Abstract
Purpose
Assessing whole-body radiation injury and absorbed dose is essential for remediation efforts following accidental or deliberate exposure in medical, industrial, military, or terrorist incidents. We hypothesize that variations in specific metabolite concentrations extracted from blood plasma would correlate with whole-body radiation injury and dose.
Methods and Materials
Groups of C57BL/6 mice (n=12 per group) were exposed to 0 Gy, 2 Gy, 4 Gy, 8 Gy, and 10.4 Gy of whole-body γ-radiation. At 24 hours post treatment all animals were euthanized and both plasma and liver biopsies obtained - the latter being used to deconvolve a distinct hepatic radiation injury response within plasma. A semi-quantitative untargeted metabolites/lipid profiling using both GC/MS and LC/MS/MS platforms was performed and identified 354 biochemicals. A second set of C57BL/6 mice (n=6 per group) were used to assess a subset of identified plasma markers beyond 24 hours.
Results
We identified a cohort of 37 biochemical compounds in plasma that yielded the optimal separation of the irradiated sample groups, with the most correlated metabolites associated with pyrimidine (positively correlated) and tryptophan (negatively correlated) metabolism. The latter were predominantly associated with indole compounds, and there was evidence to indicate that these were also correlated between liver and plasma. No evidence of saturation as a function of dose was observed, as has been noted for studies involving metabolite analysis of urine.
Conclusion
Plasma profiling of specific metabolites related to the pyrimidine and tryptophan pathways can be used to differentiate whole-body radiation injury and dose response. As the tryptophan associated indole compounds have their origin in the intestinal microbiome and subsequently the liver, these metabolites in particular represent an attractive marker for radiation injury within blood plasma.
1. Introduction
The ability to accurately discern the received whole-body radiation dose after a specific radiological event is of great relevance to the subsequent triaging of victims for treatment. Ionizing radiation induces a diversity of cellular responses across a range of tissue types and these all contribute towards an overall clinical diagnosis which has been to date traditionally classified as acute, late consequential or late [1]. The most serious of these, acute radiation syndrome (ARS), need to be identified as soon as possible so as to ensure patients can be offered appropriate and immediate medical attention, with treatments specifically designed to ameliorate the worst effects of radiation sickness [2].
The study of metabolites and lipids in biofluids such as urine or blood plasma represents a particularly attractive mechanism whereby the holistic, whole-body response to radiation may be monitored. Whilst the presence of specific gene transcripts (and to a lesser extent proteins) is tightly coupled to their tissue types of origin, metabolites represent the effective endpoints of cellular regulatory processes [3]. As such, the presence and concentration of specific metabolites are likely to be a direct consequence of the whole-body response to any perturbations, such as those from ionizing radiation. Easy to obtain, prepare and subsequently process, the isolation of such metabolites is also in many ways much less complex than the more rigorous laboratory controlled environments needed to extract and process nucleic acids or proteins, offering a potentially effective technological development pathway for an ultimately deployable ‘kit’ in the field.
Several previous studies have attempted to discern biofluid metabolites consistent with an ‘in vivo’ whole-body response to radiation using laboratory rodents, specifically blood and urine. All studies published related to analyses of the former confirm differences in metabolites in response to ionizing radiation, albeit with little concordance. Urine has been a particular focus, as metabolites are expected to accumulate in the bladder and thus can be pooled and collected over set time periods. In one series of studies, urine was collected from male C57BL6 mice 24 hours after being irradiated between 0 and 8 Gy, and was subsequently analyzed for metabolite markers for radiation injury [4,5]. These studies indicated that the the pyrimidine and purine metabolic pathway compounds thymidine, deoxyuridine and deoxyxanthosine were consistent markers for radiation exposure - however these authors noted that the response of these markers saturated beyond 3 Gy. Such saturation was also reported for the urinary derived pyrimidine deoxycytidine [6]. Subsequent studies using Wistar rats using a sham versus 3Gy whole body irradiation indicated a common upregulated pyrimidine response associated with excess thymidine, a likely consequence of increased DNA breakdown and cell turnover after exposure to irradiation [7,8]. This issue of ‘saturation’ is however a cause for some concern from a diagnostic perspective.
Whilst there has been less focus on blood plasma metabolomics, prior studies have indicated that the presence of specific metabolites and proteins in serum/plasma are indicators of radiation exposure, namely citrulline (epithelial radiation-induced small bowel damage) [9], amylase (radiation damage to the parotid gland) [10] and Flt3-ligand (bone marrow radiation damage) [11]. All three are associated with a specific point of origin and whilst they may be broadly used to confirm significant radiation exposure, they do so without any great specificity [12]. Furthermore, in each case there is no data articulating the dynamic range, both in time and in metabolite concentration, with regard to received radiation dose, making it difficult to use these as suitably robust biomarkers.
