Abstract
Members of the cadherin superfamily of proteins are involved in diverse biological processes such as morphogenesis, sound transduction, and neuronal connectivity. Key to cadherin function is their extracellular domain containing cadherin repeats, which can mediate interactions involved in adhesion and cell signaling. Recent cellular, biochemical, and structural studies have revealed that physical interaction among cadherins is more complex than originally thought. Here we review work on new cadherin complexes and discuss how the classification of the mammalian family can be used to search for additional cadherin interacting partners. We also highlight some of the challenges in cadherin research, namely, the characterization of a cadherin connectome in biochemical and structural terms, as well as the elucidation of molecular mechanisms underlying the functional diversity of non-classical cadherins in vivo.
Keywords: adhesion, neuronal connectivity, hearing, mechanotransduction, cancer, brain development
Cadherin Superfamily
The classical cadherins were discovered as glycoproteins that mediate calcium-dependent cell-cell adhesion in early vertebrate embryo development and epithelial tissues [1–6]. Subsequent identification and characterization of multiple members of the superfamily revealed function beyond adhesion, including roles in signaling [7–10], mechanotransduction [11,12], and brain morphogenesis and wiring [13–15]. Cadherins, referred to here as all members of the superfamily (including protocadherins [16–17] and others), have also been found in multiple species and categorized in several subfamilies across phyla [18–21].
Characterization of the superfamily has been difficult due to the family’s many members and their functional diversity. The classical cadherins involved in cell-cell adhesion have been studied in depth and molecular mechanisms underlying their biological function have been probed extensively in tetrapods [22]. However, less is known about the function of non-classical members and about members in other species [13,15,23–27].
Here we start with a brief, structurally-based introduction to the mammalian superfamily and its classification. We then focus on new molecular mechanisms involved in interactions among cadherin superfamily members in tetrapods and fish. We discuss the emergent view that cadherins might function as heterotypic receptor complexes (formed by more than one member of the superfamily), and on how classification of the family, sequence similarities, and structural relationships may help to identify new interactions among family members.
Cadherin Architecture and Structural Diversity
The hallmark of cadherin proteins is the cadherin extracellular repeat (EC) [28–30] (Figure 1), with cadherins defined as having at least two tandem EC repeats in their extracellular domain. In addition, cadherins usually have a transmembrane and a C-terminal cytoplasmic domain (Figure 1c). Each EC repeat consists of approximately 100 amino acids sharing a common fold, but not identical sequence, with seven β strands in a “Greek-key” motif (Figure 1a). The repeats arrange in series and linker regions feature multiple highly conserved amino acids that coordinate calcium ions in three calcium- binding sites (Figure 1b). The sequence motifs involved in calcium binding have enabled the identification of over 100 superfamily members in humans, and more throughout multiple species [31,27].
Figure 1.
Architecture and structural diversity of cadherins. (a) Representative topology diagram of two cadherin repeats (Cdh23 EC1+2). A typical EC repeat features seven β-strands labeled A to G. Three calcium ions (sites 1–3) are located at the linker region between repeats (green spheres). The ribbon diagram on the right depicts the 3D structure of the two repeats. (b) Detail of calcium binding sites at linker region between repeats. Protein side chains and backbone atoms are in stick representation for amino acids indicated within a calcium binding motif. (c) Arrangement of EC repeats for different family members. Classical and desmosomal cadherins feature a cleavable prodomain, five EC repeats, and variable cytoplasmic domains that bind catenins. Clustered protocadherins sport six EC repeats and variable cytoplasmic domains. Non-clustered protocadherins feature from two up to 34 EC repeats, and some of them have variable non-cadherin extracellular domains. (d)&(e) Illustration of cis vs trans interactions for classical cadherins mediating contact between two cells. Trans interactions facilitate the formation of cis complexes [22].
Classical cadherin proteins involved in calcium-dependent cell-cell adhesion in mammals, such as CDH1 and CDH2 (E- and N-cadherin; human nomenclature used unless otherwise noted), have been the most studied and serve as the archetypical members of the superfamily [32]. These cadherins have five extracellular EC repeats labeled EC1 to EC5, starting from the repeat most distal to the membrane at the N-terminus of the protein (Figure 1c). Cleavage of a N-terminal “prodomain” is required to enable their adhesive function [33], which is achieved by trans interactions, generally homotypic, between extracellular cadherin domains protruding from adjacent cells [34,35] (Figure 1e). Classical cadherins may also interact in parallel with partner molecules from the same cell through cis interactions [36–38], with trans bond formation preceding cis complex formation [22] (Figure 1d). The cytoplasmic domain of classical cadherins binds to catenins (p120 and β), which in turn regulate stability of the extracellular cadherin bond, endocytosis, and interaction between cadherins and the cytoskeleton through α catenin and other proteins [39–44].
Similarly, other cadherins are involved in trans and cis interactions, and use their cytoplasmic domain to bind to regulatory proteins (see Box 1). However, non-classical cadherins display a more diverse set of cytoplasmic domains, and their extracellular regions have varying number of EC repeats (Table 1). For instance, CDH13 (T-cadherin) with five EC repeats, is unique as it lacks transmembrane and cytoplasmic domains and is anchored to the membrane through a glycophosphatidylinositol (GPI) moiety [45,46]. The non-classical cadherins CDH16 and CDH17 have seven EC repeats and very short cytoplasmic domains [47–49]. On the protocadherin side, three gene clusters (α, β, and γ) code for a large number of proteins (a total of ~60 in most mammalian species) each with six EC repeats and a single pass transmembrane domain. The α and γ clusters have a variable cytoplasmic subdomain concatenated to a constant cytoplasmic region, whereas PCDHβs lack the latter [16,17,24,50] (Figure I). Other “non-clustered” protocadherins have signature sequences in their cytoplasmic domains [51–54] or feature very long extracellular domains containing up to 34 EC repeats (Table 1 [54]). Overall, members of the superfamily have distinct and varied structural features that are relevant for their sometimes poorly understood function. These features also segregate members in different subfamilies, hinting that classification of the family according to sequence and structure may provide insights into evolutionary and functional relationships. However, a consistent classification has not been straightforward.
BOX 1. Family members and signaling.
Since the discovery of interactions between classical cadherins and the catenins involved in gene transcription, it has been clear that cadherins may function in both cellular adhesion and intracellular signaling. For instance, while the classical cadherin extracellular domain provides an adhesive bond and acts as a calcium sensor, differential cadherin affinities in homotypic binding mediated by EC1 modulate GTPase signaling [112]. On the clustered protocadherin side, the α and γ subfamilies have been implicated in neuronal survival by signaling through binding of their cytoplasmic domains to kinases [17,101] (Figure I).
Other “non-clustered” protocadherins have cytoplasmic signature sequences that determine cytoplasmic signaling partners (CM1, CM2, and CM3 sequences for δ1 protocadherins and CM1 and CM2 for δ2 protocadherins [51–54]). δ1 protocadherins interact with protein phosphatase-1α (PP1α), the histone-regulating TATA-binding protein-associated factor-1 TAF1 (Pcdh7), or β-catenin (PCDH11Y). Similarly, the δ2 protocadherins interact with the serine-threonine kinase TAO2β (pcdh8), the actin regulatory complex Nap1/WAVE1 (pcdh10), and mouse disabled-1 mDab1 (pcdh18; [54] and references therein; Figure I). In addition, interaction with the WAVE regulatory complex (WRC) is mediated by WRC interacting sequences (WIRS) present in the α and δ2 subfamilies [113]. Overall, these cytoplasmic partners highlight the role of protocadherins in intracellular signaling.
Other family members, often featuring long extracellular domains and loosely grouped into the protocadherin ε subfamily (Table 1 [54]) have been implicated in signaling processes that involve interaction of their cytoplasmic domains with diverse intracellular partners [114]. Among them, dachsous 1 (Dchs1) and Fat4 play a role in brain morphogenesis and planar cell polarity in kidneys and inner ear ([114] and references therein). Fat4 interacts with MUPP1 and LIX/LIX1L. Fat1, involved in CNS and kidney development, binds to β-catenin, atrophins, Ena/VASP, HOMER, and Scribble ([114] and references therein). Fat3 has been implicated in neuronal morphology [115], but less is known about its cytoplasmic partners. Celsrs are important in planar cell polarity and neuronal morphology, but again little is known about their intracellular partners [116].
Two members of the ε subfamily stand out with rather unique functional features. PCDH15 and CDH23 are important for sensory perception [12] (hearing, balance, and sight, see main text). In hair cells of the inner ear, these cadherins participate directly in mechanotransduction by linking adjacent stereocilia from the same cell. In addition, their multiple intracellular partners (sans, whirlin, harmonin, and myosin VIIa) have been proposed to form a signaling protein network that is dysfunctional in Usher syndrome, a genetic disorder that causes deafness and blindness [117,118] (Figure I).
Despite the many cases in which cytoplasmic binding partners of cadherins have been identified, it still unclear how extracellular and intracellular domains work together to integrate and trigger corresponding signaling cascades.
Table 1.
