Abstract
The move of vertebrates to a terrestrial lifestyle required major adaptations in their locomotory apparatus and reproductive organs. While the fin-to-limb transition has received considerable attention1,2, little is known about the developmental and evolutionary origins of external genitalia. Similarities in gene expression have been interpreted as a potential evolutionary link between the two anatomical structures3-6, yet without providing any underlying developmental mechanism. Here, we have reexamined this question using micro-Computed Tomography (μCT), lineage tracing in three amniote clades and RNA-Seq transcriptional profiling. We show that the developmental origin of external genitalia has shifted through evolution, and in some taxa limbs and genitals share a common primordium. In squamates, the genitalia develop directly from the budding hindlimbs, or the remnants thereof, whereas in mice the genital tubercle originates from the ventral and tail bud mesenchyme. The recruitment of different cell populations for genital outgrowth follows a change in the relative position of the cloaca, the genitalia organizing center. Ectopic grafting of the cloaca demonstrates the conserved ability of different mesenchymal cells to respond to these genitalia-inducing signals. Our results support a limb-like developmental origin of external genitalia as the ancestral condition. Moreover, it suggests that a change in the relative position of the cloacal signaling center during evolution has led to an altered developmental route of external genitalia in mammals, while preserving parts of the ancestral limb molecular circuitry due to a common evolutionary origin.
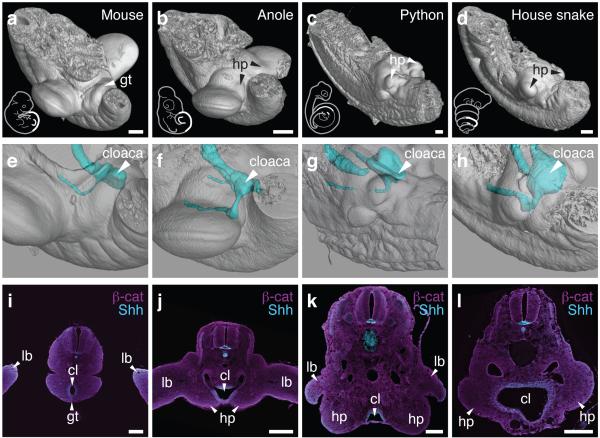
To investigate potential interdependencies in the development of limbs and external genitalia, we first determined the location of the two structures during embryogenesis. We focused on mouse4,5 and squamates (lizard and snakes), which show progressive limb-reduction7, yet maintain their external genitalia, the hemipenes8. μCT-reconstructions of mouse, anole lizard (Anolis), python and house snake embryos revealed different anterior-posterior locations of the developing external genitalia, relative to limbs. In mice, the genital tubercle is positioned caudal to the hindlimbs (Fig. 1a), whereas in squamates the paired hemipenes bud from the limbs, or remnants thereof (Fig. 1b-d). The cloaca, a signaling center important for genitalia development6,9,10, is similarly located within the limb-field of squamates (Fig. 1f-h) and expresses Shh (Fig. 1j-l). Thus in squamates all three anatomical structures, limb, hemipenis and cloaca, align at the same anterior-posterior position.
Figure 1.
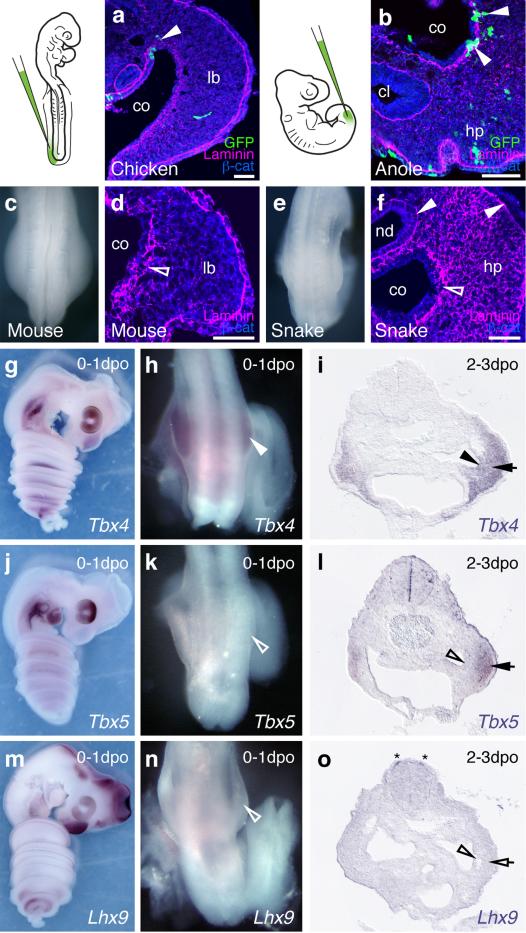
A relative positional shift of limbs, genitalia and the cloaca in squamates. μCT-scans of mouse (a), anole (b), python (c) and house snake (d) lumbo-sacral regions (highlighted in sketch in white) at embryonic stages, illustrating the position of the developing external genitalia. (e-h) 3D-reconstructions of cloacal volumes. The cloaca is located at the same anterior-posterior position as the limb in squamates (f-h), however, is positioned more posteriorly in the mouse (e). (i-l) Transversal sections stained for β-catenin and Shh, indicating the conservation of a cloacal signaling center in all four species. Scale bars, 200 μm. gt: genital tubercle; hp: hemipenis; lb: limb; cl: cloaca.
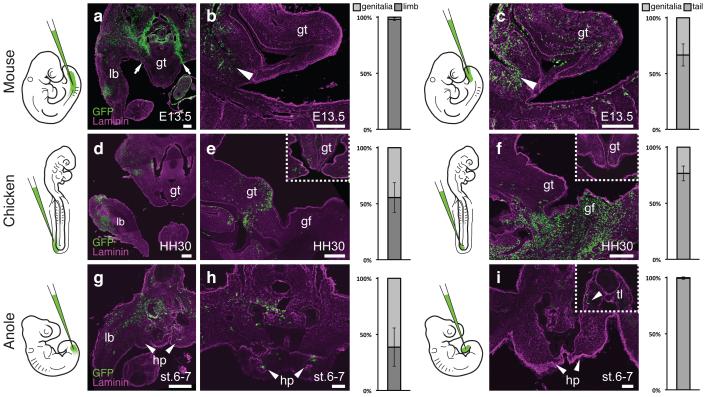
We decided to investigate whether these positional differences would reflect distinct developmental origins of external genitalia. While the cells of the vertebrate limb bud arise through an epithelial-to-mesenchymal transition (EMT) of an epithelial lateral plate mesoderm (LPM) population lining the coelomic cavity11, the developmental origin of external genitalia in vertebrates is still unclear. We developed a lenti-viral lineage tracing approach (see Methods), to systematically follow the two sources previously proposed, the LPM and tail bud10,12,13, in three amniote species, mouse, chicken and anole. Injections into the coelom of embryonic day 9.5 (E9.5) mouse embryos label cells surrounding the coelom as well as the developing hindlimb (Fig. 2a). However, no GFP-positive cells are observed in the genital tubercle, with a sharp boundary of labeled cells extending from the coelomic cavity (Fig. 2a,b). In contrast, injection into the posterior mesenchyme of mouse embryos labels the genital tubercle (Fig. 2c). Tailbud injections label the posterior half, whereas the infra-umbilical mesenchyme gives rise to its anterior part12-14 (Extended Data Fig. 1). In chicken, coelomic injection into stage HH14 embryos labels cells in both limb and genital tubercle (Fig. 2d,e), without any obvious boundary. Tailbud infection also results in GFP-positive genitalia cells, mostly in the posterior tubercle, suggesting that multiple lineages contribute to this species’ genitalia (Fig. 2f). Similar conclusions were reached in a parallel study (Embryonic origin and compartmental organization of the external genitalia, Herrera A. M. & Cohn M. J.). In Anolis, coelomic injections at stage 2-3 result in GFP-positive cells in the limb and the developing genitalia (Fig. 2g,h), whereas no labeled cells are seen in the hemipenes following tailbud injections (Fig. 2i).