There have been limited studies to date characterizing serum/plasma metabolomics’ response to ionizing radiation as a function of varying dose and time after irradiation. Using proton nuclear magnetic resonance spectroscopy (1H NMR), Khan et al. [13] studied the serum metabolites of groups of irradiated mice exposed to 3, 5, and 8 Gy respectively, with serum obtained day 1, 3, and 5 post-irradiation. These authors reported results showing increased lactate, amino acids, choline, and lipid signals as well as decreased glucose signals. Although it is difficult to absolutely quantify 1H NMR spectra, these authors did not report any saturation in the serum metabolite signals identified beyond the 3 Gy level.
A subsequent publication by the same group obtained serum and urine from a single cohort of the same mice strain, exposed to 5 Gy [14]. Samples were taken at time points designed to best reflect the phases of radiation sickness (6h, 5 days, 10 days, 15 days, 20, and 25 days post-irradiation), and subjected to 1H NMR. This data confirmed the serum observations made in their earlier publication, but also strongly implicated the presence of a gastrointestinal component in the urine metabonome, with increased trimethyl amine, hippurate, phenylalanine, and other aromatic amino acid signals. In [15], sham and 6.5 Gy irradiated Wistar rats had their blood plasma analyzed using liquid chromatography-mass spectrometry at 24 hours post-exposure. 19 metabolites were identified as associated with a broad range of metabolic pathways, including both pyrimidine and tryptophan - the latter again associated with a gastrointestinal origin.
Perturbations to the gut microbiome - by the use of germ-free or antibiotic treated mice - have been known for some time to profoundly alter the blood metabolite content, most particularly those pathways associated with aromatic amino acids. Indeed, the production of bioactive indole-containing metabolites derived from tryptophan is totally dependent on the gut microbiome [16]. The evidence from the literature to date suggests that there are in fact two convolved signals in both urine and in blood plasma associated with a whole body response to irradiation, one from the host and the second from its symbiont, the gut microbiome. This suggests that the latter undergoes some form of metabolic dysbiosis itself in response to radiation injury, presenting the possibility that its characterization could offer an alternative and promising route to radiation injury biomarkers. Furthermore, the gut microflora impacts crypt loss and survival of mice exposed to whole body irradiation [17,18].
In this study we attempted to discern a clear gut microbiome signature in an untargeted metabolomics screen of C57BL/6 murine liver and blood plasma samples taken 24 hours post-irradiation from groups of 12 animals exposed to sham (0,2,4,8,10.4 Gy) doses corresponding to different severities of ARS as determined by the IAEA and WHO: mild (1-2 Gy), moderate (2-4 Gy), severe (4-6 Gy), very severe (6-8 Gy), lethal (>8 Gy). We also wished to determine if the saturation signatures, previously reported in several urine studies, were present in the serum samples. The extracted samples were split into equal parts for processing on both GC/MS and LC/MS/MS platforms (Supplementary Figure 1) using the services of Metabolon, Inc., with the data processed using their standard analysis workflows. A total of 354 metabolites/lipids were identified, which we subsequently independently reanalyzed using in-house bioinformatics.
2. Methods and Materials
2.1. Animals and Treatments
Five groups of 12 male C57BL/6 mice were used in this study, each group member receiving a whole-body dose of 0, 2, 4, 8, or 10.4 Gy. Whole-body irradiation (WBI) was performed on anesthetized mice (intraperitoneal ketamine and xylazine 7:1 mg/ml for 100 ml/mouse) using a Shepherd137Cs -ray irradiator at a dose rate of 236cGy/min following biosafety guidelines of Albert Einstein College of Medicine (see Table 1 for details). Liver and plasma samples were collected at 24 hours, ad-lib, after the 0 Gy, sham treatment, or whole-body irradiation, and snap frozen at -80 °C. All animal studies followed protocols in accordance with the Institutional Animal Care and Use Committee (IACUC) of the Albert Einstein College of Medicine. All samples were then sent to Metabolon, Inc. (Durham, NC) for processing. A second cohort of C57BL/6 mice (n=6 per group) were used to assess response in selected plasma biomarkers at 48 and 96 hours hours post-irradiation. Additional methods detailed in Supplementary materials.
Table 1.
