Abstract
BACKGROUND
Susceptibility to asthma is influenced by genes and environment; implicated genes may indicate pathways for therapeutic intervention. Genetic risk factors may be useful in identifying subtypes of asthma and determining whether intermediate phenotypes, such as elevation of the total serum IgE level, are causally linked to disease.
METHODS
We carried out a genomewide association study by genotyping 10,365 persons with physician-diagnosed asthma and 16,110 unaffected persons, all of whom were matched for ancestry. We used random-effects pooled analysis to test for association in the overall study population and in subgroups of subjects with childhood-onset asthma (defined as asthma developing before 16 years of age), later-onset asthma, severe asthma, and occupational asthma.
RESULTS
We observed associations of genomewide significance between asthma and the following single-nucleotide polymorphisms: rs3771166 on chromosome 2, implicating IL1RL1/IL18R1 (P =3×10−9); rs9273349 on chromosome 6, implicating HLA-DQ (P = 7×10−14); rs1342326 on chromosome 9, flanking IL33 (P = 9×10−10); rs744910 on chromosome 15 in SMAD3 (P = 4×10−9); and rs2284033 on chromosome 22 in IL2RB (P = 1.1×10−8). Association with the ORMDL3/GSDMB locus on chromosome 17q21 was specific to childhood-onset disease (rs2305480, P = 6×10−23). Only HLA-DR showed a significant genomewide association with the total serum IgE concentration, and loci strongly associated with IgE levels were not associated with asthma.
CONCLUSIONS
Asthma is genetically heterogeneous. A few common alleles are associated with disease risk at all ages. Implicated genes suggest a role for communication of epithelial damage to the adaptive immune system and activation of airway inflammation. Variants at the ORMDL3/GSDMB locus are associated only with childhood-onset disease. Elevation of total serum IgE levels has a minor role in the development of asthma. (Funded by the European Commission and others.)
Asthma is a syndrome of unknown origins that is characterized by abnormal and inflamed mucosa of the airways, wheezing, and shortness of breath. In some patients, irreversible airway remodeling and intractable airflow limitation may develop. The disease has a high prevalence and a chronic relapsing course. Although some effective therapies exist for mild asthma, severe asthma remains difficult to treat. The societal cost of the disease is substantial.1,2
Asthma runs strongly in families, and its heritability has been estimated as 60%.3 Genetic studies offer a structured means of understanding the causes of asthma as well as identifying targets that can be used to treat the syndrome.
Childhood asthma is more common in boys than in girls and may persist throughout life. It is often associated with atopy, and as a consequence, much of the research into asthma has focused on atopic mechanisms.4 However, the link between atopic sensitization and asthma symptoms in children is absent in many populations,5 calling into question a causal role of IgE production in the disease. Adult-onset asthma typically develops in middle age and is more common in women than in men. It is less obviously associated with allergy and may be resistant to treatment. Occupational asthma results from workplace exposure to dusts or chemicals and is the most common occupational lung disease in the European Community.
An adequately powered genomewide association study is currently the method of choice for identifying genes that influence complex disease.6 The GABRIEL (A Multidisciplinary Study to Identify the Genetic and Environmental Causes of Asthma in the European Community) Consortium conducted a previous, comparatively small genomewide association study, which showed that multiple markers on chromosome 17q21 were strongly and reproducibly associated with childhood-onset asthma.7,8 We report here the results of a genomewide association study conducted with a sample 10 times the size of that used in our first-generation study. In addition to reporting on analyses of childhood-onset asthma, we report on analyses of later-onset and occupational asthma. Genetic studies can determine whether disease-related phenotypes cause the disease or result from it.9,10 Consequently, we wanted to determine whether there was a concordance between single-nucleotide polymorphisms (SNPs) associated with asthma and SNPs associated with total serum IgE levels. We also wanted to ascertain the extent to which the genetic variants that we observed to be associated with asthma determine the individual risk of asthma development, and we modeled the effects these variants have on the burden of disease in the populations we studied.
METHODS
SUBJECTS
The study consisted of 10,365 case subjects and 16,110 controls recruited from 23 studies (Table 1; and the Supplementary Appendix, available with the full text of this article at NEJM.org). Data on case subjects and population-matched controls were obtained from clinics and from cohort and cross-sectional population surveys in Europe (Table 1 in the Supplementary Appendix). We included some subjects from family-based studies, case subjects and controls from our first-generation genomewide association study,7 and case subjects and controls of European descent from Canadian, Australian, and U.S. surveys. Asthma was considered to be present if it had been diagnosed by a physician. We defined childhood-onset asthma as the presence of the disease in a person younger than 16 years of age and later-onset asthma as disease that developed at 16 years of age or older. Some surveys contributed samples to both age-at-onset groups. Other subgroups consisted of subjects with asthma that developed at an unknown age, subjects with occupational asthma, and subjects with severe asthma (for details, see the Supplementary Appendix). All participants or their parents provided written informed consent for their participation in the study, in accordance with the rules of local ethics committees. The study was performed in accordance with the protocol.
