Abstract
The single cell gel electrophoresis assay, also known as the comet assay, is a versatile method for measuring many classes of DNA damage, including base damage, abasic sites, single strand breaks and double strand breaks. However, limited throughput and difficulties with reproducibility have limited its utility, particularly for clinical and epidemiological studies. To address these limitations, we created a microarray comet assay. The use of a micrometer scale array of cells increases the number of analysable comets per square centimetre and enables automated imaging and analysis. In addition, the platform is compatible with standard 24- and 96-well plate formats. Here, we have assessed the consistency and sensitivity of the microarray comet assay. We showed that the linear detection range for H2O2-induced DNA damage in human lymphoblastoid cells is between 30 and 100 μM, and that within this range, inter-sample coefficient of variance was between 5 and 10%. Importantly, only 20 comets were required to detect a statistically significant induction of DNA damage for doses within the linear range. We also evaluated sample-to-sample and experiment-to-experiment variation and found that for both conditions, the coefficient of variation was lower than what has been reported for the traditional comet assay. Finally, we also show that the assay can be performed using a 4× objective (rather than the standard 10× objective for the traditional assay). This adjustment combined with the microarray format makes it possible to capture more than 50 analysable comets in a single image, which can then be automatically analysed using in-house software. Overall, throughput is increased more than 100-fold compared to the traditional assay. Together, the results presented here demonstrate key advances in comet assay technology that improve the throughput, sensitivity, and robustness, thus enabling larger scale clinical and epidemiological studies.
Introduction
The single cell gel electrophoresis assay, also known as the comet assay, has become increasingly popular since it was first developed by Ostling and Johansan in 1984 (1–3). Based on the principle that damaged DNA migrates more readily than undamaged DNA when electrophoresed, DNA damage can be quantified by analysing images of electrophoresed nuclei. The comet assay enables detection of several classes of damage, including alkali sensitive sites, abasic sites, single and double strand breaks. Although the comet assay has become a broadly accepted approach for DNA damage analysis, throughput, sensitivity and reproducibility have been suboptimal. Many researchers have worked over the past decades to modify the original protocol to cater for the requirement of large-scale studies. These modifications range from assay protocol optimisation to improve sensitivity (4–7), to inclusion of internal standards to overcome inconsistency issues across samples and laboratories (8–11). Others have worked on advancing the hardware of assay to improve and facilitate processing of larger sample sizes (12, 13). Here, we describe a new approach for improving the comet assay that involves creation of a microarray of comets. Compared to the traditional comet assay, the ‘CometChip’ offers significantly greater throughput as well as robust sensitivity.
To create a CometChip, microfabrication technologies are used to create an array of silicon posts. The mold is then pressed into molten agarose, and after gelling and removing the mold, an array of microwells is formed in agarose. A solution of mammalian cells is then placed on top of the agarose and cells are then loaded into the well by gravity. Excess cells can then be removed by sheer force and an array of cells embedded in agarose remains (14, 15) (Figure 1A). The microarray approach not only improves throughput, but it also demonstrates significant advances in data quality (14, 15). The CometChip has been used for a many applications since it was first developed. These studies range from determining specific genotoxic impact of chemicals, to screening for the DNA damaging effects of nanoparticles (16–18). The CometChip has also opened doors to epidemiological studies that require processing of dozens of samples in parallel.
Figure 1.
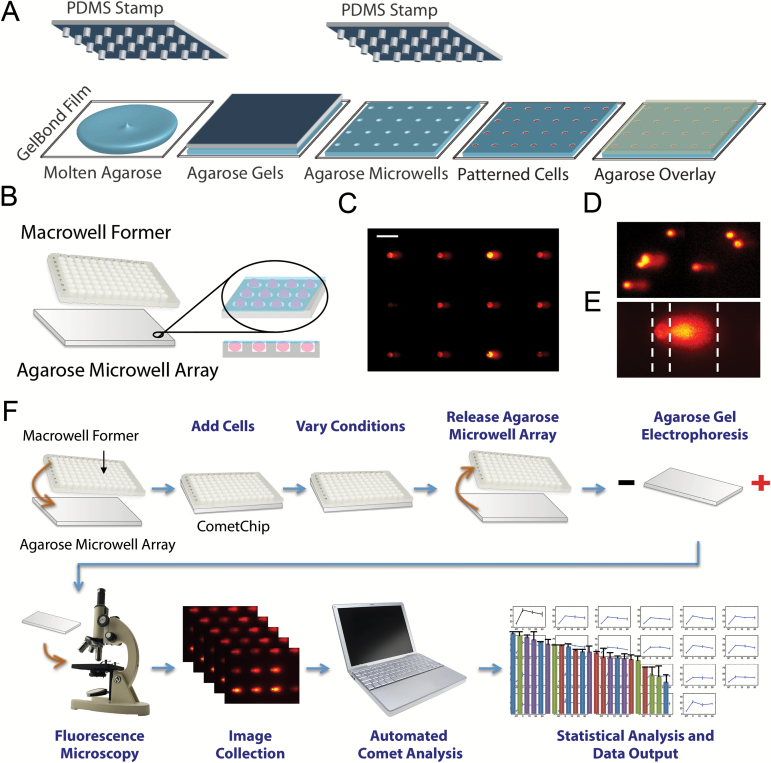
Creating and using the CometChip. (A) Microfabricated mold creates precision microwells that can be loaded with single or groups of cells. Schematics is adapted from (16). (B) Assembly of CometChip with zoomed-view of embedded wells. (C) Representative image of arrayed microwell comets collected using 10× objective lens. Scale bar indicates 100 μm. (D) Representative image of randomly distributed traditional comets. (E) Comet analysis delineating head and tail. The three dotted lines mark beginning of head, end of head/start of tail, end of tail, respectively. (F) Experimental flow of CometChip assembly and usage.