Representative members of the cadherin superfamily in humans. Members of the Homo sapiens cadherin superfamily are grouped in families and subfamilies according to multiple criteria (following [13]). Families defined through EC1-based alignments are also shown [20], with classification outliers highlighted in red.
| Superfamily | Family | Subfamily | Name | Repeats | Peculiarities |
|---|---|---|---|---|---|
|
| |||||
| Cadherin | C-1 | Type-I | CDH1 (E) | 5 | Pro-domain |
| CDH2 (N) | |||||
| CDH3 (P) | |||||
| CDH4 (R) | |||||
| CDH15 (M) | |||||
|
| |||||
| Type-II | CDH5 (VE) | 5 | Pro-domain | ||
| CDH6 (K) | |||||
| CDH7 | |||||
| CDH8 | |||||
| CDH9 (T1) | |||||
| CDH10 (T2) | |||||
| CDH11 (OB) | |||||
| CDH12 (N2) | |||||
| CDH18 | |||||
| CDH19 | |||||
| CDH20 | |||||
| CDH22 | |||||
| CDH24 | |||||
|
| |||||
| Desmosomal | DSC1 | 5 | Pro-domain | ||
| DSC2 | |||||
| DSC3 | |||||
| DSG1 | |||||
| DSG2 | |||||
| DSG3 | |||||
| DSG4 | |||||
|
| |||||
| 7D-family | CDH16 (Ksp) | 7 | |||
| CDH17 (Li) | |||||
|
| |||||
| Solitary | CDH13 (T) | 5 | Pro-domain/Lacks TM and cyto-domain | ||
| CDH26 | 5 | Pro-domain | |||
|
| |||||
| C-2 | Flamingo/CELSR | CELSR1-3 | 9 | Pro-domain/7 TM helices | |
|
| |||||
| Cadherin-related | Cr-1a | Clustered protocadherins | PCDHα1-13; αc1,2 | 6 | α and γ subfamily members share a constant domain each in the cytoplasmic region |
| PCDHβ1-16 | |||||
| PCDHγa1-12, γb1-7, γc3-5 | |||||
|
| |||||
| PCDH12 (VE2) | 6 | ||||
| PCDH20 | 7 | ||||
|
| |||||
| δ1 | PCDH1 (AXPC) | 7 | |||
| PCDH7 (BH,NF) | 7 | aa insertion | |||
| PCDH9 | 7 | ||||
| PCDH11X | 7 | ||||
| PCDH11Y | 7 | ||||
|
| |||||
| δ2 | PCDH10 (OL) | 6 | aa insertion | ||
| PCDH17 | 6 | aa insertion | |||
| PCDH18 | 6 | ||||
| PCDH19 | 6 | ||||
| PCDH8 (arcadlin, PAPC) | 6 | aa insertion | |||
|
| |||||
| Cr-1b | ε | RET | 4* | ||
| DCHS1 | 27 | ||||
| FAT4 | 34 | ||||
|
|
|
||||
| Cr-2 | CDHR1 (PCDH21) | 6 | |||
| CDHR2 (PCDH24) | 9 | ||||
| CDH23 | 27 | ||||
| CDHR5 (μPCDH) | 4 | ||||
|
|
|
||||
| Cr-3 | CLSTN1-3 | 2 | |||
| PCDH15 | 11 | ||||
| CDHR3 (CDH28) | 6 | ||||
| CDHR4 (CDH29) | 6 | ||||
| FAT1 | 34 | ||||
| FAT2 | 34 | ||||
| FAT3 | 34 | ||||
|
|
|
||||
| Solitary | DCHS2 | ? | |||
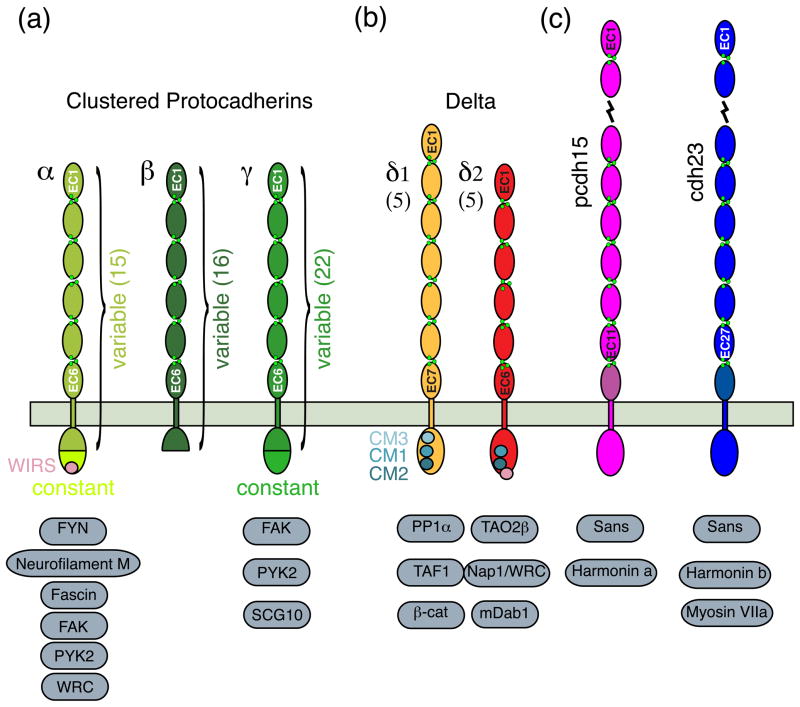
Figure I.
Domain organization and cytoplasmic partners for protocadherins involved in signaling. (a) Clustered protocadherins are divided in three subfamilies: α, β, and γ. Distinct constant cytoplasmic regions are present for all members of the α and γ subfamilies. Cytoplasmic binding partners are listed. Number of members for each family in the human genome is indicated in parenthesis. (b) The non-clustered δ protocadherins feature seven and six EC repeats for subfamilies δ1 and δ2, respectively. Cytoplasmic domains have distinct sequence motifs (CM1, CM2 and CM3 for δ1; CM1 and CM2 for δ2; WIRS for α and δ2). (c) Pcdh15 and Cdh23 have unusually long extracellular domains (11 and 27 EC repeats respectively). The two proteins and their cytoplasmic binding partners are involved in inherited deafness and blindness (Usher syndrome).
Cytoplasmic-based Classification of Cadherins
Initial attempts to classify the cadherin family used functional criteria and cytoplasmic domain sequences, rather than full-length protein sequences. This approach permitted easy identification of classical cadherins in different species having similar function and cytoplasmic domains, but varied extracellular domains. For instance, Drosophila E-cadherin (DE-cadherin [31,55]) is essential for epithelial adherens junction formation and binds Drosophila catenins, just as CDH1 functions in cell- cell adhesion and binds to mammalian catenins. However, DE-cadherin’s extracellular domain has seven EC repeats (instead of five ECs in CDH1), as well as an epidermal-growth-factor-like domain, a laminin globular domain, and a primitive proteolytic site, all three domains absent in CDH1 and in all mammalian classical cadherins [19]. Thus classification according to function and properties of the cytoplasmic domain, which does place DE-cadherin and CDH1 together in the classical subfamily, emerged as a good strategy to sort the family.
In this scheme, cadherins are pragmatically segregated into three groups: classical cadherins (able to bind catenins [19,56]), desmosomal cadherins, and protocadherins (PCDHs), which have a variable number of EC repeats (other than five) and cytoplasmic domains distinct from those of the first two subfamilies (Figure 1c).
As new members of the family in tetrapods and other species were discovered, new subfamilies, some with a single member, were incorporated and classified [13] (Table 1). In this cyto-centric extended classification, the classical subfamilies in the human genome are divided into type I and type II, the desmosomal cadherins comprise the desmocolin and desmoglein subfamilies, and the clustered protocadherins are readily separated into α, β, and γ subfamilies (Box 2 and Figure IIa). Human non-clustered protocadherins are less clearly separated in subfamilies unless other “ad-hoc” criteria are taken into account, such as the length of their cytoplasmic and extracellular domains (7D subfamily) or the specific cytoplasmic sequence signatures that define subfamilies δ1 and δ2. Finally, unclassified members are loosely grouped into the ε subfamily. Although classification according to features of cytoplasmic sequences is less effective for non-classical cadherins (Figure II), the relevance of most cadherin cytoplasmic domains in signaling and disease further validated this approach [13] (see Box 1). Such classification likely provides insights into common signaling pathways used by members within one species and perhaps across different species, but it is unclear whether it reflects accurate evolutionary relationships as well as trans and cis connectivity among cadherins.
BOX 2. Identifying New Cadherin Complexes.
To predict interactions among cadherins the EC1-3 based classification is used (Table 1 and Figure IIb). Specific rules of engagement can be defined based on our current knowledge of cadherin interactions. For instance, homotypic and heterotypic trans interactions have been reported for members within subtrees defined by type I, type II, and desmosomal cadherins, but not across these subtrees. Similarly, FAT4 groups with DCHS1 and DCHS2, and trans heterotypic interactions between Fat4 and Dchs1 have been reported. Therefore, members of a given subtree in which there is at least one confirmed trans homotypic or heterotypic interaction, may similarly engage in heterotypic trans interactions with some of the other members of the same subtree. This “subtree identity” criterion is valid for the cases mentioned above, and makes specific predictions (Figure IIc).
