ABSTRACT
After viral fusion with the cell membrane, the conical capsid of HIV-1 disassembles by a process called uncoating. We recently utilized the cyclosporine (CsA) washout assay, in which TRIM-CypA-mediated restriction of viral replication is used to detect the state of the viral capsid, to study the kinetics of uncoating in HIV-1-infected cells. Here we have extended this analysis to examine the effects of p24 capsid protein (p24CA) mutations and cellular environment on the kinetics of uncoating in infected cells. We found that p24CA mutations can significantly increase (A92E), delay (E45A and N74D), or have no effect (G94D) on the rate of uncoating and that these alterations are not due to changes in reverse transcription. Inhibition of reverse transcription delayed uncoating kinetics to an extent similar to that of the wild-type virus with all the p24CA mutant viruses tested. In addition, we observed differences in uncoating in two cell lines, which suggests that the cellular environment can differentially impact the disassembly of wild-type and mutant capsids. Collectively, these experiments suggest that viral and cellular factors are important for the process of uncoating. Finally, these data support the model whereby early steps in reverse transcription facilitate HIV-1 uncoating.
IMPORTANCE The HIV-1 capsid is a cone-shaped structure, composed of the HIV-1-encoded protein p24CA, which contains the viral RNA and other proteins needed for infection. After the virus enters a target cell, this capsid must disassemble by a process called uncoating. Uncoating is required for HIV-1 infection to progress, but the details of how this process occurs is not known. In this study, we used an in vivo assay to examine the uncoating process in HIV-1-infected cells. We determined that p24CA mutations could increase or decrease the rate of uncoating and that this rate varied in different cell lines. We also found that reverse transcription of the viral RNA altered the process of uncoating before the p24CA mutations. Collectively, these experiments provide a better understanding of how viral and cellular factors are involved with a poorly understood step in HIV-1 infection.
INTRODUCTION
After the HIV-1 membrane fuses with the target cell membrane, a viral complex is released into the cytoplasm of the cell. In this initial complex, the viral RNAs and associated proteins are enclosed by a cone-shaped capsid. This capsid is composed of monomers of the viral p24 capsid protein (p24CA) arranged in a hexameric lattice. At some point, the capsid must disassemble by a process called uncoating to release the reverse transcribing viral genome to integrate into the host cell DNA. Where, when, and how the viral capsid dissociates is poorly defined and a source of contention in the field. While it is clear that uncoating is required for HIV-1 replication, many questions remain about the viral and cellular factors involved with the process and its impact on subsequent steps in viral replication.
Two viral factors that have been shown to influence uncoating are the p24CA protein and the process of reverse transcription. Mutations in p24CA can alter capsid stability and decrease infectivity, indicating that overall capsid stability is important for optimal viral replication (1–5). In addition, the correct timing of uncoating is thought to be required for viral replication, as p24CA mutants that uncoat more rapidly and mutants that uncoat more slowly than wild-type virus in vitro both decrease infectivity (1). As many of these p24CA mutants with altered capsid stability also displayed defects in reverse transcription, initially it was thought that uncoating preceded reverse transcription (1). However, reverse transcription products can be detected in viral complexes that contain p24CA protein in the cytoplasm of infected cells (6). Furthermore, integration-competent preintegration complexes (PICs) can be generated within intact capsids when infection is restricted by TRIM5 alpha in the presence of proteasome inhibitors (7). Finally, treatment with the reverse transcriptase inhibitor nevirapine delays uncoating in HIV-1-infected cells, indicating that reverse transcription facilitates the process of uncoating (8, 9). Collectively, these data suggest that there is a complex interplay between these two essential steps in viral replication.
Recently, we developed an assay to study uncoating kinetics in HIV-1-infected cells, which is based on experiments performed by Perez-Caballero et al. (10) to characterize the restriction factor TRIM-CypA (cyclophilin A) (8, 11). In the cyclosporine (CsA) washout assay, TRIM-CypA is used to detect the presence of intact capsids in infected cells and inhibit their infectivity at various times postinfection. Using this assay, we found that the half-life of uncoating occurs within an hour of viral fusion and that reverse transcription facilitates the process of uncoating (8). The CsA washout assay is indirect because it measures viral susceptibility to TRIM-CypA restriction, which is mediated by the interaction of TRIM-CypA with a hexameric array of p24CA. However, a similar timing and effect of reverse transcription on uncoating have been confirmed by fluorescence microscopy and biochemical assays that directly detect the loss of p24CA (8, 9, 12, 13). Therefore, while indirect, the CsA washout assay is a good method to monitor the process of uncoating in infected cells.
Here we have extended our initial analysis to examine the effects of p24CA mutations on the kinetics of uncoating. We chose p24CA mutations (E45A, N74D, G89V, P90A, A92E, and G94) that are currently actively studied due to their effects on other aspects of HIV replication, including infection of nondividing cells, utilization of nuclear import pathways, and integration site selection (13–19). Often the effect of these mutations on other steps in viral replication has been proposed to be due to changes in uncoating. However, these mutations may also alter the ability of p24CA to interact with host factors that facilitate these subsequent steps in infection. While some of these mutants have been studied in in vitro uncoating assays, it is important to characterize uncoating in HIV-1-infected cells to differentiate between these two possibilities. We also test whether the cellular environment can differentially impact the uncoating of wild-type and mutant viruses. Finally, we determine the relative contributions of reverse transcription and these p24CA mutations on the process of uncoating.
MATERIALS AND METHODS
Cell lines, viruses, and pharmaceuticals.
HeLa and 293T cells were obtained from the American Type Culture Collection. Owl monkey kidney (OMK) cells were a gift from P. Bieniasz. The stable cell line HeLa-TC was made using a TRIM-CypA-expressing retroviral vector. Cells were maintained at 37°C and 5% CO2 and cultured in Dulbecco's modified Eagle medium (Cellgro), 10% fetal bovine serum (Gibco), and 1% penicillin-streptomycin-glutamine (Gibco). Vesicular stomatitis virus g protein (VSV-g) pseudotyped virus was produced by poly(ethylenimine) (PEI) transfection of 293T cells with 12 μg HIV-1 proviral plasmid HIV-GFP or p24CA mutants and 8 μg VSV-g expression plasmid. Virus was harvested 48 h posttransfection, purified through a 0.45-μm filter, and stored at −80°C until use. Cyclosporine (CsA) (Calbiochem) was prepared in ethanol and used at a final concentration of 2.5 μM. Nevirapine (NIH AIDS Research and Reference Reagent Program) was prepared in dimethyl sulfoxide (DMSO) and used at a final concentration of 5 μM. Ammonium chloride was prepared in double-distilled water (ddH2O) and used at a final concentration of 2.5 μM.
CsA washout assay.
The CsA washout assay has been previously described (8, 11). Briefly, OMK cells, HeLa-TC cells, or the parent HeLa cell line were plated in 96-well dishes with each experimental or control reaction performed on triplicate wells. Cells were spinoculated with VSV-g pseudotyped HIV-GFP or p24CA mutant virus in the presence of CsA and 10 μg/ml Polybrene for 1.5 h at 16°C. The inoculation medium was then exchanged for warm medium, and CsA was washed out of the zero time point reaction by medium exchange. Washout continued at various times postinfection. The negative control was ethanol washout. Two days postinfection, the cells were harvested with 100 μl trypsin and fixed by the addition of 100 μl fix (4:1, 1× phosphate-buffered saline [PBS]–10% formaldehyde). The percentage of green fluorescent protein (GFP)-positive cells was determined by flow cytometry using the Accuri C6 flow cytometer or BD LSR Fortessa analyzer, averaged for each triplicate reaction, and standard error was calculated. For experiments in HeLa-TC cells, the percentage of infected cells in the ethanol control was subtracted from the CsA reaction at each washout time point to determine the percentage of infected cells over background. The half-life of uncoating was determined by a best-fit line through the two data points flanking 50% uncoating, and times were averaged from multiple independent experiments. The average half-life of uncoating was compared between wild-type and mutant viruses using a Student's t test.
