SUMMARY
Combined BRAF and MEK targeted therapy improves upon BRAF inhibitor (BRAFi) therapy but is still beset by acquired resistance. We show that melanomas acquire resistance to combined BRAF and MEK inhibition by augmenting or combining mechanisms of single-agent BRAFi resistance. These double-drug resistance-associated genetic configurations significantly altered molecular interactions underlying MAPK pathway reactivation. V600EBRAF, expressed at supra-physiological levels because of V600EBRAF ultra-amplification, dimerized with and activated CRAF. In addition, MEK mutants enhanced interaction with over-expressed V600EBRAF via a regulatory interface at R662 of V600EBRAF. Importantly, melanoma cell lines selected for resistance to BRAFi+MEKi, but not those to BRAFi alone, displayed robust drug addiction, providing a potentially exploitable therapeutic opportunity.
INTRODUCTION
RAS and BRAF are frequently mutated in human malignancies. In advanced melanoma, NRAS and, less often, KRAS mutations occur in ~20% of cases and are mutually exclusively with BRAF mutations, which are present in ~50% of cases. Somatic MEK1 or MEK2 mutations, which can be concurrent with RAS or BRAF mutations, have also been detected (Hodis et al., 2012; Krauthammer et al., 2012; Nikolaev et al., 2012; Shi et al., 2012a), but their roles in pathogenesis and therapeutic responses remain ill-defined. BRAF mutations strongly predict responses to ATP-competitive BRAF inhibitors (BRAFi) such as vemurafenib and dabrafenib. Allosteric MEK1 and MEK2 inhibitors (MEKi), such as trametinib, selumetinib, cobimetinib, and binimetinib, may have anti-tumor activities against a broader melanoma segment, including those with NRAS mutations or with both WT NRAS and WT BRAF, but MEKi monotherapy for patients with BRAF mutant melanomas is associated with a narrower therapeutic window (vs. BRAFi) (Ribas and Flaherty, 2011).
Melanoma re-growth after initial response to MEKi has been attributed to a P124LMEK1 mutation (Emery et al., 2009) and acquired MEKi resistance in BRAF mutant colorectal cell lines has been linked to a F129LMEK1 mutation (Wang et al., 2011) or BRAF amplification (Corcoran et al., 2010). How these MEK mutations mechanistically account for MEKi resistance is not entirely clear. Due to the superior clinical benefits of BRAFi for melanoma patients, mechanisms of acquired BRAFi resistance have been studied extensively, and those well-validated clinically include NRAS or KRAS mutations (Nazarian et al., 2010; Shi et al., 2014), V600EBRAF amplification (Shi et al., 2012b) or alternative splicing (Poulikakos et al., 2011; Shi et al., 2012a), MEK1 or MEK2 mutations (Shi et al., 2012a; Wagle et al., 2011), CDKN2A loss (Shi et al., 2014), and genetic alterations in the PI3K-PTEN-AKT pathway (Shi et al., 2014; Van Allen et al., 2014). The convergence of multiple mechanisms to reactivate the MAPK pathway provided a strong rationale for combined BRAF and MEK targeting to overcome BRAFi resistance, a strategy that is supplanting single-agent BRAFi therapy. However, acquired resistance to BRAFi+MEKi still limits the long-term survival of patients with advanced V600E/KBRAF melanoma. A priori, the intransigence of acquired resistance in response to dual MAPK targeting may be due to preferential emergence of MAPK-redundant resistance pathways. Evidence of branched evolution, extensive inter-patient/tumor heterogeneity, and increased tumor fitness as melanoma emerges from BRAFi-imposed evolutionary selection may help explain why the BRAFi+MEKi combinatorial approach is also an “uphill battle” (Shi et al., 2014).
In this study, we investigate the genetic mechanisms of acquired BRAFi+MEKi resistance and elucidate their signaling consequences and therapeutic implications.
RESULTS
Genetic alterations underlying acquired resistance to BRAF/MEK co-targeting in melanoma
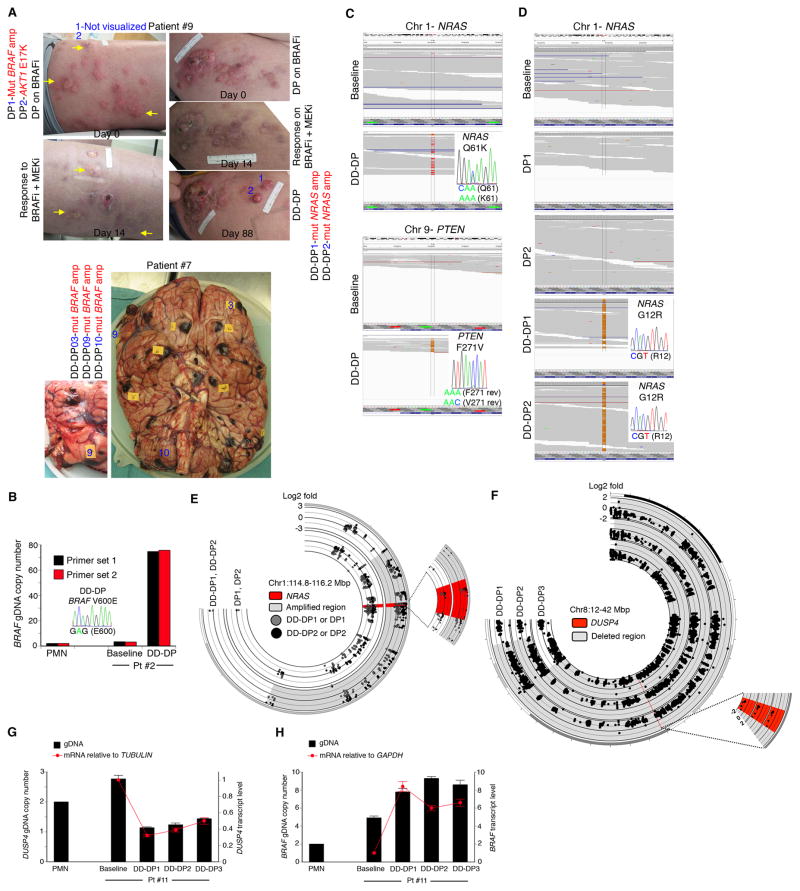
We assembled melanoma tissues with acquired resistance to BRAFi+MEKi (abbreviated as double-drug disease progression or DD-DP) (n=28 DD-DP tumors, each with patient-matched baseline tumors) from patients (n=15) treated under two distinct clinical scenarios (Figure 1A): 1) upfront BRAFi+MEKi (dabrafenib+trametinib or vemurafenib+cobimetinib) in patients (n=10) who were naïve to treatment with either BRAFi or MEKi, and 2) BRAFi+MEKi (vemurafenib+cobimetinib) in patients (n=5) who had previously responded to but progressed on BRAFi (vemurafenib) alone (Table S1). We then analyzed known mechanisms of acquired BRAFi resistance in the MAPK pathway by sequencing the most pertinent exons of BRAF, NRAS, KRAS, MEK1 and MEK2 and performing BRAF copy number analysis (Table S2). Sixteen of 28 DD-DP tumors, along with their patient-matched baseline tumors and normal tissues (n=7), were whole exome-sequenced and analyzed for MAPK and PI3K-PTEN-AKT pathway alterations as reported earlier (Shi et al., 2014) (Table S2). In 19 of 28 (68%) DD-DP tumors, we detected known mechanisms of acquired BRAFi resistance in the two core resistance pathways. These included eight DD-DP tumors harboring V600EBRAF amplification, four harboring NRAS activating mutations, one harboring a KRAS activating mutation, eight harboring CDKN2A deletions, three harboring PTEN loss-of-function (LOF) mutation (a substitution resulting in F127V; Figure S1) or deletions, and one harboring a PIK3R1 deletion. In contrast to the same alterations detected in the context of resistance to BRAFi monotherapy (Shi et al., 2014; Van Allen et al., 2014), those associated with acquired BRAFi+MEKi resistance were notable for augmented gene dosage changes, e.g. V600EBRAF ultra-amplification with 74 or 88 copies (Figure 1B, Table S2), LOF F127VPTEN mutation or homozygous PTEN deletions (Figure 1C), G12RNRAS with selective mutant allele amplification (Figure 1D and 1E), and homozygous CDKN2A deletions (Table S2). There were examples suggesting combinatorial mechanisms, e.g., concurrent heterozygous Q61KNRAS with homozygous CDKN2A deletion and LOF PTEN mutation; V600EBRAF amplification concurrent with homozygous CDKN2A deletion or hemizygous DUSP4 deletion (with related V600EBRAF up-expression and DUSP4 down-expression, Figures 1F to H); and homozygous CDKN2A deletion concurrent with homozygous PTEN deletion and hemizygous PIK3R1 deletion. Thus, genetic analysis of melanomas progressing on BRAFi+MEKi revealed a prevalence of mechanisms of acquired BRAFi resistance, but these genetic alterations often occurred in greater magnitudes or in combinations.
Figure 1. Melanomas Resistant to BRAF/MEK Inhibitors Display Exaggerated Genetic Mechanisms of BRAF Inhibitor Resistance.
(A) Clinical photos denoting specific genetic mechanisms (red) of drug resistance detected within specific tumors (blue). For patient #9, BRAFi-disease progressive melanomas responded to BRAFi+MEKi (yellow arrows) on day 14 with disease progression ensuing as evident on day 88. DP, disease progression on BRAFi; DD-DP, double drug-disease progression.
(B) Quantitative PCR (Q-PCR) and Sanger sequencing of gDNAs extracted from melanoma samples from patient #2 and peripheral mononuclear cells (PMN) as a control. The bar graph shows averages of duplicates.
(C) DD-DP melanoma from patient #6 with concurrent heterozygous Q61KNRAS (exomeSeq) and compound heterozygous of K197*PTEN (not shown) and F271VPTEN (RNASeq). Display by Integrative Genome Viewer with Sanger validation.
(D) G12RNRAS homozygosity in patient #9 DD-DP tumors.
(E) Circos plot showing NRAS copy number gains in patient #9 DD-DP tumors.
(F) Hemizygous DUSP4 deletions in all three DD-DP tumors from patient #11.
(G–H) DUSP4 (G) and BRAF (H) gDNA copy numbers and mRNA expression levels by Q-PCR and Q-RT-PCR, respectively, in tumors from patient #11. Error bars, +/− SD.
BRAFi-resistant melanoma rapidly up-regulates resistance mechanisms individually or combinatorially to overcome BRAF/MEK inhibitors
To further understand acquired BRAFi+MEKi resistance in melanoma underlying the two aforementioned clinical contexts, we generated isogenic human V600EBRAF melanoma cell lines using treatment regimens mimicking each clinical context. In the sequential resistance model, we took those isogenic sub-lines with acquired BRAFi (vemurafenib) resistance (Single-Drug Resistance or SDR), via clinically validated mechanisms such as NRAS mutation (M249R4) (Nazarian et al., 2010), V600EBRAF alternative splicing (M397R) (Shi et al., 2012b) or amplification (M395R) (Shi et al., 2012b), and generated further sub-lines with BRAFi+MEKi (vemurafenib+selumetinib) or Double-Drug Resistance (DDR). In the upfront BRAFi+MEKi resistance model, we took the same set of parental (P), drug-naïve melanoma cell lines and treated them at the outset with BRAFi+MEKi until we generated sub-lines with DDR (Figure 2A). The cell subpopulations were exposed to similar increments of inhibitor concentrations, with the duration at each inhibitor concentration dictated by successful population doubling within 3–4 days. When the time-cumulative doses to reach the full DDR phenotype (defined as 2 μM of BRAFi+MEKi) were compared between these two models, it was clear that the development of DDR from SDR was much faster than DDR directly from parental lines (Figure 2A). This observation is consistent with the hypothesis that preexisting mechanisms of BRAFi resistance could be readily augmented or tuned up to confer resistance to BRAFi+MEKi.
