Abstract
Melanin is a unique pigment with myriad functions that is found in all biological kingdoms. It is multifunctional, providing defense against environmental stresses such as ultraviolet (UV) light, oxidizing agents and ionizing radiation. Melanin contributes to the ability of fungi to survive in harsh environments. In addition, it plays a role in fungal pathogenesis. Melanin is an amorphous polymer that is produced by one of two synthetic pathways. Fungi may synthesize melanin from endogenous substrate via a 1,8-dihydroxynaphthalene (DHN) intermediate. Alternatively, some fungi produce melanin from l-3,4-dihydroxyphenylalanine (l-dopa). The detailed chemical structure of melanin is not known. However, microscopic studies show that it has an overall granular structure. In fungi, melanin granules are localized to the cell wall where they are likely cross-linked to polysaccharides. Recent studies suggest the fungal melanin may be synthesized in internal vesicles akin to mammalian melanosomes and transported to the cell wall. Potential applications of melanin take advantage of melanin's radioprotective properties and propensity to bind to a variety of substances.
Keywords: Fungi, Melanin, Cell wall, Vesicle, Chitin, Radioprotection
Introduction
Many fungal species produce melanin, a biologically important pigment. Melanin is found throughout nature, often providing a protective role such as from ultraviolet radiation. Despite its importance and ubiquity, there are many fundamental questions unanswered regarding the pigment such as the details of its chemical structure. This is due to the fact that melanin is insoluble and, therefore, cannot be studied by standard biochemical techniques. Melanin production by fungi contributes to the virulence of pathogens of humans as well as those of food crops. The pigment enhances fungal resistance to environmental damage as well. For example, radiation-resistant melanized fungi can survive harsh climates including Antarctica and contaminated nuclear reactors (Rosa et al. 2010; Zhdanova et al. 2000). Melanized fungi have even been found to survive in dishwashers where they must withstand heat and detergents (Zalar et al. 2011).
A major reason for studying fungal melanin is the pigment's contribution to virulence. A retrospective study of 18 cases of fungal infections of the CNS at a Texas hospital reveals that the majority of the fungi were dematiaceous, or melanotic, fungi (Raparia et al. 2010). Studies with animal models demonstrate that melanin is a virulence factor in several fungal pathogens. In Cryptococcus neoformans, the genes required for melanization contribute to host death (Salas et al. 1996) and dissemination from the lungs to other organs (Noverr et al. 2004). In Cryptococcus gattii, a related organism and causative agent of the current cryptococcosis outbreak in Vancouver, analysis of more virulent and less virulent strains shows that the more virulent strains have greater expression of melanin synthesis genes and produce more melanin (Ngamskulrungroj et al. 2011). Similarly, in Paracoccidioides brasiliensis, experimental infections with melanized cells result in higher fungal burdens in animals compared to nonmelanized cells (Silva et al. 2009). In addition, infection increases the expression of melanin synthesis genes in this fungus (Bailao et al. 2006). Lastly, in a sporotrichosis model, melanized fungi show greater dissemination in a mouse footpad model compared to mutants unable to make melanin (Madrid et al. 2010).
Fungal melanin can influence the immune response of the host. For example, melanin interferes with the normal function of phagocytic cells. Melanized Fonsecaea pedrosoi cells reduce the oxidative burst capacity of macrophages (Cunha et al. 2010), nitrite production (Bocca et al. 2006) and phagocytosis of the fungi (Cunha et al. 2005). Inhibition of phagocytosis has also been observed for melanized C. neoformans (Wang et al. 1995) and P. brasiliensis (da Silva et al. 2006). In Aspergillus fumigatus, melanin inhibits apoptosis in macrophages that have phagocytosed melanized conidia (Volling et al. 2011). Melanin can modulate immune function in other ways. In experimental mouse infections, cryptococcal melanin alters cytokine levels in response to infection (Mednick et al. 2005) and activates the complement system (Rosas et al. 2002). Conversely, A. fumigatus melanin inhibits cytokine production in the host, possibly by blocking pathogen-associated molecular pattern (PAMP) recognition by the immune system (Chai et al. 2010).
Melanin is critical to host invasion in plant pathogens as well. Fungi produce appressoria, structures that penetrate plant tissue, allowing the organisms to invade the host. Melanin in the cell wall of these structures provides mechanical strength to the appressoria that aids in tissue penetration. For example, in coffee berry disease caused by Colletotrichum kahawae, inhibition of melanization decreases the turgor pressure of the appressoria and consequently, the virulence of the fungus (Chen et al. 2004). Melanized appressoria have been shown to be important in the virulence of other plant pathogens including the rice blast fungus, Magnaporthe grisea (Howard and Valent 1996) and Black spot disease of roses (Diplocarpon rosae) (Gachomo et al. 2010).
This review will address the structure and synthesis of fungal melanin. Melanins are a group of related pigments that share physical and chemical traits. They are generally black or brown in color, although other colors exist. Melanins are resistant to chemical degradation by acids and insoluble in most substances. They can only be broken down by oxidation and dissolve only in alkaline solvents. Melanins are both negatively charged and hydrophobic (Nosanchuk and Casadevall 2003b). They contain unpaired electrons that can be detected by electron paramagnetic resonance (EPR) as a stable free radical signal (Enochs et al. 1993). Melanins are polymers of phenolic compounds. In some instances, the polymer subunits have been discovered; however, the exact arrangement of these subunits in the polymer remains unknown (Wakamatsu and Ito 2002).
Melanin biosynthesis
Two pathways of melanin synthesis are found in fungi. Many fungi synthesize melanin via the DHN pathway. In this pathway, the precursor molecule, acetyl coA or malonyl coA, is produced endogenously. The first step, formation of 1,3,6, 8-tetrahydroxynaphthalene (1,3,6,8-THN), is catalyzed by a polyketide synthase (PKS). After 1,3,6,8-THN production, a series of reduction and dehydration reactions produce the intermediates scytalone, 1,3,8-trihydroxynaphthalene, vermelone, and finally 1,8-dihydroxynaphthalene (DHN). Polymerization of DHN leads to formation of melanin (Butler and Day 1998a; Langfelder et al. 2003).
Polyketide synthases belong to a large enzyme family with diverse roles in production of secondary metabolites such as pigments, toxins, antibiotics and signaling molecules. These multidomain enzymes join simple subunits into long chains or cyclic compounds (Schumann and Hertweck 2006). PKS genes in fungi are frequently found in gene clusters, in which the genes encoding the various enzymes needed for the biosynthesis of a product are nearby each other on the chromosome. Phylogenetic analysis of PKS genes in ascomycetes reveals that the genes are widely distributed among the subphylum Pezizomycotina, but not in early branching ascomycetes such as Saccharomyces cerevisiae (Kroken et al. 2003).
Melanin biosynthesis gene clusters have been characterized in several fungi. In A. fumigatus, the cluster consists of six genes and spans 19 kb pairs of DNA. The genes in the cluster include alb1, the PKS gene, arp1, the scytalone reductase, arp2, the hydroxynaphthalene reductase, abr1, a multicopper oxidase, abr2, a putative laccase, and ayg1, a gene with unknown function (Tsai et al. 1999). Similarly, in Penicillium marneffei, the melanin biosynthesis gene cluster spans 25.3 kb pairs and contains a PKS gene, modifying enzymes and hypothetical proteins (Woo et al. 2010).
A few fungi synthesize melanin via l-3,4-dihyroxyphenylalanine (l-dopa), in a pathway that resembles mammalian melanin biosynthesis. There are two possible starting molecules in this second pathway, l-dopa or tyrosine. If l-dopa is the precursor molecule, it is oxidized to dopaquinone by laccase. If tyrosine is the precursor, it is first converted to l-dopa and then dopaquinone. The same enzyme, tyrosinase, carries out both steps. Dopaquinone is a highly reactive molecule. Intramolecular nucleophilic addition by the amino group produces cyclodopa. Cyclodopa is then oxidized to form dopachrome. Tautomerization of dopachrome forms dihydroxyindoles that polymerize into melanin (Land et al. 2004; Langfelder et al. 2003; Riley 1997). It is noteworthy that in both melanin pathways are found many fungi such as Sporothrix schenckii (Almeida-Paes et al. 2009), while others such as C. neoformans rely exclusively on the l-dopa pathway.
