Abstract
Linkage maps based on markers derived from genes are essential evolutionary tools for commercial marine fish to help identify genomic regions associated with complex traits and subject to selective forces at play during exploitation or selective breeding. Additionally, they allow the use of genomic information from other related species for which more detailed information is available. Sole (solea solea L.) is a commercially important flatfish species in the North Sea, subject to overexploitation and showing evidence of fisheries-induced evolutionary changes in growth- and maturation-related traits. Sole would definitely benefit from a linkage map to better understand how evolution has shaped its genome structure. This study presents a linkage map of sole based on 423 single nucleotide polymorphisms derived from expressed sequence tags and 8 neutral microsatellite markers. The total map length is 1233.8 cM and consists of 38 linkage groups with a size varying between 0 to 92.1 cM. Being derived from expressed sequence tags allowed us to align the map with the genome of four model fish species, namely medaka (Oryzias latipes), Nile tilapia (Oreochromis niloticus), three-spined stickleback (Gasterosteus aculeatus) and green spotted pufferfish (Tetraodon nigroviridis). This comparison revealed multiple conserved syntenic regions with all four species, and suggested that the linkage groups represent 21 putative sole chromosomes. The map was also compared to the linkage map of turbot (Scophthalmus maximus), another commercially important flatfish species and closely related to sole. For all putative sole chromosomes (except one) a turbot homolog was detected, confirming the even higher degree of synteny between these two flatfish species.
Introduction
Preserving the evolutionary potential of exploited marine fish species is essential to secure viable populations across a species’ full geographical and environmental range [1]. A better understanding of the strength of local and fisheries-induced adaptation hence provides important insights into the resilience and evolutionary response to environmental change and harvesting of stocks [2]. This is crucial to move towards evolutionary enlightened sustainable fisheries management [3]. In the last decade, marine fisheries have strongly declined or even collapsed [4], [5] and a growing number of fish population studies have reported significant changes in life history traits that have been associated with fisheries-induced selection [6], [7], [8]. Examples include shifts towards earlier maturation at a smaller size, increased reproductive investment and changes in growth rate, all of which are consistent with the size-selective nature of fishing. These changes, in synergy with climate change, may have drastic negative effects on fish populations and their sustainable fisheries [2]. One of the most challenging problems in studying local adaptation and fisheries-induced evolution, however, is disentangling the environmental and genetic causes behind changes in life-history traits [7], [9], [10]. Many of the adaptive traits that respond to evolution are complex and quantitative by nature. Hence, genomic tools became pivotal to reveal footprints of selection, as changes can now be studied directly at the molecular level.
Genetic linkage maps represent one of such essential evolutionary genomics tools for commercial marine fish, to help identify genomic regions associated with complex traits subject to selective forces. Additionally they are crucial during selective breeding initiatives, aiming at relieving fishery pressure on overexploited stocks, while increasing the cost-efficiency of farmed fish production. They also provide the necessary resources for genomic comparison with other fish species to understand their genome evolution and organization [11], [12] and facilitate anchoring of scaffolds to chromosomes in whole genome sequencing and assembly [13].
Sole (Solea solea L.) is a commercially important marine flatfish (Pleuronectiformes) of the family Soleidae mainly living in the Northeast Atlantic Ocean, but also in the whole Mediterranean Sea and in the Southwestern Black Sea [14]. The spawning stock biomass of the North Sea has been fluctuating around the precautionary point of 35,000 tons depending on the strength of the year classes. Periods of revival however were short and overall there has been a downward trend, putting the sole stock at risk of reduced reproductive capacity [15]. Additionally, strong evidence exists for fisheries-induced evolutionary change in the onset of sexual maturity over the last 60 years [16].
Despite the commercial importance of sole, no genomic tools are available to date [17], [18]. Currently, genetic linkage maps are available for only five other flatfish species: turbot (Scophthalmus maximus) [19], [20], brill (Scophthalmus rhombus) [21], Atlantic halibut (Hippoglossus hippoglossus) [22], half-smooth tongue sole (Cynoglossus semilaevis) [23] and olive flounder (Paralichthys olivaceus) [24], [25]. One of the major obstacles to build a linkage map for sole has been the lack of informative genetic markers. However, cutting-edge next-generation DNA sequencing technologies allow for a rapid and cost-efficient development of genetic markers across the genome of highly exploited species without any existing genomic information [26].
Here, we used a novel set of single nucleotide polymorphisms (SNPs) that were developed from expressed sequence tags (ESTs) to construct the first linkage map of sole. Additionally, we used this map for comparative mapping and establishing the syntenic relationships with four fully sequenced model species, namely medaka (Oryzias latipes), Nile tilapia (Oreochromis niloticus), three-spined stickleback (Gasterosteus aculeatus) and green spotted pufferfish (Tetraodon nigroviridis). These comparisons were ultimately used as a stepping stone to compare the sole linkage map with that of turbot (Scophthalmus maximus), which is more closely related to sole than the model fish species. The future applications of this novel molecular tool were further discussed into the context of evolutionary based fisheries management of flatfish species.
Materials and Methods
Mapping families
Two full-sib families with 46 and 35 offspring respectively were used for the linkage analysis. The sampling of the parents, broodstock management, offspring collection and DNA extractions were done by Blonk [27] and were part of a large breeding program, initiated by Solea B.V. (The Netherlands) and the Animal Breeding and Genomics Centre (Wageningen University), aiming at increased productivity of farmed sole. All procedures were in accordance with the Dutch law. The parents originated from the Southern North Sea (52°N and 2.5°E) and were collected between 2003 and 2005. Details on the management of the broodstock including these parents (called ‘B’) and their offspring can be found in [28]. Of each parent a blood sample (0.1 ml) was taken without killing them (as they were part of the sole breeding program). DNA was extracted from this sample using a Puregene DNA purification kit for non-mammalian whole blood samples (Gentra Systems). DNA extraction of the offspring was performed on 3 to 4 days old larvae using nucleospin tissue columns following the manufacturer’s guideline (96 procedure, Machery-Nagel).