As an example, given that zebrafish Pcdh19 mediates homotypic trans adhesive interactions (and assuming that this is valid for mammals), the subtree criterion used with the EC1-3 based alignment predicts potential heterotypic trans interactions among PCDH10, PCDH17, PCDHαc2, PCDHγc4, and PCDHγc5 (members of the PCDH19 subtree). Similarly, PCDH8, PCDH12, and PCDH18, as well as all δ1 protocadherins would form two groups with potential for trans homotypic and heterotypic interaction (Figure IIc). Most members of these groups do not have long N-termini, required for a handshake interaction, and lack the tryptophans required for β–strand classical complexes, suggesting an X-dimer or a novel type of interaction (except for PCDH12 and the δ1 PCDH20 that may form handshake-like complexes).
The subtree identity criterion is not applicable to clustered protocadherins, which do not seem to mediate adhesive trans interactions, and even weak trans interactions have been reported to be strictly homotypic (involving a single branch within the subtree). Similarly, CELSRs do not engage in heterotypic trans interactions.
Another plausible criterion arises when considering interactions among members of different subtrees. For instance CDH23, which groups with CDHR1 and CDHR2 (Cr-2), engages in trans heterotypic interactions with PCDH15, which groups along with CDHR3, CDHR4, CDHR5, CDH16, CDH17, FAT1, FAT2, and FAT3 (Cr-3; Figure IId). We define these subtrees as “siblings” and postulate that their members have the potential to engage in heterotypic contact. Consistently, CDHR2 and CDHR5, showing overlapping expression in the enterocyte brush border, are postulated to form a complex [119]. These two proteins share structural features important for handshake-like interactions (long N-termini, a putative CDH23-like calcium-binding site for CDHR2, and a putative PCDH15-like disulfide bond for CDHR5). Thus, this subset of cadherins may engage in handshake- like interactions. Similarly, interactions between CDH17 (Cr-3) and CDH1 (Type I) [120,121] define another set of potential interactions (Figures 3i and IId).
A refinement of these criteria may involve specific sequence motifs that facilitate known cadherin interactions. For instance, the PCDH15 “EVRIVVR” motif involved in the handshake with CDH23 is found in mouse Fat4 at the same location (“EVRVLVR”) [79]. Similarly, the CDH23 “KVNIQV” motif with interfacial residues is identical in EC1 of mouse Fat3. Thus, Fat4 and Fat3 may engage in a handshake-like interaction, and define another set of sibling subtrees.
Additional sibling subtrees can be defined when extending this criterion to cis heterotypic contacts. Pcdh19 from zebrafish and Pcdh8 from mouse form a cis complex with CDH2, defining two sets of sibling subtrees (δ2 and type I; δ2′ and type I) and thereby multiple potential complexes (Figure IIe). Similarly, the α, β and γ clustered protocadherins seem to engage promiscuously in cis heterotypic complexes. The underlying molecular mechanisms and specificity (if any) for these interactions remain unknown.
Figure II.
Sequence relationships among Homo sapiens cadherin superfamily members. (a) The sequences of all cytoplasmic domains of human cadherins aligned using MUSCLE. Tree generated using Clustal and TreeDyn. (b) Classification using sequences of repeats EC1 to EC3 aligned as in (a). Calsyntenins and RET excluded from these analyses. Non-clustered protocadherins are clearly segregated in subgroups in the EC1-EC3 based classification (Cr-2, Cr-3, δ1, δ2, and δ2′). (c) Cadherins with potential for homotypic and heterotypic trans interactions within a subtree are highlighted in the human EC1-3 tree. (d,e) Cadherins with potential for trans and cis heterotypic interactions across sibling subtrees are highlighted in (d) and (e), respectively. Members of the clustered protocadherins form cis heterotypic complexes within their subtrees as well.
EC1-based Classification of Cadherins
In parallel to cyto-centric efforts to classify the cadherin family, an alternative approach was developed based on the functional relevance of repeat EC1 [18,57]. Early research on classical cadherins showed that trans interactions were calcium-dependent and strictly homotypic, i.e, members of the subfamily interact only with identical types. Using chimeric cadherins it was shown that the EC1 repeat (most distal from the membrane) is responsible for binding specificity [35]. Over the past two decades structural [58] and biophysical [59,60] studies have further demonstrated that trans interaction of mammalian classical cadherins is mediated by the N-terminal extracellular EC1 repeat [58,61,62]. These interactions involve the exchange of secondary structure elements (short segments of β-strand A in EC1) along with exchanged-docking of one (type I) or two (type II) tryptophan side chains at the very N-terminus of the mature protein (Figure 2). While there is some variation in the surfaces involved in homotypic interactions among type I and type II members, all vertebrate classical cadherins studied so far use the same basic mechanism [22]. Moreover, extracellular domains of classical cadherins can drive adhesion by themselves (without cytoplasmic domains [61]).
Figure 2.
Strand-swapped and X-dimer trans interactions of classical cadherins [22]. (a) Structure of entire extracellular domain (EC1-5) Cdh2 (N-cadherin) engaged in a strand-swapped dimer (EC1-to-EC1 contact). Inserts show details of the exchange of tryptophans at position 2 (W2). Availability of W2 for binding is modulated by calcium (green spheres). (b) Cdh13 (T-cadherin) EC1+2 structure shown for two protomers in a X-dimer conformation. One protomer is shown in opaque surface representation, while the other is in cartoon and transparent surface representations. Inset shows details of the interaction. (c) Classical cadherins engage in a similar interaction that is thought to lead to the strand-swapped dimer (right) and facilitate cis interactions [22].
The functional relevance of the extracellular domain and the involvement of EC1 in signaling and related binding mechanisms suggested that the whole family could be classified by aligning the sequences of EC1 repeats alone [57]. This provided a way to avoid the difficulties in comparing family members with vastly different number of EC repeats or highly divergent cytoplasmic domain sequences. Remarkably, EC1-based alignments effectively segregate vertebrate cadherins in groups that are similar to those obtained using “ad-hoc” criteria along with cytoplasmic-based alignments [13,18,20] (Table 1). In this scheme, human cadherins naturally separate in groups that include type I, type II, desmosomal, and 7D cadherins within the newly defined C-1 family of the cadherin major branch. Similarly, the α, β, and γ groups of clustered protocadherins separate from δ1 and δ2 protocadherins in the Cr-1a branch of the cadherin related major branch.
However, the EC1-based classification differs in some aspects to that based on cytoplasmic properties. In the human superfamily, clustered protocadherins PCDHαc1, αc2, γc3, γc4, and γc5 feature distinct extracellular EC1 repeats that group them away from their corresponding α and β branches [24,50,63] (defined by cytoplasmic-based alignments and gene structure). The EC1-based alignment also segregates PCDH10 away from the δ2 protocadherins [20]. In addition, some cytoplasmic-based alignments that use a subset of cadherin members [20,54] suggest that PCDH15 groups with CDHR1 (PCDH21), CDH23 with CDHR2 (PCDH24), and PCDH12 with CDHR5 (μPCDH). Cytoplasmic-based alignments using the whole set of human cadherins are less conclusive about these subgroups, while the EC1-based alignment suggests the existence of three well-defined groups: the first one formed by CDHR1, CDHR2, and CDH23 (Cr-2 subfamily) and two additional single-member groups including CDHR5 and PCDH15 each [20]. Members inconsistently classified are highlighted in red in Table 1.
Importantly, new subfamilies can be identified in the EC1-based classification when considering non- mammalian sequences of cadherins [20,21]. The C-2 family, including the CELSRs in mammals, also encompasses type III cadherins like Drosophila DN-cadherin, chicken Hz, and zebrafish and Xenopus cHz-like cadherins. Type IV cadherins like Drosophila shotgun (DE-cadherin) and cricket (Gryllus bimaculatus) Gb1-cadherin also belong to the C-2 family. Type III cadherins with 13 to 17 EC repeats and type IV cadherins with seven EC repeats are not present in mammals.
The EC1-based scheme is not perfect. Proteins such as CDH1 and DE-cadherin are grouped in different subfamilies although they perform similar tasks in different species. Therefore, function may not be segregated with subfamily in this classification, which hints at more general difficulties when establishing evolutionary and sequence-structure-function relationships within a family of proteins across phyla [64]. Clearly, even close homologues may have different cellular functions and biochemical properties. Conversely, proteins that differ widely in sequence and structure may carry out similar functions in different species. Thus, evolutionary studies and sequence analyses of cadherins must be performed in the context of functional and biochemical data for members of protein families across multiple species [64].
Another shortfall of the EC1-based classification is that some cadherins use EC repeats other than EC1 to perform their function in adhesion and signaling. This could be particularly relevant for long cadherins with unusual calcium binding sites that may adopt globular shapes [65] thereby using binding mechanisms that resemble those used by the Drosophila DSCAMs [66].
