Abstract
Purpose
Mammaglobin-A (MAM-A) is overexpressed in 40–80% of primary breast cancers. We initiated a phase 1 clinical trial of a MAM-A DNA vaccine to evaluate its safety and biological efficacy.
Experimental Design
Breast cancer patients with stable metastatic disease were eligible for enrollment. Safety was monitored with clinical and laboratory assessments. The CD8 T cell response was measured by ELISPOT, flow cytometry, and cytotoxicity assays. Progression-free survival was described using the Kaplan-Meier product limit estimator.
Results
Fourteen subjects have been treated with the MAM-A DNA vaccine and no significant adverse events have been observed. Eight of fourteen subjects were HLA-A2+, and the CD8 T cell response to vaccination was studied in detail. Flow cytometry demonstrated a significant increase in the frequency of MAM-A-specific CD8 T cells following vaccination (0.9 ± 0.5% vs. 3.8 ± 1.2%, p < 0.001), and ELISPOT analysis demonstrated an increase in the number of MAM-A-specific IFN-γ-secreting T cells (41 ± 32 vs. 215 ± 67 spm, p < 0.001). Although this study was not powered to evaluate progression-free survival, preliminary evidence suggests that subjects treated with the MAM-A DNA vaccine had improved progression-free survival compared to subjects who met all eligibility criteria, were enrolled in the trial, but were not vaccinated because of HLA phenotype.
Conclusion
The MAM-A DNA vaccine is safe, capable of eliciting MAM-A-specific CD8 T cell responses, and preliminary evidence suggests improved progression-free survival. Additional studies are required to define the potential of the MAM-A DNA vaccine for breast cancer prevention and/or therapy.
Keywords: Mammaglobin-A, breast cancer, DNA vaccine, clinical trial, CD8 T cells
INTRODUCTION
Progress in basic and translational immunology has confirmed the importance of the immune system in cancer prevention, and has renewed interest in vaccine therapy for cancer (1). DNA vaccines are safe, well tolerated, and can typically be given in an ambulatory facility (2). Although breast cancer is commonly thought to be less immunogenic than melanoma or renal cell cancer, there is increasing evidence of a crosstalk between the immune system and breast cancer, and this crosstalk strongly suggests that successful development of a breast cancer vaccine could have a clinical impact. Evidence of this crosstalk includes the clinical significance of immune infiltrates in breast cancer (3), clear evidence of pre-existing immune responses to several breast cancer antigens including HER2/neu (4–6), MUC1 (7), and MAM-A (8–10), the increased prevalence of regulatory T cells in breast cancer patients (11), and upregulation of inhibitory molecules of the CD28 receptor family on breast cancer-specific T cells (12). Most breast cancer patients are diagnosed with local-regional disease, and typically have no evidence of disease following standard treatment modalities, providing a window-of-opportunity to generate effective antitumor immune responses to prevent recurrent disease (13). Peoples et al. recently confirmed the potential of breast cancer vaccine therapy in this clinical context, demonstrating that administration of a HER2/neu peptide vaccine was associated with a survival advantage in a prospective study of node-positive breast cancer patients with no evidence of disease (14). Taken together, the dynamic interaction between breast cancer and the immune system, and preliminary evidence of the efficacy of first-generation breast cancer vaccines provide strong rationale for the clinical evaluation of breast cancer vaccine strategies.
MGBA was first identified using a differential screening approach directed at the isolation of novel human breast cancer-associated genes (15). MGBA encodes MAM-A, a 10 kD glycoprotein that is related to a family of epithelial secretory proteins. Of note, MAM-A has several unique properties which make it an exceptional target for breast cancer vaccine therapy. First, MAM-A is expressed almost exclusively in breast cancer (16–19). Second, MAM-A is overexpressed in 40–80% of primary breast cancers (20–24). Overexpression of MAM-A in a significant percentage of breast cancer suggests that many breast cancer patients are likely to be candidates for vaccine therapy, and is particularly relevant for the development of vaccine strategies for the prevention of breast cancer. Third, MAM-A overexpression is evident in noninvasive, invasive, and metastatic breast cancer (25). This consistency of expression in breast cancers confirms that MAM-A is an attractive target for vaccine therapy. Finally, we have demonstrated that MAM-A is capable of eliciting an immune response in breast cancer patients (8, 26–28), and that a DNA vaccine targeting MAM-A is capable of successfully generating breast cancer immunity in preclinical models (10).
The observation that direct administration of recombinant DNA can generate potent immune responses in rodents established the field of DNA vaccines in the early 1990s (29). Since that time, DNA vaccines have remained an area of intense research interest, and vaccines targeting infectious disease and cancer have progressed into clinical trials. DNA vaccination offers several potential advantages. First, the presence of the full-length cDNA provides multiple potential epitopes, thus avoiding the need for patient selection based on MHC restriction. Second, bacterial plasmid DNA contains immunostimulatory unmethylated CpG motifs that may act as potent immune adjuvants (30, 31). Finally, DNA vaccines are relatively easy to prepare with high purity and high stability relative to proteins and other biologic agents, facilitating clinical translation of this platform, particularly in early phase clinical trials. In clinical trials of infectious disease and cancer, DNA vaccine strategies have been shown to be safe and effective in developing immune responses to malaria (32), HIV (33) and prostate cancer (34). Several reviews have recently been published summarizing progress in the field (2, 35, 36).
To explore the potential of a novel MAM-A DNA vaccine, we initiated an open-label phase 1 clinical trial in patients with metastatic breast cancer. Patients with stable metastatic disease were eligible for enrollment. Subjects were treated with three intramuscular injections of 4 mg plasmid at one month intervals using an FDA-approved carbon dioxide-powered jet delivery device. Safety was closely monitored with eight or more clinical and laboratory assessments in the first 24 weeks of the trial. The immune response to the vaccine was measured by ELISPOT analysis, multi-parameter flow cytometry, and cytotoxicity assays. Evidence of biological efficacy was assessed by measuring progression-free survival.
MATERIALS AND METHODS
Study Rationale and Objectives
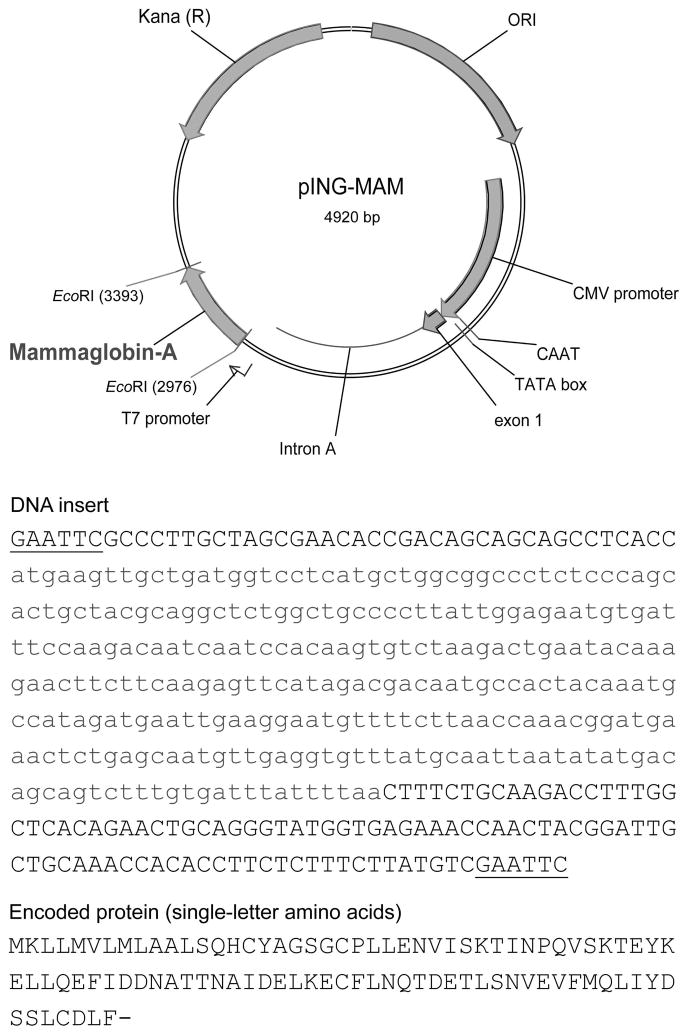
We recently initiated a single-institution, open-label, phase 1 clinical trial of a novel MAM-A DNA vaccine in breast cancer patients with stable metastatic disease (ClinicalTrials.gov identifier: NCT00807781). The primary objective of the trial was to evaluate the safety of the MAM-A DNA vaccine. Secondary and exploratory objectives included measuring the CD8 T cell response to the vaccine, and the impact on progression-free survival. The MAM-A DNA vaccine is composed of a closed circular DNA plasmid based on the pING parental vector, and is designed to express the human MAM-A breast cancer-associated antigen under a strong viral promoter (Figure 1).