Genetic markers and genotyping
Using the Roche FLX Titanium technology, in total 348,042 cDNA sequences were generated from a multiplexed sole muscle library based on seven sole individuals sampled across the East Atlantic Ocean and the Mediterranean Sea (unpublished data). A total of 11,021 contigs could be assembled and over 3,000 SNPs detected in silico. Among those, 1,536 SNPs were selected for validation using the Illumina GoldenGate™ high-throughput genotyping assay. Some of these markers have already been applied successfully in a traceability context [29]. The inheritance and the informativeness of the markers were visually checked with the GenomeStudio™ genotyping module of Illumina. Of the 1,536 genotyped SNPs, 749 were not informative within the two families. For 21 SNPs inconsistencies with mendelian inheritance were observed and for 297 SNPs the genotyping assay failed. An overview of the remaining 469 SNPs that were used to perform linkage analysis, can be found in S1 Table. In addition both families were genotyped at the following 10 microsatellite markers with the method described in Blonk et al. [28]: AF173849, AF173852, AF173854, AF173855 [30], AY950587, AY950588, AY950589, AY950591, AY950592, AY950593 [31]. In total 479 markers (469 SNPs and 10 microsatellites) were included in the linkage analysis. A high number of missing data often indicates either poor DNA quality or markers that are difficult to call, which may lead to difficulties in linkage map construction. Therefore, individuals or markers with more than 30% missing data were excluded from the dataset, being two individuals and none of the markers. The final dataset consisted of two families with respectively 46 and 33 offspring (with no more than 3% missing values per individual) and 479 markers (of which only 13 SNPs had more than 3% missing data).
Linkage map construction and genome coverage
SNPs located within the same contig were combined into haplotypes using the program Pedphase v3.0 [32]. The map was built with the program Cri-map v2.4 [33]. Initial grouping of the markers was carried out using the twopoint and autogroup option of Cri-map. The twopoint analysis was performed for all pairs of markers with a threshold of LOD = 3. Autogroup was then used to identify sets of markers which were likely located in the same linkage group (LOD ≥8). Some small linkage groups were pooled based on additional information: (1) an autogroup analysis with a lower threshold (LOD of four instead of eight) and (2) if linkage was found between the majority of the markers after performing an additional twopoint analysis with a threshold of LOD = 0.5. Secondly, the position of the SNPs and microsatellite markers within the groups was determined with the build option of Crimap in an iterative process, starting with a LOD score of 3 and through subsequent stepwise lowering of the LOD score. Finally, marker order was calibrated using the flips option with a window size of five markers. The map was drawn in the program Mapchart v2.2 [34]. Two methods were used to estimate the genome length (Ge) of sole. According to Fishman et al. [35] corrections should be made for chromosome ends by adding 2*Dav to the length of each linkage group, where Dav is the average inter-marker distance of the linkage map. Chakravarti et al. [36] suggested to multiply each linkage group by (m+1)/(m-1), where m is the number of loci in each linkage group (4th method in Chakravarti et al. [36]). The average length calculated by both methods was used as an estimate for the genome length (resp. Ge1 and Ge2).
Comparative mapping and syntenic relationships
As all SNPs in the sole map were discovered from ESTs, the assembled contigs were used as queries to perform a local blast against the genome of four fully sequenced fish species: medaka (Oryzias latipes, order Beloniformes), Nile tilapia (Oreochromis niloticus, order Cichliformes), three-spined stickleback (Gasterosteus aculeatus, order Perciformes) and green spotted pufferfish (Tetraodon nigroviridis, order Tetraodontiformes). Of all available model species, these four are most closely related to the Pleuronectiformes, each representing a different order within the Percomorphaceae [37]. Genomic information was downloaded from the genome browser Ucsc (http://genome.ucsc.edu/). A Blastn analysis was performed with NCBI-Blast under default settings with exception of the E-value (<10−10). To establish syntenic relationships with turbot (Scophthalmus maximus) a direct comparison by blast analysis was not possible, as this species is not fully sequenced yet. Therefore, the stepping stone approach as described by Sarropoulou et al. [38] was used. The four model species were used as a bridge which allowed passing from the sole linkage groups to those of turbot.
Results and Discussion
Linkage map construction and genome coverage
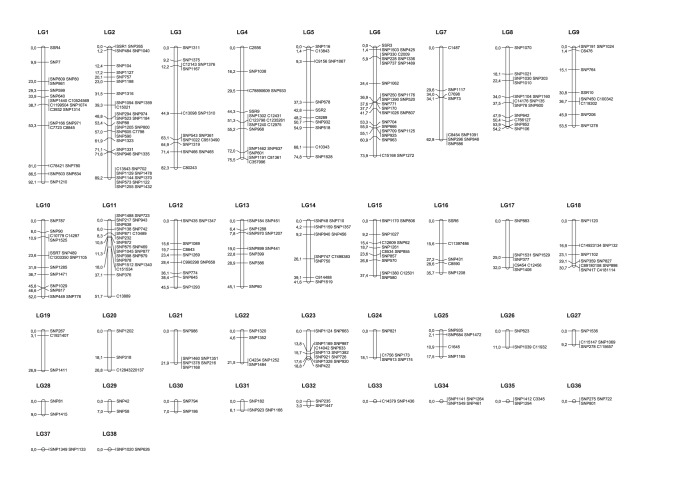
The 469 informative SNPs were distributed over 291 single-marker contigs and 73 contigs with multiple SNPs. These markers located on the same contig were combined into haplotypes, in order to compensate for the low number of informative meioses due to the low number of individuals in the mapping families and to provide a more accurate estimate of the recombination frequency between the markers [39]. This was done for all contigs with multiple SNPs, except for four of them because of evidence for recombination within that contig (S1 Table). Of the 469 SNPs and 10 microsatellite markers, 423 and 8 (∼90%) respectively were incorporated into the linkage map (Fig. 1, S1 Table). The total length of the map was estimated at 1233.8 cM and consists of 38 linkage groups (LG1 to LG38) with a size varying between 0 to 92.1 cM (Table 1, S1 Table). The number of markers per group ranges from two to 39. The average inter-marker distance (Dav) is 8.1 cM and the maximal interval is 32.7 cM. The 48 unmapped markers could not be assigned a position due to an insufficient number of informative meioses. For the same reason, the order of closely linked markers might be inaccurate or it may be impossible to separate markers.