Study Design
| Group | Number | Description |
|---|---|---|
| 0 Gy | 12 | Sham (anesthetized, no radiation) |
| 2 Gy | 12 | 2 Gy whole body radiation |
| 4 Gy | 12 | 4 Gy whole body radiation |
| 8 Gy | 12 | 8 Gy whole body radiation |
| 10.4 Gy | 12 | 10 Gy whole body radiation |
3. Results
3.1 Liver Biomarkers
For the liver samples, random forest analysis was used to classify the sham (0 Gy) and radiation-exposed groups with a predictive accuracy of 45%, with evidence of biochemicals involved in amino acid metabolism, lipid metabolism and nucleotide metabolism varying in response to radiation dose (Figure 1). Linear discriminant analysis (LDA) showed limited separation of the dose cohorts with a predictive accuracy of 38% against sham (0 Gy). Principal component analysis failed to differentiate all 5 groups. Examination of individual biochemicals showed significant elevations in essential and long-chain fatty acids in mice exposed to the highest dose of radiation, likely a consequence of increased lipolysis, and reduced glutathione in the non-sham groups which was attributed to increased glutathione synthesis, consistent with a quenching of reactive oxygen species (ROS) in response to radiation. We took the same dataset and applied sparse partial least squares-discriminant analysis (sPLS-DA) [19] to the bounding 10.4 Gy vs. 0 Gy liver samples., Using this independent technique we confirmed clear separation between sample groups, with all of the key metabolites differentiating the groups identical to those determined by the random forest analysis.
Figure 1.
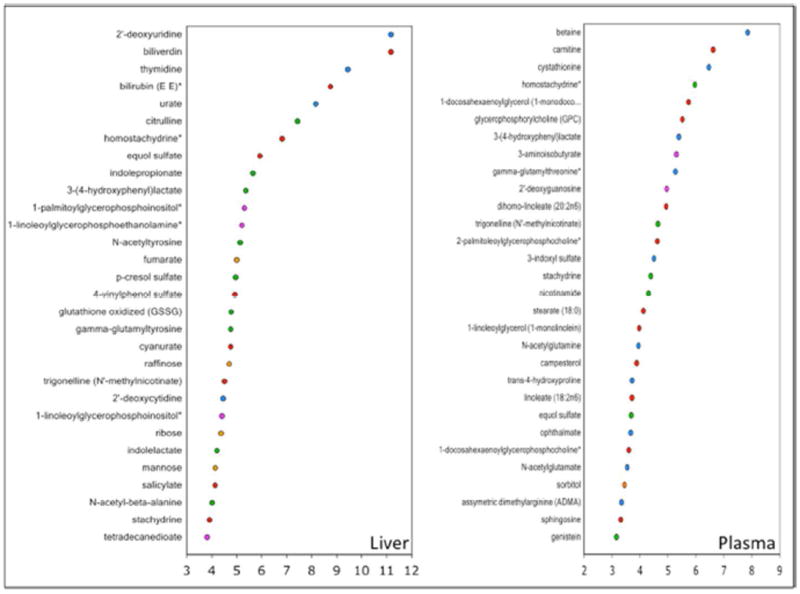
Random Forest classification analysis of liver samples (left) and plasma samples (right). In both cases the list of biochemicals associated with group separation increase along the y-axis, with the x-axis representing the mean decrease in accuracy (MDA).
3.2 Plasma Biomarkers
Random forest analysis achieved a predictive accuracy of 61% when classifying the sham (0 Gy) and radiation-exposed groups (Figure 1). The top 30 ranking biochemicals indicated important roles for nucleotide metabolism, amino acid metabolism and lipid metabolism. Linear discriminant analysis clearly segmented the sham (0 Gy) groups from the irradiated groups, although the latter lost their group identity - the overall predictive accuracy using this technique gave an accuracy of 36%. Principal component analysis failed to separate all but a subset of the 10.4 Gy group. Individual examination of biochemicals showed evidence for perturbations in pyrimidine metabolism, degradation of heme and more generally, alterations in lipid metabolism and tryptophan metabolism. In particular, stepwise changes in thymidine and 2-deoxyuridine for pyrimidine metabolism, biliverdin and bilirubin for heme degradation, certain fatty acids (lipid metabolism) and aromatic amino acids (tryptophan metabolism) were evident. The latter metabolites were of some interest, as these aromatic amino acids have been shown to have their origin within the intestinal microbiome [16,20].
