Abstract
Precise genome engineering via homologous recombination (HR)-mediated gene targeting (GT) has become an essential tool in molecular breeding as well as in basic plant science. As HR-mediated GT is an extremely rare event, positive–negative selection has been used extensively in flowering plants to isolate cells in which GT has occurred. In order to utilize GT as a methodology for precision mutagenesis, the positive selectable marker gene should be completely eliminated from the GT locus. Here, we introduce targeted point mutations conferring resistance to herbicide into the rice acetolactate synthase (ALS) gene via GT with subsequent marker excision by piggyBac transposition. Almost all regenerated plants expressing piggyBac transposase contained exclusively targeted point mutations without concomitant re-integration of the transposon, resulting in these progeny showing a herbicide bispyribac sodium (BS)-tolerant phenotype. This approach was also applied successfully to the editing of a microRNA targeting site in the rice cleistogamy 1 gene. Therefore, our approach provides a general strategy for the targeted modification of endogenous genes in plants.
Keywords: gene targeting, marker excision, piggyBac transposon, acetolactate synthase, cleistogamy 1, Oryza sativa, technical advance
Introduction
Gene targeting (GT) via HR between an endogenous gene and an exogenous targeting vector, and GT via mismatch repair with chimeric RNA/DNA oligonucleotide (chimeraplast) are established techniques that allow desired mutations to be introduced into target genes. Unless the inserted mutations confer a selectable trait, such as resistance to herbicides, HR-mediated GT with a positive–negative selection system is the current method of choice for selection of GT cells in several organisms. In flowering plants, HR-mediated GT with a positive–negative selection system has been developed as a universal approach and has been used successfully to modify several genes (Terada et al., 2002, 2007; Yamauchi et al., 2009; Moritoh et al., 2012; Ono et al., 2012; Dang et al., 2013). However, when using a positive–negative selection system, the positive selection marker gene inserted into the targeted locus should be removed completely from the GT locus if the intention is to introduce desired mutations exclusively.
Site-specific recombination systems such as the Cre/loxP and FLP/FRT system have been utilized widely to remove selectable marker genes from GT loci in mammals and plants (van der Weyden et al., 2002; Terada et al., 2010; Dang et al., 2013). However, site-specific recombination systems leave dispensable sequences, e.g., the 34-bp recognition sequences of site-specific recombinase (a single loxP and FRT site), at the excised site. Furthermore, such sequences have the potential to affect expression of adjacent genes in mammalian cells (Meier et al., 2010). In addition, it has been reported that the frequency of Cre/loxP-mediated marker elimination from a GT locus by transiently expressing Cre recombinase in rice was quite low (Terada et al., 2010; Dang et al., 2013).
In contrast, a system using a piggyBac transposon derived from the cabbage looper moth afforded precise genome modification via GT and subsequent marker excision in mammalian cells (Yusa et al., 2011a; Morioka et al., 2014; Sun and Zhao, 2014). In our previous study, we designed an assay system that allows piggyBac transposition to be visualized as luciferase luminescence in rice cells, and demonstrated that the piggyBac transposon is capable of accurate and effective transposase-mediated transposition also in plant cells (Nishizawa-Yokoi et al., 2014). In addition, we have demonstrated piggyBac transposition-mediated excision of a selectable marker from the reporter locus at high frequency without concomitant re-integration of the transposon (Nishizawa-Yokoi et al., 2014). Here, we applied piggyBac-mediated marker excision system to remove a positive selectable marker gene from a GT locus in rice.
Results
Introduction of point mutations into the ALS gene via GT using a positive–negative selection system
To date, we have successfully generated rice plants tolerant to the herbicide bispyribac sodium (BS) by introducing two point mutations specifying two amino acid changes – tryptophan (TGG) to leucine (TTG) at amino acid 548 (W548L), and serine (AGT) to isoleucine (ATT) at amino acid 627 (S627I) – in the ALS locus via GT, since the infrequent GT cells can be selected easily on medium containing BS (Endo et al., 2007). To establish a universal strategy for producing mutant plants harboring only the desired mutation in the target locus, we attempted to introduce the W548L and S627I mutations into the ALS gene locus via GT with positive–negative selection and subsequent excision of the positive selectable marker gene from the ALS gene locus using piggyBac transposon. In rice plants, a strong positive–negative selection system using the hpt gene conferring resistance to hygromycin B as positive selection marker and diphtheria toxin A subunit gene (DT-A) as a negative selection marker has been developed for the selection of GT-positive cells (Terada et al., 2002). The GT vector carries DT-A gene expression cassettes at both sides and a 6.4-kb fragment containing an ALS coding region with W548L and S627I mutations. The piggyBac transposon integrates into the host genome at TTAA elements and excises without leaving a footprint at the excised site (Cary et al., 1989). Therefore, piggyBac transposon harboring a rice actin terminator and hpt expression cassette was inserted at an HpaI site with two additional silent mutations at the junctions to make an HpaI site (gTTAAc) for piggyBac transposition (Figure 1a). The ALS gene modified with GT should be inactive since the ALS coding gene is interrupted by the hpt gene expression cassette flanked by sequences needed for piggyBac-mediated transposition.
Figure 1.

Strategy for the introduction of point mutations into the ALS locus via GT and subsequent marker excision from the GT locus using piggyBac transposon.
(a) Schematic diagram of GT at the ALS locus. The top line indicates the genomic structure of the wild-type ALS gene region. The bottom line shows the T-DNA region of the targeting vector carrying diphtheria toxin A subunit gene (DT-A) under the control of the maize polyubiquitin 1 promoter (Pubi) or rice elongation factor-1α promoter (Pef) as negative selection marker and a 6.4-kb fragment containing an ALS coding region (open box) with W548L and S627I mutations (red lines) and silent mutations (added HpaI site at 301-bp upstream of W548L; GCTGAC to GAATTC) for the insertion of piggyBac transposon (black triangle) harboring a rice actin terminator (T) and hpt gene under the control of the cauliflower mosaic virus 35S promoter (P35S) as positive selection marker. LB, left border; RB, right border.