Nevirapine addition assay and viral fusion assay.
OMK cells were plated and spinoculated with HIV-GFP or p24CA mutant virus similar to the CsA washout assay. When inoculation medium was exchanged for warm medium, medium containing drug (nevirapine or ammonium chloride) and CsA was added to the zero time point reaction. Drug addition continued in the same manner for various times postinfection. Controls included no treatment and continuous treatment with drug. Cells were harvested, and the percentage of GFP-positive cells was determined as in the CsA washout assay. For the nevirapine addition assay, the data were normalized by setting the value for the GFP-positive cells in the DMSO carrier control (no nevirapine treatment) to 100%. The percentage of GFP-positive cells at the 6-h addition time point was compared between wild-type and mutant viruses using a Student's t test. For the viral fusion assay, the half-life of fusion was determined using a best-fit line between the adjacent data points, and times were averaged from three independent experiments.
CsA addition assay.
The parent HeLa cell line used to make HeLa-TC cells was plated and spinoculated with HIV-GFP or N74D mutant virus similar to the CsA washout assay. When inoculation medium was exchanged for warm medium, medium containing CsA or the carrier control ethanol was added to the zero time point reaction. CsA or ethanol addition continued in the same manner for various times postinfection. Controls included no CsA treatment and continuous CsA treatment. Cells were harvested, and the percentage of GFP-positive cells was determined as in the CsA washout assay.
RESULTS
Mutations in p24CA affect uncoating kinetics in cultured cells.
In order to determine the effects of mutations in p24CA on the process of uncoating in HIV-1-infected cells, we performed the CsA washout assay with six previously described p24CA mutant viruses (8, 11). In this assay, uncoating is detected by the action of the cellular protein TRIM-CypA which binds to the viral capsid to restrict infection (20–22). TRIM-CypA belongs to the tripartite motif family (TRIM) family of proteins that can block viral infection (23, 24). While the exact mechanism is not known, TRIM proteins restrict retroviral infection by binding the incoming viral capsid and then either induce degradation of the capsid to block reverse transcription or block nuclear import (7, 25, 26). While it may sound counterintuitive to use a protein that may induce uncoating to study uncoating, the CsA washout assay is based on the binding of TRIM-CypA to the viral capsid and not the mechanism by which TRIM proteins disrupt the capsid. There are two characteristics of TRIM-CypA binding that allow this factor to be used in an uncoating assay. First, TRIM-CypA, like all TRIM proteins, binds to the hexameric array of the p24CA protein in the assembled HIV-1 capsid (20–22, 27). Therefore, TRIM-CypA should detect only coated viral complexes in cells. Second, the drug cyclosporine (CsA) prevents the binding of TRIM-CypA to the viral capsid in a reversible fashion (10, 20, 28, 29). Therefore, when CsA is removed, TRIM-CypA can bind any coated viral complexes and restrict infection. In contrast, any virus that has uncoated sufficiently to lack an array of p24CA protein capable of binding TRIM-CypA will be able to escape restriction and infect the cell. These two characteristics of TRIM-CypA binding allow the kinetics of HIV-1 uncoating to be monitored in infected cells using timed withdrawal of CsA (8, 10, 11).
For the CsA washout assay, owl monkey kidney (OMK) cells that endogenously express TRIM-CypA were synchronously infected with wild-type virus (HIV-GFP) or p24CA mutant GFP reporter virus in the presence of CsA. CsA was then washed out at various times postinfection by medium exchange. The cells were harvested 2 days postinfection, and flow cytometry was used to determine the percentage of infected cells at each time point, which is indicative of the percentage of uncoated virions. The data were normalized by setting the percentage of GFP-positive cells where the curve levels off to 100%, allowing a half-life of uncoating to be calculated. The uncoating kinetics of each p24CA mutant was examined in parallel with the wild-type control HIV-GFP. Most of these mutations decrease HIV infectivity, so differing dilutions of each virus were used to achieve a total infectivity of 30 to 40% in order to optimally compare uncoating kinetics.
We first examined G89V and P90A mutant viruses. These point mutations are located in the cyclophilin A (CypA) binding loop and disrupt the ability of TRIM-CypA to bind the capsid (30). Indeed, when the washout assay was performed with these mutants, we did not observe an increase in the percentage of GFP-positive cells over time, unlike the HIV-GFP positive control (Fig. 1A). In addition, there is little difference between the CsA washout curve and the carrier control ethanol (EtOH) washout curve for both mutants. Therefore, these two mutants demonstrate that changes in the percentage of GFP-positive cells over time in the CsA washout assay are dependent on the ability of TRIM-CypA to bind the viral capsid.
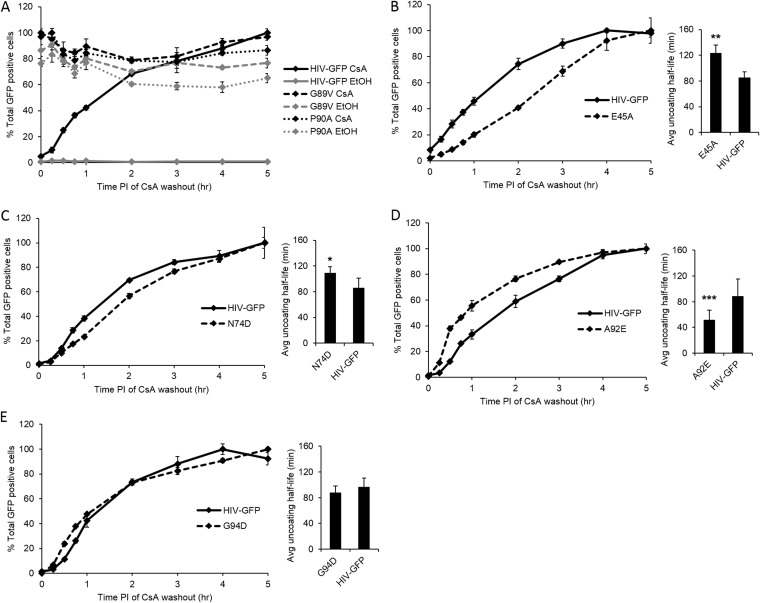
FIG 1.
p24CA mutations affect the observed rate of uncoating. The CsA washout assay was performed with HIV-GFP and several p24CA mutant viruses. Line graphs are from a representative washout assay. The time postinfection of CsA washout assay (in hours) is shown on the x axes of the line graphs. Bar graphs compare the average half-life of uncoating between HIV-GFP and the p24CA mutant virus from multiple independent experiments. Error bars denote standard errors. Values that are significantly different are indicated as follows: *, P < 0.05; **, P < 0.005; ***, P < 0.0001. (A) Uncoating cannot be observed with G89V and P90A mutant viruses, as the mutations make capsid resistant to CypA binding. (B) The E45A mutation significantly decreases the rate of uncoating compared to the wild type as measured in 9 independent experiments. (C) The N74D mutation significantly decreases the rate of uncoating compared to the wild type as measured in 6 independent experiments. (D) The A92E mutation significantly increases the rate of uncoating compared to the wild type as measured in 7 independent experiments. (E) The G94D mutation does not significantly alter the rate of uncoating compared to the wild type as measured in 7 independent experiments.