Table 1. Subjects and Populations in Studies of Patients with Asthma Onset in Childhood or Later or with Unknown Age at Onset*.
| Study or Study Group | Subjects with Asthma | Controls | |||||
|---|---|---|---|---|---|---|---|
| Country | Total | Childhood Onset | Later Onset | Total | For Childhood Onset | For Later Onset | |
| number (percent male) | |||||||
| Known age at onset | |||||||
|
| |||||||
| ALSPAC | United Kingdom | 607 | 607 (57) | — | 609 | 609 (51) | — |
|
| |||||||
| B58C | United Kingdom | 445 | 213 (56) | 232 (38) | 420 | 200 (44) | 220 (50) |
|
| |||||||
| BAMSE | Sweden | 239 | 239 (63) | — | 246 | 246 (48) | — |
|
| |||||||
| BUSSELTON | Australia | 398 | 188 (51) | 210 (33) | 809 | 390 (45) | 419 (41) |
|
| |||||||
| CAPPS and SAGE | Canada | 88 | 88 (66) | — | 182 | 182 (51) | — |
|
| |||||||
| ECRHS | Multiple countries | 664 | 279 (56) | 385 (34) | 1546 | 620 (50) | 926 (50) |
|
| |||||||
| EGEA | France | 676 | 482 (61) | 194 (49) | 1252 | 598 (48) | 654 (48) |
|
| |||||||
| GABRIEL Advanced Surveys | Germany | 841 | 841 (63) | — | 851 | 851 (52) | — |
|
| |||||||
| GAIN | Multiple countries | 1475 | 1475 (56) | — | 1668 | 1668 (50) | — |
|
| |||||||
| Karelia Allergy Study | Finland | 73 | 33 (61) | 40 (0) | 75 | 36 (61) | 39 (0) |
|
| |||||||
| KSMU | Russia | 291 | 112 (66) | 179 (39) | 277 | 116 (47) | 161 (52) |
|
| |||||||
| MAGICS | Germany | 630 | 630 (67) | — | 572 | 572 (50) | — |
|
| |||||||
| MAS | Germany | 171 | 171 (57) | — | — | — | — |
|
| |||||||
| MRCA-UKC | United Kingdom | 177 | 177 (62) | — | 399 | 399 (29) | — |
|
| |||||||
| PIAMA | Holland | 172 | 172 (59) | — | 187 | 187 (46) | — |
|
| |||||||
| SAPALDIA | Switzerland | 608 | 237 (62) | 371 (39) | 913 | 356 (52) | 557 (50) |
|
| |||||||
| SLSJ Familial Collection | Canada | 586 | 373 (50) | 213 (34) | 613 | 390 (49) | 223 (43) |
|
| |||||||
| TOMSK | Russia | 259 | 197 (65) | 62 (29) | 422 | 91 (56) | 331 (52) |
|
| |||||||
| UFA | Russia | 330 | 269 (67) | 61 (41) | 348 | 209 (49) | 139 (78) |
|
| |||||||
| Total | 8730 | 6783 | 1947 | 11,389 | 7720 | 3669 | |
|
| |||||||
| Unknown age at onset | |||||||
|
| |||||||
| CAPPS and SAGE | Canada | 179 (46) | 372 (48) | ||||
|
| |||||||
| Occupational asthma | Multiple countries | 529 (83) | 698 (85) | ||||
|
| |||||||
| AUGOSA | United Kingdom | 637 (48) | 2677 (67) | ||||
|
| |||||||
| SEVERE | United Kingdom | 290 (39) | 974 (49) | ||||
|
| |||||||
| Total | 1635 | 4721 | |||||
Data are shown for subjects who were successfully genotyped and whose genotypes passed all tests of quality control. For a list of the complete names of each study or study group, see the Supplementary Appendix.
GENOTYPING
We genotyped 582,892 SNPs for each subject, producing approximately 15 billion genotypes. See the Supplementary Appendix for a description of the handling of DNA and for further details on genotyping.
STATISTICAL ANALYSIS
The study was well powered to detect variants with an allele frequency of 10% and an odds ratio greater than 1.20 or those with an allele frequency of 45% and an odds ratio greater than 1.10. See the Supplementary Appendix for a detailed description of the statistical analyses.
After filtering the genotypes for quality, we controlled for population admixture using principal-components analysis for case subjects and controls in individual populations, removing any outliers. The principal-components analysis for European studies showed good discrimination of country of origin (Fig. 1 in the Supplementary Appendix), including subjects from multicenter studies such as the European Community Respiratory Health Survey. The mean genomic control parameter after principal-components analysis was 1.02 (range, 1.01 to 1.05).
RESULTS
ASSOCIATIONS WITH ASTHMA
We examined the full sample of persons with asthma (including subjects for whom age at onset was unknown and those with occupational asthma or severe asthma) and controls (Fig. 1 and Table 2, and Table 1 in the Supplementary Appendix). The full set of results is available at www.cng.fr/gabriel, and the results and data have been deposited in the European Genome–Phenome Archive (EGA) at the European Bioinformatics Institute (accession number, EGAS00000000077). We also tested for association among subjects with childhood-onset asthma (6783 case subjects and 7720 controls) and those with later-onset asthma (1947 case subjects and 3669 controls) (Table 2 and Fig. 2, and Tables 1 and 2 in the Supplementary Appendix). We then examined loci with genomewide significance in the full sample in the subgroups of subjects with occupational asthma and those with severe asthma (these samples were too small to justify full genomewide analyses) (Table 1 in the Supplementary Appendix). We identified loci with genomewide significance in the overall sample on chromosomes 2, 6, 9, 15, 17, and 22 (Fig. 1 and Table 2, and Table 1 in the Supplementary Appendix).
Figure 1. Manhattan Plot of Study Results.
