Abstract
The bHLH transcriptional factor BETA2/NeuroD1 is essential for the survival of photoreceptor cells in the retina. Although this gene is expressed throughout the retina, BETA2/NeuroD1 knockout mice show photoreceptor cell degeneration only in the outer nuclear layer of the retina; other retinal neurons are not affected. Previous studies on retina explants lacking three bHLH genes revealed that retinal neurons in the inner nuclear layer require multiple bHLH genes for their differentiation and survival. However, single- or double-gene mutations show no or a lesser degree of abnormalities during eye development, likely because of compensational or cooperative regulation among those genes. Because not all null mice survive until the retina is fully organized, no direct evidence of this concept has been reported. To understand the regulatory mechanisms between bHLH factors in retinal development, we performed a detailed analysis of BETA2/NeuroD1 knockout mice. BETA2/NeuroD1 was expressed in all three layers of the mouse retina, including all major types of neurons. In addition, a null mutation of BETA2/NeuroD1 resulted in up-regulation of other bHLH genes, Mash1, Neurogenin2, and Math3, in the inner nuclear layer. Our data suggest that compensatory and cross regulatory mechanisms exist among the bHLH factors during retinal development.
Keywords: BETA2/NeuroD1, Mash1, Math3, Neurogenin2, bHLH, Retina, Retinogenesis
Introduction
The retina is an excellent model system for studying the mechanism of neurogenesis because it is a relatively simple and laminated tissue. The retina is composed of distinct neuronal and glial cell populations arranged in three distinct layers: cone and rod photoreceptors are in the outer nuclear layer (ONL); bipolar cells, amacrine cells, horizontal cells, and Müller glial cells are in the inner nuclear layer (INL); and retinal ganglionic cells and displaced amacrine cells are in the ganglionic cell layer (GCL). Each cell type emerges from the retinal precursor population, and retinal precursor cell fate is influenced by both intrinsic and extrinsic factors (Cepko, 1999). Many in vivo and in vitro studies have suggested that basic helix-loop-helix (bHLH) transcription factors are intrinsic regulators for cell fate determination and differentiation (Cepko, 1999; Vetter and Brown, 2001; Hatakeyama and Kageyama, 2004; Akagi et al., 2004, Yan et al., 2005).
In developing mouse retina, bHLH genes such as Mash1, Neurogenin2, Math3, and BETA2/NeuroD1 are expressed in the retinal progenitor cells or differentiating retinal neurons (Tomita et al., 1996; Morrow et al., 1999; Pennesi et al., 2003; Ma and Wang, 2006), whereas bHLH gene Math5 is intensively expressed in retinal ganglionic cells and their progenitors (Brown et al., 2001; Wang et al., 2001; Mu et al., 2005). Although these genes are often expressed in overlapping patterns and play critical roles in cell fate determination and differentiation, the molecular mechanisms underlying the postnatal differentiation of the diverse types of retinal neurons are largely unknown. In attempts to elucidate these mechanisms, mutational analyses have been performed using homologous recombination or conditional knockout strategies (Tomita et al., 2000; Inoue et al., 2002; Akagi et al., 2004; Ma and Wang, 2006). However, targeted mutations in any of these bHLH genes lead to perinatal lethality or result in no obvious phenotypes in the retina (Tomita et al., 1996; Tomita et al., 2000; Ma and Wang, 2006). As examples, Mash1 knockout mice die soon after birth and do not show any defects in the retina (Tomita et al., 1996), and only about 40% of Math3-null mutants are viable, although little abnormality has been observed in surviving mice (Tomita et al., 2000). These results raise the possibility that other bHLH genes could be functionally redundant in the retina; that is, a single bHLH gene deletion may not be sufficient to disrupt the whole process of retinogenesis.
To determine whether multiple bHLH factors are required for the differentiation and survival of distinct cell types, double- and triple-gene knockout mice have been generated (Tomita et al., 2000; Inoue et al., 2002; Akagi et al., 2004). In those studies, retinal explant experiments were used to overcome perinatal lethality. However, retinal explants do not survive long postnatally, making it difficult to examine possible cross regulation among bHLH genes during the late stages of retinogenesis and age-related degenerative processes of the adult retina. Thus, a useful in vivo model of surviving mutants is required for more definitive analysis.