To reveal the strengths and weaknesses of the CometChip, here we have assessed the sensitivity of the assay and the variability among comets and among samples. We have determined the linear range of detection in response to H2O2 and ionizing radiation (IR). We then explored the utility of lower magnification imaging with the aim of improving throughput, and to learn about consistency, we have assessed well-to-well, chip-to-chip and experiment-to-experiment variability. Finally, as previous studies have shown that scoring more comets can improve sensitivity of the traditional comet assay (4), we have also examined the impact of the number of comets scored on the sensitivity the assay. Together, the results described here provide detailed assessment of the sensitivity, reproducibility, and robustness of the micropatterned comet assay.
Materials and methods
Cell culture
TK6 human lymphoblasts were cultured in suspension in 1× Roswell Park Memorial Institute (RPMI) medium 1640 with l-glutamine (Invitrogen) supplemented with 10% horse serum (Invitrogen) and 1% penicillin and streptomycin (100 units/ml penicillin, 100 µg/ml streptomycin) (Invitrogen) at 37°C, 5% CO2.
CometChip fabrication
CometChip were fabricated using a Polydimethylsiloxane (PDMS, Dow Corning) mold made in house, as described in Wood et al. (14) and Weingeist et al. (15) Briefly, molten 1% normal melting point agarose (Invitrogen) was applied to a sheet of GelBond film (Lonza), and allowed to gel with the PDMS mold on top. Removal of the PDMS mold reveals a ~300 μm thick gel with arrayed microwells (Figure 1A). The microwell gel was then clamped between a glass plate and a bottomless 96-well titer plate (Greiner BioOne) to create the 96-well CometChip (Figure 1B). At least 50 000 cells were added to each well of the 24-well Cometchip and allowed to settle by gravity in complete growth media at 37°C, 5% CO2. Excess cells were aspirated after 15–30min and the bottomless plate was removed in order to capture the arrayed cells in a layer of 1% low melting point agarose (Invitrogen).
Chemical treatments
Hydrogen peroxide (H2O2) (Sigma) was diluted to desired concentrations in 4°C phosphate buffered saline (PBS) immediately before usage. The bottomless 96-well plate was re-aligned to original position on the CometChip to allow dosing of individual macrowells with 100 μl H2O2. Treatment was performed at 4°C for 20min. Negative controls were dosed with PBS under same conditions.
Exposure to ionising radiation
After encapsulation in agarose, cells were irradiated on ice using a caesium irradiator, Gammacell 40Extractor various doses (1, 2, 4, 6, 8, 10, 15, 20 Gy) at a rate of 1 Gy/min.
Alkaline comet assay
The alkaline comet assay was performed as previously described.(14, 16) Briefly, gels were lysed overnight at 4°C (10mM Tris–HCl, 100mM Na2EDTA, 2.5M NaCl, pH 10 with 1% Triton X-100 added 20min before use). After lysis, gels were transferred to an electrophoresis chamber filled with alkaline unwinding buffer (0.3M NaOH and 1mM Na2EDTA) for 40min at 4°C. Electrophoresis was conducted with the same buffer at 4°C for 30min 1V/cm and 300 mA.
Fluorescence imaging and comet analysis
After electrophoresis, gels were neutralised in 0.4M Tris, pH 7.5 (2×15min) and stained with SYBR Gold (Invitrogen). Images were captured using an automated epifluorescent microscope and analysed using custom software written in MATLAB (The Mathworks) (14).
Results
Creating a 96-well CometChip
A silicon mold with pegs 40 μm in diameter on a grid with 250 μm between pegs was placed onto molten agarose. Removing the mold reveals a microwell array in agarose (Figure 1A). The slab of agarose with the microwell surface is then clamped to bottomless 96-well plate such that each well of the 96-well plate harbours ~300 microwells (Figure 1B). For clarification of nomenclature, the term ‘macrowell’ is used to refer to wells of the 96-well plate. Solutions of single cells are then added to the macrowells and cells are allowed to settle into the microwells on the bottom surface. Excess cells are then removed by aspiration and a layer of agarose is added to capture the cells. Once cells are captured, treatment is performed in the macrowells and analysis is performed according to standard conditions (Figure 1F) (19). Cells are then stained with SYBR gold and imaged under a 10× objective. Since comets are on the same plane (compared to randomly scattered traditional comets, Figure 1E), a single image is effective for capturing 9–12 analysable microwell comets (Figure 1C). The uniform head morphology, provided by the microwell geometry, also simplifies the identification of head/tail transition, ensuring consistent comet measurements (Figure 1D).
H2O2 dose response
Initial studies were aimed at assessing the limits of detection for cells exposed to oxidative damage via treatment with H2O2 on the CometChip. The ability to extrapolate a standard response curve is important for determining an effective dose range for assessing cellular responses. We previously described an X-ray IR dose response curve performed on the CometChip and showed that results were comparable to results using traditional slides (14). Here, we have extended these studies to evaluate exposure to a chemical agent and furthermore, we set out to determine the upper and lower limits of detection. Human lymphoblast TK6 cells were loaded into the macrowells of CometChip. Approximately three cells settled into each of the microwells (previous studies show that comparable results are observed for single cells and clusters of cells) (14, 15). Excess cells were then removed and a layer of agarose was added to the surface to capture the cells. Cells were then exposed to varying doses of H2O2. Here exposure was performed after cells have been encapsulated in gel to determine immediate damage caused by H2O2. Cells could also be treated in culture dishes or in vivo and then transferred to CometChip for further assay. To examine a broad range of doses, cells in this study were exposed to 13 different doses of H2O2, ranging from 5 to 120 μM. These 13 treatment concentrations, together with the negative control, corresponded to 42 macrowells of the 96-well CometChip (each condition was performed in triplicate). Cells were treated for 20min after which time the bottomless plate was removed, and cells were lysed and electrophoresed. Following staining with SYBR green, for each macrowell, 9 images were collected using a 10× objective. We analysed at least 100 microwell comets for each condition and computed the median % tail DNA. We then averaged the medians from triplicate wells. This experiment was repeated four times and Table 1 indicates the average and the standard error of the mean for the four repeats. Results show an increase in DNA damage with increasing doses of H2O2, as expected (Figure 2A). The dose response curve is sigmoidal with flatter slope at lower (<30 μM) and higher concentrations (>100 μM). To identify the linear range of detection, we performed linear regression analysis and calculated that the optimal correlation coefficient (R 2 = 0.98, Figure 2B) is for doses of H2O2 between 30 and 100 μM.
