Abstract
Chordomas are rare malignant tumors that are postulated to arise from remnants of the notochord. Currently, the interaction between chordomas and the host immune system is poorly understood. The checkpoint protein, PD-1 is expressed by circulating lymphocytes and is a marker of activation and exhaustion. Its ligands, PD-L1 (B7-H1, CD274) and PD-L2 (B7-DC, CD273), are expressed on a variety of human cancers; however this pathway has not been previously reported in chordomas. We used flow cytometric and RT-PCR analysis in three established primary and recurrent chordoma cell lines (U-CH1, U-CH2, and JHC7) as well as immunohistochemical analysis of chordoma tissues from 10 patients to identify and localize expression of PD-1 pathway proteins. PD-1 ligands are not constitutively expressed by chordoma cells, but their expression is induced in the setting of pro-inflammatory cytokines in all cell lines examined. In paraffin embedded tissues, we found that tumor infiltrating lymphocytes expressed PD-1 in 3/6 cases. We also found that, although chordoma cells did not express significant levels of PD-L1, PD-L1 expression was observed on tumor-infiltrating macrophages and tumor infiltrating lymphocytes. Our study suggests that PD-1, PD-L1, and PD-L2 are present in the microenvironment of a subset of chordomas analyzed. Future studies are needed to evaluate the contribution of the PD-1 pathway to the immunosuppressive microenvironment of chordomas.
Keywords: Chordomas, PD-1, PD-L1, B7-H1, PD-L2
Introduction
Chordomas are locally invasive malignant tumors hypothesized to be derived from remnants of the notochord. These tumors typically arise from the axial skeleton and account for 1–4 % of all primary bone tumors and 0.1–0.2 % of all intracranial neoplasms [1, 2]. Surgical resection and radiation are the mainstays of treatment as no chemotherapeutics have been approved for chordomas [3, 4]. These tumors are associated with significant morbidity and mortality with survival rates at 5, 10, and 20 years of 67, 40 and 13 %, respectively [5].
Despite extensive genetic and molecular analysis of cellular pathways implicated in chordoma pathogenesis, little is known about the interaction between chordomas and the host immune system [6–10]. Immune checkpoint proteins and their ligands are a group of surface-bound and secreted proteins that function to regulate immune responses to host proteins. This interaction serves as a potent negative signal for immune cells, rendering them exhausted or inactive (Fig. 1a) [11].
Fig. 1.
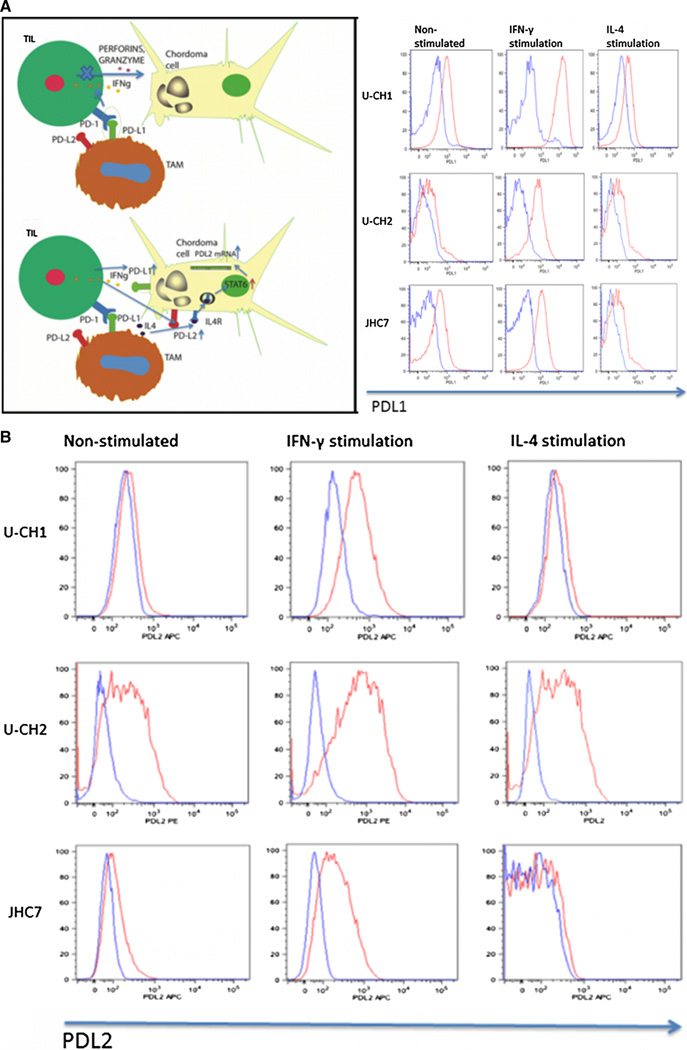
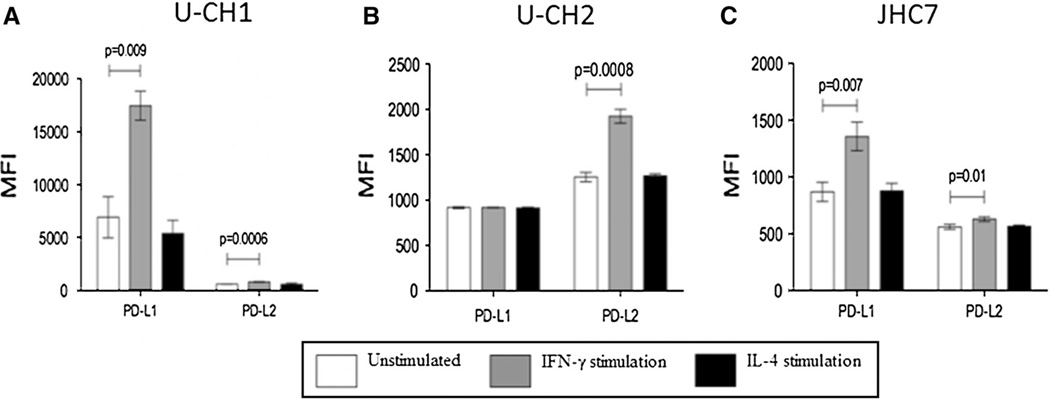
a Schematic representation of the PD-1–PD-L1 pathway and its modulation by cytokines expressed in the chordoma microenvironment and analysis of the surface expression of PD-L1 by the U-CH1, U-CH2, and JHC7 cell lines with flow cytometry. The blue curve denotes the isotype control and the red line the PD-L1 staining. Expression of PD-L1 was analyzed under basal conditions and following stimulation with IFN-γ and IL-4. The U-CH1, U-CH2, and JHC-7 cell lines all showed basal expression of PD-L1 (column 1) and inducible levels of PD-L1 by IFN-γ (column 2). b Analysis of the surface expression of PD-L2 by the U-CH1, U-CH2, and JHC7 cell lines by flow cytometry. Expression of PD-L2 was analyzed under basal conditions (column 1) and following stimulation with IFN-γ (column 2) and IL-4 (column 3). c RT-PCR demonstrating the relative expression of PD-L1 and PD-L2 in the U-CH1 and U-CH2 cell lines. c The fold expression of mRNA transcripts of PD-L1 and PD-L2 were analyzed in the basal state and following stimulation with IFN-γ and IL-4. Both PD-L1 and PD-L2 were expressed following exposure to IFN-γ in the U-CH1 and U-CH2 cell lines. Exposure to IL-4 increased expression of PD-L2 in the U-CH2 cell line