Figure 2.
Differential developmental origins of external genitalia in amniotes. (a-i) Transversal and sagittal views of GFP lentivirus injected embryos. Relative contribution of GFP-positive cells to respective organs is quantified on the right, normalized on tissue area. Error bars represent standard deviation in at least n=4 biological replicates. (a,b) Injection into the coelom at mouse E9.5 (n=48) labels the limb at E13.5, but excludes the genital tubercle (arrows). Only cells lining the peritoneal cavity are labeled (b, arrowhead), but none in the genital tubercle proper. c) Injection into the tailbud (n=101) labels cells in the genital tubercle. Accidental piercing of the coelom labels cells of the peritoneal cavity (arrowhead). (d,e) Coelom injection in HH14 chicken embryos (n=81) labels the limb and the genital tubercle at HH30. e) Sagittal and transversal close-up (inset) views. f) Sagittal and transversal close-up (inset) views of tailbud injected chick embryos (n=77), showing labeling in the genital tubercle. (g,h) Anole embryos injected into the coelom at stage 2-3 (n=94) show GFP labeling of both limb and hemipenis at stage 6-7. i) No hemipenis cells are labeled following tailbud injection (n=57), even though there are GFP-positive cells in the tail (inset, arrowhead). Scale bars, 200 μm, 50 μm (h,i). lb: limb; gt: genital tubercle; gf: genital fold; hp: hemipenis; tl: tail.
As for chicken and mouse limbs11, cells of the Anolis limb, but also the hemipenis, originate via an EMT of the coelomic epithelium (Extended Data Fig. 2a,b). In snakes, we find evidence for similar cellular dynamics. The hemipenes emerge as small buds at a “limb-like” lateral position, juxtaposed to the coelomic cavity (Extended Data Fig. 2c-f). A concomitant basement membrane breakdown, consistent with an EMT of the LPM, is seen in the budding of both mouse limbs and snake hemipenes (Extended Data Fig. 2d,f). Moreover, we find that Tbx4, a gene important for hindlimb development15 is expressed from early on, in both the coelomic epithelium and the mesenchyme of the developing hemipenis (Extended Data Fig. 2g-i). Its forelimb counterpart Tbx5 is expressed later, in the mesenchyme only, in agreement with its genitalia-expression in mammals16 (Extended Data Fig. 2j-l). This suggests that squamate external genitalia initiate with limb-like cellular dynamics, the resulting mesenchymal cell population in modern snakes being converted to a genital fate17. Thus, important differences exist in the developmental origins of external genitalia in amniotes: chicken genitals originate from both LPM and the tailbud, whereas the mouse genital tubercle consists of infra-umbilical mesenchyme and tailbud-descendant cells. In contrast, the hemipenis shares a developmental route with hindlimbs, either through secondary budding as in lizards, or by entirely recruiting the mesenchymal cell population to a genital fate in modern snakes. Given the impact of developmental lineage on an organ’s molecular architecture, we next explored the transcriptomes of emerging genitalia, in the two opposing routes of mammals and squamates.
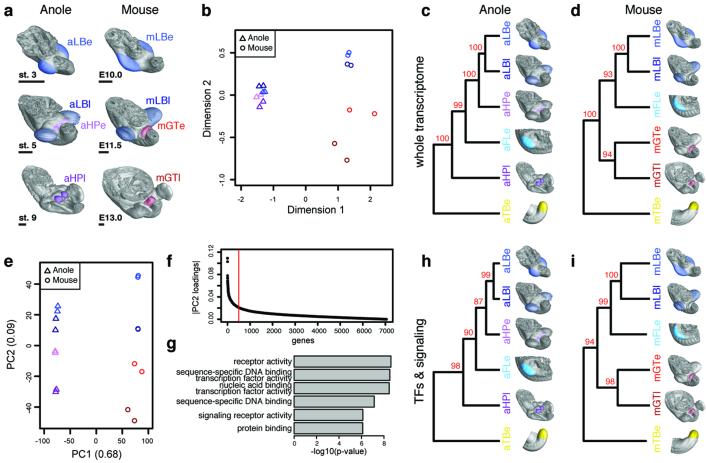
Gene expression profiling has successfully been applied to questions of developmental and evolutionary origin, of cell types and entire morphological features18-21. We thus dissected early and late stages of developing limbs and genitalia from mouse and anole embryos (Fig. 3a), for comparative RNA-seq analyses. Overall transcriptome similarities were assessed using multidimensional scaling (MDS; Fig. 3b). The transcriptomes dominantly resolve along dimension 1 in a species-dependent manner, as expected for similar tissues in evolutionary distant species22. Dimension 2, however, contains a clear organ identity signal, i.e. limb versus genitalia. This separation is virtually absent in Anolis samples, compared to mouse, reflecting similar transcriptional programs due to a common developmental origin for genitalia and hindlimbs. For hierarchical clustering, we included the early tail bud, as outgroup of the primary body axis, and forelimbs, to account for anterior-posterior differences in the two species’ genitalia (Fig. 3c,d; see Methods). In Anolis, the early hemipenis transcriptome falls within the limb clade, indicative of an almost generic limb molecular architecture, and only later differentiates into a more organ-specific signature (Fig. 3c and Extended Data Fig. 3a). In contrast, mouse genitalia transcriptomes are clearly distinct from limbs, from early on, highlighting the separate developmental origins of the two organs (Fig. 3d and Extended Data Fig. 3b).
Figure 3.
