Abstract
Rationale: Lung cancer is the leading cause of cancer death in both men and women in the United States and worldwide. Matrix metalloproteinases (MMPs) have been implicated in the development and progression of lung cancer, but their role in the molecular pathogenesis of lung cancer remains unclear. We have found that MMP19, a relatively novel member of the MMP family, is overexpressed in lung tumors when compared with control subjects.
Objectives: To test the hypothesis that MMP19 plays a significant role in the development and progression of non–small cell lung cancer (NSCLC).
Methods: We have analyzed lung cancer gene expression data, immunostained lung tumors for MMP19, and performed in vitro assays to test the effects of MMP19 in NSCLC cells.
Measurements and Main Results: We found that MMP19 gene and protein expression is increased in lung cancer tumors compared with adjacent and histologically normal lung tissues. In three independent datasets, increased MMP19 gene expression conferred a poorer prognosis in NSCLC. In vitro, we found that overexpression of MMP19 promotes epithelial–mesenchymal transition, migration, and invasiveness in multiple NSCLC cell lines. Overexpression of MMP19 with a mutation at the catalytic site did not impair epithelial–mesenchymal transition or expression of prometastasis genes. We also found that miR-30 isoforms, a microRNA family predicted to target MMP19, is markedly down-regulated in human lung cancer and regulates MMP19 expression.
Conclusions: Taken together, these findings suggest that MMP19 is associated with the development and progression of NSCLC and may be a potential biomarker of disease severity and outcome.
Keywords: MMP19, epithelial–mesenchymal transition, metastasis, non–small cell lung cancer, miR-30
At a Glance Commentary
Scientific Knowledge on the Subject
Lung cancer is the leading cause of cancer death in the United States and worldwide. Matrix metalloproteinases (MMPs) have been implicated in the development and progression of lung cancer, but their role in the molecular pathogenesis of lung cancer remains unclear.
What This Study Adds to the Field
MMP19, a poorly understood member of the MMP family, is associated with the development and progression of lung cancer and may be a potential biomarker of disease severity and outcome.
Lung cancer is the leading cause of cancer death in the United States and worldwide (1). Chemotherapies to treat lung cancer are highly toxic and largely ineffective. The overall 5-year survival for all types of lung cancer is only 16% (2). Understanding of the molecular pathology of lung cancer is central to treating this devastating disease.
Metastatic potential of tumors depends in large part on the ability of the tumor to invade the extracellular matrix, a process that requires cells to lose intercellular adhesion molecules and to express proteolytic enzymes to facilitate invasion (3, 4). Among the most widely studied class of proteolytic enzymes are the matrix metalloproteinases (MMPs), zinc-dependent endopeptidases that can degrade a wide range of substrates and are thought to be essential for basement-membrane penetration during metastasis (5–7). The roles of individual MMP family members are distinct in different pathophysiologic settings (8, 9). Therefore, understanding how individual family members contribute to disease is essential for developing MMP-targeted therapies. We have recently shown that expression of MMP19, a poorly characterized member of the MMP family, is significantly enriched in the lungs of patients with idiopathic pulmonary fibrosis (10), a disease that confers an increased risk for lung cancer (11–13) and shares many pathophysiologic mechanisms with cancer (14). Moreover, we found that expression of MMP19 in lung epithelial cells stimulated proliferation and cell migration (10). Because of the importance of cell proliferation and migration to lung cancer metastasis, we sought to understand if MMP19 played a similarly pathologic role in lung cancer. In this report, we tested the hypothesis that MMP19 expression in non–small cell lung cancer (NSCLC) is enhanced and that increased expression of MMP19 in lung cancer cells in vitro would promote cellular functions that are associated with lung cancer growth and metastasis.
Methods
The institutional review boards of the University of Pittsburgh and McMaster University approved all protocols.
Immunohistochemistry
Details are located in the Methods section of the online supplement.
Cell Culture
NSCLC cells were purchased from ATCC (Manassas, VA) and propagated in F12K (Invitrogen, Carlsbad, CA) under standard conditions.
MMP19 Wild-Type and Mutant cDNA Construction
MMP19 cDNA with vector pCMV-sport6 was from OpenBiosystems (Lafayette, CO). Mutants and adenoviral vectors were constructed as described (15). Full details are presented in the Methods section of the online supplement.
Human Tissues
Lung cancer tissue-arrays were from Pantomics (San Francisco, CA). Lung tissues for RNA extraction (stage II NSCLC, n = 22) were obtained from the University of Pittsburgh.
Quantitative Real-Time Polymerase Chain Reaction
Gene expression for MMP19 was determined using Taqman (Applied Biosystems, Foster City, CA) (16, 17). Full details are presented in the Methods section of the online supplement.
Immunoblotting
Full details are presented in the Methods section of the online supplement and have been described previously (10).
In Vitro Assays
The procedures of colony formation (18, 19) and matrigel transmigration have been described (20). Details are presented in the online supplement.
RNA Extraction
Total RNA was extracted from frozen lung in Qiazol (Qiagen, Valencia, CA) and disrupted as described (10).
Microarray Experiments
Full details are presented in the online supplement. Data are deposited in GEO (GSE47115).
Statistical Analysis
Data were analyzed by t test for comparisons between two groups. Data are presented as mean ± SD and were considered statistically significant at P less than 0.05.
MMP19 Expression and Survival
Microarray experiments were obtained from three cohorts, using two different microarray platforms: the “Challenge” (21) and the “Duke” (22) cohorts used Affymetrix, and the “Pitt cohort” used Illumina. Full details are available in the Methods section of the online supplement.
Somatic mutation polymerase chain reaction assay.
The qBiomarker Somatic Mutation PCR Array was used to screen for lung cancer disease-focused mutation profiling (SABiosciences/Qiagen, Valencia, CA). Full details are available in the online supplement.
Statistical analysis.
For overall survival analyses based on MMP19 microarray gene expression, we used the survival (23) package of the R environment (24). MMP19 expression levels were dichotomized by the median gene expression value. Full details are in the Methods section of the online supplement.
Results
MMP19 Is Overexpressed in NSCLC
To determine if MMP19 expression was regulated at the level of gene transcription, total RNA was isolated from homogenates of tumors and histologically normal lung adjacent to the tumor (n = 22) and processed for quantitative reverse-transcriptase polymerase chain reaction (RT-PCR). MMP19 gene expression was increased 33% compared with control subjects (Figure 1A).
Figure 1.
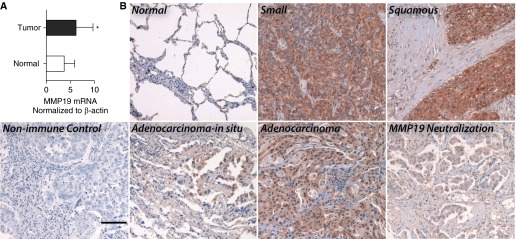
Increased expression of matrix metalloproteinase (MMP) 19 in non–small cell lung cancer (NSCLC). (A) Quantitative reverse-transcriptase polymerase chain reaction was performed for MMP19 in NSCLC (adenocarcinoma and squamous cell carcinoma) compared with corresponding nonneoplastic lung tissues. The differences in mRNA levels of MMP19 were compared by the Wilcoxon two-sample test. Data represent the mean ± SD (*P < 0.05). (B) Immunostaining was performed for MMP19 on NSCLC tissue sections as described in the Methods section (scale bar = 100 μm).