(b) Strategy for precise marker excision using piggyBac transposon from the GT locus. The top line reveals the structure of the modified ALS locus resulting from homologous recombination between the targeting vector and wild-type locus. The bottom line represents the ALS locus modified by GT and subsequent precise marker excision via piggyBac transposition. The primer sets used for PCR that identify transgenic calli in which a GT event occurred at ALS locus are shown as black arrows. White arrows indicate the primer sets used for CAPS analysis to evaluate the frequency of marker excision via piggyBac transposition. Gray arrows represent primers for PCR analysis to detect the existence of re-integrated piggyBac transposon. The numbers on each arrow reveal the length of the PCR fragments.
(c) Sequencing chromatograms of the excision site and mutation site in T0 plants.
(d–f) Southern blot analysis with probe1 (d), 2 (f), and 3 (e) shown in (a) and (b) using MfeI-digested genomic DNA of wild-type, a regenerated plant of ALS GT-A and two T0 plants each of two independent lines with ALS GT-A_hy-6 and 10 (ALS GT-A_hy-6-1, 6-2, 10-1 and 10-2).
Rice calli derived from mature seeds were inoculated with Agrobacterium harboring the GT vector and were selected on medium containing hygromycin B for 4 weeks (Figure S1). In total, 100 independent hygromycin-resistant calli were selected from 3259 (approximately 20 g) pieces of calli (Table 1) and were subjected to polymerase chain reaction (PCR) analysis with the primer sets shown in Figure 1(b) to identify transgenic calli in which GT events had occurred at the ALS locus. Both upstream and downstream junction fragments were detected in six independent selected callus lines, indicating that the positive selection marker gene was introduced into the ALS locus by homologous recombination (HR) between the GT vector and the target locus. Sequencing analysis revealed a lack of W548L or W548L/S627I mutations in the targeted ALS locus in two callus lines (ALS GT-B4 and B5), respectively. Four callus lines (ALS GT-A, B1, B2 and B3) were identified as GT calli (Table 1), and among them, two lines (ALS GT-A and B1) were used for the marker excision study.
Table 1.
Summary of GT experiments targeting the ALS locus
| Experiments | No. of Agrobacterium infected calli | No. of hygromycin- resistant calli | No. of targeted calli | No. of targeted calli with W548L/S627I mutations |
|---|---|---|---|---|
| A | 1463 (7.53 g) | 25 | 1 | 1 |
| B | 1796 (11.85 g) | 75 | 5 | 3 |
| Total | 3259 (19.38 g) | 100 | 6 | 4 |
Precise marker excision from the GT locus by piggyBac transposition
ALS GT-A and B1 calli were infected with Agrobacterium harboring a hyperactive piggyBac transposase (hyPBase) (Yusa et al., 2011a, b) expression vector driven by the constitutive maize polyubiquitin gene 1 (Ubi-1) promoter (designated ALS GT-A_hy and B1_hy, Figure S1). Transgenic calli were selected on N6D medium with geneticin and meropenem, and transferred to regeneration medium with meropenem. Batches of twenty T0 regenerated plants from five or six independent ALS GT-A_hy and B1_hy lines, respectively, were subjected to marker excision analysis by cleaved amplified polymorphic sequences (CAPS) that combined PCR amplification and restriction digestion with MfeI as described in Experimental Procedures.
The two point mutations in the targeted ALS locus generate recognition sites for the restriction enzyme MfeI (CAATTG). Therefore, MfeI digestion of PCR fragments derived from the wild-type ALS locus or GT-modified ALS locus and subsequent marker excision is expected to generate a non-digested 3.8-kb fragment or fragments of 2.9-, 0.7- and 0.2-kb, respectively (Figure 1a,b). CAPS analysis with genomic DNA extracted from leaves of ALS GT-A_hy and B1_hy revealed the expected MfeI-digested fragment in candidate marker-free rice plants containing W548L/S627I mutations in the ALS locus. More than 90% of regenerated plants (on average, 100 and 92.5% of ALS GT-A_hy and B1_hy, respectively; Tables 2 and S1) contained MfeI-digested fragments, indicating that the piggyBac has transposed effectively from the targeted ALS locus via hyPBase. Furthermore, whether the excised piggyBac transposon is re-integrated into other loci was analyzed by PCR using primers specific to the hpt coding region generating a 0.9-kb PCR fragment (Figure 1b). Only one plant of each ALS GT-A_hy and B1_hy regenerated plants harboring the MfeI-digested fragment by CAPS analysis contained hpt-specific fragments (Tables 2 and S1). Thus, 99 and 92% of ALS GT-A_hy and B1_hy regenerated plants, respectively, were found to be marker-free rice plants containing two point mutations W548L and S627I in the ALS locus. Direct sequencing of PCR fragments from eight and 16 randomly selected plants of ALS GT-A_hy and B1_hy revealed the presence of W548L/S627I mutations and silent mutations (added HpaI site) for insertion of the piggyBac transposon in all regenerated plants analyzed (Figure 1c). Therefore, our findings indicate that mutant plants containing desired point mutations and lacking residual sequences in the target locus were obtained successfully in a convenient manner.
Table 2.
PCR analysis of piggyBac excision and re-integration events in ALS GT-A_hy regenerated plants by hyPBase expression
|
piggyBac excision from OsALS locus |
Frequency of piggyBac excision (%) |
||||||
|---|---|---|---|---|---|---|---|
| Line no. | No. of T0 plants analyzed | Without marker | With marker | Total | Without re-integration | With re-integration | Total |
| 5 | 20 | 19 | 1 | 20 | 95.0 | 5.0 | 100 |
| 6 | 20 | 20 | 0 | 20 | 100 | 0 | 100 |
| 10 | 20 | 20 | 0 | 20 | 100 | 0 | 100 |
| 13 | 20 | 20 | 0 | 20 | 100 | 0 | 100 |
| 24 | 20 | 20 | 0 | 20 | 100 | 0 | 100 |
| Average | 99.0 | 1.0 | 100 | ||||
To confirm the introduction of W548L/S627I mutations via GT and the excision of the selectable marker gene by piggyBac excision in the ALS locus in T0 plants, we performed Southern blot analysis with genomic DNA extracted from wild-type, a regenerated plant of ALS GT-A and two T0 plants each of two independent lines with ALS GT-A_hy-6 and 10. Southern blot analysis of MfeI-digested DNA with probe1 (Figure 1d) showed that wild-type bands (11.8-kb) and bands indicating gene targeting (6.8 kb) were detected in ALS GT-A regenerated plants (Figure 1d), while wild-type bands (11.8 kb) and 4.8-kb bands derived from piggyBac-excised targeted ALS locus were detected using probe1 in ALS GT-A_hy regenerated plants (Figure 1d). In addition to the wild-type band (11.8-kb), 6.8-kb bands derived from the ALS locus containing W548L/S627I mutations via GT were observed using probe3 in ALS GT-A and GT-A_hy regenerated plants (Figure 1e). Furthermore, Southern blot analysis with an hpt probe revealed 2.3-kb bands derived from a positive selection marker only in ALS GT-A, indicating that excised piggyBac had not re-integrated into other loci in ALS GT-A_hy regenerated plants (Figure 1f). Consistent with these results, we confirmed the introduction of W548L/S627I mutations without any piggyBac footprint in the ALS locus in T0 plants of ALS GT-B1_hy-19 and 20 by Southern blot analysis (Figure S2a–c).