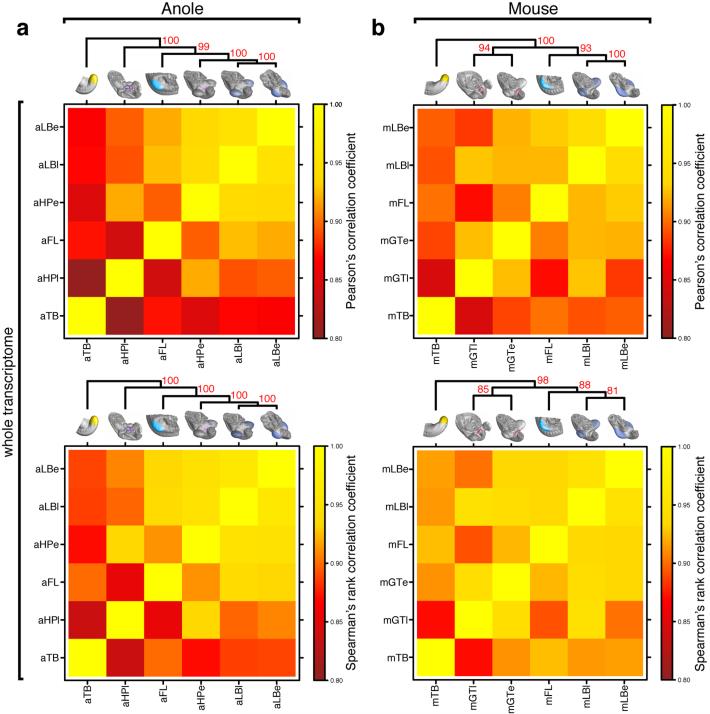
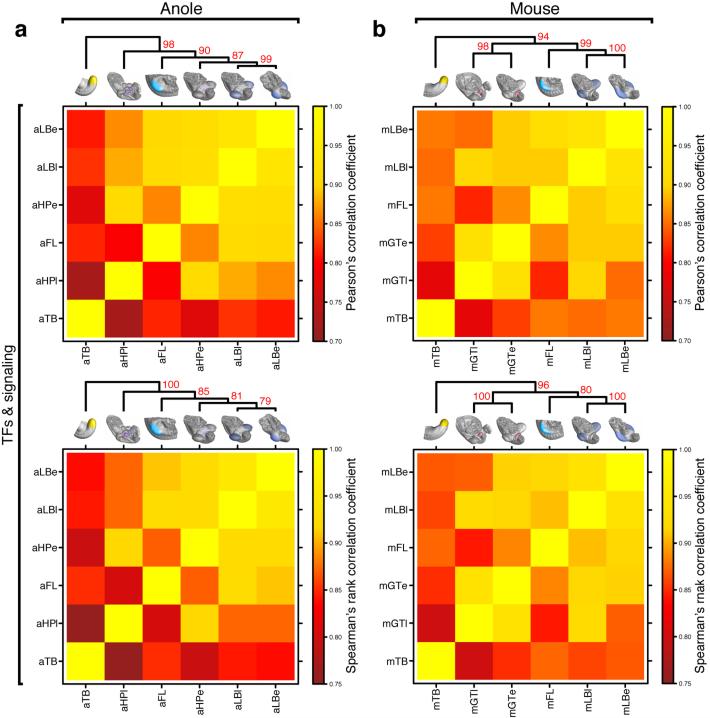
Molecular architecture of limbs and external genitalia in lizards and mice. a) Micro-dissected tissues for RNA-seq analysis, highlighted by color code and four-letter sample identifier. Early and late limb- and genitalia-buds were analyzed, for anole lizard and mouse embryos (n=2). b) Multidimensional scaling (MDS) analysis reveals greater overall transcriptome similarities in anole limb and genitalia datasets (triangles) than in their mouse counterparts (circles). (c,d) Hierarchical clustering of pairwise Pearson’s correlation coefficients, for whole transcriptome data of anole (c) and mouse (d) samples. Additional datasets are stage 2-3 anole and E9.5 mouse forelimb (turquoise) and tailbud (yellow). Numbers at nodes represent approximately unbiased p-values, obtained by multiscale bootstrap resampling. e) Principal component analysis (PCA). Species transcriptomes separate along principal component 1 (PC1), whereas organs are resolved along PC2. f) Absolute loading values for PC2, as shown in (e). g) GO term enrichment analysis using the top500 genes (red bar, f). Top hits include transcription factor- and signaling pathway-related terms. (h,i) Hierarchical clustering analysis of pairwise Pearson’s correlation coefficients, for transcription factors and signaling pathways data of anole (h) and mouse (i) samples. Sample identifiers; a: anole; m: mouse; LB: limb; HP: hemipenis; GT: genital tubercle; e: early; l: late.
To identify genes driving hindlimb- and genitalia-specific transcriptome separation, we used principal component analysis. Principal component 1 (PC1) correlates with species differences, while organ specificity is resolved along PC2 (Fig. 3e), allowing us to identify organ-specific ‘driver’ genes in a species-independent manner. We assessed the contribution of orthologous genes to PC2, according to their absolute loading values (Fig. 3f). Gene Ontology (GO) analysis23 of the top500 genes (Fig. 3f and Supplementary Table 1) identified GO-terms related to transcription factors (TFs) and signaling molecules (Fig. 3g). Gene regulatory networks thus determine limb versus genital organ transcriptomes, but also mirror their developmental origin. Strikingly, TFs/signaling molecule data is sufficient to reproduce the clustering seen with whole-transcriptomes (Fig. 3h,i and Extended Data Fig. 4a,b). Collectively, we find a clear distinction between mouse genitalia and limb transcriptomes, during early and late organogenesis. Such genitalia-specific separation is only seen in late Anolis hemipenes, arguing for a developmental repurposing to a copulatory structure. Importantly, we find transcriptional similarities between early hemipenes and hindlimbs in squamates, illustrating a common developmental origin.
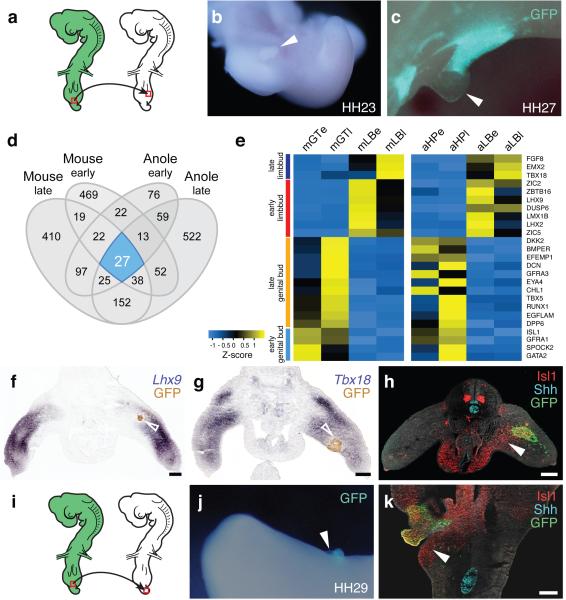
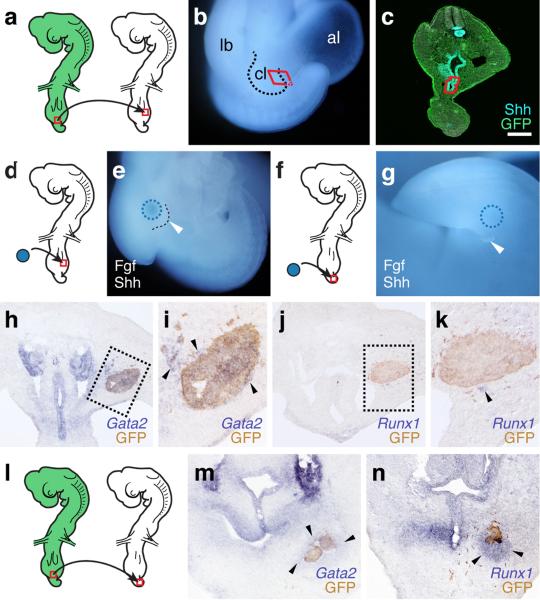
An attractive model for the varied developmental origins of amniote external genitalia would be a repositioning of the cloacal signaling center with respect to different mesenchymal cell populations with progenitor potential. Hence, bringing either hindlimb or tailbud close to the cloaca would allow these lineages to contribute to genital outgrowth. We tested this hypothesis by grafting GFP-transgenic chicken or quail cloacae into the hindlimb bud of wild-type embryos (Fig. 4a and Extended Data Fig. 5a-c). After 1-2 days of incubation, limbs showed ectopic, secondary buds (Fig. 4b,c). GFP-negative cells suggest an inductive effect of the cloaca on the surrounding mesenchyme, rather than simple over-proliferation of the graft itself (Fig. 4c). Similar buds could be induced by grafting beads soaked in known cloacal signaling molecules Shh and Fgf9,10,24,25 (Extended Data Fig. 5d-g). To assess the fate of responding cells, we re-analyzed our RNA-seq data for potential genital versus limb markers. For both species, we performed stage- and organ-specific differential expression analyses. Of the 2003 genes showing an absolute log2 fold-change greater than 1.5 (p-value < 0.05) (Extended Data Fig. 6), we identified 27 that are altered in all four comparisons, 25 of which in the same direction (Fig. 4d). Hierarchical clustering of normalized mouse expression values reveals four stage- and organ-specific signatures, which are largely conserved in Anolis (Fig. 4e). Based on expression patterns and levels (Extended Data Fig. 6 and 7), we chose marker genes to assess transcriptional changes due to ectopic cloacal signals. Indeed, limb cells close to GFP-positive cloacal grafts down-regulate limb markers Lhx9 and Tbx18 (Fig. 4f,g), and ectopically express genital markers Isl126, Gata2 and Runx1 (Fig. 4h and Extended Data Fig. 5j-m). Accordingly, when we graft cloacal tissue into the tailbud mesenchyme, ectopic budding and genital marker expression are equally induced (Fig. 4i-k and Extended Data Fig. 5l-n and 8). Hence, mesenchymal cells from different developmental origins can respond to inductive cloacal signals, generating outgrowths and genitalia-like marker gene expression. Importantly, these results support the idea that changing the relative anterior-posterior position of the cloaca could generate external genitalia with distinct developmental origins during the course of amniote evolution.
