Summary
Pollinator-mediated reproductive isolation is a major factor in driving the diversification of flowering plants. Studies of floral traits involved in reproductive isolation have focused nearly exclusively on visual signals, such as flower color. The role of less obvious signals, such as floral scent, has been studied only recently. In particular, the genetics of floral volatiles involved in mediating differential pollinator visitation remains unknown. The bumblebee-pollinated Mimulus lewisii and hummingbird-pollinated M. cardinalis are a model system for studying reproductive isolation via pollinator preference. We have shown that these two species differ in three floral terpenoid volatiles - D-limonene, β-myrcene, and E-β-ocimene - that are attractive to bumblebee pollinators. By genetic mapping and in vitro enzyme activity analysis we demonstrate that these interspecific differences are consistent with allelic variation at two loci – LIMONENE-MYRCENE SYNTHASE (LMS) and OCIMENE SYNTHASE (OS). M. lewisii LMS (MlLMS) and OS (MlOS) are expressed most strongly in floral tissue in the last stages of floral development. M. cardinalis LMS (McLMS) is weakly expressed and has a nonsense mutation in exon 3. M. cardinalis OS (McOS) is expressed similarly to MlOS, but the encoded McOS enzyme produces no E-β-ocimene. Recapitulating the M. cardinalis phenotype by reducing the expression of MlLMS by RNAi in transgenic M. lewisii produces no behavioral difference in pollinating bumblebees; however, reducing MlOS expression produces a 6% decrease in visitation. Allelic variation at the OCIMENE SYNTHASE locus likely contributes to differential pollinator visitation, and thus promotes reproductive isolation between M. lewisii and M. cardinalis. OCIMENE SYNTHASE joins a growing list of “speciation genes” (“barrier genes”) in flowering plants.
Keywords: Terpene synthase, floral volatiles, speciation, pollination, Mimulus lewisii, Mimulus cardinalis
Introduction
The rapid diversification of the world’s estimated 275,000 species of flowering plants has often been attributed to their specialized association with different animal pollinators (Grant, 1949; Stebbins, 1970). Flowering plants use a variety of signals to advertise the presence (or illusion) of a reward to their associated pollinators; the association between pollinator type and suites of signals gives rise to the concept of pollination syndromes (Fenster et al., 2004). Perhaps the most well-known and easily studied signal is floral color, which has been investigated in a variety of pollination syndromes (Rausher, 2008). Other visual signals, such as texture, pattern, orientation, anthesis time, size, and shape have been investigated to some extent in a variety of systems (Harder and Johnson, 2009; Kay and Sargent, 2009; Yuan et al., 2013a).
Floral scent – the amount, relative ratios, and identities of volatile compounds emitted by the flower – is a generally understudied signal, despite the long understanding that it may play a strong role in attracting pollinators (Raguso, 2008a). The recent development of techniques for studying floral scent, including chemical analysis of floral scent, analysis of pollinator neural activity at both the receptor and higher-order processing levels, and genetic and genomic tools, has allowed some progress in this area. However, although floral scent is frequently characterized, and genes responsible for the production of floral volatiles are occasionally identified, a synthesis of floral scent biochemistry, neurobiology, genetics, ecology, and evolution has been lacking. Those systems with well-characterized volatiles that affect pollination are separate from those with well-characterized genetics (Raguso, 2008a; Raguso, 2008b; Whitehead and Peakall, 2009; Parachnowitsch et al., 2012).
Much of the work discussing speciation involving floral volatiles has been done in extremely specialized systems where scent is crucial to plant-pollinator interactions (Raguso, 2008b), most notably the sexually deceptive orchids in the genera Chiloglottis (Schiestl and Peakall, 2005; Peakall et al., 2010) and Ophrys (Schiestl and Ayasse, 2002; Mant et al., 2005; Vereecken et al., 2010; Xu et al., 2012), as well as the non-deceptive genus Gymnadenia (Huber et al., 2005). Recent work has begun expanding this to non-orchid systems such as Silene (Waelti et al., 2008), Linanthus (Chess et al., 2008), and Petunia (Klahre et al., 2011). While there are a growing number of studies demonstrating the importance of floral volatiles in mediating these largely specialized plant-pollinator interactions, the genetic pathways controlling volatile production in these systems remain unknown.
In contrast, the genes underlying volatile production are known in a diverse range of angiosperm systems (Gang, 2005), including Clarkia (Pichersky et al., 1995; Dudareva et al., 1996; Dudareva et al., 1998; Wang and Pichersky, 1998), Antirrhinum (Dudareva et al., 2000; Dudareva et al., 2003), Petunia (Koeduka et al., 2006; Orlova et al., 2006; Dexter et al., 2007), Silene (Gupta et al., 2012), Arabidopsis (Bohlmann et al., 2000; Chen et al., 2003), and many species of agricultural importance. Although our knowledge of the genetic underpinnings of volatile production and emission has grown as a result of these systems, there is a paucity of research linking floral volatiles and plant speciation with the genetic and molecular basis for those effects.
Petunia is the only well-developed model demonstrating the role that a specific volatile plays in differential attraction of pollinators between sister species (Klahre et al., 2011). The sister species P. axillaris and P. exserta differ in their production of methyl benzoate, a volatile attractive to the hawkmoth pollinators of P. axillaris. Through QTL mapping, two regions underlying this difference were identified in the Petunia genome on chromosomes II and VII, with the P. axillaris allele at the locus on chromosome II being absolutely required for methyl benzoate production and the locus on chromosome VII substantially contributing to methyl benzoate quantity. ODO1, one of the genes hypothesized to underlie the locus on chromosome VII, encodes a MYB transcription factor that is differentially expressed in the two species. The hawkmoth Manduca sexta is attracted more strongly to near isogenic lines with high levels of methyl benzoate production, suggesting that a change in volatile production mediated by a single gene can lead to differential pollinator attraction; however, the underlying genetic mechanisms mediating species-specific volatile emission in this system have not been completely described.
The sister species M. lewisii and M. cardinalis have served as a model system for studying pollinator-mediated reproductive isolation for several decades (Hiesey et al., 1971), and the combination of ecological and genetic resources has led to the discovery of multiple loci impacting differential pollinator attraction between the two species (Bradshaw and Schemske, 2003; Yuan et al., 2013b; Yuan et al., 2013c). Mimulus lewisii is a bumblebee-pollinated alpine species, while its sister, M. cardinalis, is a lower-elevation hummingbird-pollinated species (Hiesey et al., 1971; Schemske and Bradshaw, 1999). Within areas of sympatry, pollinator fidelity is responsible for 98% of reproductive isolation between M. lewisii and M. cardinalis (Ramsey et al., 2003). Previous work has shown that three floral volatiles produced by M. lewisii – D-limonene, β-myrcene, and E-β-ocimene – are important for the attraction of bumblebee pollinators, including Bombus vosnesenskii, the native pollinator of M. lewisii in the central Sierra Nevada mountains of California. Of the three volatiles, M. cardinalis produces only D-limonene, released at just 0.9% the rate of M. lewisii (Byers et al., 2014).
Therefore, we ask some global questions, which we begin to address in this manuscript: What are the genetic underpinnings of the differential emission of floral volatiles between M. lewisii and M. cardinalis? How many genes are responsible, and how do the species differ in gene expression and protein function? What role, if any, do these scent differences play in differential pollinator visitation, and through this, reproductive isolation? In keeping with other discussions of “speciation genes” in plants (Rieseberg and Blackman, 2010), are these genes of large effect or small effect, coding or regulatory genes? Mimulus, with its known attractive volatiles, genetic and genomic tools, and well-studied ecology, is an obvious choice for filling in this missing piece of the floral scent-speciation link.