Next, we tested the E45A and N74D p24CA mutants, which have been shown to utilize an alternative nuclear import pathway from wild-type HIV (1, 2, 16). The N74D mutation also alters the integration site preference of HIV (17, 18). In the CsA washout assay, E45A mutant virus uncoated slower than the wild-type virus and had a significantly greater average half-life of uncoating (124 min) compared to HIV-GFP (85 min; Fig. 1B). N74D mutant virus uncoated slower than the wild-type virus, although it was not as delayed as E45A (Fig. 1C). The average half-life of uncoating for N74D (110 min) was also significantly increased compared to HIV-GFP (85 min; Fig. 1C). In our initial analysis of HIV-1 uncoating, we identified the p24CA mutation Q63/67A as an additional slow uncoating virus compared to the wild-type virus (8). However, the uncoating of Q63/67A mutant virus was so delayed that we did not calculate a half-life of uncoating, as by 5 h postinfection only 20% of the virus had uncoated.
Finally, we examined the uncoating kinetics of A92E and G94D mutant viruses. These mutations are located near the cyclophilin A binding loop and do not inhibit CypA binding, but instead they alter viral infectivity in response to CsA in a cell type-dependent manner (31–33). Because CypA can still bind the capsid, these mutants are proposed to alter the response of the virus to cellular CypA binding. In addition, these mutants are impaired for the ability of HIV to infect nondividing cells, indicating that they may have lost the ability to actively transport into the nucleus (14, 34). In the CsA washout assay, A92E mutant virus uncoated faster than HIV-GFP with a significantly decreased half-life of uncoating (50 min) compared to the wild-type virus (90 min; Fig. 1D). The G94D mutation did not significantly alter the kinetics of uncoating or the average half-life of uncoating (90 min) compared to HIV-GFP (97 min; Fig. 1E).
Reverse transcription does not account for the changes in uncoating kinetics.
We previously used the CsA washout assay to determine that reverse transcription facilitates the process of uncoating in cells (8). Therefore, it is possible that the changes in uncoating kinetics of these viruses may be due to alterations in reverse transcription. To examine completion of reverse transcription for each mutant virus over time in OMK cells, we performed an addition assay with the nonnucleoside reverse transcriptase inhibitor nevirapine. Using the same experimental conditions as in the CsA washout assay, OMK cells were synchronously infected with wild-type or p24CA mutant GFP reporter virus. Nevirapine was then added at times corresponding to those used in the CsA washout assay. The cells were harvested 2 days postinfection, and the percentage of infected cells was determined at each time point of nevirapine addition. These data were normalized by setting the percentage of GFP-positive cells in the DMSO carrier control to 100%. Of all the p24CA mutants examined, only E45A mutant virus showed a correlation between uncoating kinetics and reverse transcription, but this delay in reverse transcription was not statistically significant (Fig. 1 and 2). N74D and A92E mutant viruses seemed to reverse transcribe at a rate similar to the wild type, which is not consistent with uncoating slower and faster, respectively, than the wild type (Fig. 1 and 2). G94D mutant virus showed the most delayed completion of reverse transcription of all the mutants tested, which was not statistically significant, but this mutation did not alter uncoating kinetics. We have also previously observed a disparity in the kinetics of completion of reverse transcription and uncoating with the Q63/67A virus. This p24CA mutant reverse transcribed at a rate similar to that of the wild type, but it displayed much delayed uncoating (8). Therefore, these data suggest that while reverse transcription may facilitate uncoating, there are other factors involved with the process.
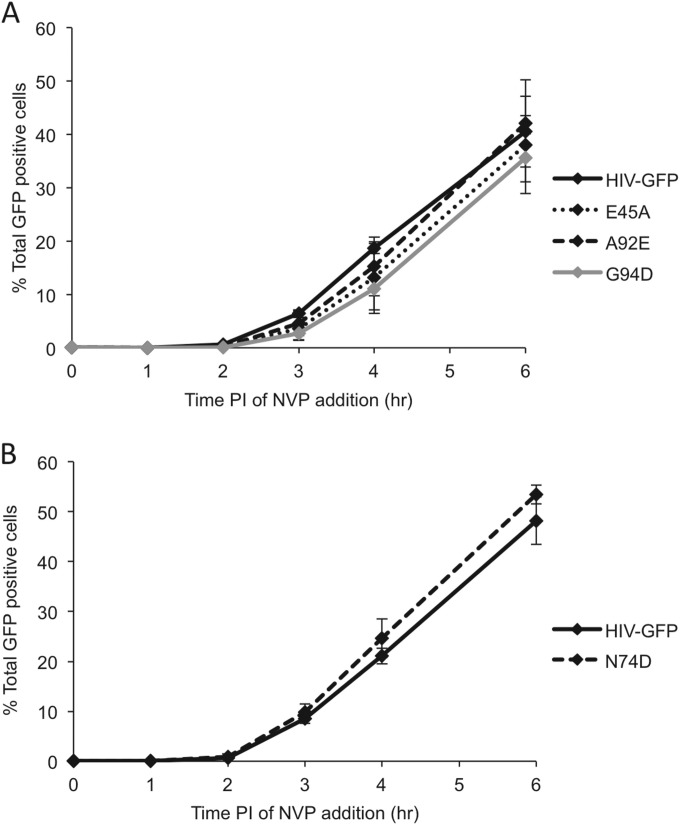
FIG 2.
Progression of reverse transcription for HIV-GFP and p24CA mutant viruses. A nevirapine (NVP) addition assay was conducted to monitor completion of reverse transcription for HIV-GFP and each of the p24CA mutant viruses. Data were normalized by setting the value for the DMSO carrier control to 100%. Values are averages ± standard errors (error bars) from three independent experiments. (A) The E45A and G94D mutations subtly decrease the rate of reverse transcription, but this change is not statistically significant at the 6-h time point. The A92E mutation does not significantly alter completion of reverse transcription relative to the wild type as assayed at the 6-h time point. (B) The N74D mutation does not significantly alter completion of reverse transcription relative to the wild type as assayed at the 6-h time point.
To determine the hierarchy between reverse transcription and the p24CA mutations on the process of uncoating, we examined the effect of reverse transcription on the uncoating kinetics of the p24CA mutant viruses. The CsA washout assay was conducted with and without a 2-h nevirapine treatment with HIV-GFP and each p24CA mutant. Treatment with nevirapine shifted the washout assay curves of all mutants similar to that observed with wild-type virus (Fig. 3). Interestingly, all viruses displayed the same rapid increase in the percentage of GFP-positive cells in the hour after nevirapine removal. Therefore, inhibition of reverse transcription has a dominant effect on uncoating compared to the p24CA mutations.
