The reaction of terbium(III) nitrate pentahydrate in acetonitrile with N,N′-bis(2-hydroxybenzyl)-N,N′-bis(pyridin-2-ylmethyl)ethylenediamine (H2bbpen), previously deprotonated with triethylamine, produced the mononuclear compound [Tb(Cbbpen)(NO3)]. The molecule lies on a twofold rotation axis and the TbIII ion is eight-coordinate with a slightly distorted dodecahedral coordination geometry.
Keywords: crystal structure; lanthanide; terbium(III); N,N′-bis(2-hydroxybenzyl)-N,N′-bis(pyridin-2-ylmethyl)ethylenediamine; mononuclear; dodecahedral.
Abstract
The reaction of terbium(III) nitrate pentahydrate in acetonitrile with N,N′-bis(2-hydroxybenzyl)-N,N′-bis(pyridin-2-ylmethyl)ethylenediamine (H2bbpen), previously deprotonated with triethylamine, produced the mononuclear compound [N,N′-bis(2-oxidobenzyl-κO)-N,N′-bis(pyridin-2-ylmethyl-κN)ethylenediamine-κ2 N,N′](nitrato-κ2 O,O′)terbium(III), [Tb(C28H28N4O2)(NO3)]. The molecule lies on a twofold rotation axis and the TbIII ion is eight-coordinate with a slightly distorted dodecahedral coordination geometry. In the symmetry-unique part of the molecule, the pyridine and benzene rings are both essentially planar and form a dihedral angle of 61.42 (7)°. In the molecular structure, the N4O4 coordination environment is defined by the hexadentate bbpen ligand and the bidentate nitrate anion. In the crystal, a weak C—H⋯O hydrogen bond links molecules into a two-dimensional network parallel to (001).
Chemical context
As far as biological and biomedical applications are concerned, complexes of polydentate ligands with a range of metal ions in different oxidation states have been synthesized to model active sites of metalloproteins and to shed light on the consequences of heavy-metal chelation in living organisms, among many other applications (Colotti et al., 2013 ▸; Nurchi et al., 2013 ▸; Sears, 2013 ▸; Happe & Hemschemeier, 2014 ▸). Pyridyl and phenolate groups have been incorporated into these ligands because of their potential to mimic the coordination environments provided by the amino acids histidine and tyrosine, respectively (Hancock, 2013 ▸; Lenze et al., 2013 ▸). In this context, the heterotrifunctional Lewis base N,N′-bis(2-hydroxybenzyl)-N,N′-bis(pyridin-2-ylmethyl)ethylenediamine (H2bbpen) is suitable for the coordination of a range of p-, d- and f-block ions because of its versatile soft donor atoms in the pyridine rings and hard donors in the amine and phenolate groups (Neves et al., 1992 ▸; Schwingel et al., 1996 ▸). Electrochemical studies of the mononuclear [Mn(bbpen)]PF6, for example, revealed that this complex mimics some of the redox features of the photosystem II (PSII) (Neves et al., 1992 ▸). Complexes of bbpen2– with vanadium(III) and oxido-vanadium(IV) have been obtained as models of the vanadium-modified transferrin, the probable vanadium-transporting protein in higher organisms (Neves et al., 1991 ▸, 1993 ▸). Iron complexes of bbpen2– modified with electron-donating and -withdrawing groups (Me, Br, NO2), in turn, have been synthesized to provide detailed chemical information on the enzymatic activity of iron-tyrosinate proteins (Lanznaster et al., 2006 ▸). This ligand has also been employed to prepare lanthanide(III), gallium(III) and indium(III) complexes for medicinal applications such as the development of new contrast agents for magnetic resonance imaging, MRI (Wong et al., 1995 ▸, 1996 ▸; Setyawati et al., 2000 ▸).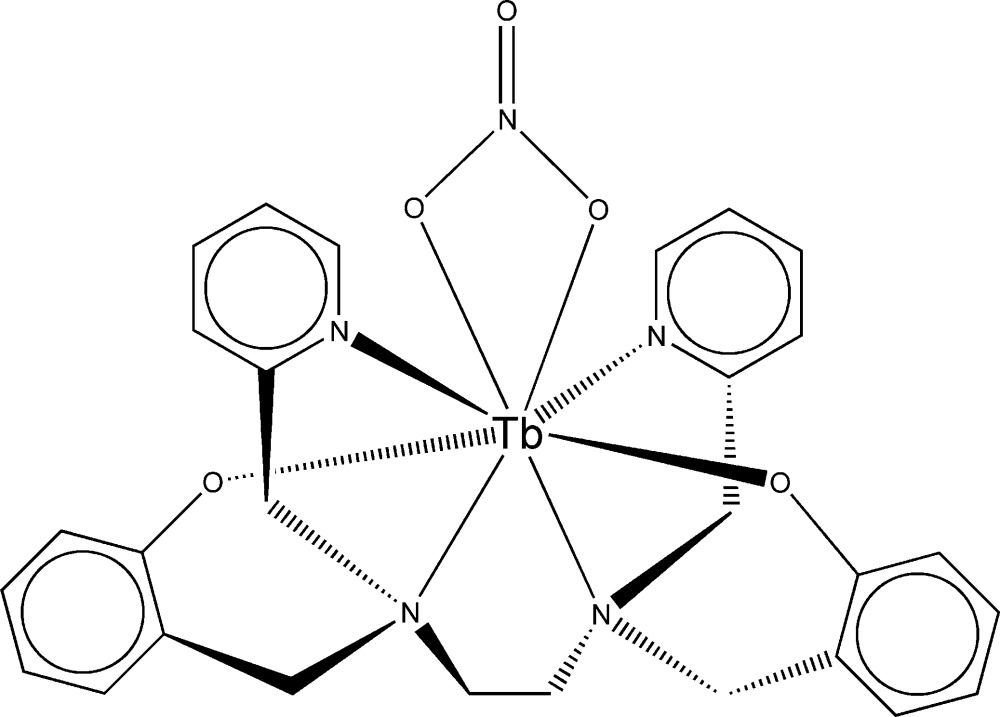