The results of genomewide association testing are shown for all subjects. The horizontal line indicates the stringent genomewide significance threshold (P≤7.2×10−8).11 Gene names are provided for loci at this level of significance and are also provided for loci at the less stringent threshold of P≤5×10−7 used by the Wellcome Trust Case-Control Consortium.6
Table 2. Loci with Genomewide Significance for Asthma and Tests of Heterogeneity across Studies*.
| Chromosome | Location | Allele | Childhood Onset | Later Onset | Total Sample† | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Gene† | Marker | Position‡ | Ref | Alt | Alt Fq | Odds Ratio (95% CI) | P Value, Random Effects | Odds Ratio (95% CI) | P Value, Random Effects | Odds Ratio (95% CI) | P Value, Random Effects | P Value, Heterogeneity | P Value, Fixed Effects | |
| Loci with genomewide significance — P random ≤7.2×10−8 | ||||||||||||||
|
| ||||||||||||||
| 2 | IL18R1 | rs3771166§ | 102352654 | G | A | 0.38 | 0.85 (0.81–0.90) | 1.1×10−8 | 0.94 (0.85–1.04) | 1.9×10−1 | 0.87 (0.83–0.91) | 3.4×10−9 | 1.8×10−1 | 3.5×10−12 |
|
| ||||||||||||||
| 6 | HLA-DQ | rs9273349 | 32733847 | T | C | 0.58 | 1.14 (1.08–1.22) | 1.9×10−5 | 1.26 (1.16–1.37) | 3.9×10−8 | 1.18 (1.13–1.24) | 7.0×10−14 | 5.0×10−1 | 7.0×10−14 |
|
| ||||||||||||||
| 9 | IL33 | rs1342326 | 6180076 | A | C | 0.16 | 1.27 (1.17–1.38) | 1.6×10−8 | 1.12 (0.99–1.26) | 6.8×10−2 | 1.20 (1.13–1.28) | 9.2×10−10 | 2.2×10−1 | 8.7×10−12 |
|
| ||||||||||||||
| 15 | SMAD3 | rs744910 | 65233839 | G | A | 0.51 | 0.89 (0.84–0.93) | 8.1×10−6 | 0.94 (0.87–1.0) | 1.4×10−1 | 0.89 (0.86–0.92) | 3.9×10−9 | 8.5×10−1 | 3.9×10−9 |
|
| ||||||||||||||
| 17 | GSDMB | rs2305480 | 35315722 | G | A | 0.45 | 0.76 (0.72–0.81) | 6.4×10−23 | 1.03 (0.94–1.13) | 4.9×10−1 | 0.85 (0.81–0.90) | 9.6×10−8 | 9.2×10−4 | — |
|
| ||||||||||||||
| 17 | GSDMA | rs3894194 | 35375519 | G | A | 0.45 | 1.26 (1.19–1.33) | 3.0×10−17 | 1.02 (0.94–1.11) | 6.0×10−1 | 1.17 (1.11–1.23) | 4.6×10−9 | 2.0×10−2 | — |
|
| ||||||||||||||
| 22 | IL2RB | rs2284033 | 35863980 | G | A | 0.44 | 0.92 (0.87–0.97) | 1.6×10−3 | 0.86 (0.80–0.94) | 4.2×10−4 | 0.89 (0.86–0.93) | 1.2×10−8 | 9.2×10−1 | 1.2×10−8 |
|
| ||||||||||||||
| Additional loci — P random ≤5×10−7 | ||||||||||||||
|
| ||||||||||||||
| 5 | SLC22A5 | rs2073643 | 131751187 | T | C | 0.55 | 0.89 (0.84–0.93) | 7.6×10−6 | 0.94 (0.87–1.0) | 1.5×10−1 | 0.90 (0.87–0.94) | 2.2×10−7 | 8.3×10−1 | 2.2×10−7 |
|
| ||||||||||||||
| 5 | IL13 | rs1295686 | 132023742 | T | C | 0.80 | 0.85 (0.79–0.90) | 3.3×10−7 | 0.94 (0.85–1.04) | 2.6×10−1 | 0.87 (0.83–0.92) | 1.4×10−7 | 3.3×10−1 | 1.4×10−8 |
|
| ||||||||||||||
| 15 | RORA | rs11071559 | 58857280 | C | T | 0.14 | 0.88 (0.81–0.95) | 1.0×10−3 | 0.78 (0.69–0.88) | 5.7×10−5 | 0.85 (0.80–0.90) | 1.1×10−7 | 8.4×10−1 | 1.1×10−7 |
Markers are identified according to their standard rs numbers. Only single-nucleotide polymorphisms (SNPs) with significant or independent effects (according to stepwise regression analysis) on the risk of asthma are shown. The total sample includes all case subjects and controls. A complete list of SNPs with genomewide significance is available in Table 2 in the Supplementary Appendix. P values are shown for tests of association under a random-effects model, tests of heterogeneity across studies with the use of Cochran’s Q test, and tests of association under a fixed-effects model when there was no significant evidence of heterogeneity. For the calculation of these odds ratios, alternate alleles (Alt) were designated as risk alleles. Alt Fq denotes alternative allele frequency, CI confidence interval, and Ref reference allele.
The nominal gene was taken from the Illumina SNP database.
Position is based on the number of base pairs from the start of the chromosome.
The rs3771166 SNP was imputed in the MRC Asthma and United Kingdom Control panel (MRCA-UKC) and in the Multicenter Asthma Genetics in Childhood Study (MAGICS) panel.