Towards Cadherin Connectomics
The EC1-based classification that emerged from work on classical cadherins remains attractive despite the pitfalls described above. It provides a simple way to deal with diverse multi-domain cadherins, while at the same time providing insights into EC1-based interaction mechanisms for cadherins. New binding mechanisms in classical and non-classical members of the family, however, suggest that the EC1-based homotypic interaction paradigm may not be applicable for all cadherins, and that the EC1-based classification should be updated. We therefore continue reviewing these new binding mechanisms observed for classical cadherins, atypical cadherins CDH23 and PCDH15, and clustered protocadherins, and suggest a new grouping based on the sequences of the first three EC repeats (Figure IIb), as well as new ways of finding potential interacting partners (Box 2; Figure IIc,d and e).
New Homotypic Trans Interaction Mechanisms Involve EC1 and EC2 Repeats
The involvement of multiple EC repeats in cell-cell adhesion mediated by classical cadherins has been highly debated over the past decade [67]. Biophysical and cell-based assays have suggested that classical cadherins could interdigitate their extracellular domains. In this arrangement, repeats EC2 to EC5 would also form part of the adhesive trans interface. However, single molecule FRET experiments [59,60] and cryo-electron tomography of desmosomes [68,69] indicate that interactions are tip-to-tip (EC1 to EC1), and reminiscent of the arrangements seen in crystals of entire extracellular domains of classical cadherins [61]. These studies further validate the approach of using EC1 sequences to classify the superfamily.
Interestingly, CDH13 (T-cadherin), a non-classical member of the cadherin family that lacks the EC1 features required for the trans EC1-to-EC1 strand-swapping interaction ([22], Figure 2a), does mediate trans homotypic cell-cell adhesion [45,46]. This unusual cadherin does not have a transmembrane or cytoplasmic domain, but it is linked to the plasma membrane through a GPI anchor. Moreover, crystallographic structures of CDH13 EC1 and EC2 repeats revealed a new mode of trans homotypic interaction that involves both its EC1 and EC2 repeats in a so-called “X-dimer” conformation [70] (Figure 2b). A series of in vitro biochemical and cell-based assays confirmed the X-dimer interface for CDH13, which was found to mediate robust aggregation of CHO-cells and to mediate inhibition of neurite outgrowth [70].
The X-dimer arrangement had been identified in previous X-ray structures of a mutant CDH1 [71], but had been regarded as either a candidate for mediating cis interactions among cadherins [71,38,72] or a crystal packing artifact (see discussion in [73]). Additional structural analyses of mutated classical cadherins and CDH13 [70,73] suggest that the X-dimer is a transient state used by all classical cadherins. This state, which is consistent with a second bonded conformation observed in single- molecule FRET experiments [60] and involves repeat EC2, may facilitate the subsequent exchange of β-strands seen in the EC1-to-EC1 strand swapped dimers [73] (Figure 2c). Thus, EC2 becomes relevant for homotypic trans interactions, and should be considered in EC-based classifications of the superfamily.
A New Trans Heterotypic Complex also Involves EC1 and EC2 Repeats
Just as the involvement of multiple EC repeats in cadherin trans interactions has been disputed, the existence of physiologically relevant heterotypic trans complexes of classical cadherins has been equally controversial. While the debate continues for classical cadherins, recent work has unequivocally shown that two non-classical cadherins, mouse Cdh23 and Pcdh15, do form a functional heterotypic complex that also involves both repeats EC1 and EC2.
The non-classical Cdh23 and Pcdh15 mouse proteins form hair-cell tip links [74–76], fine filaments essential for inner-ear mechanotransduction [12]. Tip links assemble and regenerate in a calcium- dependent manner, are constantly under tension, and connect adjacent stereocilia of the same cell (Figure 3a). The Cdh23 and Pcdh15 proteins, with 27 and 11 EC repeats, respectively, are products of deafness genes [76], are localized to tip links by antibody labeling, and their predicted length matches the length of the tip link, suggesting a tip-to-tip interaction [75,77] (Figure 3a,b). Moreover, binding experiments show robust heterotypic interaction in vitro [75,78].
Figure 3.
Inner-ear cadherin handshake and heterotypic cadherin complexes. (a) Hair cell bundle, made of actin-filled stereocilia arranged in a staircase, moves upon mechanical stimulation thereby applying tension to tip links (black box). (b) Schematic of an heterotetrameric complex of Cdh23 and Pcdh15 forming the tip link [75]. Inset shows details of interaction, with the tips of both proteins engaged in a “handshake” interaction bond (right) [79]. (c) Ribbon diagram (left) and surface representation (right) of the handshake interaction, showing calcium ions (green spheres), two protrusions (arrow heads), and the side chain of Arginine 113. (d) Detail of handshake interaction. (e) Detail of Pcdh15 N-terminus with a disulfide bond. (f) Detail of N-terminal calcium binding site 0 in Cdh23. (g–i) Hypothetical models for heterotypic interactions: (g) clustered protocadherins heterotetramers [95], (h) classical and delta-cadherin complexes [15], and (i) Cdh17 and Cdh1 [120,121].
The structures of the heterotypic complex formed by the interacting tips of mouse Cdh23 and Pcdh15 reveal that these proteins do not use any of the binding mechanisms observed for classical cadherins [78–80]. Instead, the EC1+2 protomers of each protein engage in an antiparallel extended “handshake”, with both repeats contributing to the interface [79] (Figure 3c). Consistent with this structure, an arginine residue mutated to glycine in individuals with inherited deafness [81] is located in the handshake interface (Figure 3d). The structure predicts that the arginine-to-glycine mutation would disrupt molecular interactions thereby impairing binding, and this was confirmed by size exclusion chromatography and calorimetry experiments [79], as well as by prior binding assays [75] and ex vivo functional tests [77].
The handshake interface is possible due to special structural features of both Cdh23 and Pcdh15 protomers. In both cases the N-terminal strand of EC1 is unlike that of classical cadherins: it extends towards the top of the protomer where it is tucked and secured by a novel calcium binding site in Cdh23 [78,79] and by a disulfide bond in Pcdh15 [79] (Figure 3c,e,f). Both EC1 protomers present bulges that fit in the narrower inter-repeat linker region of the adjacent molecules, thereby facilitating the overlap. Interestingly, some mouse and human PCDH15 isoforms have modified N-terminal sequences, suggesting that PCDH15 across species may form new types of complexes with itself or with other members of the cadherin family. Moreover, recent experiments suggest that immature tip links might be transiently made of mouse Pcdh15 tetramers, without any Cdh23 molecules [82]. The multiple isoforms of mouse Pcdh15 [74] may provide a pseudo-heterotypic antiparallel bond responsible for the all-Pcdh15 immature tip links observed in [82].
The consequences of preventing the Cdh23 – Pcdh15 handshake interaction in vivo were determined using the Noddy mouse model [83]. Noddy mice carry an isoleucine to asparagine mutation at position 108, also located in the handshake interface. These mice completely lack inner-ear function. In vitro experiments show that this mutation impairs binding of Pcdh15 EC1+2 I108N to Cdh23 EC1+2, without preventing proper folding of the protomer. In vivo localization is also unaffected, indicating that impaired binding between the two proteins solely causes that phenotype. Thus the heterotypic handshake interaction, extensively validated in vitro, ex vivo, and in vivo, is essential for inner-ear mechanotransduction [79,83].
The structure of the heterotypic cadherin bond formed by Cdh23 and Pcdh15 indicates that EC2 repeats can also be involved in trans binding, as in the homotypic classical X-dimer interface. Thus, the EC1-based classification should be extended to include other repeats. The new classification could be used to search for similar heterotypic binding pairs, as it is now clear that other cadherins (like the clustered protocadherins) form heterotypic complexes [15].
Protocadherin Complexes Involve Multiple Family Members and EC Repeats
The clustered protocadherins form the largest cadherin subfamily with 53 members in humans [16,17,24,50]. Many members have been studied individually, or collectively by antibody targeting of the constant domains in the cytoplasmic domains of α or γ protocadherins. Clustered protocadherins tested so far do not share the robust adhesive properties of classical cadherins, but rather cluster together in cis to form signaling receptors that mediate neuronal recognition [14,17,84] and survival [85–88] through presumably weak trans interactions.
Several issues have delayed the characterization of interactions among clustered protocadherins. First, bead-based binding experiments fail to show interaction among family members [15]. In addition, cell-based assays did not show strong calcium-dependent cell aggregation activity for multiple members of the subfamily [16,89–94]. Lastly, interpretation of these results might have been confounded by endogenous expression of γ protocadherins [95].
To overcome these difficulties and to probe interactions among mouse Pcdhγ complexes, non- adherent K562 cells, which lack endogenous expression of classical cadherins and γ protocadherins have been used [95–97]. These experiments revealed homotypic trans interaction for Pcdhγa3, b2, and c3 [95]. The interaction mediated by Pcdhγa3 was found to be only partially calcium-dependent and required repeat EC1. Additional experiments with Pcdhγa10, a12, b1, and b6 confirmed strict homotypic trans interactions. Remarkably, chimeric constructs show that the specificity of these interactions is governed by repeats EC2 and EC3 [95], not EC1 (Figure 3g). This is consistent with positively selected codon positions found mostly in EC2 and EC3 repeats when analyzing EC1-3 sequences [24].
