Abstract
The HIV-1 envelope glycoprotein (Env) trimer is responsible for receptor recognition and viral fusion with CD4+ T cells and is the sole target for neutralizing antibodies. Thus, understanding its molecular architecture is of significant interest. However, the Env trimer has proved to be a challenging target for 3D structure determination. Recent EM and X-ray structures have at last enabled us to appreciate the structural complexity and unique features of the Env trimer, and how it is recognized by an ever-expanding arsenal of potent broadly neutralizing antibodies. Here we describe our current knowledge of the Env trimer structure in the context of the exciting recent developments in identification and characterization of HIV broadly neutralizing antibodies.
Keywords: HIV-1, Envelope glycoprotein trimer, Structure, Broadly neutralizing antibodies
Introduction: Bringing the trimeric HIV-1 Env structure to fruition
The trimeric HIV-1 envelope glycoprotein (Env) is a meta-stable type I membrane fusion machine that is responsible for host cell recognition and entry of the virus into the cytoplasm. Env is expressed as a gp160 precursor that is proteolytically cleaved by furin into gp120 and gp41 heterodimers. Three such heterodimers assemble into the final trimeric Env spike. The gp120 subunit has a highly variable surface including five variable loops (V1-V5). By contrast, the gp41 subunits are more conserved in sequence as they house the fusion machinery, which is complex and has many moving parts that undergo enormous conformational rearrangements during the fusion process. The gp41 membrane proximal external region (MPER) connects the gp41 ectodomain to the transmembrane domain (TMD) and cytoplasmic domain (CTD). Perhaps the greatest challenge for structure determination (as well as immunological characterization) is that the Env trimer readily dissociates into gp120 and gp41 subunits, making Env a particularly difficult molecule to study using conventional biophysical methods.
Since the original pioneering structure of monomeric HIV-1 gp120 was determined more than 15 years ago (53), a plethora of gp120 structures have been solved in various forms. Structures of gp120 and its outer domain have been determined with soluble CD4 (sCD4) and co-receptor mimics (16, 38, 53, 57), and with different antibodies that bind the CD4bs or the gp120 outer domain (6, 9, 10, 24, 25, 29, 41, 45, 52, 56–58). These antibodies, as well as sCD4, have been essential for obtaining structural information, as they act as stabilizing agents and crystallization chaperones, although recently some unliganded g120 structures have been determined (26). All structures of gp120 exhibit a similar core fold, consisting of an inner and an outer domain (OD) connected by a bridging sheet. For successful gp120 crystallization and x-ray structure determination, the functionally important hypervariable loops V1, V2 and V3 at the trimer apex had to be deleted or severely truncated (27).
Despite the challenges presented by Env, substantial progress has been made recently in obtaining a three-dimensional structure of the HIV Env trimer as well as elucidating Env-antibody and Env-receptor interactions. With a more complete understanding of the Env trimer, a wide variety of previous observations can now be interpreted or put into the appropriate context. The Env trimer structure has also provided a basis for rational vaccine design efforts aimed at eliciting antibodies against Env (49). This review is intended to give an overview of the recent breakthroughs that led to elucidation of these soluble Env trimer structures (19, 31, 40) and enabled identification of the defining features and characteristics of the pre-fusion gp120 and gp41 subunits, the variable loops, the glycans, and the antigenic surface of this viral fusion machine.
Hitting a moving target: Strategies to study Env
Early electron tomography efforts to study the structure of the Env trimer on the viral surface (55, 59, 60) were limited in the resolution that they achieved but provided a rough outline of the molecular shape of the trimer and allowed docking of gp120 crystal structures to obtain molecular models. More recent tomograms (12, 30) at 20–30 Å resolution yielded further details through hybrid or integrative approaches that fitted the crystal structures of gp120 and/or CD4 into the low resolution EM reconstructions and enabled other portions of the trimer to be modeled for the gp120 region (30), but not for gp41. However, only limited information regarding the variable loops in gp120 could be gleaned from these low resolution models.
Many different constructs of soluble, engineered versions of Env have been pursued over the last two decades for structural studies by a large number of groups worldwide, but all but one met with failure due to a number of compounding factors. The Env trimer is not stable and readily dissociates into component subunits. One common strategy was to prevent gp120-gp41 dissociation by removal of the cleavage site on the gp160 precursor and adding trimerizing motifs, such as leucine zippers and foldons, to attempt to stabilize HIV trimers (37, 54). However, while these trimers can be visualized as a single band by gel electrophoresis, we now know that lack of cleavage between gp120 and gp41 and the presence of trimerizing motifs exerted a negative influence on the quaternary structure of the trimer (42, 48) so they do not adopt a stable, native-like fold. The heterogeneous nature of these trimers makes them nearly impossible to characterize structurally at even modest resolution, let alone atomic resolution.
A very early approach for engineering a soluble stable version of an Env trimer was through incorporation of a disulfide bond (SOS) between gp120 and gp41, together with a mutation of a residue in the N-terminal heptad repeat region (HR1) of gp41 from Isoleucine to Proline (IP) to keep gp120 and gp41 together and enhance trimerization (together known as SOSIP) (2, 44). A more susceptible cleavage site (Arg6 or R6) was also engineered between gp120 and gp41 to facilitate cleavage by furin to produce a soluble, cleaved trimer. While encouraging, the first inceptions of this construct (2, 44) that incorporated the Env sequence from an infected patient known as JR-FL (designed in the laboratory of John Moore), did not produce material that led to Env trimer crystals. More recently, screening of different genotypes with SOSIP mutations (43), truncation of the hydrophobic membrane proximal external region (MPER) at the C-terminus (22, 23), and optimizing expression and purification protocols have resulted in a high quality Env trimer that is eminently suitable for high resolution structural studies. The current version of the trimer is on a Clade A BG505 background and was selected from a panel of over 50 constructs with sequences from different strains and clades. Structural evaluation by negative stain EM showed that the BG505.664 protein (truncated at residue 664) formed compact homogeneous trimers with a ~100% yield after purification (43). Importantly, this trimer retained the antigenic and structural properties of native Env (22, 43), which have eluded other trimeric Env constructs. Retrospectively, we know now from EM studies that these earlier versions did not produce a high percentage of properly folded trimers (42, 48).