Figure 1. The MAM-A DNA vaccine is a plasmid DNA vaccine designed to drive MAM-A expression with a strong viral promoter.
The pING parental vector contains the following elements: (1) a eukaryotic promoter and enhancer from the Towne strain of CMV; (2) a polylinker region to facilitate cloning; (3) donor and acceptor splice sites and a poly adenylation signal sequence derived from the bovine growth hormone gene; (4) the ColE1 origin of replication and (5) a gene conferring kanamycin resistance.
Study Participants
The phase 1 clinical trial was approved by the Siteman Cancer Center Protocol Review and Monitoring Committee and the Washington University School of Medicine Human Studies Committee. A written informed consent was obtained from all subjects prior to enrollment/participation in the study. Men or women 18 years or older with metastatic breast cancer were eligible for enrollment. Eligible subjects had metastatic breast cancer that had been stable for at least 30 days after the last dose of chemotherapy, or that had been stable for at least 30 days on endocrine therapy, documented adequate organ and marrow function, ECOG status of less than or equal to 2, and negative urine or serum β-HCG pregnancy test for women of reproductive age. Subjects were considered ineligible if they had received an investigational drug within 30 days of enrollment, had known brain metastases, known allergy, or history of serious adverse reaction to vaccines, autoimmune disease requiring management with immunosuppression, or history of failing greater than two chemotherapy regimens for metastatic disease. Following enrollment, the subjects’ primary breast cancers were evaluated by immunohistochemistry for MAM-A expression (only subjects with MAM-A+ cancers were eligible for vaccination), and their HLA phenotype was determined (initially, only subjects with HLA-A2 and/or HLA-A3 were eligible for vaccination to facilitate immune monitoring as immunodominant epitopes have been identified for these alleles (8–10)). Screen failures were not vaccinated, but were followed for progression of disease. Subjects who were screen failures based on HLA phenotype, withdrawal of consent and inability to perform MAM-A IHC staining were considered to be appropriate for inclusion in a comparator group.
Vaccination Schedule
Study subjects were vaccinated with intramuscular injections of 4 mg plasmid DNA vaccine at day 1 (Week 1), day 29 ± 7 (Week 4), and day 57 ± 7 (Week 8) with at least 21 days between vaccinations. All vaccinations were given intramuscularly using an FDA-approved carbon dioxide-powered jet delivery device (Needle Free Biojector 2000).
Safety monitoring and study procedures
Safety was closely monitored after vaccination with eight or more clinical and laboratory assessments in the first 24 weeks of the trial. After each vaccination, study subjects were carefully observed for 60 minutes. Vital signs were monitored and adverse events were recorded. The injection site was inspected for evidence of a local reaction. Subjects were given a “Diary Card” where they recorded temperature and symptoms daily for 5 days. Laboratory tests performed on the day of vaccination included: complete blood count, comprehensive metabolic panel, and serum or urine pregnancy test (in women of childbearing potential). Patients were followed actively for 52 weeks following vaccination, and then until disease progression or death, whichever occurred first. Toxicity was graded according to the National Cancer Institute Common Terminology Criteria for Adverse Events version 3.0. Subjects had standard-of-care clinical assessments and imaging studies (CT scans) during follow-up in order to assess tumor stability.
Peripheral blood specimens were obtained for immune monitoring and other correlative studies before vaccination and at serial time points following vaccination. Peripheral blood mononuclear cells (PBMC) were isolated from heparinized blood by Ficoll–Hypaque density gradient centrifugation (Pharmacia, Uppsala, Sweden) and stored at −135 °C until evaluation. The CD8 T cells were isolated from PBMC by positive selection using a MACS bead isolation kit (Miltenyi Biotec Inc., CA) (37).
Breast cancer cell lines and cell culture
The breast cancer cell lines, UACC-812 (MAM-A+/HLA-A2+), MCF-7 (MAM-A−/HLA-A2+), MDA-MB-415 (MAM-A+/HLA-A2−), MDA-MB-134 (MAM-A−/HLA-A2−), were obtained from the American Type Culture Collection (ATCC, Manassas, VA). The cell lines were ordered in 2004 by Dr Mohanakumar laboratory. No specific authentication of the cell lines was performed in our laboratory. All experiments were performed on the cell lines below 30th passage. Breast cancer cell lines were cultured in RPMI-1640 culture medium at 37 °C in 5% CO2 incubator until they were 80% confluent. The presence of MAM-A in the cell lines was confirmed by reverse transcriptase-polymerase chain reaction (Supplementary Figure 1) and western blot analysis (data not shown). siRNA oligonucleotides and monoclonal antibodies used in the cell culture inhibition studies were obtained from Santa Cruz Biotech (Santa Cruz, CA).
ELISPOT assay
PBMC were cultured overnight in complete RPMI-1640 and viability was determined by trypan blue exclusion. PBMC with viability of at least 90% were used for ELISPOT analysis (10, 38). CD8 T cells were enriched by MACS bead negative selection using immunomagnetic separation cocktails (Stem cell Technologies, Canada). Purified CD8 T cells (2 × 105 cells, >90% purity) were cultured in triplicate with MAM-A2.1 (LIYDSSLCDL, 20 μg/mL) in 96-well ELISPOT plates (Multiscreen IP plate, Millipore, MA) pre-coated with IFN-γ mAb (4 μg/mL) in the presence of autologous-irradiated T cell-depleted PBMCs (3 × 104) for 48–72 h in humidified 5% CO2 at 37 °C. Subsequently, the plates were washed and developed to detect the number of spots in individual wells using an ImmunoSpot analyzer (Cellular Technology, Shaker Heights, OH). CD8 T cells plus autologous irradiated T cell-depleted PBMCs cultured in medium without MAM-A2.1 peptide were used as a negative control, while CD8 T cells plus autologous irradiated T cell-depleted PBMCs cultured with PHA (5 μg/mL) were used as a positive control. The number of spots in control wells was subtracted from the number of spots in experimental wells and reported in the final results as spots per million cells (spm ± SEM).
Flow cytometry
We previously demonstrated that LIYDSSLCDL, designated MAM-A2.1, is an immunodominant HLA-A2-restricted epitope (10, 38). MAM-A2.1 tetramers were developed by Beckman Coulter Immunomics (San Diego, CA) to monitor the MAM-A-specific CD8 T cell response following MAM-A DNA vaccination. An HLA-A2 tetramer incorporating an unrelated peptide from influenza (Flu), GILGFVFTL, was also prepared and used as a control. Tetramers were used to stain target cells at a concentration of 10 μL per 200 μL final volume of CD8 T cells (1 × 106 CD8 T cells/mL). Antibodies used for flow cytometry included CD8-FITC (BD Biosciences, San Jose, CA), MAM-A2.1/Tetramer-PE, and Flu-peptide/Tetramer-PE. Samples were analyzed using a FACS Calibur/LSRII flow cytometer (Becton-Dickinson, Franklin Lakes, NJ), and cell sorting was performed using a Vantage cell sorter (Becton-Dickinson). Data were analyzed using BD FACSDiva software. Gates were set according to isotype controls.
Cytotoxicity assay
The ability of MAM-A-specific CD8 T cells to lyse breast cancer cell lines was determined using an LDH cytotoxicity assay (Promega, Madison, WI). Breast cancer cells (5 × 103 cells) in 100 μL of complete medium were plated in triplicate cultures in round bottom 96-well plates in the presence of varying numbers of CD8 T cells (E:T ratios of 6.25:1 to 50:1) and incubated at 37 °C in a humidified 5% CO2 incubator for 4 h. Controls were breast cancer target cells alone or CD8 T cells isolated from normal subjects. Maximum release was determined by adding Triton X-100 (1%) to the target cells. A colorimetric measurement of the released LDH was developed as per manufacturer’s instructions and measured by spectrophotometer at 450 nm. The percent-specific lysis was calculated using the formula [(experimental LDH release − spontaneous LDH release)/(maximum LDH release − spontaneous LDH release)] × 100.
