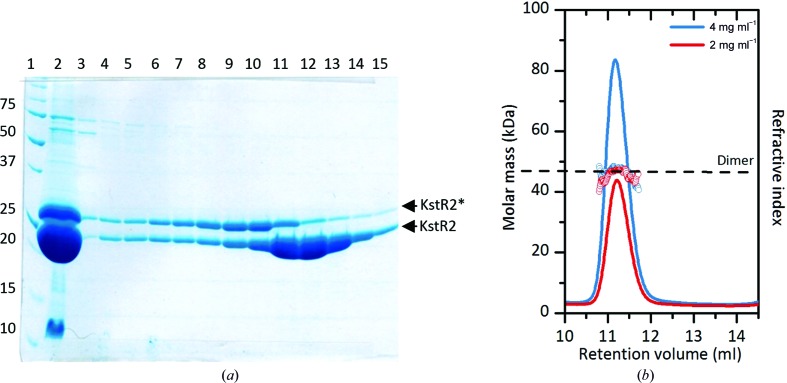
KstR2 was overexpressed in E. coli and purified as described above. The purified protein was estimated to be free from cellular protein based on SDS–PAGE analysis (Fig. 1 ▶ a). SEC-MALS analysis showed a single peak with a measured molecular mass of 47 kDa, compared with a predicted monomer molecular mass of 23.2 kDa (Fig. 1 ▶ b), indicating that KstR2 is an obligate dimer. The crystals of KstR2 belonged to space group C2, and a data set was collected to 1.86 Å resolution. A Matthews coefficient (V M) of 2.27 Å3 Da−1 predicted one molecule per asymmetric unit, with a solvent content of 55.8% (Matthews, 1968 ▶). This suggested that in the crystal the dimer sits on an axis of twofold crystallographic symmetry.
Figure 1.
Size-exclusion chromatography (SEC) of KstR2. (a) SDS–PAGE of SEC fractions. Lane 1, molecular-weight marker (labelled in kDa); lane 2, KstR2 cleaved with rTEV; lanes 3–15, fractions from SEC. Cleaved and uncleaved protein are indicated with arrows; KstR2* = His6 tag + S tag + KstR2. Fractions 11–13 were pooled and concentrated. (b) SEC-MALS analysis of purified KstR2 showing the expected refractive index of the dimer (dotted line) and the measured refractive index of the SEC peak (circles) at two concentrations of KstR2, as well as the elution volume of the peak.
Acknowledgments
Thanks to Chen Gao and Amanda Clinton for preliminary protein expression and analysis. This research was supported by the Health Research Council of New Zealand and the Foundation for Research, Science and Technology of New Zealand.
References
- Battye, T. G. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. W. (2011). Acta Cryst. D67, 271–281. [DOI] [PMC free article] [PubMed]
- Casabon, I., Zhu, S.-H., Otani, H., Liu, J., Mohn, W. W. & Eltis, L. D. (2013). Mol. Microbiol. 89, 1201–1212. [DOI] [PubMed]
- Evans, P. (2006). Acta Cryst. D62, 72–82. [DOI] [PubMed]
- Evans, P. R. (2011). Acta Cryst. D67, 282–292. [DOI] [PMC free article] [PubMed]
- French, S. & Wilson, K. (1978). Acta Cryst. A34, 517–525.
- Kapust, R. B., Tözsér, J., Fox, J. D., Anderson, D. E., Cherry, S., Copeland, T. D. & Waugh, D. S. (2001). Protein Eng. 14, 993–1000. [DOI] [PubMed]
- Kendall, S. L., Burgess, P., Balhana, R., Withers, M., ten Bokum, A., Lott, J. S., Gao, C., Uhia-Castro, I. & Stoker, N. G. (2010). Microbiology, 156, 1362–1371. [DOI] [PMC free article] [PubMed]
- Kendall, S. L., Withers, M., Soffair, C. N., Moreland, N. J., Gurcha, S., Sidders, B., Frita, R., Ten Bokum, A., Besra, G. S., Lott, J. S. & Stoker, N. G. (2007). Mol. Microbiol. 65, 684–699. [DOI] [PMC free article] [PubMed]
- Lee, C.-D., Sun, H.-C., Hu, S.-M., Chiu, C.-F., Homhuan, A., Liang, S.-M., Leng, C.-H. & Wang, T.-F. (2008). Protein Sci. 17, 1241–1248. [DOI] [PMC free article] [PubMed]
- Lee, W., VanderVen, B. C., Fahey, R. J. & Russell, D. G. (2013). J. Biol. Chem. 288, 6788–6800. [DOI] [PMC free article] [PubMed]
- Matthews, B. W. (1968). J. Mol. Biol. 33, 491–497. [DOI] [PubMed]
- Moreland, N., Ashton, R., Baker, H. M., Ivanovic, I., Patterson, S., Arcus, V. L., Baker, E. N. & Lott, J. S. (2005). Acta Cryst. D61, 1378–1385. [DOI] [PubMed]
- Pandey, A. K. & Sassetti, C. M. (2008). Proc. Natl Acad. Sci. USA, 105, 4376–4380. [DOI] [PMC free article] [PubMed]
- Ramos, J. L., Martínez-Bueno, M., Molina-Henares, A. J., Terán, W., Watanabe, K., Zhang, X., Gallegos, M. T., Brennan, R. & Tobes, R. (2005). Microbiol. Mol. Biol. Rev. 69, 326–356. [DOI] [PMC free article] [PubMed]
- Studier, F. (2005). Protein Expr. Purif. 41, 207–234. [DOI] [PubMed]
- Winn, M. D. et al. (2011). Acta Cryst. D67, 235–242.