The present soluble SOSIP trimers do not have the MPER, transmembrane and cytoplasmic domains of full-length Env (Fig. 1A). Nevertheless, structural comparisons of these soluble Env trimers with tomographic reconstructions of the Env trimer on the virus (30), and more recently, a detergent solubilized full-length Env studied using single particle EM (3), revealed remarkable similarity at the resolution of these studies (~15– 35 Å). Probing the structures further using newly discovered broadly neutralizing antibodies (bnAbs) that recognize only cleaved, native trimers (3, 8) and that interact with both gp120 and gp41 subunits illustrates that these SOSIP trimers recapitulate the highly complex and extended epitopes found on native Env. These studies also revealed the structural similarity of Env across diverse Env sequences corresponding to three different subtypes.
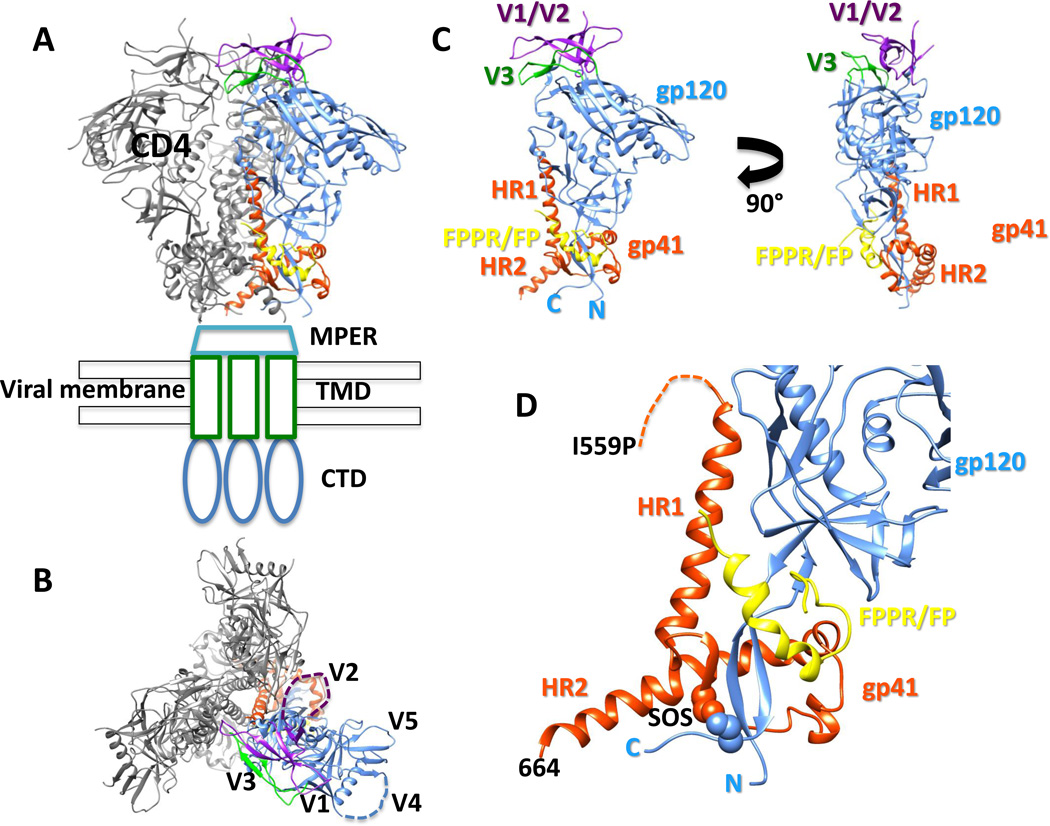
Fig. 1.
Structure of the soluble BG505 SOSIP.664 Env trimer. (A) Side and (B) top views of Env trimer (PDB 4TVP). The receptor CD4 binding site is labeled CD4. Structures of the membrane proximal external region (MPER), transmembrane domain (TMD) and cytoplasmic domain (CTD) have not yet been determined. The variable loops (V1-V5) are located at the apex of the trimer and project outward. V2 and V4 (dashed lines) are disordered in the current structures. (C) A single gp120/gp41 protomer is displayed. The gp41 components heptad repeat 1 and 2 (HR1, HR2) are positioned at the base of the trimer. The fusion peptide proximal region (FPPR) and fusion peptide (FP) are located at the gp120 interface. The N and C termini of gp120 form important interactions with gp41. (D) Close up of gp41. The stabilizing disulfide bond (SOS) introduced at residues 501 (gp120) and 605 (gp41) is shown in spheres. The isoleucine to proline (I559P) is disordered in the current structures and its approximate location is shown with a dashed line. The C terminus of gp41 in the BG505 SOSIP.664 trimer is labeled as 664.
