Abstract
The human DMD gene is the largest known to date, spanning > 2000 kb on the X chromosome. The gene size is mainly accounted for by huge intronic regions. We sequenced 190 kb of Fugu rubripes (pufferfish) genomic DNA corresponding to the complete dystrophin gene (FrDMD) and provide the first report of gene structure and sequence comparison among dystrophin genomic sequences from different vertebrate organisms. Almost all intron positions and phases are conserved between FrDMD and its mammalian counterparts, and the predicted protein product of the Fugu gene displays 55% identity and 71% similarity to human dystrophin. In analogy to the human gene, FrDMD presents several-fold longer than average intronic regions. Analysis of intron sequences of the human and murine genes revealed that they are extremely conserved in size and that a similar fraction of total intron length is represented by repetitive elements; moreover, our data indicate that intron expansion through repeat accumulation in the two orthologs is the result of independent insertional events. The hypothesis that intron length might be functionally relevant to the DMD gene regulation is proposed and substantiated by the finding that dystrophin intron gigantism is common to the three vertebrate genes.
[Supplemental material is available online at www.genome.org.]
The human DMD gene is the largest known to date, spanning >2000 kb on the X chromosome and occupying roughly 0.1% of the genome (Lander et al. 2001). The gene is composed of 79 exons that together account for only 0.6% of its sequence (Ahn and Kunkel 1993). Its main protein product, dystrophin, a member of the spectrin superfamily, is a rod-shaped 427-kD protein consisting of four domains: an N-terminal actin-binding domain, 24 spectrin-like repeats, a cystein-rich domain, and a unique C-terminal domain (Koenig et al. 1988). In skeletal muscle, dystrophin localizes to the cytoplasmic surface of the sarcolemma, where it is thought to provide a link between cytoskeletal actin and the extracellular matrix.
The DMD gene also encodes two nonmuscular full-length isoforms, each controlled by a different promoter located in the 5′region of the gene (Nudel et al. 1989; Gorecki et al. 1992), whereas at least four internal promoters located within introns drive expression of smaller products (Lederfein et al. 1992; Byers et al. 1993; D'Souza et al. 1995; Lidov et al. 1995). Alternative splicing events provide further dystrophin diversification, as the gene product is alternatively spliced throughout its coding sequence (Feener et al. 1989; Bies et al. 1992; Surono et al. 1997; Sironi et al. 2002).
In vertebrates another large gene (Love et al. 1989) encodes utrophin, a protein displaying structure conservation with dystrophin over its entire length, with higher sequence similarity in the C- and N-terminal regions (Tinsley et al. 1992; Pearce et al. 1993). It has been assumed that the two genes were separated by duplication during early vertebrate evolution. Despite high structural homology, the utrophin gene is about one-third the length of the dystrophin gene; this feature does not imply loss of coding information, as all short dystrophin isoforms have counterparts transcribed from the utrophin locus (Blake et al. 1995; Wilson et al. 1999).
Dystrophin-like proteins have been described in both C. elegans and D. melanogaster (Bessou et al. 1998; Greener and Roberts 2000); the corresponding genes have been termed dys-1 and DmDYS (also referred to as dmDRP), respectively and the latter, in analogy to the human locus, also codes for a distal short isoform (Neuman et al. 2001). A sea urchin dystrophin gene has also been described that codes for a full-length protein and a short product, as well (Wang et al.1998).
At the protein level, dystrophin/utrophin-like proteins have been shown to be highly conserved throughout metazoans, indicating a fundamental role in animal biology (Roberts and Bobrow 1998). Yet, the precise function of these proteins is unknown and the molecular mechanisms ensuring proper expression of dystrophin isoforms in different organs and correct alternative splicing events are far from clear.
In humans, mutations in the dystrophin gene are responsible for either Duchenne or Becker muscular dystrophy (DMD and BMD), and the majority of DMD and BMD patients carry deletions in the gene (den Dunnen et al. 1989). The worldwide incidence of DMD is 1 in 3500 male births, one-third of which arise from new mutations (Ahn and Kunkel 1993); it has been speculated that the size of the dystrophin gene might partially account for the high new mutation rate observed. In this view, intron expansion might be regarded as a genetic load, and the question remains open as to whether intron sequences have a role in (or are responsible for) any physiological (or pathological) process.
Comparative genomic sequence analysis offers a powerful strategy for the identification of functionally relevant gene regulatory elements in noncoding regions. Such comparative analyses between human and mouse genomic portions have frequently detected many regions of similarity (Hardison et al. 1997; Dubchak et al. 2000; Gottgens et al. 2000; Loots et al. 2000); yet it is difficult to establish whether they really represent functional elements or are merely the result of too little time for divergence. In this respect, Fugu genome analysis might be of fundamental importance, because it is hypothesized that the large evolutionary distance separating pufferfish and mammals (about 430 million years; Powers 1991) will have resulted in divergence of most sequences except for those of conserved functional importance.
In the present study we report the characterization of the Fugu rubripes dystrophin gene (FrDMD) and draw extensive sequence and gene structure comparison among the human gene and its orthologs from mouse, Fugu, Drosophila, and C. elegans.
RESULTS
Isolation of FrDMD
The Fugu dystrophin gene was isolated as described in Methods; it consists of 82 coding exons with a length varying between 39 and 269 bp and a mean of 133.24. The average intron length is about 1900 bp, with a maximum of 45921 bp (for intron 1) and a minimum of 77 bp. All except one of the introns is flanked by the canonical GT-AG splice-site nucleotide consensus. One intron, between Fugu exons 15 and 16, uses an AAG|gcaag splice donor site. This is the most commonly found ‘atypical’ splice donor site in vertebrate genes (Senapathy et al. 1990). The predicted protein product consists of 3641 residues; pairwise sequence alignment with human dystrophin revealed 55% identity and 71% similarity. The C- and N- terminal regions (with the exclusion of exon 1) display higher conservation (65% and 84% identity, respectively) compared to the rod domain (46%). Pairwise sequence alignment of pufferfish and human dystrophin proteins is available as Supplementary material (Suppl. 1). The first FrDMD exon (sequence: MAEAVRPEDYCDEPVEDEFGEIIKCRS) displays no similarity to any mammalian dystrophin exon 1, and no significant homology to any other peptide was retrieved using a BLASTp search against the NCBI protein database.