BETA2/NeuroD1 is known to be expressed in the CNS and PNS as early as E8.5 and its expression persists through adulthood (Cho and Tsai, 2004). BETA2/NeuroD1-null mice exhibit a severe loss of cone and rod photoreceptors (Pennesi et al., 2003). In wild-type mice, a high level of BETA2/NeuroD1 expression is observed in the outer half of the neuroblastic layer (NL) of the developing retina, and a lower expression level is observed in the developing INL around birth. A moderate expression level persists throughout the postnatal stages in the ONL and INL and remains at a stable level in the ONL of the adult retina (Morrow et al., 1999; Pennesi et al., 2003). In addition, in retinal explants, BETA2/NeuroD1 is detected in the precursors of various lineages and plays essential roles in the specification of many distinct neuronal cell types in cooperation with other bHLH genes (Akagi et al., 2004). Although expression of BETA2/NeuroD1 is detected in many neuronal lineages in the retina (Akagi et al., 2004), no other significant abnormalities have been found in the retina other than photoreceptor cell degeneration (Pennesi et al., 2003). It is possible that other bHLH factors may compensate for BETA2/NeuroD function in the differentiation and maintenance of these neurons. In this study, we characterized BETA2/NeuroD1 expression in all retinal neurons at various postnatal stages and found that the BETA2/NeuroD1 was expressed in all major cell types in the retina. Furthermore, BETA2/NeuroD1-null mutation led to compensatory up-regulation of Mash1 and Neurogenin2 in the INL.
Results
Expression of BETA2/NeuroD1 in the postnatal mouse retina
To address the importance of BETA2/NeuroD1 in postnatal retinal development, we examined its spatiotemporal expression by immunohistochemically analyzing the postnatal stages from P0 to P30. We had previously demonstrated that BETA2/NeuroD1 exhibits a dynamic expression pattern in the retina during embryonic stages (Pennesi et al., 2003). In that study, BETA2/NeuroD1 was expressed in the outer half of the neuroblastic layer (NL) and in a certain populations of cells in the developing INL and GCL (Morrow et al., 1999; Pennesi et al., 2003). In our current study, at P0 and P3, BETA2/NeuroD1 expression was observed in the outermost three to five layers of the NL, and a few faintly expressing retinal cells were also observed in the middle part of the NL as well as in the GCL (Figs. 1A and 1B). At P5, in addition to the strong immunoreactivity in the presumptive ONL where differentiating photoreceptors are localized, BETA2/NeuroD1 expression was observed in the outer half of the developing INL where differentiating horizontal, bipolar, and amacrine cells are localized (Fig. 1C). Notably, BETA2/NeuroD1 delineated the boundary between the outer and inner halves of the INL by its preferential expression. The expression patterns were persistently high in the ONL and outer half of the INL at P10 (Fig. 1D) and P15 (Fig. 1E) and remained at a stable level in the adult retina at P30 (Fig. 1F). In addition, punctate staining was observed in the developing inner half of the INL and GCL at all stages examined (Figs. 1C–E). This spatiotemporal expression pattern of BETA2/NeuroD1 in the INL during postnatal development raises the possibility that BETA2/NeuroD1 has functions in cell type specification in the INL, the GCL, or both.
Fig. 1.
Spatiotemporal expression of BETA2/NeuroD1 in the retina. Immunoreactivity of anti-NeuroD1 antibodies at P0 (A), P3 (B), P5 (C), P10 (D), P15 (E), and P30 (F). NL, neuroblastic layer; GCL, ganglionic cell layer; ONL, outer nuclear layer; INL, inner nuclear layer; IPL, inner plexiform layer; arrowheads, BETA2/NeuroD1-expressing neurons in the NL; arrows, BETA2/NeuroD1-expressing neurons in the INL and INL. Scale bar = 100 µm.
Characterization of BETA2/NeuroD1-expressing neurons in the retina
To define the subpopulations of BETA2/NeuroD1-expressing neurons in the retina, especially in the INL and GCL, we performed double immunofluorescence with a series of markers for retinal neurons and the anti-NeuroD1 antibodies on P15 wild-type retinas. The expression of BETA2/NeuroD1 was detected mainly in the photoreceptor cells in the ONL (Figs. 2A, 2D, and 2G), as previously reported (Pennesi et al., 2003). Immunofluorescence analysis revealed what appeared to be two different populations of cells labeled in the outer half and innermost layer of the INL and a few cells in the GCL. To determine their identity, we used antibodies against Calbindin for horizontal cells (Fig. 2B), protein kinase C α (PKCα) for rod bipolar cells (Fig. 2E), and Pax6 for amacrine cells (Fig. 2H). Anti-NeuroD1 antibody-positive cells included Rodopsin-expressing photoreceptor cells in the ONL (data not shown), Calbindinexpressing horizontal cells (Fig. 2C), PKCα-expressing bipolar cells in the outer half of the INL (Fig. 2F), and Pax6-expressing amacrine cells in the inner half of the INL and occasionally in the GCL (Fig. 2I). No Vimentin-expressing Müller glial cells were observed co-localizing with NeuroD1, as we expected (data not shown). The fact that BETA2/NeuroD1-immunopositive cells included major retinal neurons implies that BETA2/NeuroD1 expression is associated with a subpopulation of retinal neurons in the INL.