Western blot analysis
Total proteins were extracted from cells with lysis buffer (50 mM HEPES [pH 7.6], 150 mM NaCl, 1% Triton X-100, 30 mM Na4P2O7, 10% glycerol, 1 mM benzamidine, 1 mM DTT, 10 μg of leupeptin/mL, 1 mM phenylmethylsulfonyl fluoride, 50 mM NaF, 1 mM sodium orthovanadate, 10 mM sodium pyrophosphate decahydrate, and 10 mM β-glycerophosphate (Sigma Aldrich). After cell lysis, the supernatant was collected after centrifugation at 15,000 × g for 15 min at 4 °C (39). Protein concentration was determined with a Bradford’s assay kit from Bio-Rad (Munich, Germany). Total proteins were separated on a 4–12% sodium dodecyl sulfate-polyacrylamide gradient gel and transferred onto a nitrocellulose membrane. The membranes were blocked overnight at 4 °C in Tris-buffered saline with 0.05% Tween 20 (5% nonfat milk in 10 mM Tris-HCl, 100 mM NaCl, 0. 1% Tween 20, pH 7.6). The membranes were incubated first with primary antibodies diluted 1 in 1,000 in blocking buffer at room temperature for 2 h, and then with a horseradish peroxide-conjugated secondary IgG mAb diluted 1 in 5,000 for 1 h. All primary and secondary Ab were obtained from Santa Cruz Biotech. The membrane was developed using a chemiluminescence kit (Millipore) and analyzed using Bio-Rad Universal Hood II (Hercules, CA). Densitometric analysis was done using the software provided by Bio-Rad.
Gene expression analysis
Relative expression of HLA-A2, MAM-A, NKG2D, DAP-10, perforin and actin were analyzed using FAM-labeled RT-PCR primers (Applied Biosystems, Foster City, CA) as per the manufacturer’s recommendations (26). Briefly, total RNA was extracted from 1 × 106 cells using TRIzol reagent (Sigma–Aldrich). RNA samples were quantified by absorbance at 260 nm. The RNA was reverse-transcribed and RT-PCR was performed in a final reaction volume of 20 μL using iCycler 480 Probes Master (Roche Diagnostics). Each sample was analyzed in triplicate. Cycling conditions consisted of an initial denaturation step at 95 °C for 15 min, followed by 40 cycles at 95 °C for 30 s, followed by 61 °C for 1min.
Statistical analysis
As a phase 1 study, the data analysis was primarily descriptive in nature. The differences in CD8 T cell frequencies over time (pre- vs. post-vaccination) were compared using two-way ANOVA for repeated measurement data or Friedman rank-sum test as appropriate, and results were expressed in mean ± standard error of mean. Correlation analysis was performed using Spearman rank test. Progression-free survival (PFS) of the vaccinated patients was described using Kaplan-Meier product limit estimator and compared to a subset of screen failures who were considered to be appropriate for comparative analysis (HLA phenotype) by log-rank test. PFS was calculated based on the date of study registration. The distributions of demographic and clinical characteristics between two groups were also compared by two-sample t-test or Fisher’s exact test as appropriate. All analyses were two-sided and significance was set at a p-value of 0.05. Statistical analyses were performed using statistical packages SAS 9.2 (SAS Institutes, Cary, NC).
RESULTS
Study subjects and vaccine safety
A total of 53 subjects were consented and enrolled in phase 1 clinical trial between December 2009 and May 2013. Following enrollment, the HLA phenotypes of the subjects were determined, and breast cancer specimens were evaluated for MAM-A protein expression. 14 subjects met all eligibility criteria and were vaccinated. 39 subjects were screen failures. 10 of the 14 vaccinated subjects have been followed for at least 52 weeks, and detailed immune monitoring was completed in 8 of the 14 vaccinated subjects. The remaining six vaccinated subjects were not HLA-A2+, and detailed immune monitoring is ongoing in these patients, pending validation of epitopes and reagents for the specific HLA alleles expressed by these subjects. Data from all 14 vaccinated subjects is included in the safety analyses, and in the analyses of progression-free survival (PFS).
Of the 39 screen failures, 9 were excluded based on HLA phenotype, 2 were excluded because of withdrawal of consent, and one was excluded because of inability to perform IHC staining for MAM-A expression. These 12 subjects met all other inclusion/exclusion criteria and were considered suitable for comparative analysis. The 27 screen failures that were not considered suitable for comparative analysis were excluded because their breast cancers were MAM-A-negative (8/27), insurance denial (6/27), disease progression prior to completion of screening studies (4/27), lost to follow up (4/27), or ineligibility secondary to failing more than two pre-study chemotherapy regimens, dialysis, locally advanced disease, non-compliance or concurrent trastuzumab therapy (1 each). Of the vaccinated subjects, the median age was 51 years. 93% were female, 86% were white, and 100% were ER-positive. All subjects had received endocrine therapy, and the majority had bone-only metastatic disease (57%) (Table 1). Similar demographics were noted in the comparative analysis screen-failure group. None of the baseline demographic or clinical characteristics were significantly different between the vaccinated and comparative analysis screen-failure groups. Of note, screen failure and vaccinated patients received similar therapies after enrollment (Supplementary Figure 6).
TABLE 1.
Patient Demographics and baseline characteristics
| Baseline characteristics | Vaccinated n = 14 | Screen failure n = 12 | p-value |
|---|---|---|---|
| Age (years): mean ±SD (median and range) | 50.5 ± 11.1 (48.6, 33–70) | 54.5 ± 12.1 (55.0, 33–80) | 0.38 |
| Gender, n (%) | 0.99 | ||
| Male | 1 (7.1) | 0 | |
| Female | 13 (92.9) | 12 (100) | |
| Race, n (%) | 0.99 | ||
| White | 12 (85.7) | 10 (83.3) | |
| Black | 1 (7.1) | 1 (8.3) | |
| Unknown | 1 (7.1) | 1 (8.3) | |
| Biomarker profile, n (%) | |||
| ER+ | 14 (100) | 10 (83.3) | 0.20 |
| Her-2+ | 3 (21.4) | 1 (8.3) | 0.59 |
| Triple Negative | 0 (0) | 2 (16.7) | 0.20 |
| Prior Therapy, n (%) | |||
| Chemotherapy | 9 (64.3) | 9 (75) | 0.68 |
| Endocrine therapy | 14 (100) | 10 (83.3) | 0.20 |
| Radiation | 7(50) | 5 (41.7) | 0.71 |
| Surgery | 12 (85.7) | 12 (100) | 0.48 |
| Site of disease, n (%) | |||
| Bone-only | 8 (57.1) | 4 (33.3) | 0.26 |
| Viscera-only | 0 (0) | 2 (16.7) | 0.20 |
| LNs-only | 1 (7.1) | 0 (0) | 0.99 |
| Multiple sites | 4 (28.6) | 7 (58.3) | 0.23 |
| Screen Fails, n (%) | |||
| HLA typing | 9 (75) | ||
| Withdrew Consent | 2 (16.7) | ||
| Tissue Unavailable | 1 (8.3) |
In the current study, positive MAM-A expression was an inclusion criteria. MAM-A expression was quantified using the Allred scoring system (40), which is a composite score based on the proportion of cells staining positive, and the staining intensity. MAM-A IHC was performed on 36 subjects, as some subjects were identified as screen failures before MAM-A IHC could be performed. The average Allred score for the 36 subjects was 3.9 (n= 36, includes screen failures). 24/36 subjects (66%) had positive MAM-A expression (defined as Allred score 3–8). The average proportion score for the 36 subjects was 2.4 (n = 36, includes screen failures).