The fact that C. neoformans makes melanin only by the l-dopa pathway combined with an absolute dependence on exogenous substrate for melanin synthesis has made this organism an important system for studying fungal melanization, for these characteristics allow considerable control on the type of melanin made. The C. neoformans genome encodes two laccase genes, LAC1 and LAC2, with LAC1 being the main producer of melanin (Pukkila-Worley et al. 2005). The exact role of the LAC2 gene remains to be determined. Interestingly, there are differences in the cellular location of the two laccases. Laccase encoded by LAC1 is localized to the cell wall (Zhu et al. 2001). In contrast, a LAC2–GFP fusion protein is primarily cytoplasmic (Missall et al. 2005). In addition, gene expression analysis shows some differences in the regulation of the LAC1 and LAC2 genes (Missall et al. 2005).
l-Dopa is one of several melanin substrates identified in C. neoformans. Other catecholamines can serve as substrates such as dopamine and norepinephrine as well as d-dopa (Eisenman et al. 2007). The substrate list continues to grow, indicating that the fungus may use a wide array of substrates, maximizing its ability to produce melanin. Plant-derived substances including flavonoids (Fowler et al. 2011) and caffeic acid (Vidotto et al. 2004) are among the list, as is bacterially-derived homogentisic acid (Frases et al. 2007). Coculture of C. neoformans with the bacteria Klebsiella aerogenes results in melanin production by the fungus. In this interaction, dopamine is produced by the bacteria and released into a medium and subsequently used by C. neoformans as a melanin substrate (Frases et al. 2006). Despite the fact that pigments are produced from many different substrates, it is important to note that not all of the pigments produced are the same. Different properties are observed for melanins derived from different substrates. Comparison of the catecholamines l-dopa, methyldopa, epinephrine and norepinephrine shows differences in term of color, yield and thickness of the cell wall melanin layer. Additionally, the pigments vary in the strength of the stable free radical signal detectable by EPR. In fact, such a signal is absent from epinephrine-derived pigment, implying the lack of a key criteria in defining a melanin (Garcia-Rivera et al. 2005).
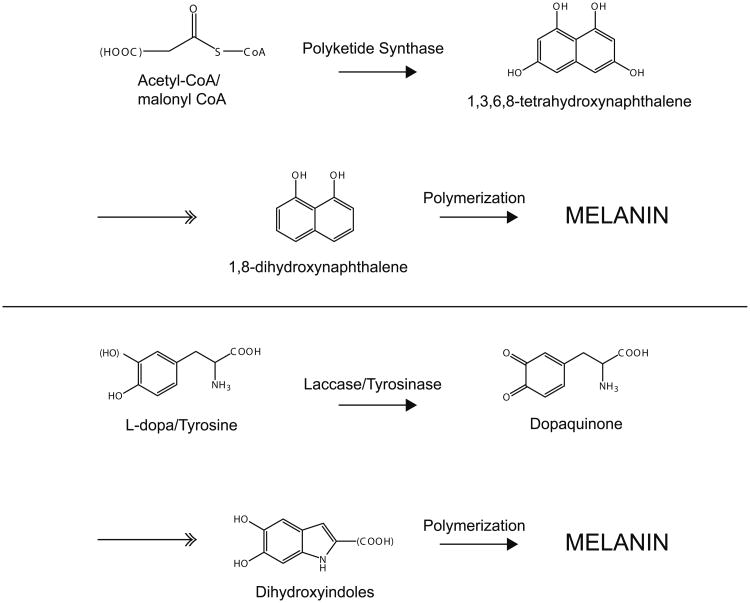
Figure 1 summarizes the key steps in melanin synthesis of each pathway. The pathways are similar in that phenolic and/or indolic precursors are polymerized into melanin. However, each step in the DHN pathway is catalyzed enzymatically, while the l-dopa pathway is believed to occur spontaneously after the initial oxidation by laccase. Additionally, melanin produced by DHN pathway contains carbon and oxygen only, while the l-dopa pathway melanins also contain nitrogen. Melanin synthesized via the l-dopa pathway is referred to as eumelanin. This pathway may also incorporate sulfur atoms to produce pheomelanins, which are red or yellow in color (Ito and Wakamatsu 2003). For a further discussion of the steps in melanin synthesis, see the review by Land et al (2004). We note that the final step leads to the word ‘melanin’ which is written out because the structure of melanin is unsolved. Current models of melanin structure envision a structure with graphitic planar sheets of covalently linked aromatic molecules (Clancy and Simon 2001; Diaz et al. 2005; Meng and Kaxiras 2008). Even with some insight into the structure we emphasize that the molecular structure could be locally variable such that the pigment has an amorphous and noncrystalline structure. Furthermore, we note that the structure of melanin is beyond our current technological horizon as the available analytical methods are insufficient to yield a structural solution.
Fig. 1.
Comparison of DNH (top) and l-dopa (bottom) melanin synthesis pathways. In the DHN pathway, the precursor, malonyl CoA or acetyl CoA, is a substrate of polyketide synthase. Several enzymatic steps are required to produce DHN, the subunit of the melanin polymer. In the l-dopa pathway, the precursor, tyrosine or l-dopa, is a substrate of tyrosinase or laccase. Several enzymatic steps are required to produce dihydroxyindole, the subunit of the melanin polymer
Copper is important for the production of melanin by both the DHN and l-dopa pathways. In Gaeumannomyces graminis, addition of copper to the medium results in dark cultures and the cells have a layer of melanin in the cell wall visible by TEM (Caesar-Tonthat et al. 1995). Similarly, variation in copper levels among commercial fungal media (potato dextrose agar) correspond to variable pigmentation in multiple fungal species including Trichoderma harzianum, Stachybotrys atra, Fusarium culmorum, and Cladosporium herbarum (Griffith et al. 2007). Molecular studies also highlight the importance of copper in melanization. Mutant screens for albino mutants have uncovered several genes involved in copper homeostasis in C. neoformans including a CLC-type chloride channel gene (Zhu and Williamson 2003) and the copper transporters CCC2 and ATX1 (Walton et al. 2005). Similarly, deletion of the copper-transporting ATPase, BcCCC2 in Botrytis cinerea reduces melanization and formation of sclerotia (Saitoh et al. 2010b). In these examples, the albino phenotype can be rescued by addition of excess copper to the growth media. The importance of copper to melanin production is likely to be due to its requirement as a cofactor for melanin biosynthesis enzymes. Lac-cases that catalyze l-dopa melanin production are copper-dependent (Williamson 1997). In addition, multi-copper oxidases and laccases have been identified in PKS gene clusters of A. fumigatus (Tsai et al. 1999), Aspergillus niger (Jorgensen et al. 2011), and Cochliobolus heterostrophus (Saitoh et al. 2010a). The function of these copper-dependent enzymes in DHN melanin synthesis is hypothesized to be in polymerization of the DHN subunits into melanin. Additional roles for copper are suggested. For example, copper has been shown to also stimulate laccase gene expression in C. neoformans (Jiang et al. 2009; Zhu and Williamson 2003).