Immunoprecipitation experiments also suggest that members of the Pcdhγ subfamily form tetrameric complexes in cis [95] (Figure 3g). The number of cis heterotypic tetramers that can be formed and made available for trans homotypic interaction is large enough to provide a rich repertoire of unique receptors that can form the basis for a neuronal recognition code [84,95]. Interestingly, recent experiments suggests that members of the α, β, and γ subfamilies interact (directly or indirectly) with each other, although not necessarily to mediate adhesion [98–102]. Thus, the cadherin code for neuronal recognition could be greatly extended.
Proteomic analysis using mouse proteins also suggest that members of the α and γ subfamilies interact with the classical cadherin Cdh2 [98], but likely in an indirect way [99]. Similarly, members of the γ subfamily may interact with other members of the superfamily, like Cdh4 and Pcdh17 [98], while members of the δ1, δ2, and ε subfamilies of non-clustered protocadherins may also form heterotypic complexes [103–106] (Figure 3g,h).
For instance, Cdh2 and Pcdh19 functionally cooperate with each other in zebrafish brain development [105], and their physical interaction has been the best characterized so far [106]. While zebrafish Pcdh19 does not mediate adhesion on its own, and does not interact in trans with zebrafish Cdh2, it is responsible for the trans adhesive interaction of the heterotypic Cdh2-Pcdh19 complex. Cdh2 seems to enable the adhesion driven by Pcdh19, suggesting a paradigm-shifting view in which classical cadherins, at least in zebrafish, act as regulators of δ2 protocadherin-mediated adhesion [15,106] (Figure 3h).
Overall, these results show an apparent heterotypic promiscuity among non-classical cadherins interacting in cis (particularly for the clustered protocadherins), while trans interactions that involve repeats EC1, EC2, and EC3 are strictly homotypic, at least for the γ subfamily. It is apparent that involvement of some cadherins in signaling and cell recognition requires a transient and perhaps weak interaction, while in other cases, a strong bond may serve for concomitant signaling and adhesive functions. Regardless of their role in signaling or in more stable adhesive contacts, most of the protocadherin interactions described so far involve repeats that go beyond repeat EC1, and most have not been complemented with structures. Moreover, there is a lack of a systematic exploration of the possible complexes that can be formed by superfamily members.
Concluding Remarks
The research summarized above has revealed functional and structural diversity among cadherins. Overall, these results highlight the need to abandon the narrow view of homotypic adhesive interactions mediated by EC1 for all cadherins and explore function beyond the classical paradigm (see Outstanding Questions). Many cadherins use alternate binding mechanisms that involve both homotypic and heterotypic interactions mediated by multiple EC repeats. These interactions may play a role in adhesion, signaling, or both. Together these results also suggest that the EC1-based classification should be extended to incorporate at least EC2 and EC3 repeats. Evolutionary relationships are unlikely to be captured by such classification, as they require more comprehensive analyses of the full-length protein sequences and of the biochemical properties of both intra and extracellular domains. However, we propose that EC-based classifications may serve a different purpose: to identify families and subfamilies, within a species, that share distinct binding mechanisms, thereby providing roadmaps to probe and build species-specific cadherin connectomes. For instance, rearrangement of the human cadherin superfamily using sequences covering EC1, EC2, and EC3 repeats reveals interesting relationships and general criteria that can be used to predict interaction candidates (Box 2). As a test, we propose two strong complex candidates for heterotypic interactions (FAT4/FAT3 and CDHR2/CDHR5), among many other possibilities that need to be systematically explored, as recently done for immunoglobulin and LRR proteins in Drosophila [107].
Ultimately, understanding how cadherins interact with each other will serve to design molecular handles that can control their function in vivo, perhaps to modify development and morphogenesis [108], or alter neuronal connectivity in a controlled fashion [109]. This may provide essential tools to probe the functional connectome of the brain [110].
Outstanding Questions BOX
The molecular mechanisms underlying function of cadherins have been elucidated in great detail for some members of the superfamily. However, multiple challenges need to be addressed to answer outstanding questions and to probe the function of many more cadherins for which data are scarce. For instance, ex vivo and in vitro assays to test cadherin interactions often give confounding results and a decade of trials suggests that five conditions must be satisfied in ex vivo cell-based binding assays: 1) experiments should show that endogenous expression of all cadherins is limited and is not interfering with the interactions being probed; 2) induced expression of cadherins and other proteins upon transfection must be checked as well; 3) control of surface expression by endocytosis and cleavage through metalloproteases must be considered when interpreting ex vivo results; 4) interactions with other non-cadherin proteins [91,111], not covered in this review, should be considered and probed; 5) in vivo and ex vivo interactions must be recapitulated in vitro to identify all components of the complexes and differentiate direct vs. indirect interactions. Similarly, in vitro bead- based binding assays should take into account the lack of cytoplasmic domains and cis partners that might be required to establish an interaction. In addition, the role of mechanical forces as modulators of binding affinity, as well as isoform diversity and glycosylation must be taken into account in the context of testing functional cadherin interactions.
Additional challenges arise because ex vivo and in vitro binding assays are often carried out with shorter recombinant versions of the wild-type proteins. This may hinder the elucidation of interaction mechanisms that involve other repeats, especially for long cadherins, which may lack an obvious EC1 repeat or which may fold in globular shapes [65]. Site-directed mutagenesis is also often used to probe molecular interaction mechanisms, but these mutations may have pleotropic effects, for example affecting folding or calcium binding affinity. Impaired calcium binding may have subtle effects on rigidity that could prevent binding, even under saturating calcium concentrations.
Addressing these issues and challenges systematically may provide a way to answer several open questions: 1) What is the exact stoichiometry and composition of cadherin complexes, especially those formed by clustered protocadherins? 2) What are the molecular mechanisms underlying cis interactions, and are there new ways to form trans complexes? 3) What determines homotypic and heterotypic specificities (or lack thereof)? 4) Are all signaling interactions among cadherins intrinsically weak? 5) How do the extracellular domains of cadherins control the binding of signaling molecules to their own cytoplasmic tails? 6) How do long cadherin extracellular domains arrange on the membrane surface?
Highlights.
Cadherin proteins and their interactions are essential for multicellular life.
Physical interaction among cadherins is more complex than originally thought.
Classification of the superfamily may guide the search for new cadherin complexes.