Programmed death 1 (PD-1) and its ligands, PD-L1 (B7-H1, CD274) and PD-L2 (B7-DC, CD273), have been implicated in promoting progression of central nervous system tumors such as glioblastoma as well as solid tumors outside the central nervous system, including lung, gastric, and pancreatic cancers, among others [12–17]. PD-1 is expressed on the surface of lymphocytes, and its ligands, PD-L1 and PD-L2, are expressed by tumor cells, tumor-associated immune cells, and stromal cells [14, 15].
To date, no study has analyzed the PD-1 pathway in human chordomas. We used established chordoma cell lines and resected chordoma tissues to evaluate the expression of these surface markers in chordoma. We also assessed the inducible expression of PD-L1 and PD-L2 in chordoma cell lines, since cytokine stimulation has been associated with up-regulated expression of these proteins in other tumors [25].
Materials and methods
Three human chordoma cell lines and archived tissue from 10 patients were used to assess the levels of PD-1, PD-L1 and PD-L2 expression in chordomas.
Cell lines
The established human chordoma cell lines U-CH1 and U-CH2 were kindly provided by Dr. Michael Kelly (Duke University School of Medicine). U-CH1 is a chordoma cell line derived from a recurrent sacrococcygeal chordoma treated with radiation and UCH-2 is derived by a recurrent sacral tumor treated with radiation. JHC7, a third human chordoma cell line, is derived by a surgically resected primary spinal chordoma and was established as previously described [18]. The U-CH1 and U-CH2 cell lines were cultured in gelatin coated flasks and were grown with IMDM–RPMI media (4:1) supplemented with 10 % FBS and 1 % penicillin–streptomycin. The JHC7 cell line was cultured in non-gelatin coated flasks. Cells were grown in MesenPRO basal media plus supplement (Invitrogen, Carlsbad, California, USA), 5 mM glutamine, and 1 % antibiotic and antimycotic for the first 2 passages and were switched to DMEM supplemented with 10 % FBS and 1 % penicillin–streptomycin in sequential passages. All three cell lines were incubated at 37 °C in a humidified atmosphere containing 5 % CO2.
Flow cytometry
Surface expression of PD-L1 and PD-L2 was assessed in the U-CH1, U-CH2, and JHC7 cell lines. Baseline expression and inducible PD-L1 and PD-L2 expression were assessed after interferon-gamma (IFN-γ) and interleukin-4 (IL-4) treatments. Cells were treated with IFN-γ at a dose of 10 ng/ml (eBioscience, San Diego, California, USA) or IL-4 (eBioscience) at a dose of 50 ng/ml for 24 h. Cells were trypsinized, washed with PBS, and stained with PD-L1 PE Cy7 (eBioscience) and PD-L2 APC or PE (eBioscience). Aqua fixable viability dye (Life Technologies, Grand Island, New York, USA) was used to exclude dead cells from the analysis. Stained cells were analyzed using a LSR II flow cytometer (BD Biosciences, San Jose, California, USA). Data analysis was performed with FlowJo software (Tree Star Inc., Ashland, Oregon, USA). We considered positive PDL1, PDL2 staining only when >5 % of the cells had a higher fluorescent intensity than the isotype control. All flow experiments have been repeated at least in triplicates.
