Abstract
In multicellular organisms, all the cells are genetically identical but turn genes on or off at the right time to promote differentiation into specific cell types. The regulation of higher-order chromatin structure is essential for genome-wide reprogramming and for tissue-specific patterns of gene expression. The complexity of the genome is regulated by epigenetic mechanisms, which act at the level of DNA, histones, and nucleosomes. Epigenetic machinery is involved in many biological processes, including genomic imprinting, X-chromosome inactivation, heterochromatin formation, and transcriptional regulation, as well as DNA damage repair. In this review, we summarize the recent understanding of DNA methylation, cytosine derivatives, active and passive demethylation pathways as well as histone variants. DNA methylation is one of the well-characterized epigenetic signaling tools. Cytosine methylation of promoter regions usually represses transcription but methylation in the gene body may have a positive correlation with gene expression. The attachment of a methyl group to cytosine residue in the DNA sequence is catalyzed by enzymes of the DNA methyltransferase family. Recent studies have shown that the Ten-Eleven translocation family enzymes are involved in stepwise oxidation of 5-methylcytosine, creating new cytosine derivatives including 5-hydroxymethylcytosine, 5-formylcytosine, and 5-carboxylcytosine. Additionally, histone variants into nucleosomes create another strategy to regulate the structure and function of chromatin. The replacement of canonical histones with specialized histone variants regulates accessibility of DNA, and thus may affect multiple biological processes, such as replication, transcription, DNA repair, and play a role in various disorders such as cancer.
Keywords: Cytosine variants, DNA methylation, DNA methyltransferases, Histone variants, Passive and active demethylation, TET family enzymes
Introduction
In the last decade, epigenetics has become an important topic of genetic research. The classical definition of epigenetics refers to the mitotically and/or meiotically heritable changes in gene activity that does not involve alterations in DNA sequence. This definition emphasizes the heritability of the cellular phenotype, and therefore, it only includes changes in the germ line that can be passed down from generation to generation and changes in dividing cells that can be transferred to daughter cells. Currently, we know that epigenetic changes can be induced by environmental factors at different times in life and are potentially reversible. In 2007, Brenda Weis proposed the broader term of epigenetics that refers to “the study of regulation of gene activity that is not dependent on gene sequence and includes heritable and non-heritable alterations in gene activity and transcriptional potential of a cell” (Brenda Weis at the “Diet, Epigenetic Events, and Cancer Prevention Symposium” on September 27th, 2007, in Washington, D.C./http://prevention.cancer.gov/files/news-events/100908_epigenetics%20meeting%20report%20Sept%202007.pdf).
Epigenetic control operates on three major levels, i.e., on DNA, histones, and nucleosomes. The relationships among these various epigenetic elements are currently being extensively investigated. In this review, data from the literature are analyzed to discuss the significance of DNA methylation and demethylation, cytosine derivatives as well as histone variants in the epigenetic regulation of the genome.
DNA Level
DNA Methylation
DNA methylation is a biochemical process crucial for normal development in higher organisms, and it is the most thoroughly studied epigenetic mark. Methylation entails the covalent attachment of a methyl (CH3) group to the C5 position of a cytosine residue, forming 5-methylcytosine (5mC).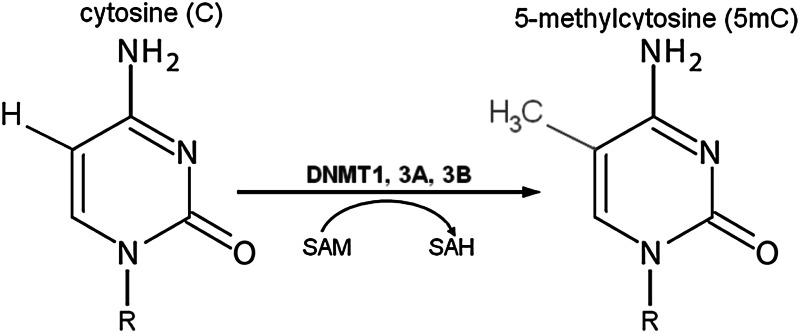
In some organisms, this modification is so frequent that it is denoted as the fifth nucleotide. The methyl group is transferred from S-adenosyl-l-methionine (SAM) to cytosine by the DNA methyltransferase (DNMT) family of enzymes: DNMT1, DNMT2, DNMT3A, DNMT3B, and DNMT3L (Jin et al. 2011). DNMT1 preferentially methylates hemimethylated cytosines in CpG dinucleotide sequences, maintaining the methylation pattern during replication (Probst et al. 2009). In contrast to DNMT1, DNMT3A and 3B prefer unmethylated CpG dinucleotides and perform de novo methylation in early development (Li 2002). Thus, DNMT1 acts primarily as a maintenance methyltransferase during DNA synthesis, and DNMT3A and DNMT3B act as de novo enzymes in development. A growing body of evidence suggests that DNMT1 may also be necessary for de novo methylation of genomic DNA (Egger et al. 2006) and that DNMT3A and DNMT3B are also responsible for the maintenance of methylation during cell replication (Riggs and Xiong 2004). It is worth noting that DNMT2 displays weak DNA methyltransferase activity but actually functions as an RNA methyltransferase. The DNMT2 enzyme specifically methylates cytosine-38 in the anticodon loop of aspartic acid transfer RNA that protects tRNAs from cleavage under stress conditions (Goll et al. 2006; Schaefer et al. 2010).
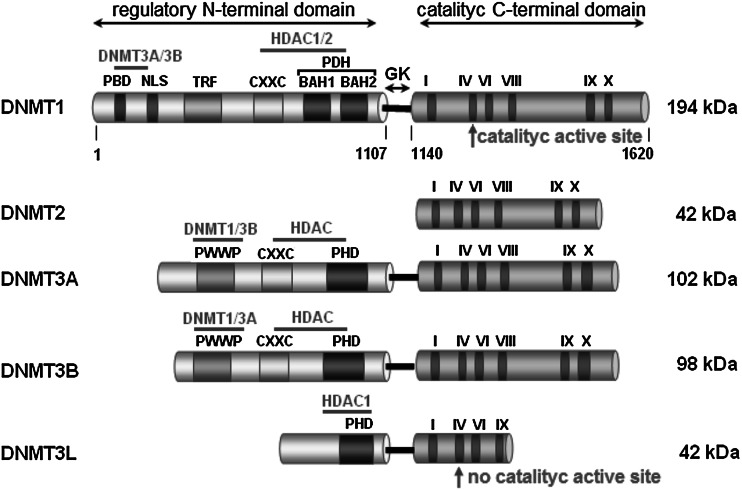
A recent finding has suggested that DNMT2 might be involved in the mammalian paramutation pathway, by protecting small RNA molecules against endonucleolytic cleavage (Adams and Meehan 2013; Kiani et al. 2013), and thus it might induce heritable epigenetic phenotypes. DNMT3L, although it shares homology with DNMT3A and 3B, has no catalytic activity. Instead, DNMT3L increases the ability of DNMT3A and B to bind to methyl groups, thus facilitating methylation in vivo (Bird 2002; Jin et al. 2011). Moreover, DNMT3L recognizes nucleosomes with an unmethylated histone H3 lysine 4 (H3K4) and recruits DNMT3A and DNMT3B to their targets (Saitou et al. 2012). Structural and functional domains of mammalian DNMTs are shown in Fig. 1.
Fig. 1.