Acknowledgments
We thank members of the Sotomayor, Corey, and Gaudet laboratories, as well as A. Krol for valuable discussions. This work was supported by the NIH (K99/R00 DC012534-01 to MS and R01 DC02281 to DPC). DPC is an investigator of the Howard Hughes Medical Institute.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Takeichi M. Functional correlation between cell adhesive properties and some cell surface proteins. Journal of Cell Biology. 1977;75:464–474. doi: 10.1083/jcb.75.2.464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hyafil F, Babinet C, Jacob F. Cell-cell interactions in early embryogenesis: a molecular approach to the role of calcium. Cell. 1981;26:447–454. doi: 10.1016/0092-8674(81)90214-2. [DOI] [PubMed] [Google Scholar]
- 3.Damsky C, Knudsen KA, Dorio RJ, Buck CA. Manipulation of cell-cell and cell-substratum interactions in mouse mammary tumor epithelial cells using broad spectrum antisera. Journal of Cell Biology. 1981;89:173–184. doi: 10.1083/jcb.89.2.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bertolotti R, Rutishauser U, Edelman GM. A cell surface molecule involved in aggregation of embryonic liver cells. Proceedings of the National Academy of Sciences, USA. 1980;77:4831–4835. doi: 10.1073/pnas.77.8.4831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Takeichi M. Cadherins: A molecular family important in selective cell-cell adhesion. Annu Rev Biochem. 1990;59:237–252. doi: 10.1146/annurev.bi.59.070190.001321. [DOI] [PubMed] [Google Scholar]
- 6.Kemler R. Classical cadherins. Semin Cell Biol. 1992;3:149–155. doi: 10.1016/s1043-4682(10)80011-x. [DOI] [PubMed] [Google Scholar]
- 7.Suzuki ST. Structural and functional diversity of cadherin superfamily: are new members of cadherin superfamily involved in signal transduction pathway? J Cell Biochem. 1996;61:531–542. doi: 10.1002/(sici)1097-4644(19960616)61:4<531::aid-jcb6>3.0.co;2-p. [DOI] [PubMed] [Google Scholar]
- 8.Vleminckx K, Kemler R. Cadherins and tissue formation: integrating adhesion and signaling. Bioessays. 1999;21:211–220. doi: 10.1002/(SICI)1521-1878(199903)21:3<211::AID-BIES5>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- 9.Wheelock MJ, Johnson KR. Cadherin-mediated cellular signaling. Curr Opin Cell Biol. 2003;15:509–514. doi: 10.1016/s0955-0674(03)00101-7. [DOI] [PubMed] [Google Scholar]
- 10.Saito M, Tucker DK, Kohlhorst D, Niessen C, Kowalczyk AP. Classical and desmosomal cadherins at a glance. J Cell Sci. 2012;125:2547–2552. doi: 10.1242/jcs.066654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Leckband DE, le Duc Q, Wang N, de Rooij J. Mechanotransduction at cadherin mediated adhesions. Curr Opin Cell Biol. 2011;23:523–530. doi: 10.1016/j.ceb.2011.08.003. [DOI] [PubMed] [Google Scholar]
- 12.Gillespie PG, Muller U. Mechanotransduction by hair cells: Models, molecules, and mechanisms. Cell. 2009;139:33–44. doi: 10.1016/j.cell.2009.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hirano S, Takeichi M. Cadherins in brain morphogenesis and wiring. Physiol Rev. 2012;92:597–634. doi: 10.1152/physrev.00014.2011. [DOI] [PubMed] [Google Scholar]
- 14.Lefebvre JL, Kostadinov D, Chen WV, Maniatis T, Sanes JR. Protocadherins mediate dendritic self- avoidance in the mammalian nervous system. Nature. 2012;488:517–521. doi: 10.1038/nature11305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Weiner JA, Jontes JD. Protocadherins, not prototypical: a complex tale of their interactions, expression, and functions. Front Mol Neurosci. 2013;6:1–4. doi: 10.3389/fnmol.2013.00004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sano K, et al. Protocadherins: a large family of cadherin-related molecules in central nervous system. EMBO J. 1993;12:2249–2256. doi: 10.1002/j.1460-2075.1993.tb05878.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kohmura N, et al. Diversity revealed by a novel family of cadherins expressed in neurons at a synaptic complex. Neuron. 1998;20:1137–1151. doi: 10.1016/s0896-6273(00)80495-x. [DOI] [PubMed] [Google Scholar]
- 18.Nollet F, Kools P, van Roy F. Phylogenetic analysis of the cadherin superfamily allows identification of six major subfamilies besides several solitary members. Journal of Molecular Biology. 2000;299:551–572. doi: 10.1006/jmbi.2000.3777. [DOI] [PubMed] [Google Scholar]
- 19.Oda H, Takeichi M. Evolution: structural and functional diversity of cadherin at the adherens junction. J Cell Biol. 2011;193:1137–1146. doi: 10.1083/jcb.201008173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hulpiau P, van Roy F. Molecular evolution of the cadherin superfamily. The International Journal of Biochemistry and Cell Biology. 2009;41:349–369. doi: 10.1016/j.biocel.2008.09.027. [DOI] [PubMed] [Google Scholar]
- 21.Hulpiau P, Gul IS, van Roy F. New insights into the evolution of metazoan cadherins and catenins. Prog Mol Biol Transl Sci. 2013;116:71–94. doi: 10.1016/B978-0-12-394311-8.00004-2. [DOI] [PubMed] [Google Scholar]
- 22.Brasch J, Harrison OJ, Honig B, Shapiro L. Thinking outside the cell: how cadherins drive adhesion. Trends Cell Biol. 2012;22:299–310. doi: 10.1016/j.tcb.2012.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tada MN, et al. Genomic organization and transcripts of the zebrafish protocadherin genes. Gene. 2004;340:197–211. doi: 10.1016/j.gene.2004.07.014. [DOI] [PubMed] [Google Scholar]
- 24.Wu Q. Comparative genomics and diversifying selection of the clustered vertebrate protocadherin genes. Genetics. 2005;169:2179–2188. doi: 10.1534/genetics.104.037606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yu WP, et al. Elephant shark sequence reveals unique insights into the evolutionary history of vertebrate genes: A comparative analysis of the protocadherin cluster. Proceedings of the National Academy of Sciences, USA. 2008;105:3819–3824. doi: 10.1073/pnas.0800398105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jiang XJ, Li S, Ravi V, Venkatesh B, Yu WP. Identification and comparative analysis of the protocadherin cluster in a reptile, the green anole lizard. PLoS One. 2009;4:e7614. doi: 10.1371/journal.pone.0007614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nichols SA, Roberts BW, Richter DJ, Fairclough SR, King N. Origin of metazoan cadherin diversity and the antiquity of the classical cadherin/β-catenin complex. Proceedings of the National Academy of Sciences, USA. 2012;109:13046–13051. doi: 10.1073/pnas.1120685109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ringwald M, et al. The structure of cell adhesion molecule uvomorulin. insights into the molecular mechanism of Ca2+-dependent cell adhesion. EMBO J. 1987;6:3647–3653. doi: 10.1002/j.1460-2075.1987.tb02697.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Overduin M, et al. Solution structure of the epithelial cadherin domain responsible for selective cell adhesion. Science. 1995;267:386–389. doi: 10.1126/science.7824937. [DOI] [PubMed] [Google Scholar]
- 30.Shapiro L, et al. Structural basis of cell-cell adhesion by cadherins. Nature. 1995;374:327–336. doi: 10.1038/374327a0. [DOI] [PubMed] [Google Scholar]
- 31.Oda H, Uemura T, Harada Y, Iwai Y, Takeichi M. A Drosophila homolog of cadherin associated with armadillo and essential for embryonic cell-cell adhesion. Dev Biol. 1994;165:716–726. doi: 10.1006/dbio.1994.1287. [DOI] [PubMed] [Google Scholar]
- 32.van Roy F, Berx G. The cell-cell adhesion molecule E-cadherin. Cell Mol Life Sci. 2008;65:3756–3788. doi: 10.1007/s00018-008-8281-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ozawa M, Kemler R. Correct proteolytic cleavage is required for the cell adhesive function of uvomorulin. Journal of Cell Biology. 1990;111:1645–1650. doi: 10.1083/jcb.111.4.1645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nagafuchi A, Shirayoshi Y, Okazaki K, Yasuda K, Takeichi M. Transformation of cell adhesion properties by exogenously introduced E-cadherin cDNA. Nature. 1987;329:341–343. doi: 10.1038/329341a0. [DOI] [PubMed] [Google Scholar]
- 35.Nose A, Tsuji K, Takeichi M. Localization of specificity determining sites in cadherin cell adhesion molecules. Cell. 1990;61:147–155. doi: 10.1016/0092-8674(90)90222-z. [DOI] [PubMed] [Google Scholar]
- 36.Brieher WM, Yap AS, Gumbiner BM. Lateral dimerization is required for the homophilic binding activity of C-cadherin. J Cell Biol. 1996;135:487–496. doi: 10.1083/jcb.135.2.487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Takeda H, Shimoyama Y, Nagafuchi A, Hirohashi S. E-cadherin functions as a cis-dimer at the cell- cell adhesive interface in vivo. Nature Structural Biology. 1999;6:310–312. doi: 10.1038/7542. [DOI] [PubMed] [Google Scholar]