We decided to implement an independent, unsupervised analysis, based around an attempt to discern a cohort of metabolites and lipids whose level of increase/decrease was consistently correlated with radiation dose. Given the full dataset we (i) used the Pearson correlation coefficient (PCC) to determine biochemicals which showed either a positive or negative correlation with increased dosage, selecting those with an absolute correlation value ≥ 0.45; (ii) the resulting 37 metabolites (Table 2) were analyzed using sPLS-DA; (iii) models with up to 10 components containing subsets of 5, 10, 15, 20, 25, 30, or 35 metabolites on each component were generated and leave one out cross-validation (LOOCV) was used to choose the best model (i.e. number of components and metabolites giving the lowest predictive error rate). We repeated this analysis removing each of the non-boundary radiation groups and noted that excluding the 4 Gy group yielded more optimal correlation scores and overall predictive accuracy. The lowest error rate was achieved using a model with 5 components and selecting 5 metabolites on each of these components. The overall error rate for this model was 0.32, representing a 68% accuracy in predicting dosage - as distinct from the 61% classification estimate determined from the random forest classifier for 0 Gy versus all irradiated groups combined. In Figure 2 we show how the samples are projected/clustered based on the multidimensional space defined by this model. Note that only the first three components are shown, but these are the ones capturing the most variance in the dataset. Not only are the irradiated groups clearly separable, but also their contiguous separation along the first component correlates with increasing radiation dose. Whilst there was general agreement between the random forest derived top ranked biochemicals and those determined from our sPLS-DA analysis, the correspondence of these 37 metabolites and increased radiation dose gives us a tractable cohort of potential biomarkers with which to not only to validate, but also to form the basis of a plasma-based metabolite panel for radiation dose estimation.
Table 2.
37 Metabolites.
| Name | Pearson Correlation |
|---|---|
| thymidine | 0.74 |
| 2’ - deoxyuridine | 0.73 |
| 2-aminobutyrate | 0.46 |
| 2-hydroxyisobutyrate | 0.45 |
| 1-eicosadienoylglycerophosphocholine* | −0.46 |
| 2-palmitoylglycerophosphocholine* | −0.46 |
| ectoine | −0.46 |
| homocitrulline | −0.46 |
| 3-hydroxypropanoate | −0.48 |
| citrulline | −0.48 |
| 1-arachidonoylglycerophosphoethanolamine* | −0.48 |
| equol glucuronide | −0.49 |
| 3-phenylpropionate (hydrocinnamate) | −0.50 |
| stachydrine | −0.50 |
| 1-palmitoleoylglycerophosphocholine* | −0.50 |
| chiro-inositol | −0.50 |
| docosahexaenoate (DHA22:6n3) | −0.50 |
| gamma-glutamylisoleucine* | −0.50 |
| 1-arachidonoylglycerophosphoinositol* | −0.51 |
| eicosapentaenoate (EPA20:5n3) | −0.52 |
| **pipecolate | −0.52 |
| dihomo-linolenate (20:3n3 or n6) | −0.53 |
| 1-myristoylglycerophosphocholine | −0.53 |
| 2-linoleoylglycerophosphoethanolamine* | −0.53 |
| salicylate | −0.54 |
| **phenol sulfate | −0.54 |
| N-acetylhistidine | −0.55 |
| 4-vinylphenol sulfate | −0.55 |
| 1-linoleoylglycerophosphoethanolamine* | −0.56 |
| **3-indoxyl sulfate | −0.56 |
| **p-cresol sulfate | −0.59 |
| **indolelactate | −0.64 |
| 1-linoleoylglycerophosphoinositol* | −0.64 |
| homostachydrine* | −0.65 |
| 1-palmitoylglycerophosphoinositol* | −0.67 |
| **equol sulfate | −0.70 |
| **indolepropionate | −0.71 |
37 biochemical compounds identified following Pearson Correlation and sPLS-DA and LOOCV validation of the full plasma data set. The metabolites preceded by ** have been previously shown to have their origin in the intestinal microbiome.
Figure 2.
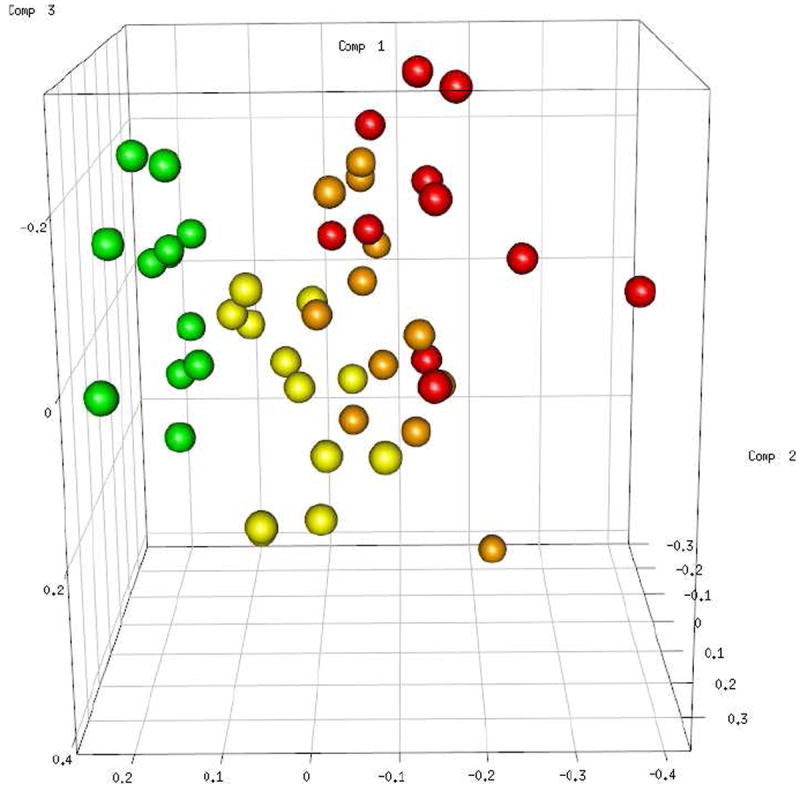
All the plasma sample groups have been projected/clustered based on the multidimensional space defined by the best sPLS-DA model determined from the 37 biochemical compounds listed in Table 1. Note that only the first three components are shown, but these are the ones capturing the most variance in the dataset. The samples are colored as follows: Green -0 Gy, Yellow -2 Gy, Orange -8 Gy, Red -10.4 Gy.