More recently, lanthanide(III) chelate complexes have also attracted attention in the field of molecular magnetism due to their highly significant single-ion magnetic anisotropy (Sessoli & Powell, 2009 ▸; Luzon & Sessoli, 2012 ▸). Accordingly, a number of examples of mononuclear lanthanide complexes that exhibit single-molecule magnet (SMM) behaviour have been reported (Rinehart & Long, 2011 ▸; Chilton et al., 2013 ▸; Ungur et al., 2014 ▸; Zhang et al., 2014 ▸). Our interest in the class of lanthanide complexes in which two coordination sites are occupied by relatively labile ligands, as in the title complex, comes from the possibility of using them as starting materials for the preparation of heteronuclear aggregates of d- and f-block ions that present SMM features. In this case, the replacement of the labile ligands by specific bidentate metalloligands can give rise to heteronuclear metal aggregates in which desirable ferromagnetic or ferrimagnetic exchange interactions are favoured (Totaro et al., 2013 ▸; Westrup et al., 2014 ▸).
Structural commentary
The molecular structure of the title compound is shown in Fig. 1 ▸. The TbIII ion is eight-coordinate with a dodecahedral array of N and O atoms (Table 1 ▸); the four N atoms of the O2N4-ligand (bbpen) form one plane, the four O atoms the other, with the phenolic O atoms in the B-sites (roughly equatorial) and the nitrate group O atoms in the A-sites (above and below the equatorial plane). The normals to the two planes are essentially perpendicular. A twofold rotation axis passes through O3 and N1 of the nitrate group, the terbium(III) atom and the mid-point of the C7—C7i bond [symmetry code (i) 1 − x, y, −z +  ]. In the symmetry-unique part of the molecule, the pyridine and benzene rings are both essentially planar and form a dihedral angle of 61.42 (7)°. The eightfold coordination pattern might also be described as a distorted bicapped trigonal prism with O1 and N2 as the capping atoms. However, this ignores the symmetry of the coordination, e.g. O1 and O1i would occupy different sites in the coordination polyhedron. Also, some of the rectangular faces of the prism are difficult to identify. In contrast, the dodecahedral pattern incorporates the twofold symmetry and the distortion from the ideal geometry is minimal.
]. In the symmetry-unique part of the molecule, the pyridine and benzene rings are both essentially planar and form a dihedral angle of 61.42 (7)°. The eightfold coordination pattern might also be described as a distorted bicapped trigonal prism with O1 and N2 as the capping atoms. However, this ignores the symmetry of the coordination, e.g. O1 and O1i would occupy different sites in the coordination polyhedron. Also, some of the rectangular faces of the prism are difficult to identify. In contrast, the dodecahedral pattern incorporates the twofold symmetry and the distortion from the ideal geometry is minimal.
Figure 1.
View of a molecule of [Tb(bbpen)(NO3)], indicating the atom-numbering scheme. H atoms have been omitted for clarity. Displacement ellipsoids are drawn at the 50% probability level [symmetry code: (i) −x + 1, y, −z +  ].
].
Table 1. Selected bond lengths ().
| Tb1O1 | 2.1947(13) | Tb1N3 | 2.5558(16) |
| Tb1O2 | 2.4764(15) | Tb1N1 | 2.891(2) |
| Tb1N2 | 2.5521(17) |
Supramolecular features
In the crystal, a weak C—H⋯O hydrogen bond (Table 2 ▸) links molecules into a two-dimensional network parallel to (001), Fig. 2 ▸.
Table 2. Hydrogen-bond geometry (, ).
| DHA | DH | HA | D A | DHA |
|---|---|---|---|---|
| C8H8BO3ii | 0.99 | 2.37 | 3.338(3) | 166 |
Symmetry code: (ii)  .
.
Figure 2.