SOSIP Env trimer structure
The SOSIP trimer structure (Fig. 1A-B) (1, 19, 31) exhibits an overall architecture similar to influenza hemagglutinin (HA), the prototypic type I membrane fusion protein. The conserved fusion machinery (HA2 for HA; gp41 for Env) is at the base or stem and is located proximal to the membrane, while the more variable head domain (HA1 for HA; gp120 for Env), which contains the receptor-binding site, is distal to the membrane and sits atop the highly conserved fusion machinery. The interaction of the gp120 monomers with gp41 helps constrain the gp41 subunits in a pre-fusion conformation, and prevent progression into the post-fusion, extended 6-helix bundle conformation (PDB 1ENV) (50). In the gp41 structure, the two main helices, HR1 and HR2 lie parallel and more tangential to the trimer axis, respectively (Fig. 1C). The three HR1 helices form a coiled coil along the trimer axis as for the central helix of influenza hemagglutinin (see fig. 3 of Julien, et al. (19)). The gp120 variable loops V1, V2, and V3 comprise the trimer apex and make stabilizing inter-protomer contacts, with V3 sequestered beneath V1-V2 (Fig. 1A-C) and behind a glycan at position N197 on the adjacent gp120 protomer. V4 and V5 project outward from the surface of gp120 and do not interact with the other variable loops. This architecture was conserved both within the crystal lattice necessary for x-ray crystallography and in the vitreous buffer milieu used for single particle cryoEM studies. A recent structure of the SOSIP trimer at higher resolution (~3.5 Å) enabled a more complete gp41 structure to be built and described fascinating new features of the gp41 domain with implications for membrane fusion (40).
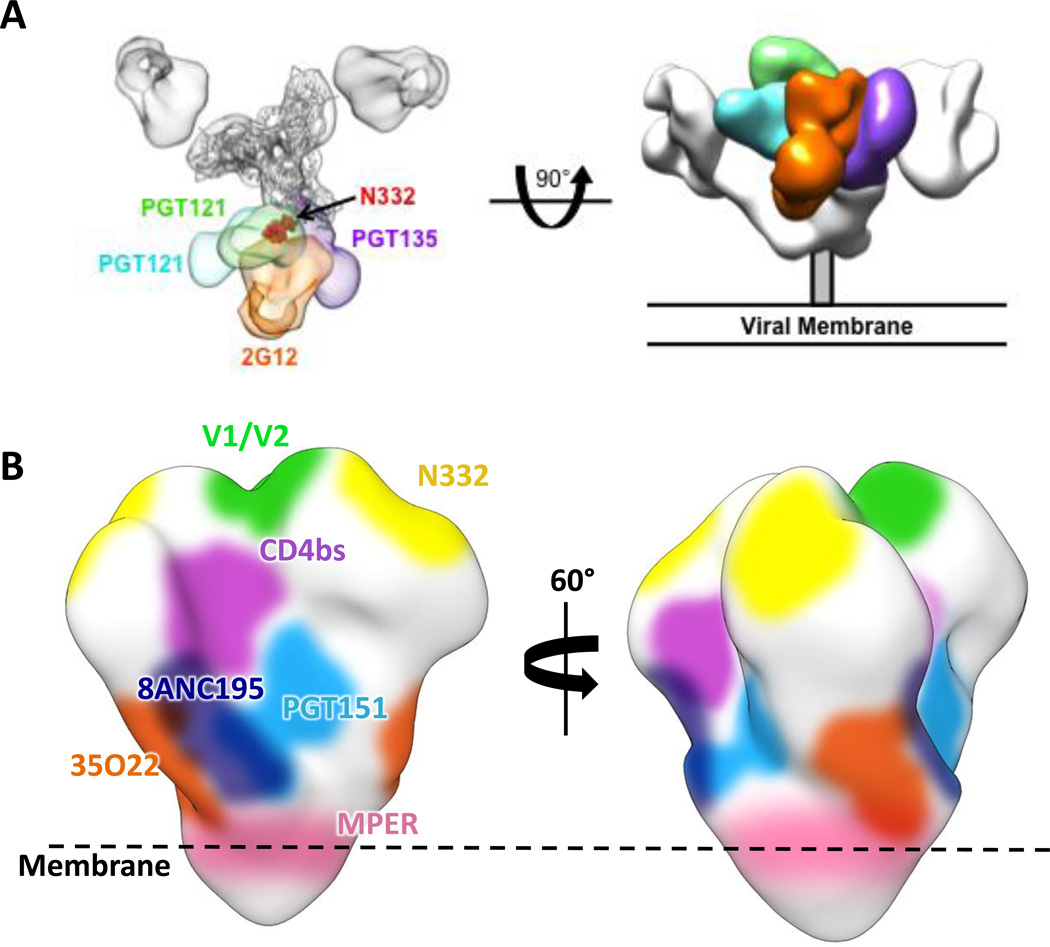
Fig. 3.
Sites of vulnerability on the surface of HIV-1 Env. (A) Top and side view HIV-1 Env trimer bound to different antibodies that all recognize the N332 glycan, but approach from different angles. The antibodies are colored separately and superimposed on the electron microscopy (EM) map EMD-5982. The crystal structure of gp120 (gray cartoon) is docked in the EM map and the N332 glycan is indicated by red spheres. (B) Known epitopes for bnAbs colored on the surface of the EM map EMD-5919.