3.3 Plasma Microbiome Signature
Of the biochemicals that significantly correlated with radiation dose, those that increased (thymidine and 2-deoxyuridine) recapitulated those identified in the context of pyrimidine metabolism (Figure 3, upper panel), and of those that decreased (< -0.55), four biochemicals in particular (indolepropionate, equol sulfate, indole lactate and 3-indoxyl sulfate) are all tryptophan compounds with a gut microbiome metabolic contribution (Figure 3, lower panel). Thus it seems clear that the predominant metabolite components of the plasma response to enhanced irradiation are those associated with DNA strand breaks/RNA damage (increases with irradiated dose) and those metabolites originating or associated directly with the gut microbiome (decreases with irradiated dose).
Figure 3.
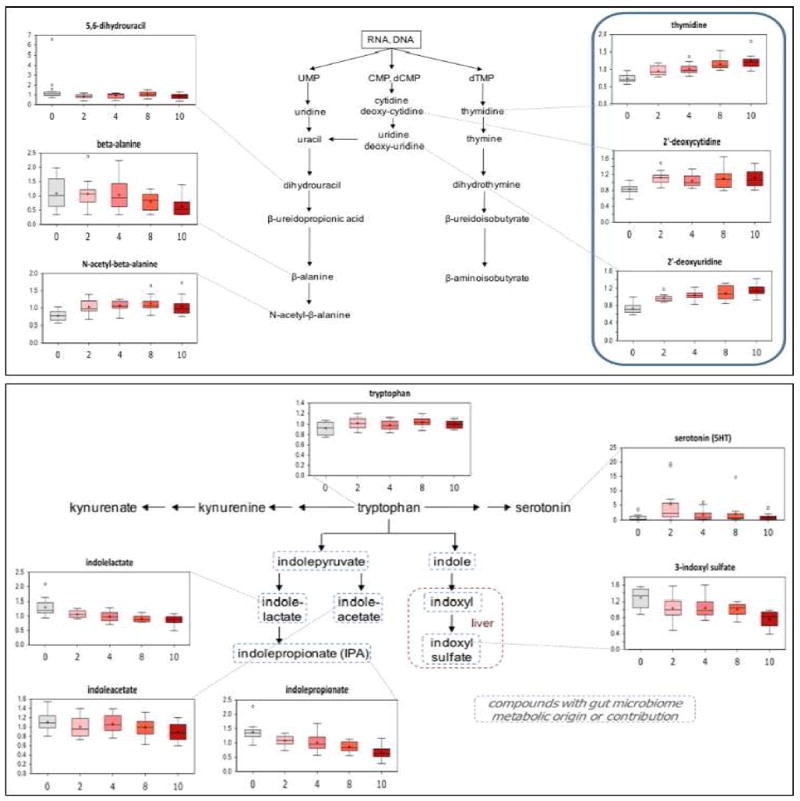
Pyrimidine (upper panel) and tryptophan (lower panel) metabolism pathways and associated metabolite measurements in plasma. The x-axis indicates received radiation dosage in Gy.
3.4 Plasma Response beyond 24 hours
To further assess this gut microbiome response, a second cohort of male C57BL/6 mice were used to validate a subset of the identified biomarkers and to examine their behavior in the ad-lib state beyond 24 hours post-irradiation. In this second experiment, selected markers were measured at either 48 or 96 hours post-irradiation in groups (n=6 per group) of mice receiving a 10.4 Gy radiation dose as compared to non-irradiated controls (Figure 4). Results from this experiment show that indole-3-pyruvate and indole-3-acetamide are also detected as being markedly reduced in irradiated samples along with the previously identified biomarkers IPA and indole lactate and that decreases in indole levels are evident at both timepoints, indicating persistent changes in tryptophan metabolism.