Vaccine safety was closely monitored following vaccination with eight or more clinical and laboratory assessments in the first 24 weeks of the trial. The most common grade 1 toxicity was malaise/flu-like symptoms (4/14). Other grade 1 toxicities potentially attributable to vaccination included vaccine site tenderness (1/14), rash (1/14), and precipitation of a shingles episode, (designated as “infection”) (1/14). A shingles episode treated with Valtrex (also designated as “infection”) was the only grade 2 toxicity (1/14). There were no grade 3 or 4 toxicities reported (Table 2).
TABLE 2.
Adverse events
| Toxicity | Grade 1 | Grade 2 | Grade 3 | Grade 4 |
|---|---|---|---|---|
| Vaccine site tenderness | 1 (7.1) | 0 | 0 | 0 |
| Malaise/flu-like symptoms | 4 (28.6) | 0 | 0 | 0 |
| Rash | 1 (7.1) | 0 | 0 | 0 |
| Infection | 1 (7.1) | 1 (7.1) | 0 | 0 |
Mammaglobin-A DNA vaccination elicits MAM-A2.1-specific CD8 T cells
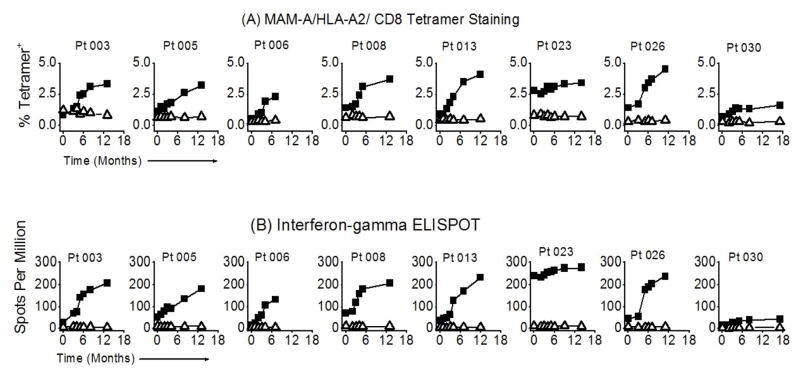
We previously demonstrated that LIYDSSLCDL, designated MAM-A2.1, is an immunodominant HLA-A2-restricted epitope derived from MAM-A (10, 38). In preclinical studies, we observed that MAM-A2.1-specific CD8 T cells expanded following MAM-A DNA vaccination of HLA-A2 transgenic mice (38). In the current study, we performed tetramer and ELISPOT analyses to measure the MAM-A2.1-specific CD8 T cell response to vaccination in HLA-A2+ subjects. The frequency of CD8+/MAM-A2.1+ T cells significantly increased following vaccination (0.9 ± 0.5% vs. 3.8 ± 1.2%, p < 0.001) (Figure 2A, top panel), with longitudinal analysis demonstrating significant responses in 6/8 subjects. Two subjects (023 and 030) did not show a significant increase in the frequency of CD8+/MAM-A2.1+ T cells. Of note, subject 023 had a significant frequency of CD8+/MAM-A2.1+ T cells prior to vaccination that increased marginally following vaccination (2.8% pre-vaccination vs. 3.4% at 12 months), while subject 030 had a low frequency of CD8+/MAM-A2.1+ T cells prior to vaccination that marginally increased after vaccination (0.7% vs. 1.6% at 12 months). ELISPOT analyses demonstrated that the number of MAM-A2.1-specific CD8 T cells capable of IFN-γ secretion increased significantly following vaccination in 6/8 subjects (41 ± 32 vs. 215 ± 67 spots per million (spm) CD8 T cells, p < 0.001) (Figure 2B, bottom panel). Consistent with the tetramer analyses, no significant change in the frequency of IFN-γ+ CD8 T cells was noted in subjects 023 (241 vs. 277 spm) or 030 (18 vs. 41 spm). Taken together, these data suggest that MAM-A DNA vaccination is capable of inducing potent MAM-A-specific CD8 T cell responses in breast cancer patients.
Figure 2. MAM-A DNA vaccination results in expansion of MAM-A-specific CD8 T cells.
PBMC were obtained prior to vaccination, and at serial time points following vaccination. The frequency and number of MAM-A-specific CD8 T cells was determined by (A) tetramer analysis, and (B) IFN-γ ELISPOT. PBMC from eight HLA-A2+ patients were stained with anti-CD8 mAb and HLA-A2/MAM-A2.1 tetramer (closed squares) or HLA-A2/Flu-M1 tetramer (open triangles). Patients were vaccinated at 0, 1, and 2 months. Data shown represent the percent CD8+/tetramer+ cells as a percentage of total CD8+ T cells. Significant increases in tetramer frequency were noted in 6/8 patients. The same PBMC samples were used to determine the number of MAM-A2.1-specific CD8 T cells producing IFN-γ by ELISPOT assay. Purified CD8 T cells were stimulated with MAM-A2.1 peptide in the presence of autologous irradiated T cell-depleted PBMCs, and the number of IFN-γ-producing cells was determined using an ELISPOT reader. Data are presented as the number of IFN-γ-producing cells per million CD8 T cells (spots per million, SPM).
MAM-A-specific CD8 T cells induced following vaccination are cytotoxic
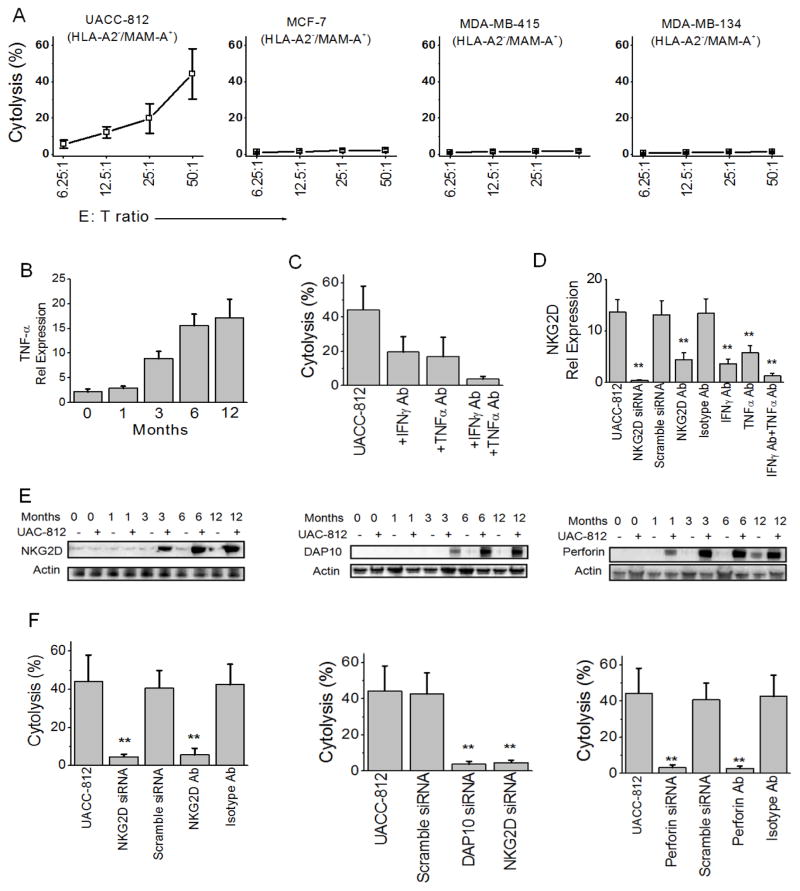
To assess whether vaccine-induced MAM-A-specific CD8 T cells are capable of lysing MAM-A+ breast cancers, we performed in vitro cytotoxicity assays against breast cancer cell lines. Purified CD8 T cells from responders were tested against a panel of breast cancer cell lines including UACC-812 (MAM-A+/HLA-A2+), MCF-7 (MAM-A−/HLA-A2+), MDA-415 (MAM-A+/HLA-A2−), and MDA-134 (MAM-A−/HLA-A2−). MAM-A and HLA-A2 expression was confirmed in breast cancer cell lines by RT-PCR (Supplementary Figure 1). CD8 T cells purified from PBMC of responders at 12 months following vaccination were able to lyse MAM-A+/HLA-A2+ UACC-812 breast cancer cells (44.3 ± 13.9% at 50:1 E/T ratio), but not control breast cancer cell lines (MCF-7, 2.2 ± 0.8%, MDA-415, 1.8 ± 0.6% or MDA-134, 1.5 ± 0.4%) (Figure 3A). These studies confirm the MAM-A specific cytotoxicity, and MHC class I restriction to HLA-A2, of the MAM-A DNA vaccine-induced CD8 T cells.