Schematic structure of mammalian DNMT family members. DNMT1, the first described methyltransferase, preferentially methylates hemimethylated DNA (Robertson 2001). DNMT2 lacks the N-terminal domain, while C-terminal domain contains the full set of sequence motifs but has not been shown to have transmethylase activity (Bestor 2000). DNMT3A and DNMT3B have similar domain arrangements and an equal preference for hemimethylated and unmethylated DNA (Robertson 2001). DNMT3L, being closely related to the catalytic domain of DNMT3A/3B, lacks canonical DNA cytosine−methyltransferase motifs (Bestor 2000). Its N-terminal regulatory domains exhibit little similarity but the catalytic domains of DNMTs are conserved. The N-terminal domain possesses: PBD—proliferating cell nuclear antigen (PCNA) binding domain, NLS—nuclear localization signal, TRF—targeting replication foci, CXXC—cysteine rich, zinc finger DNA-binding motif, PBH—polybromo homology domain, PWWP—tetrapeptide domain containing proline−tryptophan−tryptophan−proline motif. The N-terminal and C-terminal domains are linked by dinucleotide repeats: GK—glycine−lysine repeat. The C-terminal domain consists of six most conserved amino acid motifs (motif I and X form SAM binding site, motif IV binds cytosine at the active site). Mapped interaction sites of DNMTs and HDACs (histone deacetylases) are indicated in the above diagrams. The borders of the DNMT1 domains are marked according Song et al. (2011)
The level of 5mC affects gene expression, and deregulation of cytosine methylation may play a role in development, cellular differentiation, or disease (Santos-Rebouc and Pimentel 2007; Aguilera et al. 2010; Hackett and Surani 2013). The DNA methylation level can affect transcriptional activities, hypermethylation (a surplus of methyl groups) of promoter regions, and is generally associated with transcriptional silencing, but hypomethylation (a deficit of methyl groups) causes an increased level of gene expression (Crider et al. 2012). Approximately, 2–8 % of the cytosines in the mammalian genome are methylated, mostly in CpG sequences (Zhu 2009; Varriale 2014). In the human genome, CpG dinucleotides are distributed asymmetrically among GC-rich and -poor DNA regions, and not all CpG sites are methylated. The pattern of DNA methylation varies in different cell types and is tissue specific. For example, in differentiated mammalian cells, cytosine residues in GC-rich regions (which typically contain more than 50 % GC) are usually methylated. DNA regions that contain a high frequency of CpG sites are so-called CpG islands (CGIs) and represent an important feature of the mammalian genome. They are located in promoters, preferentially near the transcription start sites (TSSs) of >50 % of human genes. CGI methylation is lower at promoters and higher in gene bodies and intergenic regions. CGI-rich promoters are largely free of DNA methylation due to the abundance of GC-rich transcription factor-binding sites (Deaton and Bird 2011). Methylation of DNA cytosine residues at promoter regions usually correlates with a higher order of chromatin state and repression mRNA transcription. However, Niesen et al. (2005) revealed that a sequence-specific DNA-binding protein can facilitate transcriptional activation of methylated promoter. Interestingly, recent findings suggest that in undifferentiated stem cells, cytosines outside of CpG sites can be methylated as well, and this process is particularly important for the proper regulation of gene expression in embryonic stem cells (ESCs) (Lister et al. 2009). As previously mentioned, gene bodies are highly methylated but the role of methylation remains largely unresolved. Some studies have begun to decipher molecular implications of gene body methylation. For example, methylation in the gene body contributes to the suppression of transcriptional noise (Huh et al. 2011) and might stimulate transcription elongation (Jones 2012). A recent study has suggested that exons are methylated at higher levels than introns and possibly play a role in the regulation of mRNA splicing (Laurent et al. 2010). More details about genomic locations of DNA methylation and its consequence can be found in excellent recent reviews (Estécio and Issa 2011; Moore et al. 2013).
DNA methylation has been considered a stable, persistent and heritable mark; therefore, methyl groups are added but not removed. Recent data have indicated that transcription factors and related proteins not only protect sequences from methylation but also initiate active DNA demethylation (Stadler et al. 2011). Both passive demethylation during replication and active demethylation take place in eukaryotic cells. For example, DNA methylation patterns undergo reprogramming during the establishment of primordial germ cells (PGCs) as well as after fertilization (Branco et al. 2011; Saitou et al. 2012). Surprisingly, the establishment of DNA methylation patterns occurs during development and differentiation of the central nervous system, where it has been implicated in synaptic plasticity, learning, and memory. In the human brain, DNA methylation changes are strongly correlated with age (Hernandez et al. 2011). In turn, pathological activation of DNMTs and aberrant 5mC formation may cause neurodegradation and apoptotic neuronal death (Chestnut et al. 2011; Hernandez and Singleton 2012).
DNA methylation influences gene expression not only by impeding the binding of specific transcription factors but also by recruiting chromatin-modifying proteins. DNA methylation also determines the histone modification patterns and the DNMTs and methyl-CpG-binding domain (MBD) proteins that help to recruit repressor complexes containing histone deacetylases (HDACs) (Fuks et al. 2003). Conversely, interactions between DNMT1, G9a (methyltransferase H3K9), and the replication complex lead to dimethylation of histone H3 lysine 9 (H3K9me2), a repressive epigenetic mark. Methylated H3K9 is bound by heterochromatin protein 1 (HP1), which interacts directly with DNMT1, resulting in cytosine methylation (Smallwood et al. 2007; Saitou et al. 2012). The interaction of the H3K9 methyltransferases (SUV39H1 and ESET) with DNMT3A and DNMT3B can also cause DNA methylation at H3K9me2 (Fuks et al. 2003). Notably, chromatin organization differs between CpG and non-CpG promoters. GC-rich DNA is preferentially bound by CXXC domain proteins that can recruit chromatin modifiers, including the CXXC finger protein 1 (Cfp1) subunit of the H3K3me3 methyltransferase complex and KDM2A, an H3K36me2 demethylase (Vavouri and Lehner 2012). In addition to participating in the histone modifications, DNA methylation may influence the incorporation of histone variant H2A.Z into nucleosomes. A growing body of evidence suggests that the H2A.Z is excluded from methylated DNA and the global anticorrelation between DNA methylation and H2A.Z is observed (Conerly et al. 2010; Weber and Henikoff 2014).
Taken together, DNA methylation affects the interaction between the histone and DNA, resulting in either activation or repression of transcription. It is well known that the disruption of methylation patterns can cause many diseases including cancer, autoimmune disease, as well as chromosomal instability, and mental retardation syndromes (Dobrovic 2010; Javierre et al. 2011). In humans, mutations in genes, including DNMTs and methyl-CpG binding proteins (MBPs), could have profound impact on specific DNA methylation patterns leading to epigenetic diseases (Santos-Rebouc and Pimentel 2007). Up to now, more studies have signified that life style and environmental factors, such as nutrient supply, drugs, pollutants, pathogens, sex hormones, radiation, heavy metals, and early stress can modulate DNA methylation (Javierre et al. 2011; Lim and Song 2012). Interestingly, certain dietary constituents (e.g., folate and bioactive components) may alter genomic and gene-specific DNA methylation levels during embryonic development and adult life (Aguilera et al. 2010; Choi and Friso 2010; McKay and Mathers 2011). Concerning the reversible nature of DNA methylation, it seems to be attractive for epigenetic modulation (Egger et al. 2004; Yang et al. 2010).
Cytosine Variants
It has long been known that cytosine can exist in one of two functional states, unmethylated or methylated. Moreover, mechanisms of DNA methylation are among the best understood epigenetic phenomena. Recently, several cytosine variants, including 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC), 5-carboxylcytosine (5caC), and 3-methylcytosine (3mC), were identified.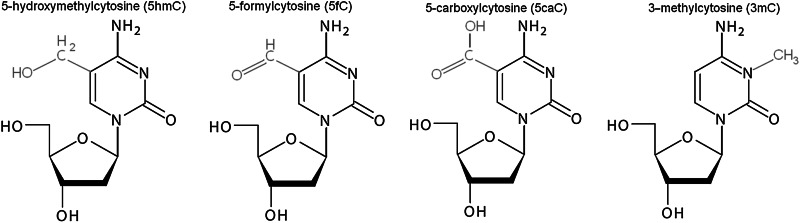
5-Hydroxymethylcytosine (5hmC)
5-Hydroxymethylcytosine was discovered 60 years ago in T2 bacteriophage (Wyatt and Cohen 1952), and 20 years later Penn et al. found 5hmC base in mammalian cells (Penn et al. 1972). These early findings could not be replicated in later studies until 2009, when two independent groups showed that 5hmC exists in mouse Purkinje neurons (Kriaucionis and Heintz 2009) and in ESCs (Tahiliani et al. 2009). Currently, 5hmC is regarded as the “sixth” base of the genome of higher organisms (Münzel et al. 2010). The levels of 5hmC in the genome are relatively low and account for ~0.4 % of all cytosines compared to the ~10 % that are 5mC (Branco et al. 2011). 5hmC constitutes approximately 40 % of the modified cytosines in mouse brain, and the amount increases during maturation in both the hippocampus and the cerebellum (Szulwach et al. 2011). Recently, it has been confirmed that 5hmC is generated by the Ten-Eleven Translocation (TET) enzymes that are Fe(II) and α-oxoglutarate-dependent dioxygenases. The TET subfamily, including TET1, TET2, and TET3, catalyzes the conversion of 5mC–5hm in vitro and in vivo (Ito et al. 2010; Branco et al. 2011) and may be engaged in the further oxidation of 5hmC–5fC and 5caC (He et al. 2011; Ito 2011) (Fig. 2). The TET proteins contain iron and oxyglutarate domains as well cysteine-rich regions that are most likely involved in DNA binding (Iyer et al. 2009). Moreover, TET1 and TET3 contain CXXC zinc finger domains, which allow binding to unmethylated, methylated and hydroxymethylated DNA.
Fig. 2.