Figure 4.
The cloacal signaling center can recruit different mesenchymal cell populations for the outgrowth of external genitalia. a) Schematic of the hindlimb grafting procedure in chicken embryos. GFP-transgenic cloacae are transplanted into the proximal-ventral portion wild-type hindlimbs. b,c) Ectopic outgrowth (arrowheads) on limbs with cloacal grafts (n=30/118). d) Venn diagram of pairwise differential expression analysis results (log(fold change) > 1.5, p-value < 0.05) of limbs versus genital tissues, for early and late budding stages in anole and mouse. e) Heat map of Z-score normalized values of core 25 genes showing consistent differential expression between limbs and genitalia. Mouse row-based hierarchical clustering was re-used for anole samples. f-h) Analysis of limb- and genital-specific markers. Expression of limb markers Lhx9 (f) and Tbx18 (g) is down-regulated near the GFP-positive cloacal graft (empty arrowheads), while genital marker Isl1 (h) is expressed ectopically (arrowhead). i) Schematic of the tail bud grafting procedure in chicken embryos. GFP-transgenic cloacae are transplanted into ventral wild-type tailbuds. j) Ectopic outgrowth on tails with cloacal grafts (n=16/87). k) Genital marker Isl1 is up-regulated ectopically in the tail, close to the GFP-positive cloacal graft (arrowhead). All gene expression assessed in at least n=3. Scale bars, 200 μm.
In summary, we show substantial variation in external genitalia development in extant amniote species. We propose that repositioning the cloaca can recruit different mesenchymal cell populations, either through spatial or heterochronic changes in the dynamics of their emergence. In squamates, the hindlimb is the dominant source, with modern snakes entirely repurposing a mesenchymal bud to a genital fate. In mice, limbs and genitals have discrete developmental origins, the LPM and the ventral and tailbud mesenchyme, respectively, with chicken showing an intermediate state. Moreover, we find that similarities in limb and genitalia transcriptomes are dependent on the cellular source of the primordia from which they emerge. Specifically, there is a higher degree of early transcriptome congruence in species deriving their intromittent organs from limb anlagen. Strikingly, the ability of different mesenchymal cell populations to respond to cloacal, genitalia-inducing signals seems conserved in extant species. It is therefore tempting to speculate that a limb-derived state could represent the ancestral condition in the evolution of external genitalia, as suggested by their position relative to limbs during turtle development17,27 and the bifid genitalia of basal mammals28,29. As such, a developmental continuity between limbs and genitalia could have turned into an ‘evolutionary continuity’ in mammals, as the two organs spatially separated due to a relative repositioning of the cloaca30. Once shared developmental trajectories could thus help to explain molecular similarities still noticeable in species that develop the two organs from distinct cellular sources3,4,19.
METHODS SUMMARY
μCT-scan reconstructions were used to visualize overall embryonic morphology. For lineage tracing, GFP-expressing lentiviruses were injected into the coelomic cavity or tailbud of mouse, chicken and Anolis embryos. RNA-seq was performed on mouse and Anolis embryonic tissue, using an Illumina HiSeq 2000. Normalized gene read-counts were used for downstream analyses. For grafting experiments, GFP-transgenic chicken or quail cloacal tissue was grafted ectopically into hindlimbs or tailbuds of wild-type recipient chicken. Marker gene expression and grafted tissue were assessed with in situ hybridization and immunohistochemistry. See full Methods for details and any associated references.
METHODS
Tissue sample collection
All embryos were collected in accordance with the appropriate Institutional Animal Care and Use Committee guidelines. Timed-pregnant CD1 and C57BL/6 females were purchased from Charles River. Gravid Anolis females were purchased from Candy Quality Reptiles, La Place, LA (IACUC #26-11 and #28-14). Anole housing and egg incubation were done as previously described31. To collect early stage pre-oviposition embryos, females were euthanized by intraperitoneal Euthasol injection and eggs dissected from the oviduct. Snake husbandry and egg collection have been described before32. Fertilized white leghorns chicken eggs were obtained from Charles River Laboratories and incubated at 38°C. For staging of mouse embryos, noon on the day of the vaginal plug was considered as embryonic day 0.5 (E0.5). For chicken and anole embryos, staging was performed according to Hamburger and Hamilton33 or Sanger and colleagues31, respectively. Embryonic tissue was dissected in cold PBS and either fixed in 4% PFA and processed for cryo-embedding or CT scanning, or else directly processed for RNA extraction or stored in RNAlater (Qiagen).
CT scanning and image processing
Embryos were fixed in 4% PFA and stored in 100% ethanol. Staining was done for 48h in 30% PTA (1% (w/v) phosphotungstic acid in water) and 70% ethanol34. Specimens were rinsed and stored in 70% ethanol until image acquisition. CT scans were carried out on a Bruker Skyscan 1173 or a Nikon (Metris) X-tek HMXST225, at 50-57kV, 115-145uA and 810-1000ms exposure time. Voxel sizes ranged from 0.0023 to 0.0056, with 1500 to 2400 total projections. Post-processing of scan data was done in VGStudio MAX 2.2 (Volume Graphics). For 3D re-construction of the cloaca, serial TIFF stacks produced by VGStudio were read into the Imaris software package (Bitplane) and the endodermal epithelium was used as a guide to manually outline the extent of the cloacal volume.
Immunohistochemistry and In Situ Hybridization
Fixed embryos were embedded in 7.5% gelatin / 15% sucrose or dehydrated in sucrose gradients and embedded in OCT. Sectioning was performed on a Leica CM3000 cryostat. For immunohistochemistry, sections were incubated with primary antibodies in PBST (PBS/BSA 0.2%, Triton 0,1% / SDS 0,02%) overnight, washed 2×10min in PBST and incubated for 1h with secondary antibodies. To detect genital tubercle and ectopic limb Isl1 expression, as well as Lmx1b, signal was amplified using the TSA Plus Cy3 kit (Perkin Elmer). Primary antibodies used were anti β (Santa Cruz), Laminin (Sigma), GFP (Abcam), QCPN, Lmx1b and Isl1 (all Developmental Studies Hybridoma Bank). In Situ Hybridization was performed using standard protocols32,35. Fluorescent images were acquired on a Zeiss LSM10 inverted confocal miscroscope. Bright-field images were acquired on a Nikon Eclipse E1000. Whole-mount images were acquired on a Leica MZ FL III. Images were globally processed for color balance and brightness using Adobe Photoshop.