Fig. 2.
Characterization of BETA2/NeuroD1-expressing cells in the retina. Anti-NeuroD1 double immunostaining with Calbindin (top row), PKCα (middle row), and Pax6 (bottom row) on wild-type (BETA2+/+) retina at P15. ONL, outer nuclear layer; INL, inner nuclear layer; IPL, inner plexiform layer; GCL, ganglionic cell layer; arrows, double-positive cells; arrowheads, single-positive cells;. Scale bar = 50 µm.
Distribution of major neurons in the BETA2/NeuroD1-null retina
Since BETA2/NeuroD1 was detected in all cell types in the INL, we asked whether these cell types in BETA2/NeuroD1-null mutants are defective. To answer this question, we performed immunohistochemical analysis with specific markers for horizontal, bipolar, and amacrine cells. Cell counting revealed that no statistically significant differences were found in the Calbindin-expressing horizontal cells (Fig. 3A, top row), PKCα-expressing bipolar cells (Fig. 3A, middle row), or Pax6-expressing amacrine cells (Fig. 3A, bottom row). The lack of defects in bipolar cells is consistent with our earlier observation that there are no differences in the number of Goα -expressing cone bipolar cells and PKCα-expressing rod bipolar cells between wild-type and BETA2/NeuroD1 knockout mice (Pennesi et al., 2003). Quantification of the current results revealed that the number of neurons in the null mice was not significantly different in the horizontal (BETA2+/+: 8.3 ± 0.9 cells; BETA2−/−: 9.0 ± 1.5 cells/section in an area of 0.15 mm2; n = 3; P > 0.05), bipolar (BETA2+/+: 58.7 ± 3.8 cells; BETA2−/−: 57.5 ± 12.4 cells/section in an area of 0.15 mm2; n ≥ 3; P > 0.05), and amacrine cells in the INL (BETA2+/+: 152.3 ± 2.2 cells; BETA2−/−: 145.3 ± 6.1 cells /section in an area of 0.15 mm2; n = 3; P > 0.05) and in the GCL (BETA2+/+: 43.3 ± 1.5 cells; BETA2−/−: 52.3 ± 3.5 cells/section in an area of 0.15 mm2; n = 3; P > 0.05) (Fig. 3B). To provide a clear reference for the number of retinal neurons in this particular animal, we also counted the total number of retinal cells in the INL and GCL and compared the numbers between two genotypes. The total cell numbers in the INL (BETA2+/+: 424.0 ± 32.0 cells; BETA2−/−: 466.0 ± 28.9 cells/section in an area of 0.15 mm2) and GCL (BETA2+/+: 44.4 ± 2.6 cells; BETA2−/−: 48.3 ± 2.6 cells/section in an area of 0.15 mm2) were not significantly different between the two genotypes (n ≥ 3; P > 0.05) (Fig. 3C). We also examined the number of Vimentin-expressing Müller glial cells in the INL, which do not express BETA2/NeuroD1, and found that the BETA2/NeuroD1-null retinas were similar in all respects to littermate controls (BETA2+/+: 133.00 ± 5.86 cells/section; BETA2−/−: 125.33 ± 4.20 cells/section in an area of 0.15 mm2; n ≥ 3; P > 0.05). These results suggest that BETA2/NeuroD1-null mutation does not affect cell fate or the distribution of major neurons in the INL and GCL. In addition, the results raise the possibility that the normal morphologic characteristics of BETA2/NeuroD1-null mice might be due to functional compensation of closely related bHLH genes such as Mash1, Neurogenin2, and Math3.
Fig. 3.
Population of major retinal neurons in the inner nuclear layer (INL) and ganglionic cell layer (GCL) of the BETA2/NeuroD1-null retina. (A) Immunohistochemical analysis with Calbindin (top row), PKCα (middle row), and Pax6 (bottom row). Scale bar = 100 µm. (B) Quantification of antibody-positive cells in the INL and GCL. (C) Total number of neurons in the INL and GCL. For B and C, data represent the means ± S.E.M. Statistical significance (P value) was determined using Student’s t-test.