Env structures using complementary techniques
Crystallographic studies typically require removal of glycans to attain better homogeneity of the protein preparation; therefore, many structures offer an incomplete view of the surface of Env. Most bnAb epitopes, however, contain one or more glycans, therefore presenting an apparent conundrum for structural studies. Creative solutions have been devised to overcome this limitation by first making complexes of gp120 with bnAbs and then digesting away accessible glycans not bound or protected by the antibodies, therefore making the complex more amenable to crystallization (25). Glycans bound by the bnAb thereby remain protected from glycosidases, thus allowing the epitopes of gp120 within the antibody footprint to be visualized in a near native glycosylated state. Most studies, however, require production in cell lines that produce uniform high mannose glycans and lack the ability to produce complex glycans, so there may be subtle differences in the observed antibody-epitope interactions if they involve the outer tips of the glycans that differ between complex and high mannose glycans.
EM, on the other hand, can be conducted on Env trimers produced in human cell lines that are capable of full glycan processing (e.g. human embryonic kidney, HEK 293T). With modern EM technologies, many of these glycans can be observed in reconstructions below 8 Å (31). If the glycans are not constrained, the extended branching sugars average out during single particle processing, but density for the core sugars can often still be visualized. Locking glycans down with a bnAb, as was previously successful in crystallographic studies, aids in visualizing additional portions of glycans. For example, the cryoEM structure of SOSIP.664 in complex with a CD4 binding site antibody PGV04 reveals that the light chain of the antibody interacts with extended regions of a glycan at position N276 (31). Interestingly, removal of this glycan increases access to the CD4 receptor binding site and increases germline antibody binding (18, 33).
The highly glycosylated surface of gp120
Glycans, which coat the entire surface of Env (Fig. 2), were originally thought to be a barrier for recognition by the immune system, but have turned out to be key features on the surface of Env for bnAb recognition. Once referred to as the silent face because of its perceived lack of reactivity with human antibodies (32), the dense high mannose patch on the outer domain of gp120 has recently been described as a supersite of vulnerability due to the diversity of bnAbs that are capable of recognizing this region of the trimer (25). In fact, this region is the most available to recognition by antibodies in terms of steric accessibility and in the variety of possible approach angles. Multiple bnAbs that recognize this site bind to the same glycan (N332), but use different angles of approach and different combinations of the surrounding glycans for interaction (9, 19, 25, 36, 41, 46) [Fig 3A]. Superposition of the structures reveals that the N332 glycan is in a very similar conformation in Env structures with different antibodies, thus presenting a somewhat structurally conserved epitope. The density of glycans within the high mannose patch may therefore constrain glycan motion (25). In addition, the high density of glycans in this patch (4) likely explains the predominance of high mannose glycans, as it interferes with glycan processing enzymes, which would normally trim the high mannose sugars down and add back other sugars to make complex glycans. This clustering thereby leads to the high mannose phenotype on the outer domain. Further constraints on glycan processing and flexibility may be imparted by the quaternary arrangement of the trimer.
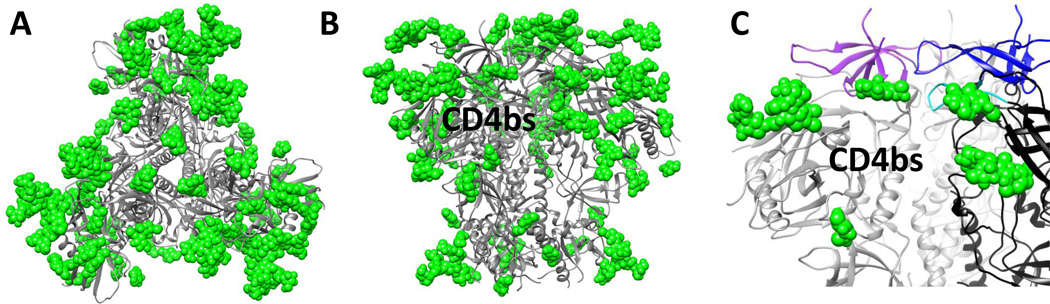
Fig. 2.
The highly glycosylated surface of Env. (A) Top and (B) side views of the Env trimer (gray cartoon) illustrating the extensive glycan shield. Shown in green are the glycans resolved in the current structures of the Env trimer. The full extent of most glycans remains to be determined so the figure here under-represents the extent of glycans. The receptor CD4 binding site is labeled. (C) Close up illustrating the variable loop and glycan constraints restricting access to the CD4 binding site. The V1/V2 regions of two different protomers are colored purple and blue. V3 is colored in cyan. Glycans are displayed as green spheres.
EM studies of the trimer in complex with bnAb PG9 revealed that glycans N156/N173 and N160 are clustered around the threefold axis at the trimer apex (20). In particular, N160 glycans are apposed and form a deep tunnel restricting access to the portions of V1 and V2 that hold the trimer together at the apex [Fig. 3B]. This arrangement results in an epitope that spans two of the three gp120 protomers, precluding more than one PG9 antibody from binding to the trimer at a time due to steric exclusion (20). Crystal structures of the PG9 and PG16 antibodies in complex with V1 and V2, engineered onto a protein scaffold (34, 39), revealed that the epitope at the trimer apex consists of both high mannose and complex glycans.
While structural studies are typically conducted on a single Env sequence, it is important to consider how Env evolves during the course of infection. For these types of studies, researchers have employed next generation sequencing (NGS) to conduct in depth studies of Env evolution in response to antibody pressure (7, 29, 51, 58). One such study highlighted the co-evolution of a particular human antibody lineage and viral Env, and illustrated how the trimer apex mutated in response to antibody pressure (7). Over time, one antibody lineage VRC26, which recognizes the trimer in a similar manner as PG9, loses its dependence on the glycans originally required for binding. While NGS is powerful, it can only provide inferred glycosylation patterns and reveals nothing about actual glycan utilization or specific glycoforms. In order to address this issue, glycan arrays that probe bnAb binding can be utilized, although they too are indirect measures outside of native Env. Therefore, tools and methodology currently under development will be critical to further understand site-specific and strain-dependent glycosylation (13–15).