The pufferfish gene contains no sequence corresponding to exon 78, and protein alignment with full-length human dystrophin stops at exon 77. This reflects the situation in the zebrafish, where there are only seven terminal amino acids after exon 77 (GGGRLNP), showing no similarity to the mammalian termini (Bolanos-Jimenez et al. 2001). We have not been able to identify this region in the Fugu gene. The identifiable C-terminal portion of the pufferfish protein (60 amino acids, sequence: QDASGLEEVMEQLNNSFPHSQGRSIGSLFH MADDLGRAMESLVSIMTDEQSAEQPEALPL) shows sequence similarity to the unique C-terminal domain of human dystrophin short isoforms derived from the omission of exon 78 from the transcript because of alternative splicing (Feener et al. 1989; Austin et al. 1995; Lidov et al. 1995). A BLASTp search revealed that this sequence also shares homology with the C-terminal region of sea urchin dystrophin (accession no.: AAK20664).
At least four short dystrophin isoforms are transcribed from the human gene and, in order to verify their presence in FrDMD, we searched for potential transcription start sites and first exons located within corresponding introns, using the NIX tool package. The Eponine program predicted the occurrence of a transcription start site within FrDMD intron 66 (corresponding to human intron 62) and, immediately downstream, a putative first exon (sequence: MREQLRK) highly homologous to human Dp71 first exon (sequence: MREQLKG) was identified by the FEX program.
Gene Structure of Dystrophin Orthologous Genes
Data concerning the genomic organization of dystrophin orthologs from F. rubripes, D. melanogaster, C. elegans, and mouse are summarized in Table 1. As expected, the invertebrate dystrophin genes display a totally different organization compared to their vertebrate orthologs, presenting fewer exons of increased length; conversely the human and mouse loci, as well as FrDMD, display striking structure conservation. In all organisms, dystrophin seems to be encoded by relatively small exons, with the exception of Drosophila exons 4 and 9 (3465 and 1041 bp, respectively). Gene length (from the first exon of the muscle isoform to termination codon) was calculated: In the five species, the dystrophin loci cover a considerable portion of total genome size (from 0.03% up to 0.07% for C. elegans and human, respectively) and encode transcripts of similar length, reflecting their high conservation at the protein level (Roberts and Bobrow 1998; this paper). Dramatic differences are evident when the percentage of coding sequence over total gene length is considered: Intron size increases together with organism complexity; less than 0.6% of the human DMD gene is represented by exon sequences, in contrast to 35.6% for dys-1. As far as the nematode gene is concerned, introns do not differ in size from the average length reported for this organism. In contrast, in the other four organisms, intron lengths are several times the respective average lengths. The Drosophila gene is constituted of 35 exons and presents five extremely long intervening sequences (over 5500 bp), whereas the remaining introns can be considered within the average size. Conversely, the Fugu gene, as well as human and mouse dystrophin loci, are characterized by a majority of huge intervening regions and a few small ones. It is well known that the compact genome of the pufferfish owes its small size (400 Mb) to short intergenic and intronic sequences; remarkably, FrDMD intron 1 covers 45 kb, whereas introns 2 and 66 span more than 8 and 12 kb respectively, and 17 introns are longer than 2 kb.
Table 1.
Genomic Organization of Dystrophin Genes
| Organism | Gene length (kb) | Genome % | CDS length | CDS % | Exon number | Mean exon length | Mean intron length | Average exon length | Average intron length |
| C. elegans | 31 | 0.031 | 11,022 | 35.61 | 48 | 232.67 | 424.11 | 299.1 | 466.6 |
| D. melanogaster | 131 | 0.072 | 10,491 | 8.03 | 35 | 314.29 | 3,536.38 | 423.3 | 563.9 |
| F. rubripes | 165 | 0.040 | 10,926 | 6.63 | 82 | 133.24 | 1,900.70 | n.d. | a79.0 |
| M. musculus | 2,256 | 0.065 | 11,034 | 0.49 | 79 | 174.85 | 27,488.59 | 156.3 | 1,321.4 |
| H. sapiens | 2,155 | 0.063 | 11,055 | 0.51 | 79 | 176.57 | 27,160.21 | 152.7 | 3,413.4 |
Mean intron and exon lengths refer to mean values in the dystrophin gene; average lengths represent mean values on available genes. Average exon and intron lengths for Drosophila, C. elegans, mouse and human had been previously reported (Deutsch and Long 1999). All lengths are expressed in bp except for gene lengths, expressed in kb. n.d.: not determined; a: this value was derived from a modal distribution and thus it represents a moda and not a mean; intron length distribution in Fugu displays 75% of introns <425 bp (in humans 75% of introns are <2609 bp) (Aparicio et al. 2002).
Interspecies Comparison
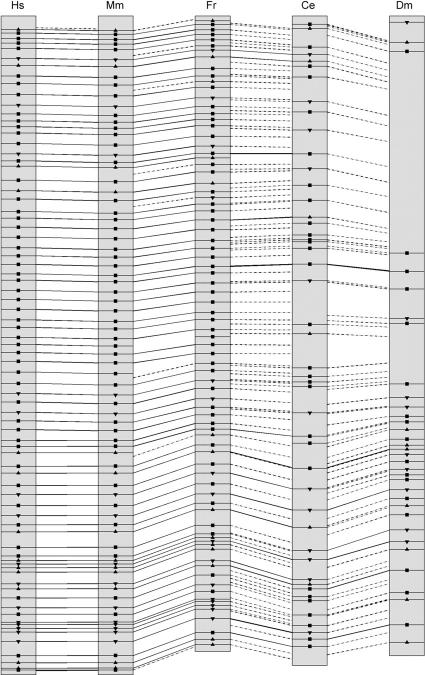
To compare the genomic organization of dystrophin orthologs, we merged gene structure data with CLUSTALW protein alignments. Intron positions with respect to dystrophin coding regions are represented in Figure 1.
Figure 1.
Gene structure comparison among vertebrate and invertebrate dystrophin genes. Intron positions are indicated as lines interrupting the coding sequences. Continuous lines across two sequences represent identical intron positions (small changes of ±10 nt were ignored; Betts et al. 2001). Hatched lines indicated intron positions that are not conserved. Exon codon phases are also reported and represented as squares or triangles. Squares indicate phase 0; up- and down-pointing triangles indicate phase 1 and phase 2, respectively.
As expected, the human and mouse genes show complete conservation of intron/exon junctions, with all introns also preserving equal boundary phases. Intron/exon junctions of the two mammalian genes are also conserved in FrDMD and, in most cases, introns preserve the same insertion pattern with respect to coding frames. Surprisingly, considering the small size of the Fugu genome, four extra introns interrupt the pufferfish gene. In particular, human/mouse exons 6,10, 20, and 23 are split into two smaller exons by insertion of short introns (size ranging from 84–332 bp). Considerably less conservation in structure is observed when the invertebrate dystrophin-like genes are considered. Yet, 15 intron positions are conserved between the nematode gene and FrDMD, with exon codon phases also conserved except in two cases. Finally, as reported above, the 5′ half of DmDYS is characterized by large exons. This feature implies that only one intron position coincides with the nematode gene in this region. Conversely, seven intron/exon boundaries in the C-terminal part of the coding sequence are coincident with those of dys-1.