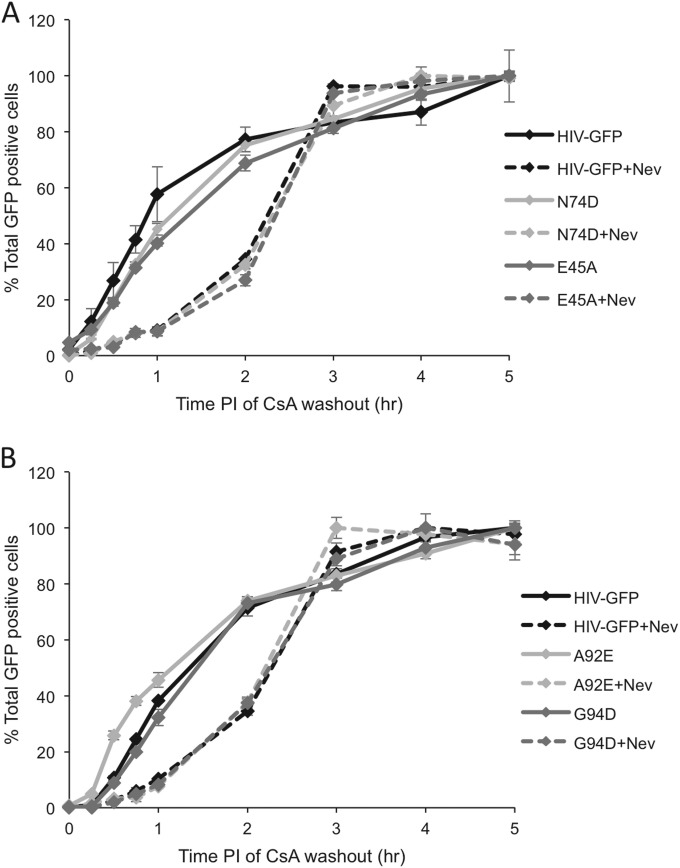
FIG 3.
Inhibition of reverse transcription delays the uncoating of p24CA mutant viruses. The CsA washout assay was performed with and without a 2-h nevirapine (Nev) treatment for HIV-GFP and each of the p24CA mutant viruses. The results of a representative assay from three independent experiments are shown. Error bars denote standard errors. Inhibition of reverse transcription similarly delays the uncoating kinetics of HIV-GFP and E45A, N74D, A92E, and G94D mutant viruses despite their differential rates of uncoating without nevirapine treatment.
The cellular milieu can impact uncoating kinetics.
We do not know to what extent the cellular environment affects uncoating in vivo. Currently, the favored hypothesis is that cellular factors are involved with facilitating the process of uncoating, but a bona fide uncoating factor has not been identified. In order to examine the effect of cellular environment on uncoating kinetics, we conducted the CsA washout assay in a HeLa cell line that was engineered to express TRIM-CypA (HeLa-TC). This assay should also reveal any differences in uncoating between owl monkey cells and human cells, a more relevant cell line with regard to HIV infection.
In parallel CsA washout assays, HIV-GFP uncoated significantly faster in HeLa-TC cells than in OMK cells (Fig. 4). We have previously shown that changes in the rate of viral fusion can account for alterations in the observed half-life of uncoating in the CsA washout assay (8). Therefore, we performed an ammonium chloride addition assay in both cell lines in parallel with a CsA washout assay to directly correlate the timing of viral fusion and uncoating. Similar to the nevirapine addition assay, cells were infected with GFP reporter virus, and then ammonium chloride was added at various times postinfection to block viral fusion. The half-life of viral fusion was greater in HeLa-TC cells than in OMK cells indicating that the difference in uncoating kinetics between the two cell lines is not due to differences in viral fusion (Fig. 4 and Table 1). We also examined the uncoating kinetics of N74D virus in both cell lines, as this virus has been the most well studied for its differential interaction with cellular host factors (16–19, 35). Similar to OMK cells, N74D mutant virus uncoated slower than the wild-type virus in HeLa-TC cells (Fig. 4). However, the delay in uncoating in HeLa-TC cells was significantly greater than that observed in OMK cells. Collectively, these results suggest that the cellular environment of HeLa-TC cells has a differential effect on the uncoating of HIV-GFP and N74D viruses. As both viruses were pseudotyped with VSV-g, it is unlikely that this discrepancy is due to differences in viral fusion.
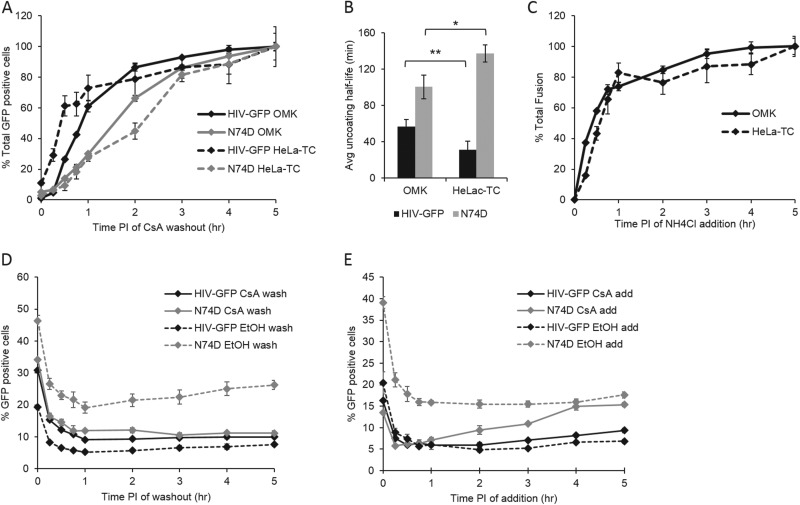
FIG 4.
Cellular factors differentially affect uncoating. (A) The CsA washout assay was performed with HIV-GFP and N74D mutant virus in OMK and HeLa-TC cells. HIV-GFP uncoats faster in HeLa-TC cells than in OMK cells. The uncoating of N74D mutant virus is delayed to greater extent in HeLa-TC cells than in OMK cells compared to the wild-type virus (HIV-GFP). The results shown are from a representative washout assay. Error bars denote standard errors. (B) The average half-life of uncoating for each virus in each cell type was determined from 4 independent experiments. There is a significant difference in uncoating half-life for HIV-GFP in OMK cells and HeLa-TC cells. There is also a significant difference in the increased half-life of uncoating due to the N74D mutation between OMK cells and HeLa-TC cells. Error bars denote standard errors. Values that are significantly different are indicated by bars and asterisks as follows: *, P < 0.05; **, P < 0.005. (C) An ammonium chloride addition assay was performed in parallel with a CsA washout assay to determine the kinetics of viral fusion in relation to uncoating in OMK and HeLa-TC cells. VSV-g pseudotyped viruses fused faster in OMK cells than in HeLa-TC cells. The results shown are of a representative assay. Error bars denote standard errors. (D) To determine the effect of CsA on viral infectivity over time in HeLa cells, the CsA washout assay was performed in the parent HeLa cell line for HIV-GFP and N74D viruses. Ethanol washout was the negative control. Compared to the ethanol control, CsA subtly increases HIV-GFP infectivity at each time point assayed. Compared to the ethanol control, CsA decreases N74D mutant virus infectivity with similar magnitude at each time point assayed. The results shown are of a representative experiment from 3 independent experiments. Error bars denote standard errors. (E) To determine the susceptibility of HIV-GFP and N74D viruses to CsA over time, a CsA addition assay was performed in HeLa cells. Ethanol addition was the negative control. HIV-GFP does not show a change in infectivity over any time of CsA addition. Compared to the ethanol control, N74D virus loses susceptibility to CsA addition between 1 and 4 h postinfection. The results shown are of a representative experiment from 3 independent experiments. Error bars denote standard errors.