The gp120 variable loops
Underneath and interspersed between the glycans on the surface of Env are the variable loops (V1-V5) that splay out around the perimeter of the gp120 core (Fig. 1B), thereby masking the more conserved regions of gp120 underneath. These loops can tolerate high mutations rates (including insertions and deletions) and exhibit high levels of glycosylation and flexibility; therefore, they are typically removed from crystallization constructs. V1 and V2 extend away from the trimer apex, with V1 projecting toward the high mannose patch on the gp120 outer domain and V2 situated above the CD4 binding site (CD4bs); both V1 and V2 readily evolve under immune pressure. For example, in longitudinal studies of HIV (7, 29, 51), the length, amino acid composition, and number of glycosylation sites of the variable loops was observed to change. Because each of these loops radiates from the periphery of the trimer core, length changes are accommodated without compromising the core structure or function of Env. These loops are also flexible, further adding to the variable nature of the trimer apex. In the available structures that contain V1 and V2 (19, 31, 34, 39), V1 exhibits different conformations while large portions of V2 are not visualized due to disorder.
The length of the V3 loop cannot vary significantly without disrupting important trimer-stabilizing contacts, because the tip of the V3 loop reaches over and makes inter-protomer contacts with the base of V2, near position 197, on the adjacent protomer. The sequence at the V3 tip is also very conserved as either GPGQ or GPGR. The remaining amino acid residues in the V3 loop determine co-receptor usage (either CCR5 or CXCR4) and conserved basic residues (lysine and arginine) are proximal to the CD4bs of the adjacent gp120 protomer. The conserved V3 glycans, N295, N301, and N332 (or N334), all reside at the base of the loop and comprise part of the high mannose patch.
The CD4 receptor binding site
All of the conserved regions of gp120, outside of the CD4bs, are at the interface of the subunits or underneath the trimer apex formed by the V1/V2/V3 cap. Although the CD4bs itself is conserved and free of glycans, its location near the interprotomer interface limits accessibility (Fig 2B-C). CD4, and CD4bs-targeting bnAbs, must approach from a particular angle in order to engage the trimer and avoid the nearby V1, V2, and V3 variable loops. The CD4bs is potentially further restricted by surrounding glycans, particularly the one at position N276 (18, 33), which delineates the bottom of the CD binding site and epitopes for CD4 binding site antibodies.
In addition to the constraints within a single gp120, the adjacent protomer also influences access to the CD4bs. Conserved glycans at positions N262, N301, and N448 project toward the neighboring CD4bs. The tip of V3 and helix α-0 also come into close proximity of the CD4bs. In fact, residues proximal to α-0 have been shown to affect CD4-induced conformational changes (21) despite its position on the opposite side of the CD4bs within the gp120 monomer. To date, there is one predominant angle of approach that was identified from a study of VRC01 and VRC01-like bnAbs (56), although other subtly different angles are also observed as exemplified by b12 (47, 57) or CH103 (29). Many non-bnAbs approach the CD4bs in gp120 at a steep vertical angle of approach, but cannot bind to gp120 in natively structured Env trimers. These non-bnAbs can neutralize some tier 1 viruses, where the trimers are more open and the V1 and V2 loops have undergone a conformational rearrangement, but they cannot neutralize more robust, tier 2 viruses, which have much more compact trimers with restricted access to the CD4bs (48).
Neutralizing antibodies targeting the CD4bs manage to navigate all of these obstacles presented by trimeric Env, suggesting that they arise against a native form of Env in natural infection. Some highly evolved CD4bs antibodies have acidic residue insertions in the typically conserved heavy framework region 3 portion of the antibody (24, 28, 29). Modeling of these antibodies in the CD4bs of the cryoEM trimer structure (31) illustrates that these acidic residues interact with highly conserved basic residues in V3. Thus, as antibodies evolve and attempt to negotiate the variability surrounding the CD4bs, they are able to acquire additional quaternary contacts in the invariant V3 co-receptor loop that is only proximal to the CD4bs in intact trimers.
The gp41 fusion machinery
The structure of gp41 in its post-fusion six-helix bundle was solved more than 17 years ago (50), and revealed that heptad repeat 1 (HR1) lies at the core of the bundle with HR2 at the periphery, both in extended helical conformations. In the structure of the prefusion trimer, HR1 forms a three-helix bundle at the center of the trimer perpendicular to the membrane, while HR2 runs diagonal to the membrane at the periphery (Fig. 1A) (19, 31). Both helices in the pre-fusion trimer are significantly shorter that the post-fusion counterpart, and HR2 forms a long curving helix associated with some smaller helical segments. The immunodominant disulfide loop region is sequestered between HR1 and HR2 near the membrane proximal region of the trimer. The conserved N- and C-terminal peptides of gp120 (C1 and C5) also meander up and down through the hydrophobic core of gp41 (Fig. 1C-D) (40). Finally, the fusion peptide is buried in a hydrophobic pocket at the gp41-gp120 interface, where it is positioned to respond to conformational changes that rearrange this interface (31, 40).