Melanin in the fungal cell wall
Melanin is associated with the cell wall of fungi. The cell wall is a multilayered structure that provides cell shape and protection from osmotic stress. It is important in virulence through direct interaction with the host. The cell wall is primarily composed of branched fibrillar polysaccharides including β-linked glucan, chitin, mannan and galactofuran, which are found in variable amounts depending on the fungal species. In addition, proteins are also present either covalently linked or more loosely associated secreted proteins (Latge 2007; Latge et al. 2005; Ruiz-Herrera et al. 2006). Melanin can be found in the inner or outer layers of the cell wall depending on the fungal species. For example, in C. neoformans, melanin is found in the innermost layer of the cell wall, near the plasma membrane (Nosanchuk and Casadevall 2003a). In contrast, in Candida albicans, melanin is found on the outer layer and in the plant pathogen G. graminis, it is in the middle of the cell wall (Caesar-Tonthat et al. 1995; Walker et al. 2010). Recent studies explore the association of melanin with cell wall components as well as the overall architecture of cell wall melanin.
Since melanin is insoluble, many traditional biochemical techniques are unsuitable for studying the pigment. Progress on melanin structure has been made with the use of solid state NMR. When purified melanin shells (called melanin “ghosts”) from C. neoformans are cultured in the presence of various 13C labeled precursors and analyzed by this technique, the data show that melanin is covalently cross-linked with polysaccharide components of the cell wall, especially those containing mannose. These results indicate that melanin may not be a discrete layer of the cell wall but rather an integrated component (Zhong et al. 2008).
Melanin has a significant impact on the overall structure and architecture of the cell wall. Comparison of melanized wild type or albino mutant conidia of A. fumigatus reveals drastic differences between melanized and nonmelanized cell surfaces. A combination of SEM and TEM shows the albino mutants lack a thick and rough outer layer of the cell wall. Further analysis reveals that the presence of melanin in the cell wall increased both the negative charge (Zeta potential) and hydrophobicity of the cells (Pihet et al. 2009).
Melanin also impacts the overall porosity of the cell wall. Jacobson and Ikeda employ a biochemical approach to study porosity. By using SDS-boiled cell walls of C. neoformans to make a “size exclusion column” and comparing the elution of dextran from columns made of melanized versus nonmelanized cells walls, they show that melanization substantially reduces the pore sizes found in the cell wall (Jacobson and Ikeda 2005). In an independent study, NMR cryoporometry is used to compare pore sizes in C. neoformans melanin ghosts. Melanin ghosts prepared from older cultures have smaller pores, indicating that melanization is associated with a decrease in porosity. The reduction in pore size combined with the absorption properties of melanin are suggested as a mechanism for the acquired drug resistance of melanized cells to polyenes and echinocandins (Eisenman et al. 2005).
Numerous microscopy studies on the overall architecture of the melanin polymer in fungi reveal that melanin forms granular particles in the cell wall. Surface imaging of C. neoformans melanin ghosts by SEM and AFM shows that the melanin is composed of irregular granules 50–80 nm in diameter (Eisenman et al. 2005). Strikingly similar granules cover the surface of melanized F. pedrosoi sclerotic cells, often in groups or patches (Franzen et al. 2006). Granular particles are also observed in the salt-tolerant black yeast, Hortaea werneckii (Kogej et al. 2007) and melanin isolated from spores of the mushroom Agaricus bisporum (Hegnauer et al. 1985). TEM cross-sectional images of the C. neoformans melanin ghosts further reveal multiple layers of melanin. The width of each layer corresponds to the diameter of the granules, suggesting that melanin in the cell wall of C. neoformans is composed of multiple layers of these particles (Eisenman et al. 2005). The organization of the melanin layer in spherical particles of approximately similar size raises the conundrum of how a polymerization reaction produces such relatively homogenous particles, a problem that can be resolved by positing that melanization occurs in vesicles (see below).
The observation that melanin in the fungal cell wall consists of small granules rather than a solid layer suggests the presence and need of a “scaffold” to keep the melanin in place. Cross-linking of melanin to cell wall components is a possible mechanism. Indeed, published reports suggest that chitin may be the scaffold for melanin in fungal cells walls. Chitin is a polymer of β(1,4)-linked N-acetylglucosamine subunits joined in antiparallel chains hydrogen bonding to produce strong microfibrils. Chitin cross-links to other cell wall polysaccharides and proteins and may form up to 40% of the fungal cell wall. Chitin is synthesized by chitin synthase (CHS) which uses UDP-N-acetylglucosamine as a substrate to add residues to the growing chitin chain. Many fungi have multiple CHS genes, with different roles in growth and budding (Munro and Gow 2001).
In Wangiella dermatitidis and C. neoformans, mutation of genes involved in chitin synthesis has a dramatic effect on the melanin phenotypes. Deletion of the WdCHS4 gene results in the release of dark pigment in culture supernatants and agar surrounding W. dermatitidis cells (Wang et al. 1999). Similarly, melanin “halos” around C. neoformans growth in agar and darkening of culture supernatants occurs when chitin-related genes are disrupted including chitin synthases, chitin regulatory genes and chitin deacetylases (Baker et al. 2007; Banks et al. 2005; Walton et al. 2005). Biochemical data suggests a direct interaction between chitin and fungal melanin. Melanized cell walls of Aspergillus nidulans do not release chitin when subjected to enzymatic lysis in contrast to nonmelanized cells. Additionally, solubilized melanin inhibits chitinase activity by binding directly to chitin (Bull 1970).
Chitin is important for melanin synthesis by C. albicans. This yeast previously was not known to produce melanin. Although it doesn't produce dark colonies, melanin-like particles are found associated with C. albicans when it is cultured with l-dopa (Morris-Jones et al. 2005). Addition of N-acetylglucosamine to cultures incubated with l-dopa increases melanin production as measured by OD310. Conversely, C. albicans chitin synthase mutants do not produce detectable melanin. TEM analysis of the mutants show melanin granules accumulate inside the cells (Walker et al. 2010). Hence, in contrast to W. dermatitidis and C. neoformans, in which chitin mutants secrete melanin, the C. albicans mutants instead fail to secrete melanin to the cell wall.
Fungal “melanosomes”
In mammalian cells, melanin is synthesized by specialized cells known as melanocytes. Melanin biosynthesis occurs in melanosomes, lysosome-related organelles. In this manner, the cells are protected from melanin and its intermediates, which are highly reactive and have a tendency to bind different substances. Melanosome formation goes through several morphological stages. In the early stages, a striated scaffold can be seen. Melanin is gradually deposited, creating the appearance of dark stripes in the melanosome. Eventually, dark, electron dense elliptical melanosomes form and are secreted to nearby keratinocytes where they have a protective role against ultraviolet radiation (Raposo and Marks 2002; Tolleson 2005).
Recent evidence supports the hypothesis that fungal melanosomes exist. Putative fungal melanosomes are best characterized in F. pedrosoi, the causative agent of chromoblastomycosis. TEM images of F. pedrosoi cells show internal organelles with fibrillar matrices that resemble mammalian melanosomes. These structures can be stimulated by treatment with bovine fetal serum, resulting in a gradual darkening of the melanosomes followed by fusion with the plasma membrane. The melanosomes are ultimately found in concentric rings in the cell wall (Franzen et al. 2008; Franzen et al. 1999). In contrast, treatment with the melanization inhibitor tricyclazole abolishes the electron-dense organelles (Franzen et al. 2006). Internal “melanosomes” are also observed in C. albicans (Walker et al. 2010), Cladosporium carrionii and Hormoconis resinae (San-Blas et al. 1996).
Studies in C. neoformans suggest that melanin may also be synthesized in vesicles in this fungus. Laccase activity is associated with extracellular vesicles secreted from C. neoformans (Rodrigues et al. 2008). When l-dopa is added to these vesicles, they melanize and increase in diameter. Interestingly, radiolabeled l-dopa associates with the vesicles, irrespective of laccase, suggesting that the lipid surface itself may be conducive to the formation of melanin. In addition, purified melanin from C. neoformans binds the lipophilic probe, DiI, suggesting there is lipid associated with the melanin (Eisenman et al. 2009). Knockdown of the secretory system of C. neoformans via RNAi of the SEC6 gene inhibits formation of melanin. Furthermore, laccase is improperly localized in these cells. It is found in intracellular vesicles, instead of the cell wall or culture supernatant (Panepinto et al. 2009). Together, these data strongly support the hypothesis that melanization in C. neoformans is vesicle-associated. Moreover, synthesis of fungal melanin in melanosomes or vesicles would shield the cell from the cytotoxic compounds produced during melanin polymerization and also account for the dimensions of the granular melanin particles described above. Figure 2 depicts a model of melanization in C. neoformans incorporating the results of these studies.