We next performed MMP19 immunostaining of neoplastic and normal human lung tissue. Expression of MMP19 was primarily observed in neoplastic cells (Figure 1B). There is a distinct absence of staining in tumor-associated stroma. Compared with lung adenocarcinoma, adenocarcinoma-in-situ (bronchioloalveolar carcinoma) exhibited less staining. In comparison, normal lung tissue exhibited less immunostaining for MMP19. By immunoblotting, we surveyed expression of MMP19 across different lung cancer cell lines (see Figure E1 in the online supplement). MMP19 was readily detected in A549 and H522 cells. In comparison, H23, H226, and H460 cells expressed less. Thus, MMP19 gene expression is increased in NSCLC.
Increased MMP19 Expression Portends a Poor Prognosis
To test the hypothesis that MMP19 expression is associated with prognosis in NSCLC, we examined microarray data from three independent cohorts of patients. The first was obtained from publically available data from the Director’s Challenge Consortium for the Molecular Classification of Lung Adenocarcinoma (referred hereafter as the “Challenge” [21]). Details of the samples are presented in the Methods section. The second cohort is from the University of Pittsburgh (Pitt). Table E1 summarizes the demographic and clinical characteristics of the study subjects in the Challenge and Pitt cohorts. The third cohort was obtained from publically available dataset, referred to as the “Duke cohort” (22). All the subjects in the Challenge cohort had lung adenocarcinomas, and squamous cell carcinomas (21.7%) were represented in the Pitt cohort. Most of the cases represented early stage disease (stage I) in these two cohorts. A total of 52.3% of the subjects in the Duke cohort had adenocarcinomas by histology, whereas 47.7% had squamous cell carcinomas. Clinical data were not available for this cohort.
A total of 32 microarray samples from the Challenge cohort were excluded from our study because they were determined to be outliers by the dChip microarray processing software. The remaining 408 samples were used for overall survival analysis. Only 291 of these subjects had available clinical and therapy information, and this group was used to perform statistical adjustments of overall survival in the Challenge cohort. Increased MMP19 gene expression by microarray levels above the median threshold was significantly associated with decreased survival in the Challenge (hazard ratio [HR], 1.59; 95% confidence interval [CI], 1.19–2.13; P = 0.001) and the Pitt (HR, 2.28; 95% CI, 1.00–5.21; P = 0.033) cohorts (Figures 2A and 2B). Median survival in subjects from the Challenge cohort with MMP19 microarray gene expression above the median threshold was 68 versus 84.5 months in subjects with MMP19 expression below the median threshold. Similarly, subjects from the Pitt cohort with MMP19 microarray gene expression above the median had shorter survival times (44.8 mo) when compared with the subjects with levels below the median (93.4 mo). Survival differences in subjects with high MMP19 persist after adjusting for age, sex, and adjuvant therapy in the Challenge (HR, 1.5; 95% CI, 1.04–2.1; P = 0.023) (Figure 2C; see Table E2) and Pitt cohort (HR, 4.1; 95% CI, 1.52–11.2; P = 0.005) (Figure 2D; see and Table E3). From the Duke cohort (see Figure E2), subjects with MMP19 expression levels above the median had worst survival than subjects with low MMP19 levels (HR, 1.67; 95% CI, 1.00–2.8; P < 0.05). The median survival of subjects with high MMP19 levels was 33.5 versus 55.3 months for subjects with low MMP19 levels.
Figure 2.
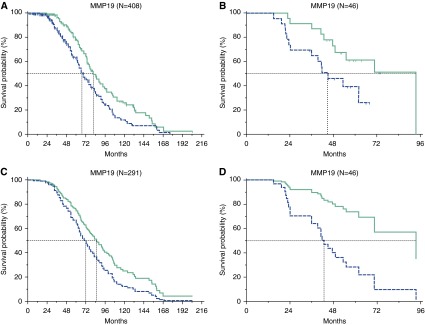
Increased matrix metalloproteinase (MMP) 19 gene expression in non–small cell lung cancer portends a worse prognosis. Overall survival analysis by Kaplan-Meier curves of MMP19 microarray gene expression levels in lung adenocarcinoma tissue obtained from 408 subjects from the Director’s Challenge Consortium for the Molecular Classification of Lung Adenocarcinoma (A) and 46 non–small cell lung cancer subjects obtained from the University of Pittsburgh (B). Adjusted overall survival differences in the (C) Challenge cohort (adjusted for age, sex, adjuvant chemotherapy, and radiotherapy) and (D) Pitt cohort (adjusted for age, sex, adjuvant chemotherapy, and radiotherapy as well as intraoperative brachytherapy). Blue line = patients with MMP19 microarray expression levels above the median value. Green line = patients with MMP19 microarray expression levels below the median value. Dotted black lines = the median survival in months for both groups.
Because of the critical associations between NSCLC and certain driver mutations (25), we next asked if MMP19 expression was associated with any of these mutations. To answer this question, we screened for 83 known mutations and 11 copy number changes in 11 genes as described in the Methods section. We do not have mutation data for six samples, but 60% of the remaining samples had one or more mutation detected. We were not powered to consider the association with survival for each mutation alone. We totaled the number of cases with a detected mutation for both arms (high or low MMP19) and found that mutation status did not influence the MMP19 expression level association with overall survival. Similarly, there was no difference in overall survival based solely on mutation status. Taken together these data suggest that high MMP19 gene expression is a negative prognostic factor in NSCLC.