TABLE 1.
Viral fusion impacts the observed rate of uncoatinga
| Cell | Avg half-life (min)b |
|
|---|---|---|
| Uncoating | Fusion | |
| OMK | 63.79 (9.48) | 18.8 (1.78) |
| HeLa-TC | 36.79 (1.96) | 28.38 (0.62) |
The CsA washout assay was performed with HIV-GFP in OMK and HeLa-TC cells in parallel with an ammonium chloride addition assay to monitor viral fusion.
The average half-lives of uncoating and viral fusion were determined from 3 independent experiments. The values shown in parentheses are standard errors.
One potential confounding factor in the assay is the cellular protein CypA that binds p24CA. The role of CypA in HIV-1 replication is not well understood, as CypA can increase, decrease, or have no effect on viral infectivity depending on the cell type (30, 36–38). In the CsA washout assay, the presence of CsA will also prevent the binding of cellular CypA. Therefore, observations made about uncoating using the CsA washout assay will not reveal the effect of cellular CypA, if any, on uncoating. However, it has been shown that the N74D mutation may affect the susceptibility of HIV-1 to cellular CypA in HeLa cells (39). Specifically, blocking CypA binding with CsA decreased infectivity of N74D virus by ∼2- to 3-fold in HeLa cells, whereas wild-type virus infectivity was slightly increased. We normalize data in the CsA washout assay using the 4- or 5-h time point independently for each virus. If cellular CypA has a differential effect on infectivity over time, normalizing the data using the 5-h time point of CsA washout could bias the resultant uncoating curves for each virus. To directly examine the effect of cellular CypA over time on wild-type and N74D mutant virus, we performed a CsA washout assay in the parent HeLa cell line that was used to generate the HeLa-TC cell line (Fig. 4D). Ethanol washout was the negative control. The presence of CsA subtly increased HIV-GFP infectivity compared to the ethanol control, and this increase was of a similar magnitude at each time point examined. For N74D mutant virus, the presence of CsA decreased infectivity by 2-fold. Importantly, this decrease in infectivity was consistent at each time point tested. Therefore, using the 5-h time point to normalize data in the CsA washout assay data will not bias the resultant uncoating kinetics of either HIV-GFP or N74D virus.
The differences between the CsA and ethanol washout curves were consistent over time, suggesting that cellular CypA may not affect uncoating of N74D or HIV-GFP virus in HeLa cells. To more directly address this idea, we performed a CsA addition assay in the parent HeLa cell line, reasoning that if CypA affected uncoating, the virus would escape from the inhibitory effect of CsA with kinetics similar to that of the uncoating (Fig. 4E). CsA addition did not affect HIV-GFP infectivity compared to the ethanol control at any time point tested. In contrast, N74D virus escaped the inhibitory effects of CsA by 4 h postinfection compared to the ethanol control. This gradual increase in infectivity is not similar to the uncoating kinetics as assayed in the CsA washout assay (Fig. 4A), suggesting that CypA is not involved in uncoating of N74D virus. Instead, these data suggest that some aspect of the N74D capsid changes by 4 h postinfection so that it is no longer susceptible to the effects of CypA binding. Given that N74D virus has been shown to utilize an alternate set of nuclear import factors, it is intriguing to speculate that the role of capsid in nuclear import of N74D mutant virus may be different from wild-type virus.
DISCUSSION
Using the CsA washout assay, we were able to observe the effect of p24CA mutations on uncoating kinetics in HIV-1-infected cells (Fig. 1). In general, there was an approximately 1.5- to 2-fold difference in the half-life of uncoating for the E45A, N74D, and A92E mutant viruses compared to the wild-type virus (HIV-GFP). While this difference may be thought of as small, this alteration in uncoating is sufficient to decrease overall infectivity of the virus (1, 2, 14, 39). Interestingly, the two p24CA mutants with the largest alterations in uncoating, E45A and Q63/67A mutant viruses, display greatest defects in infectivity at 3 to 4% of the wild-type virus (1, 2, 8). The processivity of the HIV-1 reverse transcriptase (RT) is 2 or 3 nucleotides per second, so changes in uncoating rates of as little as 10 minutes would correlate with very different states (on the order of 1.2 kb) of the reverse transcribing genome. Given the complex interplay between uncoating and reverse transcription, these changes in uncoating could therefore lead to decreased infectivity. The CsA washout assay is also based on infectivity. Therefore, the effect of each mutation on uncoating is likely more severe than the data indicate, as a positive readout in the CsA washout assay requires that the virus uncoat and establish a provirus from which to express the GFP reporter. Thus, these mutants also provide a measure of the extent to which HIV can tolerate changes in uncoating and still achieve productive infection.
As expected, G89V and P90A mutant viruses did not display a change in GFP-positive cells over time due to the fact that these mutations disrupt cyclophilin A (CypA) binding (30). In addition to serving as controls, these mutants show that the increase in infectivity seen over time in the assay is dependent on the ability of TRIM-CypA to bind the retroviral capsid. E45A mutant virus has previously been shown to have increased capsid stability and uncoat slower than the wild-type virus in an in vitro assay (1). Our results are consistent with these data, as we observed a lower rate of uncoating for E45A mutant virus. In contrast, N74D mutant virus does not alter capsid stability and uncoats similarly to the wild-type virus in an in vitro assay (35). However, in the CsA washout assay, this mutant uncoated slower than the wild type. Additionally, A92E mutant virus displays capsid stability similar to that of the wild-type virus, but it uncoats faster in the CsA washout assay (14). While there is often a general correlation between in vitro capsid stability and the rate of uncoating, these data suggest that other factors are involved with uncoating in vivo. A similar conclusion was reached in a recent study in which the R132K suppressor mutation of E45A mutant virus rescued infectivity but did not restore capsid stability (40). Likewise, we previously reported the Q63/67A p24CA mutation results in slow uncoating kinetics in the CsA washout assay while showing decreased capsid stability and accelerated uncoating kinetics in an in vitro assay (1, 8).
Why are there discrepancies between different uncoating assays? Depending on the methods used, each assay may detect a different aspect of the uncoating process. Furthermore, it is not surprising that some mutants may display a different effect on uncoating when assayed using in vivo compared to in vitro assays, given that the HIV-1 capsid is relatively difficult to isolate and inherently unstable. For in vitro uncoating assays, viral capsids are purified and incubated, and then uncoating is determined by comparing the relative amounts of particulate and soluble p24CA protein (1, 41). In the cell-based fate of the capsid assay, infected cells are lysed several hours postinfection, and uncoating is assayed by comparing the amounts of particulate and soluble forms of p24CA (42). Coated viral particles can be directly detected in the cytoplasm of infected cells by staining with an antibody to p24CA in microscopy-based uncoating assays (8, 13, 43). Differing from these assays, the CsA washout uses infectivity as a readout and thus is indirect, as it measures viral susceptibility to TRIM-CypA. However, the timing of the loss of sensitivity to TRIM-CypA restriction in the CsA washout assay correlates with the loss of p24CA in infected cells as assayed by fluorescence microscopy and Western blotting (8, 12, 13). In addition, the delay in uncoating due to inhibition of reverse transcription, first identified with the CsA washout assay and a microscopy assay, has also been confirmed biochemically by Western blotting (8, 9). Therefore, despite an indirect detection method, the CsA washout assay is a good method to monitor the process of uncoating in infected cells.