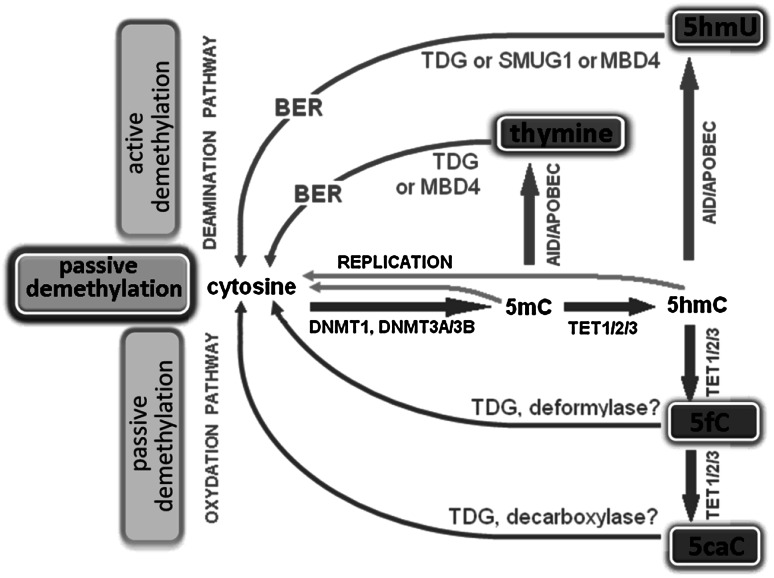
Passive and active DNA demethylation pathways. Passive DNA demethylation is caused by a reduction in activity or absence of DNMTs (yellow arrows). Active demethylation via oxidation pathway (green arrows): TET enzymes can hydroxylate methylcytosine (5mC) to form 5-hydroxymethylcytosine (5hmC); further oxidation produces 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC). 5fC and 5caC can be actively removed by the DNA glycosylases. In addition, a putative deformylase may convert 5fC to C and decarboxylase convert 5caC to C. Active demethylation via deamination pathway (red arrows): AID/APOBEC family members can deaminate 5mC or 5hmC to form thymidine or 5-hydroxymethyluracil (5hmU). These intermediates are replaced by cytosine during base excision repair (BER) mediated by the uracil-DNA glycosylase (UDG) family, like TDG or SMUG1 as well as MBD4 (specifically recognize thymine and 5hmU). AID activation-induced deaminase, APOBEC apolipoprotein B mRNA-editing enzyme complex, BER—base excision repair, DNMT1/3A/3B—DNA methyltransferase, MBD4—methyl-binding domain protein 4, SMUG1—single-strand specific monofunctional uracil-DNA glycosylase, TET1/2/3—ten-eleven methylcytosine dioxygenase family, TDG—thymine-DNA glycosylase (Color figure online)
Other CXXC-containing proteins, for example DNMT1, almost solely bind to unmethylated DNA; therefore, poor recognition of 5hmC could lead to passive demethylation (Valinluck and Sowers 2007). The level of 5hmC in adult tissues is between 0.03 and 0.69 % with the highest levels (0.4–0.7 %) in the central nervous system (Globisch et al. 2010). The biological role of 5hmC is still unclear. It has been postulated that 5hmC could be an intermediate in active DNA demethylation, and it may play an important role in gene regulation (Tahiliani et al. 2009; Wu and Zhang 2010). It has been observed that 5hmC is enriched in the body of the active genes and at the TSSs of transcriptionally inactive genes (Song et al. 2010; Wu et al. 2011a). In vitro analysis revealed that 5hmC in the gene body prevents the binding of MBD proteins, which act as transcriptional repressors (Valinluck et al. 2004; Jin et al. 2010). The level of 5hmC in the gene body might modify the accessibility of chromatin to the transcriptional machinery. Nestor et al. have demonstrated that 5hmC patterns are tissue specific. The global content of 5hmC varies markedly between tissues and does not correlate with global 5mC levels (Nestor et al. 2012). Chen et al. (2012) have demonstrated that aging increases both global- and locus-specific 5hmC content in the mouse hippocampus.
It is possible that 5hmC initiates the pathway of passive or active DNA demethylation by excluding DNMT1 and the MBD proteins from methylating cytosine, and it may recruit other unknown 5hmC-specific effector proteins (Stroud et al. 2011). Recent in vitro studies have revealed that TET proteins could contribute to the removal of methylated cytosine (He et al. 2011; Ito et al. 2011; Matarese et al. 2011). This enzyme family has the capacity to oxidize 5mC not only to 5hmC but also to 5-formylcytosine and 5-carboxylcytosine. Other researchers have shown that thymine-DNA glycosylase (TDG) belonging to the uracil-DNA glycosylase (UDG) superfamily can recognize and excise 5fC and 5caC; thus, the base excision repair (BER) system could be a trigger (Ooi and Bestor 2008; He et al. 2011; Matarese et al. 2011). The crystal structure of human TDG revealed a binding pocket that can accommodate 5caC which facilitates its cleavage (Zhang et al. 2012; Kohli and Zhang 2013). Furthermore, TDG can remove T:G or hmU:G mismatches generated by enzymatic deamination of 5mC to thymine and 5hmC to 5-hydroxymethyluracil (5hmU) (Shen et al. 2014). In addition, alternative UDG glycosylases including methyl-CpG-binding domain protein 4 (MBD4) and single-strand-selective monofunctional uracil-DNA glycosylase 1 (SMUG1) can be involved in active DNA demethylation pathway (Shen et al. 2014). Recent studies have reported that the hydroxylation of 5mC mediated by the Tet1 protein promotes active DNA demethylation in the adult brain by deaminating cytosine residue to uracil by the activation-induced deaminase (AID)/apolipoprotein B mRNA-editing enzyme complex (APOBEC) family, and then deaminated cytosine residue is excised by DNA glycosylases and repaired by the BER pathway (Guo et al. 2011). Potential mechanisms responsible for passive and active demethylation are presented in Fig. 2.
5-Formylcytosine (5fC)
5-Formylcytosine is one of the DNA base variants produced by oxidation of 5hmc by the TET family of enzymes (Ito et al. 2011). Thin layer chromatography and tandem liquid chromatography−mass spectrometry has revealed 5fC in mouse ESCs and in brain cortex (Raiber et al. 2012). The levels of 5fC are estimated to be from 0.02 to 0.002 % of the genomic DNA of ES cells and are 10- to 100-fold lower than the levels of 5hmC (Ito et al. 2011; Pfaffeneder et al. 2011). These levels seem reasonable because TET1 and TET2 are highly expressed and most likely play roles in DNA methylation reprogramming and cell differentiation (Koh et al. 2011). Indeed, during differentiation, levels of 5fC decrease, suggesting its participation in development and germ cell programming (Pfaffeneder et al. 2011). A recent study has reported that CGI promoters were more enriched in 5fC levels than in 5hmC or 5mC levels, which correlated with active gene expression. Moreover, TDG was shown to be actively involved in the removal of 5fC marks in CGIs, exons, and promoter regions (Raiber et al. 2012). Therefore, 5fC excision may help to establish correct methylation patterns during cell-specific developmental programs. Surprisingly, 5fC-enriched promoter regions overlap with H3K4me3, suggesting cross-talk between these marks in transcriptionally active genes.
5-Carboxylcytosine (5caC)
5-Carboxylcytosine is one of the intermediates in active DNA demethylation and is produced by TET-mediated enzymatic oxidation from 5fC. The TET3 protein is most likely responsible for this conversion (Gu et al. 2011). To date, 5caC has been found in ESCs and in mouse pre-implantation embryos (Inoue et al. 2011; He et al. 2011). Alioui and co-workers have shown that 5caC is detectable in the somatic cells of amphibian ovaries and is primarily localized to gene-rich euchromatic regions similar to 5hmC (Alioui et al. 2012). This study also demonstrated that TDG glycosylase can initiate the BER pathway and cleave 5caC both in vitro and in vivo, but the MBD4 enzyme exhibited no activity toward 5caC. Interestingly, 5caC levels increased when TDG was depleted in mouse ES cells; thus, TDG is most likely not the only enzyme capable of processing 5caC (He et al. 2011). It is not known whether TDG is able to recognize and excise 5caC from duplex DNA and whether additional enzymes might be engaged in the conversion of 5caC in mammalian cells.
3-Methylcytosine (3mC)
3-Methylcytosine is a DNA adduct created by spontaneous exposure to endogenous or environmental alkylating agents, leading to cytotoxicity and carcinogenesis. This mutagenic lesion can be directly repaired with the participation of the ABH3 or ABH2 DNA dioxygenases through the BER pathway in humans, or it can be dealkylated by AlkB in bacteria (Koivisto et al. 2004; Yi et al. 2012). Biochemical experiments indicate that ALKBH2 prefers double-stranded DNA (dsDNA) substrates, while ALKBH3 prefers single-stranded DNA (ssDNA) substrates, which are generated by the activating signal cointegrator complex (ASCC) (Dango et al. 2011; Yi et al. 2012). Dango et al. (2011) demonstrated that loss of ALKBH3 or ASCC3 significantly reduced cell proliferation in vitro and in vivo in xenograft models. Concurrently, the accumulation of endogenous 3meC in genomic DNA was observed. Additionally, ALKBH2 has been shown to play an efficient role in pediatric brain tumors during chemotherapy treatment, and the combination of an ALKBH2 knockdown and cisplatin chemotherapy seems to improve the efficacy of treatment (Cetica et al. 2009; Wu et al. 2011b). Taken together, these findings indicate an important role for alkylation repair in removing environmentally induced DNA lesions as well as in maintaining genome integrity and stability.