Overall, 15 intron positions are conserved in C. elegans, Fugu, mouse, and human, whereas only four intron positions, corresponding to introns 30, 52, 56, and 62 of the human gene, are preserved in all five organisms.
Intron lengths were also compared. The mean difference between corresponding intron lengths only amounted to 11.4% when the murine and human genes were considered, the main diversity being accounted for by intron 6 (6856 and 107,479 bp for human and mouse, respectively). Conversely, no correlation was evident between corresponding human and Fugu intron lengths.
We previously demonstrated that, in the human dystrophin gene, out-of-frame (OF) exons are separated by significantly longer introns compared to exons that are predicted to be in-frame (IF), significant differences being accounted for by rod-domain exons (Pozzoli et al. 2002). This same finding can be applied to the murine gene (Table 2): When the whole gene was considered, mean genomic distances amounted to 31,332 and 92,440 bp for IF and OF exons, respectively, resulting in a significant difference (one-way ANOVA; df = 1, P = 0.003). Again, if only rod-domain exons were considered, differences between genomic distances improved the significance (see Table 2). Most interestingly, these same calculations gave similar results when FrDMD was analyzed. Even though the comparison between genomic distances was not significant, there was a considerable difference between IF and OF exons: mean genomic distances being 2350 and 4390 bp, respectively; moreover, also in this case, analysis of rod-domain introns revealed increased differences between the two groups (Table 2).
Table 2.
Comparison Between In-frame and Out-of-frame Exon Genomic Distances
| DMD | Dmd | FrDMD | |||||||
| mean | p | mean | p | mean | p | ||||
| IF | OF | IF | OF | IF | OF | ||||
| whole gene | 31.07 | 72.35 | 0.0064 | 31.33 | 92.44 | 0.0030 | 2.35 | 4.39 | 0.1542 |
| rod domain | 26.80 | 81.68 | 0.0030 | 27.99 | 86.16 | 0.0019 | 1.70 | 2.51 | 0.0840 |
Genomic distances are expressed in kb.
One-way ANOVA was used to assess significance in mean values comparisons.
IF, in-frame; OF, out-of-frame
Comparative Sequence Analysis
It was our particular interest to investigate whether comparative sequence analysis would identify conserved elements within introns that might play a role in regulating dystrophin expression or splicing. For this reason comparative analysis was performed by generating pairwise global sequence alignments of human–mouse and human–Fugu dystrophin genes. The AVID program was used for this purpose. We then used a plotting program that scans the alignment with a sliding window of 100 bp, determines the percentage identity, and moves along the sequence in 25-bp increments. For the human–mouse alignment, a significance cut-off value of ≥80% identity over ≥120 bp was chosen, as previously suggested (Dubchak et al. 2000); multiple regions above threshold were detected within many intron sequences. Conversely, only a few regions displayed more than 50% identity when human and pufferfish sequences were aligned. Yet, many of these regions coincided with above-threshold human–mouse aligned segments and did not represent spurious alignments due to low complexity DNA. A total of 11 regions were detected, displaying more than 50% identity with the pufferfish gene and a significant alignment with mouse; these sequences are located in introns 1, 7, 40, 57, 58, 60, 68 (two regions), 70, 71, and 77; all of them are available as local alignments (Suppl. 2).
As previously reported (Greener et al. 2002), a region displaying high sequence conservation was detected in the 3′ untranslated region of the three genes.
Analysis of Interspersed Repeated Elements
Contributions of repetitive elements to dystrophin intron lengths in all five organisms were calculated and are shown in Table 3. We previously reported a total percentage of 32.1% for the human gene (Pozzoli et al. 2002); we now report a value of 37.43%; it should be noted that an updated version of human repeat libraries was used for the present study and that introns 51–53 have been added (they could not be included in our previous reports as they were not sequenced). Repetitive elements cover a similar fraction of the Dmd gene intron size (36.04%) and, as expected, only a small portion (3.52%) of FrDMD.
Table 3.
Analysis of Repetitive Elements in Dystrophin Intron Sequences
| Organism | Total intron length (kb) | Repeat % | ||||
| simple | species-specific | mammalian | matching | total | ||
| C. elegans | 20 | 5.44 | 10.62 | — | — | 16.06 |
| D. melanogaster | 120 | 2.31 | 0.23 | — | — | 2.54 |
| F. rubripes | 154 | 1.87 | 1.65 | — | — | 3.52 |
| M. musculus | 2,245 | 2.37 | 26.46 | 6.94 | 0.27 | 36.04 |
| H. sapiens | 2,144 | 1.04 | 25.41 | 10.59 | 0.39 | 37.43 |
Simple, simple repeats and low complexity DNA; species-specific, repeated elements specific for either the human or mouse genome; mammalian, mammalian-wide repeated elements; matching, mammalian-wide intersepersed repeats matching in global human–mouse genomic alignments.
Many interspersed repeated elements are restricted to closely related species: About half of the human repeats cannot be identified in genomes of other than primate origin; similarly, most repeats that can be detected in mouse DNA are specific to rodents. Nonetheless, repeated sequences that are common to all mammalian genomes exist, as they probably amplified before the mammalian radiation; conversely no overlap exists between mammals and fish or invertebrate repeat libraries.
We divided identified repeats on the basis of their species distribution (see Table 3), whereas simple repeats and low-complexity regions were considered as a separate group because they can originate in any genome at any time. Moreover, human and mouse genomic sequences were aligned without repeat masking to identify repeat matches; obviously, matching repeats fall into the common repeat group. Remarkably, despite the fact that a similar fraction of total intron size in the human and mouse genes is represented by repeated sequences, the great majority of them are accounted for by species-specific elements with common repeats covering less than 11% and matching sequences less than 0.4%.
We then wished to quantify the association between conservation of noncoding nonrepetitive DNA and repeat density. The human and mouse genes were used for this purpose. As previously described (Chiaromonte et al. 2001), we used a 10-kb sliding window to produce a local evaluation of the fraction of noncoding nonrepetitive nucleotides aligning in the two species, and the fraction of repetitive nucleotides. The locus-level correlation between these two functions was then calculated using the Pearson correlation formula, and a significant value (P < 0.001) of −0.26 was obtained.