Figure 2. Melanoma Cells with Acquired BRAFi Resistance Further Resist BRAFi+MEKi by Augmenting Existing or Combining Distinct Mechanisms.
(A) Relative drug exposure times required to achieve resistance to BRAFi+MEKi in three isogenic groups of V600EBRAF melanoma cell lines comparing progression from SDR->DDR vs. P->DDR (P, parental, SDR, Single Drug or BRAFi Resistant; DDR, Double Drug Resistant). [inhibitor], 0.1 to 2.0 μM.
(B) gDNA and cDNA BRAF copy numbers (average of duplicates) by Q-PCR or Q-RT-PCR (top) and by semi-quantitative PCR (bottom).
(C) gDNA and cDNA NRAS levels in the M249 P, SDR, and SDR-DDR cell lines in A and B.
(D) Sanger sequencing of cDNAs from cell lines in C with chromatograms showing detection of the WT vs. mutant NRAS transcripts (ratio estimated by peak heights).
(E) Western blot (WB) of indicated total and phospho-protein levels from three isogenic triplets (SDR sub-lines annotated with known BRAFi resistance mechanisms; FL, full-length, TR, truncated; TUBULIN, loading control). Treatments with BRAFi (SDR) or BRAFi+MEKi (DDR) (1 μM), 16 hr prior to lysate preparation. BRAF WB, both short and long exposures shown. Quantification of WBs for NRAS (M249 triplet): 1, 0.98, 1.65; for p61 BRAF (M397 triplet): 1, 2.55, 7.33; and for FL BRAF (M395 triplet): 1, 10.89, 13.63 (normalization to TUBULIN and then parental values).
(F) NRAS knockdown in the M249 SDR and SDR-DDR lines by shRNA as shown by WB 72 hr after lentiviral infections. Inhibitors were at 1 μM each. shSCR, shScrambled.
(G) Three-day MTT assays using M249 cell lines from F. [inhibitor] in μM.
(H) Ten-day clonogenic assays using M249 cell lines from F. BRAFi or BRAFi+MEKi treatments every two days were started 24 hr after plating.
(I) cDNA Sanger sequencing showing WT vs. mutant NRAS transcripts and the estimated ratio in M238 AR (SDR) cells.
(J) Stable knockdown of PTEN by lentiviral shRNA in M238 AR (SDR) (BRAFi, 1 μM) showing the levels of indicated phospho- and total proteins by WB of cellular lysates 72 hr post-transduction, compared to protein levels in the M238 parental cell line (P) treated with DMSO. GAPDH, loading control.
(K) Long-term clonogenic assays of indicated cells from J.
(L) WB showing the DUSP4 protein levels in control and stable knockdown M395 SDR.
(M) Three-day MTT assays of indicated cells from L.
Error bars, +/− SEM; n=5; normalized to DMSO as 100%. BRAFi, vemurafenib; MEKi, selumetinib.
To assess this hypothesis, we examined the SDR vs. SDR-DDR isogenic pairs of cell lines for alterations in the preexisting, defined mechanisms of BRAFi resistance (Figure 2B–E). We showed that the M397 SDR->DDR progression was associated with a dramatically upregulated level of alternatively spliced V600EBRAF mRNA (Figure 2B). Moreover, the M395 SDR->DDR progression resulted in further V600EBRAF amplification along with RNA up-expression. The M249 SDR->DDR progression upregulated mutant NRAS mRNA levels without gDNA copy number gain (Figure 2C–D). Accordingly, at the protein expression level (Figure 2E), M249 SDR-DDR expressed an increased NRAS level; M397 SDR-DDR upregulated the level of a truncated p61 V600EBRAF; and M395 SDR-DDR upregulated V600EBRAF expression further (all relative to isogenic SDR sub-lines). Moreover, full-length V600EBRAF over-expression (in M395 SDR or SDR-DDR) was associated with extensive p-CRAF levels (vs. their parental line). Thus, common mechanisms of acquired BRAFi resistance are highly tunable, by either genetic or non-genetic means, and augmentation or combination of such molecular alterations readily confers resistance to BRAFi+MEKi.
We then tested whether specific examples of gene dosage augmentation or concurrent genetic alterations from the exomic analysis of paired melanoma tissues would augment BRAFi+MEKi resistance in cell line models. Parallel to the mutant NRAS amplifications detected in both DD-DPs of patient #9 (Figures 1D–E), M249 SDR-DDR up-expressed mutant NRAS (albeit via a non-mutational mechanism) (vs. P or SDR) (Figures 2A, 2C–E). NRAS knockdown (Figure 2F) restored BRAFi sensitivity to M249 SDR, as would be expected, but it also strongly restored BRAFi+MEKi sensitivity to M249 SDR-DDR in both short- and long-term (Figure 2G–H) survival assays, indicating that over-expression of mutant NRAS drove DDR. To engineer a DDR cell line mimicking Q61KNRAS heterozygosity+F271VPTEN/K197*PTEN compound heterozygous mutations (DD-DP of patient #6; Figure 1C), we took advantage of the PTEN-expressing, Q61KNRAS-driven M238 SDR sub-line (Figure 2I), which was derived from its V600EBRAF parental line by incremental exposures to increasing doses of BRAFi, and stably introduced shPTEN. We showed that PTEN knockdown in M239 SDR increased the p-AKT level (Figure 2J) and resistance to BRAFi+MEKi (Figure 2K), indicating that each resistance mechanism (Q61KNRAS and PTEN loss) quantitatively contributed to DDR. Moreover, given that V600EBRAF amplification concurred with hemizygous DUSP4 deletion (DD-DP1, 2 of patient #11), we tested whether DUSP4 knockdown (Figure 2L) could confer double-drug resistance to the M395 SDR sub-line, which acquired BRAFi resistance via V600EBRAF amplification. As seen in Figure 2M, M395 SDR was moderately cross-resistant to BRAFi+MEKi treatments, but loss of DUSP4 expression augmented DDR.
Clonal analysis detects alternative genetic configurations in BRAFi+MEKi resistance associated with MAPK reactivation
Previous results indicate that, once sub-clones with specific BRAFi resistance mechanisms have attained clonal dominance, overcoming BRAFi resistance with the added MEKi is at best an “uphill battle.” We then sought to understand the underlying mechanism(s) of resistance to upfront BRAFi+MEKi (vemurafenib+selumetinib). A polyclonal DDR sub-line derived from M249 harbored both mutant BRAF ultra-amplification and a MEK1 mutation (F129L) (data not shown). F129LMEK1 had previously been uncovered in a colorectal sub-line bred to acquire selumetinib resistance (Wang et al., 2011). To understand the individual contributions of V600EBRAF amplification and MEK1 mutation to the DDR phenotype, we re-treated the M249 P with increments of BRAFi+MEKi but derived two single cell-derived M249 DDR sub-clones, DDR4 and DDR5. In contrast to M249 P, both DDR4 and DDR5 were highly resistant to the growth-inhibitory effect of BRAFi+MEKi in 3-day MTT assays (Figure 3A). In fact, the apparent “growth-stimulation” of DDR4 and DDR5 by BRAFi+MEKi treatment was due to a relative loss of their viability in the absence of optimal concentrations of the inhibitors. This “drug addiction” phenomenon was even more profound in long-term clonogenic assays (see Figure 4). SCH772984, an ERKi and an analog of which is being tested clinically, was inefficient to inhibit the growth of DDR4 or DDR5 by itself but was highly active against M249 P (Figure 3A). In fact, low concentrations of SCH772984 rescued DDR4 and DDR5 from drug addiction, suggesting that sub-optimal ERKi dosing to overcome DDR may paradoxically perpetuate DDR fitness. In contrast, ERKi restored BRAFi+MEKi sensitivity to DDR4 and DDR5, consistent with MAPK pathway reactivation as the major mechanism of acquired resistance to upfront BRAFi+MEKi. This was corroborated by analyzing the MAPK pathway status (pERK levels) in the M249 triplet (Figure 3B). After plating for 16 hr without both inhibitors, the triplet cell lines were treated with BRAFi+MEKi (1 hr) at increasing concentrations (Figure 3B) or with BRAFi+MEKi (1 μM) for increasing durations (up to 72 hr) (Figure S2A). Western blot analysis showed that DDR4 and DDR5, compared to M249 P, displayed higher baseline and inhibitor-treated p-ERK levels as well as faster p-ERK recovery in the continued presence of BRAFi+MEKi. Monitoring further upstream for p-MEK and downstream for p-RSK (Thr573) levels revealed a similarly rapid recovery of the MAPK pathway (Figure S2B).
Figure 3. Melanoma Cells Clonally Develop Resistance to Upfront BRAFi+MEKi via Alternative Genetic Configurations.
(A) Three-day MTT assays (error bars, +/− SEM, n=5; top, relative raw values; bottom, normalized to DMSO vehicle as 100%). Cells were plated 16 hr without inhibitors prior to treatment with indicated inhibitor(s) (in μM).
(B) Western blot (WB) of indicated total and phospho-proteins. M249 cell lines were plated 16 hr without inhibitors prior to BRAFi+MEKi treatments for 1 hr (0–10 μM in 10-fold increments). TUBULIN, loading control.
(C) BRAF copy number by gDNA Q-PCR (averages of duplicates).
(D) Sanger sequencing showing BRAF and MEK1 mutational status of M249 cell lines.
(E) Integrated Genome View snapshots of reference and mutant/variant allelic frequencies (MAFs) centered on the A to T mutation (chromsome 7:140453136; V600EBRAF) and on the C to G mutation (chromosome 15:66729179; F129LMEK1) in indicated M249 cell lines. Mutant:WT estimated from the MAFs. Note a low MAF of F129LMEK1 in M249 P.
(F) CNV display by Circos (with respect to M249 P) showing distinct BRAF amplicons in DDR4 vs. DDR5 (top) and MEK1 copy number gain in DDR5 (bottom).
(G–H) WB of indicated total and phospho-proteins from M249 cell lines plated 16 hr without inhibitors prior to ERKi treatments for 1 hr (0–10 μM) without or with BRAFi+MEKi co-treatment (1 μM) (G) or prior to BRAFi+MEKi treatment (1 μM, 1 hr) (H).
BRAFi, vemurafenib; MEKi, selumetinib; ERKi, SCH772984. See also Figure S2.
Figure 4. Achieving BRAF/MEK Inhibitor Resistance via Tuning V600EBRAF Gene Dosage with or without MEK Mutations.
(A) M249 DDR4 and DDR5 plated 16 hr with BRAFi+MEKi (1 μM each), transduced with lentiviral shVector or shBRAF for 48 hr, and treated with (+) or without (−) inhibitors at 1 μM (1 hr) were analyzed by Western blot (WB). TUBULIN, loading control.
(B) WB for CRAF or BRAF in M249 triplet 48 hr after without (−) or with (+) CRAF or BRAF knockdown, as indicated.
(C) Cells from B plated for clonogenic assays.