Fig. 2. Model depicting melanin formation in C. neoformans. Laccase containing vesicles are synthesized intracellularly and transported to the cell wall. Here, they interact with melanization substrate such as l-dopa to produce melanin granules. Cell wall polysaccharides such as chitin serve as a scaffold to which the melanin granules are cross-linked.

Melanin degradation
Melanin degradation is an area about which very little is known. However, understanding how melanin degradation occurs has practical relevance since melanin contributes to fungal virulence (for further discussion see Butler et al. (2005)). Melanin degradation has potential applications for human health. For instance, it may lead to improvements in melanoma treatment by breaking down melanin so tumors are responsive to radiation treatment. Melanin degradation is also an area of interest to the cosmetics industry for hair or skin bleaching purposes.
Experiments showing degradation of melanin by fungi are performed by growing the fungus on medium containing melanin. If degradation occurs, lightening or “bleaching” of the medium by the growing fungus can be observed. Phanerochaete chrysosporium and other white rot fungi can bleach melanin. These wood-decaying fungi contain lignin-degrading enzymes (peroxidases and laccases) that may also degrade melanin (Butler and Day 1998b). Other white rot fungal species including Bjerkandera adusta, Trametes versicolor and Trametes hirsute can degrade melanin produced by blue stain fungi, organisms that are responsible for discoloring wood (Ratto et al. 2001).
Melanin-bleaching enzymes are found in culture filtrates from some fungi such as Sporotrichum pruinosum (Mohorcic et al. 2007). For example, a partially purified lignin peroxidase from P. chrysosporium can degrade the synthetic substrate 2,2-azino-bis [3-ethylbenzthiazoline-6-sulfonic acid] (ABTS) in a hydrogen peroxide-dependent reaction (Woo et al. 2004). Similarly, a peroxidase capable of bleaching human hair melanin is found in Ceriporiopsis culture supernatants (Nagasaki et al. 2008).
Applied significance of fungal melanization
Melanin's unique properties suggest potential environmental and medical applications. An intriguing property of melanin is that it can shield organisms from ionizing radiation. Since melanin has a stable free radical population, it is thought that the radioprotective properties are due to scavenging of free radicals generated by radiation. Recent evidence suggests another mechanism of radioprotection, whereby damaging Compton recoil electrons are blocked and the energy dissipated in the melanin polymer (Schweitzer et al. 2009). Melanized fungi have been isolated from radioactively contaminated soils at the Nevada Test Site and areas around the damaged Chernobyl nuclear reactors. The ability of melanized fungi to survive in the presence of high levels of radiation, together with the large hyphal surface area of fungi in soils, has led to the suggestion that these organisms could be used for absorption and decontamination (Dighton et al. 2008). This hypothesis is supported by the discovery that melanized fungi not only survive high radiation levels but also have enhanced growth upon exposure. Experimental evidence suggests that fungal melanin can convert the energy of radiation to metabolically useful reducing power (Dadachova et al. 2007). Additional support for this notion comes from the observation that irradiation of a melanin electrode with gamma rays resulted in the formation of an electric current, thus indicating the potential for this interaction to result in energy transduction (Turick et al. 2011).
Medical treatments using radiation such as external beam radiation therapy for cancer patients or radioimmunotherapy can damage bone marrow resulting in debilitating side effects. In experimental mouse models, melanin can successfully shield bone marrow from such side effects. Melanin coated nanoparticles injected into mice lodge in the bone marrow. Treated mice have higher white blood cell and platelet counts compared to control mice after radiation treatment. Furthermore, the melanin-coated nanoparticles do not interfere with tumor treatment in these animals (Schweitzer et al. 2010). Thus, melanin's radioprotective effects have potential use in medicine.
Melanin is both hydrophobic and negatively charged, giving it the ability to bind to a broad spectrum of substances. Based on this, melanized fungi are proposed to be good candidates in bioremediation, since the organisms can potentially bind many toxic substances. This has been tested in laboratory and field experiments. A. niger can bind to sepia ink in water, thereby decolorizing it (Lamia and Neji 2010). Lichens isolated from an abandoned mine containing high levels of toxic heavy metals have uranium, iron and copper associated with melanized tissues of the lichen (Purvis et al. 2004). Similarly, in situ experiments with bacterial pyomelanin show uranium binding in contaminated soils (Turick et al. 2008). Together, these data demonstrate the potential of fungal melanin to bind to toxins in contaminated sites.
Melanin's binding capacity presents a problem in treatment of fungal infection (Nosanchuk and Casadevall 2006). Studies indicate that the melanized fungi are more resistant to antifungal therapy compared to non-melanized. When the PKS1 gene of W. dermatitidis is knocked out, cells do not produce melanin. In addition, they are more susceptible to voriconazole and amphotericin B (Paolo et al. 2006). Similarly, melanized cells of Histoplasma capsulatum and C. neoformans are more resistant to killing by caspofungin and amphotericin B (van Duin et al. 2002). Melanization can also enhance the drug resistance of C. neoformans growing in a biofilm (Martinez and Casadevall 2006).
The increased drug resistance of melanized fungi can be attributed to binding of the drugs to melanin. Addition of melanin to the medium of Madura mycetomatis, the causative agent of eumycetoma, increases the MICs of itraconazole and ketoconazole dramatically, suggesting that melanin blocks the drug (van de Sande et al. 2007). In other fungi, this phenomenon is observed with polyenes and echinocandins but not azoles. Incubation of such antifungals with melanin particles decreases activity against C. neoformans and H. capsulatum. Elemental analysis of the incubated melanins shows alteration in the melanin likely due to drug binding (van Duin et al. 2002). Melanin can extract amphotericin B, reducing the final concentration of drug in solution (Ikeda et al. 2003).
A question raised by these studies is whether inhibition of melanization would increase efficacy of antifungal drugs. Tentative evidence for this comes from the observation that administration of the melanization inhibitor glyphosate to mice infected with C. neoformans is sufficient to promote considerable clearance of infection (Nosanchuk et al. 2001). There are currently a limited number of drugs effective against fungal infections. Therefore, there is urgent need for improvements in antifungal therapy. Along similar lines, studies suggest that melanization should be considered when testing drug efficacy in vitro. In MIC assays, C. neoformans and H. capsulatum show susceptibility to the anti-fungal drug caspofungin. However, the drug is not effective in vivo. An explanation for this discrepancy is that melanization of the fungus in vivo inhibits drug activity (van Duin et al. 2002).
Conclusions
Despite the difficulties inherent in the study of melanin, it is clear that considerable progress has been made in recent years in understanding the synthesis, cell wall assembly, function and degradation of fungal melanins. Given the protean functions associated with melanin, it is likely that this pigment must possess tremendous structural complexity. The association of melanin with fungal virulence finds an echo in the difficulty in treatment of melanotic tumors such as melanoma, thus speaking to the protective properties conferred upon cells by this enigmatic pigment. Melanin research remains an area full of opportunities that remains of the great frontiers of biology.
Acknowledgments
This work was supported (in part) by a grant from The City University of New York PSC-CUNY Research Award Program to H. Eisenman. A. Casadevall is supported by the National Institutes of Health grants HL059842, AI033774, AI033142, and AI052733.
Contributor Information
Helene C. Eisenman, Email: helene.eisenman@baruch.cuny.edu, Department of Natural Sciences, Baruch College and Graduate Center, the City University of New York, 17 Lexington Avenue, Box A-506, New York, NY 10010, USA,.