Compensation of bHLH factors in BETA2/NeuroD1-null retina
Cultured retinal explants have demonstrated that the subtype specification of retinal neurons requires the bHLH genes Mash1, Neurogenin2, Math3, and BETA2/NeuroD1 (Cepko, 1999; Inoue et al., 2002; Akagi et al., 2004). To explore the in vivo mechanisms underlying the regulatory interactions associated with BETA2/NeuroD1-related genes in the absence of BETA2/NeuroD1, we examined the expression patterns of Mash1, Neurogenin2, and Math3 on wild-type and BETA2/NeuroD1-null mice at P5, P10, and P15. At P5, expression of Mash1 and Neurogenin2 were detected in the ONL and the outer half of the INL (Figs. 4A–4B’). Math3-positive cells seemed to be localized in the inner half of the INL with no clear boundary (Figs. 4C and 4C’). No differences in immunoreactivity between the two genotypes were observed at this stage (Figs. 4A–4C’). By P10, in the absence of BETA2/NeuroD1, Mash1 and Neurogenin2 expression levels were up-regulated in the retina, especially in the outer half, where horizontal cells and bipolar cells were highly localized (Figs. 4D–4E’). Mash1 and Neurogenin2 expression were observed in the inner half of the INL and GCL, where amacrine cells are localized. However, the Math3 expression level was up-regulated slightly in BETA2/NeuroD1-null retina, although to a lesser extent than that observed for Mash1 and Neurogenin2 (Figs. 4F and 4F’). Similarly, at P15, Mash1 and Neurogenin2 expression levels were up-regulated in the BETA2/NeuroD1-null retina (Figs. 4G–4H’), but the Math3 expression level was not much increased in this immunohistochemical analysis compared to those of Mash1 and Neurogenin2 (Figs. 4I and 4I’). A similar expression pattern of these genes was observed at extended time points P30 and P40 (data not shown).
Fig. 4.
Up-regulation of Mash1 and Neurogenin2 expression levels in the BETA2/NeuroD1-null retina. Immunohistochemical analysis was performed with anti-Mash1, anti-Neurogenin2, and anti-Mash1 antibodies on wild-type (BETA2+/+) and knockout (BETA2−/−) mice at P5, P10, and P15. ONL, outer nuclear layer; INL, inner nuclear layer; GCL, ganglionic cell layer. Scale bar = 50 µm.
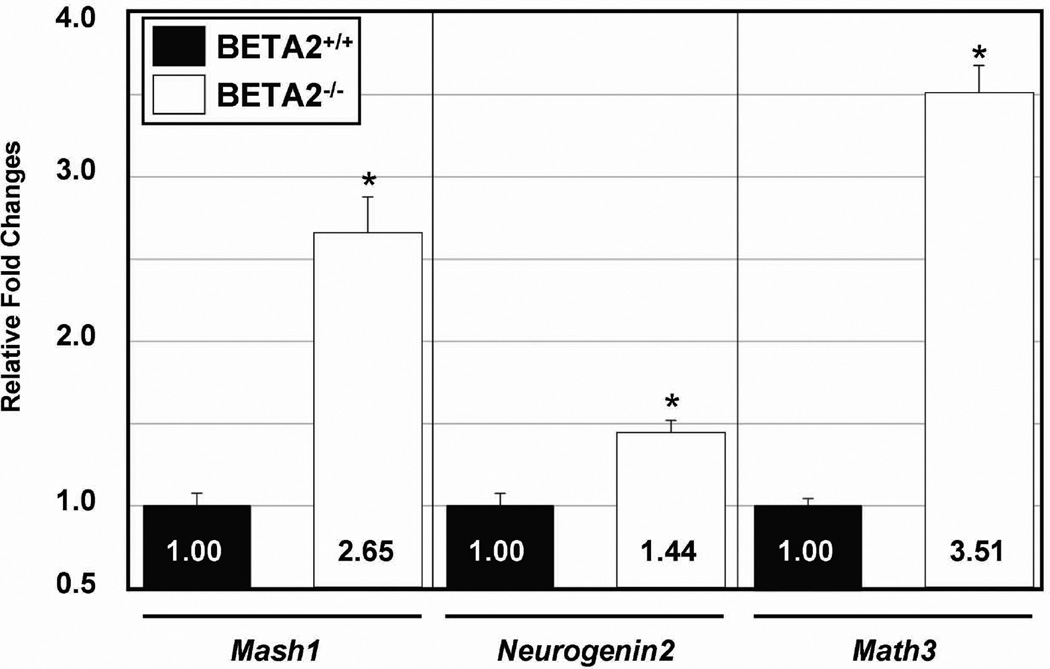
To validate our immunohistochemistry results (see Figure 4), we examined the expression of Mash1, Neurogenin2, and Math3 genes using quantitative RT-PCR (qRT-PCR). We extracted RNA from P15 wild-type littermate controls and BETA2/NeuroD1-null retinas and compared the levels of expression of those genes using four independent groups of samples for each genotype (Fig. 5). The expression levels were normalized with reference to β-actin. qRT-PCR analysis demonstrated a significant increase in the levels of all three transcripts in the BETA2/NeuroD1-null retina. Mash1 and Neurogene2 were significantly increased to 2.65 and 1.44 fold, respectively, in BETA2/NeuroD1-null retina compared to wild-type littermate controls (n=4, P<0.05) (Fig. 5). Also, Math3 expression was also increased to 3.34 fold in BETA2/NeuroD1-null retina (Fig 5). Taken together, these results suggest that to compensate for the loss of BETA2/NeuroD1, other bHLH genes are up-regulated in vivo.
