Abstract
We have determined the genomic map of the bacterium Salmonella typhi Ty2, the causal organism of typhoid fever, by using pulsed-field gel electrophoresis. Digestion of the Ty2 genome with endonucleases Xba I, Bln I, and Ceu I yielded 33, 26, and 7 fragments, respectively, that were placed in order on a circular chromosome of 4780 kb. Transposon Tn10 was inserted in specific genes of Salmonella typhimurium and transduced into S. typhi, and thus, the positions of 37 S. typhi genes were located through the Xba I and Bln I sites of the Tn10. Gene order on chromosomes of Escherichia coli K-12 and S. typhimurium LT2 is remarkably conserved; however, the gene order in S. typhi Ty2 is different, suggesting it has undergone major genomic rearrangements during its evolution. These rearrangements include inversions and transpositions in the 7 DNA fragments between the seven rrn operons for rRNA (postulated to be due to homologous recombination in these rrn genes), another inversion that covers the replication terminus region (resembling inversions found in other enteric bacteria), and at least three insertions, one as large as 118 kb. Partial digestion of genomic DNA with the intron-encoded endonuclease I-Ceu I, which cuts only in rrn genes, shows chromosomal rearrangements, apparently due to homologous recombination in the rrn genes, that were detected in all wild-type strains of S. typhi tested. These rearrangements may have been selected to compensate for the insertions that otherwise would have altered the locations of genes with respect to the origin and terminus of replication. These observations are relevant to our view of the evolution of the bacterial genome and may be significant in the virulence of S. typhi.
Full text
PDF
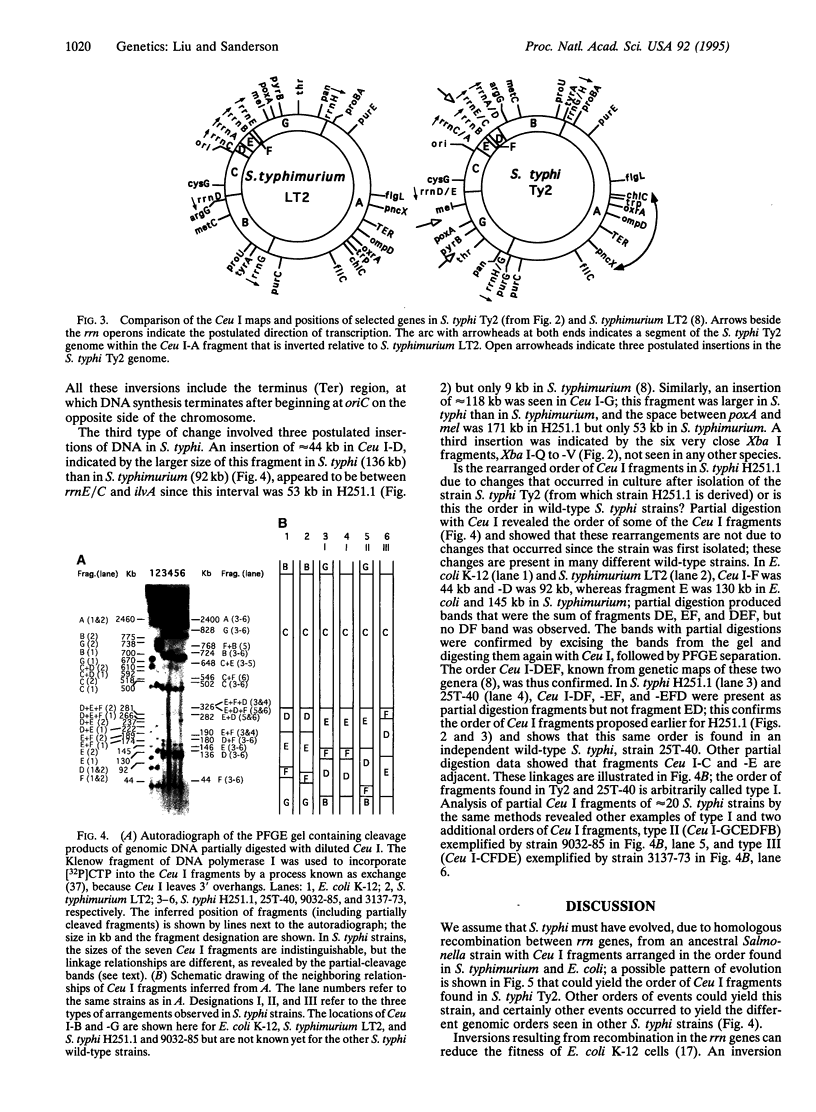


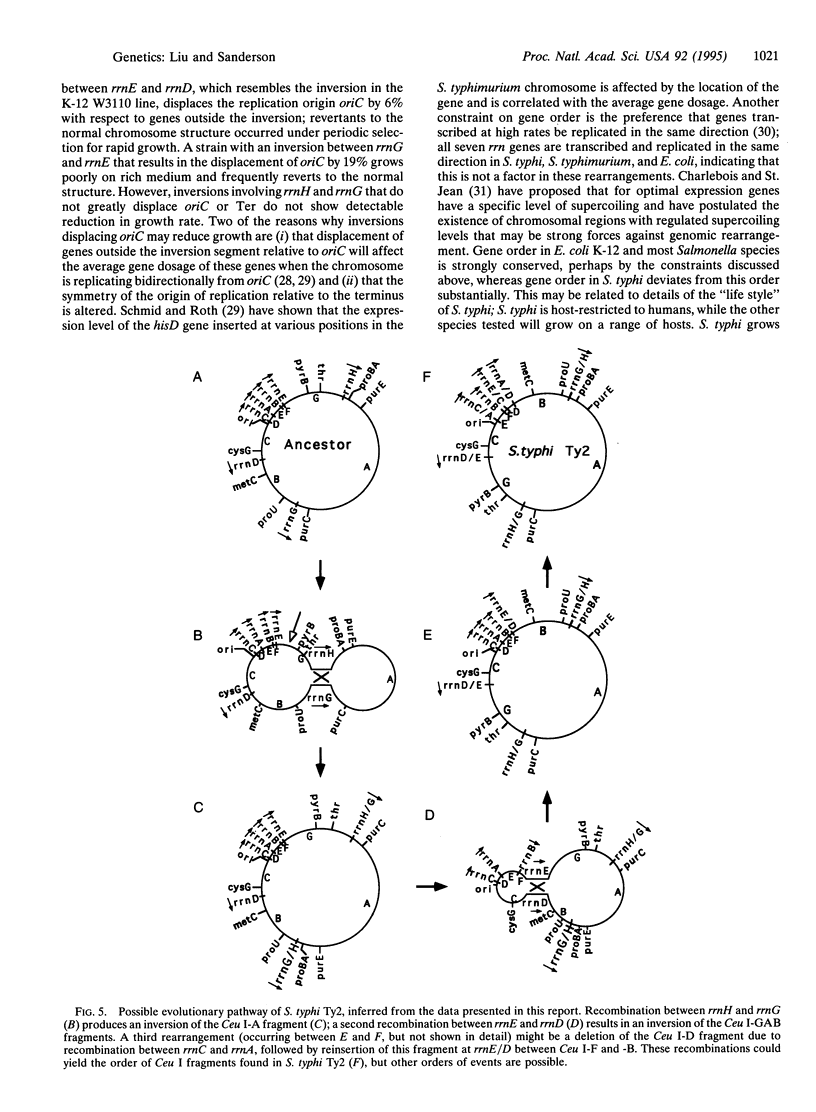

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson P., Roth J. Spontaneous tandem genetic duplications in Salmonella typhimurium arise by unequal recombination between rRNA (rrn) cistrons. Proc Natl Acad Sci U S A. 1981 May;78(5):3113–3117. doi: 10.1073/pnas.78.5.3113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anderson R. P., Roth J. R. Gene duplication in bacteria: alteration of gene dosage by sister-chromosome exchanges. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 2):1083–1087. doi: 10.1101/sqb.1979.043.01.120. [DOI] [PubMed] [Google Scholar]
- Brewer B. J. When polymerases collide: replication and the transcriptional organization of the E. coli chromosome. Cell. 1988 Jun 3;53(5):679–686. doi: 10.1016/0092-8674(88)90086-4. [DOI] [PubMed] [Google Scholar]