Table I.
TK6 Cell H2O2 dose response (n = 4)
| H2O2 dose (μM) | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 5 | 10 | 15 | 20 | 30 | 40 | 50 | 60 | 70 | 80 | 90 | 100 | 120 | |
| % Taila | 8.7 | 10 | 10.5* | 13.5* | 16.3* | 29.7** | 37.7** | 41.6** | 48.7** | 50.1** | 58.4** | 55.2** | 56.4** | 62.5** |
| SEMb (%) | 0.4 | 0.8 | 0.4 | 2.1 | 2 | 3.5 | 2.5 | 2.9 | 1.7 | 2.8 | 4.1 | 2.3 | 3.4 | 3.8 |
| CVc (%) | 10 | 14.9 | 6.6 | 31 | 24 | 23.4 | 13 | 14.1 | 7.1 | 11 | 14.1 | 8.3 | 12.1 | 12.1 |
| 0 | 5 | 10 | 15 | 20 | 30 | 40 | 50 | 60 | 70 | 80 | 90 | 100 | 120 | |
a% Tail DNA was calculated from averages of four repeats.
bSEM is standard error of the mean.
cCV is coefficient of variation. Student’s t-test was performed to compare each treated dose to negative controls (*P < 0.05, **P < 0.005).
Figure 2.
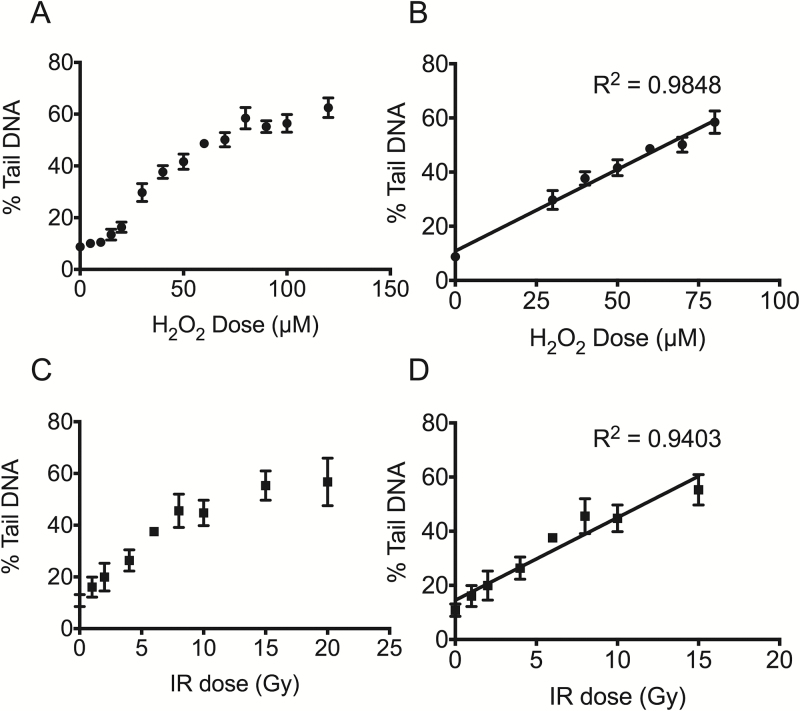
Human lymphoblast TK6 cells dose response obtained using the 96-well CometChip. (A) H2O2 dose response of TK6 cells. Cells were loaded into triplicate macrowells of a 96-well CometChip and treated at varying concentrations of H2O2 for 20min at 4°C. All 13 doses and the negative controls were conducted on the same 96-well CometChip. Data points and error bars represent averages and standard errors of the mean, respectively, of four independent experiments. At least 100 comets were scored for each condition in each experiment. (B) Linear regression of the H2O2 dose response for TK6 cells. Line of best fit was obtained using Linear Regression analysis in GraphPad Prism. R 2 = 0.98 for H2O2 doses of 0, 30, 40, 50, 60, 70 and 80 μM. (C) Irradiation dose response of TK6 cells. Cells were loaded into triplicate wells of the 96-well CometChip, cut into individual pieces and exposed to varying doses of IR on ice, respectively. Data points and error bars represent averages and standard errors of the mean, respectively, of three independent experiments. At least 100 comets were scored for each condition in each experiment. (D) Linear regression of TK6 cell IR dose response. Line of best fit was obtained using Linear Regression analysis in GraphPad Prism. R 2 = 0.94 for IR dose of 0, 1, 2, 4, 6 8, 10 and 15 Gy.
As a reference for the H2O2 studies, we performed a gamma-IR (γ-IR) dose response experiment. TK6 cells were loaded into the CometChip following identical procedures to that for H2O2 and exposed to gamma-IR varying from 1 to 20 Gy at a dose rate of 1 Gy/min. Results collected from three independent experiments are shown in Table 2 and Figure 2C. In contrast to the H2O2 dose response results, the lowest gamma-IR dose used was sufficient to induce significant damage (P < 0.05) and fell within the linear detection range. The response curve plateaued when the dose exceeded 15 Gy, indicating saturation of the assay. We performed linear regression analysis on the data from 0 to 15 Gy γ-IR, shown in Figure 2D, and obtained an optimal correlation coefficient (R 2 = 0.94). By comparing the two dose response curves, we estimate that 30 μM H2O2 induces similar levels of damage as 4 Gy γ-IR in TK6 cells when assayed by alkaline comet assay. These results demonstrate the feasibility of assessing multiple samples in parallel and the ability to extrapolate complete dose response curves from a single CometChip, which facilitates optimisation of experimental conditions. The ability to perform a parallel dose response with a known DNA damaging agent (i.e. γ-IR) can also help to delineate the impact of unknown agents on cellular DNA.
Table II.