Real time-polymerase chain reaction (RT-PCR)
RNA was extracted from U-CH1 and U-CH2 cells at baseline and following stimulation with IFN-γ (10 ng/ml) and IL-4 (50 ng/ml). Tumor RNA was extracted using the RNeasy Mini Kit (Qiagen, Valencia, California, USA) according to the manufacturer’s protocol. 1 µg of RNA was used for cDNA synthesis using the QuantiTect Reverse Transcription Kit (Qiagen) in order to synthesize 100 µl of cDNA. RT-PCR was performed using the Step-One Plus detection system (Applied Biosystems, Grand Island, New York, USA). Reactions were performed at a final volume of 25 µl which contained 1 µl cDNA, 12.5 µl TaqMan Universal Master Mix (Applied Biosystems), 1.25 µl of 18s VIC-TAMRA rRNA primer mix (Applied Biosystems), 1.25 µl of primer mix (sense and anti-sense) of the gene of interest and 9 µl of DEPC treated water. All oligonucleotide primers for RT-PCR were synthesized by Applied Biosystems (Table S1). The same thermal profile conditions were used for all primer sets. Samples were run in triplicate. The 2-ΔΔCt was used to analyze the relative gene expression of the stimulated groups versus controls. All RT-PCR experiments have been repeated three times (three independent experiments) with similar results. One RT-PCR round of the three has been used to draw statistical conclusions.
Immunohistochemistry (IHC) for PD-1 and PD-L1
IHC was performed on 10 formalin-fixed, paraffin embedded (FFPE) blocks from chordoma samples identified in the Johns Hopkins Hospital surgical pathology archives. Samples were cut in 5 µm-thick sections and stained for PD-1 or PD-L1 as previously described [19, 20]. 1.8 µg/µl of 5H1 monoclonal antibody (mAb) was used for PD-L1 staining, and 1 µg/µl of M3 mAb was used for PD-1 staining. Tonsil tissue was used as both a positive and negative control for PD-1 staining. An IgG isotype control was used for PD-L1 staining. In addition, native tissue elements also served as internal positive and negative controls for both PD-1 and PD-L1 staining. Tumor cells and infiltrating immune cells were scored separately for PD-L1 expression, with a threshold of >5 % membranous expression considered to be positive. 20 A threshold >5 % of tumor infiltrating lymphocytes (TIL) expressing PD-1 was also used to consider a case positive for PD-1. CD3 and CD68 IHC for all cases was performed according to standard automated protocol. Three board-certified pathologists (JMT, PCB, EFM) independently scored PD-1 and PD-L1 staining and the geographic co-localization of these markers within the tumor microenvironment.
Patient data
Patient information, including tumor location, previous treatment, surgical margins, and tumor recurrence were obtained by retrospective review of 10 patients treated at the Johns Hopkins Hospital. Collection of all patient information was completed with institutional review board approval from the Johns Hopkins institutional review board committee.
Statistical analysis
Statistical analysis was performed using Prism 6 (Graph-Pad Software Inc., San Diego, California, USA). Differences between groups were assessed using an unpaired, non-parametric, two-tailed t test. A p value ≤0.05 was considered statistically significant. The relative expression and the error bars for the 2−ΔΔCt method were calculated using Data Assist (Applied Biosystems).