MMP19 Promotes Colony Formation of A549 Cells
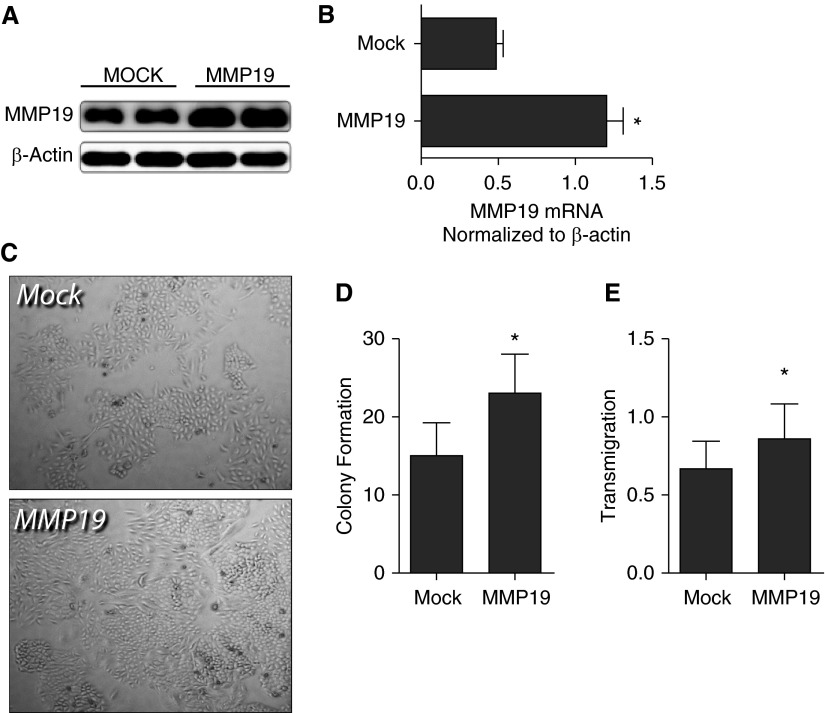
The functional consequences of increased MMP19 expression in NSCLC are unknown. To investigate the role of MMP19 in the pathogenesis of NSCLC, we perturbed expression of MMP19 in vitro and determined the effect on cellular processes relevant to cancer. In a first set of experiments we transfected full length MMP19 into A549 cells. We confirmed increased expression of MMP19 by immunoblotting and by quantitative RT-PCR (Figures 3A and 3B). Following transfection, we determined the number of A549 colonies formed on both tissue culture-treated plastic and soft agar. MMP19-transfected cells formed a significantly increased number of colonies (57%) compared with the mock-transfected control subjects (Figures 3C and 3D). We next determined if MMP19 overexpression would promote invasion into matrigel. A549 cells were transfected with MMP19 or a mock control, and the number of cells invading the matrigel matrix was quantified. A total of 33% more MMP19-transfected cells invaded matrigel compared with mock control subjects (Figure 3E). Thus MMP19 overexpression in A549 cells is associated with two potentially important cancerous processes including colony formation and invasion.
Figure 3.
Matrix metalloproteinase (MMP) 19 expression promotes colony formation and motility in vitro. (A) Immunoblotting for MMP19 protein was performed on lysates from A549 cells stably expressing MMP19 compared with vector control. (B) Quantification of band intensity was performed with National Institutes of Health ImageJ as described in the Methods section (*P < 0.05). (C) Representative image of colony formation of A549 cells transfected with MMP19 full length cDNA and vector control. Colonies were allowed to form on a hard surface over 10 days. (D) Promotion of colony formation and anchorage-independent growth of A549 cells transfected with MMP19 compared with control subjects. Colonies were allowed to form on soft agar over 21 days and quantified as described in the Methods section (*P < 0.05). (E) Following transfection of A549 cells with MMP19, cell migration and invasion into matrigel was quantified as described in the Methods section (*P < 0.05). Data represent mean of five experiments with SEM. Five high-power fields were counted for each experiment.
Loss of MMP19 Decreases Proliferation of Lung Cancer Cells
To investigate the association between MMP19 and proliferation in vitro, we transfected MMP19-targeting siRNA, MMP19 cDNA, or controls into A549, H522, and H1299 cells (see Figure E3). We found a significant reduction in proliferation as measured by the MTT assay in all three cell lines following loss of MMP19. Overexpression of MMP19 did not lead to increased proliferation in these cells. Thus, loss of MMP19 leads to decreased proliferation in several lung cancer cell lines.
MMP19 Expression Promotes Epithelial–Mesenchymal Transition In Vitro
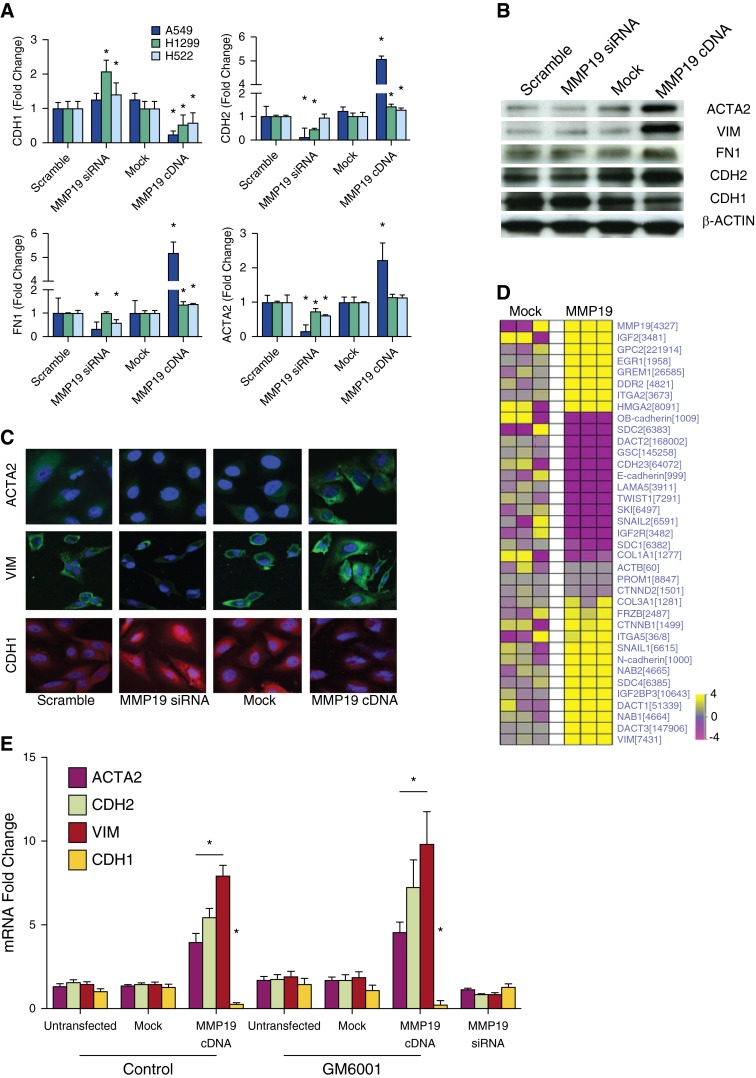
Epithelial–mesenchymal transition (EMT) is a process by which epithelial cells lose their intercellular adhesion proteins, exhibit increased motility, and express mesenchymal markers (26, 27). To define whether MMP19 expression is associated with EMT, we used loss and gain of MMP19 function strategies: we transfected MMP19 cDNA and siRNA into multiple lung cancer cell lines: A549, H1299, and H522 cells. We quantified changes in the expression of mesenchymal and epithelial cell markers (Figure 4A). Efficiency of MMP19 silencing is shown in Figure E4. We noted low baseline expression of MMP19 in H1299 and H522 cells. After silencing of MMP19 we found gene expression changes that favored the epithelial phenotype: we found increased expression of the epithelial marker CDH1 (E-cadherin) in H1299 and H522 cells. We also found decreased expression of the mesenchymal markers CDH2 (N-cadherin, in A549 and H1299 cells), FN1 (fibronectin, in A549 and H522 cells), and ACTA2 (α-smooth muscle actin). With increased expression of MMP19, we found gene expression changes favoring the mesenchymal phenotype. CDH1 was decreased in all three lines. CDH2 and FN1 gene expression were increased in all three lines. ACTA2 was increased in A549 cells. These changes were validated by immunoblotting (Figure 4B; see Figure E5).