TK6 cell IR dose response (n = 3)
| IR dose (Gy) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 2 | 4 | 6 | 8 | 10 | 15 | 20 | |
| % Taila | 10.9 | 16.1* | 19.9* | 26.4* | 37.5* | 45.6* | 44.8* | 55.3* | 56.7* |
| SEMb (%) | 2.3 | 3.9 | 5.4 | 4.1 | 1.2 | 6.5 | 4.9 | 5.6 | 9.2 |
a% Tail DNA was calculated from averages of four repeats.
bSEM is standard error of the mean.
Student’s t-test was performed to compare each treated dose to negative controls (*P < 0.05, **P < 0.005).
CometChip platform and analysis program potentiate efficacy of 4× imaging to further increase throughput
One of the most time consuming steps in the traditional comet assay is the process of acquiring a sufficient number of analysable images, since generally each comet requires its own image. This part of the analysis, besides being labour-intensive, can also introduce bias (9, 20). Previously, we described the benefit of having fully automated imaging and for image analysis for the CometChip (14,15). We estimated that CometChip can accelerate data collection and analysis by at least an order of magnitude due to the fact that ~9 comets can be analysed from a single image, made possible by the fact that comets are on the same focal plane (15). Here, we have explored the possibility of increasing throughput even more by using a 4× rather than a 10× objective, the key advantage being that more comets can be captured in a single image. We performed dose response experiments using TK6 cells treated with varying doses of H2O2, and compared the results when imaged under 10× and 4× objective lens. Because each image contained more than 50 comets under the field of view of a 4× image (Figure 3A), only two images were needed for each sample to acquire 100 analysable comets (this is compared to 100 independent images using the traditional assay). Results from the average of three independent experiments show increasing damage with increasing dose, as expected (Figure 3B). We found that between 30 and 100 μM H2O2 dose treatments again produced the best-fit linear curve (R 2 = 0.99, Figure 3C), indicating that this range is the linear detection range. Comparing results from 4× and 10× it is clear that data collected within this linear detection range using the two objectives are very similar (Figure 3D). Importantly, the absolute levels of damage are not anticipated to be identical because of disparate physical properties of 10× and 4× objectives (i.e. aperture and focal distance). The observations that the linear detection range and the extent to which damage levels change with dose show that results are very consistent for the two objectives (Figure 3D). Given that more than 50 comets can be captured in a single 4× image, the results here show that as few as two images are sufficient for analysis of DNA damage induced by 30–100 μM H2O2 using the CometChip.
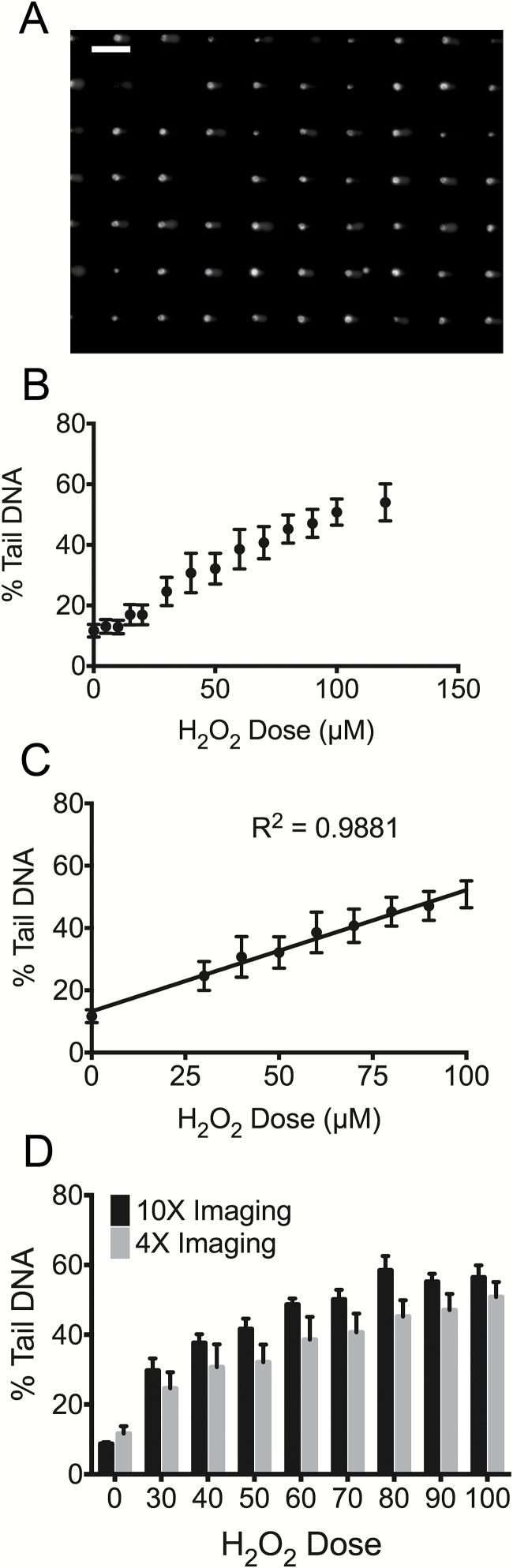
Figure 3.
Efficacy of the 4× objective for image acquisition. (A) Representative image of arrayed comets imaged under 4× objective lens. Scale bar indicates 200 μm. (B) H2O2 dose response of TK6 cells when analysed using a 4x objective. Cells were loaded into triplicate wells of the 96-well CometChip and treated at varying concentrations of H2O2 for 20min at 4°C. Untreated cells were exposed to 1× PBS under the same experimental conditions. All 13 doses and the negative control were conducted on the same 96-well CometChip. Comet images were analysed with using an in-house fully automated MatLab program (14, 15). Data points and error bars represent averages and standard errors of the mean, respectively, of three independent experiments. At least 100 comets were scored for each condition in each experiment. (C) Linear regression of TK6 cell H2O2 dose response. Line for best-fit was obtained using Linear Regression analysis in GraphPad Prism. R 2 = 0.99 for H2O2 doses of 0, 30, 40, 50, 60, 70, 80 and 100 μM. (D) Comparison of results collected using 10× and 4× objective lenses. Dose responses of linear range were plotted as re-presentation of Figures 2B and 3C.