Results
Genetic mapping of species-specific differences in floral volatiles
Construction of an F1 cross between M. lewisii inbred line LF10 and M. cardinalis inbred line CE10 revealed patterns of inheritance of loci controlling the emission of D-limonene, β-myrcene, and E-β-ocimene. The emission rate of D-limonene for the F1 (mean = 47.2 ng/flower/hr; N = 3) was similar to that of the M. lewisii parental inbred line (mean = 55.1 ng/flower/hr, N = 9; two-tailed Mann-Whitney p = 0.864, U = 15) and much higher than that of the M. cardinalis parental inbred line (mean = 0.5 ng/flower/hr, N = 9; one-tailed p = 0.005, U = 27), suggesting that high levels of D-limonene emission are inherited from the M. lewisii parent in a dominant manner. The pattern was similar for β-myrcene (F1: mean = 2.6 ng/flower/hr, M. lewisii: mean = 3.3 ng/flower/hr, M. cardinalis: mean = 0.0 ng/flower/hr; F1 vs. M. lewisii two-tailed p = 0.600, U = 17; F1 vs. M. cardinalis one-tailed p = 0.005, U = 27). For E-β-ocimene, the M. lewisii allele appears to be semidominant (F1: mean = 2.8 ng/flower/hr; M. lewisii: mean = 7.6 ng/flower/hr; M. cardinalis: mean = 0.0 ng/flower/hr; F1 vs. M. lewisii two-tailed p = 0.036, U =25; F1 vs. M. cardinalis one-tailed p = 0.005, U =27). Complete or partial dominance of the M. lewisii alleles for these floral volatiles is consistent with other traits that differ between the species (Bradshaw et al., 1998).
When a backcross (F1 x M. cardinalis) population of 100 plants was scored for the presence or absence of emission of β-myrcene and E-β-ocimene, it segregated approximately 1:1 for both volatiles (0.52:0.48 β-myrcene present:absent; 0.38:0.62 E-β-ocimene present:absent), suggesting that alleles at Mendelian loci might control the difference in emission of these monoterpenes between M. lewisii and M. cardinalis. D-limonene and β-myrcene emission rates were very highly correlated (r = 0.975), but neither was particularly highly correlated with E-β-ocimene emission rate (r = 0.474 versus D-limonene, r = 0.574 versus β-myrcene). Therefore, we considered a two-locus model for the difference in these three compounds between the two species – one locus controlling the production of D-limonene and β-myrcene, and another, unlinked, locus controlling E-β-ocimene. A larger backcross population (N = 768) was constructed to map the two loci with greater precision.
Identification and characterization of a bifunctional LIMONENE-MYRCENE SYNTHASE (LMS) in M. lewisii flowers
The locus associated with D-limonene and β-myrcene emission was mapped to a 15 cM interval between markers M02_510K and M02_1500K (Table S4), ca. 5.3 cM from M02_1500K. Using the assembled and annotated M. guttatus genome v1.1 as a reference (http://www.phytozome.net/cgi-bin/gbrowse/mimulus/), the ortholog of M02_1500K maps to M. guttatus scaffold 89 at position 201 kbp. On scaffold 89 between positions 206 kbp and 226 kbp there is a cluster of three terpene synthases/cyclases – excellent candidates for controlling D-limonene and β-myrcene emission.
Indel markers developed for two of the M. lewisii/cardinalis candidate genes in the terpene synthase cluster (Table S5) revealed no recombinations (in 768 backcross plants) between themselves or the putative LIMONENE-MYRCENE SYNTHASE (see Methods). The very tight linkage among the candidate terpene synthases within the cluster made it impractical to resolve the identity of the D-limonene and β-myrcene synthases by recombination. RT-PCR showed that, of the three candidates, only the M. lewisii ortholog (KF857265) of the M. guttatus terpene synthase gene on scaffold 89 at position 321 kbp (mgv1a003660m) is transcribed in M. lewisii flowers. The marker genotype at M02_1500 accounted for 92% of the difference between M. lewisii and M. cardinalis emissions of D-limonene and 98% of the difference in β-myrcene emissions, consistent with a single-locus model for D-limonene and β-myrcene production. No transgressive segregation was observed in the backcross population. The predicted AHI50308 gene product contains the conserved DDxx(D/E) and (N,D)Dxx(S,T,G)xxxE (NSE/DTE) motifs required for Mg2+ binding during the terpene synthesis process (Nieuwenhuizen et al., 2013), as well as the RRx8W motif required for cyclic terpene formation (Dudareva et al., 2003).
The M. lewisii cDNA (KF857264) orthologous to mgv1a003660m, designated TS321K, was overexpressed in E. coli (as in Bohlmann et al., 2000). A crude lysate from the E. coli culture was supplied with geranyl pyrophosphate (GPP) as a substrate, yielding D-limonene and β-myrcene in the same proportions as observed in the authentic headspace collection from M. lewisii flowers (Table S1, Fig. 2A). This suggests that the high correlation between D-limonene and β-myrcene emission in the backcross mapping population is due to the pleiotropic effect of a bifunctional LIMONENE-MYRCENE SYNTHASE (LMS) encoded by a single LMS gene in M. lewisii (MlLMS). This is consistent with the frequent occurrence of multi-product terpene synthases (Dudareva et al., 2004).
Figure 2.
Terpene synthase activity in vitro and in vivo. (A) Products of terpene synthases using a bacterial overexpression system and in vitro enzyme assay. Using geranyl pyrophosphate (GPP, the common monoterpene precursor), the MlLMS enzyme produces D-limonene (b) and β-myrcene (a) in the same relative proportion as in the floral volatile emission. M. lewisii MlOS produces E-β-ocimene (c), but M. cardinalis McOS does not. (B) Temporal expression of terpene synthases in vivo. MlLMS is expressed just prior to flowering and in open flowers, but McLMS is not expressed in open flowers. MlOS shows a similar expression pattern to MlLMS, and McOS is expressed at the same stages, despite producing no terpenoid volatile that we could detect. Developmental staging is according to bud size of M. lewisii, and the corresponding stage of M. cardinalis is pictured for McLMS, McOS, and McUBC.
The M. cardinalis LMS (McLMS) coding sequence was not expressed in vitro because there is a G66T transversion mutation in exon 3 of McLMS (KM659024) that results in a nonsense mutation in the McLMS protein (G201X, using MlLMS AHI50308 as the reference allele).
Of note, this is not a definitive demonstration that LMS is the gene underlying the locus responsible for the difference in D-limonene and β-myrcene emission between M. lewisii and M. cardinalis; it is formally possible that a different, tightly-linked gene instead might be responsible. Transgenic complementation of the nonfunctional M. cardinalis LMS allele by the M. lewisii allele would be necessary to show conclusively that LMS is the causal gene for the phenotypic difference.
Identification and characterization of OCIMENE SYNTHASE (OS) in M. lewisii flowers
The locus associated with E-β-ocimene emission was mapped to a 7.5 cM interval midway between markers sc4_2325K and M13_2620 (Table S4). The marker genotype at sc4_2325 accounted for 98% of the difference between M. lewisii and M. cardinalis emissions of E-β-ocimene, consistent with a single-locus model for E-β-ocimene production. No transgressive segregation was observed in the backcross population. The orthologous region of the M. guttatus genome lies in a 484 kbp interval (2325 kbp – 2809 kbp) on scaffold 4. There is a cluster of five terpene biosynthesis genes on scaffold 4 at position 2538 kbp – 2577 kbp. The gene at position 2538 kbp (mgv1a020487m) is annotated as a terpene synthase, while the other four genes are annotated as sesquiterpene cyclases (Fig. 1). An indel marker developed for the M. lewisii/cardinalis ortholog of mgv1a003660m, designated TS2538 (Table S5), revealed no recombinations (in 768 backcross plants) with the putative OCIMENE SYNTHASE (see Methods).
Figure 1.
Mimulus lewisii and M. cardinalis and their terpene synthases. (A) Mimulus lewisii and M. cardinalis. (B) Genetic maps of MlLMS and MlOS and the homologous regions in M. guttatus. Positions on the lower half of each are from the M. guttatus genome scaffolds; annotations are from queries of the M. guttatus transcripts with BLASTx. Putative terpene synthases are highlighted. M. lewisii limonene-myrcene synthase is homologous to a terpene synthase at 319,982 bp on M. guttatus scaffold 89, and M. lewisii ocimene synthase is homologous to a terpene synthase at 2,538,727 bp on scaffold 4.