Results
Inducible membranous expression of PD-L1 and PD-L2 proteins is present in chordoma cell lines
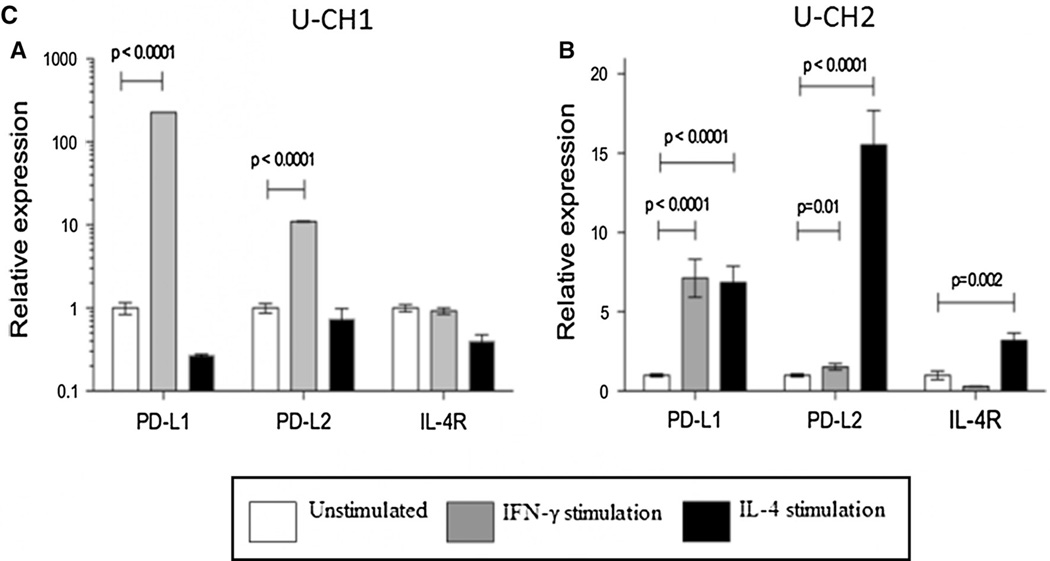
Membranous expression of PD-L1 and PD-L2 on patient derived chordoma cell lines was confirmed by flow cytometry in the U-CH1, U-CH2, and JHC7 cell lines. All cell lines expressed basal levels of PD-L1 (Fig. 1a) and PD-L2 (Fig. 1b) in <5 % of total number of cells analyzed. Stimulation of the U-CH1with IFN-γ resulted in a statistically significant up-regulation of both PD-L1 (baseline MFI = 6,941 versus IFNg stimulation MFI = 17,467, p = 0.009) and PD-L2 (baseline MFI = 617 versus IFNg stimulation MFI = 843, p = 0.0006). Stimulation of JHC7 with IFN-γ resulted in a statistically significant up-regulation of both PD-L1 (baseline MFI = 869 versus IFNg stimulation MFI = 1,365, p = 0.007) and PD-L2 (baseline MFI = 560 versus IFNg stimulation MFI = 629, p = 0.01). Stimulation with IFN-γ resulted in the up-regulation of PD-L2 (baseline MFI = 1,254 versus IFNg stimulation MFI = 1,924, p = 0.0008) but not PD-L1 in the U-CH2 cell line. Stimulation with IL-4 did not lead to a significant change in the expression of PD-L1 nor PD-L2 in any of the cell lines. Interestingly, when the media was switched from the stem cell media to the FBS supplemented media, JHC7 cells exhibited neither baseline nor inducible PD-L1 expression. The median fluorescent intensity (MFI) of PD-L1 and PD-L2 is shown for U-CH1, U-CH2, and JHC7 (Fig. 2). These data suggest that PD-L1 and PD-L2 are inducible after exposure to IFN-γ.
Fig. 2.
Comparison of PD-L1 and PD-L2 MFI of the U-CH1, U-CH2, and JHC7 cell lines by flow cytometry. MFI values of PD-L1 and PD-L2 were calculated under basal conditions (column 1) and following stimulation with IFN-γ (column 2) and IL-4 (column 3)
Basal and inducible PD-L1 and PD-L2 are expressed at the mRNA level in chordoma cell lines
RT-PCR was used to confirm expression of PD-L1 and PD-L2 at the mRNA level in the U-CH1 and U-CH2 (Fig. 1c) chordoma cell lines. U-CH1 and U-CH2 showed a significant up-regulation of PD-L1 and PD-L2 mRNA transcripts upon IFN-γ stimulation (p < 0.0001 for both cell lines). Interestingly, IL-4 stimulation led to significant up-regulation of the PD-L1, PD-L2, and the IL-4 receptor (IL-4R) mRNA transcripts in the U-CH2 cell line but not the U-CH1 cell line.
PD-1 and PD-L1 expression in the human chordoma microenvironment
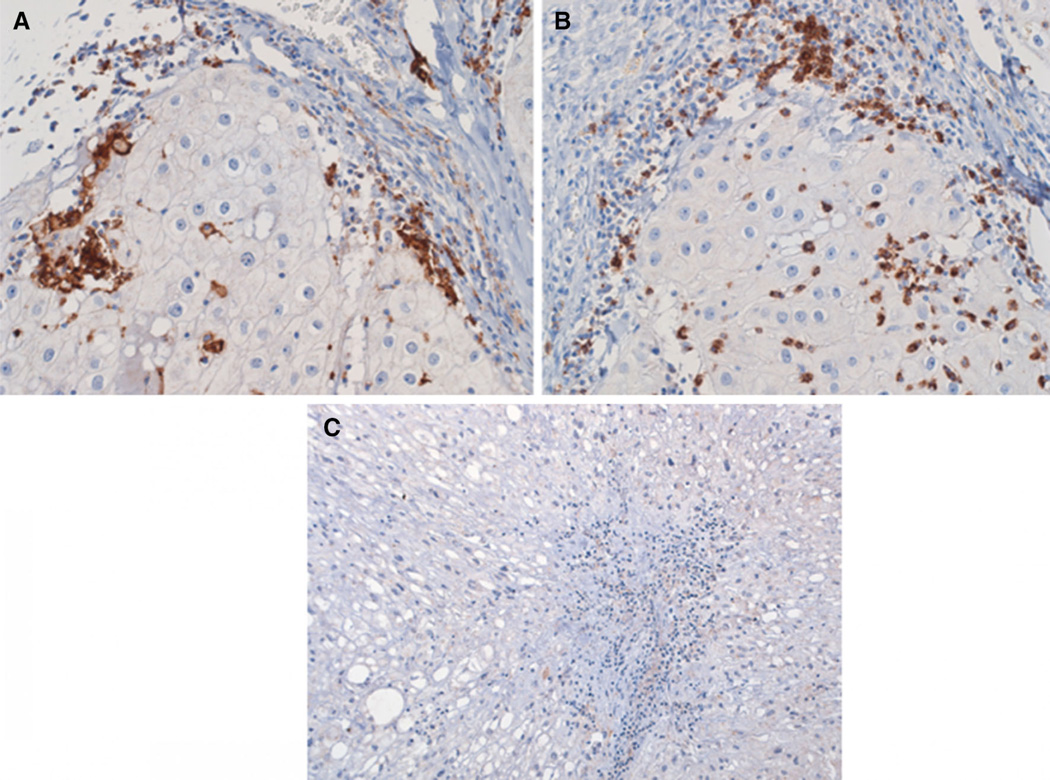
Infiltrating immune cells (tumor infiltrating lymphocytes and tumor infiltrating macrophages) were observed in FFPE archival human chordoma tissues in 6/10 cases. Of the cases with positive staining for immune cells, 3/6 demonstrated PD-1 expression by immune cells and 4/6 demonstrated membranous PD-L1 expression in areas infiltrated with tissue associated macrophages (TAM) and TILs. For the three cases that demonstrated both PD-1 and PD-L1 expression, the expression of these molecules was observed primarily at the interface between tumor and stroma (Figs. S1, 3). Scattered infiltrating immune cells were observed that co-localized with PD-1 and PD-L1 staining. None (0/10) of the cases demonstrated robust expression of PD-L1 by tumor cells. Clinical and pathologic characteristics are summarized in Table 1. Collectively, these findings suggest that a subset of chordomas express the PD-1/PD-L1 pathway.
Table 1.
Clinicopathologic features of ten chordoma cases analyzed for PD-1 and PD-L1 expression by immunohistochemistry
| Case | Tumor location |
Previous treatment |
Surgical margins |
