Abstract
The family Rickettsiaceae currently includes two genera: Orientia that contains one species, Orientiatsutsugamushi, and Rickettsia that contains 28 species.
Occidentia massiliensis gen. nov., sp. nov. strain OS118T is the type strain of O. massiliensis gen. nov., sp. nov., the type species of the new genus Occidentia gen. nov. within the family Rickettsiaceae. This strain, whose genome is described here, was isolated in France from the soft tick Ornithodoros sonrai collected in Senegal. O. massiliensis is an aerobic, rod-shaped, Gram-negative, obligate intracellular bacillus that may be cultivated in BME/CTVM2 cells.
Here we describe the features of O. massiliensis, together with the complete genomic sequencing and annotation. The 1,469,252 bp long genome (1 chromosome but no plasmid) contains 1,670 protein-coding and 41 RNA genes, including one rRNA operon.
Keywords: Occidentia massiliensis, Genome, Senegal, Soft tick, Ornithodoros sonrai, Taxonogenomics
Introduction
Occidentia massiliensis gen. nov., sp. nov. strain OS18T is the type strain of O. massiliensis gen. nov., sp. nov. This bacterium was isolated from an Ornithodoros sonrai tick collected in Senegal. It is an aerobic, rod-shaped, Gram-negative, obligate intracellular bacillus.
The family Rickettsiaceae Pinkerton, [1] currently includes two genera: Orientia Tamura et al.[2] that contains one species, Orientia tsutsugamushi (Hayashi 1920, Tamura et al. 1995), and Rickettsia (da Rocha-Lima 1916) that contains 28 species [3]. Many members of this family have been detected and identified by PCR only, and have yet to be validly published [4,5]. The family Rickettsiaceae is composed of obligate intracellular bacteria that infect the cytoplasm and sometimes the nucleus of eukaryotic cells within which they live freely [6]. In addition, both Rickettsia species and O. tsutsugamushi are rod-shaped or coccoid, Gram-negative, bacteria intimately associated with arthropod hosts [7]. To date, none of the members of this family has been cultivated axenically. Many validly published species within the family Rickettsiaceae are pathogenic for humans and other vertebrates, causing spotted fevers or various forms of typhus. These diseases are transmitted by arthropods (mostly ticks, mites, lice or fleas). Other species of undescribed pathogenicity have only been detected in arthropods. Phylogenetically, Occidentia massiliensis gen. nov., sp. nov., is most closely related to Orientia tsutsugamushi (Figure 1) [2]. By comparison with Rickettsia species, O. tsutsugamushi differs in outer envelope layers [8], antigenic properties and by the absence of peptidoglycans and lipopolysaccharides. These phenotypic differences are supported by a 16S rRNA nucleotide sequence identity < 90.6%. In 2003, Fournier et al. developed genetic criteria to classify rickettsial isolates based on comparison of 16S rRNA and other genes [9]. The development of this strategy, combining sequences from several genes, notably housekeeping genes, was motivated by the small number of phenotypic criteria available for these strictly intracellular bacteria.
Figure 1.
A consensus phylogenetic tree showing the position of Occidentia massiliensis strain OS18T relative to other strains within the family Rickettsiaceae, based on 16S rDNA sequence comparison. GenBank accession numbers are indicated in parentheses. Sequences were aligned using CLUSTALW, and phylogenetic inferences obtained using the Bayesian phylogenetic analysis [10] with the TOPALi 2.5 software (Biomathematics and Statistics Scotland, Edinburgh, UK) within the integrated MrBayes application [11], using the HKY85 + Г substitution model. Numbers at the nodes are percentages of bootstrap values from 100 replicates. Anaplasma ovis was used as the outgroup. The scale bar represents a 5% nucleotide sequence divergence.
Here we present a summary classification and a set of features for O. massiliensis gen. nov., sp. nov., strain OS18T (CSUR = P764, DSM = 24860) together with the description of the complete genomic sequencing and annotation. These characteristics support the circumscription of the genus Occidentia and its type species, O. massiliensis within the Rickettsiaceae family.