The CsA washout assay provides a direct correlation between uncoating and infectivity. Because of this, the effect of a mutation on uncoating in the CsA washout assay will be detected only in the subset of viral particles that can achieve productive infection. In the other uncoating assays outlined above, viral capsids are analyzed irrespective of their ability to infect cells. As the majority of HIV virions that enter cells do not establish productive infection, the characteristics of these uninfectious virions could account for some of the differences observed between assays. These differences highlight the importance of determining the uncoating kinetics of important p24CA mutants in infected cells and with respect to viral infectivity.
Do changes in uncoating kinetics account for the effects of the p24CA mutations on other steps of HIV-1 replication? Mutations in p24CA have been shown to influence infection of nondividing cells, utilization of nuclear import pathways, and integration site selection (13–19). The E45A, Q63/67A, and N74D mutations all decrease the rate of uncoating to different degrees, with the N74D mutant virus the most similar to the wild-type virus and the Q63/67A mutant virus the most delayed (Fig. 1) (8). These mutants also all utilize an alternative nuclear import pathway from TNPO3 and NUP358/RanBP2 which has been implicated in facilitating wild-type HIV-1 nuclear entry (16, 17). Thus, it is tempting to speculate that the rate of uncoating may affect which nuclear import pathway HIV uses during infection. In contrast, A92E and G94D mutant viruses are impaired for infection in nondividing cells in a CypA-dependent manner (14, 34). As only A92E mutant virus uncoats at a rate different from that of the wild-type virus, it seems unlikely that the inability of these mutants to infect nondividing cells is due to an altered rate of uncoating.
While our data suggest that reverse transcription facilitates uncoating, for any of the p24CA mutant viruses there was not a significant difference in reverse transcription that could account for a change in uncoating kinetics (8) (Fig. 2). This result further reinforces the conclusion that other factors, likely both viral and cellular, are involved in the uncoating process. The importance of cellular factors in uncoating is demonstrated by the experiments in HeLa-TC cells (Fig. 4). Despite a lower rate of viral fusion, uncoating of wild-type virus is faster in HeLa-TC cells (Fig. 4 and Table 1). This could potentially be due to a factor that facilitates uncoating in more abundance in HeLa-TC cells than in OMK cells or the absence of a factor that abrogates uncoating in HeLa-TC cells. Further experiments would be required to identify such a factor, but these results are important to consider when choosing time points to assay uncoating in different cell types. Furthermore, there was a more pronounced delay in uncoating due to the N74D mutation in HeLa-TC cells compared to OMK cells. This delay is even more profound when considering that uncoating of wild-type virus is faster in HeLa-TC cells. Interestingly, these results suggest that the cellular environment of HeLa-TC cells has a differential effect on uncoating by facilitating more-rapid uncoating of HIV-GFP and more-delayed uncoating of N74D mutant virus. Given that N74D virus has been shown to utilize a different set of host factors for nuclear entry than wild-type virus, it is tempting to speculate that these host factors may be involved with the differential effect in HeLa-TC cells.
The simplest hypothesis is that expression levels of the TRIM-CypA protein account for the differences in uncoating kinetics between OMK and HeLa-TC cells. However, the data do not support this explanation in two ways. First, in both cell lines, the CsA washout assay was conducted under conditions in which TRIM-CypA restriction is not saturated, meaning that in the absence of CsA, viral infectivity is completely inhibited. TRIM proteins rapidly associate with incoming virus (10). Therefore, under nonsaturating conditions, there is a sufficient amount of TRIM-CypA protein in both cell lines to bind to the viral capsid and inhibit infectivity. Any additional amount of TRIM-CypA protein present in the cells therefore becomes essentially irrelevant for restriction. Second, if different levels of TRIM-CypA expression were responsible for the altered uncoating kinetics observed in HeLa-TC cells compared to OMK cells, then both viruses should display the same shift in uncoating kinetics. This is not what we observed. Instead, wild-type virus uncoated faster in HeLa-TC cells, while N74D virus uncoated slower (Fig. 4A). Therefore, overall differences in TRIM-CypA protein levels between HeLa-TC and OMK cells do not account for the differences observed in the uncoating of wild-type and N74D viruses.
A second hypothesis is that the cellular protein CypA is responsible for the differential uncoating kinetics observed between OMK and HeLa cells. The use of CsA in the assay, which is necessary to regulate TRIM-CypA binding, will also prevent the binding of cellular cyclophilin A. The effect of CypA on HIV-1 infectivity is highly cell type specific, and the role of CypA in uncoating is not defined (30, 36–38). CypA was able to modulate uncoating in a cell type-dependent manner in the fate of the capsid assay and stabilize the capsid in an in vitro uncoating assay (35, 44). In the washout assay, observations made about uncoating are irrespective of any effect of cellular CypA on uncoating due to the presence of CsA. It was previously shown that CsA treatment decreased infectivity of N74D mutant virus by ∼2- to 3-fold in HeLa cells, whereas wild-type virus infectivity was slightly increased (39). Given that we normalize data in the CsA washout assay using infectivity at the 4- or 5-h time point, this cell type-specific alternation in infectivity could bias results if the effect of cellular CypA on infectivity changes over time. However, we found that the increase in HIV-GFP infectivity and decrease in N74D virus infectivity was consistent at all time points examined in the parent HeLa cell line (Fig. 4D). Therefore, using the 5-h time point to normalize data in the CsA washout assay data will not bias the resultant uncoating kinetics of either HIV-GFP or N74D virus. Collectively, these results suggest that cell-specific effects of CypA do not account for the differential uncoating kinetics of HIV-GFP and N74D viruses in HeLa cells compared to OMK cells.
What stage of uncoating is detected in the CsA washout assay? This question is difficult to answer, as the exact mechanism of TRIM protein restriction is not completely defined. The current model is that TRIM forms a lattice or net around the capsid. This model is based on electron microscopy experiments in which recombinant TRIM5-21R associated in a hexagonal lattice over a hexagonal array of p24CA (27). TRIM-CypA would be expected to form a similar lattice, given its high degree of similarity in the coiled-coil and B-Box 2 domains required for lattice formation (20, 27). Despite this model, we do not know how much of the viral capsid is required to be present for TRIM-CypA lattice formation and restriction. TRIM-CypA can inhibit the infectivity of virus with mixed capsids in which only 25% of p24CA monomers are susceptible to restriction (45). However, in these experiments viral capsids were exposed to TRIM-CypA upon viral entry and therefore were likely intact. A different situation exists in the CsA washout assay where viral capsids are allowed to uncoat to different extents in the cytoplasm before being exposed to TRIM-CypA binding by the withdrawal of CsA. A microscopy assay that examined the integrity of viral capsids localizing with rhesus TRIM5α cytoplasmic bodies suggested that TRIM proteins associate with intact capsids (46). On the basis of these data, we hypothesize that in the CsA washout assay TRIM-CypA is binding intact capsids or viral complexes that have just started to uncoat. Further development of the assay is needed to test this hypothesis, but this would suggest that the CsA washout assay is detecting early stages of HIV uncoating.