Presence of tumor infiltrating lymphocytes |
PD-1 expression on tumor infiltrating lymphocytes |
PD-L1 expression on chordoma cells*** |
PD-L1 expression on infiltrating immune cells (TIL and TAM) |
|---|---|---|---|---|---|---|---|
| 1 | Clivus | None | Negative | Yes | Negative | Negative | Positive |
| 2 | Sacrum | None | Positive | Yes | Positive | Negative | Positive |
| 3 | Sacrum | None | Negative | No | N/A** | Negative | Negative |
| 4 | Clivus | None | Positive | No | N/A** | Negative | Negative |
| 5 | L3–L5 vertebra | Irradiation | Negative | No | N/A** | Negative | Negative |
| 6 | Sacrum | None | Negative | Yes | Positive | Negative | Positive |
| 7* | Sacrum | None | Negative | Yes | Positive | Negative | Positive |
| 8 | Sacrum | None | Negative | Yes | Negative | Negative | Negative |
| 9 | Sacrum | None | Negative | Yes | Negative | Negative | Negative |
| 10 | Sacrum | Surgery | Negative | No | N/A** | Negative | Negative |
TIL tumor infiltrating lymphocyte
TAM tumor associated monocyte
Tumor from which JHC7 cell line was derived
No lymphocytes were seen in tissue section
>5 % tumor cell membranous staining was considered positive
Discussion
The discovery of key genes, such as brachyury, and oncogenic signaling pathways, such as the Akt/PI3 kinase pathway, has improved our understanding of chordoma pathogenesis [6, 21]. The interaction between chordomas and the host immune system, however, remains poorly understood. Many cancers have evolved mechanisms to suppress and evade the host immune system and significant progress has been made in understanding and reversing tumor-mediated immune tolerance [22]. PD-1/PD-L1 blockade is a promising approach that is being actively investigated for clinical translation in a variety of cancers [23]. Whether this pathway plays a role in chordoma pathogenesis and progression, to our knowledge, has not been previously investigated.
The lack of an immunocompetent mouse chordoma model makes the study of any immunological aspect of this tumor type difficult. Patient derived xenografts as well as human chordoma cell lines can grow in immunodeficient mice including NOD/SCID IL2rg (−/−) that lack B, T and natural killer (NK) activity and Nude mice lacking B, T cell function but maintaining intact NK cells. One approach to study chordoma immunology in mice is through the creation of a “humanized” mouse model. In this model, mice are depleted of their mouse bone marrow and human CD34+ hematopoietic stem cells are engrafted in the bone marrow instead, before human chordoma cells are implanted in the mice. This method is complex because of cost, time, issues with histocompatibility antigens, and heterogeneity in the achieved levels of the mouse “humanization” [24].
In this manuscript we study the expression of the PD-1 pathway in human chordomas, a first step that sets the basis for future studies to perform a more in depth analysis of this pathway and its involvement in chordomas. In our cohort, 67 and 50 % of patient samples that contained tumor infiltrating immune cells showed positive staining for PD-L1 and PD-1, respectively. Interestingly, expression of PD-L1 localized to tumor infiltrating immune cells rather than chordoma tumor cells despite chordoma cell lines clearly demonstrating the ability to up-regulate PD-L1 expression in the setting of pro-inflammatory cytokines. These data suggest that chordomas may not be subject to significant baseline immunologic pressure, but have the capacity to implement immunologic escape mechanisms in the setting of an active immune response.