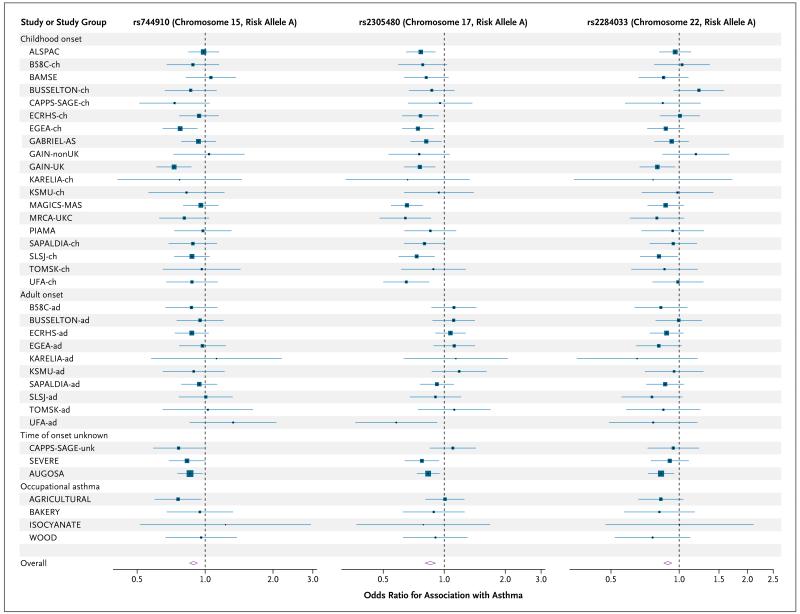
Figure 2. Forest Plots of Odds Ratios for Markers with Genomewide Association with Asthma According to Study Subgroup.
The odds ratios and 95% confidence intervals for six loci show independent effects on asthma risk. For the calculation of these odds ratios, alternate alleles were designated as risk alleles. In order to include the subjects from our original genome screen, rs3771166 was imputed for subjects from the MRCA-UKC and MAGICS panels. Sample size is indicated by the size of the square at the center of each confidence interval. For a list of the complete names of each study or study group, see the Supplementary Appendix.
The strongest association on chromosome 2 was with a SNP within the IL18R1 gene (rs3771166) (P = 3.4×10−9; odds ratio, 0.87). IL18R1 is in close proximity to IL1RL1, which also contained SNPs showing significant association with asthma (Table 1 in the Supplementary Appendix); these SNPS were in high linkage disequilibrium with rs3771166 (r2 = 0.96 for the correlation between rs3771166 and each of the three most significant SNPs on the IL1RL1 gene). A locus centered on rs9273349 in the HLA-DQ region of the major histocompatibility complex gene (MHC) also showed strong evidence of association in the overall sample (P = 7×10−14; odds ratio, 1.18). We observed significant association between disease and rs1342326 (on chromosome 9), flanking the IL33 gene (P = 9×10−10; odds ratio, 1.20) (Table 2). Other SNPs showing significant association with disease were rs744910 on chromosome 15 (P = 4×10−9; odds ratio, 0.89), within the SMAD3 gene, and rs2284033, within IL2RB on chromosome 22 (P = 1×10−8; odds ratio, 0.89) (Table 2).
The odds ratios for these loci generally suggested more pronounced effects in childhood-onset asthma than in later-onset asthma. The HLA-DQ region was observed to have a slightly stronger association with later-onset disease (P = 4×10−8; odds ratio, 1.26) than with childhood-onset disease (P = 2×10−5; odds ratio, 1.14). However, none of these differences were significant (Table 2 and Fig. 2, and Table 2 in the Supplementary Appendix).
The span of the SNPs associated with disease at the ORMDL3/GSDMB locus on chromosome 17q12 was approximately 380 kb (Table 2, and Table 1 in the Supplementary Appendix). As in previous studies,7,8 we observed for 17q12 SNPs significant evidence of heterogeneity between associations with childhood-onset and disease and associations with significant later-onset disease (P<0.001) (Fig. 2). Significant associations were confined to childhood-onset disease (Table 1 in the Supplementary Appendix), and a separate analysis of subjects with childhood-onset disease (and controls) yielded a maximum association at rs2305480 within the GSDMB gene (P = 6×10−23; odds ratio, 0.76) (Table 2). Among subjects with childhood-onset disease, forward stepwise regression established an independent association with rs3894194 within GSDMA (P = 0.0015), a member of the same gene family as GSDMB, in addition to rs2305480. We found no evidence of stronger associations with haplotypes containing either or both of these two SNPs.
Using a less stringent threshold for significance (P<5×10−7), we observed association between disease and rs2073643 near SLC22A5 on chromosome 5 (P = 2×10−7; odds ratio, 0.90) (Table 2). This locus is approximately 273 kb upstream of — and is independent of — the previously established IL13 locus,12 which contains a SNP (rs1295686) for which there was also evidence of association (P = 1×10−7) (Table 2). We also observed association with rs11071559 within RORA on chromosome 15 (P = 1×10−7; odds ratio, 0.85) (Table 2).
We did not observe any significant associations that were particular to the groups with severe or occupational asthma. We did find a suggestive association with rs1837253 on chromosome 5, situated within the TSLP gene, among patients with severe asthma in the SEVERE study (P = 3×10−6; odds ratio, 0.56), but this association was not replicated among patients with severe asthma in the AUGOSA (Asthma UK Genetics of Severe Asthma) study.
Logistic-regression models for interactions between loci showed effects of only marginal significance (P = 0.01) before correction for multiple comparisons; these findings are thus unlikely to be of biologic importance. We found no evidence of dominance, or interactions with sex, for any of the loci.
ASSOCIATIONS WITH TOTAL SERUM IgE
We tested for association with the natural logarithm of total serum IgE levels in 7087 subjects with asthma and 7667 controls, using random-effects pooled estimates (Table 3, and Fig. 2 in the Supplementary Appendix), and identified one novel locus within the class II region of MHC (P = 8×10−15). We also confirmed previously observed associations with markers at loci housing FCER1A, IL13, STAT6, and IL4R/IL21R13 (Table 3, and Fig. 2 in the Supplementary Appendix).