Figure 4.
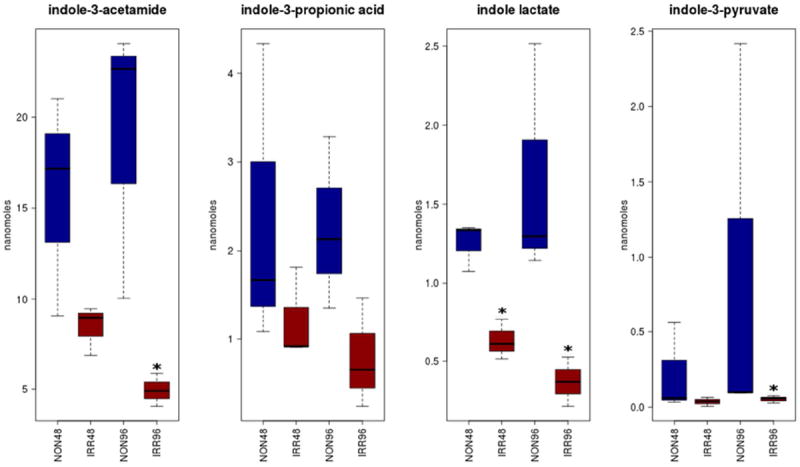
Indole metabolites measured in a second cohort of mice receiving 10.4 Gy show persistent changes at both 48 (IRR48) and 96 (IRR96) hours post-irradiation compared to their respective non-irradiated counterparts (NON48 and NON96). Asterisks identify significantly changed metabolites (p<=0.1) between irradiated samples and controls at each timepoint using a non-parametric Wilcoxon rank-sum test.
3.5 Correlation between Liver and Plasma
We attempted to discern a common metabolite/lipid association between the liver and plasma - choosing the boundary 0 Gy and 10.4 Gy groups to perform this analysis so as to best emphasize any correlated signal across the two profiles. The highest scoring biochemical, common to both liver and plasma datasets was 3-indoxyl sulfate, a sulfated gut microbiome metabolite of tryptophan processed in the liver; as 3-indoxyl sulfate is also dependent on renal function we performed additional screening of a panel of markers for renal function (Supplementary Table 1), which show relatively little change, indicating that the plasma/liver correlation is specific to those tissues.
4. Discussion
The purpose of this work was to determine if consistent trends in metabolite and lipid concentrations were discernible in blood plasma taken from whole body irradiated mice 24 hours after a range of doses equivalent to non-lethal, sub-lethal and lethal exposures. We included liver samples in our studies as the liver plays a major role in metabolism, especially in the first-pass metabolism of intestinal absorbates in the portal circulation. The lack of a discernible response in liver samples below 10.4 Gy indicates the degree to which the liver can compensate for additional oxidative stress caused by ionizing radiation. These results are in line with our expectations given the fact that in both mouse and humans, higher doses (>10 Gy) are required to induce radiation-induced liver disease (RILD).
Of the 37 plasma biochemicals whose concentrations were correlated to radiation dose, the dominant underlying metabolic pathways were those associated with pyrimidine and tryptophan metabolism. Whilst the presence of metabolite compounds associated with these two pathways is in agreement with several previous irradiation studies, our work is the first to indicate their concentration being consistently correlated with received dose. We did not see evidence for a dose-response saturation as reported by previous studies on urine derived metabolites - this serves to highlight the potential advantage in using plasma as a biofluid for such functional biodosimetry studies. Of particular interest is the extent to which we can implicate a predominantly microbiome signature evident in murine plasma in the form of several of the indole family of biochemical compounds, whose presence is inversely correlated with radiation dose from 0 Gy to 10.4 Gy. In addition we see a similar correlation between plasma and liver for 3-indoxyl sulfate, which is a by-product of hepatic processing of indole, between 0 and 10.4 Gy.
Whilst the intestinal microbial response to ionizing radiation has not been formally determined, the study of ionizing radiation on individual prokaryotes has typically focused on model strains(most notably E. coli), in isolation and under controlled conditions. Inhibition of tryptophanase, the engine of tryptophan metabolism, has been reported in the literature [21], but involved doses ≥ 103 Gy and was quantified very shortly after radiation exposure, where the presence of significant numbers of both single and double strand breaks might be expected to compromise the conventional prokaryotic transcriptome. Clearly the radiation dosage in the whole body context is unlikely to yield such a profound transcriptional response. Assuming that the colonic microbiome is in effect transcriptionally unperturbed by increased radiation doses within the range [2 -10 Gy], a possible explanation could be that radiation casues damage to the intestinal epithelium, compromising the transfer of indoles and other microbiome metabolites to the host. Indeed, the observed concentration of citrulline in our study is inversely correlated to radiation dose, consistent with damage to the intestinal epithelium. However, such damage is typically associated with an increase in epithelial permeability [22], yet despite this we see a consistent trend for a decrease in indole production. A recent study presented clear evidence for significant microbial dysbiosis in fecal pellets taken from Wistar rats exposed to whole body 10 and 18 Gy doses based on community profiling determined daily over a 21 day period post-irradiation [23].