Histone Variants
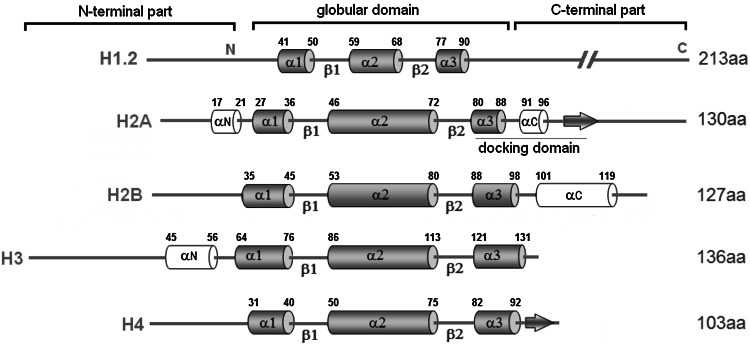
Histones are small, basic, and highly conserved proteins that serve as structural scaffolds for DNA packaging (Cooper 2000). A DNA molecule (~147 bp in length) wrapped around the octamer of a histone (two dimers of H2A–H2B and a heterotetramer (H3–H4)2) constitutes a nucleosome, the fundamental repeating unit of eukaryotic chromatin (Cooper 2000). Histone H1 binds to linker DNA (~50 bp) between nucleosomes, forming a macromolecular structure to help in further compaction of genomic DNA (Sancho et al. 2008). Histone proteins have a tripartite structure consisting of a central globular domain flanked by N- and C-terminal parts (Fig. 3). The unstructured tail located at the N-terminal portion protrudes away from the nucleosome and, therefore, is prone to a variety of post-translational modifications (PTMs) (Kouzarides 2007). The highly conserved globular domain, termed a helix-turn-helix, contains three α–helices separated by loop regions and is involved in histone–histone and histone−DNA interactions (Luger 2001). The C-terminal domains of all histones except histone H1 and H2A are relatively short (Vogler et al. 2010).
Fig. 3.
Schematic structure of the five histone proteins. The N-terminal part is flexible and positively charged and protrudes from the nucleosome. Two short helices, α-1 and α-2 have a length of 10–14 amino acid residues; central α-2 helix comprises ~28 amino acid residues (Luger 2001). The H2A-docking domain spans amino acids 82–119 and is implicated in both structural and functional properties of the nucleosome (Shukla et al. 2011). It stabilizes the wrapping of one helical turn of DNA around the histone octamer (Shukla et al. 2011) and the binding of H2A–H2B dimers to (H3–H4)2 tetramers (Bönisch and Hake 2012). In addition, the H2A C-terminus has also been found to be crucial for binding of the linker histone H1 to nucleosome (Vogler et al. 2010). α helices and β strands of the histone fold extensions are shown as open boxes and arrows, respectively
Histone tails have many positively charged amino acids (especially lysine and arginine), which facilitate their binding to the negatively charged DNA molecule and intranucleosomal interaction (Hansen 2002). The N-terminal histone tails have been studied extensively, but little is known about the function of the C-terminal part. Vogler et al. have shown that the H2A C-terminal tail plays a pivotal role in regulating chromatin structure and dynamics (Vogler et al. 2010). These experiments revealed that the H2A C-terminus is required for efficient nucleosome translocation by chromatin remodelers and acts as a novel recognition module for linker histone H1 (Vogler et al. 2010). It appears that the H2A C-terminal tail has a dual function. On the one hand, it provides stabilization of the nucleosomal core particle, and on the other hand, it participates in interactions with proteins that control chromatin dynamics and conformation.
There are highly similar forms of histones termed ‘histone variants’. It has been estimated that approximately 937 different variants of linker and core histones exist in various species. In humans, 57 histone variants are encoded by 94 genes, the majority of them being present in four clusters: cluster 1 on chromosome 6 (6p22), cluster 2 on chromosome 1 (1p21), cluster 3 on chromosome 1 (1q42), and cluster 4 on chromosome 12 (12p12–13). The incorporation of specific histone variants into nucleosomes has significant impacts on gene expression, heterochromatization, and the formation of specialized regions of the chromatin (Kamakaka and Biggins 2005; Pusarla and Bhargava 2005). The histone variants have recently emerged as important factors in regulating chromatin states and also DNA repair in response to genotoxic treatments (Malik and Henikoff 2003). Moreover, it is likely that histone variants, as potential drivers of cancer initiation and/or progression, thus may be utilized as prognostic indicators of cancer (Vardabasso et al. 2014).
Histone H1
Histone H1 proteins consist of 194–346 amino acid residues, depending on the variant. Approximately 126 different members of the H1 family have been reported from diverse species thus far (http://www.actrec.gov.in/histome/). Eleven variants of histone H1 have been described in humans; these are coded by a single gene that exhibits either replication-dependent or replication-independent expression (Table 1). Three of the variants are testis-specific (i.e., HIST1H1T, H1FNT, and HILS1), one of them is oocyte-specific (H1foo), and the others are somatic variants. Linker histone H1 is involved in chromatin compaction and plays a role in the formation of higher-order chromatin structures (Millán-Ariño et al. 2014). The specific role of histone H1 variants is still far from clear, and genomic distribution of H1 is challenging due to the lack of variant-specific antibodies (Izzo et al. 2013).
Table 1.
Variants of histone H1 in humans
| Variants | Name of genes | Genomic location (Ensembl) | Protein length (aa) (Swiss-Prot) | Function (reference) |
|---|---|---|---|---|
| H1.0b | H1F0 | 22q13.1 | 194 | Nucleosome spacing and chromatin compaction |
| H1.1a | HIST1H1A | 6p22.2 | 215 | Linker histones inhibit sliding of histone octamers, and it is postulated that they can inhibit chromatin remodeling (Clausell et al. 2009) |
| H1.2a | HIST1H1C | 6p22.2 | 213 | |
| H1.3a | HIST1H1D | 6p22.2 | 221 | However, a recent study suggested that H1.2–H1.5 are depleted from active promoters and gene regulatory elements but enriched at regions carrying repressive histone marks (Izzo et al. 2013). H1 binding might be more sensitive to initiation of transcription than to transcriptional elongation (Izzo et al. 2013). Linker histones may operate in conjunction with a ‘network’ of other chromatin-binding proteins so as to define permissive (euchromatin) and repressive (heterochromatin) DNA domains (Ausio 2006). |
| H1.4a | HIST1H1E | 6p22.2 | 219 | |
| H1.5a | HIST1H1B | 6p22.1 | 226 | |
| H1xb | H1FX | 3q21.3 | 213 | |
| H1oo | H1FOO | 3q22.1 | 346 | |
| H1t | HIST1H1T | 6p22.2 | 207 | |
| Testis-specific H1 | H1FNT | 12q13.11 | 255 | |
| Spermatid-specific H1 | HILS1 | 17q21.33 | 231 |
aGene expressed in a replication-dependent manner
bGene expressed in a replication-independent manner
Histone H2A
Histone H2A proteins are composed of ~130 amino acid residues, but atypical variants (macroH2As, H2A.X and H2A-Bbd) differ in size. Approximately 265 different members of histone H2A were identified from a variety of species (http://www.actrec.gov.in/histome/). In humans, nineteen variants of histone H2A encoded by 26 genes were reported (Table 2).
Table 2.