The IgE-related IL13 and MHC loci showed evidence of association with asthma, albeit not at the level of genomewide significance (Table 3). Moreover, mapping of the MHC loci associated with elevated IgE levels and with asthma showed that the principal genetic effects were independent of one another — near HLA-DRB1 for elevated IgE levels and near HLA-DQB1 for asthma. Linkage disequilibrium was weak between the trait-associated markers at the two MHC loci. The HLA-DQB1 markers associated with asthma remained significantly associated with asthma after adjustment for IgE level. SNPs at the FCER1A, STAT6, and IL4R/IL21R loci were not associated with asthma.
Table 3. Peak Associations with Total Serum IgE Compared with Peak Associations with Asthma*.
| Chromosome | Gene† | Marker | Position‡ | P Value, Random Effects§ | |
|---|---|---|---|---|---|
| IgE | Asthma | ||||
| Loci linked to serum IgE | |||||
|
| |||||
| 1 | FCER1A | rs2252226 | 157542777 | 6.6×10−5 | 4.7×10−1 |
|
| |||||
| 5 | IL13 | rs20541 | 132023863 | 1.0×10−6 | 2.8×10−7 |
|
| |||||
| 6 | HLA-DRB1 | rs9271300 | 32689560 | 8.3×10−15 | 3.2×10−1 |
|
| |||||
| 12 | STAT6 | rs167769 | 55790042 | 8.5×10−7 | 7.2×10−4 |
|
| |||||
| 16 | IL4-R/IL21R | rs1859308 | 27305499 | 8.2×10−6 | 3.5×10−1 |
|
| |||||
| Loci linked to asthma | |||||
|
| |||||
| 2 | IL18R1 | rs3771166 | 102352654 | 2.0×10−1 | 3.4×10−9 |
|
| |||||
| 6 | HLA-DQB1 | rs9273349 | 32733847 | 5.4×10−5 | 7.0×10−14 |
|
| |||||
| 9 | IL33 | rs1342326 | 6180076 | 6.0×10−1 | 9.2×10−10 |
|
| |||||
| 15 | SMAD3 | rs744910 | 65233839 | 1.5×10−1 | 3.9×10−9 |
|
| |||||
| 17 | GSDMB | rs2305480 | 35315722 | 6.0×10−2 | 9.6×10−8 |
|
| |||||
| 17 | GSDM1 | rs3894194 | 35375519 | 3.0×10−1 | 4.6×10−9 |
|
| |||||
| 22 | IL2RB | rs2284033 | 35863980 | 3.9×10−1 | 1.1×10−8 |
|
| |||||
| 5 | IL13 | rs1295686 | 132023742 | 2.4×10−6 | 1.4×10−7 |
|
| |||||
| 5 | SLC22A5 | rs2073643 | 131751187 | 2.0×10−2 | 2.2×10−7 |
|
| |||||
| 15 | RORA | rs11071559 | 58857280 | 3.0×10−2 | 1.1×10−7 |
Two sets of loci are shown: the most significant hits that show association with the total serum IgE level (on the basis of genomewide-association-study significance in the current study or previously published studies) and those that show association with asthma in the current study.
The nominal gene was taken from the Illumina SNP database.
Position is based on the number of base pairs from the start of the chromosome.
P values are shown for the association of marker with phenotype in a random effects model.
POPULATION BURDEN AND INDIVIDUAL RISK
For each of the seven SNPs associated with childhood asthma, we assessed the individual risk in a classification analysis, using a cutoff point of 0.5 (i.e., subjects were classified as having asthma if the predicted probability of disease was ≥0.5). The sensitivity for classifying subjects as having or not having asthma was 35%, and the specificity 75%. The area under the receiver-operating-characteristic curve was 0.58 (on a scale of 0.50 to 1.00).
Estimation of the population attributable risk fraction for the joint action of the significant loci indicated that 38% (95% confidence interval [CI], 28 to 44) of cases of childhood-onset asthma were attributable to SNP combinations at these loci (representing a higher risk than that for the controls with the lowest 10% of the risk score). In an independent population-based replication sample of 517 case subjects and 3486 controls, drawn from members of the British 1958 birth cohort who were not included in the current study (details are provided at www.b58cgene.sgul.ac.uk), the same SNPs accounted for 49% (95% CI, 27 to 65) of the lifetime asthma risk (from birth to middle age). These results indicate that although the SNPs would not be effective in classifying the individual risk of asthma, the SNPs (or rather, the downstream functions of which they are markers) do have a substantial effect on the burden of asthma in the community.
We tested for significant associations with loci previously reported in studies of persons of European ancestry with asthma (including PDE4D,14 CHI3L1,15 DPP10,16 GPR154 [NPSR1],17 ADAM33,18 PHF11,19 OPN3,20 IRAK3,21 PCDH1,22 HLA-G,23 and DENND1B24) (Table 3 in the Supplementary Appendix). We observed associations with SNPs implicating DPP10 (P≤3×10−3), PDE4D (P≤4.7×10−3), GPR154 (P≤1×10−3), and HLA-G (P≤2×10−4), but none of these associations were significant after correction for multiple comparisons. It has been suggested that genetic determinants of lung function influence susceptibility to asthma25,26; our data do not support this hypothesis (Table 4 in the Supplementary Appendix).
DISCUSSION
Our study identifies candidate genes in a pathway that initiates type 2 helper T-cell (Th2) inflammation in response to epithelial damage and points to other candidate genes that may act in a pathway that down-regulates airway inflammation and remodeling. Our study also shows that asthma is heterogeneous: later-onset cases are influenced more by the MHC than are childhood-onset cases, and our findings confirm a strong and specific effect of the chromosome 17q locus on childhood-onset disease.7,8 Moreover, our results show that genetics cannot easily be used to determine the individual risk of asthma.