- Caro L. G., Berg C. M. Chromosome replication in some strains of Escherichia coli K12. Cold Spring Harb Symp Quant Biol. 1968;33:559–573. doi: 10.1101/sqb.1968.033.01.063. [DOI] [PubMed] [Google Scholar]
- Casse F., Pascal M. C., Chippaux M. Comparison between the chromosomal maps of Escherichia coli and Salmonella typhimurium. Length of the inverted segment in the trp region. Mol Gen Genet. 1973 Aug 17;124(3):253–257. doi: 10.1007/BF00293096. [DOI] [PubMed] [Google Scholar]
- Crosa J. H., Brenner D. J., Ewing W. H., Falkow S. Molecular relationships among the Salmonelleae. J Bacteriol. 1973 Jul;115(1):307–315. doi: 10.1128/jb.115.1.307-315.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cryz S. J., Jr, Vanprapar N., Thisyakorn U., Olanratmanee T., Losonsky G., Levine M. M., Chearskul S. Safety and immunogenicity of Salmonella typhi Ty21a vaccine in young Thai children. Infect Immun. 1993 Mar;61(3):1149–1151. doi: 10.1128/iai.61.3.1149-1151.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edelman R., Levine M. M. Summary of an international workshop on typhoid fever. Rev Infect Dis. 1986 May-Jun;8(3):329–349. doi: 10.1093/clinids/8.3.329. [DOI] [PubMed] [Google Scholar]
- Edwards M. F., Stocker B. A. Construction of delta aroA his delta pur strains of Salmonella typhi. J Bacteriol. 1988 Sep;170(9):3991–3995. doi: 10.1128/jb.170.9.3991-3995.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hill C. W., Gray J. A. Effects of chromosomal inversion on cell fitness in Escherichia coli K-12. Genetics. 1988 Aug;119(4):771–778. doi: 10.1093/genetics/119.4.771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hill C. W., Harnish B. W. Transposition of a chromosomal segment bounded by redundant rRNA genes into other rRNA genes in Escherichia coli. J Bacteriol. 1982 Feb;149(2):449–457. doi: 10.1128/jb.149.2.449-457.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hone D. M., Harris A. M., Chatfield S., Dougan G., Levine M. M. Construction of genetically defined double aro mutants of Salmonella typhi. Vaccine. 1991 Nov;9(11):810–816. doi: 10.1016/0264-410x(91)90218-u. [DOI] [PubMed] [Google Scholar]
- Honeycutt R. J., McClelland M., Sobral B. W. Physical map of the genome of Rhizobium meliloti 1021. J Bacteriol. 1993 Nov;175(21):6945–6952. doi: 10.1128/jb.175.21.6945-6952.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohara Y., Akiyama K., Isono K. The physical map of the whole E. coli chromosome: application of a new strategy for rapid analysis and sorting of a large genomic library. Cell. 1987 Jul 31;50(3):495–508. doi: 10.1016/0092-8674(87)90503-4. [DOI] [PubMed] [Google Scholar]