Fig. 5.
Quantitative RT-PCR (qRT-PCR) analysis of Mash1, Neurogenin2, and Math3 genes in the BETA2/NeuroD1-null retina. qRT-PCR analyses were performed with retinas from four independent different groups of wild-type (BETA2+/+) and BETA2/NeuroD1-null (BETA2−/−). Data represent the mean ± S.D. from four independent experiments (*P < 0.05, Student's t test).
Discussion
In this study, we re-evaluated subpopulations of BETA2/NeuroD1-expressing neurons in the mouse retina and have found compensatory expression of other bHLH factors in the INL, where no obvious morphologic defects were observed. Our results suggest that BETA2/NeuroD1 knockout mice are useful for studying compensational and cross regulation among bHLH genes.
Subpopulation of BETA2/NeuroD1-expressing cells in the retina
In a previous report on a BETA2/NeuroD1-LacZ knockin allele, we performed β-galactosidase assays to examine the expression pattern of BETA2/NeuroD1 in the retina and observed a drastic change in LacZ gene expression in the outer half of the developing retina and in the ONL of postnatal and adult retina (Pennesi et al., 2003). However, no detailed analysis was performed in those experiments. In the current study, we adopted immunohistochemical techniques using anti-NeuroD1 antibodies. The sensitivity and specificity of the BETA2/NeuroD1 antibodies allowed us to explore retinal cell fate specification in more detail (Chu and Tsai, 2005; Cho and Tsai, 2006).
BETA2/NeuroD1 expression in the retina has been observed in many species (Acharya et al., 1997; Roztocil et al., 1997; Pennesi et al., 2003). On the basis of its spatiotemporal expression pattern, BETA2/NeuroD1 is highly expressed in photoreceptor cells and its precursor population during the embryonic stage. In addition, a lower level of expression has been observed in amacrine cells (Morrow et al., 1999; Inoue et al., 2002; Pennesi et al., 2003). Mice lacking the bHLH genes Neurogenin2, Math3, and BETA2/NeuroD1 exhibited a severe loss of horizontal and bipolar cell populations, and mice lacking Math1, Math3, and BETA2/NeuroD1 had fewer photoreceptor cells and more retinal ganglionic cells (Akagi et al., 2004). These observations led us to investigate BETA2/NeuroD1 expression in various cell types in the retina other than photoreceptors and amacrine cells. Although BETA2/NeuroD1 mutants did not show any changes in horizontal, bipolar, and amacrine cells, we found specific expression of BETA2/NeuroD1 not only in amacrine cells but also in horizontal and bipolar cells (Fig. 2). This result indicates that BETA2/NeuroD1 may not be a dominant regulator for the genesis of these three types of cells. However, the question remains as to whether the low expression level of BETA2/NeuroD1 has functional consequences in the production of a subset of retinal neurons in the INL.
Importance of the bHLH compensatory regulation in the retina
During postnatal development, BETA2/NeuroD1 expression is observed mainly in photoreceptor cells in the ONL (Pennesi et al., 2003). However, moderate levels of expression remain in the outer half and innermost layer of the INL of the retina as well as in a certain population of cells in the GCL (Figs. 1 and 4). Therefore, we could not rule out the possibility that BETA2/NeuroD1 has functions in cell type specification in the INL together with other bHLH genes. Both gain-of-function and loss-of-function studies have demonstrated that BETA2/NeuroD1 participates in the neuron/glia cell fate decisions, similar to other bHLH genes, including Mash1, Math3, and Math5 in retinal explants (Brown et al., 1998; Morrow et al., 1999; Tomita et al., 2000). Thus, we attempted to identify differences in the population of major cell types in BETA2/NeuroD1-null retina compared with wild-type littermate retina. However, we found no differences (Fig. 3). This result may be due to compensational regulation by other bHLH genes, such as Mash1, Math3, Neurogenin2, and Math5.