Lineage tracing analyses
For lentiviral lineage tracing, viral particles harboring ubiquitously expressing GFP cassettes (UbiC-GFP or hPGK-GFP) were produced by transient transfection in 293T cells as described elsewhere36. Viral particles were then injected into either the coelomic cavitiy, or the tailbud mesenchyme of mouse E9.5 embryos, chicken HH14 or Anolis st. 2-3. For mouse experiments, timed-pregnant CD1 females were anaesthetized using isoflurane and surgery, in utero visualization of embryos and virus injection was done as previously outlined37. For chicken embryos, eggs were lowered and windowed and virus injected using a pressure injector. For Anolis lineage tracing, we developed a novel whole-embryo ex ovo culturing system, using media conditions previously described for squamate organ cultures38. Briefly, we prepared culture dishes with an indentation, by pouring 1% Agar Noble (BD Difco) dissolved in culture medium into cell culture dishes containing a modeling clay, egg-shaped casting mold. Once solidified, the mold was removed and st. 2-3 Anolis embryos, dissected in 2× PBS, were placed with their yolk intact in the resulting cavity and covered with culture medium. 10% ink in 1× PBS/PenStrep was mouth-pipetted underneath the embryo, for better visualization, and lentiviral particles were injected using a pressure injector. To increase viral infection rate, embryo plates were kept for 12-16h at 37°C in a humidified chamber, before switching them to 28°C. Additional tailbud injections were performed using DiI. Embryos survived for up to 12 days. Only live specimens showing overall normal morphology were considered for further analysis.
To assess GFP+ cell contribution, embryos were dissected, fixed in 4% PFA, gelatin-embedded and cryo-sectioned. Sections were stained for GFP and imaged on a Zeiss LSM10 inverted confocal miscroscope. For quantifications, 4-5 embryos per condition and species were imaged on multiple sections spanning the respective organs. GFP+ cell counting was performed in ImageJ, using the “ITCN” plugin written by Thomas Kuo (UCSB). A total of 59331 GFP+ cells were counted (mouse: 33853; chicken: 23710, anole: 1768). Counts were averaged over multiple sections and normalized on tissue area measured. The resulting ratios of the two tissues are given as a percentage of total GFP+ cells per area.
RNA sample preparation and sequencing
Total RNA was extracted from freshly dissected or RNAlater-preserved tissue using the Arcturus PicoPure RNA Isolation Kit (Life Technologies) and enriched for mRNA fraction using MPG mRNA Purification Kit (PureBiotech). Multiplexed RNA-seq libraries were produced using the SPIA cDNA synthesis kit and the Ovation Ultralow DR Multiplex system (NuGen). Sequencing was done on an Illumina HiSeq 2000, with eight samples multiplexed per lane. Base calling was performed using the Illumina software. Over 561M 50bp reads were generated, with an average of 23.4 reads per sample (median = 22.0M reads). For all tissues, biological duplicates were sequenced.
Read mapping and transcriptome analyses
Initial read mapping was performed with the RNA-Seq unified mapper (RUM)39, using mouse NCBI37/mm9 and Anolis AnoCar2.0 genome assemblies, with UCSC mm9 refseq and ASU_Acar v2.140 annotation files, respectively. This resulted in 10265 orthologuos genes between mouse and Anolis. An in-house improved annotation for Anolis, generated prior to the publication of Eckalbar et al., yielded concordant results in all downstream analyses. To account for species-specific differences in non-uniquely mapping (NU) reads, we redistributed NU reads based on the number of uniquely mapping (U) reads mapping to the respective loci, following a logic outlined before41. Multidimensional scaling analysis was carried out on normalized read counts using the ‘edgeR’ bioconductor package42. Genes differentially expressed between early fore- and hindlimb samples were determined in ‘edgeR’, and genes showing consistent changes in mouse and Anolis were excluded from further analyses, to dampen potential anterior-posterior differences between the organs. For hierarchical clustering, correlation coefficient and principal component analyses, we calculated ‘transcripts per million’ (TPM)43 values, which were then log2-transformed. Hierarchical clustering was done using the ‘pvclust’ R package44, with 1000 iterations of multi-scale bootstrap re-sampling, and approximately unbiased (AU) p-values are provided in the graph. Heat maps of correlation coefficients were plotted with the ‘lattice’ R package. Principal component analysis was done with the ‘prcomp’ function in the ‘stats’ R package and ‘GOseq’45 was used for GO-term enrichment analysis. Pair-wise differential expression analysis between early and late budding stages, in mouse and Anolis limbs and genitalia, was done in ‘edgeR’, and Venn diagram of genes with an absolute log(fold-change) > 1.5 and p-value < 0.05 was visualized using ‘VennDiagram’46.
Grafting experiments
For heteroptopic, homochronic cloacal grafts, donor and recipient embryos were incubated to reach stage HH17-20. Donor embryos were either GFP-transgenic chicken47, purchased from Clemson University, or quail, purchased from Strickland GameBird Farm. Cloacas were dissected in ice-cold PBS and grafted using tungsten needles to a proximal-ventral position, to mimic the squamate configuration, and removed from the apical ectodermal ridge to avoid Shh-induced digit duplications10, or the tailbud in wild-type recipient chicken. Successful grafts were incubated for 1-3 additional days, dissected and screened for the appearance of ectopic outgrowths. Donor versus recipient tissue was discriminated using either GFP or QCPN antibody staining on cryo-sections, or GFP fluorescence for whole-mount embryos. Sham surgery or grafting of GFP-positive limb mesenchyme did not cause any comparable outgrowths. Cloaca-induced outgrowths never stained positive for Alcian blue at later stages, indicating that they were not digit duplications (data not shown). For bead experiments, Affi-Gel Blue Gel beads (150-300μm; Bio-Rad) were washed in PBS and incubated for 1-2h at room temperature, in PBS with recombinant proteins (SHH, FGF2, FGF8; all R&D Systems) at concentrations of 0.1-1μg/μl. Soaked beads were briefly washed in PBS and grafted to limb and tail buds, as outlined for the cloacal grafts. Control grafts using beads soaked in PBS with bovine serum albumin (BSA) did not yield any observable outgrowths.
Extended Data
Extended Data Figure 1.
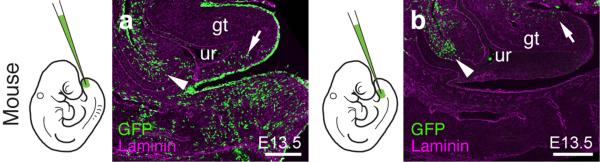
Two separable ventral cell populations give rise to the murine genital tubercle. a,b) Injection into the most distal ventral part of the embryo, the tailbud, marks cells posterior/ventral to the phallic part of the urethra (a, arrow; n=7), whereas injection closer to the allantois, into the infra-umbilical mesenchyme, labels cells anterior/dorsal to the phallic part of the urethra (b, arrow; n=4). Cells lining the peritoneal cavity are also being marked (arrowhead), due to accidental piercing of the coelom. Scale bars, 200 μm. gt: genital tubercle, ur: urethra.
Extended Data Figure 2.