When overexpressed in E. coli and supplied with GPP as a substrate, the M. lewisii TS2538 cDNA (KF857262) encodes a functional OCIMENE SYNTHASE (MlOS, AHI50306) (Fig. 2A, Table S1). However, under the same conditions the M. cardinalis TS2538 cDNA (KF857263) does not encode an enzyme (McOS, AHI50307) capable of synthesizing any monoterpene that we could detect. McOS differs from MlOS at 19 amino acid residues, including insertion of a leucine residue at position 238 in McOS and deletion of an arginine residue at position 308 in McOS (Fig. S1). Both sequences contain the same DDxx(D/E) and NSE/DTE Mg2+ binding motifs, as well as the RRx8W cyclase motif, which are unaltered by the 19 nonsynonymous amino acid substitutions (Fig. S1).
As with LMS, in the absence of a transgenic complementation test this is not a strict demonstration that MlOS is the gene underlying the locus responsible for the difference in E-β-ocimene emission between M. lewisii and M. cardinalis.
LMS and OS expression in vivo
Using RT-PCR with six different stages of flowering tissue from early bud (8d prior to anthesis, 5mm) to open flower (see Yuan et al., 2013c), we found that both MlLMS and MlOS are expressed in the last three days prior to anthesis (15mm and 20mm) of floral development, as well as in the open flower (Fig. 2B). McLMS is weakly expressed in late-stage floral buds but not expressed in open flowers. McOS is expressed similarly to MlOS.
Construction of RNAi knockdowns of MlLMS and MlOS in stably transformed M. lewisii
Using RNA interference via Agrobacterium-mediated in planta transformation of hairpin RNAi constructs into M. lewisii (Yuan et al., 2013c), we were able to knock down the expression of both MlLMS and MlOS to produce much lower floral volatile levels, comparable to those produced in plants homozygous for the M. cardinalis alleles at LMS and OS (Table S2). This allowed us both to verify LMS and OS gene function in vivo and to determine the effect of decreased emission of specific floral volatiles on pollinating bumblebees.
We recovered 24 M. lewisii (inbred line LF10) T1 plants carrying the MlLMS-RNAi transgene, and assayed each transgenic plant in triplicate for floral volatile production. All T1 plants had lower emission rates of D-limonene and β-myrcene relative to the wild-type M. lewisii LF10 (D-limonene: range = 1.2%–56.1%, mean = 10.2%; β-myrcene: range = 4.1–50.0%, mean = 12.8%). Interestingly, most T1 plants showed a decrease in emission of terpinolene (range = 0.0%–132.6%, mean = 18.0%), indicating that MlLMS may be responsible for synthesizing an additional minor compound in M. lewisii. One of these T1 transgenics (LMS321K-8) was selfed as the parent of T2 plants used for pollinator studies (for data on three other T2 lines from independent T1 transgenics, see Table S3). The original T1 LMS321K-8 had very low emission rates of D-limonene and β-myrcene, with a mean of 2.8% D-limonene production and 9.1% β-myrcene production relative to the M. lewisii LF10 T0 parent. Notably, LMS321K-8 had an increase in E-β-ocimene of 452.8% compared with the M. lewisii T0 parent (Table 1, Fig. 3). All other T1 plants had a similar increase in E-β-ocimene production relative to the wild-type parent (range = 190.4–493.9%, mean = 383.4%).
Table 1.
Volatile production in transgenic (T1 parent plants of greenhouse experiment lines) and wild-type M. lewisii and M. cardinalis. Values are an average of 2–3 independent headspace samples. Emission values in parentheses represent the 90% confidence interval. See Table S2 for complete data on all volatiles produced by M. lewisii.
| Volatile | LF10 (ng/hr) |
CE10 (ng/hr) |
MlLMS- RNAi (ng/hr) |
MlOS- RNAi (ng/hr) |
MlLMS- RNAi (% LF10) |
MlLMS- RNAi (% CE10) |
MlOS- RNAi (% LF10) |
MlOS- RNAi (% CE10 |
|---|---|---|---|---|---|---|---|---|
| β-myrcene | 2.837 (2.056, 3.793) | Absent (0.000, 0.000) | 0.257 (0.194, 0.339) | 1.116 (1.047, 1.185) | 9.06% | n/a | 39.34% | n/a |
| D-limonene | 43.228 (35.539, 50.708) | 1.024 (0.649, 1.419) | 1.216 (0.765, 1.757) | 23.820 (23.285, 24.354) | 2.81% | 118.75% | 55.10% | 2326.17 % |
| E-β-ocimene | 5.563 (4.320, 7.049) | Absent (0.000, 0.000) | 25.125 (20.591, 30.120) | 0.054 (0.042, 0.074) | 451.64% | n/a | 0.97% | n/a |
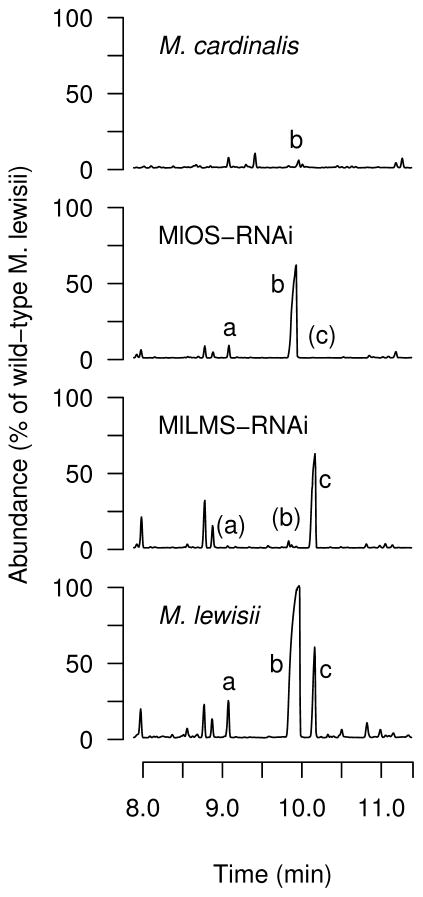
Figure 3.
Stable RNAi knockdowns of MlLMS and MlOS in M. lewisii produce plants with low emission levels of D-limonene (b)/β-myrcene (a) and E-β-ocimene (c), respectively.
A total of 71 T2 plants from the self-pollinated progeny of T1 LMS321K-8 were assayed using headspace collection of floral volatiles to select the greenhouse population for the bumblebee pollinator behavioral experiment. The 24 individuals selected for the experiment produced much less D-limonene and β-myrcene compared to the wild-type M. lewisii ancestor (D-limonene: range = 0.1–2.4%, mean = 1.9%; β-myrcene: range = 0.0–4.8%, mean = 0.6%), and more E-β-ocimene than the M. lewisii wild-type ancestor (range = 93.0–510.5%, mean = 247.2%). The D-limonene and β-myrcene levels were similar to those found in M. cardinalis (D-limonene: range = 0.03–2.8%, mean = 0.9% of wild-type M. lewisii; β-myrcene is absent from M. cardinalis).
Only two T1 plants carrying the MlOS-RNAi transgene were recovered, but both had the desired E-β-ocimene knockdown phenotype relative to M. lewisii LF10 (E-β-ocimene: range = 0.8%–2.9%, mean = 1.8%; D-limonene: range = 39.6%–58.9%, mean = 49.2%; β-myrcene: range = 28.3%–41.2%, mean = 34.8%). T1 plant TS2538-1, which was self-pollinated to create a T2 population for pollinator studies, had a much lower emission rate of E-β-ocimene (0.8%) relative to M. lewisii LF10, as well as lower emission rates of D-limonene (39.6%) and β-myrcene (28.3%)(Fig. 3). T1 plant TS2538-2 flowered substantially later than TS2538-1, so T2 lines were not created from this plant. A total of 80 T2 plants were produced from TS2538-1, and these produced similar amounts of D-limonene and β-myrcene as the M. lewisii LF10 ancestor (D-limonene: range = 61.3–127.4%, mean = 83.6%; β-myrcene: range = 57.3–144.5%, mean = 88.7%), but much less E-β-ocimene (range = 0.9–3.9%, mean = 1.9%).