Arturo Casadevall, Department of Microbiology and Immunology, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA.
References
- Almeida-Paes R, Frases S, Fialho Monteiro PC, Gutierrez-Galhardo MC, Zancope-Oliveira RM, Nosanchuk JD. Growth conditions influence melanization of Brazilian clinical Sporothrix schenckii isolates. Microbes Infect. 2009;11:554–562. doi: 10.1016/j.micinf.2009.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bailao AM, Schrank A, Borges CL, Dutra V, Molinari-Madlum EEWI, Felipe MSS, Mendes-Giannini MJS, Martins WS, Pereira M, de Almeida M, Soares C. Differential gene expression by Paracoccidioides brasiliensis in host interaction conditions: representational difference analysis identifies candidate genes associated with fungal pathogenesis. Microbes Infect. 2006;8:2686–2697. doi: 10.1016/j.micinf.2006.07.019. [DOI] [PubMed] [Google Scholar]
- Baker LG, Specht CA, Donlin MJ, Lodge JK. Chitosan, the deacetylated form of chitin, is necessary for cell wall integrity in Cryptococcus neoformans. Eukaryot Cell. 2007;6:855–867. doi: 10.1128/EC.00399-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banks IR, Specht CA, Donlin MJ, Gerik KJ, Levitz SM, Lodge JK. A chitin synthase and its regulator protein are critical for chitosan production and growth of the fungal pathogen Cryptococcus neoformans. Eukaryot Cell. 2005;4:1902–1912. doi: 10.1128/EC.4.11.1902-1912.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bocca AL, Brito PP, Figueiredo F, Tosta CE. Inhibition of nitric oxide production by macrophages in chromoblastomycosis: a role for Fonsecaea pedrosoi melanin. Mycopathologia. 2006;161:195–203. doi: 10.1007/s11046-005-0228-6. [DOI] [PubMed] [Google Scholar]
- Bull AT. Inhibition of polysaccharases by melanin: enzyme inhibition in relation to mycolysis. Arch Biochem Biophys. 1970;137:345–356. doi: 10.1016/0003-9861(70)90448-0. [DOI] [PubMed] [Google Scholar]
- Butler MJ, Day AW. Fungal melanins: a review. Can J Microbiol. 1998a;44:1115–1136. [Google Scholar]
- Butler MJ, Day AW. Destruction of fungal melanins by ligninases of Phanerochaete chrysosporium and other white rot fungi. Int J Plant Sci. 1998b;159:989–995. [Google Scholar]
- Butler MJ, Gardiner RB, Day AW. Degradation of melanin or inhibition of its synthesis: are these a significant approach as a biological control of phytopathogenic fungi? Biol Control. 2005;32:326–336. [Google Scholar]
- Caesar-Tonthat T, Van Ommen KF, Geesey GG, Henson JM. Melanin production by a filamentous soil fungus in response to copper and localization of copper sulfide by sulfide-silver staining. Appl Environ Microbiol. 1995;61:1968–1975. doi: 10.1128/aem.61.5.1968-1975.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chai LY, Netea MG, Sugui J, Vonk AG, van de Sande WW, Warris A, Kwon-Chung KJ, Kullberg BJ. Aspergillus fumigatus conidial melanin modulates host cytokine response. Immunobiology. 2010;215:915–920. doi: 10.1016/j.imbio.2009.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Z, Nunes MA, Silva MC, Rodrigues CJ., Jr Appressorium turgor pressure of Colletotrichum kahawae might have a role in coffee cuticle penetration. Mycologia. 2004;96:1199–1208. [PubMed] [Google Scholar]
- Clancy CM, Simon JD. Ultrastructural organization of eumelanin from Sepia officinalis measured by atomic force microscopy. Biochemistry. 2001;40:13353–13360. doi: 10.1021/bi010786t. [DOI] [PubMed] [Google Scholar]
- Cunha MM, Franzen AJ, Alviano DS, Zanardi E, Alviano CS, De Souza W, Rozental S. Inhibition of melanin synthesis pathway by tricyclazole increases susceptibility of Fonsecaea pedrosoi against mouse macrophages. Microsc Res Tech. 2005;68:377–384. doi: 10.1002/jemt.20260. [DOI] [PubMed] [Google Scholar]
- Cunha MM, Franzen AJ, Seabra SH, Herbst MH, Vugman NV, Borba LP, de Souza W, Rozental S. Melanin in Fonsecaea pedrosoi: a trap for oxidative radicals. BMC Microbiol. 2010;10:80. doi: 10.1186/1471-2180-10-80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva MB, Marques AF, Nosanchuk JD, Casadevall A, Travassos LR, Taborda CP. Melanin in the dimorphic fungal pathogen Paracoccidioides brasiliensis: effects on phagocytosis, intracellular resistance and drug susceptibility. Microbes Infect. 2006;8:197–205. doi: 10.1016/j.micinf.2005.06.018. [DOI] [PubMed] [Google Scholar]
- Dadachova E, Bryan RA, Huang X, Moadel T, Schweitzer AD, Aisen P, Nosanchuk JD, Casadevall A. Ionizing radiation changes the electronic properties of melanin and enhances the growth of melanized fungi. PLoS One. 2007;2:e457. doi: 10.1371/journal.pone.0000457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diaz P, Gimeno Y, Carro P, Gonzalez S, Schilardi PL, Benitez G, Salvarezza RC, Creus AH. Electrochemical self-assembly of melanin films on gold. Langmuir. 2005;21:5924–5930. doi: 10.1021/la0469755. [DOI] [PubMed] [Google Scholar]
- Dighton J, Tugay T, Zhdanova N. Fungi and ionizing radiation from radionuclides. FEMS Microbiol Lett. 2008;281:109–120. doi: 10.1111/j.1574-6968.2008.01076.x. [DOI] [PubMed] [Google Scholar]
- Eisenman HC, Frases S, Nicola AM, Rodrigues ML, Casadevall A. Vesicle-associated melanization in Cryptococcus neoformans. Microbiology. 2009;155:3860–3867. doi: 10.1099/mic.0.032854-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisenman HC, Mues M, Weber SE, Frases S, Chaskes S, Gerfen G, Casadevall A. Cryptococcus neoformans laccase catalyses melanin synthesis from both D- and l-DOPA. Microbiology. 2007;153:3954–3962. doi: 10.1099/mic.0.2007/011049-0. [DOI] [PubMed] [Google Scholar]
- Eisenman HC, Nosanchuk JD, Webber JB, Emerson RJ, Camesano TA, Casadevall A. Microstructure of cell wall-associated melanin in the human pathogenic fungus Cryptococcus neoformans. Biochemistry. 2005;44:3683–3693. doi: 10.1021/bi047731m. [DOI] [PubMed] [Google Scholar]
- Enochs WS, Nilges MJ, Swartz HM. A standardized test for the identification and characterization of melanins using electron paramagnetic resonance (EPR) spectroscopy. Pigment Cell Res. 1993;6:91–99. doi: 10.1111/j.1600-0749.1993.tb00587.x. [DOI] [PubMed] [Google Scholar]
- Fowler ZL, Baron CM, Panepinto JC, Koffas MA. Melanization of flavonoids by fungal and bacterial laccases. Yeast. 2011;28:181–188. doi: 10.1002/yea.1829. [DOI] [PubMed] [Google Scholar]
- Franzen AJ, Cunha MM, Batista EJ, Seabra SH, De Souza W, Rozental S. Effects of tricyclazole (5-methyl-1,2,4-triazol[3,4] benzothiazole), a specific DHN-melanin inhibitor, on the morphology of Fonsecaea pedrosoi conidia and sclerotic cells. Microsc Res Tech. 2006;69:729–737. doi: 10.1002/jemt.20344. [DOI] [PubMed] [Google Scholar]