The squamate hemipenis mesenchyme initiates with limb-like cellular dynamics from the coelomic epithelium through a epithelial-to-mesenchymal transition (EMT) a) Injection of GFP-expressing lentiviruses into the coelom of chicken embryos at HH14 labels cells emerging from the epithelium that contribute to the hindlimb mesenchyme (arrowhead). b) In lizards, labeled cells leaving the coelomic epithelium contribute to the hemipenis mesenchyme (arrowheads). c) Dorsal view of the hindlimb region of an E10.0 mouse embryo. d) Transversal section of a limb bud, showing EMT of the coelomic epithelium (diffuse Laminin staining, empty arrowhead), as cells contribute to the limb-bud mesenchyme. e) Dorsal view of the budding hemipenis of a snake embryo, one day after egg deposition. f) Transversal section of the hemipenis region. The basement membrane of the coelomic epithelium is breaking down (empty arrowhead), while it is intact for both the nephric duct and the surface ectoderm (arrowheads). g-o) Expression of genitalia and limb genes during hemipenis initation. g-i) Tbx4 is expressed early and late during hemipenis initiation, in both the coelomic epithelium (arrowhead) and the hemipenis mesenchyme (arrow). j-l) Tbx5 is only expressed later, in the mesenchyme (arrow), but is absent from the coelomic epithelium (empty arrowheads). m-o) Limb marker gene Lhx9 (see also Fig. 4e) is absent from both epithelium (empty arrowhead) and mesenchyme (empty arrow), but can de detected in dI1 neurons (asterisk). All gene expression assessed in at least n=3. Scale bars, 50 μm. co: coelom; lb: limb; hp: hemipenis; nd: nephric duct; cl: cloaca.
Extended Data Figure 3.
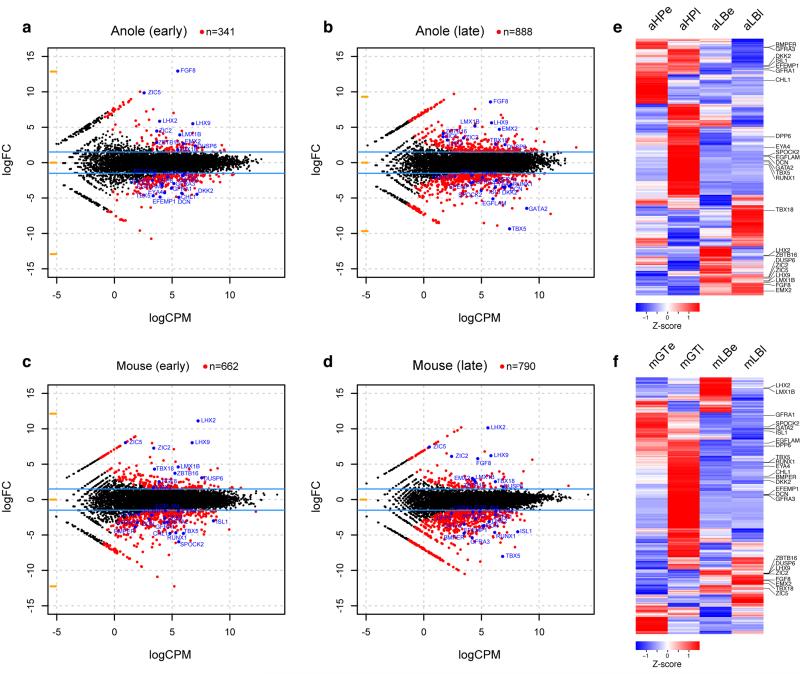
Heat maps of Pearson’s and Spearman’s rank correlation coefficients and cluster analysis of whole transcriptome data. a,b) Hierarchical clustering on pairwise correlation coefficients, for whole transcriptome data of anole (a) and mouse (b) samples. Numbers at nodes represent approximately unbiased p-values obtained by multiscale bootstrap resampling. Sample identifiers; a: anole; m: mouse; LB: limb; HP: hemipenis; GT: genital tubercle; e: early; l: late.
Extended Data Figure 4.
Heat maps of Pearson’s and Spearman’s rank correlation coefficients and cluster analysis of transcription factor and signaling pathway data. a,b) Hierarchical clustering on pairwise correlation coefficients, of transcription factor and signaling pathway data of anole (a) and mouse (b) samples. Numbers at nodes represent approximately unbiased p-values obtained by multiscale bootstrap resampling. Sample identifiers; a: anole; m: mouse; LB: limb; HP: hemipenis; GT: genital tubercle; e: early; l: late.
Extended Data Figure 5.
Heterotopic grafting of the cloacal signaling center leads to ectopic outgrowths and genitalia-like transcriptional changes. a-c) Schematics and close-up images of the cloacal grafting procedure. a) The cloaca of a stage HH17-19 GFP-transgenic chicken embryo (red rectangle) is transplanted into the proximal-ventral portion of the limb of a wild-type embryo. b,c) Only the ventral-most part of the cloaca, including the cloacal membrane, is dissected out (b, red box), and subsequently cleared of excess mesenchymal cells attached to the Shh-expressing endoderm (c, red outline). d-g) Grafting of beads soaked in Shh and Fgf can induce ectopic outgrowths on both limbs (e, n=6/48) and tail (g, n=3/31). h-k) Ectopic expression of genital markers Gata2 (h,i; arrowheads) and Runx1 (j,k; arrowhead) in limb buds, following cloaca-to-limb grafts. l-n) Ectopic expression of genital marker Gata2 (m; arrowheads) and Runx1 (n; arrowheads) in the tail region, following cloaca-to-tail grafts. All gene expression assessed in at least n=3. Scale bar, 200 μm. lb: limb; cl: cloaca; al: allantois.
Extended Data Figure 6.
Pairwise differential expression analysis of limb and genitalia transcriptomes. a-d) Smear plot visualization of differential expression analyses of early anole (a), late anole (b), early mouse (c) and late mouse (d) limb versus genitalia transcriptomes. Genes used for the Venn diagram in Fig. 4d (|log2(fold-change)| > 1.5; p-value < 0.05) are highlighted in red, core 25 marker genes (see Fig. 4e and text) are highlighted and labeled in blue. e,f) Heat map of Z-score normalized expression values for all genes fulfilling Venn diagram criteria (n=2003), for anole (e) and mouse (f) data. Row-based hierarchical clustering was employed; core 25 marker genes are indicated on the right.
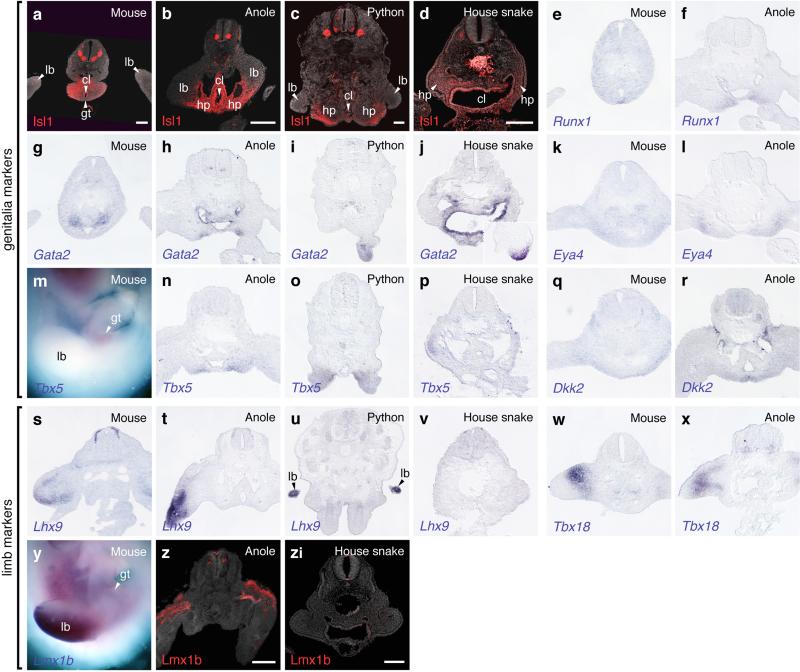
Extended Data Figure 7.