Effects of MlLMS and MlOS knockdowns on bumblebee pollinator behavior
Two experiments, one for the MlLMS-RNAi transgenics and one for the MlOS-RNAi transgenics, were performed to assay the impact of reduced monoterpene production on bumblebee (Bombus impatiens) visitation in a captive greenhouse setting. During each experiment, both preference (expressed as the proportion of total visits to each flower type) and constancy (expressed as the tendency of an individual bumblebee to deviate from random choices, exclusive of preference, see Waser, 1986) were measured.
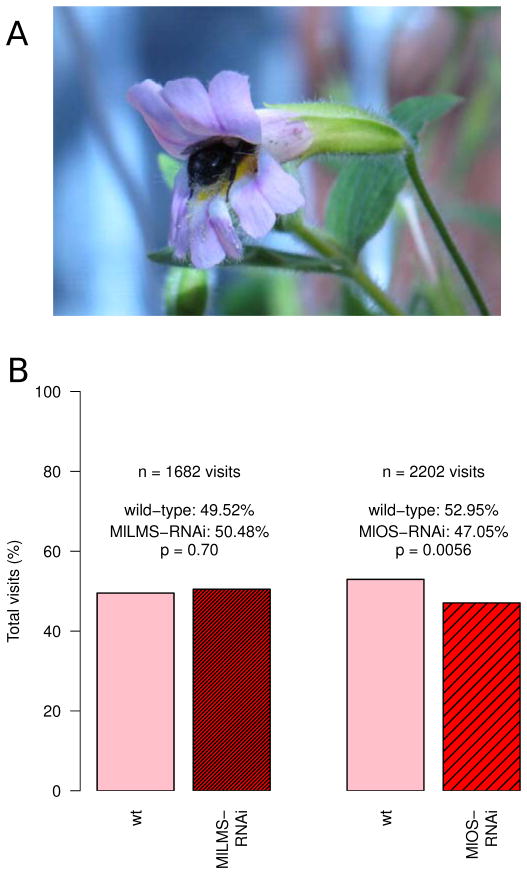
A total of 1682 visits were observed to flowers in the MlLMS-RNAi knockdown experiment. Visits were defined as observable contact with the sexual organs of the flower – i.e., the bumblebee entered the flower fully, as required to effect pollination. Of 1682 visits, 833 (49.52%) were to the wild-type M. lewisii and 849 (50.48%) were to the M. lewisii MlLMS-RNAi transgenic plants, showing no significant difference (χ2 = 0.15, p = 0.70, Fig. 4). Bumblebees appeared to show no overall qualitative behavioral difference towards either flower type.
Figure 4.
Greenhouse experiments with Bombus impatiens and M. lewisii wild-type and transgenic lines. (A) Image of a typical bumblebee visit. (B) Response of bumblebees to MlLMS RNAi knockdowns, MlOS RNAi knockdowns, and the wild-type parent. Bumblebees show the same visitation response to MlLMS knockdown transgenics as to wild-type M. lewisii. Bumblebees preferentially visit wild-type M. lewisii over MlOS knockdown transgenics.
A total of 39 bumblebee foraging bouts were assayed for constancy, with an average Bateman’s index of −0.0114 (−1 indicates complete inconstancy – regular switching between types; 0 indicates random visitation patterns; +1 indicates complete constancy, always within types). To determine whether this constancy was significantly different from random visitation, the same bumblebee foraging bouts were used with 100,000 simulated runs of randomly permuted plant locations, resulting in an average Bateman’s index of −0.1141. A total of 96,648 simulations had more divergent Bateman’s index values than the actual data, showing that bumblebees demonstrated no constancy when presented with these flowers (p = 0.97).
For the MlOS-RNAi knockdown experiment, a total of 2202 visits were observed. Of these visits, 1166 (52.95%) were to wild-type M. lewisii and 1036 (47.05%) were to the M. lewisii MlOS-RNAi plants, showing a significant preference for the wild-type M. lewisii flowers (X2 =7.67, p = 0.0056, Fig. 4). Bumblebees approaching the MlOS-RNAi flowers were noted to frequently wave their antennae and contact the flower with their antennae prior to aborting a potential visit, suggesting that E-β-ocimene may operate as a near-field olfactory cue, but this behavior was not noted for the wild-type flowers in this experiment nor for either flower type in the MlLMS-RNAi experiment. Constancy was also absent in the MlOS-RNAi experiment, with a total of 46 bumblebee foraging bouts showing an average Bateman’s index of 0.0149; the simulation described above was repeated using these foraging bouts, with an average Bateman’s index of −0.1142 (p = 0.95).
Discussion
Mimulus lewisii produces three floral volatiles with significant neurophysiological and behavioral effects on bumblebees – D-limonene, β-myrcene, and E-β-ocimene, while M. cardinalis produces only D-limonene at much lower levels (0.9% of M. lewisii) (Byers et al., 2014). These differences are likely due to mutations in two genes: LIMONENE-MYRCENE SYNTHASE (MlLMS, McLMS) and OCIMENE SYNTHASE (MlOS, McOS). In quantitative genetic terms, allelic variation at loci containing LMS and OS accounts for 92–98% of the phenotypic difference between M. lewisii and M. cardinalis in floral emission of D-limonene, β-myrcene, and E-β-ocimene. The very low level of volatile emission from M. cardinalis flowers can be explained at the molecular genetic level; McLMS is a null allele due to a nonsense mutation in exon 3, while McOS has multiple coding sequence differences that eliminate its ability to produce E-β-ocimene.
RNAi knockouts show that the loss-of-function LMS and OS alleles can recapitulate the M. cardinalis volatile emission phenotypes, and that functional copies of both genes are necessary to produce D-limonene, β-myrcene, and E-β-ocimene in vivo. However, there remains the formal possibility that the allelic variants producing differences in floral volatile emissions between M. lewisii and M. cardinalis are not in LMS or OS, but in genes very tightly linked to them. To show that the M. lewisii alleles of LMS and OS are sufficient (since we have shown that they are necessary) to produce D-limonene, β-myrcene, and E-β-ocimene, we would have to transform M. cardinalis with constructs containing the M. lewisii alleles. However, M. cardinalis is very difficult to transform, so we have not performed these definitive experiments. Although we lack conclusive evidence that these are the genes underlying these loci, we present strong circumstantial evidence (the nonsense mutation in McLMS and the lack of product from McOS activity in vitro) that is consistent with this, and RNAi knockouts show that the loss-of-function LMS and OS alleles can recapitulate the M. cardinalis volatile emission phenotypes necessary to test for differential pollinator visitation.
Surprisingly, despite the high level of production of D-limonene and β-myrcene in M. lewisii flowers, substantially knocking down emission of these two compounds produces no significant effect on bumblebee visitation in the greenhouse. In contrast, knocking down emission of E-β-ocimene results in a modest (6%) but significant decrease in bumblebee visitation, suggesting that alternative alleles of OCIMENE SYNTHASE can contribute to reproductive isolation between the bumblebee-pollinated M. lewisii and the hummingbird-pollinated M. cardinalis. Although 6% is a modest effect size in molecular genetic terms, in evolutionary genetic terms a selection coefficient (s) of 0.06 (130 greater visits to the wild-type plant / 2202 total visits) would sweep the beneficial allele to fixation very quickly in natural populations (Hartl and Clark, 1997), so we designed our pollinator visitation experiments to detect a difference in visitation as small as 5%. Assuming an infinite population size, the probability of fixation of the allele is 2s, or 12%; an effective population size greater than five individuals would allow selection to exceed drift as an evolutionary force at this locus (Ne = 1/(4s) = 4.17).
Why does the loss of D-limonene and β-myrcene have no effect on bumblebee visitation? First, the T2 plants used in the greenhouse experiment had surprisingly high levels of E-β-ocimene, perhaps due to rerouting of a common pool of the shared precursor geranyl pyrophosphate (GPP). Terpene synthesis is a flexible but complex process, and buildups of precursors can be utilized by alternate metabolic pathways (Gang, 2005). Given the much higher emission of E-β-ocimene in the MlLMS-RNAi transgenic plants, M. lewisii may be prone to this effect. As the RNAi technique used here is an analogous (but weaker) representation of the phenotypes resulting from a loss-of-function mutation in a wild population, fluctuations in volatile production as found here are reflective of the system’s physiology and the effects that might occur in a natural setting.