Each of the chordoma cell lines tested exhibited variable expression of PD-L1 and PD-L2. The U-CH1 and U-CH2 cell lines had lower basal expression of both PD-L1 and PD-L2 as compared to JHC7. Similar to previous studies, we found that expression of PD-L1 depended on culture medium [25]. The JHC7 cell line had the highest baseline expression of PD-L1 when cultured in stem cell media. When the cell line was switched to FBS containing media, the baseline and inducible expression of PD-L1 significantly decreased.
Expression of checkpoint molecules, including PD-L1 and PD-L2, is a dynamic process that is highly coordinated with changes in the tumor microenvironment. Expression of PD-L1 is responsive to the presence of Th1 related cytokines (IFN-γ) while expression of PD-L2 can be up-regulated by Th2 related cytokines (IL-4, IL-10, and IL-13). Furthermore, expression of PD-L1 can be induced in both cancer cells and TAMs by activation of the TLR4 or STAT pathways. Expression of PD-L2 is induced on TAMs and is regulated by activation of the IL-4R and STAT pathways (Fig. 1a) [26]. As previously reported in other tumors, we found that local cytokines modulated the surface expression of PD-L1. IFN-γ increased the expression of PD-L1 and PD-L2 in the U-CH1, U-CH2, and JHC-7 cell lines. Interestingly, while expression of PD-L1 or PD-L2 was not up-regulated following exposure to IL-4 in the U-CH2 cell line at the protein level, RT-PCR showed significantly increased transcription of PD-L1 and PD-L2 mRNA following exposure to IL-4. These data indicate that post-transcriptional regulation of PD-L1 and PD-L2 mRNA transcripts may play an important role in modulating activity of the PD-1 pathway.
Glioblastoma, the most aggressive central nervous system tumor has been shown to exhibit high PDL1 expression on tumor cells, TAMs and microglia. Lower grade astrocytic tumors in contrast exhibit much lower PDL1 expression, implying that the more indolent the progression of a tumor the lower the PD-L1 expression is on the tumor cells [27]. Chordomas, are slowly growing tumors that remain undetected by the immune system for many years before they are sizeable enough to have a clinical impact. In this context it does not seem surprising that chordoma tumor cells do not express surface PD-L1. However, the mechanism by which slowly growing tumors escape immune surveillance is not yet known. Myeloid infiltration in the tumor microenvironment and the expression of immune checkpoints such as PD-L1 and PD-1 early on in the disease may be a mechanism by which recruited myeloid cells shut down a T-cell reaction against cancer cells.
Immunohistochemistry primarily localized PD-L1 expression to TAMs. Previous studies have shown that increased density of TAMs correlates with decreased survival in patients with glioblastoma, renal cell carcinoma, and breast cancer, among others [28–30]. Previous studies in cultured glioblastoma cell lines have shown that monocytes exposed to the supernatant of cultured glioma cells up-regulate expression of PD-L1, likely as a result of exposure to IL-10 [14, 30–32]. Whether the release of IL-10 is related to an autocrine or paracrine process, is a topic of ongoing investigation. In addition, future work evaluating the expression of checkpoint molecules on circulating peripheral T cells and monocytes will provide a better understanding of the systemic immune function of patients with chordomas and provide further insight into the immunologic profile of these tumors.
Conclusion
The interaction between chordomas and the host immune system is poorly understood. Here, we show that human-derived chordoma cell lines demonstrate inducible expression of PD-L1 and PD-L2, and that primary chordoma tissue shows variable expression of PD-1 and PD-L1 in infiltrating immune cells. These data are the first to report the presence of PD-1-PD-L1 pathway in chordomas. Future studies are required to understand the role of PD-1 signaling in both tumor infiltrating and peripheral immune cells and the therapeutic potential of PD-1/PD-L1 blockade in chordomas.
Supplementary Material
Fig. 3.
Representative photomicrograph demonstrating the location of PD-1 and PD-L1 expression in the chordoma microenvironment. a PD-L1 expression co-localizing with TILs and TAMs in the same geographic distribution as PD-1 expression. b PD-1 staining on peritumoral and tumor infiltrating lymphocytes c IgG isotype control for PD-L1 staining. ×68 original magnification, all panels
Acknowledgments
The authors would like to thank Ada Tam and Lee Blosser from the Johns Hopkins University Flow Cytometry Core for their guidance on the execution of the flow cytometric analysis. This work was supported by the Chordoma Foundation seed grants to both the laboratories of ML and AQH.
Footnotes
The present work has been presented at the 4th Quadrennial Meeting of the World Federation of Neuro-Oncology (SNO) November 21–24, 2013.
Electronic supplementary material The online version of this article (doi:10.1007/s11060-014-1637-5) contains supplementary material, which is available to authorized users.
Conflicts of interest The authors have no conflict of interest to declare pertaining to the design, execution and submission of this work.