- 38.Pertz O, et al. A new crystal structure, Ca2+ dependence and mutational analysis reveal molecular details of E-cadherin homoassociation. EMBO Journal. 1999;18:1738–1747. doi: 10.1093/emboj/18.7.1738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Nagafuchi A, Takeichi M. Cell binding function of E-cadherin is regulated by the cytoplasmic domain. EMBO J. 1988;7:3679–3684. doi: 10.1002/j.1460-2075.1988.tb03249.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ozawa M, Baribault H, Kemler R. The cytoplasmic domain of the cell adhesion molecule uvomorulin associates with three independent proteins structurally related in different species. EMBO Journal. 1989;8:1711–1717. doi: 10.1002/j.1460-2075.1989.tb03563.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Abe K, Takeichi M. EPLIN mediates linkage of the cadherin catenin complex to F-actin and stabilizes the circumferential actin belt. Proceedings of the National Academy of Sciences, USA. 2008;105:13–19. doi: 10.1073/pnas.0710504105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pokutta S, Drees F, Yamada S, Nelson WJ, Weis WI. Biochemical and structural analysis of alpha- catenin in cell-cell contacts. Biochem Soc Trans. 2008;36:141–147. doi: 10.1042/BST0360141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Shapiro L, Weis WI. Structure and biochemistry of cadherins and catenins. Cold Spring Harb Perspect Biol. 2009;1:a003053. doi: 10.1101/cshperspect.a003053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hong S, Troyanovsky RBTS. Binding to F-actin guides cadherin cluster assembly, stability, and movement. J Cell Biol. 2013;201:131–143. doi: 10.1083/jcb.201211054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ranscht B, Dours-Zimmermann MT. T-cadherin, a novel cadherin cell adhesion molecule in the nervous system lacks the conserved cytoplasmic region. Neuron. 1991;7:391–402. doi: 10.1016/0896-6273(91)90291-7. [DOI] [PubMed] [Google Scholar]
- 46.Vestal DJ, Ranscht B. Glycosyl phosphatidylinositol-anchored T-cadherin mediates calcium- dependent, homophilic cell adhesion. J Cell Biol. 1992;119:451–461. doi: 10.1083/jcb.119.2.451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Thomson RB, et al. Isolation and cDNA cloning of Ksp-cadherin, a novel kidney specific member of the cadherin multigene family. J Biol Chem. 1995;270:17594–17601. doi: 10.1074/jbc.270.29.17594. [DOI] [PubMed] [Google Scholar]
- 48.Berndorff D, et al. Liver-intestine cadherin: molecular cloning and characterization of a novel Ca(2+)-dependent cell adhesion molecule expressed in liver and intestine. J Cell Biol. 1994;125:1353–1369. doi: 10.1083/jcb.125.6.1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Wendeler MW, Jung R, Himmelbauer H, Gessner R. Unique gene structure and paralogy define the 7D-cadherin family. Cell Mol Life Sci. 2006;63:1564–1573. doi: 10.1007/s00018-006-6014-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wu Q, Maniatis T. A striking organization of a large family of human neural cadherin like cell adhesion genes. Cell. 1999;97:779–790. doi: 10.1016/s0092-8674(00)80789-8. [DOI] [PubMed] [Google Scholar]
- 51.Wolverton T, Lalande M. Identification and characterization of three members of a novel subclass of protocadherins. Genomics. 2001;76:66–72. doi: 10.1006/geno.2001.6592. [DOI] [PubMed] [Google Scholar]
- 52.Vanhalst K, Kools P, Staes K, van Roy F, Redies C. delta-protocadherins: a gene family expressed differentially in the mouse brain. Cell Mol Life Sci. 2005;62:1247–1259. doi: 10.1007/s00018-005-5021-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Redies C, Vanhalst K, Roy F. delta-protocadherins: unique structures and functions. Cell Mol Life Sci. 2005;62:2840–2852. doi: 10.1007/s00018-005-5320-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Kim S, Yasuda S, Tanaka H, Yamagata K, Kim H. Non-clustered protocadherin. Cell Adh Migr. 2011;5:97–105. doi: 10.4161/cam.5.2.14374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Woods DF, Wu JW, Bryant PJ. Localization of proteins to the apico-lateral junctions of Drosophila epithelia. Dev Genet. 1997;20:111–118. doi: 10.1002/(SICI)1520-6408(1997)20:2<111::AID-DVG4>3.0.CO;2-A. [DOI] [PubMed] [Google Scholar]
- 56.Takeichi M. Morphogenetic roles of classic cadherins. Current Opinion in Cell Biology. 1995;7:619–627. doi: 10.1016/0955-0674(95)80102-2. [DOI] [PubMed] [Google Scholar]
- 57.Pouliot Y. Phylogenetic analysis of the cadherin superfamily. Bioessays. 1992;14:743–748. doi: 10.1002/bies.950141104. [DOI] [PubMed] [Google Scholar]
- 58.Boggon TJ, et al. C-cadherin ectodomain structure and implications for cell adhesion mechanisms. Science. 2002;296:1308–1313. doi: 10.1126/science.1071559. [DOI] [PubMed] [Google Scholar]
- 59.Zhang Y, Sivasankar S, Nelson WJ, Chu S. Resolving cadherin interactions and binding cooperativity at the single-molecule level. Proceedings of the National Academy of Sciences, USA. 2009;106:109–114. doi: 10.1073/pnas.0811350106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Sivasankar S, Zhang Y, Nelson WJ, Chu S. Characterizing the initial encounter complex in cadherin adhesion. Structure. 2009;17:1075–1081. doi: 10.1016/j.str.2009.06.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Harrison OJ, et al. The extracellular architecture of adherens junctions revealed by crystal structures of type I cadherins. Structure. 2011;19:244–256. doi: 10.1016/j.str.2010.11.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Patel SD, et al. Type II cadherin ectodomain structures: Implications for classical cadherin specificity. Cell. 2006;124:1255–1268. doi: 10.1016/j.cell.2005.12.046. [DOI] [PubMed] [Google Scholar]
- 63.Takei Y, et al. Two novel CNRs from the CNR gene cluster have molecular features distinct from those of CNR1 to 8. Genomics. 2001;72:321–330. doi: 10.1006/geno.2000.6468. [DOI] [PubMed] [Google Scholar]
- 64.Dickinson DJ, Weis WI, Nelson WJ. Protein evolution in cell and tissue development: going beyond sequence and transcriptional analysis. Dev Cell. 2011;21:32–34. doi: 10.1016/j.devcel.2011.06.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Jin X, et al. Crystal structures of Drosophila N-cadherin ectodomain regions reveal a widely used class of Ca2+-free interdomain linkers. Proceedings of the National Academy of Sciences, USA. 2012;109:E127–134. doi: 10.1073/pnas.1117538108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Sawaya MR, et al. A double S shape provides the structural basis for the extraordinary binding speciffcity of Dscam isoforms. Cell. 2008;134:1007–1018. doi: 10.1016/j.cell.2008.07.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Leckband D, Prakasam A. Mechanism and dynamics of cadherin adhesion. Annual Review of Biomedical Engineering. 2006;8:259–287. doi: 10.1146/annurev.bioeng.8.061505.095753. [DOI] [PubMed] [Google Scholar]
- 68.He W, Cowin P, Stokes DL. Untangling desmosomal knots with electron tomography. Science. 2003;302:109–113. doi: 10.1126/science.1086957. [DOI] [PubMed] [Google Scholar]
- 69.Al-Amoudi A, Diez DC, Betts MJ, Frangakis AS. The molecular architecture of cadherins in native epidermal desmosomes. Nature. 2007;450:832–837. doi: 10.1038/nature05994. [DOI] [PubMed] [Google Scholar]
- 70.Ciatto C, et al. T-cadherin structures reveal a novel adhesive binding mechanism. Nat Struct Mol Biol. 2010;17:339–347. doi: 10.1038/nsmb.1781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Nagar B, Overdulin M, Ikura M, Rini J. Structural basis of calcium-induced E-cadherin rigidification and dimerization. Nature. 1996;380:360–364. doi: 10.1038/380360a0. [DOI] [PubMed] [Google Scholar]
- 72.Haussinger D, et al. Proteolytic E-cadherin activation followed by solution NMR and X-ray crystallography. EMBO Journal. 2004;23:1699–1708. doi: 10.1038/sj.emboj.7600192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Harrison OJ, et al. Two-step adhesive binding by classical cadherins. NSMB. 2010;17:348–357. doi: 10.1038/nsmb.1784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Ahmed ZM, et al. The tip-link antigen, a protein associated with the transduction complex of sensory hair cells, is protocadherin-15. Journal of Neuroscience. 2006;26:7022–7034. doi: 10.1523/JNEUROSCI.1163-06.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Kazmierczak P, et al. Cadherin 23 and protocadherin 15 interact to form tip-link filaments in sensory hair cells. Nature. 2007;449:87–91. doi: 10.1038/nature06091. [DOI] [PubMed] [Google Scholar]
- 76.Corey DP. Stringing the fiddle: the inner ear’s two-part invention. Nature Neuroscience. 2007;10:1232–1233. doi: 10.1038/nn1007-1232. [DOI] [PubMed] [Google Scholar]
- 77.Lelli A, Kazmierczak P, Kawashima Y, Muller U, Holt JR. Development and regeneration of sensory transduction in auditory hair cells requires functional interaction between cadherin-23 and protocadherin-15. J Neurosci. 2010;30:11259–11269. doi: 10.1523/JNEUROSCI.1949-10.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Elledge HM, et al. Structure of te N terminus of cadherin 23 reveals a new adhesion mechanism for a subset of cadherin superfamily members. Proceedings of the National Academy of Sciences, USA. 2010;107:10708–100712. doi: 10.1073/pnas.1006284107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Sotomayor M, Weihofen WA, Gaudet R, Corey DP. Structure of a force-conveying cadherin bond essential for inner-ear mechanotransduction. Nature. 2012;492:128–132. doi: 10.1038/nature11590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Sotomayor M, Weihofen WA, Gaudet RG, Corey DP. Structural determinants of cadherin-23 function in hearing and deafness. Neuron. 2010;66:85–100. doi: 10.1016/j.neuron.2010.03.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Ahmed ZM, et al. Gene structure and mutant alleles of PCDH15: nonsyndromic deafness DFNB23 and type 1 Usher syndrome. Hum Genet. 2008;124:215–223. doi: 10.1007/s00439-008-0543-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Indzhykulian AA, et al. Molecular remodeling of tip links underlies mechanosensory regeneration in auditory hair cells. PLoS Biology. 2013:e1001583. doi: 10.1371/journal.pbio.1001583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Geng R, et al. Noddy, a mouse harboring a missense mutation in protocadherin-15, reveals the impact of disrupting a critical interaction site between tip-link cadherins in inner ear hair cells. J Neurosci. 2013;33:4395–4404. doi: 10.1523/JNEUROSCI.4514-12.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Yagi T. Molecular codes for neuronal individuality and cell assembly in the brain. Front Mol Neurosci. 2012;5:45. doi: 10.3389/fnmol.2012.00045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Wang X, et al. Gamma protocadherins are required for survival of spinal interneurons. Neuron. 2002;36:843–854. doi: 10.1016/s0896-6273(02)01090-5. [DOI] [PubMed] [Google Scholar]