It is also possible that the high production of D-limonene and β-myrcene in M. lewisii serves another function within the plant, such as defense against herbivores, nectar robbers, or disease (Kessler et al., 2013), as these volatiles are known anti-herbivory compounds (Levin, 1976). Although the three volatiles have similar physical properties, D-limonene and β-myrcene may serve to mediate long-distance attraction at the patch level rather than at the level of the individual flower; long-distance attraction has been shown to be important for honeybee (Apis mellifera) navigation (Bogdany and Taber, 1979). The high production of D-limonene and β-myrcene may be a remnant of some previous pollination syndrome, environmental context, or merely the byproduct of some other metabolic process within the plant. Similarly, although a significant effect on bumblebee visitation was seen with the loss of E-β-ocimene, it is possible that the main role of this volatile may lie elsewhere (Kessler et al., 2013), for example in herbivory defense (Arimura et al., 2004) with a secondary role in the attraction of bumblebee pollinators. Data on herbivory, florivory, or pathogen infestation in wild populations of M. lewisii and M. cardinalis are currently lacking, limiting our ability to speculate on these possibilities. Future field experiments will increase our understanding of the multiple roles these volatiles may be playing in M. lewisii and M. cardinalis.
Finally, it is possible that these effects differ from those that would be found with wild Bombus vosnesenskii. However, both species are generalist floral visitors, and the M. lewisii scent elicits similar olfactory responses in both bee species. Moreover, B. impatiens has been used as a model for bumblebee-flower interactions in other systems, including those involving B. vosnesenskii (Bodbyl Roels and Kelly, 2011), thus we feel that B. impatiens is an excellent model for these experiments (see SI Materials and Methods for a full explanation). Although these results differ in detail from those we found in previous behavioral experiments with artificial and extracted floral scents, in which all three monoterpenes were required for maximum bumblebee response (Byers et al., 2014), the greenhouse experiments offer a more realistic assay for the effect of scent on pollinators by allowing them to integrate multiple floral cues.
What role does scent play in pollinator interaction within this system? Many studies have shown that scent plays a strong role in landing decisions by diurnal pollinators such as bumblebees and honeybees (Butler, 1951; Galen and Kevan, 1980; Galen and Kevan, 1983; Lunau, 1992; Majetic et al., 2009; Dötterl and Vereecken, 2010) – the initial approach may be guided by patch-level visual signals, followed by a visually-guided approach to an individual flower. At that point, the final landing decision may be influenced by floral scent, especially in relatively weakly scented flowers such as M. lewisii (Dötterl and Vereecken, 2010; Parachnowitsch et al., 2012). Therefore, even in the densely-flowered greenhouse experiments, signals such as the presence or absence of E-β-ocimene may play a significant role in final landing decisions. Additionally, densities in the greenhouse experiments were similar to those found in wild populations of M. lewisii, which grows along montane streambeds in large clusters, so the dense greenhouse conditions are a better indicator of the potential effect of a single change in scent in a wild population.
How might a loss-of-function allele of OS promote a pollinator switch from bumblebees to hummingbirds? Hummingbirds have a very limited sense of smell (Ioalé and Papi, 1989), and retain scent information very poorly (Goldsmith and Goldsmith, 1982), so the loss of scent in a hummingbird-pollinated flower such as M. cardinalis (an “anti-bee” but not “pro-bird” shift, to use the language in Castellanos et al., 2004) would likely have no fitness cost, and might even increase fitness by discouraging bumblebee visitors from transferring heterospecific pollen to the stigma and carrying away nectar or pollen. In combination with the difference in visual signals and mechanical access found between M. lewisii and M. cardinalis, such a loss of E-β-ocimene might serve to reinforce visitation behavior. Whether these changes in floral volatiles evolved in allopatry or as reinforcement during secondary contact is unclear; investigating the volatile profiles and orthologous terpene synthase genes of other species in Mimulus section Erythranthe may provide some insight into this question.
The fact that the OS polymorphism between M. lewisii and M. cardinalis is in a structural gene contradicts the current thinking that genes involved in prezygotic reproductive isolation – often referred to as “speciation genes” (Coyne, 1992) or “barrier genes” (Noor and Feder, 2006) – are nearly always regulatory genes (Rieseberg and Blackman, 2010). However, the limited number of genes with known effects in prezygotic reproductive isolation should preclude any general conclusions from being drawn about this process. We would suggest, however, that the potential for structural genes to contribute to this process should not be ignored.
Although many systems used in the study of floral volatiles have relatively strong scents that are detectable by the human nose, scent can also be a factor in reproductive isolation in systems where it is easily missed, as in Mimulus. The role of strong emissions of floral volatiles in attracting nighttime pollinators from a distance is well documented (Raguso and Willis, 2003). The potential role of changes in floral scent in pollinator-mediated reproductive isolation involving generalist, daytime pollinators such as bumblebees is largely unknown, and no examples integrating floral scent genetics and pollinator reproductive isolation in sister species with generalist pollinators have been reported. Some authors have commented that the role of floral scent in reproductive isolation is questionable in generalist cases, as floral scents thus serve less as “private channels” and pollinators are attracted to multiple floral scent profiles (Schiestl and Ayasse, 2002).
Floral scent should be considered as an attractive factor even in generalist systems, along with more easily-measured visual signals such as floral color and pattern. Here, the sister species M. lewisii and M. cardinalis can be used as a model for the study of reproductive isolation involving floral volatiles – one can begin by looking at species-specific differences, identifying critical volatiles within a complex mixture via electrophysiological and behavioral assays (Riffell et al., 2013; Byers et al., 2014). Then, studies can proceed by determining the genetic basis of these phenotypic differences, creating high-resolution genetic materials (near-isogenic lines, transgenics), and, finally, assaying of the results of these genetic changes in ecologically relevant greenhouse or field settings. Nearly all previous studies of the role of floral volatiles in speciation have only answered a subset of these questions, but our work with Mimulus, an emerging model system, shows that a comprehensive, integrative study is possible.
Experimental Procedures
QTL and fine mapping
Volatiles were first assayed in triplicate in an F1 cross of M. lewisii inbred line LF10 and M. cardinalis inbred line CE10 (LF10 × CE10) and compared with previous results for the parent lines (9 samples each; see Byers et al., 2014) using a Mann-Whitney U test. A coarse mapping population consisting of 100 individuals of a cross between LF10 and CE10, backcrossed to CE10 [(LF10 × CE10) × CE10], was then constructed. Headspace volatiles were collected in the manner described in Byers et al., 2014 (see also SI Materials and Methods and methods below) and assayed for emission rates of D-limonene, β-myrcene, and E-β-ocimene. Pearson correlation coefficients were calculated pairwise for the three scents to investigate potential linkage or pleiotropy. A subset of 24 backcross plants with the two most divergent phenotypes (high D-limonene/β-myrcene and low E-β-ocimene; low D-limonene/β-myrcene and high E-β-ocimene) were screened at 34 indel markers evenly spaced across the genome (Table S4) with the intent of creating a low-resolution quantitative trait locus (QTL) map. However, it was clear from inspection of the genotypic and phenotypic data that the emission of D-limonene/β-myrcene and E-β-ocimene were, to a first approximation, segregating as Mendelian traits.
A larger backcross population (N = 768) was constructed and screened with markers flanking the putative LIMONENE-MYRCENE SYNTHASE (M02_510 and M02_1500), and flanking the putative OCIMENE SYNTHASE (sc4_2325K and M13_2620) (Table S4). Markers used in the mapping process were developed from M. lewisii and M. cardinalis genome sequences, and amplify codominant markers in the backcross. To reduce the effort required to score flowers for volatile production, only those backcross plants with informative recombinations between markers flanking LMS (N = 107) or OS (N = 52) were phenotyped for scent, using a direct extraction assay from flowers rather than the more labor- and time-intensive headspace collection method. For further details, see Supplemental Information.