- Krawiec S., Riley M. Organization of the bacterial chromosome. Microbiol Rev. 1990 Dec;54(4):502–539. doi: 10.1128/mr.54.4.502-539.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krug P. J., Gileski A. Z., Code R. J., Torjussen A., Schmid M. B. Endpoint bias in large Tn10-catalyzed inversions in Salmonella typhimurium. Genetics. 1994 Mar;136(3):747–756. doi: 10.1093/genetics/136.3.747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu S. L., Hessel A., Cheng H. Y., Sanderson K. E. The XbaI-BlnI-CeuI genomic cleavage map of Salmonella paratyphi B. J Bacteriol. 1994 Feb;176(4):1014–1024. doi: 10.1128/jb.176.4.1014-1024.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu S. L., Hessel A., Sanderson K. E. Genomic mapping with I-Ceu I, an intron-encoded endonuclease specific for genes for ribosomal RNA, in Salmonella spp., Escherichia coli, and other bacteria. Proc Natl Acad Sci U S A. 1993 Jul 15;90(14):6874–6878. doi: 10.1073/pnas.90.14.6874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu S. L., Hessel A., Sanderson K. E. The XbaI-BlnI-CeuI genomic cleavage map of Salmonella enteritidis shows an inversion relative to Salmonella typhimurium LT2. Mol Microbiol. 1993 Nov;10(3):655–664. doi: 10.1111/j.1365-2958.1993.tb00937.x. [DOI] [PubMed] [Google Scholar]
- Liu S. L., Hessel A., Sanderson K. E. The XbaI-BlnI-CeuI genomic cleavage map of Salmonella typhimurium LT2 determined by double digestion, end labelling, and pulsed-field gel electrophoresis. J Bacteriol. 1993 Jul;175(13):4104–4120. doi: 10.1128/jb.175.13.4104-4120.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mahan M. J., Roth J. R. Ability of a bacterial chromosome segment to invert is dictated by included material rather than flanking sequence. Genetics. 1991 Dec;129(4):1021–1032. doi: 10.1093/genetics/129.4.1021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall P., Lemieux C. Cleavage pattern of the homing endonuclease encoded by the fifth intron in the chloroplast large subunit rRNA-encoding gene of Chlamydomonas eugametos. Gene. 1991 Aug 15;104(2):241–245. doi: 10.1016/0378-1119(91)90256-b. [DOI] [PubMed] [Google Scholar]
- Pang T., Altwegg M., Martinetti G., Koh C. L., Puthucheary S. Genetic variation among Malaysian isolates of Salmonella typhi as detected by ribosomal RNA gene restriction patterns. Microbiol Immunol. 1992;36(5):539–543. doi: 10.1111/j.1348-0421.1992.tb02053.x. [DOI] [PubMed] [Google Scholar]