CometChip suppresses sample-to-sample variation
Each macrowell of the 96-well CometChip is comparable to a single slide using the traditional assay. To learn about the inter-sample variation using the CometChip, well-to-well variation was assessed among triplicate wells on the same CometChip for a variety of H2O2 doses (Table 3; Figure 4A). The coefficient of variance (CV) among wells ranged from 5 to 16% (Table 3). It is interesting to note that this CV can be easily achieved by novice users of the platform, whereas the standard assay often requires repeat practices to achieve corresponding CV. Typical slide-to-slide variation of standard assay ranged from 15 to 30% in published reports (11, 14). We also studied consistency from chip-to-chip and we found excellent concordance (Table 4, Figure 4B). To learn about variation among experiments, the CV was calculated for four independent experiments performed on different days (Table 1). In Figure 4C, results of each experimental repeat were plotted as individual bars. The CV ranged from 7 to 31% (Table 1). Samples treated with 15 to 30 μM were most variable (CV > 20%), which is consistent with their lower absolute values. Compared to the well-to-well CV, experiment-to-experiment is higher (9.8 versus 14.4%, respectively).
Table III.
Well-to-well variability
| H2O2 dose (μM) | ||||||
|---|---|---|---|---|---|---|
| 0 | 20 | 40 | 50 | 70 | 90 | |
| Well #1a | 8.4 | 20.9 | 36.7 | 43.1 | 43.2 | 56.8 |
| Well #2a | 7.7 | 16.7 | 39.6 | 42 | 45.1 | 48.4 |
| Well #3a | 10.4 | 16.9 | 40.4 | 45.8 | 52.6 | 48 |
| Mean ± σb | 8.9±1.4 | 18.2±2.4 | 38.9±1.9 | 43.7±2.0 | 47.0±5.0 | 51.0±5.0 |
| CVc | 15.70% | 13.20% | 5.00% | 4.50% | 10.60% | 9.80% |
aMedian % tail DNA of each well were calculated.
bMedian % tail DNA of each.
cCV is coefficient of variance.
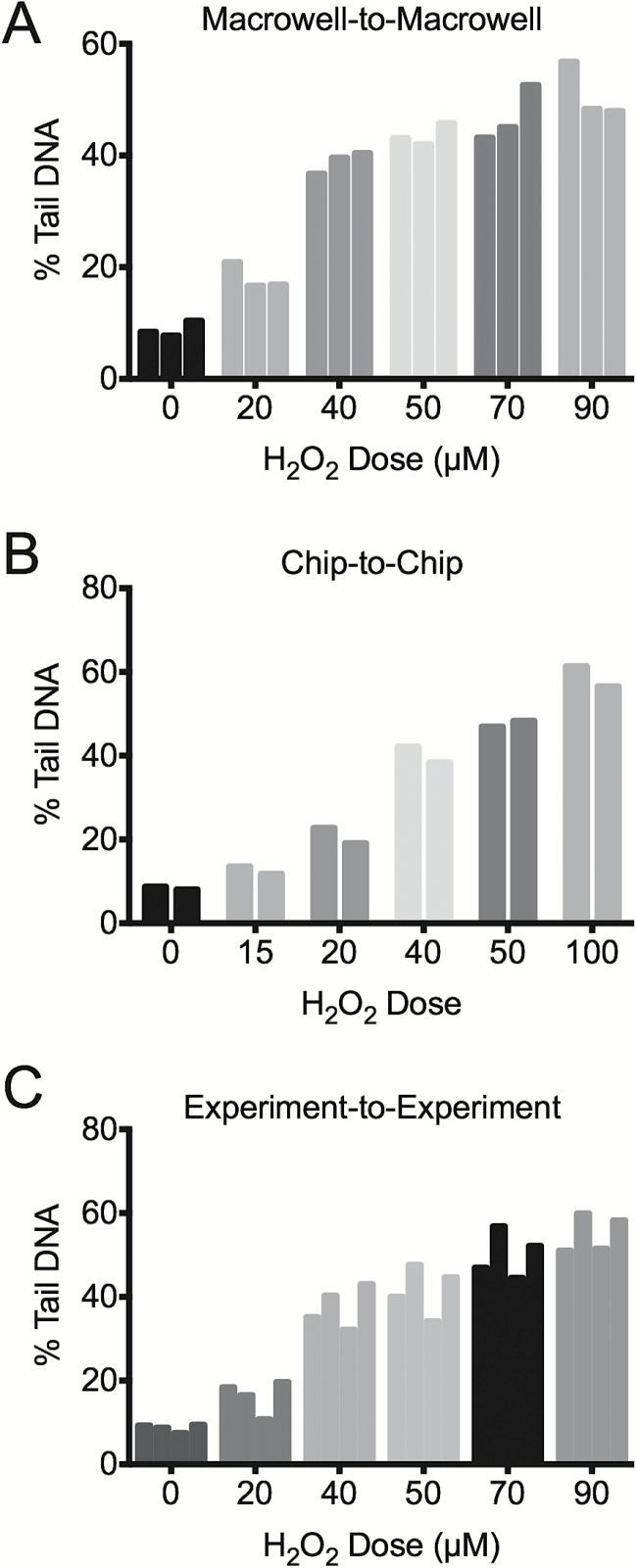
Figure 4.
Well-to-well, chip-to-chip, and experiment-to-experiment variation of CometChip results using human lymphoblast TK6 cells exposed to H2O2. Median % Tail DNA of (A) triplicate wells on the same 96-well CometChip, (B) of two wells, on different CometChips, and (C) of four repeat experiments. TK6 cells were loaded and exposed to varying H2O2 doses. Non-treated cells were exposed to 1× PBS. At least 100 comets were scored.
Table IV.
Chip-to-chip variability
| H2O2 dose (μM) | ||||||
|---|---|---|---|---|---|---|
| 0 | 15 | 20 | 40 | 50 | 100 | |
| Chip #1a | 8.7 | 13.5 | 22.8 | 42.2 | 46.9 | 61.4 |
| Chip #2a | 8.1 | 11.8 | 19.1 | 38.4 | 48.3 | 56.5 |
aMedian % Tail DNA of one well on each chip were calculated.