- Franzen AJ, Cunha MM, Miranda K, Hentschel J, Plattner H, da Silva MB, Salgado CG, de Souza W, Rozental S. Ultrastructural characterization of melanosomes of the human pathogenic fungus Fonsecaea pedrosoi. J Struct Biol. 2008;162:75–84. doi: 10.1016/j.jsb.2007.11.004. [DOI] [PubMed] [Google Scholar]
- Franzen AJ, de Souza W, Farina M, Alviano CS, Rozental S. Morphometric and densitometric study of the biogenesis of electron-dense granules in Fonsecaea pedrosoi. FEMS Microbiol Lett. 1999;173:395–402. doi: 10.1111/j.1574-6968.1999.tb13531.x. [DOI] [PubMed] [Google Scholar]
- Frases S, Chaskes S, Dadachova E, Casadevall A. Induction by Klebsiella aerogenes of a melanin-like pigment in Cryptococcus neoformans. Appl Environ Microbiol. 2006;72:1542–1550. doi: 10.1128/AEM.72.2.1542-1550.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frases S, Salazar A, Dadachova E, Casadevall A. Cryptococcus neoformans can utilize the bacterial melanin precursor homogentisic acid for fungal melanogenesis. Appl Environ Microbiol. 2007;73:615–621. doi: 10.1128/AEM.01947-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gachomo EW, Seufferheld MJ, Kotchoni SO. Melanization of appressoria is critical for the pathogenicity of Diplocarpon rosae. Mol Biol Rep. 2010;37:3583–3591. doi: 10.1007/s11033-010-0007-4. [DOI] [PubMed] [Google Scholar]
- Garcia-Rivera J, Eisenman HC, Nosanchuk JD, Aisen P, Zaragoza O, Moadel T, Dadachova E, Casadevall A. Comparative analysis of Cryptococcus neoformans acid-resistant particles generated from pigmented cells grown in different laccase substrates. Fungal Genet Biol. 2005;42:989–998. doi: 10.1016/j.fgb.2005.09.003. [DOI] [PubMed] [Google Scholar]
- Griffith GW, Easton GL, Detheridge A, Roderick K, Edwards A, Worgan HJ, Nicholson J, Perkins WT. Copper deficiency in potato dextrose agar causes reduced pigmentation in cultures of various fungi. FEMS Microbiol Lett. 2007;276:165–171. doi: 10.1111/j.1574-6968.2007.00923.x. [DOI] [PubMed] [Google Scholar]
- Hegnauer H, Nyhlén LE, Rast DM. Ultrastructure of native and synthetic Agaricus bisporus melanins—implications as to the compartmentation of melanogenesis in fungi. Exp Mycol. 1985;9:1–29. [Google Scholar]
- Howard RJ, Valent B. Breaking and entering: host penetration by the fungal rice blast pathogen Magnaporthe grisea. Annu Rev Microbiol. 1996;50:491–512. doi: 10.1146/annurev.micro.50.1.491. [DOI] [PubMed] [Google Scholar]
- Ikeda R, Sugita T, Jacobson ES, Shinoda T. Effects of melanin upon susceptibility of Cryptococcus to antifungals. Microbiol Immunol. 2003;47:271–277. doi: 10.1111/j.1348-0421.2003.tb03395.x. [DOI] [PubMed] [Google Scholar]
- Ito S, Wakamatsu K. Quantitative analysis of eumelanin and pheomelanin in humans, mice, and other animals: a comparative review. Pigment Cell Res. 2003;16:523–531. doi: 10.1034/j.1600-0749.2003.00072.x. [DOI] [PubMed] [Google Scholar]
- Jacobson ES, Ikeda R. Effect of melanization upon porosity of the cryptococcal cell wall. Med Mycol. 2005;43:327–333. doi: 10.1080/13693780412331271081. [DOI] [PubMed] [Google Scholar]
- Jiang N, Sun N, Xiao D, Pan J, Wang Y, Zhu X. A copper-responsive factor gene CUF1 is required for copper induction of laccase in Cryptococcus neoformans. FEMS Microbiol Lett. 2009;296:84–90. doi: 10.1111/j.1574-6968.2009.01619.x. [DOI] [PubMed] [Google Scholar]
- Jorgensen TR, Park J, Arentshorst M, van Welzen AM, Lamers G, Vankuyk PA, Damveld RA, van den Hondel CA, Nielsen KF, Frisvad JC, Ram AF. The molecular and genetic basis of conidial pigmentation in Aspergillus niger. Fungal Genet Biol. 2011;48:544–553. doi: 10.1016/j.fgb.2011.01.005. [DOI] [PubMed] [Google Scholar]
- Kogej T, Stein M, Volkmann M, Gorbushina AA, Galinski EA, Gunde-Cimerman N. Osmotic adaptation of the halophilic fungus Hortaea werneckii: role of osmolytes and melanization. Microbiology. 2007;153:4261–4273. doi: 10.1099/mic.0.2007/010751-0. [DOI] [PubMed] [Google Scholar]
- Kroken S, Glass NL, Taylor JW, Yoder OC, Turgeon BG. Phylogenomic analysis of type I polyketide synthase genes in pathogenic and saprobic ascomycetes. Proc Natl Acad Sci U S A. 2003;100:15670–15675. doi: 10.1073/pnas.2532165100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamia K, Neji G. Aspergillus niger is able to decolourize sepia ink contained in saline industrial wastewaters. Desalin Water Treat. 2010;20:144–153. [Google Scholar]
- Land EJ, Ramsden CA, Riley PA. Quinone chemistry and melanogenesis. Methods Enzymol. 2004;378:88–109. doi: 10.1016/S0076-6879(04)78005-2. [DOI] [PubMed] [Google Scholar]
- Langfelder K, Streibel M, Jahn B, Haase G, Brakhage AA. Biosynthesis of fungal melanins and their importance for human pathogenic fungi. Fungal Genet Biol. 2003;38:143–158. doi: 10.1016/s1087-1845(02)00526-1. [DOI] [PubMed] [Google Scholar]
- Latge JP. The cell wall: a carbohydrate armour for the fungal cell. Mol Microbiol. 2007;66:279–290. doi: 10.1111/j.1365-2958.2007.05872.x. [DOI] [PubMed] [Google Scholar]
- Latge JP, Mouyna I, Tekaia F, Beauvais A, Debeaupuis JP, Nierman W. Specific molecular features in the organization and biosynthesis of the cell wall of Aspergillus fumigatus. Med Mycol. 2005;43(Suppl 1):S15–S22. doi: 10.1080/13693780400029155. [DOI] [PubMed] [Google Scholar]
- Madrid IM, Xavier MO, Mattei AS, Fernandes CG, Guim TN, Santin R, Schuch LF, Nobre Mde O, Meireles MCA. Role of melanin in the pathogenesis of cutaneous sporotrichosis. Microbes Infect. 2010;12:162–165. doi: 10.1016/j.micinf.2009.10.004. [DOI] [PubMed] [Google Scholar]
- Martinez LR, Casadevall A. Susceptibility of Cryptococcus neoformans biofilms to antifungal agents in vitro. Antimicrob Agents Chemother. 2006;50:1021–1033. doi: 10.1128/AAC.50.3.1021-1033.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mednick AJ, Nosanchuk JD, Casadevall A. Melanization of Cryptococcus neoformans affects lung inflammatory responses during cryptococcal infection. Infect Immun. 2005;73:2012–2019. doi: 10.1128/IAI.73.4.2012-2019.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meng S, Kaxiras E. Theoretical models of eumelanin protomolecules and their optical properties. Biophys J. 2008;94:2095–2105. doi: 10.1529/biophysj.107.121087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Missall TA, Moran JM, Corbett JA, Lodge JK. Distinct stress responses of two functional laccases in Cryptococcus neoformans are revealed in the absence of the thiol-specific antioxidant Tsa1. Eukaryot Cell. 2005;4:202–208. doi: 10.1128/EC.4.1.202-208.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohorcic M, Friedrich J, Renimel I, André P, Mandin D, Chaumont JP. Production of melanin bleaching enzyme of fungal origin and its application in cosmetics. Biotechnol Bioprocess Eng. 2007;12:200–206. [Google Scholar]