Variants of histone H2A in humans
| Variants | Name of genes | Genomic location (Ensembl) | Protein length (aa) (Swiss-Prot) | Function (reference) |
|---|---|---|---|---|
| H2A type 1 | HIST1H2AI | 6p22.1 | 130 | Stabilization of the histone core octamer (Ausio 2006) |
| HIST1H2AK | ||||
| HIST1H2AL | ||||
| HIST1H2AM | ||||
| HIST1H2AG | ||||
| H2A1 type 1-A | HIST1H2AA | 6p22.2 | 131 | |
| H2A type1-B/E | HIST1H2AE | 6p22.2 | 130 | |
| HIST1H2AB | ||||
| H2A type 1-C | HIST1H2AC | 6p22.2 | 130 | |
| H2A type 1-D | HIST1H2AD | 6p22.2 | 130 | |
| H2A type 1-H | HIST1H2AH | 6p22.1 | 128 | |
| H2A type 1-J | HIST1H2A | 6p22.1 | 128 | |
| H2A type 2-A | HIST2H2AA4 | 1q21.2 | 130 | |
| HIST2H2AA4 | ||||
| H2A type 2-B | HIST2H2AB | 1q21.2 | 130 | |
| H2A type 2-C | HIST2H2AC | 1q21.2 | 129 | |
| H2A type 3 | HIST3H2A | 1q42.13 | 130 | Unknown function |
| H2A-Bbd type 1 | H2AFB1 | Xq28 | 115 | Transcription activation (Tolstorukov et al. 2012) |
| H2A-Bbd type 2/3 | H2AFB2 | Xq28 | 115 | Transcription activation (Tolstorukov et al. 2012) |
| H2AFB3 | ||||
| H2A.J | H2AFJ | 12p12.3 | 129 | unknown function |
| H2A.X | H2AFX | 11q23.3 | 143 | Genome integrity: DNA repair regulation (Pusarla and Bhargava 2005) |
| H2A.Z.1 | H2AFZ | 4q23 | 128 | Maintenance of heterochromatin, transcription repression and activation (Fan et al. 2004; Guillemette et al. 2005; Raisner et al. 2005) |
| H2A.Z.2 | H2AFV | 7p13 | 128 | |
| macroH2A.1 | H2AFY | 5q31.1 | 372 | Silencing: enriched in inactivated chromosome X (Pusarla and Bhargava 2005) |
| macroH2A.2 | H2AFY2 | 10q22.1 | 372 | Silencing: enriched in inactive X-chromosome chromatin and in senescence-associated heterochromatin (Pusarla and Bhargava 2005) |
Histone H2B
Except for four variants, the variants of histone H2B contain 126 amino acid residues. The histone H2B family contains 214 different members described from diverse species (http://www.actrec.gov.in/histome/). Histone H2B forms a dimer with histone H2A in nucleosome cores. Histone H2B has 19 variants encoded by 23 genes in humans, the majority of which are assembled in cluster 1 (i.e., 6p22.1–22.2) (Table 3). There are relatively few PTMs identified among the amino acid residues of histone H2B compared to other core histones.
Table 3.
Variants of histone H2B in humans
| Variants | Name of genes | Genomic location (Ensembl) | Protein length (aa) (Swiss-Prot) | Function (references) |
|---|---|---|---|---|
| H2B type 1-A | HIST1H2BA | 6p22.2 | 127 | Specific role of H2B variants is poorly understood. It is probable that they specialize in chromatin compaction and transcription repression, particularly during gametogenesis (Kamakaka and Biggins 2005) |
| H2B type 1-B | HIST1H2BB | 6p22.2 | 126 | |
| H2B type 1-c/E/F/G/I | HIST1H2BG | 6p22.2 | 126 | |
| HIST1H2BF | ||||
| HIST1H2BE | ||||
| HIST1H2BI | ||||
| HIST1H2BC | ||||
| H2B type 1-D | HIST1H2BD | 6p22.2 | 126 | |
| H2B type 1-H | HIST1H2BH | 6p22.2 | 126 | |
| H2B type 1-J | HIST1H2BJ | 6p22.1 | 126 | |
| H2B type 1-K | HIST1H2BK | 6p22.1 | 126 | |
| H2B type 1-L | HIST1H2BK | 6p22.1 | 126 | |
| H2B type 1-M | HIST1H2BM | 6p22.1 | 126 | |
| H2B type 1-N | HIST1H2BN | 6p22.1 | 126 | |
| H2B type 1-O | HIST1H2BN | 6p22.1 | 126 | |
| H2B type 2-E | HIST2H2BE | 1q21.2 | 126 | |
| H2B type 2-F | HIST2H2BF | 1q21.2 | 126 | |
| H2B type 3-B | HIST3H2BB | 1q42.13 | 126 | |
| H2B type F-M | H2BFM | Xq22.2 | 257 | |
| H2B type F-S | H2BFS | 21q22.3 | 126 | |
| H2B type W-T | H2BFWT | Xq22.2 | 175 | |
| putative H2B type 2-C | HIST2H2BC | 1q21.2 | 193 | |
| putative H2B type 2-D | HIST2H2BD | 1q21.2 | 164 |
Histone H3
Histone H3 consists of ~136 amino acid residues; only the centromere protein A (CENP-A) is a longer variant. The histone H3 family contains 216 different members characterized from various species (http://www.actrec.gov.in/histome/). In humans, 20 genes encode 8 variants of histone H3, most of which are clustered on chromosome 6 (Table 4). Histone H3 is the most extensively post-translationally modified of the five histones.
Table 4.
Variants of histone H3 in humans
| Variants | Name of genes | Genomic location (Ensembl) | Protein length (aa) (Swiss-Prot) | Function (references) |
|---|---|---|---|---|
| H3.1a | HIST1H3A | 6p22.2 | 136 | Replication and DNA repair (Szenker et al. 2011) |
| HIST1HAD | ||||
| HIST1H3C | ||||
| HIST1H3E | ||||
| HIST1H3I | ||||
| HIST1H3G | ||||
| HIST1HAJ | ||||
| HIST1H3H | ||||
| HIST1H3B | ||||
| HIST1H3F | ||||
| H3.1t | HIST3H3 | 1p42.3 | 136 | Chromatin reorganization during spermatogenesis (Witt et al. 1996) |
| H3.2a | HIST2H3C | 1q21.2 | 136 | Replication and DNA repair (Szenker et al. 2011) |
| HIST2H3A | ||||
| HIST2H3D | ||||
| H3.3b | H3F3A | 1p42.2 | 136 | Transcription activation (Mito et al. 2005) |
| H3F3B | 17q25.1 | |||
| H3.3C (also named H3.5) | H3F3C | 12p11.21 | 135 | Transcription activation (Schenk et al. 2011) |
| H3.Xb | H3X | 5p15.1 | Protein not detected in vivo (Wiedemann et al. 2010) | |
| H3.Yb | H3Y | 5p15.1 | Probable regulation of cellular responses to stress, transcription activation (Wiedemann et al. 2010) | |
| H3-like centromeric protein A (CENP-A) | CENPA | 2p23.3 | 140 | Proper chromosome segregation (Fukagawa 2004) |
aGene expressed in a replication-dependent manner
bGene expressed in a replication-independent manner
Histone H4
Histone H4 contains only 103 amino acid residues and forms a heterotetramer (H3–H4)2 with histone H3. The histone H4 family consists of 116 members reported from different organisms (http://www.actrec.gov.in/histome/). Interestingly, humans have a single histone H4 protein encoded by 14 genes, eleven of which are clustered on chromosome 6 (Table 5).
Table 5.
Histone H4 in humans
| Histone | Name of genes | Genomic location (Ensembl) | Protein length (aa) (Swiss-Prot) | Function (references) |
|---|---|---|---|---|
| H4 | HIST4H4 | 12p12.3 | 103 | No variants are recognized |
| HIST1H4A | 6p22.1 | |||
| HIST1H4B | 6p22.2 | |||
| HIST1H4C | 6p22.2 | |||
| HIST1H4D | 6p22.2 | |||
| HIST1H4E | 6p22.2 | |||
| HIST1H4F | 6p22.2 | |||
| HIST1H4H | 6p22.2 | H4 makes contact with the other three histones in the octamer | ||
| HIST1H4I | 6p22.1 | |||
| HIST1H4J | 6p22.1 | |||
| HIST1H4K | 6p22.1 | |||
| HIST1H4L | 6p22.1 | |||
| HIST2H4A | 1q21.2 | |||
| HIST2H4B | 1p21.2 |
Conclusions
DNA methylation is considered to be a relatively stable epigenetic modification. Recent genome-wide analyses of the DNA methylation in mammalian cells suggest that some enzymes are capable of erasing or modifying existing methylation patterns. Although DNA cytosine methylation is well-characterized, little is known about the role of cytosine derivatives in gene expression regulation. In the future, high-resolution sequencing technologies should enable creation of quantitative maps of 5hmC, 5fC, and 5caC in different cell types. Understanding the dynamics of these modifications can help to explain their role in physiological or pathological conditions. Interestingly, due to subtle sequence divergences, incorporation of histone variants may influence the stability of nucleosome and change the potential of specific histone modifications. Histone variant composition is a key player in shaping chromatin structure; this also should be considered as one of the epigenetic regulation elements. It is well known that epigenetic disturbance may lead to different phenotypes and monogenic or complex diseases as well as oncogenic transformation. We strongly believe that rapidly growing understanding of epigenetic phenomena could bring a breakthrough in the diagnosis and treatment of many disorders. Moreover, better knowledge about the epigenetic etiology of the diseases provides an opportunity to develop innovative new epigenetic drugs.
Acknowledgments
This work was supported by the Grant from the National Science Centre no. 2012/06/A/NZ3/00022 and the statutory funds of the Laboratory of Drug Addiction Pharmacology, Institute of Pharmacology PAS.
Conflict of interest
The authors declare that they have no conflict of interests.