Microwell comets exhibit low comet-to-comet variation enabling a robust assay with as few as 20 comets scored
While it is conventional to score at least 100 comets for the traditional assay, there is not a consensus about the number of comets that should be scored. Relatively few studies report the potential value of scoring more than 100 comets, due in part to the impracticality as a consequence of the time required for analysis of hundreds of comets.(4, 21–23) The CometChip offers two advantages for analysing more than 100 comets per sample. First, as described above, the comets are on the same focal plane, so many comets can be analysed in a single image. Second, in-house image analysis software automatically identifies and analyses comets for key parameters including tail length, olive tail moment and percent tail DNA, which greatly increases throughput. These features made it feasible to assess the robustness of the assay even with hundreds of comets per sample.
To learn how the number of comets per sample impacts the robustness of the analysis on the CometChip, we analysed TK6 lymphoblast cells challenged with exposure to H2O2. Of particular interest were the low doses, as we wanted to learn if increasing the number of comets might offer greater sensitivity of the assay. We scored 20, 50, 100, 200, 400 and 600 comets for each of four different doses of H2O2 and repeated the experiment four times. Table 5 lists the average results of the four repeats, which are then plotted in Figure 5A. Changing the number of comets scored in each experiment did not affect the absolute % tail DNA measured. Importantly, when % DNA damage exceeds ~15%, only 20 comets are needed to generate robust data, which is far fewer than 100 comets which are standard using the traditional assay (Figure 5B). Similar findings also applied to measurement of double strand breaks using neutral comet assay on previously published neutral CometChip (15). In contrast, at lower damage levels below the linear range (~ 9–11% tail), scoring more comets effectively increased the sensitivity of the assay. We were able to detect damage induced by 10 and 5 μM of H2O2 by scoring 100 and 400 comets respectively (Figure 5C–E). The impact that the number of comets scored has on assay sensitivity is consistent with results published by Sharma et al. (4) and can be explained by the lowered standard deviation and thereby decreased CV. At low doses (e.g. 5 and 10 μM), the signal-to-noise ratio is low and hence reducing variance increases the significance level. On the other hand, higher levels of H2O2 exposure (i.e. 20 and 30 μM) produce sufficient DNA damage that is more resistant to noise-induced variances. As a result, we observed reduced P values with increasing number of comets scored in 5 μM experiments, but not in 30 μM experiments (Figure 5F). Overall, the number of comets required to achieve sensitive measurements of DNA damage using the CometChip, either at low (100–400 comets) or high doses (20 comets), is far fewer than what is required for traditional comet assay (4).
Table V.
Comparison of average median % tail DNA results analysed from comet pools of varying sizes, n = 4
| H2O2 dose (μM) | ||||||
|---|---|---|---|---|---|---|
| # of comets Sored | 0 | 5 | 10 | 20 | 30 | |
| 20 | % Taila ± SDb | 8.8±1.7 | 10.5±1.7 | 10.4±0.8 | 15.9±4.5* | 27.4±6.6** |
| 50 | % Taila ± SDb | 8.6±1.2 | 9.8±1.7 | 10.3±0.7 | 16.0±5.6* | 28.2±8.8** |
| 100 | % Taila ± SDb | 8.4±1.1 | 9.7±1.6 | 10.4±1.0* | 15.5±4.0* | 29.2±7.9** |
| 200 | % Taila ± SDb | 8.4±0.8 | 10.2±2.0 | 10.6±0.6* | 16.5±4.5* | 29.1±7.7** |
| 400 | % Taila ± SDb | 8.5±0.9 | 10.2±1.6* | 10.4±0.5* | 16.1±4.5* | 28.7±7.8** |
| 600 | % Taila ± SDb | 8.4±0.7 | 10.1±1.6* | 10.4±0.4** | 16.3±4.5* | 28.8±7.5** |
a%Tail DNA was calculated from averaging median % tail from four repeats.
bSD is standard deviation (SD is used here instead of SEM because comet data were pooled. Student’s t-test was performed to compare each treated dose to negative controls in each pool (*P < 0.05, **P < 0.01).
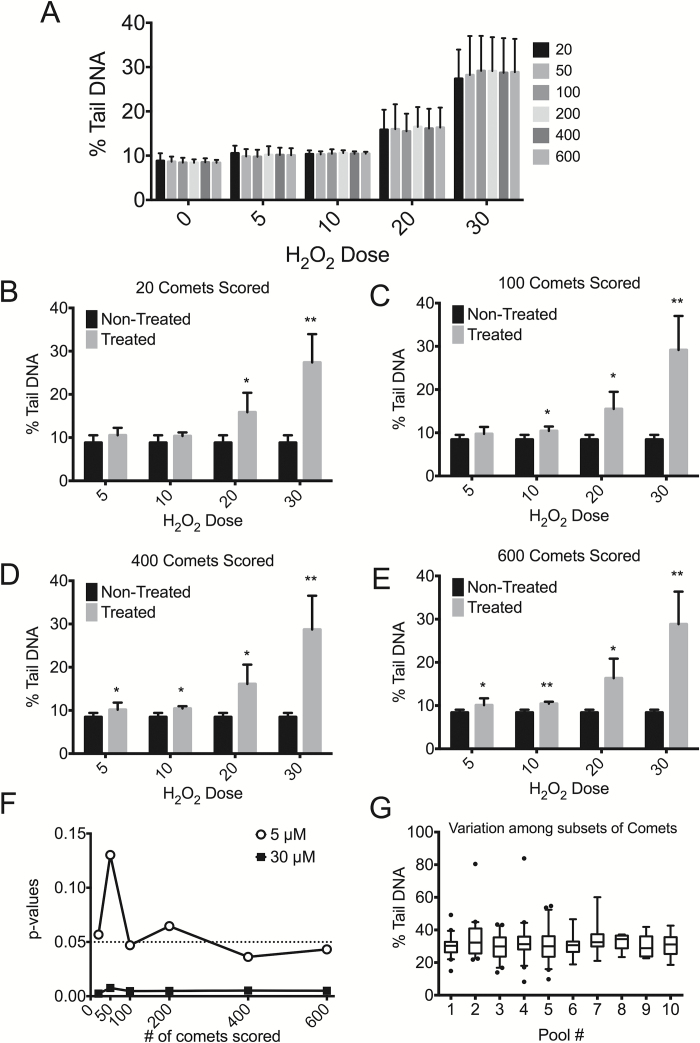
Figure 5.