We do not know how the process of uncoating occurs, but the data presented here indicate that viral and cellular factors are important. In our current model, the process of reverse transcription facilitates HIV-1 uncoating in the cytoplasm of infected cells. In order to refine this model, we examined the effect of reverse transcription on the uncoating of each p24CA mutant. Strikingly, wild-type and p24CA mutant viruses displayed similar delays in uncoating, suggesting that the effect of reverse transcription on uncoating is functionally active before the effect of the p24CA mutation (Fig. 3). In these experiments, there was also a rapid increase in the percentage of GFP-positive cells in the hour after nevirapine removal for wild-type and p24CA mutant viruses (Fig. 3) (8). This result indicates that a reverse transcription product generated in the hour after nevirapine removal is facilitating uncoating. Using quantitative PCR analysis, we previously found that early products of reverse transcription were generated in the first 2 h after viral fusion, which correlates with the timing of uncoating in the CsA washout assay (8). In addition, in the CsA washout assay the majority of uncoating occurs by 3 h postinfection, which precedes the completion of reverse transcription (Fig. 1 and 2) (8). Therefore, these data further support a model in which early steps of reverse transcription facilitate HIV-1 uncoating.
ACKNOWLEDGMENTS
We thank Paul Bieniasz and Theodora Hatziioannou for OMK cells and Chris Aiken for proviral plasmids. We also thank the NIH AIDS Research and Reference Reagent Program for nevirapine. We thank the RHLCC flow cytometry core. We also thank Cindy Danielson for critical review of the manuscript.
This work was supported by NIH grants RO1 AI47770 and P50 GM082545 to T.J.H. and grant 5F32AI089359-03 to A.E.H. The RHLCC flow cytometry core was supported by a Northwestern University Flow Cytometry Core Facility and Cancer Center support grant (NCI CA060553).
REFERENCES
- 1.Forshey BM, von Schwedler U, Sundquist WI, Aiken C. 2002. Formation of a human immunodeficiency virus type 1 core of optimal stability is crucial for viral replication. J Virol 76:5667–5677. doi: 10.1128/JVI.76.11.5667-5677.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.von Schwedler UK, Stray KM, Garrus JE, Sundquist WI. 2003. Functional surfaces of the human immunodeficiency virus type 1 capsid protein. J Virol 77:5439–5450. doi: 10.1128/JVI.77.9.5439-5450.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Fitzon T, Leschonsky B, Bieler K, Paulus C, Schroder J, Wolf H, Wagner R. 2000. Proline residues in the HIV-1 NH2-terminal capsid domain: structure determinants for proper core assembly and subsequent steps of early replication. Virology 268:294–307. doi: 10.1006/viro.1999.0178. [DOI] [PubMed] [Google Scholar]
- 4.Tang S, Murakami T, Agresta BE, Campbell S, Freed EO, Levin JG. 2001. Human immunodeficiency virus type 1 N-terminal capsid mutants that exhibit aberrant core morphology and are blocked in initiation of reverse transcription in infected cells. J Virol 75:9357–9366. doi: 10.1128/JVI.75.19.9357-9366.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Tang S, Murakami T, Cheng N, Steven AC, Freed EO, Levin JG. 2003. Human immunodeficiency virus type 1 N-terminal capsid mutants containing cores with abnormally high levels of capsid protein and virtually no reverse transcriptase. J Virol 77:12592–12602. doi: 10.1128/JVI.77.23.12592-12602.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.McDonald D, Vodicka MA, Lucero G, Svitkina TM, Borisy GG, Emerman M, Hope TJ. 2002. Visualization of the intracellular behavior of HIV in living cells. J Cell Biol 159:441–452. doi: 10.1083/jcb.200203150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Anderson JL, Campbell EM, Wu X, Vandegraaff N, Engelman A, Hope TJ. 2006. Proteasome inhibition reveals that a functional preintegration complex intermediate can be generated during restriction by diverse TRIM5 proteins. J Virol 80:9754–9760. doi: 10.1128/JVI.01052-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hulme AE, Perez O, Hope TJ. 2011. Complementary assays reveal a relationship between HIV-1 uncoating and reverse transcription. Proc Natl Acad Sci U S A 108:9975–9980. doi: 10.1073/pnas.1014522108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yang Y, Fricke T, Diaz-Griffero F. 2013. Inhibition of reverse transcriptase activity increases stability of the HIV-1 core. J Virol 87:683–687. doi: 10.1128/JVI.01228-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Perez-Caballero D, Hatziioannou T, Zhang F, Cowan S, Bieniasz PD. 2005. Restriction of human immunodeficiency virus type 1 by TRIM-CypA occurs with rapid kinetics and independently of cytoplasmic bodies, ubiquitin, and proteasome activity. J Virol 79:15567–15572. doi: 10.1128/JVI.79.24.15567-15572.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hulme AE, Hope TJ. 2014. The cyclosporin A washout assay to detect HIV-1 uncoating in infected cells. Methods Mol Biol 1087:37–46. doi: 10.1007/978-1-62703-670-2_4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kutluay SB, Perez-Caballero D, Bieniasz PD. 2013. Fates of retroviral core components during unrestricted and TRIM5-restricted infection. PLoS Pathog 9:e1003214. doi: 10.1371/journal.ppat.1003214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yamashita M, Perez O, Hope TJ, Emerman M. 2007. Evidence for direct involvement of the capsid protein in HIV infection of nondividing cells. PLoS Pathog 3:1502–1510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Qi M, Yang R, Aiken C. 2008. Cyclophilin A-dependent restriction of human immunodeficiency virus type 1 capsid mutants for infection of nondividing cells. J Virol 82:12001–12008. doi: 10.1128/JVI.01518-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ylinen LM, Schaller T, Price A, Fletcher AJ, Noursadeghi M, James LC, Towers GJ. 2009. Cyclophilin A levels dictate infection efficiency of human immunodeficiency virus type 1 capsid escape mutants A92E and G94D. J Virol 83:2044–2047. doi: 10.1128/JVI.01876-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lee K, Ambrose Z, Martin TD, Oztop I, Mulky A, Julias JG, Vandegraaff N, Baumann JG, Wang R, Yuen W, Takemura T, Shelton K, Taniuchi I, Li Y, Sodroski J, Littman DR, Coffin JM, Hughes SH, Unutmaz D, Engelman A, KewalRamani VN. 2010. Flexible use of nuclear import pathways by HIV-1. Cell Host Microbe 7:221–233. doi: 10.1016/j.chom.2010.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Schaller T, Ocwieja KE, Rasaiyaah J, Price AJ, Brady TL, Roth SL, Hue S, Fletcher AJ, Lee K, KewalRamani VN, Noursadeghi M, Jenner RG, James LC, Bushman FD, Towers GJ. 2011. HIV-1 capsid-cyclophilin interactions determine nuclear import pathway, integration targeting and replication efficiency. PLoS Pathog 7:e1002439. doi: 10.1371/journal.ppat.1002439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Koh Y, Wu X, Ferris AL, Matreyek KA, Smith SJ, Lee K, KewalRamani VN, Hughes SH, Engelman A. 2013. Differential effects of human immunodeficiency virus type 1 capsid and cellular factors nucleoporin 153 and LEDGF/p75 on the efficiency and specificity of viral DNA integration. J Virol 87:648–658. doi: 10.1128/JVI.01148-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Matreyek KA, Engelman A. 2011. The requirement for nucleoporin NUP153 during human immunodeficiency virus type 1 infection is determined by the viral capsid. J Virol 85:7818–7827. doi: 10.1128/JVI.00325-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sayah DM, Sokolskaja E, Berthoux L, Luban J. 2004. Cyclophilin A retrotransposition into TRIM5 explains owl monkey resistance to HIV-1. Nature 430:569–573. doi: 10.1038/nature02777. [DOI] [PubMed] [Google Scholar]