A sheet of molecules, lying in a plane normal to the c axis, linked through short ‘weak hydrogen bonds’, as C8—H8B⋯O3ii [symmetry codes: (ii) x +  , y +
, y +  , z; (iii) x −
, z; (iii) x −  , y −
, y −  , z].
, z].
Database survey
Some examples of complexes with bbpen2– and related ligands with d-block metal ions appear in the literature (Xu et al., 2000 ▸; dos Anjos et al., 2006 ▸; Lanznaster et al., 2006 ▸; Golchoubian & Gholamnezhad, 2009 ▸; Thomas et al., 2010 ▸) as well as p-block metal(III) compounds (Wong et al., 1995 ▸, 1996 ▸) and related yttrium(III) and lanthanide(III) complexes (Setyawati et al., 2000 ▸; Yamada et al., 2010 ▸).
Synthesis and crystallization
Tb(NO3)3·5H2O, ethylenediamine, salicylaldehyde, sodium borohydride, 2-picolyl-chloride hydrochloride and triethylamine were purchased from Aldrich and used without purification. N,N′-bis(salicylidene)ethylenediamine (H2salen) (Diehl et al., 2007 ▸), N,N′-bis(2-hydroxybenzyl)ethylenediamine (H2bben) and N,N′-bis(2-hydroxybenzyl)-N,N′-bis(2-pyridylmethyl)ethylenediamine (H2bbpen) (Neves et al., 1992 ▸) were prepared as described in the literature. The preparation of the title complex was carried out under N2(g) using standard Schlenk and glove-box techniques. Acetonitrile was dried with CaH2 and distilled prior to use. A solution containing triethylamine (300 µl, 2.15 mmol) in acetonitrile (10 ml) was added to a suspension of H2bbpen (0.454 g, 1.00 mmol) in acetonitrile (25 ml) under stirring, giving a clear light-orange solution. After 15 min, this solution was added to a colourless solution of Tb(NO3)3·5H2O (0.434 g, 0.998 mmol) in acetonitrile (25 ml). A pale-yellow solution was obtained, which gave a 65% yield of the solid of the title compound upon cooling at 253 K for 2–3 days. Recrystallization of this solid by vapor diffusion of dimethoxyethane into the reaction mixture gave pale-pink crystals after two weeks at room temperature. These crystals are air-stable and insoluble in all common organic solvents.
Refinement
Crystal data, data collection and structure refinement details are summarized in Table 3 ▸. Hydrogen atoms were included in idealized positions (with C—H distances set at 0.97 and 0.93 Å for the methylene and trigonal–planar groups, respectively) and their U iso values were set to ride (1.2×) on the U eq values of the parent carbon atoms.
Table 3. Experimental details.
| Crystal data | |
| Chemical formula | [Tb(C28H28N4O2)(NO3)] |
| M r | 673.47 |
| Crystal system, space group | Orthorhombic, C2221 |
| Temperature (K) | 100 |
| a, b, c () | 8.5947(6), 18.2401(17), 16.9272(13) |
| V (3) | 2653.6(4) |
| Z | 4 |
| Radiation type | Mo K |
| (mm1) | 2.71 |
| Crystal size (mm) | 0.43 0.20 0.20 |
| Data collection | |
| Diffractometer | Bruker D8 Venture/Photon 100 CMOS |
| Absorption correction | Multi-scan (SADABS2014/2; Bruker, 2014 ▸) |
| T min, T max | 0.581, 0.746 |
| No. of measured, independent and observed [I > 2(I)] reflections | 75009, 3320, 3289 |
| R int | 0.020 |
| (sin /)max (1) | 0.668 |
| Refinement | |
| R[F 2 > 2(F 2)], wR(F 2), S | 0.010, 0.027, 1.15 |
| No. of reflections | 3320 |
| No. of parameters | 178 |
| H-atom treatment | H-atom parameters constrained |
| max, min (e 3) | 0.87, 0.30 |
| Absolute structure | Flack x determined using 1431 quotients [(I +)(I )]/[(I +)+(I )] (Parsons Flack, 2004 ▸) |
| Absolute structure parameter | 0.0107(19) |
Supplementary Material
Crystal structure: contains datablock(s) I. DOI: 10.1107/S2056989014026826/lh5741sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S2056989014026826/lh5741Isup2.hkl
CCDC reference: 1037922
Additional supporting information: crystallographic information; 3D view; checkCIF report
Acknowledgments
Financial support from the Brazilian agencies CNPq (grant No. 307592/2012–0) and CAPES (grant PVE A099/2013) is gratefully acknowledged. The authors also thank CNPq, CAPES and Fundação Araucária (Brazil) for fellowships.