SNPs at the chromosome 17q21 locus associated with asthma are also strongly associated with variation in the expression of ORMDL37 and GSDMB,27,28 meaning that it is feasible that the locus contributes to asthma through the differential regulation of both genes. Changes in the expression of the homologous ORM gene in yeast cause dysregulation of sphingolipid metabolism.29 Evidence that similar pathways are involved in humans would provide a potential mechanism (and therapeutic target) for the modulation of airway inflammation.30
The association between SNPs flanking IL33 on chromosome 9 and atopic asthma has been reported previously.31 IL33 was originally detected in airway epithelial cells.32 It activates nuclear factor κB (NF-κB) and mitogen-activated protein (MAP) kinases and drives production of Th2-associated cytokines32 such as interleukin-4, interleukin-5, and interleukin-13. It is detected in inflamed tissues.33 IL33 is constitutively expressed in endothelial and epithelial cells and may function as an endogenous danger signal (an “alarm-in”) to alert the immune system of epithelial damage during trauma or infection.34
The locus at chromosome 2, implicating IL1RL1 and IL18R1, has also previously been shown to be associated with asthma.31 The effect at this locus has been attributed to IL1RL1 (encoding the receptor for interleukin-33), but we were not able to dissect the association to the extent of being able to clearly implicate IL1RL1 over IL18R1. IL18 is closely related to IL3332 and synergizes with IL12 to induce the production of interferon-γ and to promote Th1 responses.35 The expression of IL18R1 is also concentrated within the respiratory epithelium.
SMAD3 is a transcriptional modulator activated by transforming growth factor β, a polypeptide that controls proliferation, differentiation, and other functions in many cell types, including regulatory T cells.36 Mice deficient in Smad3 have increased levels of proinflammatory cytokines in their lungs.37
IL2 controls the survival and proliferation of regulatory T cells and is implicated in the differentiation and homeostasis of effector T-cell subgroups, including Th1, Th2, Th17, and memory CD8+ T cells. 38 The interleukin-2 receptor is composed of α (CD25), β (CD122), and γ chains.38 The β chain (interleukin-2 receptor β [RB]) identified in our study is a signal-transduction element that is also present in the interleukin-15 receptor. Thus, the IL33 and IL18R1 loci may modify the inflammatory response to epithelial damage, whereas SMAD3 and IL2RB may regulate homeostatic and healing pathways.
HLA-DQ was the first asthma-susceptibility locus to be identified,39 and extended haplotypes encompassing HLA-DQ and HLA-DR have been studied for their effects on specific allergen sensitization40,41 and on the formation of tumor necrosis factor and related gene products.42 Our results indicate that HLA-DQ may be fundamental to later-onset disease. Adult-onset asthma is thought to have a minor atopic component; HLA-DQ may restrict the response to bacterial or other antigens that do not act as classical allergens in these patients.
Suggestive data point to two other genes, SLC22A5 and RORA. SLC22A5 encodes a carnitine transporter and, like ORMDL3/GSDMB and IL18R1/IL1RL1, sports variants that are associated with Crohn’s disease.43 Asthma and Crohn’s disease may therefore have shared mechanisms, perhaps involving a modulation of microbial interactions with the mucosa. RORA encodes a member of the NR1 subfamily of nuclear hormone receptors. It is expressed at high levels in keratinocytes, together with a cluster of genes that form the structural and innate immune defenses of the epithelial barrier.44
We found little overlap between the principal loci that confer susceptibility to asthma and those that regulate total serum IgE levels, with the exception of IL13 and the HLA region (which contains multiple loci affecting these phenotypes). This suggests that elevation of the IgE level is probably an inconstant secondary effect of asthma rather than its cause, a conclusion that is consistent with the absence of a relationship between atopic sensitization and asthma in many populations.5 Genetic studies of children with atopic dermatitis have shown that defects in barrier proteins such as Flaggrin (FLG)45 and the serine protease inhibitor SPINK546 commonly confer a predisposition to the disease, indicating that increases in IgE levels may be the result of barrier failure.
Many loci have been reported to have an association with asthma47; most of these associations were not significant in our well-powered study. The relative risks observed with our markers will not be useful as predictors of disease in individual patients, no doubt because of variable environmental effects. These loci nevertheless contribute substantially to the asthma burden in the general population. Further studies are needed to identify the specific genetic variants that contribute to disease and to establish their biologic function. Such studies may help to identify the causes of airway epithelial damage that initiate the asthmatic process. In the mean time, ORMDL3 and GSDMB, IL33 and IL1RL1, IL18R1, SMAD3, and IL2RB already provide a focus in the search for more effective therapies for asthma.
Supplementary Material
Acknowledgments
Supported by a contract from the European Commission (018996) and grants from the French Ministry of Research, the Wellcome Trust (WT084703MA), and Asthma UK.
We thank Dana Finch, GABRIEL project manager, for her administrative support of the study and the GABRIEL Consortium.
Footnotes
Disclosure forms provided by the authors are available with the full text of this article at NEJM.org.