In summary, our data suggests that incident ionizing radiation perturbs the murine gut microbial metabolome, manifested as suppression of this super-organism’s collective tryptophan metabolism, offering an attractive means to infer a likely whole-body radiation dose based on the concentrations of indole compounds in a given blood sample. Confirming the robustness of this gut microbial metabolome to radiation in the context of an overall metabolic response, both directly and as a consequence of the host-microbiome interaction, will be critical in being able to distil the 37 compounds identified in this study into a subset capable of robustly estimating prior radiation exposure for functional biodosimetry.
Supplementary Material
Footnotes
Conflict of Interest: None
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Contributor Information
Pilib Ó Broin, Department of Genetics, Albert Einstein College of Medicine
Bhavapriya Vaitheesvaran, Department of Medicine, Diabetes Center, Stable Isotope and Metabolomics Core Facility, Albert Einstein College of Medicine
Subhrajit Saha, Department of Radiation Oncology, Albert Einstein College of Medicine
Kirsten Hartil, Department of Medicine, Diabetes Center, Stable Isotope and Metabolomics Core Facility, Albert Einstein College of Medicine
Emily I. Chen, Department of Pharmacology, Proteomics Shared Resource, Herbert Irving Comprehensive Cancer Center, Columbia University Medical Center
Devorah Goldman, Department of medicine, OHSU
William Harv Fleming, Department of Medicine, OHSU.
Irwin J. Kurland, Department of Medicine, Diabetes Center, Stable Isotope and Metabolomics Core Facility, Albert Einstein College of Medicine
Chandan Guha, Department of Radiation Oncology, Albert Einstein College of Medicine
Aaron Golden, Department of Genetics, Albert Einstein College of Medicine, and Department of Mathematical Sciences, Yeshiva University
References
- 1.Stone HB, Coleman CN, Anscher MS, McBride WH. Effects of radiation on normal tissue: Consequences and mechanisms. The lancet oncology. 2003;4:529–536. doi: 10.1016/s1470-2045(03)01191-4. [DOI] [PubMed] [Google Scholar]
- 2.Friesecke I, Beyrer K, Fliedner TM system MtMtpfravaabfacg. How to cope with radiation accidents: The medical management. Br J Radiol. 2001;74:121–122. doi: 10.1259/bjr.74.878.740121. [DOI] [PubMed] [Google Scholar]
- 3.Fiehn O. Metabolomics--the link between genotypes and phenotypes. Plant molecular biology. 2002;48:155–171. [PubMed] [Google Scholar]
- 4.Tyburski JB, Patterson AD, Krausz KW, Slavik J, Fornace AJ, Jr, Gonzalez FJ, Idle JR. Radiation metabolomics. 1. Identification of minimally invasive urine biomarkers for gamma-radiation exposure in mice. Radiat Res. 2008;170:1–14. doi: 10.1667/RR1265.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Tyburski JB, Patterson AD, Krausz KW, Slavik J, Fornace AJ, Jr, Gonzalez FJ, Idle JR. Radiation metabolomics. 2. Dose- and time-dependent urinary excretion of deaminated purines and pyrimidines after sublethal gamma-radiation exposure in mice. Radiat Res. 2009;172:42–57. doi: 10.1667/RR1703.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Parizek J, Arient M, Dienstbier Z, Skoda J. Deoxycytidine in urine as an indicator of changes after irradiation. Nature. 1958;182:721–722. doi: 10.1038/182721a0. [DOI] [PubMed] [Google Scholar]
- 7.Lanz C, Patterson AD, Slavik J, Krausz KW, Ledermann M, Gonzalez FJ, Idle JR. Radiation metabolomics. 3. Biomarker discovery in the urine of gamma-irradiated rats using a simplified metabolomics protocol of gas chromatography-mass spectrometry combined with random forests machine learning algorithm. Radiat Res. 2009;172:198–212. doi: 10.1667/RR1796.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Johnson CH, Patterson AD, Krausz KW, Lanz C, Kang DW, Luecke H, Gonzalez FJ, Idle JR. Radiation metabolomics. 4. Uplc-esi-qtofms-based metabolomics for urinary biomarker discovery in gamma-irradiated rats. Radiat Res. 2011;175:473–484. doi: 10.1667/RR2437.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lutgens LC, Deutz NE, Gueulette J, Cleutjens JP, Berger MP, Wouters BG, von Meyenfeldt MF, Lambin P. Citrulline: A physiologic marker enabling quantitation and monitoring of epithelial radiation-induced small bowel damage. Int J Radiat Oncol Biol Phys. 2003;57:1067–1074. doi: 10.1016/s0360-3016(03)00781-8. [DOI] [PubMed] [Google Scholar]