Organism information
In June 2009, 20 adult Ornithodoros sonrai soft ticks were collected from rodent burrows in the Soulkhou Thissé village (a rural village in the Guinean-Sudanian zone in Senegal, close to the city of Tambacounda) as part of a prospective study on tick-borne relapsing fever in West Africa. Ticks were kept alive until further processed. All ticks were disinfected, ground in Rinaldini solution and inoculated in a tick cell line monolayer (BME/CTVM2 cells from Rhipicephalus microplus) [12] using the shell-vial technique [13]. Shell vials were further incubated in an aerobic atmosphere at 28°C. Culture medium (Leibovitz-15 medium supplemented with 10% heat-inactivated fetal calf serum, 10% tryptose phosphate broth and 2 mM glutamine) was changed once a week. Strain OS18 (Table 1) was isolated in 2009 after one month of culture (Figure 2).
Table 1.
Classification and general features of Occidentia massiliensis strain OS18T according to the MIGS recommendations [14]
| MIGS ID | Property | Term | Evidence codea |
|---|---|---|---|
| |
Current classification |
Domain Bacteria |
TAS [15] |
| |
Phylum Proteobacteria |
TAS [16] |
|
| |
Class Alphaproteobacteria |
TAS [17] |
|
| |
Order Rickettsiales |
TAS [6,18,19] |
|
| |
Family Rickettsiaceae |
TAS [1,18,19] |
|
| |
Genus Occidentia |
IDA |
|
| |
Species Occidentia massiliensis |
IDA |
|
| |
Type strain OS18T |
IDA |
|
| |
Gram stain |
Negative |
IDA |
| |
Cell shape |
Rod |
IDA |
| |
Motility |
Unknown |
IDA |
| |
Sporulation |
Nonsporulating |
IDA |
| |
Temperature range |
Mesophilic |
IDA |
| |
Optimum temperature |
28°C |
IDA |
| MIGS-6.3 |
Salinity |
Unknown |
IDA |
| MIGS-22 |
Oxygen requirement |
Aerobic |
IDA |
| |
Carbon source |
Unknown |
NAS |
| |
Energy source |
Unknown |
NAS |
| MIGS-6 |
Habitat |
Ornithodoros sonrai |
IDA |
| MIGS-15 |
Biotic relationship |
Obligate intracellular |
IDA |
| |
Pathogenicity |
Unknown |
|
| |
Biosafety level |
2 |
|
| MIGS-14 |
Isolation |
Ornithodoros sonrai |
IDA |
| MIGS-4 |
Geographic location |
Senegal |
IDA |
| MIGS-5 |
Sample collection time |
June 2009 |
IDA |
| MIGS-4.1 |
Latitude |
14.05 |
IDA |
| MIGS-4.2 |
Longitude |
−15.516667 |
IDA |
| MIGS-4.3 |
Depth |
0.5 m below surface |
IDA |
| MIGS-4.4 | Altitude | 45 m above sea level | IDA |
aEvidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample but based on a generally accepted property for the species or anecdotal evidence). Evidence codes come from the Gene Ontology project [20]. If the evidence is IDA, then the property was directly observed for a live isolate by one of the authors or an expert mentioned in the acknowledgements.
Figure 2.

Gimenez stain of O. massiliensis strain OS18 T grown in BME/CTVM2 cells.
Five other morphologically and genetically indistinguishable isolates were recovered from five other O. sonrai ticks from the same batch. The 16S rRNA nucleotide sequence (GenBank accession number GU937608) of Occidentia massiliensis strain OS18T was 93.7% similar to Orientia tsutsugamushi strain Gilliam (GenBank accession number D38622), the phylogenetically closest species, but formed a separate, well-supported (bootstrap value 98%) sister branch to the O. tsutsugamushi species (Figure 1). This value was lower than the 95% 16S rRNA gene sequence threshold recommended by Stackebrandt and Elbers to delineate a new genus without carrying out DNA-DNA hybridization [21].