Mash1, Math3, and Neurogenin2 are known to be expressed in the developing retina and act as positive regulators (Guillemot and Joyner, 1993; Gradwohl et al., 1996; Sommer et al., 1996; Morrow et al., 1999; Tomita et al., 2000). Together with homeodomain factors such as Pax6 and Crx, these factors play important roles in cell type specification during early development (Marquardt et al., 2001; Hatakeyama and Kageyama, 2004). For example, Mash1 and Math3 are expressed predominantly in bipolar cells, and double knockouts of these genes decreases the bipolar cell population while increasing the Müller glial cell population (Tomita et al., 2000). Neurogenin2 is also transiently expressed in all major neuron types in the mouse retina, and its expression is required for photoreceptor cells, horizontal cells, and bipolar cells (Akagi et al., 2004; Ma and Wang, 2006). In contrast, BETA2/NeuroD1 is transiently expressed in differentiating amacrine cells (Fig. 2I and Morrow et al., 1999; Pennesi et al., 2003). Although BETA2/NeuroD1-null mutation shows delayed amacrine cell development at earlier stages, the number of amacrine cells eventually is the same as that found in wildtype retinas (Fig. 3B and Morrow et al., 1999). However, in double-knockout mutations with BETA2/NeuroD1 and Math3, the number of amacrine cells is decreased and that of retinal ganglionic cells is increased. Interestingly, amacrine cells adopt the ganglion cells’ fate in this BETA2/NeuroD1;Math3 double-knockout mutant (Inoue et al., 2002). In addition, the triple bHLH knockouts Mash1;Neurogenin2;Math3 and Math3;Neurogenin2; BETA2/NeuroD1 have been shown to have fewer horizontal cells, but any combination of double mutations of Mash1 or Math3 or BETA2/NeuroD1 with Neurogenin2 display abnormalities in retina development (Akagi et al., 2004). Furthermore, single-knockout mutations of the genes barely affect the neuronal cell population in the INL and show no retinal abnormalities. Taken together, these results suggest that the bHLH factors cross regulate each others’ expression and can specify neuronal subtypes cooperatively during late retinogenesis, especially in progenitor cells in the INL, to generate various subtypes of retinal neurons. Although the precise mechanism for retina cell type specification remains to be determined, our results provide further support for cooperative and compensational regulatory specification during postnatal retinogenesis.
Experimental Procedures
Animals
BETA2/NeuroD1 knockout mice were generated in the 129SvJ background as described previously (Liu et al., 2000). All genotypes described were confirmed by polymerase chain reaction (Naya et al., 1997). The animals were handled in accordance with the policies on the treatment of laboratory animals of The University of Texas M. D. Anderson Cancer Center and the Baylor College of Medicine.
Immunohistochemical analysis
12 µm cryostat sections were obtained from 10% buffered formalin-fixed eye samples from postnatal day 5 (P5) to P30. Endogenous peroxidase activity and nonspecific antibody binding were blocked with 3% hydrogen peroxide (H2O2) in methanol for 25 min and 3% normal serum with 3% bovine serum albumin (BSA) in phosphate-buffered saline (PBS, pH 7.2–7.4) for 1 hr. Goat anti-NeuroD1 antibodies (N-19 and G-20), rabbit anti-Math3 antibodies (Santa Cruz Biotechnology, Santa Cruz, CA) were used at 1:500 dilution. Rabbit anti-Mash1 antibody (Chemicon, Temecula, CA) was used at 1:100 dilution. Rabbit anti-Neurogenin2 (Chemicon), affinity-purified rabbit anti-Calbindin-D-28K (Sigma, St. Louis, MO), anti-Pax6 (Chemicon), and anti- PKCα (Sigma) antibodies were used at 1:200 dilution. All antibodies were diluted in 3% normal serum in PBS containing 3% BSA, and samples were incubated with antibodies at 4°C for 16 hr. Immunoreactivity was visualized using an ABC kit (Vector Laboratories, Burlingame, CA) for colorimetric visualization (brown) and counterstained with methyl green (green). For immunofluorescence staining, donkey cy3-conjugated (red) anti-rabbit and donkey fluorescein isothiocyanate-conjugated (green) anti-goat antibodies were used at 1:400 dilutions (Molecular Probes, Eugene, OR).
Cell number counting
The total number of antibody-positive cells in a 0.15-mm2 area was counted from at least three independently prepared retinas at P15. Mean ± S.E.M. values were evaluated between wild-type (BETA2+/+) and BETA2/NeuroD1 knockout (BETA2−/−) animal groups, and significance was determined by Student’s t-test. A P value < 0.05 was considered statistically significant (a 95% confidence level).
Quantitative RT-PCR analysis
Two micrograms of total RNA from P15 retina of wild-type (BETA2+/+) or BETA2/NeuroD1 knockout (BETA2−/−) mice was reserved transcribed with Superscript reverse transcriptase II (Invitrogene, Carlsbad, CA). cDNA equivalent to 40ng was used for qPCR amplification (Applied Biosystems, Foster City, CA) using SYBR green master mix (Applied Biosystems). The relative expression levels were normalized to those of β-actin. qRT-PCR data were calculated using the comparative Ct method (Applied Biosystems). Standard deviations from the mean of the [Δ]Ct values were calculated from four independent groups of cDNA samples. The PCR primers used in this experiment are listed in Table 1.