To further verify our GT-mediated precision mutagenesis system, a single base substitution (CAGCAGCA/GTCATCACGATTCC) was introduced by GT into the microRNA targeting site of the rice cleistogamy 1 (Oscly1) gene (Chen, 2004; Nair et al., 2010) – an orthologue of Arabidopsis thaliana AP2 transcription factor involved in the specification of floral organ identity (Jofuku et al., 1994) – and the selectable marker was subsequently excised from the GT locus using piggyBac transposon (Figure S3a,b). Four independent callus lines were identified as GT calli from 5139 (approximately 25.6 g) pieces of calli by PCR analysis with the primer sets shown in Figure S3(b) and subsequent sequence analysis (Table S2). Two independent lines (cly1 GT-1 and -2) were infected with Agrobacterium harboring a hyPBase expression vector (designated cly1 GT-1_hy and GT-2_hy) and were subjected to marker excision analysis. Via PCR with the primer sets shown in Figure S3(b) and sequence analysis, we confirmed that 91% and 98% of cly1 GT-1_hy and GT-2_hy regenerated plants, respectively, contained the desired A to G mutation in the microRNA targeting site of the Oscly1 gene but not the positive selectable marker gene (Figure S3a–c and Tables S3 and S4). Furthermore, Southern blot analysis revealed that all the regenerated plants so far analyzed were marker-free plants, and that there was no re-integration of piggyBac transposon (Figures S3d–f). Taken together, these results validated accurate and effective piggyBac-mediated marker excision from the Oscly1 locus. Detailed characterization of Oscly1-mutated rice will be presented elsewhere.
Analysis of marker-free T1 progeny harboring an ALS gene modified via GT
T1 progeny plants were obtained from self-pollinating ALS GT-A_hy-10 T0 plants. Genomic DNA was extracted from four progenies of ALS GT-A_hy-10 T0 regenerated plants and subjected to Southern blot analysis. Analysis of MfeI-digested DNA with probe1 or probe3 (Figure 1b) showed that a plant with wild-type ALS locus (ALS GT-A_hy 10-1), heterozygous plants (ALS GT-A_hy 10-2, -3), and a homozygous (ALS GT-A_hy 10-4) plant with the modified ALS locus were obtained (Figure 2a,b). All of these T1 progeny lacked the hpt-specific fragment, indicating that the piggyBac transposon with the selectable marker gene had transposed and excised from the ALS locus without re-integration (Figure 2c). Furthermore, a hyPBase-specific fragment segregated out and was not detected in ALS GT-A_hy 10-3 calli by Southern blot analysis (Figure 2d,e).
Figure 2.
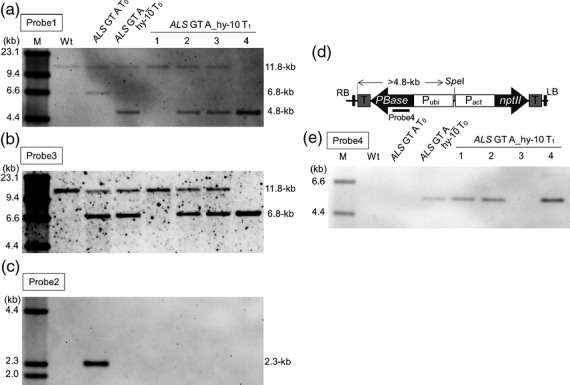
Segregation of the ALS gene harboring W548L/S627I mutations and hyPBase expression vector in T1 progeny.
(a–c) Southern blot analysis with probe 1 (a), 2 (c), 3 (b) shown in Figure 1(a,b) using MfeI-digested genomic DNA of wild-type, a regenerated plant of ALS GT-A and ALS GT-A_hy-10, and four T1 progeny of ALS GT-A_hy-10 (ALS GT-A_hy-10-1, 2, 3, and 4). Southern blot analysis revealed that the ALS locus with W548L/S627I segregated in T1 progenies of ALS GT-A_hy-10 (wild-type ALS gene, line no. 1; heterozygous modification of the ALS gene, line no. 2 and 3; homozygous modification of ALS gene, line no. 4).
(d) Schematic diagram of hyPBase expression vector. The hyPBase expression vector carries a rice Ubi-1 promoter (Pubi)::hyPBase (PBase) cassette and a rice actin promoter (Pact)::nptII expression cassette. T, terminator; LB, left border; RB, right border.
(e) Southern blot analysis with probe 4 shown in (d) using SpeI-digested genomic DNA of wild-type, a regenerated plant of ALS GT-A and ALS GT-A_hy-10, and four T1 progeny of ALS GT-A_hy-10 (ALS GT-A_hy-10-1, 2, 3, and 4). The hyPBase expression vector segregated out in ALS GT-A_hy 10-3.
To examine whether the introduction of W548L and S627I mutations into the ALS gene via GT confers herbicide BS-tolerant phenotype in these T1 progeny plants, ALS gene transcript levels in rice calli derived from ALS GT-A_hy 10-1 (wild-type ALS locus, ALS-wt), 10-2, 10-3 (heterozygous plants, GT-hetero), and 10-4 (homozygous with the modified ALS locus, GT-homo) lines were analyzed by reverse transcription-polymerase chain reaction (RT-PCR) and BS-susceptibility test. We confirmed that there was no change in ALS gene transcription levels between ALS-wt, GT-hetero and GT-homo calli (Figure 3a–c). After 3-weeks of culture on callus induction medium containing 0.75 μm BS, GT-hetero and GT-homo calli with the modified ALS gene showed BS tolerance (Figure S4). Furthermore, the acquisition of BS tolerance was confirmed in segregated T1 progeny plants (Figure 3d).