- 21.Sebastian S, Luban J. 2005. TRIM5alpha selectively binds a restriction-sensitive retroviral capsid. Retrovirology 2:40. doi: 10.1186/1742-4690-2-40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Forshey BM, Shi J, Aiken C. 2005. Structural requirements for recognition of the human immunodeficiency virus type 1 core during host restriction in owl monkey cells. J Virol 79:869–875. doi: 10.1128/JVI.79.2.869-875.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Towers GJ. 2005. Control of viral infectivity by tripartite motif proteins. Hum Gene Ther 16:1125–1132. doi: 10.1089/hum.2005.16.1125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Nisole S, Stoye JP, Saib A. 2005. TRIM family proteins: retroviral restriction and antiviral defence. Nat Rev Microbiol 3:799–808. doi: 10.1038/nrmicro1248. [DOI] [PubMed] [Google Scholar]
- 25.Stremlau M, Owens CM, Perron MJ, Kiessling M, Autissier P, Sodroski J. 2004. The cytoplasmic body component TRIM5alpha restricts HIV-1 infection in Old World monkeys. Nature 427:848–853. doi: 10.1038/nature02343. [DOI] [PubMed] [Google Scholar]
- 26.Wu X, Anderson JL, Campbell EM, Joseph AM, Hope TJ. 2006. Proteasome inhibitors uncouple rhesus TRIM5alpha restriction of HIV-1 reverse transcription and infection. Proc Natl Acad Sci U S A 103:7465–7470. doi: 10.1073/pnas.0510483103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ganser-Pornillos BK, Chandrasekaran V, Pornillos O, Sodroski JG, Sundquist WI, Yeager M. 2011. Hexagonal assembly of a restricting TRIM5alpha protein. Proc Natl Acad Sci U S A 108:534–539. doi: 10.1073/pnas.1013426108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nisole S, Lynch C, Stoye JP, Yap MW. 2004. A Trim5-cyclophilin A fusion protein found in owl monkey kidney cells can restrict HIV-1. Proc Natl Acad Sci U S A 101:13324–13328. doi: 10.1073/pnas.0404640101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Towers GJ, Hatziioannou T, Cowan S, Goff SP, Luban J, Bieniasz PD. 2003. Cyclophilin A modulates the sensitivity of HIV-1 to host restriction factors. Nat Med 9:1138–1143. doi: 10.1038/nm910. [DOI] [PubMed] [Google Scholar]
- 30.Gamble TR, Vajdos FF, Yoo S, Worthylake DK, Houseweart M, Sundquist WI, Hill CP. 1996. Crystal structure of human cyclophilin A bound to the amino-terminal domain of HIV-1 capsid. Cell 87:1285–1294. doi: 10.1016/S0092-8674(00)81823-1. [DOI] [PubMed] [Google Scholar]
- 31.Braaten D, Aberham C, Franke EK, Yin L, Phares W, Luban J. 1996. Cyclosporine A-resistant human immunodeficiency virus type 1 mutants demonstrate that Gag encodes the functional target of cyclophilin A. J Virol 70:5170–5176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Aberham C, Weber S, Phares W. 1996. Spontaneous mutations in the human immunodeficiency virus type 1 gag gene that affect viral replication in the presence of cyclosporins. J Virol 70:3536–3544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hatziioannou T, Perez-Caballero D, Cowan S, Bieniasz PD. 2005. Cyclophilin interactions with incoming human immunodeficiency virus type 1 capsids with opposing effects on infectivity in human cells. J Virol 79:176–183. doi: 10.1128/JVI.79.1.176-183.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yamashita M, Emerman M. 2009. Cellular restriction targeting viral capsids perturbs human immunodeficiency virus type 1 infection of nondividing cells. J Virol 83:9835–9843. doi: 10.1128/JVI.01084-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Shah VB, Shi J, Hout DR, Oztop I, Krishnan L, Ahn J, Shotwell MS, Engelman A, Aiken C. 2013. The host proteins transportin SR2/TNPO3 and cyclophilin A exert opposing effects on HIV-1 uncoating. J Virol 87:422–432. doi: 10.1128/JVI.07177-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Luban J, Bossolt KL, Franke EK, Kalpana GV, Goff SP. 1993. Human immunodeficiency virus type 1 Gag protein binds to cyclophilins A and B. Cell 73:1067–1078. doi: 10.1016/0092-8674(93)90637-6. [DOI] [PubMed] [Google Scholar]
- 37.Franke EK, Yuan HE, Luban J. 1994. Specific incorporation of cyclophilin A into HIV-1 virions. Nature 372:359–362. doi: 10.1038/372359a0. [DOI] [PubMed] [Google Scholar]
- 38.Thali M, Bukovsky A, Kondo E, Rosenwirth B, Walsh CT, Sodroski J, Gottlinger HG. 1994. Functional association of cyclophilin A with HIV-1 virions. Nature 372:363–365. doi: 10.1038/372363a0. [DOI] [PubMed] [Google Scholar]
- 39.Ambrose Z, Lee K, Ndjomou J, Xu H, Oztop I, Matous J, Takemura T, Unutmaz D, Engelman A, Hughes SH, KewalRamani VN. 2012. Human immunodeficiency virus type 1 capsid mutation N74D alters cyclophilin A dependence and impairs macrophage infection. J Virol 86:4708–4714. doi: 10.1128/JVI.05887-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yang R, Shi J, Byeon IJ, Ahn J, Sheehan JH, Meiler J, Gronenborn AM, Aiken C. 2012. Second-site suppressors of HIV-1 capsid mutations: restoration of intracellular activities without correction of intrinsic capsid stability defects. Retrovirology 9:30. doi: 10.1186/1742-4690-9-30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Aiken C. 2009. Cell-free assays for HIV-1 uncoating. Methods Mol Biol 485:41–53. doi: 10.1007/978-1-59745-170-3_4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Stremlau M, Perron M, Lee M, Li Y, Song B, Javanbakht H, Diaz-Griffero F, Anderson DJ, Sundquist WI, Sodroski J. 2006. Specific recognition and accelerated uncoating of retroviral capsids by the TRIM5alpha restriction factor. Proc Natl Acad Sci U S A 103:5514–5519. doi: 10.1073/pnas.0509996103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Arhel NJ, Souquere-Besse S, Munier S, Souque P, Guadagnini S, Rutherford S, Prevost MC, Allen TD, Charneau P. 2007. HIV-1 DNA Flap formation promotes uncoating of the pre-integration complex at the nuclear pore. EMBO J 26:3025–3037. doi: 10.1038/sj.emboj.7601740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Li Y, Kar AK, Sodroski J. 2009. Target cell type-dependent modulation of human immunodeficiency virus type 1 capsid disassembly by cyclophilin A. J Virol 83:10951–10962. doi: 10.1128/JVI.00682-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Shi J, Friedman DB, Aiken C. 2013. Retrovirus restriction by TRIM5 proteins requires recognition of only a small fraction of viral capsid subunits. J Virol 87:9271–9278. doi: 10.1128/JVI.00713-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yu Z, Dobro MJ, Woodward CL, Levandovsky A, Danielson CM, Sandrin V, Shi J, Aiken C, Zandi R, Hope TJ, Jensen GJ. 2013. Unclosed HIV-1 capsids suggest a curled sheet model of assembly. J Mol Biol 425:112–123. doi: 10.1016/j.jmb.2012.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]