supplementary crystallographic information
Crystal data
| [Tb(C28H28N4O2)(NO3)] | Dx = 1.686 Mg m−3 |
| Mr = 673.47 | Mo Kα radiation, λ = 0.71073 Å |
| Orthorhombic, C2221 | Cell parameters from 9558 reflections |
| a = 8.5947 (6) Å | θ = 2.9–28.3° |
| b = 18.2401 (17) Å | µ = 2.71 mm−1 |
| c = 16.9272 (13) Å | T = 100 K |
| V = 2653.6 (4) Å3 | Prism, pale pink |
| Z = 4 | 0.43 × 0.20 × 0.20 mm |
| F(000) = 1344 |
Data collection
| Bruker D8 Venture/Photon 100 CMOS diffractometer | 3320 independent reflections |
| Radiation source: fine-focus sealed tube | 3289 reflections with I > 2σ(I) |
| Graphite monochromator | Rint = 0.020 |
| Detector resolution: 10.4167 pixels mm-1 | θmax = 28.4°, θmin = 2.9° |
| φ and ω scans | h = −11→11 |
| Absorption correction: multi-scan (SADABS2014/2; Bruker, 2014) | k = −24→24 |
| Tmin = 0.581, Tmax = 0.746 | l = −22→22 |
| 75009 measured reflections |
Refinement
| Refinement on F2 | Hydrogen site location: inferred from neighbouring sites |
| Least-squares matrix: full | H-atom parameters constrained |
| R[F2 > 2σ(F2)] = 0.010 | w = 1/[σ2(Fo2) + (0.0139P)2 + 1.0428P] where P = (Fo2 + 2Fc2)/3 |
| wR(F2) = 0.027 | (Δ/σ)max = 0.004 |
| S = 1.15 | Δρmax = 0.87 e Å−3 |
| 3320 reflections | Δρmin = −0.30 e Å−3 |
| 178 parameters | Absolute structure: Flack x determined using 1431 quotients [(I+)-(I-)]/[(I+)+(I-)] (Parsons & Flack, 2004) |
| 0 restraints | Absolute structure parameter: −0.0107 (19) |
| Primary atom site location: structure-invariant direct methods |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| Tb1 | 0.5000 | 0.51826 (2) | 0.2500 | 0.01172 (4) | |
| O1 | 0.59854 (16) | 0.54526 (8) | 0.13398 (8) | 0.0167 (3) | |
| O2 | 0.4357 (2) | 0.39605 (9) | 0.30473 (10) | 0.0312 (4) | |
| O3 | 0.5000 | 0.29285 (11) | 0.2500 | 0.0586 (9) | |
| N1 | 0.5000 | 0.35974 (11) | 0.2500 | 0.0293 (6) | |
| N2 | 0.7540 (2) | 0.49068 (9) | 0.32221 (10) | 0.0183 (3) | |
| N3 | 0.66305 (19) | 0.63257 (8) | 0.27777 (9) | 0.0130 (3) | |
| C1 | 0.8272 (3) | 0.42575 (12) | 0.32538 (14) | 0.0249 (4) | |
| H1 | 0.7809 | 0.3851 | 0.2993 | 0.030* | |
| C2 | 0.9672 (2) | 0.41490 (13) | 0.36483 (14) | 0.0275 (5) | |
| H2 | 1.0155 | 0.3681 | 0.3656 | 0.033* | |
| C3 | 1.0342 (2) | 0.47389 (13) | 0.40282 (14) | 0.0253 (5) | |
| H3 | 1.1304 | 0.4685 | 0.4298 | 0.030* | |
| C4 | 0.9590 (2) | 0.54124 (13) | 0.40111 (12) | 0.0203 (4) | |
| H4 | 1.0022 | 0.5823 | 0.4278 | 0.024* | |
| C5 | 0.8200 (2) | 0.54783 (11) | 0.35990 (10) | 0.0151 (3) | |
| C6 | 0.7349 (2) | 0.62034 (12) | 0.35643 (12) | 0.0160 (4) | |
| H6A | 0.6528 | 0.6212 | 0.3975 | 0.019* | |
| H6B | 0.8088 | 0.6606 | 0.3679 | 0.019* | |
| C7 | 0.5652 (2) | 0.69987 (10) | 0.28069 (12) | 0.0156 (3) | |