Figure 4.
Matrix metalloproteinase (MMP) 19 induces epithelial–mesenchymal transition (EMT) in vitro. (A) Quantitative reverse-transcriptase polymerase chain reaction was performed for markers of EMT in A549, H1299, and H522 cells 24 hours after transient transfection with MMP19 siRNA and full length MMP19 cDNA compared with control subjects (ACTA2 = α-smooth muscle actin; CDH2 = N-cadherin; FN1 = fibronectin 1; VIM = vimentin). Data were analyzed by t test, *P < 0.05, MMP19 silencing or overexpression compared with the scrambled control, or MMP19 overexpression compared with the mock control. (B) Immunoblots for the epithelial marker (CDH1 = E-cadherin) and mesenchymal markers (ACTA2, VIM, FN1, and CDH2) in A549 cell lysates from MMP19 cDNA and siRNA transiently transfected A549 cells and mock control. (C) Immunofluorescent staining of A549 cells for ACTA2 (green, top) and VIM (green, middle), and CDH1 (red). Cells were counterstained with DAPI (blue; original magnification ×400). (D) Microarray heatmap of EMT-relevant gene expression in stable MMP19-transfected A549 cells and mock control as described in the Methods section. (E) Quantitative reverse-transcriptase polymerase chain reaction was performed for markers of EMT following silencing or overexpression of MMP19 in the presence of the MMP inhibitor GM6001. GM6001 did not block differential expression of these EMT markers (*P < 0.05 compared with mock transfection control).
Microscopically, we observed that MMP19-transfected A549 cells exhibited a spindle-shaped morphology, more characteristic of mesenchymal cells. These MMP19-transfected cells expressed lower levels of CDH1 and increased ACTA2 and VIM by immunofluorescence (Figure 4C). Conversely, silencing of MMP19 by siRNA in A549 cells led to attenuated expression of VIM and ACTA2 and enhanced expression of CDH1. To understand the global effect of MMP19 expression in A549 cells on “EMT signature” genes (28, 29), we reanalyzed gene expression microarray data (GSE34994) performed in MMP19-transfected cells compared with mock transfection control subjects (10). We found differential expression of several EMT-relevant genes including genes coding for cell-surface proteins, cytoskeletal markers, extracellular matrix proteins, and EMT transcription factors (Figure 4D). Thus, overexpression of MMP19 leads to global changes consistent with EMT in A549 cells.
We next determined if chemical inhibition of MMP19 would mimic silencing of MMP19 and attenuate EMT (Figure 4E). We transfected A549 cells with MMP19 of the MMP inhibitor GM6001, a hydroxamic acid inhibitor of MMPs (30). We found that incubation of A549 cells with GM6001 failed to block EMT in A549 cells. Thus MMP19-induced EMT is not blocked by the MMP inhibitor GM6001.
Mutated MMP19 Catalytic Domain Does Not Affect Migration or EMT
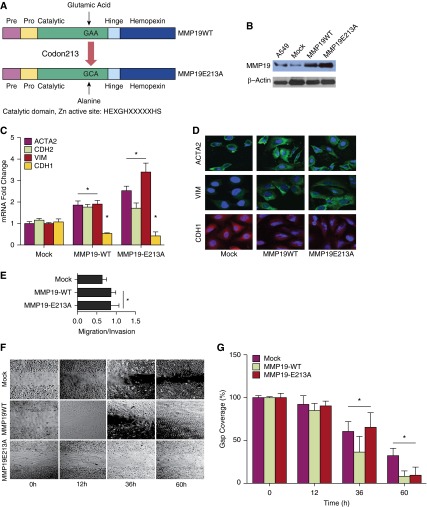
Because we found that the MMP inhibitor GM6001 did not block EMT, we next determined if a functional MMP19 catalytic site was necessary for the phenotypes that we have observed following transfection of MMP19: EMT and migration. To answer this question, we generated a mutated MMP19 construct at codon 213 (Figure 5A) that has no enzymatic activity to determine the effects of MMP19 catalytic domain on EMT, colony formation, and invasiveness. Loss of catalytic activity of the MMP19E213A construct is shown in Figure E6. The MMP19E213A construct is detectable by immunoblotting and showed similar expression levels to the wild-type construct (Figure 5B). Overexpression of both the wild-type and mutant MMP19 constructs increased expression of ACTA2, CDH2, and VIM expression in A549 cells and decreased expression of CDH1 by quantitative RT-PCR (Figure 5C) and by immunofluorescence (Figure 5D).
Figure 5.
Effect of matrix metalloproteinase (MMP) 19 catalytic domain on A549 in vitro phenotypes: epithelial–mesenchymal transition, migration, and invasion. (A) Schematic representation of the E213A MMP19 point mutation in the catalytic domain. (B) Pitt cohort of MMP19 expression in protein in A549 cells transfected with wild-type (WT) and the E213A mutated MMP19 construct by immunoblotting. (C) Quantitative reverse-transcriptase polymerase chain reaction was performed for markers of epithelial–mesenchymal transition. Data represent the mean ± SEM (*P < 0.05 compared with mock transfection control). (D) Immunofluorescent staining of mesenchymal markers (anti-ACTA2 and anti-VIM, green) and epithelial marker (anti-CDH1, red). Cells were counterstained with DAPI (blue). (E) Induction of A549 cell migration and invasion with expression of MMP19 WT and MMP19E213A in matrigel. Data represent mean of five experiments with SEM. Five high-power fields were counted for each experiment (*P < 0.05 compared with mock transfection control). (F) Scratch assay was performed on A549 following transfection of the mock control, MMP19 WT, and the E213A MMP mutant. (G) Quantification of the gap coverage as described in the Methods section (*P < 0.05, compared with time = 0 control). ACTA2 = α-smooth muscle actin; CDH1 = E-cadherin; VIM = vimentin.
Then we analyzed the effect of MMP19 wild-type and MMP19E213A overexpression on the invasiveness of A549 cells with the matrigel assay. We found that both wild-type and mutant MMP19 overexpression stimulated migration and invasion of A549 cells compared with mock control (P < 0.05) (Figure 5E). We next performed a scratch assay to determine the effect of the mutated catalytic domain of MMP19 on migration (Figures 5F and 5G). After the scratch, the wild-type MMP19 construct augmented coverage of the gap to a greater extent than the mock transfection control or the E213A MMP19 mutant at 36 hours. However, by 60 hours, both the wild-type and E213A MMP19 constructs covered the gap. Thus MMP19-driven EMT and migration seem to be independent of the catalytic site.