- 86.Junghans D, Haas IG, Kemler R. Mammalian cadherins and protocadherins: about cell death, synapses and processing. Curr Opin Cell Biol. 2005;17:446–452. doi: 10.1016/j.ceb.2005.08.008. [DOI] [PubMed] [Google Scholar]
- 87.Lefebvre JL, Zhang Y, Meister M, Wang X, Sanes JR. gamma-Protocadherins regulate neuronal survival but are dispensable for circuit formation in retina. Development. 2008;135:4141–4151. doi: 10.1242/dev.027912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Chen WV, et al. Functional significance of isoform diversification in the protocadherin gamma gene cluster. Neuron. 2012;75:402–409. doi: 10.1016/j.neuron.2012.06.039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Sago H, et al. Cloning, expression, and chromosomal localization of a novel cadherin related protein, protocadherin-3. Genomics. 1995;29:631–640. doi: 10.1006/geno.1995.9956. [DOI] [PubMed] [Google Scholar]
- 90.Triana-Baltzer GB, Blank M. Cytoplasmic domain of protocadherin-alpha enhances homophilic interactions and recognizes cytoskeletal elements. J Neurobiol. 2006;66:393–407. doi: 10.1002/neu.20228. [DOI] [PubMed] [Google Scholar]
- 91.Mutoh T, Hamada S, Senzaki K, Murata Y, Yagi T. Cadherin-related neuronal receptor 1 (CNR1) has cell adhesion activity with beta1 integrin mediated through the RGD site of CNR1. Exp Cell Res. 2004;294:494–508. doi: 10.1016/j.yexcr.2003.11.019. [DOI] [PubMed] [Google Scholar]
- 92.Morishita H, et al. Structure of the cadherin-related neuronal receptor/protocadherin-α first extracellular cadherin domain reveals diversity across cadherin families. Journal of Biological Chemistry. 2006;281:33650–33663. doi: 10.1074/jbc.M603298200. [DOI] [PubMed] [Google Scholar]
- 93.Frank M, et al. Diferential expression of individual gamma-protocadherins during mouse brain development. Mol Cell Neurosci. 2005;29:603–616. doi: 10.1016/j.mcn.2005.05.001. [DOI] [PubMed] [Google Scholar]
- 94.Fernandez-Monreal M, Kang S, Phillips GR. Gamma-protocadherin homophilic interaction and intracellular trafficking is controlled by the cytoplasmic domain in neurons. Mol Cell Neurosci. 2009;40:344–353. doi: 10.1016/j.mcn.2008.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Schreiner D, Weiner JA. Combinatorial homophilic interaction between gamma protocadherin multimers greatly expands the molecular diversity of cell adhesion. Proceedings of the National Academy of Sciences, USA. 2010;107:14893–14898. doi: 10.1073/pnas.1004526107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Reiss K, et al. Regulated ADAM10-dependent ectodomain shedding of gammaprotocadherin C3 modulates cell-cell adhesion. J Biol Chem. 2006;281:21735–21744. doi: 10.1074/jbc.M602663200. [DOI] [PubMed] [Google Scholar]
- 97.Ozawa M, Kemler R. Altered cell adhesion activity by pervanadate due to the dissociation of alpha- catenin from the E-cadherin. catenin complex J Biol Chem. 1998;273:6166–6170. doi: 10.1074/jbc.273.11.6166. [DOI] [PubMed] [Google Scholar]
- 98.Han MH, Lin C, Meng S, Wang X. Proteomics analysis reveals overlapping functions of clustered protocadherins. Mol Cell Proteomics. 2010;9:71–83. doi: 10.1074/mcp.M900343-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Murata Y, Hamada S, Morishita H, Mutoh T, Yagi T. Interaction with protocadheringamma regulates the cell surface expression of protocadherin-alpha. J Biol Chem. 2004;279:49508–49516. doi: 10.1074/jbc.M408771200. [DOI] [PubMed] [Google Scholar]
- 100.Bonn S, Seeburg PH, Schwarz MK. Combinatorial expression of alpha- and gamma- protocadherins alters their presenilin-dependent processing. Mol Cell Biol. 2007;27:4121–4132. doi: 10.1128/MCB.01708-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Chen J, et al. alpha- and gamma-protocadherins negatively regulate PYK2. J Biol Chem. 2009;284:2880–2890. doi: 10.1074/jbc.M807417200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Biswas S, Emond MR, Jontes JD. The clustered protocadherins Pcdhα and Pcdhγ form a heteromeric complex in zebrafish. Neuroscience. 2012;219:280–289. doi: 10.1016/j.neuroscience.2012.05.058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Blevins CJ, Emond MR, Biswas S, Jontes JD. Differential expression, alternative splicing, and adhesive properties of the zebrafish δ1-protocadherins. Neuroscience. 2011;199:523–534. doi: 10.1016/j.neuroscience.2011.09.061. [DOI] [PubMed] [Google Scholar]
- 104.Yamagata K, et al. Arcadlin is a neural activity-regulated cadherin involved in long term potentiation. J Biol Chem. 1999;274:19473–19479. doi: 10.1074/jbc.274.27.19473. [DOI] [PubMed] [Google Scholar]
- 105.Biswas S, Emond MR, Jontes JD. Protocadherin-19 and N-cadherin interact to control cell movements during anterior neurulation. J Cell Biol. 2010;191:1029–1041. doi: 10.1083/jcb.201007008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Emond MR, Biswas S, Blevins CJ, Jontes JD. A complex of protocadherin-19 and N-cadherin mediates a novel mechanism of cell adhesion. J Cell Biol. 2011;195:1115–1121. doi: 10.1083/jcb.201108115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Ozka E, et al. An Extracellular Interactome of Immunoglobulin and LRR Proteins Reveals Receptor-Ligand Networks. 2013 doi: 10.1016/j.cell.2013.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Xiong F, et al. Specified neural progenitors sort to form sharp domains after noisy shh signaling. Cell. 2013;153:550–561. doi: 10.1016/j.cell.2013.03.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Demireva EY, Shapiro LS, Jessell TM, Zampieri N. Motor neuron position and topographic order imposed by β- and γ-catenin activities. Cell. 2011;147:641–652. doi: 10.1016/j.cell.2011.09.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Morgan JL, Lichtman JW. Why not connectomics? Nature Methods. 2013;10:494–500. doi: 10.1038/nmeth.2480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Li Y, et al. Molecular and functional interaction between protocadherin-γC5 and GABAA receptors. J Neurosci. 2012;32:11780–11797. doi: 10.1523/JNEUROSCI.0969-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Tabdili H, et al. Cadherin point mutations alter cell sorting and modulate GTPase signaling. J Cell Sci. 2012;125:3299–3309. doi: 10.1242/jcs.087395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Chen B, et al. The WAVE regulatory complex links diverse receptors to the actin cytoskeleton. Cell. 2014;156:195–207. doi: 10.1016/j.cell.2013.11.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Sadeqzadeh E, de Bock CE, Thorne RF. Sleeping giants: Emerging roles for the Fat cadherins in health and disease. Med Res Rev. 2013;34:190–221. doi: 10.1002/med.21286. [DOI] [PubMed] [Google Scholar]
- 115.Deans MR, et al. Control of neuronal morphology by the atypical cadherin Fat3. Neuron. 2011;71:820–832. doi: 10.1016/j.neuron.2011.06.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Boutin C, Goffinet AM, Tissir F. Celsr1-3 cadherins in PCP and brain development. Curr Top Dev Biol. 2012;101:161–183. doi: 10.1016/B978-0-12-394592-1.00010-7. [DOI] [PubMed] [Google Scholar]
- 117.Friedman TB, Schultz JM, Ahmed Z, Tsilou ET, CBC Usher syndrome: hearing loss with vision loss. Adv Otorhinolaryngol. 2011;70:56–65. doi: 10.1159/000322473. [DOI] [PubMed] [Google Scholar]
- 118.Bonnet CEAA. Usher syndrome (sensorineural deafness and retinitis pigmentosa): pathogenesis, molecular diagnosis and therapeutic approaches. Curr Opin Neurol. 2012;25:42–49. doi: 10.1097/WCO.0b013e32834ef8b2. [DOI] [PubMed] [Google Scholar]
- 119.McConnell RE, Benesh AE, Mao S, Tabb DL, Tyska MJ. Proteomic analysis of the enterocyte brush border. Am J Physiol Gastrointest Liver Physiol. 2011;300:G914–926. doi: 10.1152/ajpgi.00005.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Baumgartner W, et al. Heterotypic trans-interaction of LI- and E-cadherin and their localization in plasmalemmal microdomains. J Mol Biol. 2008;378:44–54. doi: 10.1016/j.jmb.2008.02.023. [DOI] [PubMed] [Google Scholar]
- 121.Bartolmas T, Hirschfeld-Ihlow C, Jonas S, Schaefer M, Gessner R. LI-cadherin cis dimerizes in the plasma membrane Ca(2+) independently and forms highly dynamic transcontacts. Cell Mol Life Sci. 2012;69:3851–3862. doi: 10.1007/s00018-012-1053-y. [DOI] [PMC free article] [PubMed] [Google Scholar]