| H7A | 0.6323 | 0.7432 | 0.2719 | 0.019* | |
| H7B | 0.5190 | 0.7043 | 0.3340 | 0.019* | |
| C8 | 0.7933 (2) | 0.64051 (11) | 0.21953 (12) | 0.0162 (4) | |
| H8A | 0.8566 | 0.5952 | 0.2212 | 0.019* | |
| H8B | 0.8607 | 0.6814 | 0.2372 | 0.019* | |
| C9 | 0.7479 (2) | 0.65450 (12) | 0.13510 (12) | 0.0161 (4) | |
| C10 | 0.6559 (2) | 0.60220 (10) | 0.09514 (11) | 0.0155 (4) | |
| C11 | 0.6273 (2) | 0.61288 (12) | 0.01400 (12) | 0.0188 (4) | |
| H11 | 0.5663 | 0.5782 | −0.0142 | 0.023* | |
| C12 | 0.6873 (3) | 0.67359 (13) | −0.02515 (12) | 0.0232 (4) | |
| H12 | 0.6664 | 0.6801 | −0.0798 | 0.028* | |
| C13 | 0.7777 (3) | 0.72492 (13) | 0.01476 (14) | 0.0251 (4) | |
| H13 | 0.8186 | 0.7663 | −0.0123 | 0.030* | |
| C14 | 0.8073 (2) | 0.71496 (11) | 0.09491 (12) | 0.0199 (4) | |
| H14 | 0.8688 | 0.7498 | 0.1225 | 0.024* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| Tb1 | 0.01260 (5) | 0.01031 (5) | 0.01225 (5) | 0.000 | 0.00008 (7) | 0.000 |
| O1 | 0.0177 (6) | 0.0180 (6) | 0.0145 (6) | −0.0021 (5) | 0.0023 (5) | −0.0009 (5) |
| O2 | 0.0439 (9) | 0.0209 (7) | 0.0287 (8) | −0.0086 (7) | −0.0077 (7) | 0.0083 (6) |
| O3 | 0.096 (2) | 0.0096 (8) | 0.0704 (19) | 0.000 | −0.046 (3) | 0.000 |
| N1 | 0.0418 (14) | 0.0119 (9) | 0.0343 (13) | 0.000 | −0.028 (2) | 0.000 |
| N2 | 0.0170 (8) | 0.0191 (8) | 0.0188 (8) | 0.0028 (7) | −0.0021 (6) | 0.0002 (6) |
| N3 | 0.0124 (7) | 0.0136 (7) | 0.0128 (6) | 0.0003 (6) | −0.0002 (5) | 0.0006 (5) |
| C1 | 0.0244 (10) | 0.0208 (10) | 0.0294 (11) | 0.0046 (8) | −0.0059 (9) | −0.0027 (8) |
| C2 | 0.0250 (14) | 0.0275 (10) | 0.0298 (10) | 0.0106 (8) | −0.0037 (8) | 0.0042 (8) |
| C3 | 0.0180 (13) | 0.0364 (11) | 0.0214 (9) | 0.0023 (8) | −0.0036 (7) | 0.0102 (8) |
| C4 | 0.0176 (11) | 0.0282 (10) | 0.0153 (8) | −0.0033 (7) | −0.0028 (6) | 0.0061 (8) |
| C5 | 0.0147 (8) | 0.0196 (9) | 0.0110 (8) | 0.0001 (7) | 0.0013 (6) | 0.0032 (7) |
| C6 | 0.0162 (9) | 0.0181 (9) | 0.0138 (9) | −0.0009 (8) | −0.0022 (7) | −0.0010 (8) |
| C7 | 0.0161 (8) | 0.0110 (8) | 0.0198 (8) | −0.0006 (7) | −0.0002 (7) | −0.0012 (7) |
| C8 | 0.0124 (8) | 0.0210 (9) | 0.0153 (8) | −0.0023 (7) | 0.0007 (7) | 0.0021 (7) |
| C9 | 0.0136 (9) | 0.0209 (10) | 0.0139 (9) | 0.0017 (8) | 0.0009 (7) | 0.0012 (8) |
| C10 | 0.0130 (8) | 0.0175 (9) | 0.0158 (8) | 0.0028 (7) | 0.0024 (7) | 0.0001 (7) |
| C11 | 0.0181 (9) | 0.0231 (10) | 0.0153 (9) | 0.0024 (8) | 0.0000 (7) | −0.0021 (8) |
| C12 | 0.0266 (11) | 0.0286 (11) | 0.0142 (9) | 0.0029 (9) | −0.0004 (8) | 0.0042 (8) |
| C13 | 0.0303 (11) | 0.0238 (10) | 0.0210 (11) | −0.0032 (9) | 0.0013 (9) | 0.0080 (8) |
| C14 | 0.0196 (9) | 0.0208 (9) | 0.0194 (10) | −0.0012 (8) | 0.0002 (8) | 0.0022 (8) |
Geometric parameters (Å, º)
| Tb1—O1i | 2.1947 (13) | C3—H3 | 0.9500 |