Effect of MMP19 on the Expression of Genes Associated with Metastasis and Prognosis
To begin to address the mechanism by which MMP19 controls the invasiveness of lung cancer cells, we reexamined gene expression analysis of MMP19-transfected A549 cells. We queried the dataset for genes enriched in metastasis (Figure 6A; see Tables E4 and E5) (31, 32) and prognosis (Figure 6B; see Tables E6 and E7) (33, 34). Of the metastasis genes, syntaxin 1A, hypoxia inducible factor 1α, and chaperonin-containing TCP1 subunit 3 are all increased in MMP19-transfected cells. These genes were common to a three- and six-gene prognostic signature for NSCLC and were among the top 0.02% of signatures with maximum verifiability (35). Of the “prognosis” genes, overexpression of MMP19 gene led to differential expression of 46 genes. A total of 32 genes were markedly up-regulated and 14 genes were down-regulated. Most notably, we observed that overexpression of MMP19 in A549 cells caused a significant increase (>95%) in the expression of several proproliferative growth factors and other MMPs. Taken together, high MMP19 gene expression in NSCLC is associated with a global gene expression pattern linked to metastasis and poor prognosis.
Figure 6.
Effect of matrix metalloproteinase (MMP) 19 overexpression on gene signatures associated with metastasis and prognosis gene in non–small cell lung cancer. Agilent microarray heatmaps of metastasis (A) and prognosis (B) gene expression signatures in A549 cells following stable transfection with MMP19 and vector control (P < 0.05).
Low miR-30 Expression Is Associated with Increased MMP19 in NSCLC
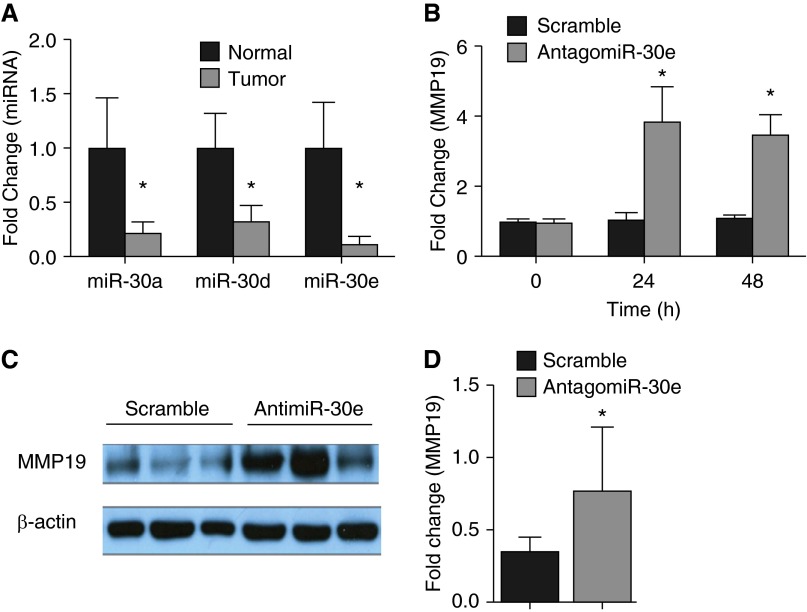
Accumulating evidence indicates that miRNAs are critical regulators of gene expression in cancer (36–38). Public databases suggest that the miR-30 family targets MMP19 (39). The expression of miR-30 in lung cancer is unknown. In this experiment, we tested the hypothesis that expression of miR-30 regulates expression of MMP19 in NSCLC. To investigate the expression of miR-30 isoforms in NSCLC, quantitative RT-PCR analysis was performed. We found that miR-30a, -30d, and -30e were significantly down-regulated in NSCLC (Figure 7A). These data suggest that the relative lack of these miRNAs may explain increased expression of MMP19. To test the hypothesis that MMP19 is a target gene of miR-30, we transfected a miR-30e antagonist (antimiR-30e) into A549 cells and found that MMP19 was up-regulated by both mRNA (Figure 7B) and protein (Figures 7C and 7D). These data suggest that miR-30 may target MMP19 and that down-regulation of miR-30 may lead to MMP19 overexpression in lung cancer.
Figure 7.
miR-30 regulates matrix metalloproteinase (MMP) 19 expression in A549 cells. (A) Expression of miR-30a, -30d, and -30e were down-regulated in human non–small cell lung cancer compared with corresponding normal control subjects as detected by Taqman quantitative reverse-transcriptase polymerase chain reaction (*P < 0.05; n = 15). (B) MMP19 gene expression was determined by reverse-transcriptase polymerase chain reaction in A549 cells 24 hours following transfection with antagomiR-30e oligonucleotides or controls. Data represent the mean ± SEM (*P < 0.05, n = 4). (C) MMP19 protein expression was determined by immunoblotting lysates of A549 cells 48 hours following transfection with antagomiR-30e oligonucleotides or control. (D) Quantification of MMP19 band intensity in C in ImageJ. Data represent mean ± SEM (*P < 0.05, n = 3).
Discussion
The MMPs likely represent the most prominent family of proteases involved in tumor invasion (40). A challenge to researchers studying all 23 unique MMP proteins (5) is to understand the divergent and redundant functions of all the members of the MMP family and how they may be differentially regulated in cancer (41). In this study, we have investigated the role of a lesser-known member of the MMP family, MMP19, in promoting the malignant behaviors of NSCLC. Our key observation, using genomic analysis of three independent cohorts of patients with NSCLC, is that increased expression of MMP19 is associated with decreased survival in NSCLC patients. To further explore the mechanism of our clinical observations, we have shown that MMP19 expression in multiple lung cancer cell lines is associated with invasive phenotypes including migration and EMT. Our data support previous studies that have shown that MMP19 expression is associated with increased invasiveness in vitro in different cancers (42–44). These results point to a crucial role for MMP19 in the evolution of NSCLC.
Little is known about the regulation of MMP19 expression (44). Regulation of MMPs may occur at multiple levels from transcription, enzymatic activation, cellular compartmentalization, and inactivation by protein inhibitors (40). In lung cancer, MMP gene expression may be regulated by variations in copy number (45). To examine this question, we have analyzed the Pitt cohort and found that MMP19 expression is independent of a selected number of mutations and copy number variation. We have shown a relatively novel mechanism regulating MMP expression in lung cancer. MicroRNAs have been increasingly recognized as critical regulators of MMP expression in cancer (46, 47). We found that the regulation of MMP19 depends, at least in part, on the expression of the miR-30 family. Whether or not MMP19 is principally regulated by the miR-30 family in NSCLC is unclear. We found that miR-30 isoforms are down-regulated in NSCLC and that blockade of miR-30e dramatically stimulated MMP19 expression at transcriptional and translational levels in vitro. Down-regulation of miR-30 may lead to increased expression of multiple proteins, including MMP19, that promote tumorigenesis and metastasis. The application of microRNA mimics or antagonists is currently being studied as treatment for NSCLC (17, 48).