Figure 3.
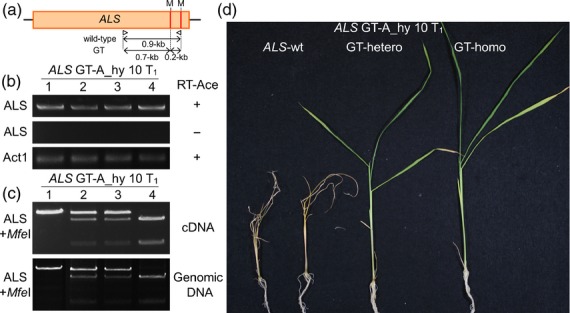
Analysis of the ALS gene harboring W548L/S627I mutations in T1 progeny.
(a) Diagram showing the targeted ALS locus. Arrowheads indicate the primers used for the reverse transcriptase-polymerase chain reaction (RT-PCR, b) and CAPS (c). The expected band sizes of the RT-PCR (0.9-kb) and CAPS (wild-type, 0.9-kb; GT 0.7-kb and 0.2-kb) are shown.
(b) Transcript levels of the ALS gene in T1 plants carrying the wild-type (line no. 1) and modified ALS gene (heterozygous, lines no. 2, 3 or homozygous, line no. 4). Top and middle panels show RT-PCR analysis using the ALS gene-specific primers with (top) or without (middle) reverse transcriptase (RT-Ace + or −, respectively). The Actin 1 (Act1) gene was used as an internal control (bottom panel).
(c) CAPS analysis combining PCR analysis using ALS gene-specific primers with cDNA (top) or genomic DNA (bottom) and MfeI digestion in T1 plants carrying the wild-type (line no. 1) or modified ALS gene (heterozygous, lines no. 2, 3 or homozygous, line no. 4).
(d) Herbicide bispyribac (BS)-tolerant phenotype of T1 plants. GT line A_hy T1 plants carrying the modified ALS gene [either heterozygous (GT-hetero) or homozygous (GT-homo)] showed BS tolerance after 3 weeks of BS treatment, but not T1 plants carrying the wild-type ALS gene (ALS-wt).
Discussion
The efficiency of positive–negative selection with the hpt and DT-A genes (GT-positive calli per hygromycin-resistant calli: 4.0%, Table 1) in this study was comparable to previous results (0.053–5.3%) (Terada et al., 2002, 2007; Yamauchi et al., 2009; Moritoh et al., 2012; Ono et al., 2012; Dang et al., 2013). Our previous report showed that approximately 1500 rice calli were transformed by Agrobacterium harboring a GT vector carrying the 5′-truncated ALS coding region with two mutations (W548L/S627I) and the downstream region of the ALS gene – namely a target gene-dependent selection – and 66 independent GT plants were obtained by selection on medium containing BS (Endo et al., 2007). Differences in the GT frequency between the previous study and this present work (GT-positive calli per callus infected by Agrobacterium: 4/3259) might be attributed to the integration of multiple T-DNA copies into the host genome, since random integration of the GT vector concomitant with sequence-specific integration via HR kills GT cells due to expression of the DT-A gene. In fact, extra bands in addition to the true GT fragment were detected by Southern blot analysis in GT lines obtained without negative selection (Endo et al., 2007). Saika and Toki (2010) reported that single copy T-DNA integration was observed to have occurred in <40% of isolated secondary calli of Nipponbare (on average, 2.6 T-DNA copies per transgenic callus). In addition, DT-A proteins directly cause lethality in cells, suggesting that transient expression of DT-A might be one of the causes of loss of GT frequency.
Using a GT procedure with positive–negative selection, an entire hpt expression cassette should be inserted into the ALS locus by HR regardless of the presence or absence of the two point mutations. In circumstances where GT occurs in somatic plant cells, DNA double-strand breaks (DSBs) are thought to be repaired mainly by synthesis-dependent strand annealing (SDSA) using the targeting vector as a repair template (Puchta and Fauser, 2013). In SDSA, single-strand overhangs with the 3′ end of the DSB site invade the repair template to form a D-loop structure and copy genetic information. Subsequently, 3′ ends are detached from the D-loop structure and re-anneal with the second strand at the DSB site. According to the SDSA model, the exogenous DNA information – hpt expression cassette – on the repair template is thought to inhibit homology search and homologous pairing between the 3′ end of the DSB site in the ALS locus and the GT vector at the step of strand invasion. In addition, the existence of a large exogenous DNA stretch within sequences homologous to the ALS locus may lead to a one-sided invasion, in which one end of the DSB is repaired by HR and the other end is rejoined by non-homologous end-joining (NHEJ). Because DNA synthesis is halted to allow copying of the repair template and resolution of the D-loop structure on hpt expression cassette is assumed to occur, the 3′ end of the DSB cannot re-anneal with the second strand at the DSB site. Thus, the extent of exogenous DNA information that needs to be copied/pasted into the target locus using positive–negative selection is larger than that needed in target gene-dependent selection, resulting in a decreased frequency of GT with positive–negative selection.
Consistent with previous reports (Endo et al., 2007; Johzuka-Hisatomi et al., 2008; Saika et al., 2011), the lack of either or both of the desired W548L/S627I mutations in the ALS gene were found in two callus lines out of six independently selected callus lines harboring the positive selection marker in the ALS locus (Table 1). Therefore, our findings could be explained by the occurrence of mismatch correction of the heteroduplex molecules formed between the genomic DNA and targeting vector or resolution of D-loop structures between the positive selection marker cassette and the W548L mutation or W548L and S627I mutations.
About 30% of piggyBac excision events from GT loci were reported to be accompanied by re-integration of piggyBac in mammalian cells under negative selection (Yusa et al., 2011a, b). In contrast, re-integration of excised piggyBac from the targeted ALS locus was observed in only 1% of calli analyzed (Tables 2 and S1). Our results might derive from expression of hyPBase protein from the stably integrated hyPBase gene under a strong promoter, i.e. re-integrated piggyBac transposon could be transferred again before finally being deleted from the plant chromosome. While, excision of a selection marker via hyPBase-induced piggyBac transposition from a reporter locus was detected previously in 72 and 91% of cases in two independent lines of transgenic T0 plants (Nishizawa-Yokoi et al., 2014). In this report, the frequency of excision of the piggyBac transposon from the GT locus was consistently over 90% (Tables 2, S1, S3 and S4). Our data raise the possibility that piggyBac can transpose easily from the GT locus regardless of the location of piggyBac insertion in the genome. This hypothesis can be explained by the findings of two previous reports: (i) piggyBac transposition has been reported to be inhibited by DNA methylation in mouse cells (Wang et al., 2008); and (ii) the DNA methylation profile of genomic loci modified by GT is altered in Arabidopsis (Lieberman-Lazarovich et al., 2013).