(D) Whole exome-based phylogenetic relationships of the M249 triplet cell lines. Branch lengths proportional to the number of heterozygous (het) single nucleotide variants (SNVs) and small insertion-deletions (INDELs) private to each cell line with respect to the theoretical common ancestral cell sub-population (#1). The DDR-unique copy number variations of indicated genes also shown.
(E, F) WB of total and phospho-protein levels in M249 triplet and M249 P engineered to express V600EBRAF and F129LMEK1 (E) or Q56PMEK1 (F). Selected cell lines treated with BRAFi+MEKi (1 μM) for 16 hr and then washed free of inhibitors for 8 hr.
(G) WB analysis of M249 P engineered to express vector, WTBRAF, or V600E/R509HBRAF (without inhibitors) or M249 DDR4 (BRAFi+MEKi, 1 μM, 16 hr).
(H) Clonogenic assays of M249 P engineered to express the indicated levels of WT vs. mutant BRAF and/or MEK1 and their relative resistance to BRAFi+MEKi over inhibitor concentrations and time.
(I) Temporal genetic clonal evolution of MAPKi resistance with magnitudes matching graded selective pressures and with augmented gene dosage vs. combinatorial genetic alterations proposed as distinct pathways. Distinct V600EBRAF amplicons indicative of convergent evolution. Each circle, dominant sub-clone.
BRAFi, vemurafenib; MEKi, selumetinib. See also Figures S3 and Table S3.
Consistent with the BRAF protein levels (Figure 3B), we found that DDR4 harbored V600EBRAF ultra-amplification (47.4 fold or >160 copies), while DDR5 harbored low copy-number V600EBRAF gain (4.6 fold or 20 copies) along with F129LMEK1 (Figures 3C and 3D). V600EBRAF copy number gains quantified by gDNA Q-PCR were corroborated by Sanger sequencing, which showed a BRAF mutant to WT ratio of 2:1 in the parental line (or about 3 BRAF copies) and apparent V600EBRAF homozygosity in DDR4 and DDR5 resulting from selective V600EBRAF amplification (Figure 3D). Moreover, whole exome sequence (WES) analysis of the M249 triplet cell lines confirmed that the DDR-associated altered mutant/variant allelic frequencies (MAFs) of V600EBRAF and F129LMEK1 (Figure 3E) were likely due to mutant allele-selective copy number gains (Figure 3F). WES analysis also detected a low F129LMEK1 MAF (4%) in the parental polyclonal line, suggesting preexistence of this drug-resistant sub-clone (Figure 3E). In addition, copy number variation (CNV) analysis revealed distinct BRAF amplicons in M249 DDR4 vs. DDR5, suggesting convergent evolution (Figure 3F, top). Since the concurrence of V600EBRAF amplification and F129LMEK1 in M249 DDR5 could be selected by distinct inhibitor concentrations, we derived two additional M249 DDR sub-lines (M249 DDR2 and M249 DDR3) by treatments from the outset with a higher concentration of BRAFi+MEKi (0.5 μM). In a pattern suggestive of convergent evolution, both M249 DDR2 and DDR3 displayed low-copy number gains of both V600EBRAF and F129LMEK1 (Figures S2C–E).
Using a WTBRAF cell line, human 293T, we then tested the impact of MEK mutants associated with MAPKi resistance on cellular substrate levels (i.e., pERK) and the p-ERK IC50 of MEKi (Figures S2F–G). We over-expressed F129LMEK1, C121SMEK1 (which confers BRAFi resistance) (Wagle et al., 2011) and several MEK mutants (Q56PMEK1, K59delMEK1, and E203KMEK1) associated with clinical resistance to MAPK targeting and compared their impacts on baseline pERK levels as well as the sensitivities of p-ERK to MEKi (selumetinib). Although over-expression of these MEK1 mutants (vs. WTMEK1) variably increased the baseline p-ERK level, their cellular p-ERK IC50 to MEKi did not differ appreciably, arguing against allosteric MEKi binding defect as the shared mechanism of action of MEK mutants. Their concurrence with V600EBRAF amplification argues for a possible cooperative biochemical mechanism of resistance.
To further understand the impact of ERKi on survival of DDR4 and DDR5 cells (Figure 3A), we withdrew DDR4 and DDR5 (16 hr off) from BRAFi+MEKi and then treated them with either ERKi (1 hr) alone or BRAFi+MEKi+ERKi (1 hr) (Figure 3G). ERKi alone was ineffective at suppressing the p-ERK rebound following double-drug withdrawal (Figure 3G). However, once BRAFi+MEKi were re-introduced, additional treatment with ERKi was highly effective in suppressing the p-ERK levels (Figure 3G). Thus, ERKi treatment alone of some melanoma cells previously selected for resistance by BRAFi+MEKi would be ineffective unless very high ERKi doses were delivered, which is unlikely achievable clinically. Thus, clonal M249 DDR4 and DDR5 melanoma sub-lines harbor salient but distinct genetic alterations that represent tunable and combinatorial modes of resistance to BRAFi+MEKi reversible by combining ERKi.
Distinct mechanisms of resistance driven by V600EBRAF ultra-amplification or V600EBRAF amplification+F129LMEK1
Earlier, we noted a robust upregulation of p-CRAF in the M395 SDR and SDR-DDR sub-lines that harbor V600EBRAF amplification (Figure 2E). Hence, we probed the p-CRAF levels in the M249 triplet lines. DDR4 and DDR5, freshly treated with BRAFi+MEKi (1 h), displayed robust elevated p-CRAF levels (DDR4 > DDR5 ≫ P; Figure 3H). Up-regulated p-CRAF levels in DDR4 and DDR5 did not require the continued presence of both inhibitors, as their withdrawal for up to 20 hours after an overnight (16 hr) treatment did not diminish the p-CRAF levels (Figure S2H). We hypothesized that this strong CRAF up-regulation in DDR4 (and a weaker one in DDR5) may be driven by supra-physiologic V600EBRAF over-expression, the degree of which positively correlated with that of p-CRAF up-regulation (Figures 3B, 3H and 4A). To test this hypothesis, we knocked down BRAF levels in DDR4 and DDR5, with or without BRAFi+MEKi, and found that BRAF knockdown effectively down-regulated p-CRAF levels (Figure 4A). BRAF knockdown also reduced p-CRAF levels in the V600EBRAF-amplified M395 SDR-DDR sub-line (Figure 2B; Figure S3A). We also knocked down CRAF directly (Figure 4B) and tested the individual contributions of BRAF vs. CRAF to the clonogenic (i.e., long-term) growth/survival of the M249 triplet (Figure 4C). As expected, M249 P growth/survival was not sensitive to CRAF knockdown but highly sensitive to BRAF knockdown. Consistent with prior short-term assays (Figure 3A), both M249 DDR4 and DDR5 displayed dramatic “drug addiction.” Importantly, in the presence of both inhibitors, the growth/survival of DDR4 and DDR5 was highly dependent on either CRAF or BRAF, suggesting functional and physical interaction.
To assess whether there are likely additional genetic underpinnings of p-CRAF upregulation (and DDR) in DDR4 and DDR5, we analyzed the phylogenetic relationship of the M249 triplet (Figure 4D) and assessed the genetic alterations shared by DDR4 and DDR5 (Table S3). From this WES-based phylogeny, it was apparent that DDR4 and DDR5 single-cell clones represent minor sub-clones in the parental, polyclonal population, since they each harbors a large number of private mutations, which escaped detection in the mixed parental population. In fact, the number of shared genetic alterations between DDR4 and DDR5 was exceedingly small (Table S3), suggesting that these few alterations (aside from V600EBRAF amplification) were unlikely drivers of DDR. As the M249 P majority population does not harbor the DDR4 or DDR5-private mutations, we reasoned that the ability of salient genetic feature shared by DDR2, DDR3, DDR4, and DDR5 (V600EBRAF amplification) and by DDR2, DDR3, and DDR5 (F129LMEK1 and its low copy number gain) to reconstitute DDR (and their biochemical features) would establish sufficiency (in light of necessity established earlier).
We then directly tested whether supra-physiologic V600EBRAF over-expression, mimicking the DDR4 and DDR5 levels, would be sufficient to up-regulate p-CRAF levels. We engineered the M249 P to express stably and homogeneously the empty vector, F129LMEK1 (Figure 4E) or Q56PMEK1 (Figure 4F) (identified in clinical MEKi (Villanueva et al., 2013) or BRAFi+MEKi (Wagle et al., 2014) resistance), V600EBRAF high over-expression, and V600EBRAF low over-expression concurrent with a MEK1 mutation. Regardless of double-drug treatment (16 hr) or subsequent withdrawal (8 hr), V600EBRAF high over-expression induced a robust DDR4-like p-CRAF level, while V600EBRAF low over-expression concurrent with a MEK1 mutation induced a lower, DDR5-like p-CRAF level. Neither vector control nor MUTMEK1 alone had any impact on the p-CRAF level. Also, supra-physiologic expression of WTBRAF or V600E/R509HBRAF (known to disrupt BRAF-CRAF dimerization) in M249 P only marginally up-regulated p-CRAF (Figure 4G). However, the M249 P engineered cell lines (vs. the spontaneously resistant DDR4 and DDR5 sub-lines), displayed a slower pERK recovery (with or without BRAFi+MEKi; Figures 4E–F). This difference (a few hours) was minimal compared to the extremely slow p-ERK recovery observed in the parental line (not detectable by 3 days; Figure S2A) and appeared to be due to prior MAPKi exposure or pre-conditioning, which abolished the small difference in the p-ERK recovery rate between the M249 P engineered lines vs. DDR4 and DDR5 (Figure S3B).
We then assessed the relative potencies of individual alterations observed in M249 DDR4 and DDR5 to confer BRAFi+MEKi resistance in M249 P using both short- (Figures S3C–D) and long-term (Figures 4H) survival assays. V600EBRAF high over-expression or V600EBRAF low over-expression concurrent with a MEK1 mutant (F129L or Q56P) conferred more than one-log (short-term) or two-log (long-term) increases in MAPKi resistance. Interestingly, pre-conditioning of the engineered M249 lines conferred double-drug addiction (Figure S3E–F). As was noted previously for DDR4 and DDR5 sub-lines (Figures 3A, 4C), the double-drug addiction phenotype also exaggerated the apparent double-drug resistance phenotype of pre-conditioned M249 P engineered with each genetic configuration (Figure S3D). These data together (Figures 4E–F; Figure S3B–F) thus suggest a mechanistic link between double-drug addiction and p-ERK rebound (see below in Figure 7). Moreover, supra-physiologically expressed V600E/R509HBRAF, defective in p-CRAF induction (Figure 4G), was also compromised in its ability to resist repeated treatments with BRAFi+MEKi (1 μM, 24 days). V600EBRAF low over-expression or MEK1 mutation alone was individually able to confer BRAFi+MEKi resistance but only to an extent appreciably weaker than achieved by their combination (see growth at 0.1 μM vs. 1.0 μM of drugs at days 15 and 24) (Figure 4H). The combinatorial effects of over-expressed V600EBRAF and MEK1 mutants on promoting the double-drug resistance phenotype could also be observed in a different cell line (Figure S3 G–I). Thus, ERKi-sensitive, acquired resistance to BRAFi+MEKi observed in DDR4 and DDR5 is causally attributable to either supra-physiologic over-expression of V600EBRAF or a lower degree of V600EBRAF and MUTMEK over-expression (Figure 4I; Figure S3). Mechanistically, excess V600EBRAF proteins promote dimerization with CRAF and CRAF activation/dependency.