Figure 3. MAM-A DNA vaccination induces MAM-A-specific CD8 T cells capable of specifically lysing MAM-A+ breast cancer cell lines in a NKG2D/DAP10/perforin-dependent manner.
(A) CD8 T cells isolated from HLA-A2+ subjects 12 months following vaccination were tested for cytolytic activity on a panel of breast cancer cell lines at various E:T ratios. Only the HLA-A2+/MAM-A+ breast cancer cell line, UACC-812, was effectively lysed. (B) CD8 T cells were purified from PBMC at the indicated timepoints following vaccination, and TNF-α production in response to stimulation with UACC-812 breast cancer cells was determined by qRT-PCR. Similar results were observed for IFN-γ production (data not shown). (C) UACC-812-specific cytolytic activity was determined in the presence of antibodies to isotype control, IFN-γ, or TNF-α. Error bars represent the mean ± SEM from the six HLA-A2+ responders. (D) UACC-812-specific cytolytic activity was determined in the presence of antibodies to isotype control, NKG2D, IFN-γ, or TNF-α, or control or NKG2D siRNA. Error bars represent the mean ± SEM from the six HLA-A2+ responders. (E) Purified CD8 T cells from subjects vaccinated with the MAM-A DNA vaccine were stimulated with UACC-812 breast cancer cells at a 50:1 T cell to breast cancer cell ratio as indicated. Representative immunoblots for NKG2D (left panel); DAP10 (middle panel), and perforin (right panel). Actin expression represents a protein loading control. (F) Purified CD8 T cells were tested for UACC-812-specific cytolytic activity in the presence of isotype control, NKG2D, DAP-10 or perforin Ab, or control, NKG2D, DAP10 or perforin siRNA. Error bars represent the mean ± SEM from the six HLA-A2+ responders.
IFN-γ and TNF-α contribute to the cytolytic activity of MAM-A-specific CD8 T cells
IFN-γ and TNF-α have been shown to contribute to the cytolytic activity of antigen-specific CD8 T cells (41). To determine the impact of MAM-A DNA vaccination on TNF-α expression in MAM-A-specific CD8 T cells, purified CD8 T cells from vaccinated subjects were co-cultured with UACC-812 breast cancer cells, and TNF-α expression was determined by RT-PCR and immunoblot analysis. Within the sensitivity of our detection system we were not able to perform ELISA based approach to TNF-α detection. TNF-α expression increased 7.8-fold following DNA vaccination (relative expression compared to actin 2.2 ± 0.6 vs. 17.1 ± 3.7) (Figure 3B). Similar results were observed when TNF-α expression was assessed by immunoblot (Supplementary Figure 2). Both IFN-γ and TNF-α contribute to the cytolytic activity of MAM-A-specific CD8 T cells as the ability to lyse UACC-812 cells was significantly reduced following incubation with mAb specific for IFN-γ (16.6 ± 7.1% vs. 43 ± 15% for isotype control), TNF-α (14.3 ± 8.5%) or both (2.4 ± 1.1%) (Figure 3C). Taken together, these results demonstrate that MAM-A DNA vaccination is capable of inducing MAM-A specific CD8 T cells that are cytolytic, and IFN-γ and TNF-α contribute to this cytolytic activity.
NKG2D signaling contributes to the cytolytic activity of MAM-A specific CD8 T cells
NKG2D has been shown to be upregulated upon activation of CD8 T cells (42). Following vaccination, NKG2D mRNA expression was upregulated in purified CD8 T cells cultured with UACC-812 breast cancer cells (0.3 ± 0.2 relative expression compared to actin prior to vaccination vs. 13.7 ± 2.4 at 12 months, p < 0.05) (Figure 3D). Similar increases in NKG2D protein expression were also observed (Figure 3E). Of note, NKG2D mRNA and protein expression were downregulated in the presence of anti-IFN-γ mAb (3.7 ± 0.9 relative expression compared to actin), anti-TNF-α mAb (5.8 ± 1.4), or both (1.3 ± 0.5) suggesting important roles for IFN-γ and TNF-α in the induction of NKG2D expression (Figure 3D, and Supplementary Figure 3). Specific ablation or antibody neutralization of NKG2D significantly reduced the cytolytic activity of MAM-A-specific CD8 T cells (Figure 3F).
The NKG2D adapter protein DAP-10 has been shown to be an important downstream mediator of NKG2D-induced cytolytic activity (43), and perforin has been identified as a key cytotoxic effector molecule released by activated CD8 T cells (44). To determine if DAP-10 and/or perforin contribute to NKG2D-mediated cytolytic activity following vaccination, we assessed DAP-10 and perforin expression in MAM-A-specific CD8 T cells following co-culture with UACC-812 breast cancer cells. Expression of DAP-10 and perforin were increased in purified CD8 T cells following vaccination (DAP-10: 0.5 ± 0.3 relative expression compared to actin prior to vaccination vs. 9.3 ± 2.7 at 12 months, p < 0.05; perforin: 1.7 ± 0.6 relative expression compared to actin prior to vaccination vs. 16.5 ± 4.1 at 12 months, p < 0.05) (Figure 3E, middle and right panels, and Supplementary Figures 4 and 5). Specific ablation or antibody neutralization of NKG2D was associated with downregulation of DAP-10 expression (Supplementary Figure 4), and reduced perforin expression (Supplementary Figure 5), suggesting a critical role for NKG2D in the regulation of DAP-10 and perforin expression in MAM-A-specific CD8 T cells following vaccination. Similarly, antibodies to IFN-γ, TNF-α, or both strongly inhibited the expression of both DAP-10 (Supplementary Figure 4) and perforin (Supplementary Figure 5) in CD8 T cells cultured with UACC-812 cells.
Specific ablation of NKG2D or DAP10 expression was associated with a significant reduction in MAM-A specific CD8 T cell cytolytic activity (DAP10: 44.3 ± 13.9% vs. 3.8 ± 1.6%, p < 0.05; NKG2D: 44.3 ± 13.9% vs. 4.7 ± 1.3%, p < 0.05) (Figure 3F). Cytotoxicity was mediated almost exclusively through perforin, as specific ablation or antibody neutralization of perforin reduced cytolytic activity to almost zero. Based on these results we conclude that MAM-A DNA vaccination is capable of inducing MAM-A-specific CD8 T cells with significant cytolytic activity. This cytolytic activity is dependent on NKG2D, DAP10 and perforin expression.
Preliminary evidence suggests that MAM-A DNA vaccination is associated with improved progression-free survival
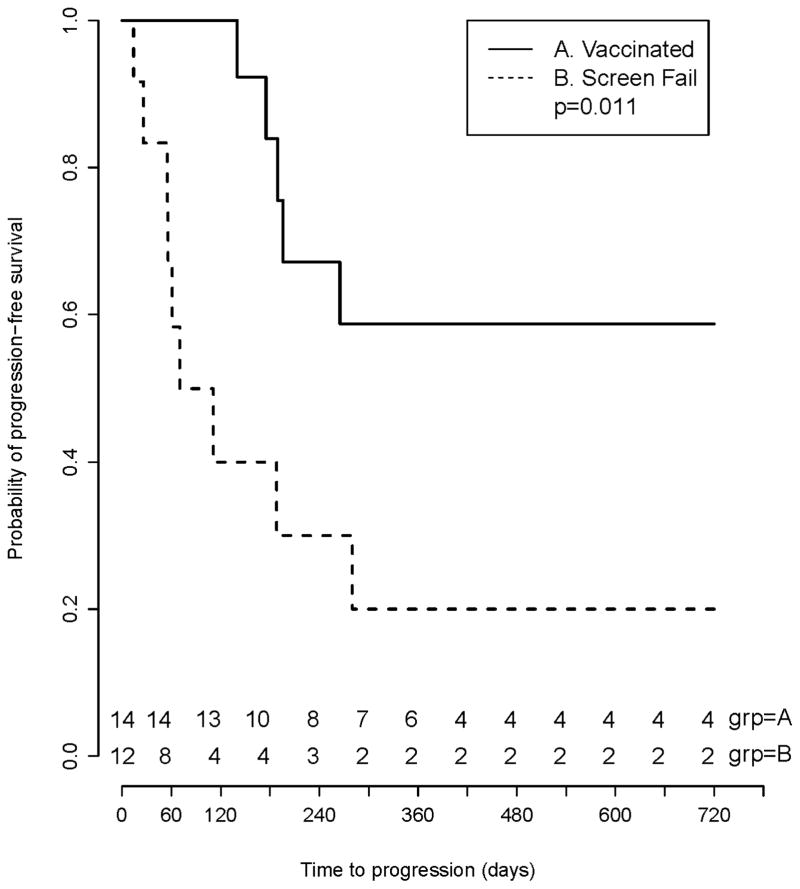
Although this study was not powered to evaluate the impact of MAM-A vaccination on progression-free survival, clinical outcome was one of the exploratory objectives. We compared progression-free survival between subjects who received the vaccine, and subjects who met all eligibility criteria and were enrolled in the trial but were screen failures because of HLA phenotype. Progression-free survival was determined by blinded review of case report forms, medical records, and diagnostic imaging results. Progression-free survival (PFS) of the vaccinated patients was described using Kaplan-Meier product limit estimator and compared to a subset of screen failures by log-rank test. MAM-A vaccination was associated with a prolonged progression-free survival (6-month PFS 53% vs. 33%, p = 0.011) (Figure 4).