Figure 1. Sex-averaged linkage map of sole.
Map distances are calculated using the Kosambi mapping function and shown in centimorgans. Combined SNPs are indicated with a ‘C’ at the beginning of their name.
Table 1. Number of markers, the corresponding number of distinct contigs and map length for each linkage group of sole.
| LG | No. Markers | No. contigs | Length (cM) |
| LG1 | 30 | 21 | 92.1 |
| LG2 | 39 | 35 | 89.2 |
| LG3 | 21 | 15 | 82.3 |
| LG4 | 32 | 17 | 75.5 |
| LG5 | 15 | 10 | 74.8 |
| LG6 | 28 | 25 | 73.9 |
| LG7 | 12 | 9 | 62.8 |
| LG8 | 17 | 15 | 54.2 |
| LG9 | 15 | 9 | 53.5 |
| LG10 | 18 | 14 | 52 |
| LG11 | 27 | 17 | 51.7 |
| LG12 | 14 | 10 | 45.5 |
| LG13 | 10 | 10 | 45.1 |
| LG14 | 14 | 11 | 41.6 |
| LG15 | 16 | 13 | 37.4 |
| LG16 | 8 | 4 | 35.7 |
| LG17 | 9 | 6 | 32 |
| LG18 | 16 | 10 | 30.7 |
| LG19 | 4 | 3 | 26.9 |
| LG20 | 7 | 3 | 26.8 |
| LG21 | 6 | 6 | 21.9 |
| LG22 | 6 | 5 | 21.5 |
| LG23 | 14 | 13 | 18.8 |
| LG24 | 6 | 5 | 18.1 |
| LG25 | 6 | 5 | 17.5 |
| LG26 | 4 | 3 | 11 |
| LG27 | 8 | 5 | 9.2 |
| LG28 | 2 | 2 | 9 |
| LG29 | 2 | 2 | 7 |
| LG30 | 2 | 2 | 7 |
| LG31 | 3 | 3 | 6.1 |
| LG32 | 2 | 2 | 3 |
| LG33 | 3 | 2 | 0 |
| LG34 | 4 | 4 | 0 |
| LG35 | 4 | 3 | 0 |
| LG36 | 3 | 3 | 0 |
| LG37 | 2 | 2 | 0 |
| LG38 | 2 | 2 | 0 |
The estimated genome length according to Fishman et al. [35] (Ge1) is 1849.4 cM. However, to obtain a more accurate estimate, two further corrections were made: (1) The distance between two markers in a linkage map should not exceed 30 cM [40]. As the haploid chromosome number of sole is 21 [41], there are 17 linkage groups in excess. For these 17, no correction was made for chromosome ends, but instead 17*30 cM was added to the map length of 1233.8 cM. (2) For the number of acrocentric chromosomes, 13 in the case of sole [41], only Dav instead of 2Dav was added (105.3 cM instead of 210.6 cM). For the other 8 chromosomes 2Dav was added (129.6 cM). In total, the estimated genome length (Ge1) after both corrections is 1978.7 cM. The estimated genome length (Ge2) according to the fourth method of Chakravarti et al. [36] is 1490.2 cM. The average of both values (Ge1 and Ge2) is 1734.45 cM, corresponding to approximately 70% genome coverage. Half of the linkage groups are larger than 30 cM, a size similar to those found in maps of other (flat)fish species [20], [22], [24], [42]. Twelve linkage groups are smaller than 10 cM, therefore covering only part of the chromosomes represented by these linkage groups. Several of these linkage groups most likely represent different parts of the same chromosomes.
Comparative mapping with model fish species
The 423 mapped SNPs originated from 326 different contigs (S1 Table). The sequences for these contigs were aligned against the genome of stickleback, pufferfish, medaka and tilapia using Blastn. One-hundred and eighty contigs showed a significant sequence homology above the threshold (E-value <10−10) in at least one of the four fish species examined: 152 in stickleback, 126 in tilapia, 105 in medaka and 97 in pufferfish. Fifty-eight contigs showed a significant sequence homology in all four model species. A detailed overview of the Blastn results can be found in S2–S5 Tables. The highest levels of sequence similarity were found with the stickleback genome (∼47%), followed by the tilapia genome (∼39%) and the lowest levels with the medaka (∼31%) and pufferfish (∼30%) genome. However, these results are not in concordance with the phylogenetic position of the Pleuronectiformes among other Percomorphaceae, [37], [43]. Flatfish belong to the Carangimorphariae, which are most closely related to the Ovalentariae, which include medaka and tilapia, and more distantly related to Percomorpharia, which include stickleback and pufferfish. Similar results were observed for turbot [20]. Despite the closer relationship between turbot and medaka, more sequence similarity was observed with stickleback (∼50%) than with medaka and pufferfish (∼40%). Bouza et al. [20] suggested that this reflected phylogenetic discordances between the use of mitochondrial and nuclear genes, as the proposed phylogeny was based on the mitogenome [44], [45]. However, recent phylogenetic studies using a combination of both marker types do not support such discordance [37], [43], [46]. Another possible explanation for this apparent discrepancy is provided by the different evolutionary rates among closely related species. Medaka and pufferfish (and teleosts in general) evolved faster than other vertebrate species [45], [47], [48], [49], hence they might also have evolved faster than sole, stickleback and tilapia. Support for this is presented by Betancur et al. [43], which show that the branch lengths in their evolutionary tree are longer for medaka and pufferfish in comparison to the other three species. S6 Table lists the number and length of the conserved syntenic regions (i.e. markers on the same linkage group that are also on the same chromosome, regardless the order) with each of the four model species. Most syntenic sequences were found with stickleback (23 syntenic regions, containing on average 5.3 sequences), followed by tilapia, medaka and pufferfish (resp. 23, 19 and 18 regions, containing 4.3, 4.3 and 3.6 sequences). These numbers are in concordance with the observed levels of sequences similarity for each species (see above). The total length of the syntenic regions was smaller for stickleback (200.1 Mb) than for tilapia (257.8 Mb) and medaka (224.7 Mb), but larger when compared to pufferfish (89.2 Mb). This suggests that the stickleback genome is more compact in comparison to tilapia and medaka, but larger than pufferfish, which is consistent with their genome size. Tilapia has the largest genome at about 820 Mb, followed by medaka (∼700 Mb), stickleback (∼460 Mb) and pufferfish (∼350 Mb).