Figure 7. Resistance to Combined BRAF/MEK Inhibition Results in Exquisite Drug Addiction.
(A) Clonogenic survival of M249 DDR cell lines plated in BRAFi+MEKi, 1 μM each, for 72 hr then cultured for 9 days with or without specific inhibitor withdrawal (representative of three independent repeats).
(B, C) Clonogenic/drug addiction assays as in (A) except for the indicated high (B) or low (C) ERKi doses starting at 72 hr after plating.
(D) Clonogenic assays comparing SDR vs. DDR cell lines of distinct genetic backgrounds and resistance mechanisms (amp, amplification; spl, splicing).
(E) Western blot analysis of p-ERK levels without or with acute BRAFi (SDR) or BRAFi+MEKi (DDR) withdrawal for 4 and 24 hr. TUBULIN, loading control. Quantification of p-ERK signals normalized to TUBULIN levels is shown for each cell line relative to the baseline signals (no inhibitor withdrawal).
(F) Correlation between changes in p-ERK levels (E, 4 hr vs. 0 hr) and in clonogenic growths (D) upon inhibitor(s) withdrawal.
BRAFi, vemurafenib; MEKi, selumetinib; ERKi, SCH772984. See also Figures S6 and Table S4.
Next, to dissect mechanistically how MUTMEK1 aids over-expressed V600EBRAF in establishing a full double-drug resistance phenotype, we posited that over-expressed V600EBRAF and MUTMEK physically and functionally interact in a complex facilitated by (i) the MEK mutant conformation, and (ii) a kinase-independent regulatory role of V600EBRAF. This complex facilitates MEK phosphorylation/activation by CRAF, akin to a modeled MEK-KSR2-BRAF regulatory complex (Brennan et al., 2011). Hence, we tested whether F129LMEK1 in DDR5 (but not DDR4) would be more abundantly associated physically with V600EBRAF. Accordingly, we immunoprecipitated BRAF in the M249 triplet and probed for MEK1 and MEK2 in the immunoprecipitates. Consistently, much more MEK1 and MEK2 were detected in complex with BRAF in F129LMEK1-harboring DDR5 (Figure 5A). We then specifically immunoprecipitated MEK1 and detected a dramatically higher BRAF level bound to MEK1 in DDR5 (Figure 5B). However, the pattern of BRAF-MEK2 binding was reversed; we detected more BRAF bound to MEK2 in DDR4 (Figure 5C). MEK2 in DDR4 was also associated with the highest level of activation-associated phosphorylation at Ser226, consistent with MEK2 recruitment to and activation by a BRAF-containing complex. Under the same conditions, we were unable to detect CRAF or KSR2 in BRAF, MEK1, or MEK2 immunoprecipitates (data not shown). These data suggest that the supra-physiologic level of V600EBRAF in DDR4 recruits both WTMEK1 and WTMEK2 whereas the V600EBRAF level over-expressed to a lesser extent in DDR5 recruits F129LMEK1 preferentially over WTMEK2.
Figure 5. Distinct MEK Mutants Share Enhanced Interaction with V600EBRAF.
(A–C) The M249 triplet cell lines were plated without (P) or with (DDR4 and DDR5) BRAFi+MEKi (1 μM, 16 hr), and lysates were subjected to immunoprecipitation (IP) using a control antibody (IgG) or BRAF− (A), MEK1- (B), or MEK2- (C) specific antibodies. The IP and total fractions were probed by Western blot (WB). TUBULIN, loading control.
(D) WB analysis of total and phospho-MEK1/MEK2 and -ERK levels in the M249 triplet cell lines.
(E) Structure of MEK1 (two views, 180° rotated) with the locations of MEK1 mutations, or residues homologous to MEK2 mutations, indicated in yellow, except that Q56 and F129 are indicated in green. All mutations, except I111S and P124S, have been detected in melanomas with clinical acquired MAPKi resistance.
(F) M249 P engineered to express vector or FLAG-WTMEK1, -F129LMEK1, -Q56PMEK1 concurrent with over-expression of either HA-WTBRAF or HA-V600EBRAF were plated with BRAFi+MEKi (1 μM, 16 hr; except vector control), and the lysates were subjected to IP (anti-IgG or -FLAG). WB of IP and total fractions.
BRAFi, vemurafenib; MEKi, selumetinib. See also Figure S4 and Movie S1.
We also assessed the relative phosphorylation status of MEK1 and MEK2 in DDR4 and DDR5 16 hr after treatment with BRAFi+MEKi vs. M249 P treated with DMSO. Interestingly, we observed that only DDR4, but not DDR5, harbored an enhanced level of activation-associated MEK1 and MEK2 phosphorylation (Figure 5D). In both DDR4 and DDR5, MEK1 displayed increased levels of ERK-dependent negative feedback phosphorylation on Thr291 within its proline-rich region of the kinase domain, which is not present on MEK2, suggesting that the time-cumulative ERK activities are far greater in DDR4 and DDR5 despite BRAFi+MEKi treatment than in parental M249. DDR5 harbored the highest level of p-MEK1 Thr291, which has been shown to reduce MEK1-MEK2 hetero-dimerization and MEK2 Ser226 phosphorylation (Catalanotti et al., 2009) and may also explain the reduced p-MEK1 Ser222 level (Figures 5D).
We then sought to reconstitute F129LMEK1-V600EBRAF interaction and its functional role in DDR. We had observed that the majority of MEK1 and MEK2 mutations thus far detected specifically in melanomas with clinically acquired BRAFi, MEKi, or BRAFi+MEKi resistance (Emery et al., 2009; Shi et al., 2014; Van Allen et al., 2014; Villanueva et al., 2013; Wagle et al., 2011; Wagle et al., 2014) cluster three-dimensionally in or proximal to helix A and C (Figure 5E; Movie S1). Specifically, in M249 P, we minimally over-expressed a series of FLAG-tagged MEK1 constructs and co-expressed either HA-tagged WTBRAF or V600EBRAF, both at high levels akin to DDR5 (Figure 5F; Figure S4). We then immunoprecipitated protein complexes via FLAG and detected MEK1, BRAF and HA levels. Importantly, both F129LMEK1 and Q56PMEK1, which is homolog to Q60PMEK2, displayed dramatically enhanced and preferential interaction with over-expressed V600EBRAF relative to WTMEK1. Anti-BRAF signals detected in the FLAG-immunoprecipitates presumably contained both endogenous and exogenous V600EBRAF. Thus, these data support the notion that BRAFi+MEKi treatment in melanoma selects for MEK1 or MEK2 mutations that impact a discrete structural sub-domain and leads to a conformation favoring physical association with over-expressed V600EBRAF.
To assess the functional relevance of a V600EBRAF-MUTMEK complex, we searched for clues of a BRAF-MEK physical interaction interface (Figure S5). Based on prior structural data of MEK1-BRAF (Haling et al., 2014), vemurafenib-bound V600EBRAF (Bollag et al., 2010), and MEK1-KSR2 (Brennan et al., 2011) and structural alignments of vemurafenib-bound V600EBRAF with BRAF or KSR2, we hypothesized a regulatory V600EBRAF-MUTMEK complex where V600EBRAF arginine 662 makes critical contacts with MEK residues in one complex interface (Figure 6A–B). We predicted that the R662L substitution in V600EBRAF would disrupt this face-to-face V600EBRAF-MUTMEK interaction and attenuate the DDR phenotype. Ectopic expression of vector, HA-WTBRAF, HA-V600EBRAF and HA-V600E/R662LBRAF in WTBRAF human 293T cells revealed that the R662L substitution did not interfere with the V600EBRAF kinase activation status in the absence of MAPKi (Figure 6C). We then engineered M249 P to stably express a FLAG-F129LMEK1 or FLAG-Q56PMEK1 along with HA-tagged WT or various mutant BRAF at levels akin to M249 DDR5 (Figure 6D). After BRAFi+MEKi treatment (1 μM, 16 hr), anti-FLAG immunoprecipitation followed by Western blots revealed that both MEK1 mutants most abundantly interacted with V600EBRAF, consistent with previous results (Figure 5F). Importantly, the R662L mutation in the context of V600EBRAF strongly abolished this enhanced V600EBRAF-MUTMEK1 complex and reduced the overall p-ERK levels. V600E/R509HBRAF also appeared to display reduced interaction with MUTMEK1 but without a reduction in the p-ERK levels, suggesting that this apparent reduction was due to loss of BRAF dimers (Figure 6A) (Haling et al., 2014) or higher-order oligomers (Nan et al., 2013) brought down by anti-FLAG. Consistently, whereas engineered M249 P lines highly over-expressing V600EBRAF or minimally over-expressing V600E/R509HBRAF together with a MEK1 mutant were able to resist robustly BRAFi+MEKi at 1 μM, those cell lines expressing V600E/R662LBRAF or WTBRAF along with a MEK1 mutant grew poorly over 28 or 32 day treatments with BRAFi+MEKi (Figure 6E). Taken together, these studies (Figures 4–6; Figure S3–S5) highlighted a critical role of upstream MAPK reactivation, i.e., upregulation of the V600EBRAF-CRAF-MEK complex, in the MAPKi resistance phenotype. Buildup of this plastic complex is dependent on the degree of BRAF and/or MEK inhibition and likely other cell context determinants. In the extreme case of DDR, alternative mechanisms to upregulate this complex can be achieved by V600EBRAF (variably over-expressed) interacting with WTCRAF or with MUTMEK.
Figure 6. A BRAF-MEK Interface Critical for V600EBRAF-MUTMEK1 Interaction and Cooperative Double Drug Resistance.
(A) A predicted MEK1 kinase domain (KD)-V600EBRAF KD complex with yellow color highlighting the locations of (i) MEK1 residues mutated in melanomas with acquired MAPKi resistance, (ii) V600EBRAF R509, critical for RAF-RAF dimerization, (iii) V600EBRAF R662, structurally homologous to KSR2 A879 critical for MEK1-KSR2 interaction, and (iv) V600EBRAF I617, critical for MEK-BRAF dimerization.
(B) Zoomed-in details of a MEK1-V600EBRAF interfaces, highlighting MEK1 activation segment residues (blue, M219, S222, and V224) interacting with V600EBRAF R662 (yellow), I617 (magenta) and I666 (magenta) and interactions predicted to be abolished by a R662L (red) mutation.
(C) Western blot of indicated proteins in human 293T cells transfected with vector or indicated HA-tagged BRAF constructs. TUBULIN, loading control.
(D) M249 P engineered to moderately over-express HA-BRAF or the indicated BRAF mutants along with either FLAG-MEK1 mutant (F129L or Q56P). Experiments were performed as described for Figure 5F.
(E) Clonogenic assays of M249 P engineered to express WT or indicated mutant BRAF, MEK1 mutants, and/or their empty vectors (Vec). Relative resistance to BRAFi+MEKi assessed over the indicated concentration range and time points. Three repeats (for 0.1 and 1.0 μM) are shown for the longest time points (28 and 32 days), and growths were quantified (1 μM; n=3; normalization relative to V600EBRAF+MUTMEK1 transduced cells as 100%; means and error bars, +/− SD; *p < 0.05, ***p < 0.001, ns, not significant based on ANOVA). BRAFi, vemurafenib; MEKi, selumetinib.
See also Figure S5.