| Tb1—O1 | 2.1947 (13) | C4—C5 | 1.389 (3) |
| Tb1—O2i | 2.4764 (15) | C4—H4 | 0.9500 |
| Tb1—O2 | 2.4764 (15) | C5—C6 | 1.513 (3) |
| Tb1—N2i | 2.5521 (17) | C6—H6A | 0.9900 |
| Tb1—N2 | 2.5521 (17) | C6—H6B | 0.9900 |
| Tb1—N3i | 2.5558 (16) | C7—C7i | 1.529 (4) |
| Tb1—N3 | 2.5558 (16) | C7—H7A | 0.9900 |
| Tb1—N1 | 2.891 (2) | C7—H7B | 0.9900 |
| O1—C10 | 1.324 (2) | C8—C9 | 1.503 (3) |
| O2—N1 | 1.266 (2) | C8—H8A | 0.9900 |
| O3—N1 | 1.220 (3) | C8—H8B | 0.9900 |
| N1—O2i | 1.266 (2) | C9—C14 | 1.393 (3) |
| N2—C1 | 1.342 (3) | C9—C10 | 1.412 (3) |
| N2—C5 | 1.347 (3) | C10—C11 | 1.409 (3) |
| N3—C6 | 1.485 (2) | C11—C12 | 1.390 (3) |
| N3—C7 | 1.489 (2) | C11—H11 | 0.9500 |
| N3—C8 | 1.498 (2) | C12—C13 | 1.391 (3) |
| C1—C2 | 1.391 (3) | C12—H12 | 0.9500 |
| C1—H1 | 0.9500 | C13—C14 | 1.392 (3) |
| C2—C3 | 1.380 (3) | C13—H13 | 0.9500 |
| C2—H2 | 0.9500 | C14—H14 | 0.9500 |
| C3—C4 | 1.388 (3) | ||
| O1i—Tb1—O1 | 154.07 (7) | C8—N3—Tb1 | 111.56 (11) |
| O1i—Tb1—O2i | 128.52 (6) | N2—C1—C2 | 123.4 (2) |
| O1—Tb1—O2i | 77.36 (6) | N2—C1—H1 | 118.3 |
| O1i—Tb1—O2 | 77.36 (6) | C2—C1—H1 | 118.3 |
| O1—Tb1—O2 | 128.52 (6) | C3—C2—C1 | 118.3 (2) |
| O2i—Tb1—O2 | 51.64 (9) | C3—C2—H2 | 120.9 |
| O1i—Tb1—N2i | 98.20 (5) | C1—C2—H2 | 120.9 |
| O1—Tb1—N2i | 86.89 (5) | C2—C3—C4 | 119.11 (19) |
| O2i—Tb1—N2i | 80.45 (6) | C2—C3—H3 | 120.4 |
| O2—Tb1—N2i | 79.10 (6) | C4—C3—H3 | 120.4 |
| O1i—Tb1—N2 | 86.89 (5) | C3—C4—C5 | 119.2 (2) |
| O1—Tb1—N2 | 98.20 (5) | C3—C4—H4 | 120.4 |
| O2i—Tb1—N2 | 79.10 (6) | C5—C4—H4 | 120.4 |
| O2—Tb1—N2 | 80.45 (6) | N2—C5—C4 | 122.21 (19) |
| N2i—Tb1—N2 | 157.26 (8) | N2—C5—C6 | 117.05 (16) |
| O1i—Tb1—N3i | 76.70 (5) | C4—C5—C6 | 120.74 (19) |
| O1—Tb1—N3i | 82.18 (5) | N3—C6—C5 | 111.54 (16) |
| O2i—Tb1—N3i | 141.99 (5) | N3—C6—H6A | 109.3 |
| O2—Tb1—N3i | 132.91 (6) | C5—C6—H6A | 109.3 |
| N2i—Tb1—N3i | 66.65 (5) | N3—C6—H6B | 109.3 |
| N2—Tb1—N3i | 135.88 (5) | C5—C6—H6B | 109.3 |
| O1i—Tb1—N3 | 82.18 (5) | H6A—C6—H6B | 108.0 |
| O1—Tb1—N3 | 76.70 (5) | N3—C7—C7i | 113.05 (13) |
| O2i—Tb1—N3 | 132.91 (6) | N3—C7—H7A | 109.0 |
| O2—Tb1—N3 | 141.99 (5) | C7i—C7—H7A | 109.0 |
| N2i—Tb1—N3 | 135.88 (5) | N3—C7—H7B | 109.0 |
| N2—Tb1—N3 | 66.65 (5) | C7i—C7—H7B | 109.0 |
| N3i—Tb1—N3 | 70.67 (7) | H7A—C7—H7B | 107.8 |
| O1i—Tb1—N1 | 102.97 (4) | N3—C8—C9 | 116.63 (16) |
| O1—Tb1—N1 | 102.97 (4) | N3—C8—H8A | 108.1 |
| O2i—Tb1—N1 | 25.82 (4) | C9—C8—H8A | 108.1 |
| O2—Tb1—N1 | 25.82 (4) | N3—C8—H8B | 108.1 |
| N2i—Tb1—N1 | 78.63 (4) | C9—C8—H8B | 108.1 |
| N2—Tb1—N1 | 78.63 (4) | H8A—C8—H8B | 107.3 |
| N3i—Tb1—N1 | 144.67 (4) | C14—C9—C10 | 120.42 (18) |
| N3—Tb1—N1 | 144.67 (4) | C14—C9—C8 | 120.25 (19) |
| C10—O1—Tb1 | 139.80 (12) | C10—C9—C8 | 119.08 (19) |