Contributor Information
Dimitrios Mathios, Department of Neurosurgery, The Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Jacob Ruzevick, Department of Neurosurgery, The Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Christopher M. Jackson, Department of Neurosurgery, The Johns Hopkins University School of Medicine, Baltimore, MD, USA
Haiying Xu, Department of Dermatology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Sagar Shah, Department of Biomedical Engineering, The Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Janis M. Taube, Department of Dermatology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA
Peter C. Burger, Department of Pathology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA
Edward F. McCarthy, Department of Pathology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA
Alfredo Quinones-Hinojosa, Department of Neurosurgery, The Johns Hopkins University School of Medicine, Baltimore, MD, USA; Department of Oncology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Drew M. Pardoll, Department of Pathology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA Department of Oncology, The Johns Hopkins University School of Medicine, Baltimore, MD, USA; Department of Medicine, The Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Michael Lim, Email: mlim3@jhmi.edu, Department of Neurosurgery, The Johns Hopkins University School of Medicine, Baltimore, MD, USA; Department of Neurosurgery, Johns Hopkins Hospital, 600 N. Wolfe Street, Neurosurgery - Phipps 123, Baltimore, MD 21287, USA.
References
- 1.Salisbury JR. The pathology of the human notochord. J Pathol. 1993;171:253–255. doi: 10.1002/path.1711710404. doi: 10.1002/path.1711710404. [DOI] [PubMed] [Google Scholar]
- 2.Feigl GC, et al. Evaluation of a new concept for the management of skull base chordomas and chondrosarcomas. J Neurosurg. 2005;102(Suppl):165–170. doi: 10.3171/jns.2005.102.s_supplement.0165. [DOI] [PubMed] [Google Scholar]
- 3.Abdulrauf SI. Decision-making process for the treatment of Intracranial chordomas. World Neurosurg. 2013 doi: 10.1016/j.wneu.2013.07.117. doi: 10.1016/j.wneu.2013.07.117. [DOI] [PubMed] [Google Scholar]
- 4.Siu IM, et al. Erlotinib inhibits growth of a patient-derived chordoma xenograft. PLoS One. 2013;8:e78895. doi: 10.1371/journal.pone.0078895. doi: 10.1371/journal.pone.0078895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.McMaster ML, Goldstein AM, Bromley CM, Ishibe N, Parry DM. Chordoma: incidence and survival patterns in the United States, 1973–1995. Cancer Causes Control. 2001;12:1–11. doi: 10.1023/a:1008947301735. [DOI] [PubMed] [Google Scholar]
- 6.Pillay N, et al. A common single-nucleotide variant in T is strongly associated with chordoma. Nat Genet. 2012;44:1185–1187. doi: 10.1038/ng.2419. doi: 10.1038/ng.2419. [DOI] [PubMed] [Google Scholar]
- 7.Bruderlein S, et al. Molecular characterization of putative chordoma cell lines. Sarcoma. 2010;2010:630129. doi: 10.1155/2010/630129. doi: 10.1155/2010/630129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Tamborini E, et al. Analysis of receptor tyrosine kinases (RTKs) and downstream pathways in chordomas. Neuro-oncology. 2010;12:776–789. doi: 10.1093/neuonc/noq003. doi: 10.1093/neuonc/noq003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Siu IM, et al. Establishment and characterization of a primary human chordoma xenograft model. J Neurosurg. 2012;116:801–809. doi: 10.3171/2011.12.JNS111123. doi: 10.3171/2011.12.JNS111123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Trucco MM, et al. A novel chordoma xenograft allows in vivo drug testing and reveals the importance of NF-kappaB signaling in chordoma biology. PLoS One. 2013;8:e79950. doi: 10.1371/journal.pone.0079950. doi: 10.1371/journal.pone.0079950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Nirschl CJ, Drake CG. Molecular pathways: coexpression of immune checkpoint molecules: signaling pathways and implications for cancer immunotherapy. Clin Cancer Res. 2013;19:4917–4924. doi: 10.1158/1078-0432.CCR-12-1972. doi: 10.1158/1078-0432.CCR-12-1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Akbay EA, et al. Activation of the PD-1 pathway contributes to immune escape in EGFR-driven lung tumors. Cancer Discov. 2013 doi: 10.1158/2159-8290.CD-13-0310. doi: 10.1158/2159-8290.CD-13-0310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bigelow E, et al. Immunohistochemical staining of B7-H1 (PD-L1) on paraffin-embedded slides of pancreatic adenocarcinoma tissue. J Vis Exp. 2013 doi: 10.3791/4059. doi: 10.3791/4059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bloch O, et al. Gliomas promote immunosuppression through induction of B7-H1 expression in tumor-associated macrophages. Clin Cancer Res. 2013;19:3165–3175. doi: 10.1158/1078-0432.CCR-12-3314. doi: 10.1158/1078-0432.CCR-12-3314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fang L, et al. The immune cell infiltrate populating meningiomas is composed of mature, antigen-experienced T and B cells. Neuro-oncology. 2013;15:1479–1490. doi: 10.1093/neuonc/not110. doi: 10.1093/neuonc/not110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Saito H, Kuroda H, Matsunaga T, Osaki T, Ikeguchi M. Increased PD-1 expression on CD4+ and CD8+ T cells is involved in immune evasion in gastric cancer. J Surg Oncol. 2013;107:517–522. doi: 10.1002/jso.23281. doi: 10.1002/jso.23281. [DOI] [PubMed] [Google Scholar]