References
- Adams IR, Meehan RR. From paramutation to paradigm. PLOS Genet. 2013;9(5):e1003537. doi: 10.1371/journal.pgen.1003537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aguilera O, Fernández AF, Muñoz A, Fraga MF. Epigenetics and environment: a complex relationship. J Appl Physiol. 2010;109:243–251. doi: 10.1152/japplphysiol.00068.2010. [DOI] [PubMed] [Google Scholar]
- Alioui A, Wheldon LM, Abakir A, Ferjentsik Z, Johnson AD, Ruzov A. 5-carboxylcytosine is localized to euchromatic regions in the nuclei of follicular cells in axolotl ovary. Nucleus. 2012;3(6):565–569. doi: 10.4161/nucl.22799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ausio J. Histone variants—the structure behind the function. Brief Funct Genom Proteom. 2006;5(3):228–243. doi: 10.1093/bfgp/ell020. [DOI] [PubMed] [Google Scholar]
- Bestor TH. The DNA methyltransferase of mammals. Hum Mol Genet. 2000;9(16):2395–2402. doi: 10.1093/hmg/9.16.2395. [DOI] [PubMed] [Google Scholar]
- Bird A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002;16:6–21. doi: 10.1101/gad.947102. [DOI] [PubMed] [Google Scholar]
- Bönisch C, Hake SB. Histone H2A variants in nucleosomes and chromatin: more or less stable? Nucleic Acids Res. 2012;40(21):10719–10741. doi: 10.1093/nar/gks865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Branco MR, Ficz G, Reik W. Uncovering the role of 5-hydroxymethylcytosine in the epigenome. Nat Rev Genet. 2011;13(1):7–13. doi: 10.1038/nrg3080. [DOI] [PubMed] [Google Scholar]
- Cetica V, Genitori L, Giunti L, Sanzo M, Bernini G, Massimino M, Sardi I. Pediatric brain tumors: mutations of two dioxygenases (hABH2 and hABH3) that directly repair alkylation damage. J Neurooncol. 2009;94(2):195–201. doi: 10.1007/s11060-009-9837-0. [DOI] [PubMed] [Google Scholar]
- Chen H, Dzitoyeva S, Manev H. Effect of aging on 5-hydroxymethylcytosine in the mouse hippocampus. Restor Neurol Neurosci. 2012;30(3):237–245. doi: 10.3233/RNN-2012-110223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chestnut BA, Chang Q, Price A, Lesuisse C, Wong M, Martin LM. Epigenetic regulation of motor neuron cell death through DNA methylation. J Neurosci. 2011;31(46):16619–16636. doi: 10.1523/JNEUROSCI.1639-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi SW, Friso S. Epigenetics: A new bridge between nutrition and health. Adv Nutr. 2010;1:8–16. doi: 10.3945/an.110.1004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clausell J, Happel N, Hale TK, Doenecke D, Beato M. Histone H1 subtypes differentially modulate chromatin condensation without preventing ATP-Dependent remodeling by SWI/SNF or NURF. PLoS ONE. 2009;4(10):e0007243. doi: 10.1371/journal.pone.0007243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conerly ML, Teves SS, Diolaiti D, Ulrich M, Eisenman RN, Henikoff S. Changes in H2A.Z occupancy and DNA methylation during B-cell lymphomagenesis. Genome Res. 2010;20:1383–1390. doi: 10.1101/gr.106542.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper GM (2000) The cell: a molecular approach 2nd edn. Chapter 4, the organization of cellular genomes. Sinauer Associates, Sunderland
- Crider KS, Yang TP, Berry RJ, Bailey LB. Folate and DNA methylation: a review of molecular mechanisms and the evidence for folate’s role. Adv Nutr. 2012;3:1–38. doi: 10.3945/an.111.000992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dango S, Mosammaparast N, Sowa ME, Xiong LJ, Wu F, Park K, Rubin M, Gygi S, Harper KW, Shi Y. DNA unwinding by ASCC3 helicase is coupled to ALKBH3-dependent DNA alkylation repair and cancer proliferation. Mol Cell. 2011;44(3):373–384. doi: 10.1016/j.molcel.2011.08.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deaton AM, Bird A. CpG islands and the regulation of transcription. Genes and Dev. 2011;25:1010–1022. doi: 10.1101/gad.2037511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dobrovic A. Methods for analysis DNA methylation. In: Coleman WB, Tsongalis GJ, editors. Molecular diagnostics for the clinical laboratorian. 2. New York: Humana Press; 2010. pp. 149–160. [Google Scholar]
- Egger G, Liang G, Aparicio A, Jones PA. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004;429(6990):457–463. doi: 10.1038/nature02625. [DOI] [PubMed] [Google Scholar]
- Egger G, Jeong S, Escobar SG, Cortez CC, Li TW, Saito Y, Yoo CB, Jones PA, Liang G. Identification of DNMT1 (DNA methyltransferase 1) hypomorphs in somatic knockouts suggests an essential role for DNMT1 in cell survival. Proc Natl Acad Sci USA. 2006;103(38):14080–14085. doi: 10.1073/pnas.0604602103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Estécio MRH, Issa JPJ. Dissecting DNA hypermethylation in cancer. FEBS Lett. 2011;585:2078–2086. doi: 10.1016/j.febslet.2010.12.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fan JY, Rangasamy D, Luger K, Tremethick DJ. H2AZ alters the nucleosome surface to promote HP1alpha–mediated chromatin fibre folding. Mol Cell. 2004;16(4):655–661. doi: 10.1016/j.molcel.2004.10.023. [DOI] [PubMed] [Google Scholar]
- Fukagawa T. Centromere DNA proteins and kinetochore assembly in vertebrate cells. Chromosome Res. 2004;12(6):557–567. doi: 10.1023/B:CHRO.0000036590.96208.83. [DOI] [PubMed] [Google Scholar]
- Fuks F, Hurd PJ, Deplus R, Kouzarides T. The DNA methyltransferases associate with HP1 and the SUV39H1 histone methyltransferase. Nucleic Acids Res. 2003;31(9):2305–2312. doi: 10.1093/nar/gkg332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Globisch D, Münzel M, Müller M, Michalakis S, Wagner M, Koch S, Brückl T, Biel M, Carell T. Tissue distribution of 5-hydroxymethylcytosine and search for active demethylation intermediates. PLoS ONE. 2010;5(12):e15367. doi: 10.1371/journal.pone.0015367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goll MG, Kirpekar F, Maggert KA, Yoder JA, Hsieh CL, Zhang X, Golic KG, Jacobsen SE, Bestor TH. Methylation of tRNA Asp by the DNA methyltransferase homolog Dnmt2. Science. 2006;311(5759):395–398. doi: 10.1126/science.1120976. [DOI] [PubMed] [Google Scholar]
- Gu TP, Guo F, Yang H, Wu HP, Xu GF, Liu W, Xie ZG, Shi L, He X, Jin S, Iqbal K, Shi YG, Deng Z, Szabò PE, Pfeifer GP, Li J, Xu GL. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature. 2011;477(7366):606–610. doi: 10.1038/nature10443. [DOI] [PubMed] [Google Scholar]
- Guillemette B, Bataille AR, Gevry N, Adam M, Blanchette M, Robert F, Gaudreau L. Variant histone H2AZ is globally localized to the promoters of inactive yeast genes and regulates nucleosome positioning. PLoS Biol. 2005;3(12):e384. doi: 10.1371/journal.pbio.0030384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo JU, Su Y, Zhong C, Ming GL, Song H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell. 2011;145(3):423–434. doi: 10.1016/j.cell.2011.03.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hackett JA, Surani MA. DNA methylation dynamics during the mammalian life cycle. Phil Trans R Soc B. 2013;368:20110328. doi: 10.1098/rstb.2011.0328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen JC. Conformational dynamics of the chromatin fiber in solution: determinants mechanisms and functions. Ann Rev Biophys Biomol Struct. 2002;31:361–392. doi: 10.1146/annurev.biophys.31.101101.140858. [DOI] [PubMed] [Google Scholar]