DISCUSSION
This study represents, to our knowledge, the first report of gene structure and sequence comparison among dystrophin genomic sequences from different organisms. We sequenced 190 kb of Fugu rubripes genomic DNA corresponding to the complete dystrophin gene and thus allow comparisons among three vertebrate species (human, mouse, and pufferfish) to be performed. The Fugu genome was recently sequenced to over 95% coverage (Aparicio et al. 2002), and it has been reported to be about eight times smaller than that of mammals but to contain a similar number of genes. Beyond an overall paucity of repeated DNA, the pufferfish exhibits a substantial compaction of both introns and intergenic regions. Nonetheless, previous reports indicated that gene structures appear to be conserved between Fugu and human, many intron positions being conserved in orthologous genes (Baxendale et al. 1995; Elgar et al. 1995; Macrae and Brenner 1995). In line with this observation, although FrDMD is about 14 times smaller than the human gene, almost all intron positions and phases are preserved. The only divergence in gene organization is represented by the occurrence of four small extra introns, all located in the 5′ region of the gene. This is not surprising, because both intron gain and loss have already been described in the Fugu lineage (Venkatesh et al. 1999), and differences in intron number per gene between human/pufferfish orthologous pairs have been reported (Aparicio et al. 2002).
In analogy with previous reports of orthologous gene comparison (Baxendale et al. 1995; Mason et al. 1995; Maheshwar et al. 1996), the human and pufferfish dystrophin proteins display 71% similarity, and higher levels of conservation are detectable when C-terminal domains are compared. This observation substantiates previous reports that indicated an extraordinary evolutionary sequence conservation of C-terminal domains in dystrophin family members (Roberts and Bobrow 1998). Conversely, no homology was found between FrDMD exon 1 and any other dystrophin (either vertebrate or invertebrate) first exon. Interestingly, the FrDMD gene lacks any sequence corresponding to exon 78; in humans, this exon is alternatively spliced in short dystrophin isoforms (Feener et al. 1989; Austin et al. 1995; Lidov et al. 1995), and these alternative splicing events cause a frame shift which replaces the 13 C-terminal dystrophin amino acids encoded by exon 79 with 31 new ones. Indeed, the last FrDMD exon shows sequence similarity to the unique C-terminal domain of human dystrophin short isoforms that derive from omission of exon 78 from the transcript. Interestingly, Wang et al. (1998) reported the identification of a sea urchin dystrophin ortholog that also lacks exon 78 and displays sequence homology to FrDMD dystrophin C-terminus; those authors suggested that the minus exon 78 gene might represent the evolutionary original dystrophin form.
Comparative analysis of dystrophin loci gene structure was extended to the murine gene and to two invertebrates. Four introns were found to be preserved in all five organisms and to correspond to introns 30, 52, 54, and 62 of the human/mouse genes. This latter intervening sequence contains the promoter and first exon of short human dystrophin isoform Dp71 that is ubiquitously expressed (Lederfein et al. 1992). At least three additional isoforms are expressed in mammalian species: Dp140, Dp116, and Dp260. The former represents an embryonic isoform that might be important to neural development (Lidov et al. 1995), and the latter two are specific to Schwann cells and retina, respectively (Byers et al. 1993; D'Souza et al. 1995). Transcription of a shorter product from the dmDYS gene has been reported to be driven by a promoter located in intron 16 (Neuman et al. 2001). Wang et al. (1998) reported that the sea urchin gene, which encodes a dystrophin-related protein, also codes for at least one short distal isoform sharing higher similarity to Dp116. To verify whether short dystrophin isoforms might also be predicted to be transcribed from the Fugu gene, we searched for transcription start sites in FrDMD introns. The NIX tool package only allowed prediction of a putative transcription start site within FrDMD intron 66 (corresponding to DMD intron 62) where a first exon with high sequence similarity to human Dp71 first exon could be identified. Thus it might be argued that the only transcribed short isoforms in the pufferfish might be orthologous to Dp71. Recent biochemical evidence (Bolanos-Jimenez et al. 2001) indicated that whole zebrafish embryos seem to express dystrophin proteins corresponding to all four shorter products. Yet, it should be considered that currently available computational methods might not be sensitive enough to perform specialized searches in nonmammalian species (due to the relatively low availability of known fish regulatory elements). Alternatively, the two teleosts might have evolved different dystrophin functions, perhaps reflecting different functional requirements.
An outstanding feature of the human dystrophin gene is its enormous intron size. Our data indicate that intron expansion also occurs in the murine dystrophin gene and that a striking conservation is observed between corresponding intron lengths, as they differ, on average, less than 11.4%. A previous human–mouse comparative analysis of orthologous gene pairs (Batzoglou et al. 2000) indicated that, although both exon number and length are quite well conserved, corresponding intron size tended to vary considerably, the mean ratio of the larger to the small length being 1.5 (i.e., a relative difference of 50%). Fugu rubripes is a teleost fish that separated from the mammalian lineage more than 430 million years ago; this makes the pufferfish one of our most distant extant vertebrate ancestors (Powers 1991). This evolutionarily remote organism displays several times longer than average dystrophin introns, as well. Remarkably, intron expansion in vertebrate dystrophin loci does not proceed at random; in the three organisms, out-of-frame exons are separated by longer introns compared to exons that are predicted to be in-frame. Even if these comparisons reach statistical significance only when the human and mouse genes are considered, differences between IF and OF exons in FrDMD are striking. It was proposed (Bell et al. 1998) that intron length has been exploited in the evolution of genomic structures to represent a further regulatory mechanism of alternative splicing events. As far as dystrophin loci are concerned, this hypothesis might be supported by the finding that differences between genomic distances separating in- and out-of-frame exons are paralleled by differences in splice site strength (Pozzoli et al. 2002); this same finding holds true for both the mouse and pufferfish gene (data not shown) with out-of-frame exons always displaying, on average, stronger splice sites.
Analysis of intron sequences of the human and murine dystrophin genes revealed that a similar fraction of total intron size is represented by repetitive elements. We previously reported that augmented intron size resulting from each repeat insertion in the human gene might have favored further insertions, indicating that accumulation of repetitive elements might be at least in part responsible for intron gigantism (Pozzoli et al. 2002). A similar conclusion can be drawn for the murine ortholog. Yet, our data suggest that the process that led to intron expansion through repeat accumulation in the two orthologs is the result of independent insertional events: Detailed analysis of interspersed repeats revealed that more than 70% of repeated sequences are accounted for by species-specific elements, with less than 0.5% of mammalian-wide repeats matching in corresponding introns. This indicates that, despite the striking conservation of corresponding intron size in the human and murine genes, at least 33% of intron length has been accumulated through independent events. Two hypotheses can be drawn to explain this observation: Either the base composition of dystrophin introns might favor repeated DNA insertion, or intervening sequences might be tolerant of insertional events. A previous study (Chiaromonte et al. 2001) indicated that, if the latter were the case, a correlation between divergence in aligning noncoding nonrepetitive sequences and repeat density should be detected. Those authors proposed that some genome segments are more tolerant of changes of any sort (both point mutations and transpositions) whereas others are rigid and allow only a few modifications. This means that a local negative correlation between repeat density and the number of nucleotides aligning in human and mouse sequences should be verified. This observation can be quantified at the locus level by calculation of the overall correlation between these two functions. When the DMD locus was analyzed, we obtained a significant negative correlation value of −0.26 that substantiates this view.