Melanoma cells with acquired resistance to BRAFi+MEKi display exquisite dual drug addiction
It has been reported recently that patient-derived xenografts with acquired resistance to BRAFi driven by V600EBRAF amplification or RNA over-expression could potentially be counter-selected during periods of BRAFi withdrawal (Thakur et al., 2013). We thus tested the degree to which M249 DDR4 and DDR5 were addicted to each (BRAFi or MEKi) or both (BRAFi+MEKi) inhibitors during long-term clonogenic growth. Three days after seeding, DDR4 and DDR5 cells were kept continuously on both inhibitors, washed from both, or replenished with only one of the two inhibitors. Both DDR4 and DDR4 were strongly addicted to continuous treatment with BRAFi+MEKi (Figure 7A). The loss of viability after acute BRAFi+MEKi washout could not be rescued by a dose of ERKi (1 μM) sufficient to strongly suppress the rebound in p-ERK resulting from drug withdrawal (Figure 7B; Figure S6A). Additionally, this “high” dose of ERKi could re-sensitize DDR4 and DDR5 to either BRAFi or BRAFi+MEKi, consistent with prior short-term MTT results (Figure 3A). Notably, the anti-growth/survival effect of double-drug withdrawal was comparable to that of ERKi alone or ERKi plus BRAFi (Figure 7B). However, ERKi at a sub-optimal dose (0.1 μM), which could suppress the rebound p-ERK levels induced by acute double-drug withdrawal, Figure S6B), completely rescued the anti-growth/survival effects of BRAFi+MEKi withdrawal and partially “erased” the anti- growth/survival effects of single BRAFi or MEKi withdrawal (Figure 7C). Importantly, a sub-optimal dose of ERKi could be anti-growth/survival only if DDR4 and DDR5 were continuously treated with BRAFi+MEKi. We then sought evidence consistent with melanoma regression in patients who have been discontinued on MAPK-targeted therapies due to disease progression or acquired drug resistance. From evaluable patients with melanoma who were treated with BRAFi+MEKi (n=15) or single-agent BRAFi (n=16) therapies (Table S4), we retrospectively collated radiologic images before and/or during disease progression and compared them to images, when available or feasible, after a variable time off therapies (Figure S6 C–D). Although specific clinical examples of tumor regression after cessation of BRAFi+MEKi therapy could be identified, overall disease stabilization or uniform tumor regression leading clinical remission could not be achieved. Moreover, only cases of tumor growth deceleration could be observed for melanomas after cessation of single-agent BRAFi therapy. Thus, the drug addiction phenotype can be readily elicited in DDR cell lines only if MAPK inhibition were reversed acutely and completely, and additional factors may modulate or mitigate this phenotype clinically.
Given the strong degree of double-drug addiction noted with both DDR4 and DDR5, we asked whether this would be generalizable across different cellular contexts and to melanoma cells with acquired resistance to BRAFi treatment alone. Interestingly, we found that melanoma cell lines adapted to growth with BRAFi+MEKi far more consistently displayed drug addiction (Figure 7D). Also consistent was the observation that melanoma cell lines with DDR displayed a greater rebound in p-ERK levels after drug washout (Figure 7E). This greater rebound was not necessarily due to the maximal p-ERK levels upon withdrawal of drugs but rather due to the very low p-ERK levels in the presence of both drugs (i.e., stronger on-target pathway suppression). Quantification of the fold changes in p-ERK levels (Figure 7E) and in clonogenic growths (Figure 7D) showed that they are strongly negatively correlated. Thus, melanoma cells with DDR displayed a stronger rebound in p-ERK levels and drug addiction upon drug withdrawal, when compared to melanoma cells with single-drug resistance withdrawn from BRAFi (Figure 7F). This p-ERK rebound is indicative of drug addiction since a sub-optimal dose of ERKi could rescue cells from double-drug withdrawal-induced loss of fitness (Figure 7C).
DISCUSSION
The understanding of how BRAF mutant melanomas frequently acquire BRAFi resistance via several distinct mechanisms, which thematically reactivate the MAPK pathway, has provided foundational rationale to combined BRAF/MEK inhibition to suppress such mechanisms. The ensuing translational effort has led to this combination supplanting BRAFi monotherapy in the clinic. This study of genetic alterations in melanomas with acquired BRAFi+MEKi resistance has provided unexpected insights. First, we detected alterations affecting similar genes known to be responsible for acquired BRAFi resistance, which suggests that the gene dosage or concurrence of these mutations may impart altered molecular interactions promoting BRAFi+MEKi resistance. The exaggerated genetic configurations encompassed GOF (e.g., V600EBRAF ultra-amplification, G12RNRAS amplification) and LOF (e.g. F127VPTEN, deletions affecting PTEN, CDKN2A, DUSP4) alterations, and their combinations. Second, focusing on MAPK-reactivation, we uncovered a highly plastic or tunable RAF-MEK complex resulting from mutations (SNVs and/or CNVs). For instance, supra-physiologic levels of V600EBRAF allosterically relay oncogenic MAPK signaling via back-to-back interactions with CRAF. Moreover, moderately over-expressed levels of V600EBRAF likely regulate MEK1 and MEK2 activation via a face-to-face complex. These altered molecular interactions underscore an intrinsic limitation of combined BRAF and MEK inhibition and predict potential limitations of further downstream inhibitors (e.g., ERKi) in overcoming acquired BRAFi+MEKi resistance.
Thus, we have shown how (1) quantitative genetic alterations or gene dosage impact qualitative modes of signaling, and (2) combinatorial alterations might be selected to impact survival signaling cooperatively (Figure 8). MEK1 and MEK2 mutants with alterations residing in or proximal to the helix A/C sub-structure share an increased ability to form an activation-associated complex with V600EBRAF, especially when both BRAF and MEK mutants are moderately over-expressed. Moreover, a proposed MUTMEK-V600EBRAF hetero-dimer interface strongly suggests that such a face-to-face physical interaction involves predominantly a kinase-independent or regulatory function of V600EBRAF. Together, these data indicate a V600EBRAF-CRAF-MEK1/MEK2 signaling loop that is highly susceptible to up-regulation via single or multiple convergent genetic (and likely non-genetic) alterations.
Figure 8. Alterations in a V600EBRAF-CRAF-MEK Complex with Opposite Impacts on Melanoma Fitness Contingent on the Presence of BRAF and MEK Inhibitors.
Alternative configurations of a RAF-MEK resistance-related complex consisting of (1) a supra-physiologic level of V600EBRAF, which activates CRAF, or (2) a moderately over-expressed V600EBRAF level concomitant with a mutant MEK1/2, which leads to increased V600EBRAF-MUTMEK interaction. Both signaling configurations strongly favor ERK activation, leading to growth/survival finely tuned to the BRAFi+MEKi level. Paradoxically, acute removal of BRAFi+MEKi disrupts this fine-tuning and results in a p-ERK rebound favoring cell arrest/death (i.e., drug addiction). WT (grey) and mutant (red) proteins; BRAFi or MEKi, blue circles.
Our study of melanoma cell lines with acquired resistance to combined BRAF and MEK inhibition has revealed insights into recent clinical studies. For instance, melanoma cell lines with preexisting BRAFi resistance augment preexisting mechanisms quickly as they adapt to combined BRAF and MEK inhibition. This is consistent with the clinical observation that patients who progressed on BRAFi or MEKi monotherapies infrequently respond to the addition of the other inhibitor, and, for those who do respond sequentially, the responses are generally highly transient. Furthermore, the importance of a MAPKi resistance-related complex has certain translational implications.
Successful strategies targeting this tunable-combinatorial signaling complex may include those inhibiting CRAF function (e.g., omni- or pan-RAF inhibitors), V600EBRAF-CRAF interaction, V600EBRAF-MUTMEK interaction/scaffolding, and MEK activation (e.g., phosphorylation by RAF). These strategies could be built around continued inhibition of mutant BRAF and MEK or alternating regimens. In our studies, the efficacy of an ERK inhibitor in overcoming acquired BRAFi+MEKi resistance was nuanced and depended on the experimental contexts, e.g., ERKi alone at lower concentrations promoted survival/growth of BRAFi+MEKi resistant melanoma cells in both short- and long-term assays.
While the build-up of a V600EBRAF-CRAF-MEK complex ultimately limited the efficacy of combined BRAF and MEK inhibition in melanoma, this signaling complex appeared to be poised to deliver a lethal dose of signaling once both inhibitors were efficiently and acutely removed (Figure 8). Melanoma cells with fully acquired BRAFi+MEKi resistance were much more sensitive to drug withdrawal than those with acquired resistance to BRAFi alone. It is possible that in vivo factors, such as tumor heterogeneity (e.g., sub-populations with reversible drug-tolerance but without drug addiction), three-dimensional cell-cell contacts, microenviromental signals, and/or host pharmacokinetic considerations, could render drug addiction a clinically intractable phenotype. In this light, the hypothesis of intermittent therapy with combined BRAFi and MEKi to delay acquired resistance will be tested prospectively within a large randomized clinical trial (SWOG/CTEP S1320) for the treatment of patients with BRAF mutant metastatic melanoma.
EXPERIMENTAL PROCEDURES
Patients, Tumor Samples and Genomic Analysis
Melanoma tissues and patient-matched normal tissues were collected with the approval of Institutional Review Boards at UCLA, UCSF and Vanderbilt and informed consents of each patient. Patients were enrolled in GlaxoSmithKline or Roche/Genetech clinical trials or treated per standard clinical management. We evaluated 45 tumor samples (27 DD-DP, four DP and 14 baseline or early on-treatment melanoma biopsies) from 14 patients who were either treated with BRAFi+MEKi upfront or with this combination after progression on BRAFi. In each tumor, genetic mechanisms (excluding PI3K-PTEN-AKT genetic hits) known to confer clinical resistance to BRAFi were detected by gDNA Q-PCR and/or Sanger sequencing. Twenty-three baseline and DD-DP tumors from seven patients along with normal tissues were whole exome sequence (WES)-analyzed to detect somatic alterations which are in the MAPK and PI3K-PTEN-AKT pathways and which are specific to drug-resistant tumors. Pair-end sequences with read length of 2×100 bps using the Illumina HiSeq2000 platform were generated. SNVs, INDELs and CNVs were analyzed and visualized as described (Shi et al., 2014).
Targeted Sequencing, Copy Number Quantification, and WES of Cell Lines
BRAF, NRAS, and DUSP4 cDNA levels were quantified by real time RT-PCR using TUBULIN and GAPDH levels for normalization. Relative expressions were calculated using the delta-Ct method. BRAF, NRAS, and DUSP4 gDNA relative copy numbers were quantified by real time PCR with total gDNA content estimated by assaying the β-globin gene in each sample. All primer sequences are available upon request. Sanger sequencing was performed using purified PCR via BigDye v1.1 (Applied Biosystems) in combination with a 3730 DNA Analyzer (Applied Biosystems). WES of M249 triple cell lines were analyzed for shared and distinct genetic alterations and their phylogenetic relationship.