Syntenic relationships between sole and model fish species
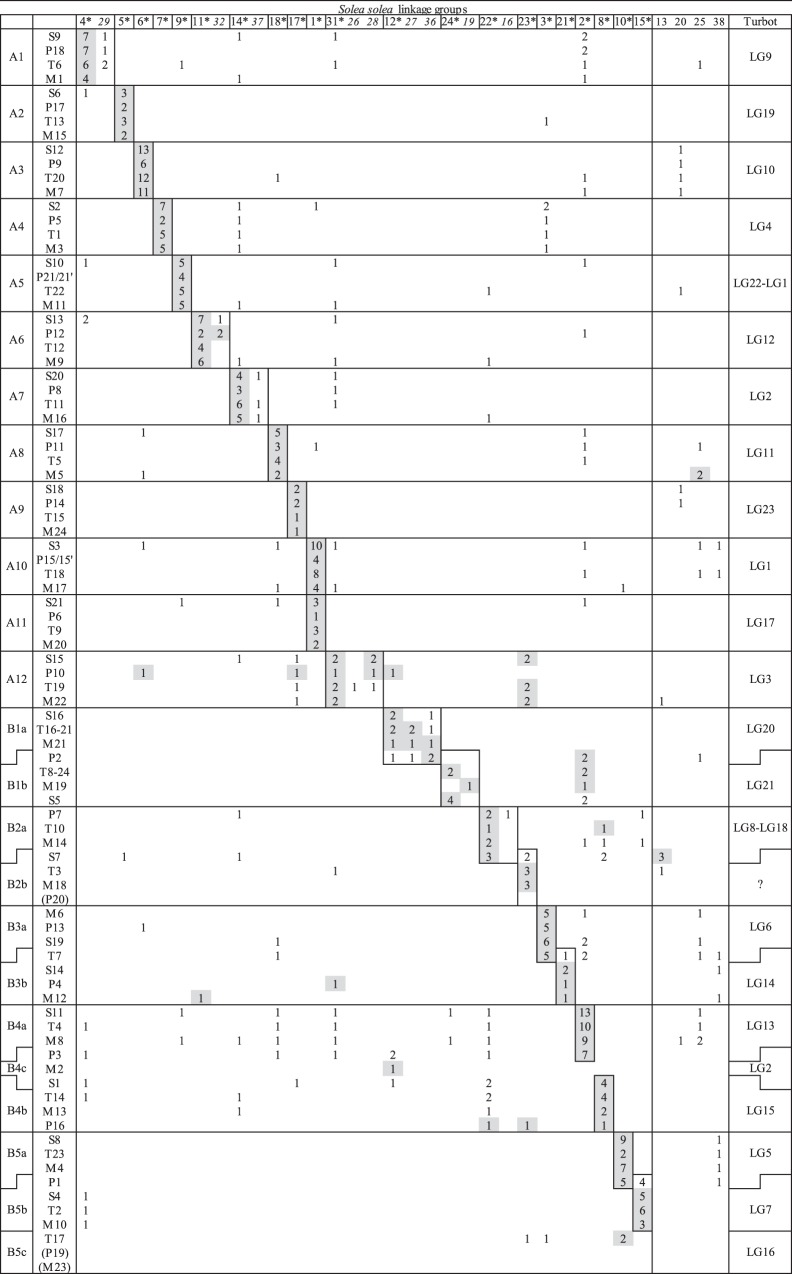
The distribution of the sequence homology between the model species chromosomes and the sole linkage groups is shown in Fig. 2. The chromosomes of the four model species are organized in A-groups (with one-to-one relationships) and B-groups (with inter-chromosomal rearrangements) according to their syntenic relationships as described in Sarropoulou et al. [38], Kai et al. [13] and Guyon et al. [50] (see left column). The middle section shows the number of contigs where sequence homology was found between the sole linkage groups and the chromosomes of the four model species. For each chromosome the sole linkage group with the largest number of homologous sequences is highlighted in grey. For the chromosomes belonging to the same A-group or B-subgroup this was mostly the same linkage group, pointing to the chromosome counterpart (or at least part of it) in sole. In total, twenty-one linkage groups were suggested as chromosome counterpart (marked with *). The remaining 17 linkage groups were in surplus as there are only 21 sole chromosomes [41]. Even with a limited number of blast hits (because of their smaller size), nine of these 17 (in italics) were suggested to be on the same chromosome as some of the other 21 linkage groups, as only hits were found with a single A- or B-subgroup.
Figure 2. Syntenic relationships between sole and five other (flat)fish.