- 10.Barrett A, Jacobs A, Kohn J, Raymond J, Powles RL. Changes in serum amylase and its isoenzymes after whole body irradiation. British medical journal. 1982;285:170–171. doi: 10.1136/bmj.285.6336.170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bertho JM, Demarquay C, Frick J, Joubert C, Arenales S, Jacquet N, Sorokine-Durm I, Chau Q, Lopez M, Aigueperse J, Gorin NC, Gourmelon P. Level of flt3-ligand in plasma: A possible new bio-indicator for radiation-induced aplasia. Int J Radiat Biol. 2001;77:703–712. doi: 10.1080/09553000110043711. [DOI] [PubMed] [Google Scholar]
- 12.Guipaud O. Serum and plasma proteomics and its possible use as detector and predictor of radiation diseases. Adv Exp Med Biol. 2013;990:61–86. doi: 10.1007/978-94-007-5896-4_4. [DOI] [PubMed] [Google Scholar]
- 13.Khan AR, Rana P, Devi MM, Chaturvedi S, Javed S, Tripathi RP, Khushu S. Nuclear magnetic resonance spectroscopy-based metabonomic investigation of biochemical effects in serum of gamma-irradiated mice. Int J Radiat Biol. 2011;87:91–97. doi: 10.3109/09553002.2010.518211. [DOI] [PubMed] [Google Scholar]
- 14.Khan AR, Rana P, Tyagi R, Kumar I, Devi MM, Javed S, Tripathi RP, Khushu S. Nmr spectroscopy based metabolic profiling of urine and serum for investigation of physiological perturbations during radiation sickness. Metabolomics : Official journal of the Metabolomic Society. 2011;7:583–592. [Google Scholar]
- 15.Liu Y, Lin Z, Tan G, Chu Z, Lou Z, Zhang J, Hong Z, Chai Y. Metabonomic studies on potential plasma biomarkers in rats exposed to ionizing radiation and the protective effects of hong shan capsule. Metabolomics : Official journal of the Metabolomic Society. 2013;9:1082–1095. [Google Scholar]
- 16.Wikoff WR, Anfora AT, Liu J, Schultz PG, Lesley SA, Peters EC, Siuzdak G. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc Natl Acad Sci U S A. 2009;106:3698–3703. doi: 10.1073/pnas.0812874106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Crawford PA, Gordon JI. Microbial regulation of intestinal radiosensitivity. Proc Natl Acad Sci U S A. 2005;102:13254–13259. doi: 10.1073/pnas.0504830102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Onoue M, Uchida K, Yokokura T, Takahashi T, Mutai M. Effect of intestinal microflora on the survival time of mice exposed to lethal whole-body gamma irradiation. Radiat Res. 1981;88:533–541. [PubMed] [Google Scholar]
- 19.Le Cao KA, Boitard S, Besse P. Sparse pls discriminant analysis: Biologically relevant feature selection and graphical displays for multiclass problems. BMC bioinformatics. 2011;12:253. doi: 10.1186/1471-2105-12-253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Grison S, Martin JC, Grandcolas L, Banzet N, Blanchardon E, Tourlonias E, Defoort C, Fave G, Bott R, Dublineau I, Gourmelon P, Souidi M. The metabolomic approach identifies a biological signature of low-dose chronic exposure to cesium 137. Journal of radiation research. 2012;53:33–43. doi: 10.1269/jrr.11071. [DOI] [PubMed] [Google Scholar]
- 21.Pollard EC, Lydersen BK. The action of ionizing radiation on the synthesis of rna and two inducible enzymes in escherichia coli. Radiat Res. 1972;50:293–300. [PubMed] [Google Scholar]
- 22.MacNaughton WK. Review article: New insights into the pathogenesis of radiation-induced intestinal dysfunction. Alimentary pharmacology & therapeutics. 2000;14:523–528. doi: 10.1046/j.1365-2036.2000.00745.x. [DOI] [PubMed] [Google Scholar]
- 23.Lam V, Moulder JE, Salzman NH, Dubinsky EA, Andersen GL, Baker JE. Intestinal microbiota as novel biomarkers of prior radiation exposure. Radiat Res. 2012;177:573–583. doi: 10.1667/rr2691.1. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