Comparative marker gene expression analysis in mouse and squamate embryos. Genitalia markers Isl1 (a-d), Runx1 (e,f), Gata2 (g-j), Eya (k,l), Tbx5 (m-p) and Dkk2 (q,r). Gata2 only becomes visibly expressed at later stages of house snake hemipenis development (j, inset). Limb markers Lhx9 (s-v), Tbx18 (w,x) and Lmx1b (y-zi). All gene expression assessed in at least n=3. Scale bar, 200 μm. lb: limb; cl: cloaca; gt: genital tubercle; hp: hemipenis.
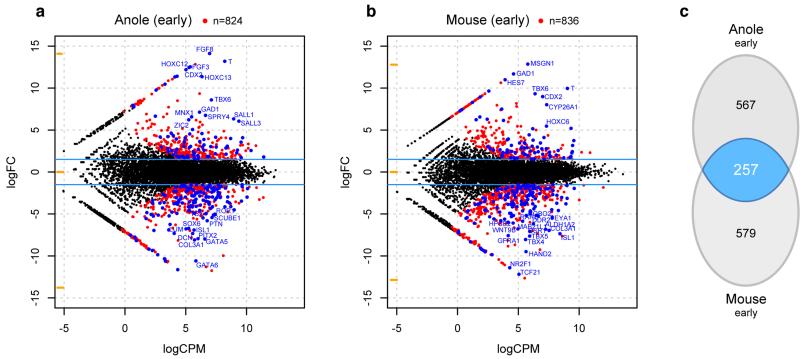
Extended Data Figure 8.
Pairwise differential expression analysis of tailbud and genitalia transcriptomes. a,b) Smear plot visualization of differential expression analyses of early anole (a) and early mouse (b) tailbud versus genitalia transcriptomes. Genes used as input for the Venn diagram in (c) (|log2(fold-change)| > 1.5; p-value < 0.05) are highlighted in red, overlapping 257 marker genes are highlighted in blue. Top 25 genes in the two species, based on logCPM and logFC, are labeled. c) Venn diagram showing overlap of pairwise differential expression analysis results (log(fold change) > 1.5, p-value < 0.05) of tailbud versus genital tissues, for early budding stages in both anole and mouse.
Supplementary Material
Acknowledgements
The authors thank D. Duboule, H. Kaessmann, A. Necsulea, B. Okaty, G. Rey and G.P. Wagner for discussions, M.A. de Bakker for the snake Tbx5 probe and A.M. Herrera and M.J. Cohn for discussing and sharing unpublished results. μCT scans were performed at the Center for Nanoscale Systems, Harvard University (supported by NSF award ECS-0335765) and at the Museum of Comparative Zoology. Next-Generation Sequencing was performed at the HMS Biopolymers Facility and computational analyses were run on the Orchestra Cluster, HMS Research Computing. P.T. was supported by post-doctoral fellowships from the Swiss National Science Foundation, EMBO and the Human Frontiers Science Program. A.C.G. was supported by a post-doctoral fellowship from the Swiss National Science Foundation. This work was supported by NIH grant R37-HD032443 to C.J.T.
Footnotes
Supplementary Information is linked to the online version of the paper at www.nature.com/nature.
Author Contributions
P.T., J.G. and C.J.T conceived the project and designed the experiments. P.T. performed most experiments and computational analyses. E.S. prepared CT scans and helped with statistical analyses. T.S. helped with CT scans, Anolis husbandry and embryo collection. A.C.G. produced lenti-viruses and A.C.A. helped with grafting experiments. J.K.H., O.P. and J.G. initiated snake analyses. O.P. contributed snake embryos. J.G. contributed to chick lineage tracing experiments. P.T., J.G. and C.J.T wrote the paper, with comments from co-authors.
Author Information
Sequencing data has been deposited in the Gene Expression Omnibus under accession number GSE60373. Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests. Readers are welcome to comment on the online version of the paper.
REFERENCES
- 1.Shubin N, Tabin C, Carroll S. Fossils, genes and the evolution of animal limbs. Nature. 1997;388:639–648. doi: 10.1038/41710. [DOI] [PubMed] [Google Scholar]
- 2.Wagner GP, Chiu CH. The tetrapod limb: a hypothesis on its origin. J. Exp. Zool. 2001;291:226–240. doi: 10.1002/jez.1100. [DOI] [PubMed] [Google Scholar]
- 3.Kondo T, Zakany J, Innis JW, Duboule D. Of fingers, toes and penises. Nature. 1997;390:29. doi: 10.1038/36234. [DOI] [PubMed] [Google Scholar]
- 4.Yamada G, et al. Molecular genetic cascades for external genitalia formation: An emerging organogenesis program. Dev. Dyn. 2006;235:1738–1752. doi: 10.1002/dvdy.20807. [DOI] [PubMed] [Google Scholar]
- 5.Cohn MJ. Development of the external genitalia: Conserved and divergent mechanisms of appendage patterning. Dev. Dyn. 2011;240:1108–1115. doi: 10.1002/dvdy.22631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lin C, et al. Delineating a conserved genetic cassette promoting outgrowth of body appendages. PLoS Genet. 2013;9:e1003231. doi: 10.1371/journal.pgen.1003231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Greer AE. Limb reduction in squamates: identification of the lineages and discussion of the trends. Journal of Herpetology. 1991 [Google Scholar]
- 8.Cope ED. On the hemipenes of the Sauria. Proceedings of the Academy of Natural Sciences of Philadelphia. 1896:461–467. [Google Scholar]
- 9.Haraguchi R, et al. Unique functions of Sonic hedgehog signaling during external genitalia development. Development. 2001;128:4241–4250. doi: 10.1242/dev.128.21.4241. [DOI] [PubMed] [Google Scholar]
- 10.Perriton CL, Powles N, Chiang C, Maconochie MK, Cohn MJ. Sonic hedgehog signaling from the urethral epithelium controls external genital development. Dev. Biol. 2002;247:26–46. doi: 10.1006/dbio.2002.0668. [DOI] [PubMed] [Google Scholar]
- 11.Gros J, Tabin CJ. Vertebrate limb bud formation is initiated by localized epithelial-to-mesenchymal transition. Science. 2014;343:1253–1256. doi: 10.1126/science.1248228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ohta S, Suzuki K, Tachibana K, Tanaka H, Yamada G. Cessation of gastrulation is mediated by suppression of epithelial-mesenchymal transition at the ventral ectodermal ridge. Development. 2007;134:4315–4324. doi: 10.1242/dev.008151. [DOI] [PubMed] [Google Scholar]
- 13.Suzuki K, Economides A, Yanagita M, Graf D, Yamada G. New horizons at the caudal embryos: coordinated urogenital/reproductive organ formation by growth factor signaling. Current Opinion in Genetics & Development. 2009;19:491–496. doi: 10.1016/j.gde.2009.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Matsumaru D, et al. Genetic analysis of the role of Alx4 in the coordination of lower body and external genitalia formation. European Journal of Human Genetics. 2013:1–8. doi: 10.1038/ejhg.2013.160. doi:10.1038/ejhg.2013.160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Naiche LA. Loss of Tbx4 blocks hindlimb development and affects vascularization and fusion of the allantois. Development. 2003;130:2681–2693. doi: 10.1242/dev.00504. [DOI] [PubMed] [Google Scholar]
- 16.Chapman DL, Garvey N, Hancock S, Alexiou M. Expression of the T-box family genes, Tbx1-Tbx5, during early mouse development. Developmental …. 1996 doi: 10.1002/(SICI)1097-0177(199608)206:4<379::AID-AJA4>3.0.CO;2-F. [DOI] [PubMed] [Google Scholar]