It has been speculated (den Dunnen et al. 1989), with precise reference to the DMD gene, that evolution would be expected to promote shortening of noncoding sequences that are prone to pathological rearrangements, unless functional elements are located within them. Yet, it has been demonstrated that hypermutable introns in the gene do not necessarily coincide with longer ones (Nobile et al. 1997). In addition, to date only a few deletion/duplication breakpoints in the dystrophin gene have been sequenced and associated with homologous unequal recombination between repeated elements (McNaughton et al. 1998; Sironi et al. 2003). These observations indicate that intron length and repeated element accumulation in dystrophin introns might not be disadvantageous with respect to locus stability. The observation that huge intron sequences are common to three vertebrate dystrophin genes seems to indicate that this feature might be relevant to some unknown cellular process. If expansion of intron sequences in vertebrate dystrophin loci was positively selected for still unknown reasons, insertion of repeated elements in mammalian species might be regarded as a powerful molecular device to rapidly expand intervening regions.
The entry of introns in eukaryotic genomes has been indicated (Mattick 1994, 2001) as the initiation of a new round of molecular evolution that might have paralleled that of protein sequences without interfering with it. This idea was based on the observation that organism complexity has been increasing together with intron number and size, a possible indication of positive selection. Given the small size of most Fugu intervening sequences (Elgar et al. 1996), it has been speculated that not all introns should necessarily be relevant in terms of information content, and the pufferfish was indicated as “an uncluttered system for the identification of and analysis of those introns important for vertebrate gene regulation and development” (Mattick 1994). This concept was recently emphasized (Aparicio et al. 2002) by the finding that the pufferfish genome contains a few giant genes with long introns that might provide insight into the molecular evolution of noncoding regions. Following this line, and given the data reported here, it can be proposed that, given their relatively huge size, dystrophin introns in Fugu rubripes might harbor an informative content, that is, a functional role. Indeed, the analysis and description of other huge genes in pufferfish might help to clarify the role (if any) of such long introns, or explain the selective forces underlying their presence.
The huge size of human dystrophin introns has hampered, until now, the development of any in vitro assay to evaluate the putative presence of functional elements located within them. Comparative genomic sequence analysis offers a powerful strategy for the identification of functionally relevant gene regulatory elements that may be later subjected to experimental evaluation. Conservation of regulatory sequences in noncoding regions between pufferfish and mammalian orthologs has been found in a number of genes, with some elements also displaying equivalent function (Aparicio et al. 1995; Popperl et al. 1995; Barton et al. 2001). On the other hand, such comparative analyses between human and mouse genomic portions frequently result in the detection of many regions of similarity (Hardison et al. 1997; Dubchak et al. 2000; Gottgens et al. 2000; Loots et al. 2000) for which functional relevance is difficult to establish. A previous report (Dubchak et al. 2000) indicated that the cut-off criteria for defining conserved noncoding sequences in pairwise alignments vary depending on the two species that are being compared; for human–mouse genomic alignments, a threshold of ≥80% identity over ≥120 bp was suggested. No threshold for mammalian–fish comparison has ever been indicated. Some examples have been reported of genes in which sequence comparison between Fugu and human orthologous pairs have not revealed any noncoding region of homology, despite the genes having conserved expression patterns and regulatory pathways (Sathasivam et al. 1997; Gellner and Brenner 1999); this may reflect the low sensitivity of currently available computational methods in detecting relatively short conserved regions in a background of extensive sequence divergence. When human–mouse dystrophin global alignments were analyzed, we detected many regions located within introns that satisfied the above cut-off criteria. Conversely, only a few regions displayed ≥50% identity when human–fish alignment was performed. However, many of them also represented above-threshold segments in human–mouse alignments; this is certainly not sufficient to indicate functional relevance for these elements, nonetheless their pattern of conservation can hardly be accidental. It is worth noting that five out of 10 conserved sequences are located in intron regions that are known to be involved in regulated alternative splicing events, namely introns 68, 70, 71, and 77, further substantiating their potential functional role. Experimental studies will be required to demonstrate whether they encode biologically important sequences. Analysis of a fourth genomic sequence would also be of help; in particular, a species at an intermediate evolutionary distance such as chicken (300 million years from mammals) might be informative.
METHODS
Isolation of Fugu rubripes Dystrophin Gene
An FrDMD exon 13 (equivalent to human exon 11) probe was generated by PCR using degenerate primers LRP1F (5′ agg gnt way tga tgg any 3′) and LRP1R (5′ act tns wyt gyt tyt cca t 3′). This was used to probe a Fugu genomic lambda 2001 phage library (G. Elgar, unpubl.). The 3′ end of the gene was isolated from the same phage library using a human exon 75 probe. Subsequently, genomic walks were made from these regions using cosmid and BAC libraries (http://www.hgmp.mrc.ac.uk/geneservice/reagents/products/genomic_resources_non_human/index.shtml), and a contig of the entire gene was constructed. Both the 5′ and 3′ ends were extensively sequenced, and any gaps have now been filled using the Fugu draft assembly (Aparicio et al. 2002).
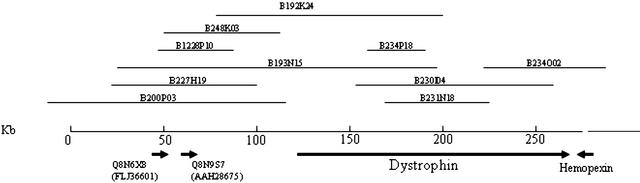
A detailed clone map of the Fugu dystrophin gene region is given in Figure 2.
Figure 2.
BAC clone map of the Fugu dystrophin gene region. The upstream region shows conserved synteny with the human region but the gene immediately downstream, Hemopexin, maps to chromosome 11p15 in human. BAC clones are available through MRC Geneservice (http://www.hgmp.mrc.ac.uk/geneservice/reagents/products/index.shtml).
Identification of FrDMD Exon 1
Despite generating a contig that includes FrDMD flanking genes (the nearest 5′ flanking gene is over 50 kb distal to exon 2), we were unable to identify the first coding exon by homology searches. The first exon of the fish gene was thus identified through 5′-RACE.