The role of tumor microenvironment on MMP19 gene expression is similarly unknown. We found that MMP19 gene expression is modestly increased compared with histologically normal tissue adjacent to the tumor. So-called “field effects,” the concept that suggests that cells in proximity to cancer cells are premalignant and should exhibit at least some, but not all, of the genetic alterations that are present in the fully developed cancer (49–52), may explain the only modest differences in MMP19 expression between tumor and adjacent “normal” tissue. Further study is necessary to elucidate extracellular cues that may lead to increased expression of MMP19.
How does MMP19 promote the invasive behaviors of NSCLC cell lines? MMPs are now known to control more than proteolysis (40). Critical, nonenzymatic functions of MMPs have been recognized (53). To address these questions, we used a catalytic site mutant. We found that several phenotypes induced by MMP19, including EMT and migration, were independent of MMP19’s enzymatic function. These data suggest that there is an as yet uncharacterized nonenzymatic function of MMP19 and support the increasing importance of nonenzymatic functions of MMPs. For example, the hemopexin domain is necessary for MMP9-induced migration (54, 55) and for the antimicrobial function of MMP12 (56). Further studies are necessary to identify the noncatalytic mechanisms and the potential MMP19 hemopexin domain-binding partners that drive the invasive phenotype of MMP19-overexpressing cells.
Even with our observations that MMP19 promotes invasiveness in vitro, we do acknowledge that our proposed mechanism for the prometastatic behavior of MMP19 is limited by corroborating evidence from in vivo modeling of lung cancer. The correlation of MMP19-driven invasive behaviors in vitro with MMP19 as a biomarker of poor prognosis in NSCLC does provide evidence that MMP19 overexpression is at least one factor driving tumor growth and invasiveness. However, establishing MMP19 as a therapeutic target in NSCLC would require in vivo evidence showing that MMP19 overexpression promotes lung cancer and that loss of MMP19 is protective. Possible approaches include testing the MMP19 knockout mouse in model of lung cancer that has been associated with metastasis (57). Such studies would more firmly support a causal role for MMP19 as a driver of metastasis and poor prognosis in NSCLC.
In summary, the key finding of our study is the association of increased MMP19 expression with increased mortality. The strength of our findings is bolstered by the observation of increased mortality in high MMP19-expressing tumors across three independent datasets from different institutions and analyzed on different platforms. How can these data be translated to patients? We suggest that future studies focus on testing MMP19 as a potential biomarker to identify patients with a resected stage I tumor with a worse prognosis. The identification of genomic signatures to define stage I patients at risk for a poorer prognosis remains an active area of research precisely because it is hoped that there may be personalized interventions in the future to enhance survival for stage I patients who are deemed to be at high risk for recurrence after surgery for NSCLC (58, 59). Can MMP19 be targeted for therapy? Although design of a specific MMP19 inhibitor would be challenging given the overlap with other MMP family members, the identification of the miR-30 family as regulators of MMP19 expression could lead to treatment with miR-30 mimics to drive down expression of MMP19 specifically.
Footnotes
Supported by the Dorothy P. and Richard P. Simmons Endowed Chair for Pulmonary Research; National Institutes of Health grants R01HL107883 and P30CA047904; and the Lung Cancer Genomics project, University of Pittsburgh Medical Center.
Author Contributions: G.Y., J.D.H.-M., T.N., M.R., A.P., B.M.J.-G., J.R., J.G., and J.M.S. performed experiments. G.Y., J.D.H.-M., N.K., and D.J.K. designed the study. G.Y., J.D.H.-M., and D.J.K. analyzed data and wrote the manuscript.
This article has an online supplement, which is accessible from this issue's table of contents at www.atsjournals.org
Originally Published in Press as DOI: 10.1164/rccm.201310-1903OC on September 24, 2014
Author disclosures are available with the text of this article at www.atsjournals.org.
References
- 1.United States Cancer Statistics1999–2009 Incidence and Mortality Web-based Report. Department of Health and Human Services, Centers for Disease Control and Prevention, and National Cancer Institute, 2013[accessed 2013 Mar 6]. Available from: http://www.cdc.gov/cancer/lung/statistics/index.htm
- 2.Surveillance, Epidemiology, and End Results ProgramTurning cancer data into discovery. National Cancer Institute, 2013[accessed 2014 Apr 3]. Available from: http://seer.cancer.gov
- 3.Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674. doi: 10.1016/j.cell.2011.02.013. [DOI] [PubMed] [Google Scholar]
- 4.Talmadge JE, Fidler IJ. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res. 2010;70:5649–5669. doi: 10.1158/0008-5472.CAN-10-1040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Khokha R, Murthy A, Weiss A. Metalloproteinases and their natural inhibitors in inflammation and immunity. Nat Rev Immunol. 2013;13:649–665. doi: 10.1038/nri3499. [DOI] [PubMed] [Google Scholar]
- 6.Shuman Moss LA, Jensen-Taubman S, Stetler-Stevenson WG. Matrix metalloproteinases: changing roles in tumor progression and metastasis. Am J Pathol. 2012;181:1895–1899. doi: 10.1016/j.ajpath.2012.08.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Stallings-Mann ML, Waldmann J, Zhang Y, Miller E, Gauthier ML, Visscher DW, Downey GP, Radisky ES, Fields AP, Radisky DC, et al. Matrix metalloproteinase induction of Rac1b, a key effector of lung cancer progression. Sci Translat Med. 2012;4:142ra95. doi: 10.1126/scitranslmed.3004062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Delassus GS, Cho H, Hoang S, Eliceiri GL. Many new down- and up-regulatory signaling pathways, from known cancer progression suppressors to matrix metalloproteinases, differ widely in cells of various cancers. J Cell Physiol. 2010;224:549–558. doi: 10.1002/jcp.22157. [DOI] [PubMed] [Google Scholar]