Table 1.
Oligonucleotide primers used for quantitative RT-PCR amplification
| Genes | Forward primer | Reverse primer | Annealing temperature |
|---|---|---|---|
| Mash1 | ACTTGAACTCTATGGCGGGTT | CCAGTTGGTAAAGTCCAGCAG | 56°C |
| Neurogene2 | AACTCCACGTCCCCATACAG | GAGGCGCATAACGATGCTTCT | 56°C |
| Math3 | AGCTGGTCAACACACAATCCT | GTTCCGAGCATTCCATAAGAGC | 56°C |
| β-actin | CAATAGTGATGACCTGGCCGT | AGAGGGAAATCGTGCGTGAC | 56°C |
Photography
Micrographs were photographed with a digital camera (Axiovision 3.1) mounted on an Axioskop2 microscope (Carl Zeiss, Thornwood, NY) or a Confocal laser scanning microscope (Olympus Fluoview laser microscope, Olympus Optical Co., Japan). Photographs were assembled in Adobe Photoshop version CS2 (Adobe Systems, San Jose, CA). Contrast, brightness, and color balance were adjusted to obtain optimum photographs.
Acknowledgments
We thank Drs. Sophia Y. Tsai and Debra E. Bramblett for helpful discussions and critical reading of this manuscript. We also thank Eric Meadows for confocal microscopy and qRT-PCR analysis, Roberto Carlos Reynaga, Wen Chen, and Xue-Fei Tong for excellent technical help. This work is supported by NIH/NEI grant EY011930 (WHK), Robert A. Welch Foundation grant G-0010 (WHK), and NIH grant HD17379 (MJT).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Acharya HR, Dooley CM, Thoreson WB, Ahmad I. cDNA cloning and expression analysis of NeuroD mRNA in human retina. Biochem. Biophys. Res. Commun. 1997;233:459–463. doi: 10.1006/bbrc.1997.6483. [DOI] [PubMed] [Google Scholar]
- Akagi T, Inoue T, Miyoshi G, Bessho Y, Takahashi M, Lee JE, Guillemot F, Kageyama R. Requirement of multiple basic helix-loop-helix genes for retinal neuronal subtype specification. J Biol Chem. 2004;279:28492–28498. doi: 10.1074/jbc.M400871200. [DOI] [PubMed] [Google Scholar]
- Brown NL, Kanekar S, Vetter ML, Tucker PK, Gemza DL, Glaser T. Math5 encodes a murine basic helix-loop-helix transcription factor expressed during early stages of retinal neurogenesis. Development. 1998;125:4821–4833. doi: 10.1242/dev.125.23.4821. [DOI] [PubMed] [Google Scholar]
- Brown NL, Patel S, Brzezinski J, Glaser T. Math5 is required for retinal ganglion cell and optic nerve formation. Development. 2001;128:2497–2508. doi: 10.1242/dev.128.13.2497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cepko CL. The roles of intrinsic and extrinsic cues and bHLH genes in the determination of retinal cell fates. Curr Opin Neurobiol. 1999;9:37–46. doi: 10.1016/s0959-4388(99)80005-1. [DOI] [PubMed] [Google Scholar]
- Cho JH, Tsai MJ. The role of BETA2/NeuroD1 in the development of the nervous system. Mol. Neurobiol. 2004;30:35–47. doi: 10.1385/MN:30:1:035. [DOI] [PubMed] [Google Scholar]
- Cho JH, Tsai MJ. Preferential posterior cerebellum defect in BETA2/NeuroD1 knockout mice is the result of differential expression of BETA2/NeuroD1 along anterior-posterior axis. Dev Biol. 2006;290:125–138. doi: 10.1016/j.ydbio.2005.11.024. [DOI] [PubMed] [Google Scholar]
- Chu K, Tsai MJ. Neuronatin, a downstream target of BETA2/NeuroD1 in the pancreas, is involved in glucose-mediated insulin secretion. Diabetes. 2005;54:1064–1073. doi: 10.2337/diabetes.54.4.1064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gradwohl G, Fode C, Guillemot F. Restricted expression of a novel murine atonal-related bHLH protein in undifferentiated neural precursors. Dev Biol. 1996;180:227–241. doi: 10.1006/dbio.1996.0297. [DOI] [PubMed] [Google Scholar]
- Guillemot F, Joyner AL. Dynamic expression of the murine Achaete-Scute homologue Mash-1 in the developing nervous system. Mech Dev. 1993;42:171–185. doi: 10.1016/0925-4773(93)90006-j. [DOI] [PubMed] [Google Scholar]
- Hatakeyama J, Kageyama R. Retinal cell fate determination and bHLH factors. Semin Cell Dev Biol. 2004;15:83–89. doi: 10.1016/j.semcdb.2003.09.005. [DOI] [PubMed] [Google Scholar]