The chromosomes of four model fish species, namely stickleback (S), tilapia (T), pufferfish (P) and medaka (M), were grouped in A- and B-groups according to their syntenic relationships as described in Sarropoulou et al. (2008), Kai et al. (2011) and Guyon et al. (2012) (left column). The numbers in the grid indicate the number of contigs where sequence homology was found between the sole linkage groups and the chromosomes of the four model species. For each chromosome the sole linkage group with the largest number of homologous sequences is highlighted in grey. Marked with *: the 21 linkage groups that are suggested as chromosome counterpart for sole (or at least part of it). In italics: linkage groups likely to be on the same chromosome as the linkage group marked with * to the left of it. For all 21 putative sole chromosomes (except for LG23) a homologous turbot linkage group is suggested (right column).
The comparative analysis implies a high degree of conserved synteny, in addition to several chromosomal rearrangements. Twelve of the 21 putative sole chromosomes predominantly showed a one-to-one syntenic relationship with the model species, namely LG4/29-A1, LG5-A2, LG6-A3, LG7-A4, LG9-A5, LG11/32-A6, LG14/37-A7, LG18-A8, LG17-A9, LG12/27/36-B1a, LG2-B4a and LG8-B4b. Six other putative sole chromosomes also showed a one-to-one syntenic relationship, with the exception of one model species: LG22/16 and LG23, both showed homology with chromosome seven of stickleback (S7), indicating that the fusion event leading to S7 in the stickleback ancestor [38], did probably not occur in the sole ancestor. A similar observation was found between LG3, LG21 and chromosome seven of tilapia (T7), and between LG10, LG15 and chromosome one of pufferfish (P1), indicating that the fusion events leading to T7 and P1 [13], [50] did not occur in sole. Beside these three inter-chromosomal rearrangements between sole and one of the four model species, interestingly, one major inter-chromosomal rearrangement between sole and all four model species was observed, namely LG1, which corresponded to two A-groups (A10 and A11), suggesting a fusion in the lineage leading to sole. Finally, the remaining two putative sole chromosomes, LG31/26/28 and LG24/19, showed conserved synteny with the chromosomes belonging to the synteny groups, A12 and B1b, respectively. However, for A12 additional hits were found with LG23 and for B1b with LG2. The homology of LG23, LG2 and several other linkage groups with multiple chromosomes can be attributed to the presence of contigs that were annotated with multigene families (e.g. myosine heavy chain, alpha-actinin-3-like, ryanodine receptor, major histocompatibility complex and tubulin genes). Paralogous sequences from such families are dispersed among chromosomes and therefore interfere with the true syntenic relationships. For all 21 putative sole chromosomes one synteny group (or two in the case of LG1) was suggested, leaving B4c and B5c. In medaka (n = 24) they represent two additional chromosomes, M2 and M23 [47]. However, M2 merged together with M8 in pufferfish (P3) and with M13 in stickleback (S1), and M23 with M10 in stickleback (S4) [13], [38]. Also in tilapia several arguments suggested that M2 merged with M4 into T23 [50]. Unfortunately, for sole no blast hits were found with M2 nor M23 (except one between M2 and LG12).
Syntenic relationships between sole and turbot
Combining linkage groups in sole chromosomes based on synteny with distantly related model species (i.e. all belonging to different orders within the Percomorphaceae), has to be done with cautious. Therefore, the syntenic relationships were also studied with another flatfish species, and thus much closer related. The sole linkage map was built exclusively with EST-based SNP markers (with the exception of eight anonymous microsatellite markers). Given their higher evolutionary conserved status than anonymous markers [51], [52], they provide a suitable framework for comparative mapping with fully sequenced species, as illustrated in the present study. However, as all markers are sole-specific and not present in the turbot or any other flatfish map, a direct comparison, similar to turbot and brill (Scophthalmus rhombus) [21], is impossible. The syntenic relationships between sole and turbot were established indirectly through evolutionary conservation with the model species [38], [53]. Bouza et al. [20] studied the syntenic relationships between turbot and stickleback, medaka and pufferfish. We used these three stepping stone species to anchor the sole linkage groups to turbot. For all 21 putative sole chromosomes (except for LG23) a homologous turbot linkage group was found (see right column in Fig. 2), suggesting a high degree of conserved synteny between these two flatfish. Additionally, these results support the approach of combining linkage groups based on synteny with model species and putting forward 21 putative sole chromosomes. However, also some differences were discussed below. For LG22/16 two turbot linkage groups (LG8-LG18) were observed, but it was suggested that these two probably are located on the same turbot chromosome. This was justified as LG8 and LG18 were syntenic to a single chromosome in all model species studied by Bouza et al. [20]. Moreover, the turbot map contains 24 linkage groups, so there are still two in surplus, as there are only 22 turbot chromosomes. Next, Bouza et al. [20] suggested a translocation between LG1 and LG22 in the lineage leading to turbot. This was not observed between the corresponding linkage groups of sole (resp. LG1 and LG9). For LG23 of sole (homologous with the chromosomes of synteny group B2b), no turbot homolog was found, whereas for LG16 of turbot (homologous with the chromosomes of synteny group B5c), no sole counterpart was suggested. These findings could be a consequence of different chromosomal rearrangements in the lineages leading to both species, however, more evidence is needed to confirm this. Finally, the fusion that was suggested leading to LG1 of sole, was clearly not observed for turbot. Interestingly, this is consistent with the karyotype of both species and might explain why sole has one chromosome less than turbot [54].