| N1—O2—Tb1 | 95.73 (12) | O1—C10—C11 | 121.83 (18) |
| O3—N1—O2 | 121.55 (11) | O1—C10—C9 | 120.05 (17) |
| O3—N1—O2i | 121.55 (11) | C11—C10—C9 | 118.13 (18) |
| O2—N1—O2i | 116.9 (2) | C12—C11—C10 | 120.67 (19) |
| O3—N1—Tb1 | 180.0 | C12—C11—H11 | 119.7 |
| O2—N1—Tb1 | 58.45 (11) | C10—C11—H11 | 119.7 |
| O2i—N1—Tb1 | 58.45 (11) | C11—C12—C13 | 120.8 (2) |
| C1—N2—C5 | 117.83 (17) | C11—C12—H12 | 119.6 |
| C1—N2—Tb1 | 126.43 (14) | C13—C12—H12 | 119.6 |
| C5—N2—Tb1 | 115.74 (12) | C12—C13—C14 | 119.2 (2) |
| C6—N3—C7 | 109.19 (15) | C12—C13—H13 | 120.4 |
| C6—N3—C8 | 107.09 (15) | C14—C13—H13 | 120.4 |
| C7—N3—C8 | 111.34 (15) | C13—C14—C9 | 120.8 (2) |
| C6—N3—Tb1 | 105.69 (12) | C13—C14—H14 | 119.6 |
| C7—N3—Tb1 | 111.67 (11) | C9—C14—H14 | 119.6 |
| Tb1—O2—N1—O3 | 180.000 (1) | Tb1—N3—C7—C7i | −38.2 (2) |
| Tb1—O2—N1—O2i | 0.000 (1) | C6—N3—C8—C9 | −179.19 (19) |
| C5—N2—C1—C2 | −0.5 (3) | C7—N3—C8—C9 | −59.9 (2) |
| Tb1—N2—C1—C2 | 178.82 (17) | Tb1—N3—C8—C9 | 65.61 (19) |
| N2—C1—C2—C3 | 0.2 (4) | N3—C8—C9—C14 | 125.3 (2) |
| C1—C2—C3—C4 | 0.8 (3) | N3—C8—C9—C10 | −60.3 (3) |
| C2—C3—C4—C5 | −1.3 (3) | Tb1—O1—C10—C11 | −142.18 (16) |
| C1—N2—C5—C4 | 0.0 (3) | Tb1—O1—C10—C9 | 37.5 (3) |
| Tb1—N2—C5—C4 | −179.43 (14) | C14—C9—C10—O1 | −179.29 (18) |
| C1—N2—C5—C6 | −179.33 (18) | C8—C9—C10—O1 | 6.3 (3) |
| Tb1—N2—C5—C6 | 1.2 (2) | C14—C9—C10—C11 | 0.4 (3) |
| C3—C4—C5—N2 | 0.9 (3) | C8—C9—C10—C11 | −174.00 (18) |
| C3—C4—C5—C6 | −179.80 (18) | O1—C10—C11—C12 | 179.21 (19) |
| C7—N3—C6—C5 | 173.14 (16) | C9—C10—C11—C12 | −0.5 (3) |
| C8—N3—C6—C5 | −66.2 (2) | C10—C11—C12—C13 | 0.3 (3) |
| Tb1—N3—C6—C5 | 52.88 (17) | C11—C12—C13—C14 | −0.2 (4) |
| N2—C5—C6—N3 | −38.3 (2) | C12—C13—C14—C9 | 0.1 (3) |
| C4—C5—C6—N3 | 142.41 (18) | C10—C9—C14—C13 | −0.2 (3) |
| C6—N3—C7—C7i | −154.7 (2) | C8—C9—C14—C13 | 174.1 (2) |
| C8—N3—C7—C7i | 87.2 (2) |
Symmetry code: (i) −x+1, y, −z+1/2.
Hydrogen-bond geometry (Å, º)
| D—H···A | D—H | H···A | D···A | D—H···A |
| C8—H8B···O3ii | 0.99 | 2.37 | 3.338 (3) | 166 |
Symmetry code: (ii) x+1/2, y+1/2, z.
References
- Anjos, A. dos, Bortoluzzi, A. J., Caro, M. S. B., Peralta, R. A., Friedermann, G. R., Mangrich, A. S. & Neves, A. (2006). J. Braz. Chem. Soc. 17, 1540–1550.
- Bruker (2010). APEX2 and SAINT. Bruker AXS Inc., Madison, Wisconsin, USA.
- Bruker (2014). SADABS. Bruker AXS Inc., Madison, Wisconsin, USA.
- Chilton, N. F., Langley, S. K., Moubaraki, B., Soncini, A., Batten, S. R. & Murray, K. S. (2013). Chem. Sci. 4, 1719–1730.
- Colotti, G., Ilari, A., Boffi, A. & Morea, V. (2013). Mini Rev. Med. Chem. 13, 211–221. [PubMed]
- Diehl, H., Hach, C. C. & Bailar, J. C. (2007). Inorganic Synthesis, pp. 196–201. New York: John Wiley & Sons Inc.