References
- 1.Smith DH, Malone DC, Lawson KA, Okamoto LJ, Battista C, Saunders WB. A national estimate of the economic costs of asthma. Am J Respir Crit Care Med. 1997;156:787–93. doi: 10.1164/ajrccm.156.3.9611072. [DOI] [PubMed] [Google Scholar]
- 2.Asthma agenda. National Asthma Campaign; London: 1998. [Google Scholar]
- 3.Duffy DL, Martin NG, Battistutta D, Hopper JL, Mathews JD. Genetics of asthma and hay fever in Australian twins. Am Rev Respir Dis. 1990;142:1351–8. doi: 10.1164/ajrccm/142.6_Pt_1.1351. [DOI] [PubMed] [Google Scholar]
- 4.Kay AB. Allergy and allergic diseases: first of two parts. N Engl J Med. 2001;344:30–7. doi: 10.1056/NEJM200101043440106. [DOI] [PubMed] [Google Scholar]
- 5.Weinmayr G, Weiland SK, Björkstén B, et al. Atopic sensitization and the international variation of asthma symptom prevalence in children. Am J Respir Crit Care Med. 2007;176:565–74. doi: 10.1164/rccm.200607-994OC. [DOI] [PubMed] [Google Scholar]
- 6.Wellcome Trust Case Control Consortium Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447:661–78. doi: 10.1038/nature05911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Moffatt MF, Kabesch M, Liang L, et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature. 2007;448:470–3. doi: 10.1038/nature06014. [DOI] [PubMed] [Google Scholar]
- 8.Bouzigon E, Corda E, Aschard H, et al. Effect of 17q21 variants and smoking exposure in early-onset asthma. N Engl J Med. 2008;359:1985–94. doi: 10.1056/NEJMoa0806604. [DOI] [PubMed] [Google Scholar]
- 9.Sheehan NA, Didelez V, Burton PR, Tobin MD. Mendelian randomisation and causal inference in observational epidemiology. PLoS Med. 2008;5(8):e177. doi: 10.1371/journal.pmed.0050177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Elliott P, Chambers JC, Zhang W, et al. Genetic loci associated with C-reactive protein levels and risk of coronary heart disease. JAMA. 2009;302:37–48. doi: 10.1001/jama.2009.954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dudbridge F, Gusnanto A. Estimation of significance thresholds for genome wide association scans. Genet Epidemiol. 2008;32:227–34. doi: 10.1002/gepi.20297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Konstantinidis AK, Barton SJ, Sayers I, et al. Genetic association studies of interleukin-13 receptor alpha1 subunit gene polymorphisms in asthma and atopy. Eur Respir J. 2007;30:40–7. doi: 10.1183/09031936.00025706. [DOI] [PubMed] [Google Scholar]
- 13.Weidinger S, Gieger C, Rodriguez E, et al. Genome-wide scan on total serum IgE levels identifies FCER1A as novel susceptibility locus. PLoS Genet. 2008;4(8):e1000166. doi: 10.1371/journal.pgen.1000166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Himes BE, Hunninghake GM, Baurley JW, et al. Genome-wide association analysis identifies PDE4D as an asthma-susceptibility gene. Am J Hum Genet. 2009;84:581–93. doi: 10.1016/j.ajhg.2009.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ober C, Tan Z, Sun Y, et al. Effect of variation in CHI3L1 on serum YKL-40 level, risk of asthma, and lung function. N Engl J Med. 2008;358:1682–91. doi: 10.1056/NEJMoa0708801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Allen M, Heinzmann A, Noguchi E, et al. Positional cloning of a novel gene influencing asthma from chromosome 2q14. Nat Genet. 2003;35:258–63. doi: 10.1038/ng1256. [DOI] [PubMed] [Google Scholar]
- 17.Laitinen T, Polvi A, Rydman P, et al. Characterization of a common susceptibility locus for asthma-related traits. Science. 2004;304:300–4. doi: 10.1126/science.1090010. [DOI] [PubMed] [Google Scholar]
- 18.Van Eerdewegh P, Little RD, Dupuis J, et al. Association of the ADAM33 gene with asthma and bronchial hyperresponsiveness. Nature. 2002;418:426–30. doi: 10.1038/nature00878. [DOI] [PubMed] [Google Scholar]
- 19.Zhang Y, Leaves NI, Anderson GG, et al. Positional cloning of a quantitative trait locus on chromosome 13q14 that influences immunoglobulin E levels and asthma. Nat Genet. 2003;34:181–6. doi: 10.1038/ng1166. [DOI] [PubMed] [Google Scholar]
- 20.White JH, Chiano M, Wigglesworth M, et al. Identification of a novel asthma susceptibility gene on chromosome 1qter and its functional evaluation. Hum Mol Genet. 2008;17:1890–903. doi: 10.1093/hmg/ddn087. [DOI] [PubMed] [Google Scholar]
- 21.Balaci L, Spada MC, Olla N, et al. IRAK-M is involved in the pathogenesis of early-onset persistent asthma. Am J Hum Genet. 2007;80:1103–14. doi: 10.1086/518259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Koppelman GH, Meyers DA, Howard TD, et al. Identification of PCDH1 as a novel susceptibility gene for bronchial hyperresponsiveness. Am J Respir Crit Care Med. 2009;180:929–35. doi: 10.1164/rccm.200810-1621OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nicolae D, Cox NJ, Lester LA, et al. Fine mapping and positional candidate studies identify HLA-G as an asthma susceptibility gene on chromosome 6p21. Am J Hum Genet. 2005;76:349–57. doi: 10.1086/427763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Sleiman PM, Flory J, Imielinski M, et al. Variants of DENND1B associated with asthma in children. N Engl J Med. 2010;362:36–44. doi: 10.1056/NEJMoa0901867. [DOI] [PubMed] [Google Scholar]