Several new antibodies that bind gp41 or an interface between gp41 and gp120 are beginning to reveal more details of the pre-fusion gp41 (3, 17, 45) [Fig. 3]. Unlike MPER antibodies, which recognize short peptides or helices that may only exist transiently, these antibodies recognize complex, extended epitopes, including glycans, in the pre-fusion state. Thus, despite close proximity to the viral membrane, the base of the trimer appears to be still accessible.
Glycan data on gp41 are sparse, but the recent EM structure of bnAb PGT151 in complex with full-length Env suggests that gp41 glycans form important components of bnAb epitopes (3, 8). Glycan array studies also indicated that the carbohydrates at positions N637 and N611 are complex (either tri- or tetra-antennary) and adjacent in space, consistent with the predicted positions from the trimer structures. Recent studies of Env in complex with the bnAbs 8ANC195 and 35O22 reveal a glycan patch that extends from gp120 into gp41 (17, 45). In particular, gp120 glycans N88, N234, and N241 are adjacent to N625 and potentially N616 (or N618 in some strains).
Concluding Remarks
The sequence of Env, and by extension its structural and antigenic properties, are shaped by evolutionary pressure exerted by antibodies. Much of what is known about Env has in fact been determined using antibodies to probe its surface, as described throughout this review. The recent structures of soluble HIV-1 Env trimers, solved in complex with antibodies, represent the most complete picture of this elusive and moving target. More than twenty years of Env research can now be interpreted in the context of the Env trimer assembly. At the same time, these structures provide a basis for driving new hypotheses and moving towards an HIV vaccine.
These structural data on Env are also of high utility for designing immunogens with the goal of eliciting bnAbs through a process known as reverse vaccinology (49). One hurdle to reverse vaccinology, as revealed by the trimer structures, is the complexity of the bnAb epitopes on the surface of the trimer, many of which incorporate glycans. A bnAb therefore faces extraordinary constraints so as to be able to neutralize a wide range of viruses. These constraints likely explain the high level of somatic hypermutation that many bnAbs undergo, oftentimes taking years to develop after being exposed to a diverse, evolving population of HIV-1 that occurs in natural infection. While many challenges lay ahead, researchers now have a platform for accelerating HIV-1 vaccine research and hurdling the previously insurmountable roadblocks.
While rational vaccine design continues to move forward, it is important to remember that the trimer structures, as determined to date, represent the most populated state of what we know to be a dynamic molecule. Techniques that can measure the dynamics of trimers in solution or on the surface of virions (5, 11, 35) are essential for extending our biological knowledge of the trimer and of the consequences of apparently reversible transitions that can take place in native or ground-state trimers prior to CD4 binding. Additionally, structural and functional data on other trimers are required, particularly on the different subtypes (clades) of HIV-1 to ascertain what, if any, structural differences and variations occur that may be of value for vaccine design and generation of a broadly neutralizing response. Further structures of intermediate states will also be required to delineate the various steps and conformational rearrangements associated with the viral fusion process and to identity possible modes of intervention. Not surprisingly, these exciting new Env structures have provided fascinating new insights and raised previously unforeseen questions, thereby opening up further avenues for research and providing tantalizing and often unexpected answers to the major outstanding questions in the HIV-1 field.
Outstanding questions.
How many biologically relevant conformations and intermediates of the Env trimer exist and what are the trajectories of their interconversion?
What influence does the MPER, TM, and CTD have on Env structure and dynamics?
What are the molecular determinants that enhance stable trimer formation?
If the human immune system is capable of targeting nearly the entire surface of Env, why is it so difficult to produce a vaccine?
Highlights.
We examine the recent structures of the HIV-1 envelope glycoprotein trimer.
Broadly neutralizing antibodies play a key role in HIV-1 Env structural studies.
The antigenic surface of HIV-1 Env is a complex landscape of glycan and peptide.
Acknowledgements
Figures 3A and 3B were created by Charles D. Murin and by Jeong Hyun Lee, respectively, of The Scripps Research Institute.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Bartesaghi A, et al. Prefusion structure of trimeric HIV-1 envelope glycoprotein determined by cryo-electron microscopy. Nat. Struct. Mol. Biol. 2013;20:1352–1357. doi: 10.1038/nsmb.2711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Binley JM, et al. A recombinant human immunodeficiency virus type 1 envelope glycoprotein complex stabilized by an intermolecular disulfide bond between the gp120 and gp41 subunits is an antigenic mimic of the trimeric virion-associated structure. J. Virol. 2000;74:627–643. doi: 10.1128/jvi.74.2.627-643.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Blattner C, et al. Structural delineation of a quaternary, cleavage-dependent epitope at the gp41-gp120 interface on intact HIV-1 Env trimers. Immunity. 2014;40:669–680. doi: 10.1016/j.immuni.2014.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bonomelli C, et al. The glycan shield of HIV is predominantly oligomannose independently of production system or viral clade. PLoS One. 2011;6:e23521. doi: 10.1371/journal.pone.0023521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Davenport TM, et al. Isolate-specific differences in the conformational dynamics and antigenicity of HIV-1 gp120. J. Virol. 2013;87:10855–10873. doi: 10.1128/JVI.01535-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Diskin R, et al. Increasing the potency and breadth of an HIV antibody by using structure-based rational design. Science. 