- Morris-Jones R, Gomez BL, Diez S, Uran M, Morris-Jones SD, Casadevall A, Nosanchuk JD, Hamilton AJ. Synthesis of melanin pigment by Candida albicans in vitro and during infection. Infect Immun. 2005;73:6147–6150. doi: 10.1128/IAI.73.9.6147-6150.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munro CA, Gow NA. Chitin synthesis in human pathogenic fungi. Med Mycol. 2001;39(Suppl 1):41–53. [PubMed] [Google Scholar]
- Nagasaki K, Kumazawa M, Murakami S, Takenaka S, Koike K, Aoki K. Purification, characterization, and gene cloning of Ceriporiopsis sp. strain MD-1 peroxidases that decolorize human hair melanin. Appl Environ Microbiol. 2008;74:5106–5112. doi: 10.1128/AEM.00253-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ngamskulrungroj P, Price J, Sorrell T, Perfect JR, Meyer W. Cryptococcus gattii virulence composite: candidate genes revealed by microarray analysis of high and less virulent Vancouver island outbreak strains. PLoS One. 2011;6:e16076. doi: 10.1371/journal.pone.0016076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nosanchuk JD, Casadevall A. Budding of melanized Cryptococcus neoformans in the presence or absence of l-dopa. Microbiology. 2003a;149:1945–1951. doi: 10.1099/mic.0.26333-0. [DOI] [PubMed] [Google Scholar]
- Nosanchuk JD, Casadevall A. The contribution of melanin to microbial pathogenesis. Cell Microbiol. 2003b;5:203–223. doi: 10.1046/j.1462-5814.2003.00268.x. [DOI] [PubMed] [Google Scholar]
- Nosanchuk JD, Casadevall A. Impact of melanin on microbial virulence and clinical resistance to antimicrobial compounds. Antimicrob Agents Chemother. 2006;50:3519–3528. doi: 10.1128/AAC.00545-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nosanchuk JD, Ovalle R, Casadevall A. Glyphosate inhibits melanization of Cryptococcus neoformans and prolongs survival of mice after systemic infection. J Infect Dis. 2001;183:1093–1099. doi: 10.1086/319272. [DOI] [PubMed] [Google Scholar]
- Noverr MC, Williamson PR, Fajardo RS, Huffnagle GB. CNLAC1 is required for extrapulmonary dissemination of Cryptococcus neoformans but not pulmonary persistence. Infect Immun. 2004;72:1693–1699. doi: 10.1128/IAI.72.3.1693-1699.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panepinto J, Komperda K, Frases S, Park YD, Djordjevic JT, Casadevall A, Williamson PR. Sec6-dependent sorting of fungal extracellular exosomes and laccase of Cryptococcus neoformans. Mol Microbiol. 2009;71:1165–1176. doi: 10.1111/j.1365-2958.2008.06588.x. [DOI] [PubMed] [Google Scholar]
- Paolo WF, Jr, Dadachova E, Mandal P, Casadevall A, Szaniszlo PJ, Nosanchuk JD. Effects of disrupting the polyketide synthase gene WdPKS1 in Wangiella [Exophiala] dermatitidis on melanin production and resistance to killing by antifungal compounds, enzymatic degradation, and extremes in temperature. BMC Microbiol. 2006;6:55. doi: 10.1186/1471-2180-6-55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pihet M, Vandeputte P, Tronchin G, Renier G, Saulnier P, Georgeault S, Mallet R, Chabasse D, Symoens F, Bouchara JP. Melanin is an essential component for the integrity of the cell wall of Aspergillus fumigatus conidia. BMC Microbiol. 2009;9:177. doi: 10.1186/1471-2180-9-177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pukkila-Worley R, Gerrald QD, Kraus PR, Boily MJ, Davis MJ, Giles SS, Cox GM, Heitman J, Alspaugh JA. Transcriptional network of multiple capsule and melanin genes governed by the Cryptococcus neoformans cyclic AMP cascade. Eukaryot Cell. 2005;4:190–201. doi: 10.1128/EC.4.1.190-201.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purvis OW, Bailey E, McLean J, Kasama T, Williamson BJ. Uranium biosorption by the lichen Trapelia involuta at a uranium mine. Geomicrobiol J. 2004;21:159–167. [Google Scholar]
- Raparia K, Powell SZ, Cernoch P, Takei H. Cerebral mycosis: 7-year retrospective series in a tertiary center. Neuropathology. 2010;30:218–223. doi: 10.1111/j.1440-1789.2009.01067.x. [DOI] [PubMed] [Google Scholar]
- Raposo G, Marks MS. The dark side of lysosome-related organelles: specialization of the endocytic pathway for melanosome biogenesis. Traffic. 2002;3:237–248. doi: 10.1034/j.1600-0854.2002.030401.x. [DOI] [PubMed] [Google Scholar]
- Ratto M, Chatani M, Ritschkoff AC, Viikari L. Screening of micro-organisms for decolorization of melanins produced by bluestain fungi. Appl Microbiol Biotechnol. 2001;55:210–213. doi: 10.1007/s002530000516. [DOI] [PubMed] [Google Scholar]
- Riley PA. Melanin. Int J Biochem Cell Biol. 1997;29:1235–1239. doi: 10.1016/s1357-2725(97)00013-7. [DOI] [PubMed] [Google Scholar]
- Rodrigues ML, Nakayasu ES, Oliveira DL, Nimrichter L, Nosanchuk JD, Almeida IC, Casadevall A. Extracellular vesicles produced by Cryptococcus neoformans contain protein components associated with virulence. Eukaryot Cell. 2008;7:58–67. doi: 10.1128/EC.00370-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosa LH, Vieira LMA, Santiago IF, Rosa CA. Endophytic fungi community associated with the dicotyledonous plant Colobanthus quitensis (Kunth) Bartl. (Caryophyllaceae) in Antarctica. FEMS Microbiol Ecol. 2010;73:178–189. doi: 10.1111/j.1574-6941.2010.00872.x. [DOI] [PubMed] [Google Scholar]
- Rosas AL, MacGill RS, Nosanchuk JD, Kozel TR, Casadevall A. Activation of the alternative complement pathway by fungal melanins. Clin Diagn Lab Immunol. 2002;9:144–148. doi: 10.1128/CDLI.9.1.144-148.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruiz-Herrera J, Elorza MV, Valentin E, Sentandreu R. Molecular organization of the cell wall of Candida albicans and its relation to pathogenicity. FEMS Yeast Res. 2006;6:14–29. doi: 10.1111/j.1567-1364.2005.00017.x. [DOI] [PubMed] [Google Scholar]
- Saitoh Y, Izumitsu K, Morita A, Shimizu K, Tanaka C. ChMCO1 of Cochliobolus heterostrophus is a new class of metallo-oxidase, playing an important role in DHN-melanization. Mycoscience. 2010a;51:327–336. [Google Scholar]
- Saitoh Y, Izumitsu K, Morita A, Tanaka C. A copper-transporting ATPase BcCCC2 is necessary for pathogenicity of Botrytis cinerea. Mol Gen Genomics. 2010b;284:33–43. doi: 10.1007/s00438-010-0545-4. [DOI] [PubMed] [Google Scholar]
- Salas SD, Bennett JE, Kwon-Chung KJ, Perfect JR, Williamson PR. Effect of the laccase gene CNLAC1, on virulence of Cryptococcus neoformans. J Exp Med. 1996;184:377–386. doi: 10.1084/jem.184.2.377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- San-Blas G, Guanipa O, Moreno B, Pekerar S, San-Blas F. Cladosporium carrionii and Hormoconis resinae (C. resinae): cell wall and melanin studies. Curr Microbiol. 1996;32:11–16. doi: 10.1007/s002849900003. [DOI] [PubMed] [Google Scholar]