Future applications
The fisheries-induced evolutionary change observed in North Sea sole over the last 60 years [16] might significantly impact the productivity of the stock, and eventually lead to stock collapse. From a conservation point of view, evolutionary insights are of paramount importance to avoid irreversible loss of adaptive genetic diversity. This linkage map based on gene-linked markers allowed for a first exploration of the sole genome. By studying the syntenic relationships with other (flat)fish, these genetic markers were positioned and 21 sole chromosomes were suggested. This knowledge is highly relevant for evolutionary studies and can serve as a roadmap to help identify and locate genes involved in local adaptive differentiation and fisheries-induced selection. Using genome scans and if selection is strong enough, genetic markers in the proximity of these genes will display reduced variation and higher degrees of differentiation (e.g. “selective sweeps”) [10], [55]. Identifying such genomic regions is impossible without knowledge on the location of the genetic markers. From a commercial point of view, a linkage map is an essential tool for selective breeding initiatives, aiming at relieving fishery pressure, while increasing the cost-efficiency of farmed fish production. The long generation time, slow growth and the occurrence of infections such as Black Patch Necrosis, have slowed down the large-scale aquaculture of sole [56]. However, research on the culture of various sole species has gained momentum [18], [57], leading to novel breeding strategies, optimised sustainable feed and improved disease resistance. The promising results from recent experimental selective breeding initiatives of sole [58], [59] further stimulate the development of genomic tools in the field of aquaculture. Maximizing growth is a focal goal in aquaculture. For turbot, a more successful flatfish in aquaculture than sole, knowledge on the locations of several growth-related traits is already available. Given the comparison between turbot and sole, that was provided by the present study, the available genomic information of turbot might point in the direction of growth-related genomic regions in sole, similar as was recently illustrated by Hermida et al. [21] between the two flatfish, turbot and brill.
Conclusion
This study constitutes the first mapping effort of the sole genome using gene-linked markers. This linkage map provides a good genome coverage with a homogenous distribution of the markers. Although genomic resource are limited for sole, this map allowed us to explore genomic architecture through comparative mapping with other fish species. The EST-linked markers offered a useful framework for a comparison with four model species. A remarkable degree of conserved synteny was observed, enabling to reconstruct 21 putative sole chromosomes with a high degree of confidence. Several cases of inter-chromosomal rearrangements were suggested as well, including a fusion in the lineage leading to sole. The stepping stone approach, used to compare the sole and turbot genome, confirmed the even higher degree of conserved synteny between the more related flatfish species. For all putative sole chromosomes (except one) it was possible to detect a turbot homolog. The fusion in sole was not observed in turbot, which is consistent with their karyotypes. This first sole map and comparative analysis represents a good starting point to identify functional genomic regions and associated candidate genes of evolutionary and commercial interest in sole, but also in other Pleuronectiformes and teleosts.
Supporting Information
Details on the genetic markers: marker name, linkage group, position (cM), corresponding contig name and accession number.
(XLSX)
Blast results of sole sequences against the stickleback genome.
(XLSX)
Blast results of sole sequences against the pufferfish genome.
(XLSX)
Blast results of sole sequences against the medaka genome.
(XLSX)
Blast results of sole sequences against the tilapia genome.
(XLSX)
Summary of sole linkage groups and homologous stickleback, pufferfish, medaka and tilapia chromosomes that share at least two markers (i.e. conserved syntenic regions).
(XLSX)
Acknowledgments
The authors wish to thank R. Blonk for providing the DNA samples and SSR genotypes of the two aquaculture families of sole, as well as M. Elferink, R. Crooijman and N. van Bers for helping with linkage map construction.
Funding Statement
Research was funded by the European Science Foundation programme Advances in Farm Animal Genomic Resources (LESC) (grant n° EX/3285). The development of the SNP markers was funded by the European Community seventh Framework Programme during the FishPopTrace project (grant agreement n° KBBE-212399). E.D. acknowledges a PhD grant from the Research Foundation-Flanders (FWO-Vlaanderen). G.E.M. was a post-doctoral researcher funded by the Research Foundation-Flanders (FWO-Vlaanderen) during the writing of this article and is now lecturer at James Cook University (Australia). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Kenchington E, Heino M, Nielsen EE (2003) Managing marine genetic diversity: Time for action? ICES J Mar Sci 60:1172–1176. [Google Scholar]
- 2. Allendorf FW, England PR, Luikart G, Ritchie PA, Ryman N (2008) Genetic effects of harvest on wild animal populations. Trends in Ecology and Evolution 23:327–337. [DOI] [PubMed] [Google Scholar]
- 3. Walsh MR, Munch SB, Chiba S, Conover DO (2006) Maladaptive changes in multiple traits caused by fishing: impediments to population recovery. Ecol Lett 9:142–148. [DOI] [PubMed] [Google Scholar]
- 4. Myers RA, Worm B (2003) Rapid worldwide depletion of predatory fish communities. Nature 423:280–283. [DOI] [PubMed] [Google Scholar]
- 5.Hutchings JA, Reynolds JD (2004) Marine fish population collapses: consequences for recovery and extinction risk. Bioscience 54.
- 6. Jørgensen C, Enberg K, Dunlop ES, Arlinghaus R, Boukal DS, et al. (2007) Ecology - Managing evolving fish stocks. Science 318:1247–1248. [DOI] [PubMed] [Google Scholar]
- 7. Kuparinen A, Merilä J (2007) Detecting and managing fisheries-induced evolution. Trends in Ecology and Evolution 22:652–659. [DOI] [PubMed] [Google Scholar]
- 8. Sharpe DMT, Hendry AP (2009) Life history change in commercially exploited fish stocks: an analysis of trends across studies. Evol Appl 2:260–275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Conover DO, Clarke LM, Munch SB, Wagner GN (2006) Spatial and temporal scales of adaptive divergence in marine fishes and the implications for conservation. J Fish Biol 69:21–47. [Google Scholar]
- 10. Nielsen EE, Hemmer-Hansen J, Larsen PF, Bekkevold D (2009) Population genomics of marine fishes: identifying adaptive variation in space and time. Mol Ecol 18:3128–3150. [DOI] [PubMed] [Google Scholar]
- 11. Wenne R, Boudry P, Hemmer-Hansen J, Lubieniecki KP, Was A, et al. (2007) What role for genomics in fisheries management and aquaculture? Aquat Living Resour 20:241–255. [Google Scholar]
- 12. Canario AVM, Bargelloni L, Volckaert F, Houston RD, Massault C, et al. (2008) Genomics toolbox for farmed fish. Rev Fish Sci 16:3–15. [Google Scholar]
- 13. Kai W, Kikuchi K, Tohari S, Chew AK, Tay A, et al. (2011) Integration of the genetic map and genome assembly of fugu facilitates insights into distinct features of genome evolution in teleosts and mammals. Genome Biology and Evolution 3:424–442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gibson NR (2005) Flatfishes: biology and exploitation. Oxford: Wiley-Blackwell. 416 p.