2011;334:1289–1293. doi: 10.1126/science.1213782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Doria-Rose NA, et al. Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies. Nature. 2014;509:55–62. doi: 10.1038/nature13036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Falkowska E, et al. Broadly neutralizing HIV antibodies define a glycan-dependent epitope on the prefusion conformation of gp41 on cleaved envelope trimers. Immunity. 2014;40:657–668. doi: 10.1016/j.immuni.2014.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Garces F, et al. Structural Evolution of Glycan Recognition by a Family of Potent HIV Antibodies. Cell. 2014;159:69–79. doi: 10.1016/j.cell.2014.09.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Georgiev IS, et al. Delineating antibody recognition in polyclonal sera from patterns of HIV-1 isolate neutralization. Science. 2013;340:751–756. doi: 10.1126/science.1233989. [DOI] [PubMed] [Google Scholar]
- 11.Guttman M, et al. CD4-induced activation in a soluble HIV-1 Env trimer. Structure. 2014;22:974–984. doi: 10.1016/j.str.2014.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Harris A, et al. Trimeric HIV-1 glycoprotein gp140 immunogens and native HIV-1 envelope glycoproteins display the same closed and open quaternary molecular architectures. Proc. Natl. Acad. Sci. USA. 2011;108:11440–11445. doi: 10.1073/pnas.1101414108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Harvey DJ, et al. MALDI-MS/MS with traveling wave ion mobility for the structural analysis of N-linked glycans. J. Am. Soc. Mass Spectrom. 2012;23:1955–1966. doi: 10.1007/s13361-012-0425-8. [DOI] [PubMed] [Google Scholar]
- 14.Harvey DJ, et al. Travelling wave ion mobility and negative ion fragmentation for the structural determination of N-linked glycans. Electrophoresis. 2013;34:2368–2378. doi: 10.1002/elps.201200669. [DOI] [PubMed] [Google Scholar]
- 15.Harvey DJ, et al. Ion mobility mass spectrometry for extracting spectra of N-glycans directly from incubation mixtures following glycan release: application to glycans from engineered glycoforms of intact, folded HIV gp120. J. Am. Soc. Mass Spectrom. 2011;22:568–581. doi: 10.1007/s13361-010-0053-0. [DOI] [PubMed] [Google Scholar]
- 16.Huang CC, et al. Structure of a V3-containing HIV-1 gp120 core. Science. 2005;310:1025–1028. doi: 10.1126/science.1118398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huang J, et al. Broad and potent HIV-1 neutralization by a human antibody that binds the gp41-gp120 interface. Nature. 2014 doi: 10.1038/nature13601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jardine J, et al. Rational HIV immunogen design to target specific germline B cell receptors. Science. 2013;340:711–716. doi: 10.1126/science.1234150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Julien JP, et al. Crystal structure of a soluble cleaved HIV-1 envelope trimer. Science. 2013;342:1477–1483. doi: 10.1126/science.1245625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Julien JP, et al. Asymmetric recognition of the HIV-1 trimer by broadly neutralizing antibody PG9. Proc. Natl. Acad. Sci. USA. 2013;110:4351–4356. doi: 10.1073/pnas.1217537110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kassa A, et al. Transitions to and from the CD4-bound conformation are modulated by a single-residue change in the human immunodeficiency virus type 1 gp120 inner domain. J. Virol. 2009;83:8364–8378. doi: 10.1128/JVI.00594-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Khayat R, et al. Structural characterization of cleaved, soluble HIV-1 envelope glycoprotein trimers. J. Virol. 2013;87:9865–9872. doi: 10.1128/JVI.01222-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Klasse PJ, et al. Influences on trimerization and aggregation of soluble, cleaved HIV-1 SOSIP envelope glycoprotein. J. Virol. 2013;87:9873–9885. doi: 10.1128/JVI.01226-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Klein F, et al. Somatic mutations of the immunoglobulin framework are generally required for broad and potent HIV-1 neutralization. Cell. 2013;153:126–138. doi: 10.1016/j.cell.2013.03.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kong L, et al. Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nat. Struct. Mol. Biol. 2013;20:796–803. doi: 10.1038/nsmb.2594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kwon YD, et al. Unliganded HIV-1 gp120 core structures assume the CD4-bound conformation with regulation by quaternary interactions and variable loops. Proc. Natl. Acad. Sci. USA. 2012;109:5663–5668. doi: 10.1073/pnas.1112391109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kwong PD, et al. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature. 1998;393:648–659. doi: 10.1038/31405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li Y, et al. HIV-1 neutralizing antibodies display dual recognition of the primary and coreceptor binding sites and preferential binding to fully cleaved envelope glycoproteins. J. Virol. 2012;86:11231–11241. doi: 10.1128/JVI.01543-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Liao HX, et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature. 2013;496:469–476. doi: 10.1038/nature12053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Liu J, et al. Molecular architecture of native HIV-1 gp120 trimers. Nature. 2008;455:109–113. doi: 10.1038/nature07159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lyumkis D, et al. Cryo-EM structure of a fully glycosylated soluble cleaved HIV-1 envelope trimer. Science. 2013;342:1484–1490. doi: 10.1126/science.1245627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.McCaffrey RA, et al. N-linked glycosylation of the V3 loop and the immunologically silent face of gp120 protects human immunodeficiency virus type 1 SF162 from neutralization by anti-gp120 and anti-gp41 antibodies. J. Virol. 2004;78:3279–3295. doi: 10.1128/JVI.78.7.3279-3295.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.McGuire AT, et al. Engineering HIV envelope protein to activate germline B cell receptors of broadly neutralizing anti-CD4 binding site antibodies. J. Exp. Med. 2013;210:655–663. doi: 10.1084/jem.20122824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.McLellan JS, et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature. 