- 9.Vanlaere I, Libert C. Matrix metalloproteinases as drug targets in infections caused by gram-negative bacteria and in septic shock. Clin Microbiol Rev. 2009;22:224–239. doi: 10.1128/CMR.00047-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Yu G, Kovkarova-Naumovski E, Jara P, Parwani A, Kass D, Ruiz V, Lopez-Otín C, Rosas IO, Gibson KF, Cabrera S, et al. Matrix metalloproteinase-19 is a key regulator of lung fibrosis in mice and humans. Am J Respir Crit Care Med. 2012;186:752–762. doi: 10.1164/rccm.201202-0302OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zatuchni J, Campbell WN, Zarafonetis CJ. Pulmonary fibrosis and terminal bronchiolar (alveolar-cell) carcinoma in scleroderma. Cancer. 1953;6:1147–1158. doi: 10.1002/1097-0142(195311)6:6<1147::aid-cncr2820060605>3.0.co;2-l. [DOI] [PubMed] [Google Scholar]
- 12.Desmeules R, Giroux M, Beaulieu M, Richard P. [Neoplastic aspect of a case of interstitial pulmonary fibrosis] Laval Med. 1955;20:1315–1325. [PubMed] [Google Scholar]
- 13.Williams WJ. Alveolar metaplasia: its relationship to pulmonary fibrosis in industry and the development of lung cancer. Br J Cancer. 1957;11:30–42. doi: 10.1038/bjc.1957.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Rabinovich EI, Kapetanaki MG, Steinfeld I, Gibson KF, Pandit KV, Yu G, Yakhini Z, Kaminski N. Global methylation patterns in idiopathic pulmonary fibrosis. PLoS ONE. 2012;7:e33770. doi: 10.1371/journal.pone.0033770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ng P, Parks RJ, Cummings DT, Evelegh CM, Sankar U, Graham FL. A high-efficiency Cre/loxP-based system for construction of adenoviral vectors. Hum Gene Ther. 1999;10:2667–2672. doi: 10.1089/10430349950016708. [DOI] [PubMed] [Google Scholar]
- 16.Beck IMRR, Rückert R, Brandt K, Mueller MS, Sadowski T, Brauer R, Schirmacher P, Mentlein R, Sedlacek R. MMP19 is essential for T cell development and T cell-mediated cutaneous immune responses. PLoS ONE. 2008;3:e2343. doi: 10.1371/journal.pone.0002343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pandit KV, Corcoran D, Yousef H, Yarlagadda M, Tzouvelekis A, Gibson KF, Konishi K, Yousem SA, Singh M, Handley D, et al. Inhibition and role of let-7d in idiopathic pulmonary fibrosis. Am J Respir Crit Care Med. 2010;182:220–229. doi: 10.1164/rccm.200911-1698OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Akagi T, Shishido T, Murata K, Hanafusa H. v-Crk activates the phosphoinositide 3-kinase/AKT pathway in transformation. Proc Natl Acad Sci USA. 2000;97:7290–7295. doi: 10.1073/pnas.140210297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Yu G, Tseng GC, Yu YP, Gavel T, Nelson J, Wells A, Michalopoulos G, Kokkinakis D, Luo JH. CSR1 suppresses tumor growth and metastasis of prostate cancer. Am J Pathol. 2006;168:597–607. doi: 10.2353/ajpath.2006.050620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yu YP, Yu G, Tseng G, Cieply K, Nelson J, Defrances M, Zarnegar R, Michalopoulos G, Luo JH. Glutathione peroxidase 3, deleted or methylated in prostate cancer, suppresses prostate cancer growth and metastasis. Cancer Res. 2007;67:8043–8050. doi: 10.1158/0008-5472.CAN-07-0648. [DOI] [PubMed] [Google Scholar]
- 21.Shedden K, Taylor JM, Enkemann SA, Tsao MS, Yeatman TJ, Gerald WL, Eschrich S, Jurisica I, Giordano TJ, Misek DE, et al. Director’s Challenge Consortium for the Molecular Classification of Lung Adenocarcinoma. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat Med. 2008;14:822–827. doi: 10.1038/nm.1790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Bild AH, Yao G, Chang JT, Wang Q, Potti A, Chasse D, Joshi MB, Harpole D, Lancaster JM, Berchuck A, et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature. 2006;439:353–357. doi: 10.1038/nature04296. [DOI] [PubMed] [Google Scholar]
- 23.Therneau TMGP. New York: Springer; 2000. Modeling survival data: extending the Cox model. [Google Scholar]
- 24.Ihaka R, Gentleman R. R: a language for data analysis and graphics. J Comput Graph Stat. 1996;5:299–314. [Google Scholar]
- 25.Cardarella S, Johnson BE. The impact of genomic changes on treatment of lung cancer. Am J Respir Crit Care Med. 2013;188:770–775. doi: 10.1164/rccm.201305-0843PP. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sabe H. Cancer early dissemination: cancerous epithelial-mesenchymal transdifferentiation and transforming growth factor β signalling. J Biochem. 2011;149:633–639. doi: 10.1093/jb/mvr044. [DOI] [PubMed] [Google Scholar]
- 27.Minn AJ, Gupta GP, Siegel PM, Bos PD, Shu W, Giri DD, Viale A, Olshen AB, Gerald WL, Massagué J. Genes that mediate breast cancer metastasis to lung. Nature. 2005;436:518–524. doi: 10.1038/nature03799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zeisberg M, Neilson EG. Biomarkers for epithelial-mesenchymal transitions. J Clin Invest. 2009;119:1429–1437. doi: 10.1172/JCI36183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Invest. 2009;119:1420–1428. doi: 10.1172/JCI39104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Leppert D, Waubant E, Galardy R, Bunnett NW, Hauser SL. T cell gelatinases mediate basement membrane transmigration in vitro. J Immunol. 1995;154:4379–4389. [PubMed] [Google Scholar]
- 31.Albini A, Mirisola V, Pfeffer U. Metastasis signatures: genes regulating tumor-microenvironment interactions predict metastatic behavior. Cancer Metastasis Rev. 2008;27:75–83. doi: 10.1007/s10555-007-9111-x. [DOI] [PubMed] [Google Scholar]
- 32.Ramaswamy S, Ross KN, Lander ES, Golub TR. A molecular signature of metastasis in primary solid tumors. Nat Genet. 2003;33:49–54. doi: 10.1038/ng1060. [DOI] [PubMed] [Google Scholar]
- 33.Coate LE, John T, Tsao MS, Shepherd FA. Molecular predictive and prognostic markers in non-small-cell lung cancer. Lancet Oncol. 2009;10:1001–1010. doi: 10.1016/S1470-2045(09)70155-X. [DOI] [PubMed] [Google Scholar]
- 34.Singhal S, Vachani A, Antin-Ozerkis D, Kaiser LR, Albelda SM. Prognostic implications of cell cycle, apoptosis, and angiogenesis biomarkers in non-small cell lung cancer: a review. Clin Cancer Res. 2005;11:3974–3986. doi: 10.1158/1078-0432.CCR-04-2661. [DOI] [PubMed] [Google Scholar]