- 15.ICES (2012) Report of the ICES Advisory Committee, 2012. Book 6: North Sea.
- 16. Mollet FM, Kraak SBM, Rijnsdorp AD (2007) Fisheries-induced evolutionary changes in maturation reaction norms in North Sea sole Solea solea . Marine Ecology-Progress Series 351:189–199. [Google Scholar]
- 17. Cerda J, Manchado M (2013) Advances in genomics for flatfish aquaculture. Genes and Nutrition 8:5–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Cerda J, Douglas S, Reith M (2010) Genomic resources for flatfish research and their applications. J Fish Biol 77:1045–1070. [DOI] [PubMed] [Google Scholar]
- 19. Ruan X, Wang W, Kong J, Yu F, Huang X (2010) Genetic linkage mapping of turbot (Scophthalmus maximus L.) using microsatellite markers and its application in QTL analysis. Aquaculture 308:89–100. [Google Scholar]
- 20.Bouza C, Hermida M, Pardo BG, Vera M, Fernandez C, et al. (2012) An Expressed Sequence Tag (EST)-enriched genetic map of turbot (Scophthalmus maximus): a useful framework for comparative genomics across model and farmed teleosts. BMC Genet 13. [DOI] [PMC free article] [PubMed]
- 21. Hermida M, Rodríguez-Ramilo ST, Hachero-Cruzado I, Herrera M, Sciara AA, et al. (2014) First genetic linkage map for comparative mapping and QTL screening of brill (Scophthalmus rhombus). Aquaculture 420–421 Supplement 1 S111–S120. [Google Scholar]
- 22. Reid DP, Smith C-A, Rommens M, Blanchard B, Martin-Robichaud D, et al. (2007) A genetic linkage map of Atlantic halibut (Hippoglossus hippoglossus L.). Genetics 177:1193–1205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Liao X, Ma H-Y, Xu G-B, Shao C-W, Tian Y-S, et al. (2009) Construction of a genetic linkage map and mapping of a female-specific DNA marker in half-smooth tongue sole (Cynoglossus semilaevis). Mar Biotechnol 11:699–709. [DOI] [PubMed] [Google Scholar]
- 24.Castano-Sanchez C, Fuji K, Ozaki A, Hasegawa O, Sakamoto T, et al. (2010) A second generation genetic linkage map of Japanese flounder (Paralichthys olivaceus). BMC Genomics 11. [DOI] [PMC free article] [PubMed]
- 25. Kang J-H, Kim W-J, Lee W-J (2008) Genetic linkage map of olive flounder, Paralichthys olivaceus . Int J Biol Sci 4:143–149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Mardis ER (2008) The impact of next-generation sequencing technology on genetics. Trends Genet 24:133–141. [DOI] [PubMed] [Google Scholar]
- 27.Blonk RJW (2010) Selecting Sole: Breeding programs for natural-mating populations [Doctoral thesis]: Wageningen University. 174 p.
- 28. Blonk RJW, Komen J, Kamstra A, Crooijmans RPMA, van Arendonk JAM (2009) Levels of inbreeding in group mating captive broodstock populations of Common sole, (Solea solea), inferred from parental relatedness and contribution. Aquaculture 289:26–31. [Google Scholar]
- 29. Nielsen EE, Cariani A, Aoidh EM, Maes GE, Milano I, et al. (2012) Gene-associated markers provide tools for tackling illegal fishing and false eco-certification. Nat Commun 3:851. [DOI] [PubMed] [Google Scholar]
- 30. Iyengar A, Piyapattanakorn S, Stone DM, Heipel DA, Howell BR, et al. (2000) Identification of microsatellite repeats in turbot (Scophthalmus maximus) and Dover sole (Solea solea) using a RAPD-based technique: characterization of microsatellite markers in Dover sole. Mar Biotechnol 2:49–56. [DOI] [PubMed] [Google Scholar]
- 31. Garoia F, Marzola S, Guarniero I, Trentini M, Tinti F (2006) Isolation of polymorphic DNA microsatellites in the common sole Solea vulgaris . Mol Ecol Notes 6:144–146. [Google Scholar]
- 32. Li X, Li J (2009) An almost linear time algorithm for a general haplotype solution on tree pedigrees with no recombination and its extensions. J Bioinf Comput Biol 7:521–545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Green P, Falls K, Crooks S (1990) Documentation for cri-map. St Louis, MI: Washington University School of Medicine.