- Perkins J. D., Heath J. D., Sharma B. R., Weinstock G. M. XbaI and BlnI genomic cleavage maps of Escherichia coli K-12 strain MG1655 and comparative analysis of other strains. J Mol Biol. 1993 Jul 20;232(2):419–445. doi: 10.1006/jmbi.1993.1401. [DOI] [PubMed] [Google Scholar]
- Reeves M. W., Evins G. M., Heiba A. A., Plikaytis B. D., Farmer J. J., 3rd Clonal nature of Salmonella typhi and its genetic relatedness to other salmonellae as shown by multilocus enzyme electrophoresis, and proposal of Salmonella bongori comb. nov. J Clin Microbiol. 1989 Feb;27(2):313–320. doi: 10.1128/jcm.27.2.313-320.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanderson K. E. Genetic relatedness in the family Enterobacteriaceae. Annu Rev Microbiol. 1976;30:327–349. doi: 10.1146/annurev.mi.30.100176.001551. [DOI] [PubMed] [Google Scholar]
- Sanderson K. E., Hall C. A. F-prime factors of Salmonella typhimurium and an inversion between S. typhimurium and Escherichia coli. Genetics. 1970 Feb;64(2):215–228. doi: 10.1093/genetics/64.2.215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmid M. B., Roth J. R. Gene location affects expression level in Salmonella typhimurium. J Bacteriol. 1987 Jun;169(6):2872–2875. doi: 10.1128/jb.169.6.2872-2875.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmid M. B., Roth J. R. Genetic methods for analysis and manipulation of inversion mutations in bacteria. Genetics. 1983 Nov;105(3):517–537. doi: 10.1093/genetics/105.3.517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Selander R. K., Beltran P., Smith N. H., Helmuth R., Rubin F. A., Kopecko D. J., Ferris K., Tall B. D., Cravioto A., Musser J. M. Evolutionary genetic relationships of clones of Salmonella serovars that cause human typhoid and other enteric fevers. Infect Immun. 1990 Jul;58(7):2262–2275. doi: 10.1128/iai.58.7.2262-2275.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith C. L., Econome J. G., Schutt A., Klco S., Cantor C. R. A physical map of the Escherichia coli K12 genome. Science. 1987 Jun 12;236(4807):1448–1453. doi: 10.1126/science.3296194. [DOI] [PubMed] [Google Scholar]
- Zahrt T. C., Mora G. C., Maloy S. Inactivation of mismatch repair overcomes the barrier to transduction between Salmonella typhimurium and Salmonella typhi. J Bacteriol. 1994 Mar;176(5):1527–1529. doi: 10.1128/jb.176.5.1527-1529.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]