- 25.Repapi E, Sayers I, Wain LV, et al. Genome-wide association study identifies five loci associated with lung function. Nat Genet. 2010;42:36–44. doi: 10.1038/ng.501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hancock DB, Eijgelsheim M, Wilk JB, et al. Meta-analyses of genome-wide association studies identify multiple loci associated with pulmonary function. Nat Genet. 2010;42:45–52. doi: 10.1038/ng.500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Cookson W, Liang L, Abecasis G, Moffatt M, Lathrop M. Mapping complex disease traits with global gene expression. Nat Rev Genet. 2009;10:184–94. doi: 10.1038/nrg2537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Verlaan DJ, Berlivet S, Hunninghake GM, et al. Allele-specific chromatin remodeling in the ZPBP2/GSDMB/ORMDL3 locus associated with the risk of asthma and autoimmune disease. Am J Hum Genet. 2009;85:377–93. doi: 10.1016/j.ajhg.2009.08.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Breslow DK, Collins SR, Bodenmiller B, et al. Orm family proteins mediate sphingolipid homeostasis. Nature. 2010;463:1048–53. doi: 10.1038/nature08787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nixon GF. Sphingolipids in inflammation: pathological implications and potential therapeutic targets. Br J Pharmacol. 2009;158:982–93. doi: 10.1111/j.1476-5381.2009.00281.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Gudbjartsson DF, Bjornsdottir US, Halapi E, et al. Sequence variants affecting eosinophil numbers associate with asthma and myocardial infarction. Nat Genet. 2009;41:342–7. doi: 10.1038/ng.323. [DOI] [PubMed] [Google Scholar]
- 32.Schmitz J, Owyang A, Oldham E, et al. IL-33, an interleukin-1-like cytokine that signals via the IL-1 receptor-related protein ST2 and induces T helper type 2-associated cytokines. Immunity. 2005;23:479–90. doi: 10.1016/j.immuni.2005.09.015. [DOI] [PubMed] [Google Scholar]
- 33.Carriere V, Roussel L, Ortega N, et al. IL-33, the IL-1-like cytokine ligand for ST2 receptor, is a chromatin-associated nuclear factor in vivo. Proc Natl Acad Sci U S A. 2007;104:282–7. doi: 10.1073/pnas.0606854104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Moussion C, Ortega N, Girard JP. The IL-1-like cytokine IL-33 is constitutively expressed in the nucleus of endothelial cells and epithelial cells in vivo: a novel ‘alarmin’? PLoS ONE. 2008;3(10):e3331. doi: 10.1371/journal.pone.0003331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fukao T, Matsuda S, Koyasu S. Synergistic effects of IL-4 and IL-18 on IL-12-dependent IFN-gamma production by dendritic cells. J Immunol. 2000;164:64–71. doi: 10.4049/jimmunol.164.1.64. [DOI] [PubMed] [Google Scholar]
- 36.Nouri-Aria KT, Durham SR. Regulatory T cells and allergic disease. Inflamm Allergy Drug Targets. 2008;7:237–52. doi: 10.2174/187152808786848405. [DOI] [PubMed] [Google Scholar]
- 37.Anthoni M, Wang G, Leino MS, Lauerma AI, Alenius HT, Wolff HJ. Smad3 -signalling and Th2 cytokines in normal mouse airways and in a mouse model of asthma. Int J Biol Sci. 2007;3:477–85. doi: 10.7150/ijbs.3.477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Létourneau S, Krieg C, Pantaleo G, Boyman O. IL-2- and CD25-dependent immunoregulatory mechanisms in the homeostasis of T-cell subsets. J Allergy Clin Immunol. 2009;123:758–62. doi: 10.1016/j.jaci.2009.02.011. [DOI] [PubMed] [Google Scholar]
- 39.Marsh DG, Meyers DA, Bias WB. The epidemiology and genetics of atopic allergy. N Engl J Med. 1981;305:1551–9. doi: 10.1056/NEJM198112243052603. [DOI] [PubMed] [Google Scholar]
- 40.Marsh DG, Blumenthal MN, Ishikawa T, et al. HLA and specific immune responsiveness to allergens. In: Tsuji K, Aizawa M, Sazazuki T, editors. Proceedings of the Eleventh International Histocompatibility Workshop and conference. Oxford University Press; Oxford, England: 1992. pp. 765–7. [Google Scholar]
- 41.Moffatt MF, Schou C, Faux JA, et al. Association between quantitative traits underlying asthma and the HLA-DRB1 locus in a family-based population sample. Eur J Hum Genet. 2001;9:341–6. doi: 10.1038/sj.ejhg.5200636. [DOI] [PubMed] [Google Scholar]
- 42.Moffatt MF, Cookson WO. Tumour necrosis factor haplotypes and asthma. Hum Mol Genet. 1997;6:551–4. doi: 10.1093/hmg/6.4.551. [DOI] [PubMed] [Google Scholar]
- 43.Barrett JC, Hansoul S, Nicolae DL, et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn’s disease. Nat Genet. 2008;40:955–62. doi: 10.1038/NG.175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Taylor JM, Street TL, Hao L, et al. Dynamic and physical clustering of gene expression during epidermal barrier formation in differentiating keratinocytes. PLoS ONE. 2009;4(10):e7651. doi: 10.1371/journal.pone.0007651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Palmer CN, Irvine AD, Terron-Kwiatkowski A, et al. Common loss-of-function variants of the epidermal barrier protein filaggrin are a major predisposing factor for atopic dermatitis. Nat Genet. 2006;38:441–6. doi: 10.1038/ng1767. [DOI] [PubMed] [Google Scholar]
- 46.Walley AJ, Chavanas S, Moffatt MF, et al. Gene polymorphism in Netherton and common atopic disease. Nat Genet. 2001;29:175–8. doi: 10.1038/ng728. [DOI] [PubMed] [Google Scholar]
- 47.Vercelli D. Advances in asthma and allergy genetics in 2007. J Allergy Clin Immunol. 2008;122:267–71. doi: 10.1016/j.jaci.2008.06.008. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.