Recent developments in sequence-specific nucleases (SSNs) such as zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs) and the bacterial clustered regularly interspaced short palindromic repeats (CRISPR)/Cas system, have enabled targeted DNA modification of plant genomes (Chen and Gao, 2013; Gaj et al., 2013; Voytas, 2013; Kim and Kim, 2014). DSBs induced by SSNs are rejoined by the NHEJ pathway, which can occasionally introduce mutations at the DSB sites. NHEJ-induced mutations are largely short deletions, but large deletions, insertions and nucleotide substitutions also occur infrequently (Chen and Gao, 2013; Gaj et al., 2013; Voytas, 2013; Kim and Kim, 2014). Therefore, it is difficult to introduce mutations designed to modify the properties of a target gene by SSNs-induced DSBs.
Because GT frequency in higher eukaryotes can be enhanced by DSBs created by SSNs (Puchta and Fauser, 2013), a negative selectable marker may not be absolutely necessary for the generation of GT plants. However, a positive selectable marker is required for selection of transformed cells from a large proportion of non-transformed cells. Therefore, a precise system of marker excision from GT loci using piggyBac transposon will become an indispensable technology for targeted gene modification leaving no residual ectopic sequences in the plant genome.
Experimental procedures
Vector construction
The GT vector was constructed as follows. The binary vector pKOD4 was constructed using the vector pKO3 (Osakabe et al., 2014) with two cytosine deaminase (codA) gene expression cassettes. A 0.6-kb diphtheria toxin (DT-A) (Terada et al., 2002) fragment was replaced by codA genes in the pKO3 vector, yielding pKOD4 with two DT-A gene expression cassettes (maize polyubiquitin 1 promoter + DT-A + rice heat shock protein (hsp) 16.9a terminator and rice elongation factor-1α promoter + DT-A + rice hsp 16.9b terminator). A 6.4-kb fragment containing the ALS gene carrying two point mutations of W548L and S627I (Endo et al., 2007) was digested with HpaI/EcoRI and integrated into the SnaBI/EcoRI site of pENTR L1/L2 (Life Technologies, http://www.lifetechnologies.com). The piggyBac inverted-repeat transposable element (IVR) harboring meganuclease I–SceI site (Nishizawa-Yokoi et al., 2014) was cloned into a HpaI site (silent mutations at 301-bp upstream of W548L; GCTGAC to GAATTC) introduced by Expression-PCR (Lanar and Kain, 1994) using the primers 5′-tgctggatgagttaacgaaaggtgagg-3′ and 5′-cctcacctttcgttaactcatccagca-3′ in the ALS gene, yielding pE(L1-L2)mALSpb.
A 6.0-kb fragment containing the Oscly1 gene was amplified by PCR from rice genomic DNA (Nipponbare) using the primers 5′-ttggcgcgccttgtcgtcacgcgccagttc-3′ (AscI site in italics) and 5′-ccttaattaatccagggaaatccaccactactact-3′ (PacI site in italics) and integrated into the AscI/PacI site of pENTR L1/L2 (Life Technologies), yielding pE(L1-L2)Oscly1. A 429-bp artificially synthesized fragment containing the 10th exon and 3′-UTR of Oscly1 harboring a single base substitution (CAGCAGCATCATCACGATTCC to CAGCAGCGTCATCACGATTCC in the putative microRNA target site at 88-bp upstream of the stop codon) carrying the piggyBac IVR in the TTAA site of the Oscly1 3′-UTR was replaced by the wild-type Oscly1 gene in pE(L1-L2)Oscly1, yielding pE(L1-L2)Oscly1pb.
A 4.3 kb fragment containing the rice actin terminator, cauliflower mosaic virus 35S promoter, hygromycin phosphotransferase gene (HPT) and rice heat shock protein 17.3 terminator was digested with I–SceI and integrated into pE(L1-L2)mALSpb or pE(L1-L2)Oscly1pb, yielding pE(L1-L2)mALSpHPTb and pE(L1-L2)Oscly1pHPTb, respectively. The fragment of mALSpHPTb or Oscly1pHPTb was re-cloned into the gene targeting binary vector pKOD4 using a Gateway LR clonase II reaction (Life Technologies), yielding pKOD4/mALS and pKOD4/Oscly1pHPTb, respectively. pKOD4/mALS or pKOD4/Oscly1pHPTb vector was transferred into Agrobacterium tumefaciens strain EHA105 (Hood et al., 1993) by electroporation.
Plant materials and Agrobacterium-mediated transformation
Agrobacterium-mediated transformation of rice was performed as described previously (Toki et al., 2006). Rice (Nipponbare) was used for GT transformation following a method described previously (Saika et al., 2011). Calli transformed with Agrobacterium harboring pKOD4/mALS were selected on callus induction (N6D) medium solidified with 0.4% gelrite (Wako Pure Chemical Industries) containing 50 mg/L hygromycin and 25 mg/L meropenem (Wako Pure Chemical Industries, http://www.wako-chem.co.jp/). GT candidate calli were transferred to N6D medium without hygromycin and meropenem and cultured for 1 month. For marker excision, GT candidate calli were transformed with Agrobacterium to introduce an expression vector encoding hyPBase, and were selected on N6D medium with 35 mg/L geneticin (Nacalai tesque, https://www.nacalai.co.jp) and 25 mg/L meropenem. For regeneration, transgenic calli were transferred to regeneration medium with 25 mg/L meropenem, and shoots arising from callus were transferred to Murashige and Skoog (MS) medium (Murashige and Skoog, 1962) without phytohormones. The hyPBase-expressing regenerated plants were subjected to marker excision analysis. The T1 progeny plants were obtained from self-pollinating marker-free T0 plants containing two point mutations, W548L and S627I, in the ALS locus and were subjected to further analysis.