Cell Culture, Constructs, Infections, and Transfections
All cell lines were maintained in DMEM with 10% heat-inactivated fetal bovine serum, 2 mmol/L glutamine in a humidified 5% CO2 incubator, with the addition of 10 ng/mL doxycycline and/or puromycin, when applicable. Stocks and dilutions of PLX4032/vemurafenib (Plexxikon, Berkeley, CA, USA), AZD6244/selumetinib (Selleck Chemicals) and SCH772984 (MERCK) were made in DMSO. Cell proliferation experiments were performed in a 96-well format (5 replicates per sample); drug treatments were initiated 24 hours post-seeding for 72 hour; and cell survival quantified using CellTiter-GLO (Promega). Clonogenic assays were performed by plating cells at single cell density in 6-well plates with fresh media and drug replenished every two days. Colonies were fixed in 4% paraformaldehyde and stained with 0.05% crystal violet. shBRAF, shCRAF, shPTEN and shNRAS were subcloned into the lentiviral vector pLL3.7; shDUSP4/pLK0.1 vectors were obtained commercially (GE Dharmacon). All WT and mutant MEK1 and BRAF constructs were epitope-tagged and sub-cloned into the doxycycline-repressible lentiviral vector pLVX-Tight-Puro (Clontech, Inc). Viral supernatants were generated by third-generation lentiviral packaging using human embryonic kideny (HEK) 293T cells. HEK293T cells were transfected using BioT (Bioland).
Protein Detection, Interaction, and Structure
Cell lysates were made in RIPA buffer (Sigma) for direct Western blotting or in a PNE buffer (PBS:H2O at 1 :1, 0.5% NP40, 5 mM EDTA, 5% glycerol) for immunoprecipitation, with both buffers supplemented with protease (Roche) and phosphatase (Santa Cruz Biotechnology) inhibitor cocktails. Western blots and immunoprecipitations were performed using the following antibodies : p-ERK1/2 (T202/Y204), p-MEK1/2 (S217/221), p-AKT (Thr308), p-CRAF (S338), total ERK1/2, MEK1/2, MEK1, MEK2, AKT, CRAF, DUSP4 and HA (Cell Signaling Technology), TUBULIN and FLAG (Sigma), BRAF (F-7), BRAF (C-19), p-MEK1 (Thr291), p-MEK1 (S222) (Santa Cruz Biotechnology) and p-MEK2 (S226) (US Biological). Western blot quantification was performed using NIH Image J. The three-dimension structures of MEK1 (3EQC) and PTEN mutants were modeled by the I-TASSER online server. Modeling the V600EBRAF-MUTMEK1 dimer interface was based on the crystal structure of the WTBRAF-WTMEK1 dimer (4MNE), the MEK1-KSR2 dimer (2Y4I), and the asymmetric, vemurafenib-bound, V600EBRAF dimer (3GO7). Protein structures were visualized using Pymol™
Supplementary Material
Structure of MEK1 with the locations of MAPKi resistance-related mutations, or residues homologous to MEK2 mutations, indicated in yellow, except that Q56 and F129 are indicated in green.
Figure S1. Related to Figure 1.
(A) Structure of PTEN (WT, magenta; F271V mutant, blue) showing locations of V271 (red) and F271 (yellow) as well as those of amino acids within 4Å (green).
(B) Western blot of indicated phospho-proteins and proteins in the PTEN non-expressing M249 V600EBRAF melanoma cell line transduced with vector, WTPTEN or F271V PTEN lentiviruses. TUBULIN, loading control.
Figure S2. Related to Figure 3.
(A–B) Western blot (WB) of indicated phospho-proteins and proteins in M249 P, DDR4, and DDR5 to follow the spontaneous recovery of p-ERK (A) and p-RSK (B) levels after a fresh dose of BRAFi and MEKi, each at 1 μM. Prior to DMSO or inhibitor treatment, cells were plated in the absence of BRAFi and MEKi for 16 hr. TUBULIN, loading control.
(C–E) (C) BRAF copy numbers (averages of duplicates; two primer sets) by Q-PCR in normal control gDNA (peripheral mononuclear cells) and in gDNA from M249 P and DDR2 and DDR3 polyclonal sub-lines derived by chronic adaptation to growth in 0.5 μM of BRAFi and MEKi. (D) Sanger gDNA sequencing chromatograms showing MEK1 mutational status and the approximate allelic ratios in M249 DDR2 and DDR3. (E) MEK1 and MEK2 gDNA Q-PCR showing copy numbers in M249 P and DDR sub-lines derived by two methods as indicated. Values represent averages of four measurements using two primer sets and performed in two independent experiments; error bars, standard deviation. MEK1 p-values (primer sets 1/2): 0.006/0.002 (DDR2 vs. P); 0.09/0.002 (DDR3 vs. P); 0.29/0.59 (DDR4 vs. P); 0.04/0.003 (DDR5 vs. P).
(F–G) (F) WB of indicated phospho-proteins (p-ERK exposurs shown in seconds) and proteins in WTBRAF HEK293T cells 72 hr after transfection with indicated FLAG-tagged MEK1 constructs and followed by 1 hr treatment with MEKi at 0, 0.01, 0.1, 1.0 and 10 μM. (G) Quantification of p-ERK WB signals in (F), with normalization to TUBULIN levels, to estimate cellular p-ERK IC50 associated with WTMEK1 vs. MUTMEK1.
(H) WB of indicated p-CRAF, p-ERK, their total levels and TUBULIN (loading control) in M249 DDR4 and DDR5, which were plated in the presence of BRAFi and MEKi, at 1 μM, for 20 hr followed by drug withdrawal for the indicated durations.
Inhibitors used, BRAFi (vemurafenib) and MEKi (selumetinib or AZD6244).
Figure S3. Related to Figure 4.
(A) Western blot (WB) of phospho-protein and proteins in M395 DDR cells infected with either shVECTOR (−) or shBRAF (+) lentivirus and cultured in the presence (+) or absence (−) of indicated inhibitor at 1 μM. TUBULIN, loading control.
(B) WB of p-ERK, ERK, and TUBULIN (loading control) in M249 P, DDR4, DDR5 and M249 P infected with the indicated lentivirus(es). Cells were plated for 24 hr without inhibitors, treated for 16 hr (except Parental or Parental+Vector) with BRAFi and MEKi at 1 μM, and followed by inhibitor wash-out and continued incubation for 8 hr. Grey bar indicates engineered M249 P cells subjected to BRAFi+MEKi pre-conditioning, which involved withdrawal from doxycycline (to induced gene expression) and treatment with BRAFi+MEKi at 1 μM for 28 days prior to the WB experiment.
(C) Three-day MTT assays (n=5; normalized to DMSO vehicle as 100%) of the M249 P, DDR4, DDR5 cell lines plated 16 hr without inhibitors and treated with BRAFi+MEKi at the indicated concentrations in μM. Experiment performed in parallel with that in D. Error bars, +/− SEM.
(D) Three-day MTT assays (n=5; normalized to DMSO vehicle as 100%) for the M249 P cell line engineered with the indicated construct(s). Expression of constructs was initiated by a two-day withdrawal from doxycycline. A set of M249 P engineered cell lines was pre-conditioned with BRAFi+MEKi treatments as described in B. Cells were plated and treated as described in C. Error bars, +/− SEM.
(E) Scatter plot showing measurements (n=5) of the relative viability and growth (using three-day MTT assays) of M249 P engineered lines (as in B and D) cultured in the absence of BRAFi and MEKi (− or + indicates prior pre-conditioning). All cell lines were removed from doxycycline, and values for growth of pre-conditioned cells were expressed relative to an arbitrary value chosen from the non-preconditioned group set as 100%.
(F) Long-term (10 day clonogenic assay) and short-term (3 day MTT assay) growth capacities of indicated cell lines in the absence of BRAFi and MEKi. Raw MTT values (with background or media only values subtracted) at day 1 (24 hr after seeding) and day 4 for all cell lines are plotted as means (n=5) and their errors (+/− SD). Numeric values represent the fold changes in growth of each cell line (day 4 mean divided by day 1 mean). V, vector.
(G) WB of indicated phospho- and total proteins in M395 P, M395 R (V600EBRAF-amplified), and M395 R engineered to express the indicated FLAG-tagged MEK1 constructs. Cells as indicated (+) were treated with BRAFi at 1 μM or vehicle (DMSO) for 48 hr.
(H) WB of inidcated protein levels in cell lines described in G were either treated with DMSO (P) or BRAFi+MEKi for 16 hr at 1 μM or washed free of inhibitors followed by additional 8 hr of culture. Tx, treatment.
(I) Three-day MTT (top) and twelve-day clonogenic (bottom) survival assays of indicated cell lines treated with BRAFi alone, MEKi alone, or both BRAFi+MEKi (in μM) (M395 R+either MUTMEK1 vs. M395 R, p < 0.001 for both assays at 1 μM of BRAFi+MEKi). Error bars, +/− SEM.
Inhibitors used, BRAFi (vemurafenib) and MEKi (selumetinib).
Figure S4. Related to Figure 5.
M249 P engineered to express vector or indicated FLAG-MEK1 constructs concurrent with over-expression of either HA-WTBRAF or HA-V600EBRAF were plated with BRAFi+MEKi (1 μM, 16 hr; except vector control), and the lysates were subjected to IP (anti-IgG or -FLAG). Western blot of immunoprecipitated (IP) and total fractions.
Figure S5. Related to Figure 6. A MEK1-V600EBRAF complex with arginine 662 at its interface.
(A) Structure of the WTMEK1-WTBRAF heterodimer (KD, kinase domain) (PDB4MNE) showing (i) the face-to-face configuration of the complex, (ii) locations (amino acid residues highlighted in yellow) of MEK1 residues mutated in melanomas with acquired BRAFi or BRAFi+MEKi resistance, and (iii) locations (yellow highlights) of BRAF V600 and R509 (i.e., the RAF dimerization interface). MEK1 residues 1–65 were not resolved in the crystal structure of 4MNE. Hence, structure of this N-terminal MEK1 region was modeled by alignment to the MEK1 structure in PDB2Y4I.
(B) Structure of the WTMEK1-V600EBRAF heterodimer modeled using the vemurafenib-bound V600EBRAF structure (PDB3OG7) aligned with the WTBRAF shown in (A) (root mean square distance or RMSD, 0.511 Å). Circled, αC helix pulled up into a “closed” conformation; BRAF E600 residue, not resolved in PDB4MNE.
(C) A hydrophobic core critical for MEK1-KSR2 interaction (PDB2Y4I) showing the αG helix of KSR2 and residue A879 critical for this interaction (i.e., A879L mutation disrupts this interaction). Structure alignment of BRAF KD and KSR2 KD (RMSD, 2.3 Å) showed that BRAF R662 would be equivalent to KSR2 A879.
(D) Structure of the WTMEK1-V600EBRAF heterodimer highlighting two orientations of the R662 side chain based on WTMEK1-WTBRAF dimer co-crystal structure (yellow) vs. the vemurafenib-bound V600EBRAF crystal structure (orange). The yellow (i.e., interacting) R662 side chain orientation was used to model a putative MEK-BRAF heterodimer interface; R662 was nominated for mutagenesis and additional studies.
Figure S6. Related to Figure 7.
(A) Western blots of p-ERK, total ERK, and TUBULIN (loading control) in M249 DDR4 and DDR5 cell lines cultured either with (“on”) or without (“off”) BRAFi+MEKi, at 1 μM (BRAFi, vemurafenib; MEKi, selumetinib). Cultures off BRAFi+MEKi were treated with ERKi (SCH772984) at the indicated concentrations for 1 hr prior to cell lysis.
(B) Western blots of p-ERK, total ERK, and TUBULIN (loading control) in M249 DDR4 cell line which was withdrawn from BRAFi+MEKi at 1 μM for the indicated hr and then treated prior to cell lysis (“add back”) with BRAFi+MEKi at 1 μM, DMSO (none), or ERKi at 0.1 μM for 1 hr.