Influence of the number of microwell comets scored on the sensitivity and statistical robustness of the CometChip assay. (A) Comparison of average median % tail DNA results analysed from comet pools of varying sizes, n = 4. TK6 cells were embedded in CometChip and treated at varying concentrations of H2O2 for 20min at 4°C. Untreated cells were exposed to 1× PBS under the same experimental conditions. Comet images were collected in a fully automated fashion. 20, 50, 100, 200, 400 and 600 comets were scored randomly to obtain median % tail DNA. Average of four repeats were plotted as column graphs with error bars indicating standard deviation of mean. Each column colour indicates a different sample size, explained by legend of the graph. (B–E) Replots of median % tail DNA results for different sample pool sizes. Student’s t-test was performed to compare each treated dose to corresponding negative controls (*P < 0.05, **P < 0.01). (F) Plot of P-values as a function of the number of comets scored. The 0.05 P value line is marked as dotted line in graph. (G) Box-and-whisker plot of median % tail DNA analysed from randomly scoring 20 comets that were exposed to 30 μM H2O2. Each box represents the lower and upper quartiles with median shown as the middle line. Whiskers show 10 to 90 percentile. Points below and above the whiskers are indicated.
As previously described, the microwell comets contain multiple cells and are self-calibrated by the morphology of the microwells (14,15). One possible reason for reduced variance using the CometChip is that the comet-to-comet variance is greatly reduced because each microwell comet is anticipated to show the average amount of damage for the cells in that well, thus reducing variance compared to single cell comets. Using the CometChip, our initial analysis shows that as few as 20 microwell comets are representative of the whole sample population. To further explore representation, we created 10 subsets of samples treated with 30 μM H2O2, each containing 20 randomly scored microwell comets. Median, quartiles, and 10–90 percentile of analysed data were plotted in Figure 5G. We found that varying the comets for each analysis did not significantly impact the median % tail DNA. Given that a typical image captured under 10× objective lens generally contain 9–12 microwell comets, it is possible to collect robust data using just two images (one under 4×). On the other hand, in situations where very low levels of DNA damage are being studied, users can achieve better sensitivity by scoring more comets. The amount of materials, time, and labour required to score more comets, nonetheless, is significantly less than traditional assay.
Discussion
Here we have analysed the detection range as well as inter and intra-sample variation of the CometChip. This novel technology, with the combination of microarray technology and fully automated image analysis was estimated to increase throughput by two orders of magnitude (15). Here we show that the CometChip also has the added advantage of requiring fewer comets to achieve the same sensitivity as the traditional assay.
The CometChip overcomes two key limitations of existing comet assay: (i) difficulty of processing multiple samples in parallel, both physically and in terms of time required; and (ii) sample-to-sample variability, which deteriorates as more samples are needed to be processed in parallel. Published studies using a large glass slide with multiple minigels on the surface have been shown to improve throughput and reproducibility (12, 13). The CometChip incorporates this advance using an easy-to-handle 96-well plate. Processing 96 samples on a single CometChip is far easier than handling 96 slides, which is beyond the capacity of most researchers. In addition to being easier to handle, by settling cells into wells with a uniform depth, the CometChip avoids problems with out-of-focus or overlapping comets (Figure 1D), making it possible to image as many as 63 analysable comets in a single image (under 4× objective, Figure 3A), which reduces comet-to-comet and sample-to-sample variation, as shown here. Being able to process many samples in parallel makes it easier to detect the linear range of genotoxin exposure, which greatly accelerates identification of doses that give rise to DNA damage levels that are not saturated, a particularly difficult optimisation step using the traditional assay.
A key advantage of reduced experimental variation is that the CometChip can potentially obviate the need for internal standards. Tremendous effort has gone toward the development of standards that help to control for intergel, inter-experiment and inter-laboratory variability (11, 24). Inter-sample variation can range from 10 to 20%, (11, 14, 15) and analysis performed by different scorers or laboratories can vary as high as 80% (6, 8, 9). Inclusion of standards adds complexity to the protocol and has limitations that may lower throughput and versatility of the assay. In contrast, the CometChip, with its reduced inter-sample variability, may obviate the need to normalise across gels. Although we did not formally access inter-scorer or inter-laboratory variation, preliminary data show excellent concordance, making the CometChip a promising alternative to internal standards.
CometChip was developed with the goal of ease of use. The hardware is designed to allow easy assembly, straightforward procedures and uniform quality. These features provide a very low barrier of entry for researchers who are unfamiliar with the assay. Indeed, we found that novice researchers were able to collect publishable quality data within several runs. An optimised version of the platform that can accommodate both 24 and 96-well plates is currently under development at Trevigen, Inc.
With its combination of higher throughput and high sensitivity, the CometChip is potentially valuable in a variety of research settings. The technology can easily be implemented for epidemiology and clinical studies that were previous inaccessible due to the limitations of traditional comet assay. Previous and ongoing studies in our own laboratory show utility of the CometChip for studying variations in repair kinetics among individuals, exploring gene-targeted cancer therapies, and evaluating DNA damage response of populations exposed to prevalent environmental genotoxins. Taken together, the CometChip significantly broadens the utility of the comet assay for a broad range of studies across many disciplines.
Funding
This work was supported by the National Institute of Environmental Healthy Studies (NIEHS) (Grant U01-ES016-45, R43-ES021116-01). D.M.W was supported by the NIEHS Training Grant (T32-ES0702). D.N.C and J.L.F were supported by MIT Undergraduate Research Opportunity Program.
Acknowledgements
We thank Bob Kirby for IR facilities and MIT Center for Environmental Health Sciences for research equipment. The authors declare no conflicts of interest.