- He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, Ding J, Jia Y, Chen Z, Li L, Sun Y, Li X, Dai Q, Song CX, Zhang K, He C, Xu GL. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science. 2011;333(6047):1303–1307. doi: 10.1126/science.1210944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernandez DG, Singleton AB. Using DNA methylation to understand biological consequences of genetic variability. Neurodegener Dis. 2012;9(2):53–59. doi: 10.1159/000333097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernandez DG, Nalls MA, Gibbs JR, Arepalli S, van der Brug M, Chong S, Moore M, Longo DL, Cookson MR, Traynor BJ, Singleton AB. Distinct DNA methylation changes highly correlated with chronological age in the human brain. Hum Mol Genet. 2011;20(6):1164–1172. doi: 10.1093/hmg/ddq561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huh I, Zeng J, Park T, Yi SV. DNA methylation and transcriptional noise. Epigenet Chromatin. 2011;6:9. doi: 10.1186/1756-8935-6-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inoue A, Shen L, Dai Q, He C, Zhang Y. Generation and replication-dependent dilution of 5fC and 5caC during mouse preimplantation development. Cell Res. 2011;21(12):1670–1676. doi: 10.1038/cr.2011.189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito S, D’Alessio AC, Taranova OV, Hong K, Lawrence C, Sowers L, Zhang Y. Role of Tet proteins in 5mC to 5hmC conversion ES-cell self-renewal and inner cell mass specification. Nature. 2010;466(7310):1129–1133. doi: 10.1038/nature09303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito S, Shen L, Dai Q, Wu SC, Collins LB, Swenberg JA, He C, Zhang Y. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science. 2011;333(6047):1300–1303. doi: 10.1126/science.1210597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iyer LM, Tahiliani M, Rao A, Aravind L. Prediction of novel families of enzymes involved in oxidative and other complex modifications of bases in nucleic acids. Cell Cycle. 2009;8(11):1698–1710. doi: 10.4161/cc.8.11.8580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Izzo A, Kamieniarz-Gdula K, Ramírez F, Noureen N, Kind J, Manke T, van Steensel B, Schneider R. The genomic landscape of the somatic linker histone subtypes H1.1 to H1.5 in human cells. Cell Rep. 2013;3:2142–2154. doi: 10.1016/j.celrep.2013.05.003. [DOI] [PubMed] [Google Scholar]
- Javierre BM, Hernando H, Ballestar E. Environmental triggers and epigenetic deregulation in autoimmune disease. Discov Med. 2011;12(67):535–545. [PubMed] [Google Scholar]
- Jin SG, Kadam S, Pfeifer GP. Examination of the specificity of DNA methylation profiling techniques towards 5-methylcytosine and 5-hydroxymethylcytosine. Nucleic Acids Res. 2010;38(11):e125. doi: 10.1093/nar/gkq223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin B, Li Y, Robertson KD. DNA Methylation: superior or subordinate in the epigenetic hierarchy? Genes Cancer. 2011;2(6):607–617. doi: 10.1177/1947601910393957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13(7):484–492. doi: 10.1038/nrg3230. [DOI] [PubMed] [Google Scholar]
- Kamakaka RT, Biggins S. Histone variants: deviants? Genes Dev. 2005;19(3):295–310. doi: 10.1101/gad.1272805. [DOI] [PubMed] [Google Scholar]
- Kiani J, Grandjean V, Liebers R, Tuorto F, Ghanbarian H, Lyko F, Cuzin F, Rassoulzadegan M. RNA–mediated epigenetic heredity requires the cytosine methyltransferase Dnmt2. PLOS Gene. 2013;9(5):e1003498. doi: 10.1371/journal.pgen.1003498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koh KP, Yabuuchi A, Rao S, Huang Y, Cunniff K, Nardone J, Laiho A, Tahiliani M, Sommer CA, Mostoslavsky G, Lahesmaa L, Orkin SH, Rodig SJ, Daley GQ. Rao A Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell. 2011;8(2):200–213. doi: 10.1016/j.stem.2011.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohli RM, Zhang Y. TET enzymes, TDG and dynamics of DNA demethylation. Nature. 2013;502(7472):472–479. doi: 10.1038/nature12750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koivisto P, Robins P, Lindahl T, Sedgwick B. Demethylation of 3-methylthymine in DNA by bacterial and human DNA dioxygenases. J Biol Chem. 2004;279(39):40470–40474. doi: 10.1074/jbc.M407960200. [DOI] [PubMed] [Google Scholar]
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128(4):693–705. doi: 10.1016/j.cell.2007.02.005. [DOI] [PubMed] [Google Scholar]
- Kriaucionis S, Heintz N. The nuclear DNA base 5-hydroxymethylcytosine is present in brain and enriched in Purkinje neurons. Science. 2009;324(5929):929–930. doi: 10.1126/science.1169786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laurent L, Wong E, Li G, Huynh T, Tsirigos A, Ong CT, Low HM, Sung KWK, Rigoutsos I, Loring J, Wei CL. Dynamic changes in the human methylome during differentiation. Genome Res. 2010;20(3):320–331. doi: 10.1101/gr.101907.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li E (2002) Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3(9):662–673 [DOI] [PubMed]
- Lim U, Song MA (2012) Dietary and lifestyle factors of DNA methylation. Methods Mol Biol 863:359–376 [DOI] [PubMed]
- Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature. 2009;462:315–322. doi: 10.1038/nature08514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luger K (2001) Nucleosomes: structure and function. In: Encyclopedia of life sciences. Nature Publishing Group, New York, pp 1–8
- Malik HS, Henikoff S. Phylogenomics of the nucleosome. Nat Struct Biol. 2003;10:882–891. doi: 10.1038/nsb996. [DOI] [PubMed] [Google Scholar]
- Matarese F, Carrillo-de Santa Pau E, Stunnenberg HG. 5-hydroxymethylcytosine: a new kid on the epigenetic block? Mol Syst Biol. 2011;7:562. doi: 10.1038/msb.2011.95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKay JA, Mathers JC. Diet induced epigenetic changes and their implications for health. Acta Physiol. 2011;202:103–118. doi: 10.1111/j.1748-1716.2011.02278.x. [DOI] [PubMed] [Google Scholar]
- Millán-Ariño L, Islam ABMMK, Izquierdo-Bouldstridge A, Mayor R, Terme JM, Luque N, Sancho M, López-Bigaq N, Jordan A. Mapping of six somatic linker histone H1 variants in human breast cancer cells uncovers specific features of H1.2. Nucleic Acid Res. 2014;42(7):4474–4493. doi: 10.1093/nar/gku079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mito Y, Henikoff JG, Henikoff S. Genome-scale profiling of histone H3.3 replacement patterns. Nat Genet. 2005;37:1090–1097. doi: 10.1038/ng1637. [DOI] [PubMed] [Google Scholar]
- Moore LD, Le T, Fan G. DNA methylation and its basic function. Neuropsychopharmacology Reviews. 2013;38:23–38. doi: 10.1038/npp.2012.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Münzel M, Globisch D, Brückl T, Wagner M, Welzmiller V, Michalakis S, Müller M, Biel M, Carell T. Quantification of the sixth DNA base hydroxymethylcytosine in the brain. Angew Chem Int Ed Engl. 2010;49(31):5375–5377. doi: 10.1002/anie.201002033. [DOI] [PubMed] [Google Scholar]
- Nestor CE, Ottaviano R, Reddington J, Sproul D, Reinhardt D, Dunican D, Katz E, Dixon JM, Harrison DJ, Meehan RR. Tissue type is a major modifier of the 5–hydroxymethylcytosine content of human genes. Genome Res. 2012;22(3):467–477. doi: 10.1101/gr.126417.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niesen MI, Osborne AR, Yang H, Rastogi S, Chellappan S, Cheng JQ, Boss JM, Blanck G. Activation of a methylated promoter mediated by a sequence-specific DNA-binding protein, RFX. J Biol Chem. 2005;280:38914–38922. doi: 10.1074/jbc.M504633200. [DOI] [PubMed] [Google Scholar]
- Ooi SK, Bestor TH. The colorful history of active DNA demethylation. Cell. 2008;133(7):1145–1148. doi: 10.1016/j.cell.2008.06.009. [DOI] [PubMed] [Google Scholar]
- Penn NW, Suwalski R, O’Riley C, Bojanowski K, Yura R. The presence of 5-hydroxymethylcytosine in animal deoxyribonucleic acid. Biochem J. 1972;126(4):781–790. doi: 10.1042/bj1260781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfaffeneder T, Hackner B, Truss M, Münzel M, Müller M, Deiml CA, Hagemeier C, Carell T. The discovery of 5-formylcytosine in embryonic stem cell. Angew Chem Int Ed Engl. 2011;50(31):7146–7150. doi: 10.1002/anie.201103899. [DOI] [PubMed] [Google Scholar]
- Probst AV, Dunleavy E, Almouzni G. Epigenetic inheritance during the cell cycle. Nat Rev Mol Cell Biol. 2009;10(3):192–206. doi: 10.1038/nrm2640. [DOI] [PubMed] [Google Scholar]
- Pusarla RH, Bhargava P. Histones in functional diversification core histone variants. FEBS J. 2005;272(20):5149–5168. doi: 10.1111/j.1742-4658.2005.04930.x. [DOI] [PubMed] [Google Scholar]
- Raiber E, Beraldi D, Ficz G, Burgess HE, Branco MR, Murat P, Oxley P, Booth MJ, Reik W, Balasubramanian S. Genome-wide distribution of 5-formylcytosine in embryonic stem cells is associated with transcription and depends on thymine DNA glycosylase. Genome Biol. 2012;13(8):R69. doi: 10.1186/gb-2012-13-8-r69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raisner RM, Hartley PD, Meneghini MD, Bao MZ, Liu CL, Schreiber SL, Rando OJ, Madhani HD. Histone variant H2AZ marks the 5` ends of both active and inactive genes in euchromatin. Cell. 2005;123(2):233–248. doi: 10.1016/j.cell.2005.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riggs AD, Xiong Z. Methylation and epigenetic fidelity. Proc Natl Acad Sci USA. 2004;101(1):4–5. doi: 10.1073/pnas.0307781100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robertson KD. DNA methylation methyltransferases and cancer. Oncogene. 2001;20(24):3139–3155. doi: 10.1038/sj.onc.1204341. [DOI] [PubMed] [Google Scholar]