Figure 4. Preliminary evidence suggests that MAM-A DNA vaccination is associated with improved progression-free survival.
A total of 53 patients were consented and enrolled in the phase 1 clinical trial. After enrollment, 39 patients were considered screen failures and 14 patients were vaccinated. Of the 39 screen failures, 12 were excluded based on HLA type, withdrawal of consent or inability to perform MAM-A IHC and were considered suitable for comparative analysis. Progression-free survival was compared between the two groups using Kaplan-Meier product limit estimator and log-rank test. Vaccinated patients are designated with a solid line, and screen failure controls are designated with a dashed line. MAM-A vaccination was associated with a prolonged progression-free survival (6-month PFS 53% vs. 33%, p = 0.011).
DISCUSSION
Preclinical studies by our group (10) and others (45–47) have demonstrated that MAM-A is an exceptional target for breast cancer vaccine therapy based on its exquisite tissue specificity (16–19), overexpression in 40–80% of breast cancers (20–24), and evidence of pre-existing immunity in breast cancer patients (8, 26–28). Based on these preclinical studies, we initiated a phase 1 clinical trial to evaluate the safety and biological efficacy of a novel plasmid MAM-A DNA vaccine. The MAM-A DNA vaccine is based on the pING vector, driving MAM-A expression by a strong viral promoter. Breast cancer patients with stable metastatic disease were eligible for participation. To date, fourteen subjects have been successfully treated with the vaccine, including eight HLA-A2+ subjects that were studied with in-depth immune monitoring analyses using PBMC collected before, during, and after vaccination.
The results of the phase 1 clinical trial demonstrate the safety of the MAM-A DNA vaccine and compelling preliminary evidence of biological efficacy. Specifically, we have made three important observations: (1) the MAM-A DNA vaccine is safe, with no significant adverse events observed in the trial. The most common grade 1 toxicity was malaise/flu-like symptoms; (2) there was a significant increase in the frequency and number of MAM-A-specific CD8 T cells following vaccination, as measured by tetramer analysis and ELISPOT; and (3) Preliminary evidence suggests that subjects who were vaccinated had improved progression-free survival compared to subjects who were enrolled in the trial but were not vaccinated because of HLA type.
Induction of effective antitumor immunity requires that a cancer vaccine be capable of eliciting robust type 1 immunity. In the last two decades considerable progress has been made in understanding the complex regulatory and signaling networks that control immune responses. It is increasingly clear that CD8 T cell responses are tightly regulated by the immune system, presumably to avoid deleterious autoimmune responses. In the context of breast cancer, regulatory networks such as the increased prevalence of T regulatory cells (Treg) (11), and upregulation of inhibitory molecules on breast cancer-specific T cells (12), are present that restrain breast cancer immune responses. These regulatory networks may also limit the T cell response to therapeutic cancer vaccines. In a previous study evaluating the CD4 T cell response to vaccination in seven of the first nine patients of this cohort we demonstrated that the frequency of Treg was significantly decreased following MAM-A DNA vaccination whereas the frequency of CD4+ICOShi MAM-A-specific T cells was increased (37). These CD4+ICOShi T cells produced higher levels of IFN-γ, and lower levels of IL-10 following vaccination.
We present evidence here that the MAM-A DNA vaccine is able to induce type 1 immune responses despite the presence of these regulatory networks. MAM-A DNA vaccination results in induction of MAM-A-specific CD8 T cells that are capable of producing both IFN-γ and TNF-α, and lysing MAM-A+ breast cancer cells. MAM-A-specific CD8 T cells also have increased NKG2D, DAP-10 and perforin expression. Of note, in CD8 T cells, NKG2D signaling has been shown to augment proliferation and cytotoxicity upon antigen encounter, suggesting that NKG2D functions as a costimulatory molecule (48). Our studies demonstrate that following MAM-A DNA vaccination there is specific upregulation of NKG2D and DAP-10 expression (Figure 5). Although this response is not unique to MAM-A vaccination, it does provide strong supporting evidence for the efficacy of the MAM-A DNA vaccine. Taken together, the results of these two studies strongly suggest that the MAM-A DNA vaccine is capable of inducing strong type 1 immune responses. The humoral response to vaccination was not measured.
Current cancer vaccine clinical development paradigms emphasize the early assessment of cancer vaccine efficacy in an appropriate clinical context (49). These paradigms have been endorsed in the literature and in FDA guidance documents. Although treatment of patients with metastatic disease is appropriate for first-in-human studies of a novel biologic therapeutic, cancer vaccine treatment of patients with metastatic disease does have unique limitations. First, it is very difficult to assess the biological efficacy of cancer vaccines in patients with metastatic disease, as the time interval from vaccine administration to subsequent disease progression in patients with metastatic disease may be too short to develop a productive antitumor immune response. Second, patients with metastatic disease have typically received multiple prior cancer treatments, which may be detrimental to the immune system. The fact that preliminary evidence suggests that the MAM-A DNA vaccine was associated with improved progression-free survival in patients with metastatic breast cancer is notable, supporting ongoing clinical development of the MAM-A DNA vaccine. Studies to define the clinical potential of a MAM-A DNA vaccine are particularly important given the safety of the vaccine, evidence of biological efficacy, exquisite tissue specificity, and near-universal expression of MAM-A in breast cancer, and the potential of MAM-A as a target for a breast cancer prevention vaccine.
Supplementary Material
Statement of translational relevance.
MAM-A is an important breast cancer-associated antigen with exquisite tissue specificity. We initiated a phase 1 clinical trial to evaluate the safety and efficacy of a MAM-A DNA vaccine. Breast cancer patients with stable metastatic disease were eligible. We demonstrate that (1) the MAM-A DNA vaccine is safe, with no significant adverse events observed, (2) there is a significant increase in the frequency and number of MAM-A-specific CD8 T cells following vaccination, and (3) preliminary evidence suggests improved progression-free survival. Additional studies are required to define the clinical potential of a MAM-A DNA vaccine for breast cancer treatment or prevention.
Acknowledgments
FUNDING
This project was funded by research grants DOD/CDMRP-BCRP W81XWH-06-1-0677 (WEG), and Gateway for Cancer Research P-06-016 (WEG). VT, PG, and TM are funded by the Foundation for Barnes-Jewish Hospital. NT was supported by a NCI grant T32 CA 009621. We thank the Alvin J. Siteman Cancer Center at Washington University School of Medicine and Barnes-Jewish Hospital in St. Louis, MO., for the use of the Biologic Therapy Core Facility; the Clinical Trials Core, and Biostatistics Core which provided critical support for the vaccine manufacture and validation; design and performance of the clinical trial, and data analysis, respectively. The Siteman Cancer Center is supported in part by NCI Cancer Center Support Grant #P30 CA91842. We are also extremely grateful to George and Diana Holway, whose generous gift provided the resources necessary to help us realize the promise of new discovery.