- 34. Voorrips RE (2002) MapChart: Software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. [DOI] [PubMed] [Google Scholar]
- 35. Fishman L, Kelly AJ, Morgan E, Willis JH (2001) A genetic map in the Mimulus guttatus species complex reveals transmission ratio distortion due to heterospecific interactions. Genetics 159:1701–1716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Chakravarti A, Lasher LK, Reefer JE (1991) A maximum-likelihood method for estimating genome length using genetic-linkage data. Genetics 128:175–182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Betancur RR, Broughton RE, Wiley EO, Carpenter K, Lopez JA, et al. (2013) The tree of life and a new classification of bony fishes. PLoS Curr 18. [DOI] [PMC free article] [PubMed]
- 38. Sarropoulou E, Nousdili D, Magoulas A, Kotoulas G (2008) Linking the genomes of nonmodel teleosts through comparative genomics. Mar Biotechnol 10:227–233. [DOI] [PubMed] [Google Scholar]
- 39. Moen T, Delghandi M, Wesmajervi MS, Westgaard JI, Fjalestad KT (2009) A SNP/microsatellite genetic linkage map of the Atlantic cod (Gadus morhua). Anim Genet 40:993–996. [DOI] [PubMed] [Google Scholar]
- 40. Postlethwait JH, Johnson SL, Midson CN, Talbot WS, Gates M, et al. (1994) A genetic-linkage map for the zebrafish. Science 264:699–703. [DOI] [PubMed] [Google Scholar]
- 41. Pardo BG, Bouza C, Castro J, Martinez P, Sanchez L (2001) Localization of ribosomal genes in Pleuronectiformes using Ag-, CMA(3)-banding and in situ hybridization. Heredity 86:531–536. [DOI] [PubMed] [Google Scholar]
- 42. Song WT, Miao GD, Zhao YW, Niu YZ, Pang RY, et al. (2013) Construction of a microsatellite-based genetic linkage map for half-smooth tongue sole Cynoglossus semilaevis . Curr Zool 59:99–108. [Google Scholar]
- 43. Betancur RR, Li C, Munroe TA, Ballesteros JA, Orti G (2013) Addressing gene tree discordance and non-stationarity to resolve a multi-locus phylogeny of the flatfishes (teleostei: pleuronectiformes). Syst Biol 62:763–785. [DOI] [PubMed] [Google Scholar]
- 44.Mabuchi K, Miya M, Azuma Y, Nishida M (2007) Independent evolution of the specialized pharyngeal jaw apparatus in cichlid and labrid fishes. BMC Evol Biol 7. [DOI] [PMC free article] [PubMed]
- 45. Setiamarga DHE, Miya M, Yamanoue Y, Azuma Y, Inoue JG, et al. (2009) Divergence time of the two regional medaka populations in Japan as a new time scale for comparative genomics of vertebrates. Biol Lett 5:812–816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Meynard CN, Mouillot D, Mouquet N, Douzery EJP (2012) A phylogenetic perspective on the evolution of mediterranean Teleost fishes. PLoS One 7. [DOI] [PMC free article] [PubMed]
- 47. Kasahara M, Naruse K, Sasaki S, Nakatani Y, Qu W, et al. (2007) The medaka draft genome and insights into vertebrate genome evolution. Nature 447:714–719. [DOI] [PubMed] [Google Scholar]
- 48. Brunet FG, Crollius HR, Paris M, Aury J-M, Gibert P, et al. (2006) Gene loss and evolutionary rates following whole-genome duplication in Teleost fishes. Mol Biol Evol 23:1808–1816. [DOI] [PubMed] [Google Scholar]
- 49. Lee AP, Kerk SY, Tan YY, Brenner S, Venkatesh B (2011) Ancient vertebrate conserved noncoding elements have been evolving rapidly in Teleost fishes. Mol Biol Evol 28:1205–1215. [DOI] [PubMed] [Google Scholar]
- 50. Guyon R, Rakotomanga M, Azzouzi N, Coutanceau JP, Bonillo C, et al. (2012) A high-resolution map of the Nile tilapia genome: a resource for studying cichlids and other percomorphs. BMC Genomics 13:222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Yu JK, La Rota M, Kantety RV, Sorrells ME (2004) EST derived SSR markers for comparative mapping in wheat and rice. Mol Genet Genomics 271:742–751. [DOI] [PubMed] [Google Scholar]
- 52. Molina-Luzon MJ, Lopez JR, Navajas-Perez R, Robles F, Ruiz-Rejon C, et al. (2012) Validation and comparison of microsatellite markers derived from Senegalese sole (Solea senegalensis, Kaup) genomic and expressed sequence tags libraries. Mol Ecol Resour 12:956–966. [DOI] [PubMed] [Google Scholar]
- 53. Guyon R, Senger F, Rakotomanga M, Sadequi N, Volckaert FAM, et al. (2010) A radiation hybrid map of the European sea bass (Dicentrarchus labrax) based on 1581 markers: Synteny analysis with model fish genomes. Genomics 96:228–238. [DOI] [PubMed] [Google Scholar]
- 54. Bouza C, Sanchez L, Martínez P (1994) Karotypic characterization of turbot (Scophthalmus maximus) with conventional, fluorochrome and restriction endonuclease-banding techniques. Mar Biol 120:609–613. [Google Scholar]
- 55. Storz JF (2005) Using genome scans of DNA polymorphism to infer adaptive population divergence. Mol Ecol 14:671–688. [DOI] [PubMed] [Google Scholar]
- 56. Howell BR (1997) A re-appraisal of the potential of the sole, Solea solea (L.), for commercial cultivation. Aquaculture 155:355–365. [Google Scholar]
- 57. Imsland AK, Foss A, Conceicao LEC, Dinis MT, Delbare D, et al. (2003) A review of the culture potential of Solea solea and Solea senegalensis . Rev Fish Biol Fish 13:379–407. [Google Scholar]
- 58. Blonk RJW, Komen H, Kamstra A, van Arendonk JAM (2010) Estimating breeding values with molecular relatedness and reconstructed pedigrees in natural mating populations of common sole, Solea Solea . Genetics 184:213–219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Blonk RJW, Komen H, Kamstra A, van Arendonk JAM (2010) Effects of grading on heritability estimates under commercial conditions: A case study with common sole, Solea solea . Aquaculture 300:43–49. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Details on the genetic markers: marker name, linkage group, position (cM), corresponding contig name and accession number.
(XLSX)
Blast results of sole sequences against the stickleback genome.
(XLSX)
Blast results of sole sequences against the pufferfish genome.
(XLSX)
Blast results of sole sequences against the medaka genome.
(XLSX)
Blast results of sole sequences against the tilapia genome.
(XLSX)
Summary of sole linkage groups and homologous stickleback, pufferfish, medaka and tilapia chromosomes that share at least two markers (i.e. conserved syntenic regions).
(XLSX)