- Inoue T, Hojo M, Bessho Y, Tano Y, Lee JE, Kageyama R. Math3 and NeuroD regulate amacrine cell fate specification in the retina. Development. 2002;129:831–842. doi: 10.1242/dev.129.4.831. [DOI] [PubMed] [Google Scholar]
- Liu M, Pleasure SJ, Collins AE, Noebels JL, Naya FJ, Tsai MJ, Lowenstein DH. Loss of BETA2/NeuroD leads to malformation of the dentate gyrus and epilepsy. Proc Natl Acad Sci U S A. 2000;97:865–870. doi: 10.1073/pnas.97.2.865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma W, Wang SZ. The final fates of neurogenin2-expressing cells include all major neuron types in the mouse retina. Mol Cell Neurosci. 2006;31:463–469. doi: 10.1016/j.mcn.2005.10.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marquardt T, Ashery-Padan R, Andrejewski N, Scardigli R, Guillemot F, Gruss P. Pax6 is required for the multipotent state of retinal progenitor cells. Cell. 2001;105:43–55. doi: 10.1016/s0092-8674(01)00295-1. [DOI] [PubMed] [Google Scholar]
- Morrow EM, Furukawa T, Lee JE, Cepko CL. NeuroD regulates multiple functions in the developing neural retina in rodent. Development. 1999;126:23–36. doi: 10.1242/dev.126.1.23. [DOI] [PubMed] [Google Scholar]
- Mu X, Fu X, Sun H, Beremand PD, Thomas TL, Klein WH. A gene network downstream of transcription factor Math5 regulates retinal progenitor cell competence and ganglion cell fate. Dev Biol. 2005;280:467–481. doi: 10.1016/j.ydbio.2005.01.028. [DOI] [PubMed] [Google Scholar]
- Naya FJ, Huang HP, Qiu Y, Mutoh H, DeMayo FJ, Leiter AB, Tsai MJ. Diabetes, defective pancreatic morphogenesis, and abnormal enteroendocrine differentiation in BETA2/neuroD-deficient mice. Genes Dev. 1997;11:2323–2334. doi: 10.1101/gad.11.18.2323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pennesi ME, Cho JH, Yang Z, Wu SH, Zhang J, Wu SM, Tsai MJ. BETA2/NeuroD1 null mice: a new model for transcription factor-dependent photoreceptor degeneration. J Neurosci. 2003;23:453–461. doi: 10.1523/JNEUROSCI.23-02-00453.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roztocil T, Matter-Sadzinski L, Alliod C, Ballivet M, Matter JM. NeuroM, a neural helix-loop-helix transcription factor, defines a new transition stage in neurogenesis. Development. 1997;124:3263–3272. doi: 10.1242/dev.124.17.3263. [DOI] [PubMed] [Google Scholar]
- Sommer L, Ma Q, Anderson DJ. neurogenins, a novel family of atonal-related bHLH transcription factors, are putative mammalian neuronal determination genes that reveal progenitor cell heterogeneity in the developing CNS and PNS. Mol Cell Neurosci. 1996;8:221–241. doi: 10.1006/mcne.1996.0060. [DOI] [PubMed] [Google Scholar]
- Tomita K, Moriyoshi K, Nakanishi S, Guillemot F, Kageyama R. Mammalian achaete-scute and atonal homologs regulate neuronal versus glial fate determination in the central nervous system. EMBO J. 2000;19(20):5460–5472. doi: 10.1093/emboj/19.20.5460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tomita K, Nakanishi S, Guillemot F, Kageyama R. Mash1 promotes neuronal differentiation in the retina. Genes Cells. 1996;1(8):765–774. doi: 10.1111/j.1365-2443.1996.tb00016.x. [DOI] [PubMed] [Google Scholar]
- Vetter ML, Brown NL. The role of basic helix-loop-helix genes in vertebrate retinogenesis. Semin Cell Dev Biol. 2001;12(6):491–498. doi: 10.1006/scdb.2001.0273. [DOI] [PubMed] [Google Scholar]
- Wang SW, Kim BS, Ding K, Wang H, Sun D, Johnson RL, Klein WH, Gan L. Requirement for math5 in the development of retinal ganglion cells. Genes Dev. 2001;15:24–29. doi: 10.1101/gad.855301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan RT, Ma W, Liang L, Wang SZ. bHLH genes and retinal cell fate specification. Mol Neurobiol. 2005;32(2):157–171. doi: 10.1385/MN:32:2:157. [DOI] [PMC free article] [PubMed] [Google Scholar]