- Saitou M, Kagiwada S, Kurimoto K. Epigenetic reprogramming in mouse pre-implantation development and primordial germ cells. Development. 2012;139(1):15–31. doi: 10.1242/dev.050849. [DOI] [PubMed] [Google Scholar]
- Sancho M, Diani E, Beato M, Jordan A. Depletion of human H1 variants uncovers specific roles in gene expression and cell growth. PLoS Genet. 2008;4(10):e1000227. doi: 10.1371/journal.pgen.1000227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santos-Rebouc CB, Pimentel MMG. Implication of abnormal epigenetic patterns for human diseases. Eur J Hum Genet. 2007;15:10–17. doi: 10.1038/sj.ejhg.5201727. [DOI] [PubMed] [Google Scholar]
- Schaefer M, Pollex T, Hanna K, Tuorto F, Meusburger M, Helm M, Lyko F. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 2010;24(15):1590–1595. doi: 10.1101/gad.586710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schenk R, Jenke A, Zilbauer M, Wirth S, Postberg J. H3.5 is a novel hominid-specific histone H3 variant that is specifically expressed in the seminiferous tubules of human testes. Chromosoma. 2011;120(3):275–285. doi: 10.1007/s00412-011-0310-4. [DOI] [PubMed] [Google Scholar]
- Shen L, Song CX, He C, Zhang Y. Mechanism and function of oxidative reversal of DNA and RNA methylation. Annu Rev Biochem. 2014;83:585–614. doi: 10.1146/annurev-biochem-060713-035513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shukla MS, Syed SH, Goutte-Gattat D, Richard JL, Montel F, Hamiche A, Travers A, Faivre-Moskalenko C, Bednar J, Hayes JJ, Angelov D, Dimitrov S. The docking domain of histone H2A is required for H1 binding and RSC-mediated nucleosome remodeling. Nucleic Acids Res. 2011;39(7):2559–2570. doi: 10.1093/nar/gkq1174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smallwood A, Estève PO, Pradhan S, Carey M. Functional cooperation between HP1 and DNMT1 mediates gene silencing. Genes Dev. 2007;21(10):1169–1178. doi: 10.1101/gad.1536807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song CX, Szulwach KE, Fu Y, Dai Q, Yi C, Li X, Li Y, Chen CH, Zhang W, Jian X, Wang J, Zhang L, Looney TJ, Zhang B, Godley LA, Hicks LM, Lahn BT, Jin P, He C. Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat Biotechnol. 2010;29(1):68–72. doi: 10.1038/nbt.1732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song J, Rechkoblit O, Bestor TH, Patel DJ. Structure of DNMT1-DNA complex reveals a role for autoinhibition in maintenance DNA methylation. Science. 2011;331(6020):1036–1040. doi: 10.1126/science.1195380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stadler MB, Murr R, Burger L, Ivanek R, Lienert F, Scholer A, van Nimwegen E, Wirbelauer C, Oakeley EJ, Gaidatzis D, Tiwari VK, Schübeler D. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature. 2011;480(7378):490–495. doi: 10.1038/nature10716. [DOI] [PubMed] [Google Scholar]
- Stroud H, Feng S, Kinney SM, Pradhan S, Jacobsen SE. 5-Hydroxymethylcytosine is associated with enhancers and gene bodies in human embryonic stem cells. Genome Biol. 2011;12(6):R5433. doi: 10.1186/gb-2011-12-6-r54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szenker E, Ray-Gallet D, Almouzni G. The double face of the histone variant H33. Cell Res. 2011;21(3):421–434. doi: 10.1038/cr.2011.14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szulwach KE, Li X, Li Y, Song CX, Wu H, Dai Q, Irier H, Upadhyay AK, Gearing M, Levey AI, Vasanthakumar A, Godley LA, Chang Q, Cheng X, He C, Jin P. 5-hmC-mediated epigenetic dynamics during postnatal neurodevelopment and aging. Nat Neurosci. 2011;14(12):1607–1616. doi: 10.1038/nn.2959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, Agarwal S, Iyer LM, Liu DR, Aravind L, Rao A. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324(5929):930–935. doi: 10.1126/science.1170116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tolstorukov MY, Goldman JA, Gilbert C, Ogryzko V, Kingston RE, Park PJ. Histone variant H2ABbd is associated with active transcription and mRNA processing in human cells. Mol Cell. 2012;47(4):596–607. doi: 10.1016/j.molcel.2012.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valinluck V, Sowers LC. Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res. 2007;67(3):946–950. doi: 10.1158/0008-5472.CAN-06-3123. [DOI] [PubMed] [Google Scholar]
- Valinluck V, Tsai HH, Rogstad DK, Burdzy A, Bird A, Sowers LC. Oxidative damage to methyl-CpG sequences inhibits the binding of the methyl-CpG binding domain (MBD) of methyl-CpG binding protein 2 (MeCP2) Nucleic Acids Res. 2004;32(14):4100–4108. doi: 10.1093/nar/gkh739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vardabasso C, Hasson D, Ratnakumar K, Chung CY, Duarte LF, Bernstein E. Histone variants: emerging players in cancer biology. Cell Mol Life Sci. 2014;71(3):379–404. doi: 10.1007/s00018-013-1343-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varriale A (2014) DNA methylation, epigenetics, and evolution in vertebrates: facts and challenges. Int J Evol Biol 475981:7 [DOI] [PMC free article] [PubMed]
- Vavouri T, Lehner B. Human genes with CpG island promoters have a distinct transcription–associated chromatin organization. Genome Biol. 2012;13(11):R110. doi: 10.1186/gb-2012-13-11-r110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogler C, Huber C, Waldmann T, Ettig R, Braun L, Izzo A, Daujat S, Chassignet I, Lopez-Contreras AJ, Fernandez-Capetillo O, Dundr M, Rippe K, Längst G, Schneider R. Histone H2A C-terminus regulates chromatin dynamics remodeling and histone H1 binding. PLoS Genet. 2010;6(12):e1001234. doi: 10.1371/journal.pgen.1001234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber CM, Henikoff S. Histone variants: dynamic punctuation in transcription. Genes Dev. 2014;28:672–682. doi: 10.1101/gad.238873.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wiedemann SM, Mildner SN, Bönisch C, Israel L, Maiser A, Matheisl S, Straub T, Merkl R, Leonhardt H, Kremmer E, Schermelleh L, Hake SB. Identification and characterization of two novel primate-specific histone H3 variants H3X and H3Y. J Cell Biol. 2010;190(5):777–791. doi: 10.1083/jcb.201002043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Witt O, Albig W, Doenecke D. Testis-specific expression of a novel human H3 histone gene. Exp Cell Res. 1996;229(2):301–306. doi: 10.1006/excr.1996.0375. [DOI] [PubMed] [Google Scholar]
- Wu SC, Zhang Y. Active DNA demethylation: many roads lead to Rome. Nat Rev Mol Cell Biol. 2010;11(9):607–620. doi: 10.1038/nrm2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H, D’Alessio AC, Ito S, Wang Z, Cui K, Zhao K, Sun YE, Zhang Y. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev. 2011;25(7):679–684. doi: 10.1101/gad.2036011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu SS, Xu W, Liu S, Chen B, Wang X, Wang Y, Liu S, Wu J. Down-regulation of ALKBH2 increases cisplatin sensitivity in H1299. Acta Pharmacol Sin. 2011;32(3):393–398. doi: 10.1038/aps.2010.216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wyatt GR, Cohen SS. A new pyrimidine base from bacteriophage nucleic acids. Nature. 1952;170:1072–1073. doi: 10.1038/1701072a0. [DOI] [PubMed] [Google Scholar]
- Yang X, Lay F, Han H, Jones PA. Targeting DNA methylation for epigenetic therapy. Trends Pharmacol Sci. 2010;31(11):536–546. doi: 10.1016/j.tips.2010.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi C, Chen B, Qi B, Zhang W, Jia G, Zhang L, Li CJ, Dinner AR, Yang CG, He C. Duplex interrogation by a direct DNA repair protein in search of base damage. Nat Struct Mol Biol. 2012;19(7):671–676. doi: 10.1038/nsmb.2320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L, Lu X, Lu J, Liang H, Dai Q, Xu GL, Luo C, Jiang H, He C. Thymine DNA glycosylase specifically recognizes 5-carboxylcytosine-modified DNA. Nat Chem Biol. 2012;8:328–330. doi: 10.1038/nchembio.914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu JK. Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet. 2009;43:143–166. doi: 10.1146/annurev-genet-102108-134205. [DOI] [PMC free article] [PubMed] [Google Scholar]