(C) Regression of disease progressive melanomas after discontinuation of combined BRAF/MEK targeted therapy. Cases of observed tumor regression subsequent to therapy cessation due to acquired drug resistance among evaluable patients (defined as patients with at least a partial objective response, without intervening treatment after disease progression and cessation of therapy, and with available radiological scans after withdrawal of BRAF and MEK inhibitors; n=6, Vanderbilt; n=8, UCSF; n=1, UCLA; total n=15; Table S4). Not shown, the majority of DP tumors followed continued to progress while off treatment. Pt, patient; Tx, treatment; DP, disease progression. White arrows point to tumors with measurements at the bottom of each CT image.
(D) Disease progressive melanomas with tumor growth rate deceleration after discontinuation of single-agent BRAF targeted therapy. Cases of tumor growth rate reduction following discontinuation of BRAF inhibitor therapy due to acquired drug resistance. Tumor growth rates (TGRs) were measured as a percentage change (in the greatest diameter of the tumors indicated by the white arrows) per week. TGRs of indicated tumors are shown from all three patients whose melanomas (i.e., target lesions) displayed an overall decelerated TGR (sum of the greatest diameters) (total evaluable patients n=16; Table S4). Not shown, no tumor was observed to regress (i.e., with a negative TGR) after vemurafenib withdrawal (range of observation interval without treatment=34–100 days).
Table S1. Related to Figure 1. Clinical characteristics of patients and their responses to BRAF + MEK inhibitors.
Table S2. Related to Figure 1. Genetic alterations detected in core resistance pathways.
Table S3. Related to Figure 3. Provided as an Excel file.
Table S4. Related to Figure 7. Clinical characteristics of patients followed for melanoma tumor regression or growth deceleration after cessation of MAPK targeted therapies.
Significance.
The understanding that BRAF mutant melanomas frequently acquire BRAFi resistance via MAPK pathway reactivation has guided the development of combined BRAF and MEK targeted therapy. Our finding that acquired resistance to BRAF and MEK co-targeting is driven by highly tunable-combinatorial mechanisms of resistance underscores the intrinsic limitation of dual MAPK pathway targeting. Mechanistic studies highlight V600EBRAF-WTCRAF and V600EBRAF-MUTMEK interactions as a basis for ERK-reactivation. Additionally, we demonstrate that melanoma cells with acquired BRAFi+MEKi resistance are exquisitely sensitive to acute drug withdrawal. Exploiting melanoma addiction to BRAFi+MEKi for therapeutic gain is being tested, via intermittent drug dosing, in the clinic (SWOG/CTEP S1320).
Acknowledgments
We are grateful to G. Bollag (Plexxikon Inc.) for providing PLX4032, B. Chmielowski and J. Glaspy for coordinated patient care, Art Villanueva, Jackie Hernandez, Elizabeth Seja, and Christine Kivork for coordinating clinical trials and specimen collection, and all patient volunteers. This work has been funded by Burroughs Wellcome Fund (to R.S.L), Stand Up To Cancer (to R.S.L.), Melanoma Research Alliance (to R.S.L. and A.A.), the National Institutes of Health (1R01CA176111 to R.S.L and 1P01CA168585 to A.R. and R.S.L.), the Ressler Family Foundation (to R.S.L. and A.R.), the Seaver institute (to R.S.L. and A.R.), the American Skin Association (to H.S. and R.S.L.), the Harry J. Lloyd Charitable Trust (to R.S.L.), the Ian Copeland Melanoma Fund (to R.S.L.), the Steven C. Gordon Family Foundation (to R.S.L. and A.R.), the NIH (5K24CA097588-09 to J.A.S.), and the Royal Marsden (to J.L.).
Footnotes
Accession Number
The SRA accession number for the exome sequence data reported in this paper is SRP049746.
AUTHOR CONTRIBUTIONS
G.M., W.H., A.H., H.S., X.K., C.C.Y. and R.S.L designed and performed experiments and analyzed data. R.C.K., A.A.S., N.K., J.B., K.B.D., D.B.J., A.A., J.A.S., A.R., and R.S.L. contributed clinical samples or key reagents. K.R., H.S., J.L., D.B.J., A.A., J.A.S., A.R., and R.S.L. analyzed clinical data. R.S.L. wrote the paper. G.M., W.H., A.H., H.S., X.K., D.B.J., A.A., J.A.S. and A.R. contributed to writing the paper. G.M., W.H., A.H., and H.S. contributed equally to this work.
No author declared conflicts of interest.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Bollag G, Hirth P, Tsai J, Zhang J, Ibrahim PN, Cho H, Spevak W, Zhang C, Zhang Y, Habets G, et al. Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature. 2010;467:596–599. doi: 10.1038/nature09454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brennan DF, Dar AC, Hertz NT, Chao WC, Burlingame AL, Shokat KM, Barford D. A Raf-induced allosteric transition of KSR stimulates phosphorylation of MEK. Nature. 2011;472:366–369. doi: 10.1038/nature09860. [DOI] [PubMed] [Google Scholar]
- Catalanotti F, Reyes G, Jesenberger V, Galabova-Kovacs G, de Matos Simoes R, Carugo O, Baccarini M. A Mek1-Mek2 heterodimer determines the strength and duration of the Erk signal. Nat Struct Mol Biol. 2009;16:294–303. doi: 10.1038/nsmb.1564. [DOI] [PubMed] [Google Scholar]
- Corcoran RB, Dias-Santagata D, Bergethon K, Iafrate AJ, Settleman J, Engelman JA. BRAF gene amplification can promote acquired resistance to MEK inhibitors in cancer cells harboring the BRAF V600E mutation. Sci Signal. 2010;3:ra84. doi: 10.1126/scisignal.2001148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emery CM, Vijayendran KG, Zipser MC, Sawyer AM, Niu L, Kim JJ, Hatton C, Chopra R, Oberholzer PA, Karpova MB, et al. MEK1 mutations confer resistance to MEK and B-RAF inhibition. Proc Natl Acad Sci U S A. 2009;106:20411–20416. doi: 10.1073/pnas.0905833106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haling JR, Sudhamsu J, Yen I, Sideris S, Sandoval W, Phung W, Bravo BJ, Giannetti AM, Peck A, Masselot A, et al. Structure of the BRAF-MEK Complex Reveals a Kinase Activity Independent Role for BRAF in MAPK Signaling. Cancer Cell. 2014 doi: 10.1016/j.ccr.2014.07.007. [DOI] [PubMed] [Google Scholar]
- Hodis E, Watson IR, Kryukov GV, Arold ST, Imielinski M, Theurillat JP, Nickerson E, Auclair D, Li L, Place C, et al. A landscape of driver mutations in melanoma. Cell. 2012;150:251–263. doi: 10.1016/j.cell.2012.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krauthammer M, Kong Y, Ha BH, Evans P, Bacchiocchi A, McCusker JP, Cheng E, Davis MJ, Goh G, Choi M, et al. Exome sequencing identifies recurrent somatic RAC1 mutations in melanoma. Nat Genet. 2012;44:1006–1014. doi: 10.1038/ng.2359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nan X, Collisson EA, Lewis S, Huang J, Tamguney TM, Liphardt JT, McCormick F, Gray JW, Chu S. Single-molecule superresolution imaging allows quantitative analysis of RAF multimer formation and signaling. Proc Natl Acad Sci U S A. 2013;110:18519–18524. doi: 10.1073/pnas.1318188110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nazarian R, Shi H, Wang Q, Kong X, Koya RC, Lee H, Chen Z, Lee MK, Attar N, Sazegar H, et al. Melanomas acquire resistance to B-RAF(V600E) inhibition by RTK or N-RAS upregulation. Nature. 2010;468:973–977. doi: 10.1038/nature09626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikolaev SI, Rimoldi D, Iseli C, Valsesia A, Robyr D, Gehrig C, Harshman K, Guipponi M, Bukach O, Zoete V, et al. Exome sequencing identifies recurrent somatic MAP2K1 and MAP2K2 mutations in melanoma. Nat Genet. 2012;44:133–139. doi: 10.1038/ng.1026. [DOI] [PubMed] [Google Scholar]
- Poulikakos PI, Persaud Y, Janakiraman M, Kong X, Ng C, Moriceau G, Shi H, Atefi M, Titz B, Gabay MT, et al. RAF inhibitor resistance is mediated by dimerization of aberrantly spliced BRAF(V600E) Nature. 2011;480:387–390. doi: 10.1038/nature10662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ribas A, Flaherty KT. BRAF targeted therapy changes the treatment paradigm in melanoma. Nat Rev Clin Oncol. 2011;8:426–433. doi: 10.1038/nrclinonc.2011.69. [DOI] [PubMed] [Google Scholar]
- Shi H, Hugo W, Kong X, Hong A, Koya RC, Moriceau G, Chodon T, Guo R, Johnson DB, Dahlman KB, et al. Acquired Resistance and Clonal Evolution in Melanoma during BRAF Inhibitor Therapy. Cancer Discov. 2014 doi: 10.1158/2159-8290.CD-13-0642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H, Moriceau G, Kong X, Koya RC, Nazarian R, Pupo GM, Bacchiocchi A, Dahlman KB, Chmielowski B, Sosman JA, et al. Preexisting MEK1 exon 3 mutations in V600E/KBRAF melanomas do not confer resistance to BRAF inhibitors. Cancer Discov. 2012a;2:414–424. doi: 10.1158/2159-8290.CD-12-0022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H, Moriceau G, Kong X, Lee MK, Lee H, Koya RC, Ng C, Chodon T, Scolyer RA, Dahlman KB, et al. Melanoma whole-exome sequencing identifies (V600E)B-RAF amplification-mediated acquired B-RAF inhibitor resistance. Nat Commun. 2012b;3:724. doi: 10.1038/ncomms1727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thakur MD, Salangsang F, Landman AS, Sellers W, Pryer NK, Levesque MP, Dummer R, McMahon M, Stuart DD. Modeling vemurafenib resistance in melanoma reveals a strategy to forestall drug resistance. Nature. 2013 doi: 10.1038/nature11814. Published online 09 January 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Allen EM, Wagle N, Sucker A, Treacy DJ, Johannessen CM, Goetz EM, Place CS, Taylor-Weiner A, Whittaker S, Kryukov GV, et al. The genetic landscape of clinical resistance to RAF inhibition in metastatic melanoma. Cancer Discov. 2014;4:94–109. doi: 10.1158/2159-8290.CD-13-0617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villanueva J, Infante JR, Krepler C, Reyes-Uribe P, Samanta M, Chen HY, Li B, Swoboda RK, Wilson M, Vultur A, et al. Concurrent MEK2 mutation and BRAF amplification confer resistance to BRAF and MEK inhibitors in melanoma. Cell Rep. 2013;4:1090–1099. doi: 10.1016/j.celrep.2013.08.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagle N, Emery C, Berger MF, Davis MJ, Sawyer A, Pochanard P, Kehoe SM, Johannessen CM, Macconaill LE, Hahn WC, et al. Dissecting therapeutic resistance to RAF inhibition in melanoma by tumor genomic profiling. J Clin Oncol. 2011;29:3085–3096. doi: 10.1200/JCO.2010.33.2312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagle N, Van Allen EM, Treacy DJ, Frederick DT, Cooper ZA, Taylor-Weiner A, Rosenberg M, Goetz EM, Sullivan RJ, Farlow DN, et al. MAP Kinase Pathway Alterations in BRAF-Mutant Melanoma Patients with Acquired Resistance to Combined RAF/MEK Inhibition. Cancer Discov. 2014 doi: 10.1158/2159-8290.CD-13-0631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H, Daouti S, Li WH, Wen Y, Rizzo C, Higgins B, Packman K, Rosen N, Boylan JF, Heimbrook D, Niu H. Identification of the MEK1(F129L) activating mutation as a potential mechanism of acquired resistance to MEK inhibition in human cancers carrying the B-RafV600E mutation. Cancer Res. 2011;71:5535–5545. doi: 10.1158/0008-5472.CAN-10-4351. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Structure of MEK1 with the locations of MAPKi resistance-related mutations, or residues homologous to MEK2 mutations, indicated in yellow, except that Q56 and F129 are indicated in green.