- 17.Yang W, Song Y, Lu YL, Sun JZ, Wang HW. Increased expression of programmed death (PD)-1 and its ligand PD-L1 correlates with impaired cell-mediated immunity in high-risk human papillomavirus-related cervical intraepithelial neoplasia. Immunology. 2013;139:513–522. doi: 10.1111/imm.12101. doi: 10.1111/imm.12101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hsu W, et al. Generation of chordoma cell line JHC7 and the identification of Brachyury as a novel molecular target. J Neurosurg. 2011;115:760–769. doi: 10.3171/2011.5.JNS11185. doi: 10.3171/2011.5.JNS11185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lipson EJ, et al. Durable cancer regression off-treatment and effective reinduction therapy with an anti-PD-1 antibody. Clin Cancer Res. 2013;19:462–468. doi: 10.1158/1078-0432.CCR-12-2625. doi: 10.1158/1078-0432.CCR-12-2625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Taube JM, et al. Colocalization of inflammatory response with B7-h1 expression in human melanocytic lesions supports an adaptive resistance mechanism of immune escape. Sci Transl Med. 2012;4:127ra137. doi: 10.1126/scitranslmed.3003689. doi: 10.1126/scitranslmed.3003689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.de Castro CV, et al. Tyrosine kinase receptor expression in chordomas: phosphorylated AKT correlates inversely with outcome. Hum Pathol. 2013;44:1747–1755. doi: 10.1016/j.humpath.2012.11.024. doi: 10.1016/j.humpath.2012.11.024. [DOI] [PubMed] [Google Scholar]
- 22.Jackson C, Ruzevick J, Phallen J, Belcaid Z, Lim M. Challenges in immunotherapy presented by the glioblastoma multiforme microenvironment. Clin Dev Immunol. 2011;2011:732413. doi: 10.1155/2011/732413. doi: 10.1155/2011/732413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chen DS, Irving BA, Hodi FS. Molecular pathways: next-generation immunotherapy—inhibiting programmed death-ligand 1 and programmed death-1. Clin Cancer Res. 2012;18:6580–6587. doi: 10.1158/1078-0432.CCR-12-1362. doi: 10.1158/1078-0432.CCR-12-1362. [DOI] [PubMed] [Google Scholar]
- 24.Shultz LD, et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2Rγnull mice engrafted with mobilized human hemopoietic stem cells. J Immunol. 2005;174(10):6477–6489. doi: 10.4049/jimmunol.174.10.6477. doi: 10.4049/jimmunol.174.10.6477. [DOI] [PubMed] [Google Scholar]
- 25.Andorsky DJ, et al. Programmed death ligand 1 is expressed by non-hodgkin lymphomas and inhibits the activity of tumor-associated T cells. Clini Cancer Res. 2011;17:4232–4244. doi: 10.1158/1078-0432.CCR-10-2660. doi: 10.1158/1078-0432.CCR-10-2660. [DOI] [PubMed] [Google Scholar]
- 26.Loke P, Allison JP. PD-L1 and PD-L2 are differentially regulated by Th1 and Th2 cells. Proc Natl Acad Sci USA. 2003;100:5336–5341. doi: 10.1073/pnas.0931259100. doi: 10.1073/pnas.0931259100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Yao Y, et al. B7-H1 is correlated with malignancy-grade gliomas but is not expressed exclusively on tumor stem-like cells. Neuro Oncol. 2009;11(6):757–766. doi: 10.1215/15228517-2009-014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Poli A, et al. Targeting glioblastoma with NK cells and mAb against NG2/CSPG4 prolongs animal survival. Oncotarget. 2013;4:1527–1546. doi: 10.18632/oncotarget.1291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Dannenmann SR, et al. Tumor-associated macrophages subvert T-cell function and correlate with reduced survival in clear cell renal cell carcinoma. Oncoimmunology. 2013;2:e23562. doi: 10.4161/onci.23562. doi: 10.4161/onci.23562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Campbell MJ, et al. The prognostic implications of macrophages expressing proliferating cell nuclear antigen in breast cancer depend on immune context. PLoS One. 2013;8:e79114. doi: 10.1371/journal.pone.0079114. doi: 10.1371/journal.pone.0079114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Rodrigues JC, et al. Normal human monocytes exposed to glioma cells acquire myeloid-derived suppressor cell-like properties. Neuro-oncology. 2010;12:351–365. doi: 10.1093/neuonc/nop023. doi: 10.1093/neuonc/nop023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Dziurzynski K, et al. Glioma-associated cytomegalovirus mediates subversion of the monocyte lineage to a tumor propagating phenotype. Clinical Cancer Res. 2011;17:4642–4649. doi: 10.1158/1078-0432.CCR-11-0414. doi: 10.1158/1078-0432.CCR-11-0414. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.