5′-RACE was performed on both Tetraodon nigroviridis and Fugu rubripes skeletal muscle cDNA using SMART™ RACE cDNA amplification kit (Clontech) and random hexamers to prime cDNA synthesis. For amplification, a reverse primer was located in exon 3 and a nested reverse in exon 2. Tetraodon and Fugu dystrophin exons 2 and 3 display 96% and 97% identity at the DNA level, respectively, and thus the same primers were used for both organisms, being designed in regions of complete identity. Primer sequences were as follows: Ex3 Rev 5′-GTCGTCCATCACACAGGTCTGAGAACA-3′, Ex2 Rev 5′-CCCATTTTGTGAAAGTCTTCTTCTGAACA-3′.
No product was obtained in the case of Fugu (probably due to the low amount of available RNA), whereas a single RACE band of about 400 bp amplified when T. nigroviridis RNA was used. The PCR product was purified using ExoSAP-IT (Amersham) and directly sequenced with the same primers used for amplification and BigDye™ Terminator Cycle Sequencing (PE Applied Biosystems). Sequences were run on an ABI PRISM 310 Genetic Analyzer. The sequence corresponding to T. nigroviridis dystrophin exon 1 and 5′ UTR is located in FS_CONTIG_18077_1 (http://www.genoscope.cns.fr/externe/tetraodon).
The corresponding Fugu sequence was easily identified by BLASTn search (http://fugu.hgmp.mrc.ac.uk). The predicted peptides in Fugu and Tetraodon display 96% identity.
Tetraodon and Fugu dystrophin exon 1 alignments as well as predicted translations are available (Suppl. 3).
Sequence Retrieval and Analysis
Human and mouse dystrophin genomic sequences can be freely accessed at the UCSC genome pages (http://genome.ucsc.edu/) (June 2002 and February 2002 releases, respectively). Intron/exon boundaries were mapped by BLASTn analysis of cDNAs (accession: NM_004006 for human and NM_007868 for mouse) against genomic sequences.
D. melanogaster and C. elegans dystrophin sequences, as well as intron/exon boundaries, were obtained through BLASTn analysis of cDNAs (accession: NM_079681 and AJ012469, respectively) against corresponding genomic sequences. T. nigroviridis dystrophin sequences are publicly available at the Tetraodon nigroviridis genome analysis pages (http://www.genoscope.cns.fr/externe/tetraodon). To compare genes structures, we identified corresponding exons by mapping genomic sequences onto protein sequence CLUSTALW (Higgins et al. 1994) multiple alignment. We have developed and used programs written in MATLAB to perform all the required tasks and to produce the structure comparison images.
Pairwise global sequence alignment was performed using the AVID program (http://baboon.math.berkeley.edu/∼syntenic/avid.html), and conserved regions were identified by calculating the percentage of identical nucleotides within a 100-nt window moved in 25-nt increments across the alignments.
In all alignments, the muscular human and mouse full-length dystrophin sequences were used.
Analysis of interspersed repetitive elements was performed using a recent update of the RepeatMasker program (http://repeat masker.genome.washington.edu) run under sensitive settings. Specialized repeated element databases were used for each organism under analysis (http://www.girinst.org). Transcription start sites were searched for using the NIX tool package (http://www.hgmp.mrc.ac.uk/Registered/Webapp/nix).
WEB SITE REFERENCES
http://genome.ucsc.edu; UCSC Genome Bioinformatics Site.
http://www.girinst.org; Genetic Information Research Institute Web page.
http://repeatmasker.genome.washington.edu; RepeatMasker Web page.
http://www.hgmp.mrc.ac.uk/geneservice/reagents/products/genomic_resources_non_human/index.shtml; MRC geneservice.
http://www.fugu.hgmp.mrc.ac.uk; HGMP resource centre.
http://baboon.math.berkeley.edu/∼syntenic/avid.html; The AVID alignment program.
http://www.genoscope.cns.fr/externe/tetraodon; Tetraodon nigroviridis genome analysis pages.
Acknowledgments
We are especially grateful to Dr. R. Giorda for precious technical advice and scientific overview. We also thank Dr. M.T. Bassi for useful discussion.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
EMAIL upozzoli@bp.lnf.it; FAX 39 031 877499.
Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.776503.
REFERENCES
- 1.Ahn A.H. and Kunkel, L.M. 1993. The structural and functional diversity of dystrophin. Nat. Genet. 3 283-291. [DOI] [PubMed] [Google Scholar]
- 2.Aparicio S., Morrison, A., Gould, A., Gilthorpe, J., Chaudhuri, C., Rigby, P., Krumlauf, R., and Brenner, S. 1995. Detecting conserved regulatory elements with the model genome of the Japanese puffer fish, Fugu rubripes. Proc. Natl. Acad. Sci. 92 1684-1688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Aparicio S., Chapman, J., Stupka, E., Putnam, N., Chia, J.M., Dehal, P., Christoffels, A., Rash, S., Hoon, S., Smit, A., et al. 2002. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297 1301-1310. [DOI] [PubMed] [Google Scholar]
- 4.Austin R.C., Howard, P.L., D'Souza, V.N., Klamut, H.J., and Ray, P.N. 1995. Cloning and characterization of alternatively spliced isoforms of Dp71. Hum. Mol. Genet. 4 1475-1483. [DOI] [PubMed] [Google Scholar]
- 5.Barton L.M., Göttgens, B., Gering, M., Gilbert, J.G.R, Grafham, D., Rogers, J., Bentley, D., Patient, R., and Green, A.R. 2001. Regulation of the stem cell leukemia (SCL) gene: A tale of two fishes. Proc. Natl. Acad. Sci. 98 6747-6752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Batzoglou S., Pachter, L., Mesirov, J.P., Berger, B., and Lander, E.S. 2000. Human and mouse gene structure: Comparative analysis and application to exon prediction. Genome Res. 10 950-958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Baxendale S., Abdulla, S., Elgar, G., Buck, D., Berks, M., Micklem, G., Durbin, R., Bates, G., Brenner, S., and Beck, S. 1995. Comparative sequence analysis of the human and pufferfish Huntington's disease genes. Nat. Genet. 10 67-76. [DOI] [PubMed] [Google Scholar]