- 35.Boutros PC, Lau SK, Pintilie M, Liu N, Shepherd FA, Der SD, Tsao MS, Penn LZ, Jurisica I. Prognostic gene signatures for non-small-cell lung cancer. Proc Natl Acad Sci USA. 2009;106:2824–2828. doi: 10.1073/pnas.0809444106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shi W, Gerster K, Alajez NM, Tsang J, Waldron L, Pintilie M, Hui AB, Sykes J, P’ng C, Miller N, et al. MicroRNA-301 mediates proliferation and invasion in human breast cancer. Cancer Res. 2011;71:2926–2937. doi: 10.1158/0008-5472.CAN-10-3369. [DOI] [PubMed] [Google Scholar]
- 37.Eder M, Scherr M. MicroRNA and lung cancer. N Engl J Med. 2005;352:2446–2448. doi: 10.1056/NEJMcibr051201. [DOI] [PubMed] [Google Scholar]
- 38.Patnaik SK, Yendamuri S. Overexpression of microRNA miR-30a or miR-191 in A549 lung cancer or BEAS-2B normal lung cell lines does not alter phenotype. PLoS One. 2010;5:e9219. doi: 10.1371/journal.pone.0009219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB. Prediction of mammalian microRNA targets. Cell. 2003;115:787–798. doi: 10.1016/s0092-8674(03)01018-3. [DOI] [PubMed] [Google Scholar]
- 40.Kessenbrock K, Plaks V, Werb Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell. 2010;141:52–67. doi: 10.1016/j.cell.2010.03.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Delassus GS, Cho H, Eliceiri GL. New signaling pathways from cancer progression modulators to mRNA expression of matrix metalloproteinases in breast cancer cells. J Cell Physiol. 2011;226:3378–3384. doi: 10.1002/jcp.22694. [DOI] [PubMed] [Google Scholar]
- 42.Muller M, Beck IM, Gadesmann J, et al. MMP19 is upregulated during melanoma progression and increases invasion of melanoma cells. Modern Pathol. 2010;23:511–521. doi: 10.1038/modpathol.2009.183. [DOI] [PubMed] [Google Scholar]
- 43.Lettau I, Hattermann K, Held-Feindt J, Brauer R, Sedlacek R, Mentlein R. Matrix metalloproteinase-19 is highly expressed in astroglial tumors and promotes invasion of glioma cells. J Neuropathol Exp Neurol. 2010;69:215–223. doi: 10.1097/NEN.0b013e3181ce9f67. [DOI] [PubMed] [Google Scholar]
- 44.Zhao H, Yang Z, Wang X, Zhang X, Wang M, Wang Y, Mei Q, Wang Z. Triptolide inhibits ovarian cancer cell invasion by repression of matrix metalloproteinase 7 and 19 and upregulation of E-cadherin. Exp Mol Med. 2012;44:633–641. doi: 10.3858/emm.2012.44.11.072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Dehan E, Ben-Dor A, Liao W, Lipson D, Frimer H, Rienstein S, Simansky D, Krupsky M, Yaron P, Friedman E, et al. Chromosomal aberrations and gene expression profiles in non-small cell lung cancer. Lung Cancer. 2007;56:175–184. doi: 10.1016/j.lungcan.2006.12.010. [DOI] [PubMed] [Google Scholar]
- 46.Fang JH, Zhou HC, Zeng C, Yang J, Liu Y, Huang X, Zhang JP, Guan XY, Zhuang SM. MicroRNA-29b suppresses tumor angiogenesis, invasion, and metastasis by regulating matrix metalloproteinase 2 expression. Hepatology. 2011;54:1729–1740. doi: 10.1002/hep.24577. [DOI] [PubMed] [Google Scholar]
- 47.Zhang H, Qi M, Li S, Qi T, Mei H, Huang K, Zheng L, Tong Q. microRNA-9 targets matrix metalloproteinase 14 to inhibit invasion, metastasis, and angiogenesis of neuroblastoma cells. Mol Cancer Ther. 2012;11:1454–1466. doi: 10.1158/1535-7163.MCT-12-0001. [DOI] [PubMed] [Google Scholar]
- 48.Kass DJ, Kaminski N. Evolving genomic approaches to idiopathic pulmonary fibrosis: moving beyond genes. Clin Transl Sci. 2011;4:372–379. doi: 10.1111/j.1752-8062.2011.00287.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Gomperts BN, Walser TC, Spira A, Dubinett SM. Enriching the molecular definition of the airway “field of cancerization:” establishing new paradigms for the patient at risk for lung cancer. Cancer Prev Res (Phila) 2013;6:4–7. doi: 10.1158/1940-6207.CAPR-12-0470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kadara H, Shen L, Fujimoto J, Saintigny P, Chow CW, Lang W, Chu Z, Garcia M, Kabbout M, Fan YH, et al. Characterizing the molecular spatial and temporal field of injury in early-stage smoker non-small cell lung cancer patients after definitive surgery by expression profiling. Cancer Prev Res (Phila) 2013;6:8–17. doi: 10.1158/1940-6207.CAPR-12-0290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Pipinikas CP, Kiropoulos TS, Teixeira VH, Brown JM, Varanou A, Falzon M, Capitanio A, Bottoms SE, Carroll B, Navani N, et al. Cell migration leads to spatially distinct but clonally related airway cancer precursors. Thorax. 2014;69:548–557. doi: 10.1136/thoraxjnl-2013-204198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Stearman RS, Dwyer-Nield L, Grady MC, Malkinson AM, Geraci MW. A macrophage gene expression signature defines a field effect in the lung tumor microenvironment. Cancer Res. 2008;68:34–43. doi: 10.1158/0008-5472.CAN-07-0988. [DOI] [PubMed] [Google Scholar]
- 53.Bauvois B. New facets of matrix metalloproteinases MMP-2 and MMP-9 as cell surface transducers: outside-in signaling and relationship to tumor progression. Biochim Biophys Acta. 2012;1825:29–36. doi: 10.1016/j.bbcan.2011.10.001. [DOI] [PubMed] [Google Scholar]
- 54.Dufour A, Sampson NS, Zucker S, Cao J. Role of the hemopexin domain of matrix metalloproteinases in cell migration. J Cell Physiol. 2008;217:643–651. doi: 10.1002/jcp.21535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Dufour A, Sampson NS, Li J, Kuscu C, Rizzo RC, Deleon JL, Zhi J, Jaber N, Liu E, Zucker S, et al. Small-molecule anticancer compounds selectively target the hemopexin domain of matrix metalloproteinase-9. Cancer Res. 2011;71:4977–4988. doi: 10.1158/0008-5472.CAN-10-4552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Houghton AM, Hartzell WO, Robbins CS, Gomis-Rüth FX, Shapiro SD. Macrophage elastase kills bacteria within murine macrophages. Nature. 2009;460:637–641. doi: 10.1038/nature08181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Farago AF, Snyder EL, Jacks T. SnapShot: Lung cancer models. Cell. 2012;149:246–, e1. doi: 10.1016/j.cell.2012.03.015. [DOI] [PubMed] [Google Scholar]
- 58.Lau SK, Boutros PC, Pintilie M, Blackhall FH, Zhu CQ, Strumpf D, Johnston MR, Darling G, Keshavjee S, Waddell TK, et al. Three-gene prognostic classifier for early-stage non small-cell lung cancer. J Clin Oncol. 2007;25:5562–5569. doi: 10.1200/JCO.2007.12.0352. [DOI] [PubMed] [Google Scholar]
- 59.Boutros PC, Lau SK, Pintilie M, Liu N, Shepherd FA, Der SD, Tsao MS, Penn LZ, Jurisica I. Prognostic gene signatures for non-small-cell lung cancer. Proc Natl Acad Sci USA. 2009;106:2824–2828. doi: 10.1073/pnas.0809444106. [DOI] [PMC free article] [PubMed] [Google Scholar]