- Farrugia, L. J. (2012). J. Appl. Cryst. 45, 849–854.
- Golchoubian, H. & Gholamnezhad, P. (2009). X-Ray Struct. Anal. Online, 25, 95–96.
- Hancock, R. D. (2013). Chem. Soc. Rev. 42, 1500–1524.
- Happe, T. & Hemschemeier, A. (2014). Trends Biotechnol. 32, 170–176. [DOI] [PubMed]
- Johnson, C. K. (1976). ORTEPII. Report ORNL-5138. Oak Ridge National Laboratory, Tennessee, USA.
- Lanznaster, M., Neves, A., Bortoluzzi, A. J., Assumpção, A. M. C., Vencato, I., Machado, S. P. & Drechsel, S. M. (2006). Inorg. Chem. 45, 1005–1011. [DOI] [PubMed]
- Lenze, M., Sedinkin, S. L. & Bauer, E. B. (2013). J. Mol. Catal. A Chem. 373, 161–171.
- Luzon, J. & Sessoli, R. (2012). Dalton Trans. 41, 13556–13567. [DOI] [PubMed]
- Neves, A., Ceccato, A. S., Erthal, S. M. D., Vencato, I., Nuber, B. & Weiss, J. (1991). Inorg. Chim. Acta, 187, 119–121.
- Neves, A., Ceccatto, A. S., Erasmus-Buhr, C., Gehring, S., Haase, W., Paulus, H., Nascimento, O. R. & Batista, A. A. (1993). J. Chem. Soc. Chem. Commun. pp. 1782–1784.
- Neves, A., Erthal, S. M. D., Vencato, I., Ceccato, A. S., Mascarenhas, Y. P., Nascimento, O. R., Horner, M. & Batista, A. A. (1992). Inorg. Chem. 31, 4749–4755.
- Nurchi, V. M., Crespo-Alonso, M., Toso, L., Lachowicz, J. I. & Crisponi, G. (2013). Mini Rev. Med. Chem. 13, 1541–1549. [DOI] [PubMed]
- Parsons, S. & Flack, H. (2004). Acta Cryst. A60, s61.
- Rinehart, J. D. & Long, J. R. (2011). Chem. Sci. 2, 2078–2085.
- Schwingel, E. W., Arend, K., Zarling, J., Neves, A. & Szpoganicz, B. (1996). J. Braz. Chem. Soc. 7, 31–37.
- Sears, M. E. (2013). Sci. World J., article no. 219840.
- Sessoli, R. & Powell, A. K. (2009). Coord. Chem. Rev. 253, 2328–2341.
- Setyawati, I. A., Liu, S., Rettig, S. J. & Orvig, C. (2000). Inorg. Chem. 39, 496–507. [DOI] [PubMed]
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Thomas, F., Arora, H., Philouze, C. & Jarjayes, O. (2010). Inorg. Chim. Acta, 363, 3122–3130.
- Totaro, P., Westrup, K. C. M., Boulon, M.-E., Nunes, G. G., Back, D. F., Barison, A., Ciattini, S., Mannini, M., Sorace, L., Soares, J. F., Cornia, A. & Sessoli, R. (2013). Dalton Trans. 42, 4416–4426. [DOI] [PubMed]
- Ungur, L., Le Roy, J. J., Korobkov, I., Murugesu, M. & Chibotaru, L. F. (2014). Angew. Chem. Int. Ed. 53, 4413–4417. [DOI] [PubMed]
- Westrup, K. C. M., Boulon, M.-E., Totaro, P., Nunes, G. G., Back, D. F., Barison, A., Jackson, M., Paulsen, C., Gatteschi, D., Sorace, L., Cornia, A., Soares, J. F. & Sessoli, R. (2014). Chem. Eur. J. 20, 13681–13691. [DOI] [PubMed]
- Wong, E., Caravan, P., Liu, S., Rettig, S. J. & Orvig, C. (1996). Inorg. Chem. 35, 715–724.
- Wong, E., Liu, S., Rettig, S. & Orvig, C. (1995). Inorg. Chem. 34, 3057–3064.
- Xu, L., Setyawati, I. A., Pierreroy, J., Pink, M., Young, V. G., Patrick, B. O., Rettig, S. J. & Orvig, C. (2000). Inorg. Chem. 39, 5958–5963. [DOI] [PubMed]
- Yamada, Y., Takenouchi, S. I., Miyoshi, Y. & Okamoto, K. I. (2010). J. Coord. Chem. 63, 996–1012.
- Zhang, P., Zhang, L., Wang, C., Xue, S. F., Lin, S. Y. & Tang, J. K. (2014). J. Am. Chem. Soc. 136, 4484–4487. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablock(s) I. DOI: 10.1107/S2056989014026826/lh5741sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S2056989014026826/lh5741Isup2.hkl
CCDC reference: 1037922
Additional supporting information: crystallographic information; 3D view; checkCIF report