- 8.Bell M.V., Cowper, A.E., Lefranc, M.P., Bell, J.I., and Screaton, G.R. 1998. Influence of intron length on alternative splicing of CD44. Mol. Cell. Biol. 18 5930-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bessou C., Giugia, J.B., Franks, C.J., Holden-Dye, L., and Segalat, L. 1998. Mutations in the Caenorhabditis elegans dystrophin-like gene dys-1 lead to hyperactivity and suggest a link with cholinergic transmission. Neurogenetics. 2 61-72. [DOI] [PubMed] [Google Scholar]
- 10.Betts M.J., Guigo, R., Agarwal, P., and Russell, R.B. 2001. Exon structure conservation despite low sequence similarity: A relic of dramatic events in evolution? EMBO J. 20 5354-5360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bies R.D., Phelps, S.F., Cortez, M.D., Roberts, R., Caskey, C.T., and Chamberlain, J.S. 1992. Human and murine dystrophin mRNA transcripts are differentially expressed during skeletal muscle, heart, and brain development. Nucleic Acids Res. 20 1725-1731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Blake D.J., Schofield, J.N., Zuellig, R.A., Gorecki, D.C., Phelps, S.R., Barnard, E.A., Edwards, Y.H., and Davies, K.E. 1995. G-utrophin, the autosomal homologue of dystrophin Dp116, is expressed in sensory ganglia and brain. Proc Natl. Acad. Sci. 92 3697-3701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bolanos-Jimenez F., Bordais, A., Behra, M., Strahle, U., Mornet, D., Sahel, J., and Rendon, A. 2001. Molecular cloning and characterization of dystrophin and Dp71, two products of the Duchenne Muscular Dystrophy gene, in zebrafish. Gene 274 217-226. [DOI] [PubMed] [Google Scholar]
- 14.Byers T.J., Lidov, H.G.W., and Kunkel, L. M. 1993. An alternative dystrophin transcript specific to peripheral nerve. Nat. Genet. 4 77-81. [DOI] [PubMed] [Google Scholar]
- 15.Chiaromonte F., Yang, S., Elnitski, L., Yap, V.B., Miller, W., and Hardison, R.C. 2001. Association between divergence and interspersed repeats in mammalian noncoding genomic DNA. Proc. Natl. Acad. Sci. 98 14503-14508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.den Dunnen J.T., Grootscholten, P.M., Bakker, E., Blonden, L.A., Ginjaar, H.B., Wapenaar, M.C., van Paassen, H.M., van Broeckhoven, C., Pearson, P.L., and van Ommen, G.J. 1989. Topography of the Duchenne muscular dystrophy (DMD) gene: FIGE and cDNA analysis of 194 cases reveals 115 deletions and 13 duplications. Am. J. Hum. Genet. 45 835-847. [PMC free article] [PubMed] [Google Scholar]
- 17.Deutsch M. and Long, M. 1999. Intron-exon structures of eukaryotic model organisms. Nucleic Acids Res. 27 3219-3228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.D'Souza V.N., Thi Man, N., Morris, G.E., Karges, W., Pillers, D.A.M., and Ray, P.N. 1995. A novel dystrophin isoform is required for normal retinal electrophysiology. Hum. Mol. Genet. 4 837-842. [DOI] [PubMed] [Google Scholar]
- 19.Dubchak I., Brudno, M., Loots, G.G., Pachter, L., Mayor, C., Rubin, E.M., and Frazer, K.A. 2000. Active conservation of noncoding sequences revealed by three-way species comparisons. Genome Res. 10 1304-1306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Elgar G., Rattray, F., Greystrong, J., and Brenner, S. 1995. Genomic structure and nucleotide sequence of the p55 gene of the puffer fish Fugu rubripes. Genomics. 27 442-446. [DOI] [PubMed] [Google Scholar]
- 21.Elgar G., Sandford, R., Aparicio, S., Macrae, A., Venkatesh, B., and Brenner, S. 1996. Small is beautiful: Comparative genomics with the pufferfish (Fugu rubripes). Trends Genet. 12 145-150. [DOI] [PubMed] [Google Scholar]
- 22.Feener C.A., Koenig, M., and Kunkel, L.M. 1989. Alternative splicing of human dystrophin mRNA generates isoforms at the carboxy terminus. Nature 338 509-511. [DOI] [PubMed] [Google Scholar]
- 23.Gellner K. and Brenner, S. 1999. Analysis of 148 kb of genomic DNA around the wnt1 locus of Fugu rubripes. Genome Res. 9 251-258. [PMC free article] [PubMed] [Google Scholar]
- 24.Gorecki D.C., Monaco, A.P., Derry, J.M.J., Walker, A.P., Barnard, E.A., and Barnard, P.J. 1992. Expression of four alternative dystrophin transcripts in brain regions regulated by different promoters. Hum. Mol. Genet. 1 505-510. [DOI] [PubMed] [Google Scholar]
- 25.Gottgens B., Barton, L.M., Gilbert, J.G., Bench, A.J., Sanchez, M.J., Bahn, S., Mistry, S., Grafham, D., McMurray, A., Vaudin, M., et al. 2000. Analysis of vertebrate SCL loci identifies conserved enhancers. Nat. Biotechnol. 18 181-186. [DOI] [PubMed] [Google Scholar]
- 26.Greener M.J. and Roberts, R.G. 2000. Conservation of components of the dystrophin complex in Drosophila. FEBS Lett. 482 13-18. [DOI] [PubMed] [Google Scholar]
- 27.Greener M.J., Sewry, C.A., Muntoni, F., and Roberts, R.G. 2002. The 3-untranslated region of the dystrophin gene—Conservation and consequences of loss. Eur. J. Hum. Genet. 7 413-420. [DOI] [PubMed] [Google Scholar]
- 28.Hardison R.C., Oeltjen, J., and Miller, W. 1997. Long human–mouse sequence alignments reveal novel regulatory elements: A reason to sequence the mouse genome. Genome Res. 7 959-966. [DOI] [PubMed] [Google Scholar]
- 29.Higgins D., Thompson, J., Gibson, T., Thompson, J.D., Higgins, D.G., and Gibson, T.J. 1994. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22 4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Koenig M., Monaco, A.P., and Kunkel, L.M. 1988. The complete sequence of dystrophin predicts a rod-shaped cytoskeletal protein. Cell 53 219-228. [DOI] [PubMed] [Google Scholar]
- 31.Lander E.S., Linton, L.M., Birren, B., Nusbaum, C., Zody, M.C., Baldwin, J., Devon, K., Dewar, K., Doyle, M., FitzHugh, W., et al. 2001. Initial sequencing and analysis of the human genome. Nature 409 860-921. [DOI] [PubMed] [Google Scholar]