The Mimulus guttatus genomic region corresponding to the M. lewisii region containing LMS or OS was examined, and candidate genes were identified based upon their map position relative to the flanking molecular markers and the M. guttatus annotation. For the LMS locus controlling D-limonene and β-myrcene emission, primers were designed to amplify indel polymorphisms in two of the terpene synthases/cyclases on M. guttatus scaffold 89. The candidate genes were designated LC250K and TS306K (see Table S5 for all primers). No recombination events were observed among the two candidate genes and the putative LMS, defining a candidate region of less than 0.1 cM. For the OS locus controlling E-β-ocimene emission, primers were designed to amplify an indel polymorphism in a terpene synthase designated TS2538 on M. guttatus scaffold 4. No recombination events were observed between TS2538 and OS, defining a candidate region of less than 0.1 cM.
In vitro assay for terpene synthase activity
For details, see Supplemental Information and Fäldt et al. (2003).
Terpene synthase expression in vivo
Total RNA was extracted from flower buds collected at 5, 8, 10, 15, and 20mm stages in M. lewisii and equivalent stages in M. cardinalis. Total RNA was extracted from open flowers of both species. cDNA was prepared from total RNA extracts using the SuperScript III First-Strand Synthesis System (Invitrogen). RT-PCR for both loci for both species was performed with these cDNA, using MlUBC as a control for background expression levels as described in Yuan et al. (2013c). Primers used were the following: MlLMS RNAi forward/reverse for MlLMS, McLMS forward/reverse for McLMS; MlOS RNAi sense forward/reverse for MlOS and McOS; and MlUBC foward/reverse for MlUBC.
Construction of MlLMS and MlOS RNAi transgenic M. lewisii
Transgenesis was done in the M. lewisii background, as insect pollination is inferred to be the ancestral state in this clade (Beardsley et al., 2003). Hairpin RNA interference (RNAi) transgenes targeted to knock down the expression of MlLMS or MlOS were constructed in pFGC5941 (Kerschen et al., 2004; Arabidopsis Biological Resource Center, CD3-447) as described in (Yuan et al., 2013c). In each case, target specificity of the RNAi fragment was assured by BLAST search against the M. lewisii LF10 genome sequence. For MlLMS, a 106 bp fragment of M. lewisii cDNA was amplified and directionally cloned into the pFGC5941 NcoI/AscI (sense) and BamHI/XbaI (antisense) sites. For MlOS, a 289 bp sense fragment was amplified and directionally cloned into the NcoI/AscI site of pFGC5941. A 180 bp antisense fragment (entirely within the 289 bp NcoI/AscI amplicon) was amplified and directionally cloned into the BamHI/XbaI site. Constructs were verified by sequencing, then electroporated separately into Agrobacterium tumefaciens strain GV3101 and used for in planta transformation of M. lewisii LF10 following (Yuan et al., 2013c).
Greenhouse experiments
For details of experimental design, see Supplemental Information. The layout of the greenhouse experiments, including randomized plant positions, is depicted in Figure S2.
Observations of bumblebee behavior were recorded for the first six hours of the first three days by two observers using voice recorders, each following one or two bumblebees at a time. At the start of each day prior to the first bumblebee activity, old flowers were removed and newly opened flowers were counted and equalized between the two types of plants (wild-type and RNAi transgenic) to ensure that bumblebees had an equal chance of encountering a given flower of each type on each day.
Data were transcribed and analyzed for preference (proportion of total visits) and constancy. For constancy, visits were “collapsed” to the plant level – i.e., multiple visits to one plant in sequence were reduced to a single visit, since flower numbers were unequal between plants and flowers were often tightly clustered. Bumblebees were only used for constancy analysis if they visited ten or more plants in a foraging bout. Constancy was calculated using Bateman’s method (described in Waser, 1986), which is independent of preference; equalizing flowers at the start of the day gave pollinators equal access to each type, as required by this metric. To determine if observed constancy was different from the null expectation, the same foraging data were used in a permutation test with shuffled plant identities, repeated 100,000 times; the fraction of the simulations with a greater than observed deviation from zero (complete randomness) was used to estimate the p-value.
Supplementary Material
Figure S1: Protein sequences of MlOS and McOS with differences highlighted.
Figure S2: Schematic of the greenhouse experimental setup.
Table S1: Products of the in vitro terpene synthase assays.
Table S2: Volatile production in the 2–4 best T1 plants recovered from RNAi experiments.
Table S3: Volatile production in T2 plants from four separate T1 parents recovered from RNAi knockdown of MlLMS.
Table S4: Molecular markers used during QTL and fine mapping.
Table S5: Sequences of oligonucleotide primers used.
Acknowledgments
We thank D. Ewing, P. Beeman, E. Forbush, J. Milne, N. Kurashige, M. Kovic, and B. Watson for greenhouse plant care, and M. Sargent for assistance in greenhouse experiments. The manuscript was improved greatly by helpful and thorough critiques from D. Tholl and two anonymous reviewers. This work was supported by a National Institutes of Health grant to HDB (5R01GM088805), National Science Foundation grants to HDB (FIBR 0328636; IOS 1209340), JAR (IOS 0822709; DBI 1121692), and KJRPB (IOS 1209340), and a NSF Graduate Research Fellowship to KJRPB (DGE 0718124). KJRPB is also supported by an ARCS Foundation Fellowship from the Seattle Chapter of the Achievement Rewards for College Scientists Foundation and the Melinda Denton Writing Fellowship.
Footnotes
Accession numbers for sequences:
KF857265, M. lewisii LIMONENE-MYRCENE SYNTHASE gene, complete cds
KF857264, M. lewisii LIMONENE-MYRCENE SYNTHASE mRNA, complete cds
AHI50308, M. lewisii LIMONENE-MYRCENE SYNTHASE protein product
KM659024, M. cardinalis LIMONENE-MYRCENE SYNTHASE mRNA, exon3
KF857262, M. lewisii OCIMENE SYNTHASE mRNA, complete cds
AHI50306, M. lewisii OCIMENE SYNTHASE protein product
KF857263, M. cardinalis OCIMENE SYNTHASE mRNA, complete cds
AHI50307, M. cardinalis OCIMENE SYNTHASE protein product
Supplemental Materials and Methods: Description of volatile headspace collection and analysis, volatile extraction collection and scoring, in vitro assays for terpene synthase activity, and greenhouse experiments.
Contributor Information
Kelsey J.R.P. Byers, Email: kelsey.byers@systbot.uzh.ch.
James P. Vela, Email: jamesv_42@hotmail.com.
Foen Peng, Email: peng.foen@gmail.com.
Jeffrey A. Riffell, Email: jriffell@uw.edu.