Screening of GT candidate by PCR
After a 4-week selection period, hygromycin-resistant calli were subjected to screening by PCR. Genomic DNA was extracted from small pieces of rice calli using Agencourt chloropure (Beckman Coulter, https://www.beckmancoulter.com) according to the manufacturer's protocol. PCR amplifications were performed with KOD FX or KOD FX neo (TOYOBO) using the primer sets as follows: for 5′ amplification of ALS, ALS GT-F (5′-gacatgacaaccagtcatccgattaggttt-3′) and Tact-R (5′-ctgacgatgagaatatatctgatgctgtga-3′); for 3′ amplification of ALS, Thsp17.3-F (5′-acatacccatccaacaatgttcaatccctt-3′) and ALS GT-R (5′-tctggagatagcatacttgctttgcttggt-3′); for 5′ amplification of Oscly1, Oscly1 GT-F (5′-tcggtcggctaaggtttgctactaaaaaca-3′) and Tact-R (5′-ctgacgatgagaatatatctgatgctgtga-3′); for 3′ amplification of Oscly1, Thsp17.3-F (5′-acatacccatccaacaatgttcaatccctt-3′) and Oscly1 GT-R (5′-cttgcacgacggttctacaggagattagtg-3′).
Sequence analysis
A 4302-bp or 3943-bp fragment was amplified using primer sets ALS GT-F/Tact-R and Thsp17.3-F/ALS GT-R from ALS GT-A or -B, respectively. A 3942-bp or 4257-bp fragment was amplified using primer sets Oscly1 GT-F/Tact-R and Thsp17.3-F/Oscly1 GT-R from Oscly1 GT, respectively. These fragments were cloned into the vector pCR-Blunt II-TOPO using TOPO cloning methods (Life Technologies) and sequenced using universal primers M13-R (5′-caggaaacagctatgac-3′) and M13-F (5′-gtaaaacgacggccagt-3′) to confirm whether the junction sequence corresponded to the anticipated junction fragment. Primers ALS-R1 (5′-acttgggatcataggcagca-3′) and ALS-R2 (5′-ccttagcagtcaggaatagcttg-3′) were used to check the presence of the W548L and S627I mutations on a fragment amplified by Thsp17.3-F/ALS GT-R primers. Primer Oscly1 Seq-5101F (5′-cgaccagaactcgaaccatc-3′) was used to check the presence of the single base substitution in the putative microRNA target site on a fragment amplified by Oscly1 GT-F/Tact-R.
piggyBac excision analysis by PCR
To evaluate the frequency of marker excision via piggyBac transposition from ALS GT locus, we performed CAPS analysis, which is PCR analysis coupled with MfeI digestion (Endo et al., 2007). A 3.8-kb piggyBac-excised fragment was amplified with PrimeSTAR GXL DNA polymerase (TaKaRa, http://www.clontech.com/takara) using primers ALS-F1 (5′-gtacgcaaattatgccgtgga-3′), which anneals to the 359 bp upstream of the insertion site of piggyBac, and ALS GT-R, which is specific for the endogenous ALS locus (3478 bp downstream of the insertion site of piggyBac (Figure 1a). If piggyBac transposon is eliminated from the targeted ALS locus by hyPBase expression, PCR analysis and subsequent MfeI digestion generates fragments of 2.9-, 0.7- and 0.2-kb (Figure 1a,b). In addition to these fragments, 3.8-kb fragments that could not be digested by MfeI were detected in T0 regenerated plants owing to its heterozygosity. However, if piggyBac transposon remains at the targeted ALS locus in hyPBase-expressing regenerated plants, the expected amplified fragments including the selectable marker gene sequence should be 8.7-kb, which cannot be amplified with the PCR conditions used here (Figure 1b). To assess whether the selectable marker was excised precisely from the Oscly1 GT locus via piggyBac transposition, we performed PCR analysis using primer sets Oscly1 GT-F/Tact-R and Oscly1 3′ UTR-F (5′-ggatgctattcttttgctctaccttttt -3′), which anneals 380-bp upstream of the insertion site of piggyBac, and Oscly1 3′ UTR-R (5′-ttactttagtaccaacatctagaaggacga-3′), which anneals 223-bp downstream of the insertion site of piggyBac. If the piggyBac transposon is eliminated from the targeted Oscly1 locus by hyPBase expression, a 0.6-kb fragment is amplified using the primer sets Oscly1 3′ UTR-F/-R; however, a 5.5-kb and 3.9-kb fragment including the selectable marker is not amplified using the primer sets Oscly1 3′ UTR-F/-R and Oscly1 GT-F/Tact-R, respectively. To analyze the re-integration frequency of piggyBac, PCR analysis was performed with PrimeSTAR GXL DNA polymerase (TaKaRa) using primers HPT-F (5′-caaagatcgttatgtttatcggcactttg-3′) and HPT-R (5′-ctcgagctatttctttgccctc-3′) (Figures 1b and S3).
Southern blot analysis
Genomic DNA was extracted from leaves of seedlings using the Nucleon Phytopure extraction kit (GE Healthcare, http://www3.gehealthcare.com/) according to the manufacturer's protocol. Two μg genomic DNA was digested with MfeI and fractionated in a 1.0% agarose gel. Southern blot analysis was performed according to the digoxigenin (DIG) Application Manual (Roche Diagnostics, http://www.roche.com/). Specific DNA probes for the ALS and Oscly1 locus were synthesized with a PCR DIG probe synthesis kit (Roche Diagnostics) according to the manufacturer's protocol, using the primers ALS probe-1 (5′-ttctttttcaatactttcctcgcttgctct-3′ and 5′-attcagccacttatcttgacacaaccattt-3′); and ALS probe-3 (5′-caaagatcgttatgtttatcggcactttg-3′ and 5′-ctcgagctatttctttgccctc-3′); Oscly1 probe-1 (5′-ggttccattccctgacccggcccacct-3′ and 5′-cagtgaatgatgcaacatgagaccgaaca-3′); Oscly1 probe-3 (5′-cagtcatctggacttgttggaattg-3′ and 5′-catcggatgagaccacattaactt-3′); probe-2 (5′-tgtgacagcccagtcatcat-3′ and 5′-cgttggatcgacatcatcag-3′); probe-4 (5′-atcagatgcctgaggatggac-3′ and 5′-acctgctgttcttcaggacctc-3′).