- Schumann J, Hertweck C. Advances in cloning, functional analysis and heterologous expression of fungal polyketide synthase genes. J Biotechnol. 2006;124:690–703. doi: 10.1016/j.jbiotec.2006.03.046. [DOI] [PubMed] [Google Scholar]
- Schweitzer AD, Howell RC, Jiang Z, Bryan RA, Gerfen G, Chen CC, Mah D, Cahill S, Casadevall A, Dadachova E. Physicochemical evaluation of rationally designed melanins as novel nature-inspired radioprotectors. PLoS One. 2009;4:e7229. doi: 10.1371/journal.pone.0007229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schweitzer AD, Revskaya E, Chu P, Pazo V, Friedman M, Nosanchuk JD, Cahill S, Frases S, Casadevall A, Dadachova E. Melanin-covered nanoparticles for protection of bone marrow during radiation therapy of cancer. Int J Radiat Oncol Biol Phys. 2010;78:1494–1502. doi: 10.1016/j.ijrobp.2010.02.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silva MB, Thomaz L, Marques AF, Svidzinski AE, Nosanchuk JD, Casadevall A, Travassos LR, Taborda CP. Resistance of melanized yeast cells of Paracoccidioides brasiliensis to antimicrobial oxidants and inhibition of phagocytosis using carbohydrates and monoclonal antibody to CD18. Mem Inst Oswaldo Cruz. 2009;104:644–648. doi: 10.1590/s0074-02762009000400019. [DOI] [PubMed] [Google Scholar]
- Tolleson WH. Human melanocyte biology, toxicology, and pathology. J Environ Sci Heal C Environ Carcinog Ecotoxicol Rev. 2005;23:105–161. doi: 10.1080/10590500500234970. [DOI] [PubMed] [Google Scholar]
- Tsai HF, Wheeler MH, Chang YC, Kwon-Chung KJ. A developmentally regulated gene cluster involved in conidial pigment biosynthesis in Aspergillus fumigatus. J Bacteriol. 1999;181:6469–6477. doi: 10.1128/jb.181.20.6469-6477.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turick CE, Ekechukwu AA, Milliken CE, Casadevall A, Dadachova E. Gamma radiation interacts with melanin to alter its oxidation-reduction potential and results in electric current production. Bioelectrochemistry. 2011;82:69–73. doi: 10.1016/j.bioelechem.2011.04.009. [DOI] [PubMed] [Google Scholar]
- Turick CE, Knox AS, Leverette CL, Kritzas YG. In situ uranium stabilization by microbial metabolites. J Environ Radioact. 2008;99:890–899. doi: 10.1016/j.jenvrad.2007.11.020. [DOI] [PubMed] [Google Scholar]
- van de Sande WW, de Kat J, Coppens J, Ahmed AO, Fahal A, Verbrugh H, van Belkum A. Melanin biosynthesis in Madurella mycetomatis and its effect on susceptibility to itraconazole and ketoconazole. Microbes Infect. 2007;9:1114–1123. doi: 10.1016/j.micinf.2007.05.015. [DOI] [PubMed] [Google Scholar]
- van Duin D, Casadevall A, Nosanchuk JD. Melanization of Cryptococcus neoformans and Histoplasma capsulatum reduces their susceptibilities to amphotericin B and caspofungin. Antimicrob Agents Chemother. 2002;46:3394–3400. doi: 10.1128/AAC.46.11.3394-3400.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vidotto V, Aoki S, Ponton J, Quindos G, Koga-Ito CY, Pugliese A. A new caffeic acid minimal synthetic medium for the rapid identification of Cryptococcus neoformans isolates. Rev Iberoam Micol. 2004;21:87–89. [PubMed] [Google Scholar]
- Volling K, Thywissen A, Brakhage AA, Saluz HP. Phagocytosis of melanized Aspergillus conidia by macrophages exerts cytoprotective effects by sustained PI3K/Akt signalling. Cell Microbiol. 2011;13:1130–1148. doi: 10.1111/j.1462-5822.2011.01605.x. [DOI] [PubMed] [Google Scholar]
- Wakamatsu K, Ito S. Advanced chemical methods in melanin determination. Pigment Cell Res. 2002;15:174–183. doi: 10.1034/j.1600-0749.2002.02017.x. [DOI] [PubMed] [Google Scholar]
- Walker CA, Gomez BL, Mora-Montes HM, Mackenzie KS, Munro CA, Brown AJ, Gow NA, Kibbler CC, Odds FC. Melanin externalization in Candida albicans depends on cell wall chitin structures. Eukaryot Cell. 2010;9:1329–1342. doi: 10.1128/EC.00051-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walton FJ, Idnurm A, Heitman J. Novel gene functions required for melanization of the human pathogen Cryptococcus neoformans. Mol Microbiol. 2005;57:1381–1396. doi: 10.1111/j.1365-2958.2005.04779.x. [DOI] [PubMed] [Google Scholar]
- Wang Y, Aisen P, Casadevall A. Cryptococcus neoformans melanin and virulence: mechanism of action. Infect Immun. 1995;63:3131–3136. doi: 10.1128/iai.63.8.3131-3136.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Zheng L, Hauser M, Becker JM, Szaniszlo PJ. WdChs4p, a homolog of chitin synthase 3 in Saccharomyces cerevisiae, alone cannot support growth of Wangiella (Exophiala) dermatitidis at the temperature of infection. Infect Immun. 1999;67:6619–6630. doi: 10.1128/iai.67.12.6619-6630.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williamson PR. Laccase and melanin in the pathogenesis of Cryptococcus neoformans. Front Biosci. 1997;2:e99–e107. doi: 10.2741/a231. [DOI] [PubMed] [Google Scholar]
- Woo PC, Tam EW, Chong KT, Cai JJ, Tung ET, Ngan AH, Lau SK, Yuen KY. High diversity of polyketide synthase genes and the melanin biosynthesis gene cluster in Penicillium marneffei. FEBS J. 2010;277:3750–3758. doi: 10.1111/j.1742-4658.2010.07776.x. [DOI] [PubMed] [Google Scholar]
- Woo SH, Suk Cho JS, Seok Lee BS, Kim EK. Decolorization of melanin by lignin peroxidase from Phanerochaete chrysosporium. Biotechnol Bioprocess Eng. 2004;9:256–260. [Google Scholar]
- Zalar P, Novak M, de Hoog GS, Gunde-Cimerman N. Dishwashers—a man-made ecological niche accomodating human opportunistic fungal pathogens. Fungal Biol. 2011;115:997–1007. doi: 10.1016/j.funbio.2011.04.007. [DOI] [PubMed] [Google Scholar]
- Zhdanova NN, Zakharchenko VA, Vember VA, Nakonechnaya LT. Fungi from Chernobyl: mycobiota of the inner regions of the containment structures of the damaged nuclear reactor. Mycology Research. 2000;104:1421–1426. [Google Scholar]
- Zhong J, Frases S, Wang H, Casadevall A, Stark RE. Following fungal melanin biosynthesis with solid-state NMR: biopolymer molecular structures and possible connections to cell-wall polysaccharides. Biochemistry. 2008;47:4701–4710. doi: 10.1021/bi702093r. [DOI] [PubMed] [Google Scholar]
- Zhu X, Gibbons J, Garcia-Rivera J, Casadevall A, Williamson PR. Laccase of Cryptococcus neoformans is a cell wall-associated virulence factor. Infect Immun. 2001;69:5589–5596. doi: 10.1128/IAI.69.9.5589-5596.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X, Williamson PR. A CLC-type chloride channel gene is required for laccase activity and virulence in Cryptococcus neoformans. Mol Microbiol. 2003;50:1271–1281. doi: 10.1046/j.1365-2958.2003.03752.x. [DOI] [PubMed] [Google Scholar]