Figure S1. Related to Figure 1.
(A) Structure of PTEN (WT, magenta; F271V mutant, blue) showing locations of V271 (red) and F271 (yellow) as well as those of amino acids within 4Å (green).
(B) Western blot of indicated phospho-proteins and proteins in the PTEN non-expressing M249 V600EBRAF melanoma cell line transduced with vector, WTPTEN or F271V PTEN lentiviruses. TUBULIN, loading control.
Figure S2. Related to Figure 3.
(A–B) Western blot (WB) of indicated phospho-proteins and proteins in M249 P, DDR4, and DDR5 to follow the spontaneous recovery of p-ERK (A) and p-RSK (B) levels after a fresh dose of BRAFi and MEKi, each at 1 μM. Prior to DMSO or inhibitor treatment, cells were plated in the absence of BRAFi and MEKi for 16 hr. TUBULIN, loading control.
(C–E) (C) BRAF copy numbers (averages of duplicates; two primer sets) by Q-PCR in normal control gDNA (peripheral mononuclear cells) and in gDNA from M249 P and DDR2 and DDR3 polyclonal sub-lines derived by chronic adaptation to growth in 0.5 μM of BRAFi and MEKi. (D) Sanger gDNA sequencing chromatograms showing MEK1 mutational status and the approximate allelic ratios in M249 DDR2 and DDR3. (E) MEK1 and MEK2 gDNA Q-PCR showing copy numbers in M249 P and DDR sub-lines derived by two methods as indicated. Values represent averages of four measurements using two primer sets and performed in two independent experiments; error bars, standard deviation. MEK1 p-values (primer sets 1/2): 0.006/0.002 (DDR2 vs. P); 0.09/0.002 (DDR3 vs. P); 0.29/0.59 (DDR4 vs. P); 0.04/0.003 (DDR5 vs. P).
(F–G) (F) WB of indicated phospho-proteins (p-ERK exposurs shown in seconds) and proteins in WTBRAF HEK293T cells 72 hr after transfection with indicated FLAG-tagged MEK1 constructs and followed by 1 hr treatment with MEKi at 0, 0.01, 0.1, 1.0 and 10 μM. (G) Quantification of p-ERK WB signals in (F), with normalization to TUBULIN levels, to estimate cellular p-ERK IC50 associated with WTMEK1 vs. MUTMEK1.
(H) WB of indicated p-CRAF, p-ERK, their total levels and TUBULIN (loading control) in M249 DDR4 and DDR5, which were plated in the presence of BRAFi and MEKi, at 1 μM, for 20 hr followed by drug withdrawal for the indicated durations.
Inhibitors used, BRAFi (vemurafenib) and MEKi (selumetinib or AZD6244).
Figure S3. Related to Figure 4.
(A) Western blot (WB) of phospho-protein and proteins in M395 DDR cells infected with either shVECTOR (−) or shBRAF (+) lentivirus and cultured in the presence (+) or absence (−) of indicated inhibitor at 1 μM. TUBULIN, loading control.
(B) WB of p-ERK, ERK, and TUBULIN (loading control) in M249 P, DDR4, DDR5 and M249 P infected with the indicated lentivirus(es). Cells were plated for 24 hr without inhibitors, treated for 16 hr (except Parental or Parental+Vector) with BRAFi and MEKi at 1 μM, and followed by inhibitor wash-out and continued incubation for 8 hr. Grey bar indicates engineered M249 P cells subjected to BRAFi+MEKi pre-conditioning, which involved withdrawal from doxycycline (to induced gene expression) and treatment with BRAFi+MEKi at 1 μM for 28 days prior to the WB experiment.
(C) Three-day MTT assays (n=5; normalized to DMSO vehicle as 100%) of the M249 P, DDR4, DDR5 cell lines plated 16 hr without inhibitors and treated with BRAFi+MEKi at the indicated concentrations in μM. Experiment performed in parallel with that in D. Error bars, +/− SEM.
(D) Three-day MTT assays (n=5; normalized to DMSO vehicle as 100%) for the M249 P cell line engineered with the indicated construct(s). Expression of constructs was initiated by a two-day withdrawal from doxycycline. A set of M249 P engineered cell lines was pre-conditioned with BRAFi+MEKi treatments as described in B. Cells were plated and treated as described in C. Error bars, +/− SEM.
(E) Scatter plot showing measurements (n=5) of the relative viability and growth (using three-day MTT assays) of M249 P engineered lines (as in B and D) cultured in the absence of BRAFi and MEKi (− or + indicates prior pre-conditioning). All cell lines were removed from doxycycline, and values for growth of pre-conditioned cells were expressed relative to an arbitrary value chosen from the non-preconditioned group set as 100%.
(F) Long-term (10 day clonogenic assay) and short-term (3 day MTT assay) growth capacities of indicated cell lines in the absence of BRAFi and MEKi. Raw MTT values (with background or media only values subtracted) at day 1 (24 hr after seeding) and day 4 for all cell lines are plotted as means (n=5) and their errors (+/− SD). Numeric values represent the fold changes in growth of each cell line (day 4 mean divided by day 1 mean). V, vector.
(G) WB of indicated phospho- and total proteins in M395 P, M395 R (V600EBRAF-amplified), and M395 R engineered to express the indicated FLAG-tagged MEK1 constructs. Cells as indicated (+) were treated with BRAFi at 1 μM or vehicle (DMSO) for 48 hr.
(H) WB of inidcated protein levels in cell lines described in G were either treated with DMSO (P) or BRAFi+MEKi for 16 hr at 1 μM or washed free of inhibitors followed by additional 8 hr of culture. Tx, treatment.
(I) Three-day MTT (top) and twelve-day clonogenic (bottom) survival assays of indicated cell lines treated with BRAFi alone, MEKi alone, or both BRAFi+MEKi (in μM) (M395 R+either MUTMEK1 vs. M395 R, p < 0.001 for both assays at 1 μM of BRAFi+MEKi). Error bars, +/− SEM.
Inhibitors used, BRAFi (vemurafenib) and MEKi (selumetinib).
Figure S4. Related to Figure 5.
M249 P engineered to express vector or indicated FLAG-MEK1 constructs concurrent with over-expression of either HA-WTBRAF or HA-V600EBRAF were plated with BRAFi+MEKi (1 μM, 16 hr; except vector control), and the lysates were subjected to IP (anti-IgG or -FLAG). Western blot of immunoprecipitated (IP) and total fractions.
Figure S5. Related to Figure 6. A MEK1-V600EBRAF complex with arginine 662 at its interface.
(A) Structure of the WTMEK1-WTBRAF heterodimer (KD, kinase domain) (PDB4MNE) showing (i) the face-to-face configuration of the complex, (ii) locations (amino acid residues highlighted in yellow) of MEK1 residues mutated in melanomas with acquired BRAFi or BRAFi+MEKi resistance, and (iii) locations (yellow highlights) of BRAF V600 and R509 (i.e., the RAF dimerization interface). MEK1 residues 1–65 were not resolved in the crystal structure of 4MNE. Hence, structure of this N-terminal MEK1 region was modeled by alignment to the MEK1 structure in PDB2Y4I.
(B) Structure of the WTMEK1-V600EBRAF heterodimer modeled using the vemurafenib-bound V600EBRAF structure (PDB3OG7) aligned with the WTBRAF shown in (A) (root mean square distance or RMSD, 0.511 Å). Circled, αC helix pulled up into a “closed” conformation; BRAF E600 residue, not resolved in PDB4MNE.
(C) A hydrophobic core critical for MEK1-KSR2 interaction (PDB2Y4I) showing the αG helix of KSR2 and residue A879 critical for this interaction (i.e., A879L mutation disrupts this interaction). Structure alignment of BRAF KD and KSR2 KD (RMSD, 2.3 Å) showed that BRAF R662 would be equivalent to KSR2 A879.
(D) Structure of the WTMEK1-V600EBRAF heterodimer highlighting two orientations of the R662 side chain based on WTMEK1-WTBRAF dimer co-crystal structure (yellow) vs. the vemurafenib-bound V600EBRAF crystal structure (orange). The yellow (i.e., interacting) R662 side chain orientation was used to model a putative MEK-BRAF heterodimer interface; R662 was nominated for mutagenesis and additional studies.
Figure S6. Related to Figure 7.
(A) Western blots of p-ERK, total ERK, and TUBULIN (loading control) in M249 DDR4 and DDR5 cell lines cultured either with (“on”) or without (“off”) BRAFi+MEKi, at 1 μM (BRAFi, vemurafenib; MEKi, selumetinib). Cultures off BRAFi+MEKi were treated with ERKi (SCH772984) at the indicated concentrations for 1 hr prior to cell lysis.
(B) Western blots of p-ERK, total ERK, and TUBULIN (loading control) in M249 DDR4 cell line which was withdrawn from BRAFi+MEKi at 1 μM for the indicated hr and then treated prior to cell lysis (“add back”) with BRAFi+MEKi at 1 μM, DMSO (none), or ERKi at 0.1 μM for 1 hr.
(C) Regression of disease progressive melanomas after discontinuation of combined BRAF/MEK targeted therapy. Cases of observed tumor regression subsequent to therapy cessation due to acquired drug resistance among evaluable patients (defined as patients with at least a partial objective response, without intervening treatment after disease progression and cessation of therapy, and with available radiological scans after withdrawal of BRAF and MEK inhibitors; n=6, Vanderbilt; n=8, UCSF; n=1, UCLA; total n=15; Table S4). Not shown, the majority of DP tumors followed continued to progress while off treatment. Pt, patient; Tx, treatment; DP, disease progression. White arrows point to tumors with measurements at the bottom of each CT image.
(D) Disease progressive melanomas with tumor growth rate deceleration after discontinuation of single-agent BRAF targeted therapy. Cases of tumor growth rate reduction following discontinuation of BRAF inhibitor therapy due to acquired drug resistance. Tumor growth rates (TGRs) were measured as a percentage change (in the greatest diameter of the tumors indicated by the white arrows) per week. TGRs of indicated tumors are shown from all three patients whose melanomas (i.e., target lesions) displayed an overall decelerated TGR (sum of the greatest diameters) (total evaluable patients n=16; Table S4). Not shown, no tumor was observed to regress (i.e., with a negative TGR) after vemurafenib withdrawal (range of observation interval without treatment=34–100 days).
Table S1. Related to Figure 1. Clinical characteristics of patients and their responses to BRAF + MEK inhibitors.
Table S2. Related to Figure 1. Genetic alterations detected in core resistance pathways.
Table S3. Related to Figure 3. Provided as an Excel file.
Table S4. Related to Figure 7. Clinical characteristics of patients followed for melanoma tumor regression or growth deceleration after cessation of MAPK targeted therapies.