RNA extraction and reverse transcriptase-polymerase chain reaction analysis
Total RNA was extracted from rice calli using an RNeasy Plant Mini Kit (Qiagen, http://www.qiagen.com/). First-strand cDNA was synthesized using ReverTra Ace (Toyobo, http://www.toyobo-global.com/) with oligo(dT20) primer. The cDNA encoding ALS was amplified from the first-strand cDNA by PCR using specific primers, 5′-gtacgcaaattatgccgtgag-3′ and 5′-acttgggatcataggcagca-3.’ To distinguish between cDNA from wild-type ALS gene and modified ALS gene, the first-strand cDNA was digested with MfeI.
BS-susceptibility test
Four-week-old rice calli derived from ALS GT-A_hy 10 T1 were transferred to callus induction medium containing 0.75 μm BS and cultured in a growth chamber under normal conditions. Three-week-old ALS GT-A_hy 10 T1 plants were transferred to ½MS medium (Murashige and Skoog, 1962) containing 1.5 μm BS and grown in a growth chamber at 29°C under continuous light. Photographs were taken 3 weeks after BS treatment in both cases.
Acknowledgments
We acknowledge Dr. B. Hohn for critical reading of the manuscript. We are grateful to Dr. R. Terada (Meijo University) and Dr. S. Iida (Shizuoka Pref. University) for providing the DT-A gene, Dr K. Uchino and Dr H. Sezutsu (National Institute of Agrobiological Sciences) for providing hyPBase, Dr. H. Rothnie for English editing, and Dr. K. Osakabe for insightful discussions and comments. We thank Ms. K. Amagai, Ms. A. Nagashii and Ms. F. Suzuki for general experimental technical support. This research was supported by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Genomics for Agricultural Innovation PGE1001) and KAKENHI (23658012 and 23310142).
SUPPORTING INFORMATION
Additional Supporting Information may be found in the online version of this article.
Figure S1. Experimental strategy for precise genome modification using the piggyBac transposon.
Figure S2. Molecular analysis of ALS GT-B1 and ALS GT-B1_hy T0 plants.
Figure S3. Strategy for the introduction of point mutations into Oscly1 locus via GT and subsequent marker excision from the GT locus using piggyBac transposon.
Figure S4. Herbicide bispyribac (BS)-tolerant phenotype of T1 calli.
Table S1. PCR analysis of piggyBac excision and re-integration events in ALS GT-B1_hy regenerated plants by hyPBase expression
Table S2. Summary of GT experiments targeting the Oscly1 locus.
Table S3. PCR analysis of piggyBac excision and re-integration events in cly1 GT-1_hy plants regenerated by hyPBase expression
Table S4. PCR analysis of piggyBac excision and re-integration events in cly1 GT-2_hy regenerated plants by hyPBase expression
References
- Cary LC, Goebel M, Corsaro BG, Wang HG, Rosen E, Fraser MJ. Transposon mutagenesis of baculoviruses: analysis of Trichoplusia ni transposon IFP2 insertions within the FP-locus of nuclear polyhedrosis viruses. Virology. 1989;172:156–169. doi: 10.1016/0042-6822(89)90117-7. [DOI] [PubMed] [Google Scholar]
- Chen X. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science. 2004;303:2022–2025. doi: 10.1126/science.1088060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen K, Gao C. TALENs: customizable molecular DNA scissors for genome engineering of plants. J. Genet. Genomics. 2013;40:271–279. doi: 10.1016/j.jgg.2013.03.009. [DOI] [PubMed] [Google Scholar]
- Dang TT, Shimatani Z, Kawano Y, Terada R, Shimamoto K. Gene editing a constitutively active OsRac1 by homologous recombination-based gene targeting induces immune responses in rice. Plant Cell Physiol. 2013;54:2058–2070. doi: 10.1093/pcp/pct147. [DOI] [PubMed] [Google Scholar]
- Endo M, Osakabe K, Ono K, Handa H, Shimizu T, Toki S. Molecular breeding of a novel herbicide-tolerant rice by gene targeting. Plant J. 2007;52:157–166. doi: 10.1111/j.1365-313X.2007.03230.x. [DOI] [PubMed] [Google Scholar]
- Gaj T, Gersbach CA, Barbas CF. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013;31:397–405. doi: 10.1016/j.tibtech.2013.04.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hood E, Gelvin S, Melchers L, Hoekema A. New Agrobacterium helper plasmids for gene-transfer to plants. Transgenic Res. 1993;2:208–218. [Google Scholar]
- Jofuku KD, den Boer BG, Van Montagu M, Okamuro JK. Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell. 1994;6:1211–1225. doi: 10.1105/tpc.6.9.1211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johzuka-Hisatomi Y, Terada R, Iida S. Efficient transfer of base changes from a vector to the rice genome by homologous recombination: involvement of heteroduplex formation and mismatch correction. Nucleic Acids Res. 2008;36:4727–4735. doi: 10.1093/nar/gkn451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim H, Kim JS. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 2014;15:321–334. doi: 10.1038/nrg3686. [DOI] [PubMed] [Google Scholar]
- Lanar DE, Kain KC. Expression-PCR (E-PCR): overview and applications. PCR Methods Appl. 1994;4:S92–S96. doi: 10.1101/gr.4.2.s92. [DOI] [PubMed] [Google Scholar]
- Lieberman-Lazarovich M, Melamed-Bessudo C, de Pater S, Levy A. Epigenetic alterations at genomic loci modified by gene targeting in Arabidopsis thaliana. PLoS One. 2013;8:e85383. doi: 10.1371/journal.pone.0085383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meier ID, Bernreuther C, Tilling T, Neidhardt J, Wong YW, Schulze C, Streichert T, Schachner M. Short DNA sequences inserted for gene targeting can accidentally interfere with off-target gene expression. FASEB J. 2010;24:1714–1724. doi: 10.1096/fj.09-140749. [DOI] [PubMed] [Google Scholar]
- Morioka Y, Fujihara Y, Okabe M. Generation of precise point mutation mice by footprintless genome modification. Genesis. 2014;52:68–77. doi: 10.1002/dvg.22727. [DOI] [PubMed] [Google Scholar]