Footnotes
Disclosure of Potential Conflict of Interest: None of the authors have any conflicts of interest to disclose.
Breast cancer cell lines and cell culture. The breast cancer cell lines, UACC-812 (MAM-A+/HLA-A2+), MCF-7 (MAM-A−/HLA-A2+), MDA-MB-415 (MAM-A+/HLA-A2−), MDA-MB-134 (MAM-A−/HLA-A2−), were obtained from the American Type Culture Collection (ATCC, Manassas, VA). The cell lines were ordered in 2004 by Dr Mohanakumar laboratory. No specific authentication of the cell lines was performed in our laboratory. All experiments were performed on the cell lines below 30th passage. Breast cancer cell lines were cultured in RPMI-1640 culture medium at 37 °C in 5% CO2 incubator until they were 80% confluent. The presence of MAM-A in the cell lines was characterized and confirmed by reverse transcriptase-polymerase chain reaction and western blot analysis.
References
- 1.Senovilla L, Vacchelli E, Garcia P, Eggermont A, Fridman WH, Galon J, et al. Trial watch: DNA vaccines for cancer therapy. Oncoimmunology. 2013;2:e23803. doi: 10.4161/onci.23803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ferraro B, Morrow MP, Hutnick NA, Shin TH, Lucke CE, Weiner DB. Clinical applications of DNA vaccines: current progress. Clin Infect Dis. 2011;53:296–302. doi: 10.1093/cid/cir334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.DeNardo DG, Brennan DJ, Rexhepaj E, Ruffell B, Shiao SL, Madden SF, et al. Leukocyte complexity predicts breast cancer survival and functionally regulates response to chemotherapy. Cancer Discov. 2011;1:54–67. doi: 10.1158/2159-8274.CD-10-0028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Disis ML, Calenoff E, McLaughlin G, Murphy AE, Chen W, Groner B, et al. Existent T-cell and antibody immunity to HER-2/neu protein in patients with breast cancer. Cancer Res. 1994;54:16–20. [PubMed] [Google Scholar]
- 5.Disis ML, Knutson KL, Schiffman K, Rinn K, McNeel DG. Pre-existent immunity to the HER-2/neu oncogenic protein in patients with HER-2/neu overexpressing breast and ovarian cancer. Breast Cancer Res Treat. 2000;62:245–52. doi: 10.1023/a:1006438507898. [DOI] [PubMed] [Google Scholar]
- 6.Disis ML, Pupa SM, Gralow JR, Dittadi R, Menard S, Cheever MA. High-titer HER-2/neu protein-specific antibody can be detected in patients with early-stage breast cancer. J Clin Oncol. 1997;15:3363–7. doi: 10.1200/JCO.1997.15.11.3363. [DOI] [PubMed] [Google Scholar]
- 7.von Mensdorff-Pouilly S, Snijdewint FG, Verstraeten AA, Verheijen RH, Kenemans P. Human MUC1 mucin: a multifaceted glycoprotein. Int J Biol Markers. 2000;15:343–56. doi: 10.1177/172460080001500413. [DOI] [PubMed] [Google Scholar]
- 8.Jaramillo A, Majumder K, Manna PP, Fleming TP, Doherty G, Dipersio JF, et al. Identification of HLA-A3-restricted CD8+ T cell epitopes derived from mammaglobin-A, a tumor-associated antigen of human breast cancer. Int J Cancer. 2002;102:499–506. doi: 10.1002/ijc.10736. [DOI] [PubMed] [Google Scholar]
- 9.Jaramillo A, Narayanan K, Campbell LG, Benshoff ND, Lybarger L, Hansen TH, et al. Recognition of HLA-A2-restricted mammaglobin-A-derived epitopes by CD8+ cytotoxic T lymphocytes from breast cancer patients. Breast Cancer Res Treat. 2004;88:29. doi: 10.1007/s10549-004-8918-1. [DOI] [PubMed] [Google Scholar]
- 10.Narayanan K, Jaramillo A, Benshoff ND, Campbell LG, Fleming TP, Dietz JR, et al. Response of established human breast tumors to vaccination with mammaglobin-A cDNA. J Natl Cancer Inst. 2004;96:1388–96. doi: 10.1093/jnci/djh261. [DOI] [PubMed] [Google Scholar]
- 11.Liyanage UK, Moore TT, Joo HG, Tanaka Y, Herrmann V, Doherty G, et al. Prevalence of regulatory T cells is increased in peripheral blood and tumor microenvironment of patients with pancreas or breast adenocarcinoma. J Immunol. 2002;169:2756–61. doi: 10.4049/jimmunol.169.5.2756. [DOI] [PubMed] [Google Scholar]
- 12.Ghebeh H, Mohammed S, Al-Omair A, Qattan A, Lehe C, Al-Qudaihi G, et al. The B7-H1 (PD-L1) T lymphocyte-inhibitory molecule is expressed in breast cancer patients with infiltrating ductal carcinoma: correlation with important high-risk prognostic factors. Neoplasia. 2006;8:190–8. doi: 10.1593/neo.05733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Aslan N, Yurdaydin C, Wiegand J, Greten T, Ciner A, Meyer MF, et al. Cytotoxic CD4 T cells in viral hepatitis. J Viral Hepat. 2006;13:505–14. doi: 10.1111/j.1365-2893.2006.00723.x. [DOI] [PubMed] [Google Scholar]
- 14.Peoples GE, Gurney JM, Hueman MT, Woll MM, Ryan GB, Storrer CE, et al. Clinical trial results of a HER2/neu (E75) vaccine to prevent recurrence in high-risk breast cancer patients. J Clin Oncol. 2005;23:7536–45. doi: 10.1200/JCO.2005.03.047. [DOI] [PubMed] [Google Scholar]
- 15.Watson MA, Fleming TP. Isolation of differentially expressed sequence tags from human breast cancer. Cancer Res. 1994;54:4598–602. [PubMed] [Google Scholar]
- 16.Fleming TP, Watson MA. Mammaglobin, a breast-specific gene, and its utility as a marker for breast cancer. Ann N Y Acad Sci. 2000;923:78–89. doi: 10.1111/j.1749-6632.2000.tb05521.x. [DOI] [PubMed] [Google Scholar]
- 17.Goedegebuure PS, Watson MA, Viehl CT, Fleming TP. Mammaglobin-based strategies for treatment of breast cancer. Curr Cancer Drug Targets. 2004;4:531. doi: 10.2174/1568009043332862. [DOI] [PubMed] [Google Scholar]
- 18.Watson MA, Darrow C, Zimonjic DB, Popescu NC, Fleming TP. Structure and transcriptional regulation of the human mammaglobin gene, a breast cancer associated member of the uteroglobin gene family localized to chromosome 11q13. Oncogene. 1998;16:817. doi: 10.1038/sj.onc.1201597. [DOI] [PubMed] [Google Scholar]
- 19.Watson MA, Fleming TP. Mammaglobin, a mammary-specific member of the uteroglobin gene family, is overexpressed in human breast cancer. Cancer Res. 1996;56:860–5. [PubMed] [Google Scholar]
- 20.Wang Z, Spaulding B, Sienko A, Liang Y, Li H, Nielsen G, et al. Mammaglobin, a valuable diagnostic marker for metastatic breast carcinoma. Int J Clin Exp Pathol. 2009;2:384–9. [PMC free article] [PubMed] [Google Scholar]
- 21.Watson MA, Dintzis S, Darrow CM, Voss LE, DiPersio J, Jensen R, et al. Mammaglobin expression in primary, metastatic, and occult breast cancer. Cancer Res. 1999;59:3028–31. [PubMed] [Google Scholar]
- 22.Sasaki E, Tsunoda N, Hatanaka Y, Mori N, Iwata H, Yatabe Y. Breast-specific expression of MGB1/mammaglobin: an examination of 480 tumors from various organs and clinicopathological analysis of MGB1-positive breast cancers. Mod Pathol. 2007;20:208–14. doi: 10.1038/modpathol.3800731. [DOI] [PubMed] [Google Scholar]
- 23.Lewis GH, Subhawong AP, Nassar H, Vang R, Illei PB, Park BH, et al. Relationship between molecular subtype of invasive breast carcinoma and expression of gross cystic disease fluid protein 15 and mammaglobin. Am J Clin Pathol. 2011;135:587–91. doi: 10.1309/AJCPMFR6OA8ICHNH. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Luo MH, Huang YH, Ni YB, Tsang JY, Chan SK, Shao MM, et al. Expression of mammaglobin and gross cystic disease fluid protein-15 in breast carcinomas. Hum Pathol. 2013;44:1241–50. doi: 10.1016/j.humpath.2012.10.009. [DOI] [PubMed] [Google Scholar]