References
- Arimura G, Ozawa R, Kugimiya S, Takabayashi J, Bohlmann J. Herbivore-induced defense response in a model legume. Two-spotted spider mites induce emission of (E)-β-ocimene and transcript accumulation of (E)-β-ocimene synthase in Lotus japonicas. Plant Physiol. 2004;135(4):1976–1983. doi: 10.1104/pp.104.042929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beardsley PM, Yen A, Olmstead RG. AFLP phylogeny of Mimulus section Erythranthe and the evolution of hummingbird pollination. Evolution. 2003;57(6):1397–1410. doi: 10.1111/j.0014-3820.2003.tb00347.x. [DOI] [PubMed] [Google Scholar]
- Bodbyl Roels SA, Kelly JK. Rapid evolution caused by pollinator loss in Mimulus guttatus. Evolution. 2011;65(9):2541–2552. doi: 10.1111/j.1558-5646.2011.01326.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogdany FJ, Taber S., III The significance of odor for bees orienting across a canyon. Apidologie. 1979;10(1):55–62. [Google Scholar]
- Bohlmann J, Martin D, Oldham NJ, Gershenzon J. Terpenoid secondary metabolism in Arabidopsis thaliana: cDNA cloning, characterization, and functional expression of a myrcene/(E)-β-ocimene synthase. Arch Biochem Biophys. 2000;375(2):261–269. doi: 10.1006/abbi.1999.1669. [DOI] [PubMed] [Google Scholar]
- Bradshaw HD, Jr, Otto KG, Frewen BE, McKay JK, Schemske DW. Quantitative trait loci affecting differences in floral morphology between two species of monkey flower (Mimulus) Genetics. 1998;149:367–382. doi: 10.1093/genetics/149.1.367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradshaw HD, Jr, Schemske DW. Allele substitution at a flower colour locus produces a pollinator shift in monkeyflowers. Nature. 2003;426:176–178. doi: 10.1038/nature02106. [DOI] [PubMed] [Google Scholar]
- Butler CG. The importance of perfume in the discovery of food by the worker honeybee (Apis mellifera L.) Proc R Soc Lond B Biol Sci. 1951;138:403–413. [Google Scholar]
- Byers KJRP, Bradshaw HD, Jr, Riffell JA. Three floral volatiles contribute to differential pollinator attraction in monkeyflowers (Mimulus) J Exp Biol. 2014;217(4):614–623. doi: 10.1242/jeb.092213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castellanos MC, Wilson P, Thomson JD. ‘Anti-bee’ and ‘pro-bird’ changes during the evolution of hummingbird pollination in Penstemon flowers. J Evol Biol. 2004;17(4):876–885. doi: 10.1111/j.1420-9101.2004.00729.x. [DOI] [PubMed] [Google Scholar]
- Chen F, Tholl D, D’Auria JC, Farooq A, Pichersky E, Gershenzon J. Biosynthesis and emission of terpenoid volatiles from Arabidopsis flowers. Plant Cell. 2003;15:481–494. doi: 10.1105/tpc.007989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chess SKR, Raguso RA, LeBuhn G. Geographic divergence in floral morphology and scent in Linanthus dichotomus (Polemoniaceae) Am J Bot. 2008;95(12):1652–1659. doi: 10.3732/ajb.0800118. [DOI] [PubMed] [Google Scholar]
- Coyne JA. Genetics and speciation. Nature. 1992;355:511–515. doi: 10.1038/355511a0. [DOI] [PubMed] [Google Scholar]
- Dexter R, Qualley A, Kish CM, Ma CJ, Koeduka T, Nagegowda DA, Dudareva N, Pichersky E, Clark D. Characterization of a petunia acetyltransferase involved in the biosynthesis of the floral volatile isoeugenol. Plant J. 2007;49(2):265–275. doi: 10.1111/j.1365-313X.2006.02954.x. [DOI] [PubMed] [Google Scholar]
- Dötterl S, Vereecken NJ. The chemical ecology and evolution of bee-flower interactions: a review and perspectives. Can J Zool. 2010;88:668–697. [Google Scholar]
- Dudareva N, Cseke L, Blanc VM, Pichersky E. Evolution of floral scent in Clarkia: novel patterns of S-linalool synthase gene expression in the C. breweri flower. Plant Cell. 1996;8:1137–1148. doi: 10.1105/tpc.8.7.1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dudareva N, D’Auria JC, Nam KH, Raguso RA, Pichersky E. Acetyl-CoA: benzylalcohol acetyltransferase - an enzyme involved in floral scent production in Clarkia breweri. Plant J. 1998;14(3):297–304. doi: 10.1046/j.1365-313x.1998.00121.x. [DOI] [PubMed] [Google Scholar]
- Dudareva N, Martin D, Kish CM, Kolosova N, Gorenstein N, Fäldt J, Miller B, Bohlmann J. (E)-β-Ocimene and myrcene synthase genes of floral scent biosynthesis in snapdragon: function and expression of three terpene synthase genes of a new terpene synthase subfamily. Plant Cell. 2003;15(5):1227–1241. doi: 10.1105/tpc.011015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dudareva N, Murfitt LM, Mann CJ, Gorenstein N, Kolosova N, Kish CM, Bohman C, Wood K. Developmental regulation of methyl benzoate biosynthesis and emission in snapdragon flowers. Plant Cell. 2000;12(6):949–961. doi: 10.1105/tpc.12.6.949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dudareva N, Pichersky E, Gershenzon J. Biochemistry of plant volatiles. Plant Physiol. 2004;135:1893–1902. doi: 10.1104/pp.104.049981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fäldt J, Martin D, Miller B, Rawat S, Bohlmann J. Traumatic resin defense in Norway spruce (Picea abies): Methyl jasmonate-induced terpene synthase gene expression, and cDNA cloning and functional characterization of (+)-3-carene synthase. Plant Mol Biol. 2003;51:119–133. doi: 10.1023/a:1020714403780. [DOI] [PubMed] [Google Scholar]
- Fenster CB, Armbruster WS, Wilson P, Dudash MR, Thomson JD. Pollination syndromes and floral specialization. Annu Rev Ecol Evol Syst. 2004;35:375–403. [Google Scholar]
- Galen C, Kevan PG. Scent and color, floral polymorphisms and pollination biology in Polemonium viscosum Nutt. The American Midland Naturalist. 1980;104(2):281–289. [Google Scholar]
- Galen C, Kevan PG. Bumblebee foraging and floral scent dimorphism: Bombus kirbyellus Curtis (Hymenoptera: Apidae) and Polemonium viscosum Nutt. (Polemoniaceae) Can J Zool. 1983;61:1207–1213. [Google Scholar]
- Gang DR. Evolution of flavors and scents. Annu Rev Plant Biol. 2005;56:301–325. doi: 10.1146/annurev.arplant.56.032604.144128. [DOI] [PubMed] [Google Scholar]
- Goldsmith KM, Goldsmith TH. Sense of smell in the Black-chinned Hummingbird. Condor. 1982;84:237–238. [Google Scholar]
- Grant V. Pollination systems as isolating mechanisms in angiosperms. Evolution. 1949;3:82–97. doi: 10.1111/j.1558-5646.1949.tb00007.x. [DOI] [PubMed] [Google Scholar]
- Gupta AK, Akhtar T, Widmer A, Pichersky E, Schiestl FP. Identification of white campion (Silene latifolia) guaiacol O-methyltransferase involved in the biosynthesis of veratrole, a key volatile for pollinator attraction. BMC Plant Biol. 2012;12:158. doi: 10.1186/1471-2229-12-158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harder LD, Johnson SD. Darwin’s beautiful contrivances: evolutionary and functional evidence for floral adaptation. New Phytol. 2009;183(3):530–545. doi: 10.1111/j.1469-8137.2009.02914.x. [DOI] [PubMed] [Google Scholar]
- Hartl DL, Clark AG. Principles of Population Genetics. 4. Sinaur Associates Inc; Sunderland, MA: 1997. [Google Scholar]
- Hiesey WM, Nobs MA, Björkman O. Experimental Studies on the Nature of Species. V Biosystematics, genetics, and physiological ecology of the Erythranthe section of Mimulus. Carnegie Inst Washington Publ. 1971;628:1–213. [Google Scholar]
- Huber FK, Kaiser R, Sauter W, Schiestl FP. Floral scent emission and pollinator attraction in two species of Gymnadenia (Orchidaceae) Oecologia. 2005;142(4):564–575. doi: 10.1007/s00442-004-1750-9. [DOI] [PubMed] [Google Scholar]
- Ioalé P, Papi F. Olfactory bulb size, odor discrimination and magnetic insensitivity in hummingbirds. Physiol Behav. 1989;45:995–999. doi: 10.1016/0031-9384(89)90227-8. [DOI] [PubMed] [Google Scholar]
- Kay KM, Sargent RD. The role of animal pollination in plant speciation: integrating ecology, geography, and genetics. Ann Rev Ecol Evol Syst. 2009;40:637–656. [Google Scholar]
- Kerschen A, Napoli CA, Jorgensen RA, Müller AE. Effectiveness of RNA interference in transgenic plants. FEBS Letters. 2004;566:223–228. doi: 10.1016/j.febslet.2004.04.043. [DOI] [PubMed] [Google Scholar]