- Moritoh S, Eun CH, Ono A, Asao H, Okano Y, Yamaguchi K, Shimatani Z, Koizumi A, Terada R. Targeted disruption of an orthologue of DOMAINS REARRANGED METHYLASE 2, OsDRM2, impairs the growth of rice plants by abnormal DNA methylation. Plant J. 2012;71:85–98. doi: 10.1111/j.1365-313X.2012.04974.x. [DOI] [PubMed] [Google Scholar]
- Murashige T, Skoog F. A revised medium for the rapid growth and bioassay with tobacco tissue cultures. Physiol. Plant. 1962;15:473–497. [Google Scholar]
- Nair SK, Wang N, Turuspekov Y, et al. Cleistogamous flowering in barley arises from the suppression of microRNA-guided HvAP2 mRNA cleavage. Proc. Natl Acad. Sci. USA. 2010;107:490–495. doi: 10.1073/pnas.0909097107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishizawa-Yokoi A, Endo M, Osakabe K, Saika H, Toki S. Precise marker excision system using an animal-derived piggyBac transposon in plants. Plant J. 2014;77:454–463. doi: 10.1111/tpj.12367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ono A, Yamaguchi K, Fukada-Tanaka S, Terada R, Mitsui T, Iida S. A null mutation of ROS1a for DNA demethylation in rice is not transmittable to progeny. Plant J. 2012;71:564–574. doi: 10.1111/j.1365-313X.2012.05009.x. [DOI] [PubMed] [Google Scholar]
- Osakabe K, Nishizawa-Yokoi A, Ohtsuki N, Osakabe Y, Toki S. A mutated cytosine deaminase gene, codA (D314A), as an efficient negative selection marker for gene targeting in rice. Plant Cell Physiol. 2014;55:658–665. doi: 10.1093/pcp/pct183. [DOI] [PubMed] [Google Scholar]
- Puchta H, Fauser F. Gene targeting in plants: 25 years later. Int. J. Dev. Biol. 2013;57:629–637. doi: 10.1387/ijdb.130194hp. [DOI] [PubMed] [Google Scholar]
- Saika H, Toki S. Mature seed-derived callus of the model indica rice variety Kasalath is highly competent in Agrobacterium-mediated transformation. Plant Cell Rep. 2010;29:1351–1364. doi: 10.1007/s00299-010-0921-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saika H, Oikawa A, Matsuda F, Onodera H, Saito K, Toki S. Application of gene targeting to designed mutation breeding of high-tryptophan rice. Plant Physiol. 2011;156:1269–1277. doi: 10.1104/pp.111.175778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun N, Zhao H. Seamless correction of the sickle cell disease mutation of the HBB gene in human induced pluripotent stem cells using TALENs. Biotechnol. Bioeng. 2014;111:1048–1053. doi: 10.1002/bit.25018. [DOI] [PubMed] [Google Scholar]
- Terada R, Urawa H, Inagaki Y, Tsugane K, Iida S. Efficient gene targeting by homologous recombination in rice. Nat. Biotechnol. 2002;20:1030–1034. doi: 10.1038/nbt737. [DOI] [PubMed] [Google Scholar]
- Terada R, Johzuka-Hisatomi Y, Saitoh M, Asao H, Iida S. Gene targeting by homologous recombination as a biotechnological tool for rice functional genomics. Plant Physiol. 2007;144:846–856. doi: 10.1104/pp.107.095992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terada R, Nagahara M, Furukawa K, Shimamoto M, Yamaguchi K, Iida S. Cre-loxP mediated marker elimination and gene reactivation at the waxy locus created in rice genome based on strong positive–negative selection. Plant Biotechnol. 2010;27:29–37. [Google Scholar]
- Toki S, Hara N, Ono K, Onodera H, Tagiri A, Oka S, Tanaka H. Early infection of scutellum tissue with Agrobacterium allows high-speed transformation of rice. Plant J. 2006;47:969–976. doi: 10.1111/j.1365-313X.2006.02836.x. [DOI] [PubMed] [Google Scholar]
- Voytas DF. Plant genome engineering with sequence-specific nucleases. Annu. Rev. Plant Biol. 2013;64:327–350. doi: 10.1146/annurev-arplant-042811-105552. [DOI] [PubMed] [Google Scholar]
- Wang W, Lin C, Lu D, Ning Z, Cox T, Melvin D, Wang X, Bradley A, Liu P. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA. 2008;105:9290–9295. doi: 10.1073/pnas.0801017105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Weyden L, Adams DJ, Bradley A. Tools for targeted manipulation of the mouse genome. Physiol. Genomics. 2002;11:133–164. doi: 10.1152/physiolgenomics.00074.2002. [DOI] [PubMed] [Google Scholar]
- Yamauchi T, Johzuka-Hisatomi Y, Fukada-Tanaka S, Terada R, Nakamura I, Iida S. Homologous recombination-mediated knock-in targeting of the MET1a gene for a maintenance DNA methyltransferase reproducibly reveals dosage-dependent spatiotemporal gene expression in rice. Plant J. 2009;60:386–396. doi: 10.1111/j.1365-313X.2009.03947.x. [DOI] [PubMed] [Google Scholar]
- Yusa K, Rashid ST, Strick-Marchand H, et al. Targeted gene correction of α1-antitrypsin deficiency in induced pluripotent stem cells. Nature. 2011a;478:391–394. doi: 10.1038/nature10424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yusa K, Zhou L, Li M, Bradley A, Craig N. A hyperactive piggyBac transposase for mammalian applications. Proc. Natl Acad. Sci. USA. 2011b;108:1531–1536. doi: 10.1073/pnas.1008322108. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1. Experimental strategy for precise genome modification using the piggyBac transposon.
Figure S2. Molecular analysis of ALS GT-B1 and ALS GT-B1_hy T0 plants.
Figure S3. Strategy for the introduction of point mutations into Oscly1 locus via GT and subsequent marker excision from the GT locus using piggyBac transposon.
Figure S4. Herbicide bispyribac (BS)-tolerant phenotype of T1 calli.
Table S1. PCR analysis of piggyBac excision and re-integration events in ALS GT-B1_hy regenerated plants by hyPBase expression
Table S2. Summary of GT experiments targeting the Oscly1 locus.
Table S3. PCR analysis of piggyBac excision and re-integration events in cly1 GT-1_hy plants regenerated by hyPBase expression
Table S4. PCR analysis of piggyBac excision and re-integration events in cly1 GT-2_hy regenerated plants by hyPBase expression