2011;480:336–343. doi: 10.1038/nature10696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Munro JB, et al. Conformational dynamics of single HIV-1 envelope trimers on the surface of native virions. Science. 2014 doi: 10.1126/science.1254426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Murin CD, et al. Structure of 2G12 Fab2 in complex with soluble and fully glycosylated HIV-1 Env by negative-stain single-particle electron microscopy. J. Virol. 2014;88:10177–10188. doi: 10.1128/JVI.01229-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Pancera M, et al. Soluble mimetics of human immunodeficiency virus type 1 viral spikes produced by replacement of the native trimerization domain with a heterologous trimerization motif: characterization and ligand binding analysis. J. Virol. 2005;79:9954–9969. doi: 10.1128/JVI.79.15.9954-9969.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Pancera M, et al. Structure of HIV-1 gp120 with gp41-interactive region reveals layered envelope architecture and basis of conformational mobility. Proc. Natl. Acad. Sci. USA. 2010;107:1166–1171. doi: 10.1073/pnas.0911004107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pancera M, et al. Structural basis for diverse N-glycan recognition by HIV-1-neutralizing V1-V2-directed antibody PG16. Nat. Struct. Mol. Biol. 2013;20:804–813. doi: 10.1038/nsmb.2600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pancera M, et al. Structure and immune recognition of trimeric prefusion HIV-1 Env. Nature. 2014;514:455–461. doi: 10.1038/nature13808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Pejchal R, et al. A potent and broad neutralizing antibody recognizes and penetrates the HIV glycan shield. Science. 2011;334:1097–1103. doi: 10.1126/science.1213256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ringe RP, et al. Cleavage strongly influences whether soluble HIV-1 envelope glycoprotein trimers adopt a native-like conformation. Proc. Natl. Acad. Sci. USA. 2013;110:18256–18261. doi: 10.1073/pnas.1314351110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sanders RW, et al. A next-generation cleaved, soluble HIV-1 Env Trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 2013;9:e1003618. doi: 10.1371/journal.ppat.1003618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sanders RW, et al. Stabilization of the soluble, cleaved, trimeric form of the envelope glycoprotein complex of human immunodeficiency virus type-1. J. Virol. 2002;76:8875–8889. doi: 10.1128/JVI.76.17.8875-8889.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Scharf L, et al. Antibody 8ANC195 reveals a site of broad vulnerability on the HIV-1 envelope spike. Cell Rep. 2014;7:785–795. doi: 10.1016/j.celrep.2014.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sok D, et al. Promiscuous glycan site recognition by antibodies to the high-mannose patch of gp120 broadens neutralization of HIV. Sci. Transl. Med. 2014;6:236ra263. doi: 10.1126/scitranslmed.3008104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tran EE, et al. Structural mechanism of trimeric HIV-1 envelope glycoprotein activation. PLoS Pathog. 2012;8:e1002797. doi: 10.1371/journal.ppat.1002797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Tran K, et al. Vaccine-elicited primate antibodies use a distinct approach to the HIV-1 primary receptor binding site informing vaccine redesign. Proc. Natl. Acad. Sci. USA. 2014;111:E738–E747. doi: 10.1073/pnas.1319512111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Walker LM, Burton DR. Rational antibody-based HIV-1 vaccine design: current approaches and future directions. Curr. Opin. Immunol. 2010;22:358–366. doi: 10.1016/j.coi.2010.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Weissenhorn W, et al. Atomic structure of the ectodomain from HIV-1, gp41. Nature. 1997;387:426–430. doi: 10.1038/387426a0. [DOI] [PubMed] [Google Scholar]
- 51.Wibmer CK, et al. Viral escape from HIV-1 neutralizing antibodies drives increased plasma neutralization breadth through sequential recognition of multiple epitopes and immunotypes. PLoS Pathog. 2013;9:e1003738. doi: 10.1371/journal.ppat.1003738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Wu X, et al. Focused evolution of HIV-1 neutralizing antibodies revealed by structures and deep sequencing. Science. 2011;333:1593–1602. doi: 10.1126/science.1207532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wyatt R, et al. The antigenic structure of the HIV gp120 envelope glycoprotein. Nature. 1998;393:705–711. doi: 10.1038/31514. [DOI] [PubMed] [Google Scholar]
- 54.Yang X, et al. Highly stable trimers formed by human immunodeficiency virus type 1 envelope glycoproteins fused with the trimeric motif of T4 bacteriophage fibritin. J. Virol. 2002;76:4634–4642. doi: 10.1128/JVI.76.9.4634-4642.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Zanetti G, et al. Cryo-electron tomographic structure of an immunodeficiency virus envelope complex in situ. PLoS Pathog. 2006;2:e83. doi: 10.1371/journal.ppat.0020083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Zhou T, et al. Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science. 2010;329:811–817. doi: 10.1126/science.1192819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhou T, et al. Structural definition of a conserved neutralization epitope on HIV-1 gp120. Nature. 2007;445:732–737. doi: 10.1038/nature05580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zhou T, et al. Multidonor analysis reveals structural elements, genetic determinants, and maturation pathway for HIV-1 neutralization by VRC01-class antibodies. Immunity. 2013;39:245–258. doi: 10.1016/j.immuni.2013.04.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Zhu P, et al. Distribution and three-dimensional structure of AIDS virus envelope spikes. Nature. 2006;441:847–852. doi: 10.1038/nature04817. [DOI] [PubMed] [Google Scholar]
- 60.Zhu P, et al. Cryoelectron tomography of HIV-1 envelope spikes: further evidence for tripod-like legs. PLoS Pathog. 2008;4:e1000203. doi: 10.1371/journal.ppat.1000203. [DOI] [PMC free article] [PubMed] [Google Scholar]