- Kessler D, Diezel C, Clark DG, Colquhoun TA, Baldwin IT. Petunia flowers solve the defence/apparency dilemma of pollinator attraction by deploying complex floral blends. Ecol Letters. 2013;16(3):299–306. doi: 10.1111/ele.12038. [DOI] [PubMed] [Google Scholar]
- Klahre U, Gurba A, Hermann K, Saxenhofer M, Bossolini E, Guerin PM, Kuhlemeier C. Pollinator choice in Petunia depends on two major genetic loci for floral scent production. Curr Biol. 2011;21:730–739. doi: 10.1016/j.cub.2011.03.059. [DOI] [PubMed] [Google Scholar]
- Koeduka T, Fridman E, Gang DR, Vassão DG, Jackson BL, Kish CM, Orlova I, Spassova SM, Lewis NG, Noel JP, Baiga TJ, Dudareva N, Pichersky E. Eugenol and isoeugenol, characteristic aromatic constituents of spices, are biosynthesized via reduction of a coniferyl alcohol ester. Proc Natl Acad Sci USA. 2006;103(26):10128–10133. doi: 10.1073/pnas.0603732103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levin DA. The chemical defenses of plants to pathogens and herbivores. Annu Rev Ecol Syst. 1976;7:121–159. [Google Scholar]
- Lunau K. Innate recognition of flowers by bumble bees: orientation of antennae to visual stamen signals. Can J Zool. 1992;70:2139–2144. [Google Scholar]
- Majetic CJ, Raguso RA, Ashman TL. The sweet smell of success: floral scent affects pollinator attraction and seed fitness in Hesperis matronalis. Funct Ecol. 2009;23:480–487. [Google Scholar]
- Mant J, Peakall R, Schiestl FP. Does selection on floral odor promote differentiation among populations and species of the sexually deceptive orchid genus Ophrys? Evolution. 2005;59(7):1449–1463. [PubMed] [Google Scholar]
- Nieuwenhuizen NJ, Green SA, Chen X, Bailleul EJD, Matich AJ, Wang MY, Atkinson RG. Functional genomics reveals that a compact terpene synthase gene family can account for terpene volatile production in apple. Plant Physiol. 2013;161:787–804. doi: 10.1104/pp.112.208249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noor MAF, Feder JL. Speciation genetics: evolving approaches. Nat Rev Genet. 2006;7:851–861. doi: 10.1038/nrg1968. [DOI] [PubMed] [Google Scholar]
- Orlova I, Marshall-Colón A, Schnepp J, Wood B, Varbanova M, Fridman E, Blakeslee JJ, Peer WA, Murphy AS, Rhodes D, Pichersky E, Dudareva N. Reduction of benzenoid synthesis in petunia flowers reveals multiple pathways to benzoic acid and enhancement in auxin transport. Plant Cell. 2006;18(12):3458–3475. doi: 10.1105/tpc.106.046227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parachnowitsch AL, Raguso RA, Kessler A. Phenotypic selection to increase floral scent emission, but not flower size or colour in bee-pollinated Penstemon digitalis. New Phytol. 2012;195:667–675. doi: 10.1111/j.1469-8137.2012.04188.x. [DOI] [PubMed] [Google Scholar]
- Peakall R, Ebert D, Poldy J, Barrow RA, Francke W, Bower CC, Schiestl FP. Pollinator specificity, floral odour chemistry and the phylogeny of Australian sexually deceptive Chiloglottis orchids: implications for pollinator driven speciation. New Phytol. 2010;188(2):437–450. doi: 10.1111/j.1469-8137.2010.03308.x. [DOI] [PubMed] [Google Scholar]
- Pichersky E, Lewinsohn E, Croteau R. Purification and characterization of S-linalool synthase, an enzyme involved in the production of floral scent in Clarkia breweri. Arch Biochem Biophys. 1995;316(2):803–807. doi: 10.1006/abbi.1995.1107. [DOI] [PubMed] [Google Scholar]
- Raguso RA. Start making scents: the challenge of integrating chemistry into pollination ecology. Entomol Exp Appl. 2008a;128(1):196–207. [Google Scholar]
- Raguso RA. Wake up and smell the roses: the ecology and evolution of floral scent. Annu Rev Ecol Evol Syst. 2008b;39:549–569. [Google Scholar]
- Raguso RA, Willis MA. Hawkmoth pollination in Arizona’s Soronan Desert: Behavioral responses to floral traits. In: Boggs CL, et al., editors. Evolution and ecology taking flight, Butterflies as model systems. University of Chicago Press; Chicago: 2003. pp. 43–65. [Google Scholar]
- Ramsey J, Bradshaw HD, Jr, Schemske DW. Components of reproductive isolation between the monkeyflowers Mimulus lewisii and M. cardinalis (Phrymaceae) Evolution. 2003;57(7):1520–1534. doi: 10.1111/j.0014-3820.2003.tb00360.x. [DOI] [PubMed] [Google Scholar]
- Rausher MD. Evolutionary transitions in floral color. Int J Plant Sci. 2008;169(1):7–21. [Google Scholar]
- Rieseberg LH, Blackman BK. Speciation genes in plants. Ann Bot. 2010;106(3):439–455. doi: 10.1093/aob/mcq126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riffell JA, Lei H, Abrell L, Hildebrand JG. Neural basis of a pollinator’s buffet: olfactory specialization and learning in Manduca sexta. Science. 2013;339(6116):200–204. doi: 10.1126/science.1225483. [DOI] [PubMed] [Google Scholar]
- Schemske DW, Bradshaw HD., Jr Pollinator preference and the evolution of floral traits in monkeyflowers (Mimulus) Proc Natl Acad Sci USA. 1999;96(21):11910–11915. doi: 10.1073/pnas.96.21.11910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiestl FP, Ayasse M. Do changes in floral odor cause speciation in sexually deceptive orchids? Plant Syst Evol. 2002;234:111–119. [Google Scholar]
- Schiestl FP, Peakall R. Two orchids attract different pollinators with the same floral odour compound: ecological and evolutionary implications. Funct Ecol. 2005;19(4):674–680. [Google Scholar]
- Stebbins GL. Adaptive radiation of reproductive characteristics in angiosperms, I, Pollination mechanisms. Annu Rev Ecol Syst. 1970;1:307–326. [Google Scholar]
- Vereecken NJ, Cozzolino S, Schiestl FP. Hybrid floral scent novelty drives pollinator shift in sexually deceptive orchids. BMC Evol Biol. 2010;10(1):103. doi: 10.1186/1471-2148-10-103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waelti MO, Muhlemann JK, Widmer A, Schiestl FP. Floral odour and reproductive isolation in two species of Silene. J Evol Biol. 2008;21(1):111–121. doi: 10.1111/j.1420-9101.2007.01461.x. [DOI] [PubMed] [Google Scholar]
- Wang J, Pichersky E. Characterization of S-adenosyl-l-methionine: (iso)eugenol O-methyltransferase involved in floral scent production in Clarkia breweri. Arch Biochem Biophys. 1998;349(1):153–160. doi: 10.1006/abbi.1997.0452. [DOI] [PubMed] [Google Scholar]
- Waser NM. Flower constancy: definition, cause, and measurement. Am Nat. 1986;127(5):593–603. [Google Scholar]
- Whitehead MR, Peakall R. Integrating floral scent, pollination ecology and population genetics. Funct Ecol. 2009;23:863–874. [Google Scholar]
- Xu S, Schlüter PM, Grossniklaus U, Schiestl FP. The genetic basis of pollinator adaptation in a sexually deceptive orchid. PLoS Genet. 2012;8(8):e1002889. doi: 10.1371/journal.pgen.1002889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan YW, Byers KJRP, Bradshaw HD., Jr The genetic control of flower-pollinator specificity. Curr Opin Plant Biol. 2013a;16(4):422–428. doi: 10.1016/j.pbi.2013.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan YW, Sagawa JM, Di Stilio VS, Bradshaw HD., Jr Bulk segregant analysis of an induced floral mutant identifies a MIXTA-like R2R3 MYB controlling nectar guide formation in Mimulus lewisii. Genetics. 2013b;194(2):523–528. doi: 10.1534/genetics.113.151225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuan YW, Sagawa JM, Young RC, Christensen BJ, Bradshaw HD., Jr Genetic dissection of a major anthocyanin QTL contributing to pollinator-mediated reproductive isolation between sister species of Mimulus. Genetics. 2013c;194(1):255–263. doi: 10.1534/genetics.112.146852. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1: Protein sequences of MlOS and McOS with differences highlighted.
Figure S2: Schematic of the greenhouse experimental setup.
Table S1: Products of the in vitro terpene synthase assays.
Table S2: Volatile production in the 2–4 best T1 plants recovered from RNAi experiments.
Table S3: Volatile production in T2 plants from four separate T1 parents recovered from RNAi knockdown of MlLMS.
Table S4: Molecular markers used during QTL and fine mapping.
Table S5: Sequences of oligonucleotide primers used.