- 17.Raynaud A, Pieau C, Gans C, Billett FS. Biology of the Reptilia, Development B. Vol. 15. John Wiley & Sons, Inc.; 1985. pp. 149–300. [Google Scholar]
- 18.Arendt D. Genes and homology in nervous system evolution: comparing gene functions, expression patterns, and cell type molecular fingerprints. Theory Biosci. 2005;124:185–197. doi: 10.1007/BF02814483. [DOI] [PubMed] [Google Scholar]
- 19.Wagner GP. The developmental genetics of homology. Nat Rev Genet. 2007;8:473–479. doi: 10.1038/nrg2099. [DOI] [PubMed] [Google Scholar]
- 20.Shubin N, Tabin C, Carroll S. Deep homology and the origins of evolutionary novelty. Nature. 2009;457:818–823. doi: 10.1038/nature07891. [DOI] [PubMed] [Google Scholar]
- 21.Wang Z, Young RL, Xue H, Wagner GP. Transcriptomic analysis of avian digits reveals conserved and derived digit identities in birds. Nature. 2011;477:583–586. doi: 10.1038/nature10391. [DOI] [PubMed] [Google Scholar]
- 22.Merkin J, Russell C, Chen P, Burge CB. Evolutionary Dynamics of Gene and Isoform Regulation in Mammalian Tissues. Science. 2012;338:1593–1599. doi: 10.1126/science.1228186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ashburner M, et al. The Gene Ontology Consortium Gene ontology: tool for the unification of biology. Nature Genetics. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Haraguchi R, et al. Molecular analysis of external genitalia formation: the role of fibroblast growth factor (Fgf) genes during genital tubercle formation. Development. 2000;127:2471–2479. doi: 10.1242/dev.127.11.2471. [DOI] [PubMed] [Google Scholar]
- 25.Miyagawa S, et al. Dosage-dependent hedgehog signals integrated with Wnt/ - catenin signaling regulate external genitalia formation as an appendicular program. Development. 2009;136:3969–3978. doi: 10.1242/dev.039438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Suzuki K, et al. Reduced BMP signaling results in hindlimb fusion with lethal pelvic/urogenital organ aplasia: a new mouse model of sirenomelia. PLoS ONE. 2012;7:e43453. doi: 10.1371/journal.pone.0043453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Larkins CE, Cohn MJ. Phallus Development in the Turtle Trachemys scripta. Sex Dev. 2014 doi: 10.1159/000363631. doi:10.1159/000363631. [DOI] [PubMed] [Google Scholar]
- 28.Griffiths M. The Biology of the Monotremes. Academic Press; 1978. [Google Scholar]
- 29.De Barros MA, et al. Marsupial morphology of reproduction: South America opossum male model. Microsc. Res. Tech. 2013;76:388–397. doi: 10.1002/jemt.22178. [DOI] [PubMed] [Google Scholar]
- 30.Jurberg AD, Aires R, Varela-Lasheras I, Novoa A, Mallo M. Switching Axial Progenitors from Producing Trunk to Tail Tissues in Vertebrate Embryos. Developmental Cell. 2013;25:451–462. doi: 10.1016/j.devcel.2013.05.009. [DOI] [PubMed] [Google Scholar]
- 31.Sanger TJ, Losos JB, Gibson-Brown JJ. A developmental staging series for the lizard genusAnolis: A new system for the integration of evolution, development, and ecology. J. Morphol. 2008;269:129–137. doi: 10.1002/jmor.10563. [DOI] [PubMed] [Google Scholar]
- 32.Gomez C, et al. Control of segment number in vertebrate embryos. Nature. 2008;454:335–339. doi: 10.1038/nature07020. [DOI] [PubMed] [Google Scholar]
- 33.Hamburger V, Hamilton HL. A Series of Normal Stages in the Development of the Chick Embryo. Vol. 88. Journal of Morphology; 1951. pp. 49–92. [PubMed] [Google Scholar]
- 34.Metscher BD. MicroCT for developmental biology: A versatile tool for high-contrast 3D imaging at histological resolutions. Dev. Dyn. 2009;238:632–640. doi: 10.1002/dvdy.21857. [DOI] [PubMed] [Google Scholar]
- 35.McGlinn E, Mansfield JH. Detection of gene expression in mouse embryos and tissue sections. Methods Mol. Biol. 2011;770:259–292. doi: 10.1007/978-1-61779-210-6_10. [DOI] [PubMed] [Google Scholar]
- 36.Barde I, Salmon P, Trono D. Production and Titration of Lentiviral Vectors. John Wiley & Sons, Inc.; 2001. doi:10.1002/0471142301.ns0421s53. [DOI] [PubMed] [Google Scholar]
- 37.Punzo C, Cepko CL. Ultrasound-guided in utero injections allow studies of the development and function of the eye. Dev. Dyn. 2008;237:1034–1042. doi: 10.1002/dvdy.21500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Buchtová M, et al. Initiation and patterning of the snake dentition are dependent on Sonic Hedgehog signaling. Dev. Biol. 2008;319:132–145. doi: 10.1016/j.ydbio.2008.03.004. [DOI] [PubMed] [Google Scholar]
- 39.Grant GR, et al. Comparative Analysis of RNA-Seq Alignment Algorithms and the RNA-Seq Unified Mapper (RUM) Bioinformatics. 2011 doi: 10.1093/bioinformatics/btr427. doi:10.1093/bioinformatics/btr427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Eckalbar WL, et al. Genome reannotation of the lizard Anolis carolinensis based on 14 adult and embryonic deep transcriptomes. BMC Genomics. 2013;14:49. doi: 10.1186/1471-2164-14-49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Brawand D, et al. The evolution of gene expression levels in mammalian organs. Nature. 2011;478:343–348. doi: 10.1038/nature10532. [DOI] [PubMed] [Google Scholar]
- 42.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26:139–140. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wagner GP, Koryu K, Lynch VJ. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. 2012:1–5. doi: 10.1007/s12064-012-0162-3. doi:10.1007/s12064-012-0162-3. [DOI] [PubMed] [Google Scholar]
- 44.Suzuki R, Shimodaira H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics. 2006;22:1540–1542. doi: 10.1093/bioinformatics/btl117. [DOI] [PubMed] [Google Scholar]
- 45.Young MD, Wakefield MJ, Smyth GK, Oshlack A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 2010;11:R14. doi: 10.1186/gb-2010-11-2-r14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Chen H, Boutros PC. VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinformatics. 2011;12:1–28. doi: 10.1186/1471-2105-12-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.McGrew MJ, et al. Localised axial progenitor cell populations in the avian tailbud are not committed to a posterior Hox identity. Development. 2008;135:2289–2299. doi: 10.1242/dev.022020. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.