References
- 1. Ostling O., Johanson K. J. (1984). Microelectrophoretic study of radiation-induced DNA damages in individual mammalian cells. Biochem. Biophys. Res. Commun., 123, 291–298. [DOI] [PubMed] [Google Scholar]
- 2. Tice R. R., Agurell E., Anderson D., et al. (2000). Single cell gel/comet assay: guidelines for in vitro and in vivo genetic toxicology testing. Environ. Mol. Mutagen., 35, 206–221. [DOI] [PubMed] [Google Scholar]
- 3. Valverde M., Rojas E. (2009). Environmental and occupational biomonitoring using the Comet assay. Mutat. Res., 681, 93–109. [DOI] [PubMed] [Google Scholar]
- 4. Sharma A. K., Soussaline F., Sallette J., Dybdahl M. (2012). The influence of the number of cells scored on the sensitivity in the comet assay. Mutat. Res., 749, 70–75. [DOI] [PubMed] [Google Scholar]
- 5. Azqueta A., Gutzkow K. B., Brunborg G., Collins A. R. (2011). Towards a more reliable comet assay: optimising agarose concentration, unwinding time and electrophoresis conditions. Mutat. Res., 724, 41–45. [DOI] [PubMed] [Google Scholar]
- 6. Ersson C., Möller L. (2011). The effects on DNA migration of altering parameters in the comet assay protocol such as agarose density, electrophoresis conditions and durations of the enzyme or the alkaline treatments. Mutagenesis, 26, 689–695. [DOI] [PubMed] [Google Scholar]
- 7. Azqueta A., Meier S., Priestley C., Gutzkow K. B., Brunborg G., Sallette J., Soussaline F., Collins A. (2011). The influence of scoring method on variability in results obtained with the comet assay. Mutagenesis, 26, 393–399. [DOI] [PubMed] [Google Scholar]
- 8. Forchhammer L., Bräuner E. V., Folkmann J. K., Danielsen P. H., Nielsen C., Jensen A., Loft S., Friis G., Møller P. (2008). Variation in assessment of oxidatively damaged DNA in mononuclear blood cells by the comet assay with visual scoring. Mutagenesis, 23, 223–231. [DOI] [PubMed] [Google Scholar]
- 9. Forchhammer L., Ersson C., Loft S., et al. (2012). Inter-laboratory variation in DNA damage using a standard comet assay protocol. Mutagenesis, 10.1093/mutage/ges032. [DOI] [PubMed] [Google Scholar]
- 10. Møller P., Friis G., Christensen P. H., et al. (2004). Intra-laboratory comet assay sample scoring exercise for determination of formamidopyrimidine DNA glycosylase sites in human mononuclear blood cell DNA. Free Radic. Res., 38, 1207–1214. [DOI] [PubMed] [Google Scholar]
- 11. Zainol M., Stoute J., Almeida G. M., Rapp A., Bowman K. J., Jones G. D; ECVAG. (2009). Introducing a true internal standard for the Comet assay to minimize intra- and inter-experiment variability in measures of DNA damage and repair. Nucleic Acids Res., 37, e150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Kiskinis E. (2002). High throughput Comet assay using 96-well plates. Mutagenesis, 17, 37–43. [DOI] [PubMed] [Google Scholar]
- 13. Gutzkow K. B., Langleite T. M., Meier S., Graupner A., Collins A. R., Brunborg G. (2013). High-throughput comet assay using 96 minigels. Mutagenesis, 28, 333–340. [DOI] [PubMed] [Google Scholar]
- 14. Wood D. K., Weingeist D. M., Bhatia S. N., Engelward B. P. (2010). Single cell trapping and DNA damage analysis using microwell arrays. Proc. Natl. Acad. Sci. U. S. A., 107, 10008–10013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Weingeist D. M., Ge J., Wood D. K., Mutamba J. T., Huang Q., Rowland E. A., Yaffe M. B., Floyd S., Engelward B. P. (2013). Single-cell microarray enables high-throughput evaluation of DNA double-strand breaks and DNA repair inhibitors. Cell Cycle, 12, 907–915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Ge J., Wood D. K., Weingeist D. M., Prasongtanakij S., Navasumrit P., Ruchirawat M., Engelward B. P. (2013). Standard fluorescent imaging of live cells is highly genotoxic. Cytometry. A, 83, 552–560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Chao M. W., Kim M. Y., Ye W., et al. (2012). Genotoxicity of 2,6- and 3,5-dimethylaniline in cultured mammalian cells: the role of reactive oxygen species. Toxicol. Sci., 130, 48–59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Watson C., Ge J., Cohen J., Pyrgiotakis G., Engelward B. P., Demokritou P. (2014). High-throughput screening platform for engineered nanoparticle-mediated genotoxicity using CometChip technology. ACS Nano, 8, 2118–2133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Olive P. L., Banáth J. P. (2006). The comet assay: a method to measure DNA damage in individual cells. Nat. Protoc., 1, 23–29. [DOI] [PubMed] [Google Scholar]
- 20. Wood R. D., Mitchell M., Sgouros J., Lindahl T. (2001). Human DNA repair genes. Science, 291, 1284–1289. [DOI] [PubMed] [Google Scholar]
- 21. Dehon G., Catoire L., Duez P., Bogaerts P., Dubois J. (2008). Validation of an automatic comet assay analysis system integrating the curve fitting of combined comet intensity profiles. Mutat. Res., 650, 87–95. [DOI] [PubMed] [Google Scholar]
- 22. Stang A., Brendamour M., Schunck C., Witte I. (2010). Automated analysis of DNA damage in the high-throughput version of the comet assay. Mutat. Res., 698, 1–5. [DOI] [PubMed] [Google Scholar]
- 23. Smith C. C., Adkins D. J., Martin E. A., O’Donovan M. R. (2008). Recommendations for design of the rat comet assay. Mutagenesis, 23, 233–240. [DOI] [PubMed] [Google Scholar]
- 24. De Boeck M., Touil N., De Visscher G., Vande P. A., Kirsch-Volders M. (2000). Validation and implementation of an internal standard in comet assay analysis. Mutat. Res., 469, 181–197. [DOI] [PubMed] [Google Scholar]