- 25.Watson MA, Dintzis S, Darrow CM, Voss LE, DiPersio J, Jensen R, et al. Mammaglobin expression in primary, metastatic, and occult breast cancer. Cancer Res. 1999;59:3028. [PubMed] [Google Scholar]
- 26.Tiriveedhi V, Sarma NJ, Subramanian V, Fleming TP, Gillanders WE, Mohanakumar T. Identification of HLA-A24-restricted CD8(+) cytotoxic T-cell epitopes derived from mammaglobin-A, a human breast cancer-associated antigen. Hum Immunol. 2012;73:11–6. doi: 10.1016/j.humimm.2011.10.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ilias Basha H, Tiriveedhi V, Fleming TP, Gillanders WE, Mohanakumar T. Identification of immunodominant HLA-B7-restricted CD8+ cytotoxic T cell epitopes derived from mammaglobin-A expressed on human breast cancers. Breast Cancer Res Treat. 2011;127:81–9. doi: 10.1007/s10549-010-0975-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jaramillo A, Narayanan K, Campbell LG, Benshoff ND, Lybarger L, Hansen TH, et al. Recognition of HLA-A2-restricted mammaglobin-A-derived epitopes by CD8+ cytotoxic T lymphocytes from breast cancer patients. Breast Cancer Res Treat. 2004;88:29–41. doi: 10.1007/s10549-004-8918-1. [DOI] [PubMed] [Google Scholar]
- 29.Fioretti D, Iurescia S, Fazio VM, Rinaldi M. DNA vaccines: developing new strategies against cancer. J Biomed Biotechnol. 2010;2010:174378. doi: 10.1155/2010/174378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Klinman DM, Yi AK, Beaucage SL, Conover J, Krieg AM. CpG motifs present in bacteria DNA rapidly induce lymphocytes to secrete interleukin 6, interleukin 12, and interferon gamma. Proc Natl Acad Sci U S A. 1996;93:2879–83. doi: 10.1073/pnas.93.7.2879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sato Y, Roman M, Tighe H, Lee D, Corr M, Nguyen MD, et al. Immunostimulatory DNA sequences necessary for effective intradermal gene immunization. Science. 1996;273:352–4. doi: 10.1126/science.273.5273.352. [DOI] [PubMed] [Google Scholar]
- 32.Chuang I, Sedegah M, Cicatelli S, Spring M, Polhemus M, Tamminga C, et al. DNA prime/Adenovirus boost malaria vaccine encoding P. falciparum CSP and AMA1 induces sterile protection associated with cell-mediated immunity. PLoS One. 2013;8:e55571. doi: 10.1371/journal.pone.0055571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kalams SA, Parker SD, Elizaga M, Metch B, Edupuganti S, Hural J, et al. Safety and comparative immunogenicity of an HIV-1 DNA vaccine in combination with plasmid interleukin 12 and impact of intramuscular electroporation for delivery. J Infect Dis. 2013;208:818–29. doi: 10.1093/infdis/jit236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pavlenko M, Roos AK, Lundqvist A, Palmborg A, Miller AM, Ozenci V, et al. A phase I trial of DNA vaccination with a plasmid expressing prostate-specific antigen in patients with hormone-refractory prostate cancer. Br J Cancer. 2004;91:688–94. doi: 10.1038/sj.bjc.6602019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Rice J, Ottensmeier CH, Stevenson FK. DNA vaccines: precision tools for activating effective immunity against cancer. Nat Rev Cancer. 2008;8:108–20. doi: 10.1038/nrc2326. [DOI] [PubMed] [Google Scholar]
- 36.Kutzler MA, Weiner DB. DNA vaccines: ready for prime time? Nat Rev Genet. 2008;9:776–88. doi: 10.1038/nrg2432. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tiriveedhi V, Fleming TP, Goedegebuure PS, Naughton M, Ma C, Lockhart C, et al. Mammaglobin-A cDNA vaccination of breast cancer patients induces antigen-specific cytotoxic CD4+ICOShi T cells. Breast Cancer Res Treat. 2013;138:109–18. doi: 10.1007/s10549-012-2110-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Bharat A, Benshoff N, Fleming TP, Dietz JR, Gillanders WE, Mohanakumar T. Characterization of the role of CD8+T cells in breast cancer immunity following mammaglobin-A DNA vaccination using HLA-class-I tetramers. Breast Cancer Res Treat. 2008;110:453–63. doi: 10.1007/s10549-007-9741-2. [DOI] [PubMed] [Google Scholar]
- 39.Tiriveedhi V, Gelman AE, Mohanakumar T. HIF-1alpha signaling by airway epithelial cell K-alpha1-tubulin: role in fibrosis and chronic rejection of human lung allografts. Cell Immunol. 2012;273:59–66. doi: 10.1016/j.cellimm.2011.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Allred DC, Harvey JM, Berardo M, Clark GM. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod Pathol. 1998;11:155–68. [PubMed] [Google Scholar]
- 41.Ratner A, Clark WR. Role of TNF-alpha in CD8+ cytotoxic T lymphocyte-mediated lysis. J Immunol. 1993;150:4303–14. [PubMed] [Google Scholar]
- 42.Chu T, Tyznik AJ, Roepke S, Berkley AM, Woodward-Davis A, Pattacini L, et al. Bystander-Activated Memory CD8 T Cells Control Early Pathogen Load in an Innate-like, NKG2D-Dependent Manner. Cell Rep. 2013;3:701–8. doi: 10.1016/j.celrep.2013.02.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Burgess SJ, Marusina AI, Pathmanathan I, Borrego F, Coligan JE. IL-21 down-regulates NKG2D/DAP10 expression on human NK and CD8+ T cells. J Immunol. 2006;176:1490–7. doi: 10.4049/jimmunol.176.3.1490. [DOI] [PubMed] [Google Scholar]
- 44.Kremer M, Suezer Y, Volz A, Frenz T, Majzoub M, Hanschmann KM, et al. Critical role of perforin-dependent CD8+ T cell immunity for rapid protective vaccination in a murine model for human smallpox. PLoS Pathog. 2012;8:e1002557. doi: 10.1371/journal.ppat.1002557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.O’Brien N, Maguire TM, O’Donovan N, Lynch N, Hill AD, McDermott E, et al. Mammaglobin a: a promising marker for breast cancer. Clin Chem. 2002;48:1362–4. [PubMed] [Google Scholar]
- 46.Cui H, Zhang W, Hu W, Liu K, Wang T, Ma N, et al. Recombinant mammaglobin A adenovirus-infected dendritic cells induce mammaglobin A-specific CD8+ cytotoxic T lymphocytes against breast cancer cells in vitro. PLoS One. 2013;8:e63055. doi: 10.1371/journal.pone.0063055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lerret NM, Rogozinska M, Jaramillo A, Marzo AL. Adoptive transfer of Mammaglobin-A epitope specific CD8 T cells combined with a single low dose of total body irradiation eradicates breast tumors. PLoS One. 2012;7:e41240. doi: 10.1371/journal.pone.0041240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Jamieson AM, Diefenbach A, McMahon CW, Xiong N, Carlyle JR, Raulet DH. The role of the NKG2D immunoreceptor in immune cell activation and natural killing. Immunity. 2002;17:19–29. doi: 10.1016/s1074-7613(02)00333-3. [DOI] [PubMed] [Google Scholar]
- 49.Hoos A, Parmiani G, Hege K, Sznol M, Loibner H, Eggermont A, et al. A clinical development paradigm for cancer vaccines and related biologics. J Immunother (1997) 2007;30:1–15. doi: 10.1097/01.cji.0000211341.88835.ae. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.