- 32.Lederfein D., Levy, Z., Augier, N., Mornet, D., and Morris, G. 1992. A 71-kd protein is a major product of the Duchenne muscular dystrophy gene in brain and other nonmuscle tissues. Proc. Natl. Acad. Sci. 89 5346-5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lidov H.G.W., Selig, S., and Kunkel, L.M. 1995. Dp140: A novel 140-kDA CNS transcript from the dystrophin locus. Hum. Mol. Genet. 4 329-335. [DOI] [PubMed] [Google Scholar]
- 34.Loots G.G., Locksley, R.M., Blankespoor, C.M., Wang, Z.E., Miller, W., Rubin, E.M., and Frazer, K.A. 2000. Identification of a coordinate regulator of interleukins 4, 13, and 5 by cross-species sequence comparisons. Science 288 136-140. [DOI] [PubMed] [Google Scholar]
- 35.Love D.R., Hill, D.F., Dickson, G., Spurr, N.K., Byth, B.C., Marsden, R.F., Walsh, F.S., Edwards, Y.H., and Davies, K.E. 1989. An autosomal transcript in skeletal muscle with homology to dystrophin. Nature 339 55-58. [DOI] [PubMed] [Google Scholar]
- 36.Macrae A.D. and Brenner, S. 1995. Analysis of the dopamine receptor family in the compact genome of the puffer fish Fugu rubripes. Genomics 25 436-446. [DOI] [PubMed] [Google Scholar]
- 37.Maheshwar M.M., Sandford, R., Nellist, M., Cheadle, J.P., Sgotto, B., Vaudin, M., and Sampson, J.R. 1996. Comparative analysis and genomic structure of the tuberous sclerosis 2 (TSC2) gene in human and pufferfish. Hum. Mol. Genet. 5 562. [PubMed] [Google Scholar]
- 38.Mason P.J., Stevens, D.J., Luzzatto, L., Brenner, S., and Aparicio, S. 1995. Genomic structure and sequence of the Fugu rubripes glucose-6-phosphate dehydrogenase gene (G6PD). Genomics 26 587-591. [DOI] [PubMed] [Google Scholar]
- 39.Mattick J.S. 1994. Introns: Evolution and function. Curr. Opin. Genet. Dev. 4 823-831. [DOI] [PubMed] [Google Scholar]
- 40.Mattick J.S. 2001. Noncoding RNAs: The architects of eukaryotic complexity. EMBO Rep. 2 986-991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.McNaughton J.C., Cockburn, D.J., Hughes, G., Jones, W.A., Laing, N.G., Ray, P.N., Stockwell, P.A., and Petersen, G.B. 1998. Is gene deletion in eukaryotes sequence-dependent? A study of nine deletion junctions and nineteen other deletion breakpoints in intron 7 of the human dystrophin gene. Gene 222 41-51. [DOI] [PubMed] [Google Scholar]
- 42.Neuman S., Kaban, A., Volk, T., Yaffe, D., and Nudel, U. 2001. The dystrophin/utrophin homologues in Drosophila and in sea urchin. Gene 263 17-29. [DOI] [PubMed] [Google Scholar]
- 43.Nobile C., Marchi, J., Nigro, V., Roberts, R.G., and Danieli, G.A. 1997. Exon-intron organization of the human dystrophin gene. Genomics 45 421-424. [DOI] [PubMed] [Google Scholar]
- 44.Nudel U., Zuk, D., Einat, P., Zeelon, E., Levy, Z., Neuman, S., and Yaffe, D. 1989. Duchenne muscular dystrophy gene product is not identical in muscle and brain. Nature 337 76-78. [DOI] [PubMed] [Google Scholar]
- 45.Pearce M., Blake, D.J., Tinsley, J.M., Byth, B.C., Campbell, L., Monaco, A.P., and Davies, K.E. 1993. The utrophin and dystrophin genes share similarities in genomic structure. Hum. Mol. Genet. 2 1765-1772. [DOI] [PubMed] [Google Scholar]
- 46.Popperl H., Bienz, M., Studer, M., Chan, S.K., Aparicio, S., Brenner, S., Mann, R.S., and Krumlauf, R. 1995. Segmental expression of Hoxb-1 is controlled by a highly conserved autoregulatory loop dependent upon exd/pbx. Cell 81 1031-1042. [DOI] [PubMed] [Google Scholar]
- 47.Powers D.A. 1991. Evolutionary genetics of fish. Adv. Genet. 29 119-228. [DOI] [PubMed] [Google Scholar]
- 48.Pozzoli U., Sironi, M., Cagliani, R., Comi, G.P., Bardoni, A., and Bresolin, N. 2002. Comparative analysis of the human dystrophin and utrophin gene structures. Genetics 160 793-798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Roberts R.G. and Bobrow, M. 1998. Dystrophins in vertebrates and invertebrates. Hum. Mol. Genet. 7 589-595. [DOI] [PubMed] [Google Scholar]
- 50.Sathasivam K., Baxendale, S., Mangiarini, L., Bertaux, F., Hetherington, C., Kanazawa, I., Lehrach, H., and Bates, G.P. 1997. Aberrant processing of the Fugu HD (FrHD) mRNA in mouse cells and in transgenic mice. Hum. Mol. Genet. 6 2141-2149. [DOI] [PubMed] [Google Scholar]
- 51.Senapathy P., Shapiro, M.B., and Harris, N.L. 1990. Splice junctions, branch point sites, and exons: Sequence statistics, identification, and applications to genome project. Methods Enzymol. 183 252-278. [DOI] [PubMed] [Google Scholar]
- 52.Sironi M., Cagliani, R., Pozzoli, U., Bardoni, A., Comi, G.P., Giorda, R., and Bresolin, N. 2002. The dystrophin gene is alternatively spliced throughout its coding sequence. FEBS Lett. 517 163-166. [DOI] [PubMed] [Google Scholar]
- 53.Sironi M., Pozzoli, U., Cagliani, R., Giorda, R., Comi, G.P., Bardoni, A., Menozzi, G., and Bresolin, N. 2003. Relevance of sequence and structure elements for deletion events in the dystrophin gene major hot-spot. Hum. Genet. 112 272-288. [DOI] [PubMed] [Google Scholar]
- 54.Surono A., Takeshima, Y., Wibawa, T., Premono, Z.A., and Matsuo, M. 1997. Six novel transcripts that remove a huge intron ranging from 250 to 800 kb are produced by alternative splicing of the 5′ region of the dystrophin gene in human skeletal muscle. Biochem. Biophys. Res. Commun. 239 895-899. [DOI] [PubMed] [Google Scholar]
- 55.Tinsley J.M., Blake, D.J., Roche, A., Fairbrother, U., and Riss, J. 1992. Primary structure of dystrophin-related protein. Nature 360 591-593. [DOI] [PubMed] [Google Scholar]
- 56.Venkatesh B., Ning, Y., and Brenner, S. 1999. Late changes in spliceosomal introns define clades in vertebrate evolution. Proc. Natl. Acad. Sci. 96 10267-10271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Wang J., Pansky, A., Venuti, J.M., Yaffe, D., and Nudel, U. 1998. A sea urchin gene encoding dystrophin-related proteins. Hum. Mol. Genet. 7 581-588. [DOI] [PubMed] [Google Scholar]
- 58.Wilson J., Putt, W., Jimenez, C., and Edwards, Y.H. 1999. Up71 and Up140, two novel transcripts of utrophin that are homologues of short forms of dystrophin. Hum. Mol. Genet. 8 1271-1278. [DOI] [PubMed] [Google Scholar]