Growth was attempted at 28°C in an aerobic atmosphere, which were the culture conditions required for the BME/CTVM2 cell line, and at 37°C in an aerobic atmosphere in L929 and XTC cell lines. Bacteria grew in BME/CTVM2 cells but no growth was obtained in L929 and XTC cell lines. Bacterial cells grown inside BME/CTVM2 cells were Gimenez-positive but weakly Gram-negative. Scanning electron microscopy revealed that cells were rod-shaped with one “rounded” end and another “blunt” end (Figure 3). A monotrichous flagellum was attached to the “blunt” end. Cells had a mean length and width of 1.23 ± 0.19 μm and 0.42 ± 0.06 μm, respectively. Bacteria were abundant within the cytoplasm but not the nucleus of tick cells (Figure 2). Typically, the highest concentration of bacteria was seen around mitochondria within cells (Figure 4). Contrary to Rickettsia species and O. tsutsugamushi, we did not identify evident differences between the inner and outer leaflets of the cell wall (Figure 5), although the periplasmic space was unusually large (0.028 ± 0.007 μm).
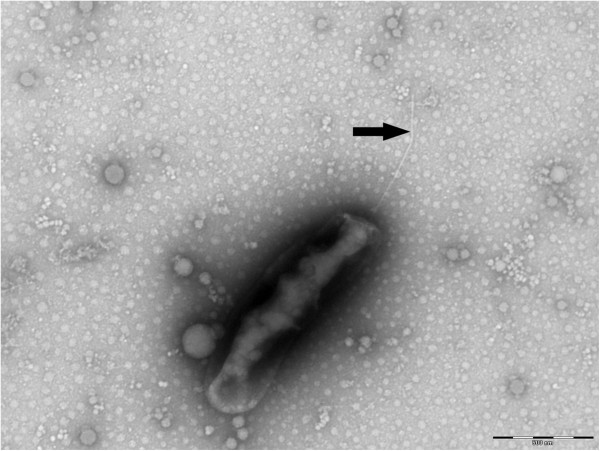
Figure 3.
Scanning electron micrograph of O. massiliensis strain OS18T, made using a Morgagni 268D (Philips) transmission electron microscope at an operating voltage of 60 kV. The arrow shows a monotrichous flagellum. The scale bar represents 500 nm.
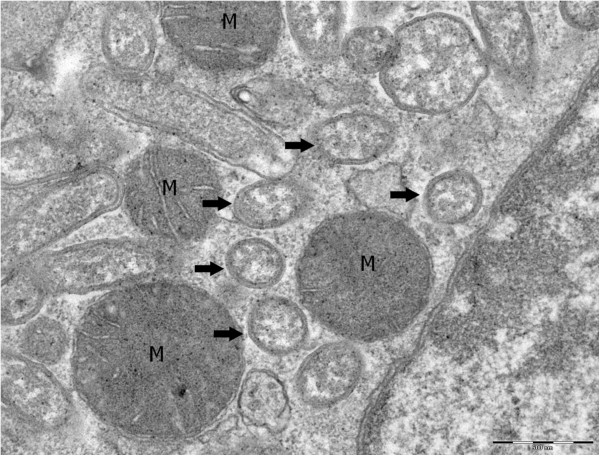
Figure 4.
Transmission electron micrograph of O. massiliensis strain OS18T grown in BME/CTVM2 cells made using uranyl acetate staining and a Morgagni 268D (Philips) at an operating voltage of 60 kV. The bacteria (arrows) characteristically surround mitochondria (M). The scale bar represents 500 nm.
Figure 5.
Transmission electron micrograph of a single O. massiliensis strain OS18 T cell made using uranyl acetate staining and a Morgagni 268D (Philips) at an operating voltage of 60 kV. The scale bar represents 500 nm.
Genome sequencing information
Genome project history
The organism was selected for sequencing on the basis of its phylogenetic position and 16S rRNA similarity to members of the family Rickettsiaceae. Nucleotide sequence similarity levels of these genes suggested that strain OS18T represents a new genus within the family Rickettsiaceae. It was the first genome of Occidentia massiliensis gen. nov., sp. nov. The Genbank accession number is CANJ00000000 and consists of 47 large contigs (>1.5 kb) in 18 scaffolds. Table 2 shows the project information and its association with MIGS version 2.0 compliance [14].
Table 2.
Project information
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 |
Finishing quality |
High-quality draft |
| MIGS-28 |
Libraries used |
One paired-end 3-kb library |
| MIGS-29 |
Sequencing platforms |
454 GS FLX Titanium |
| MIGS-31.2 |
Fold coverage |
23.2× |
| MIGS-30 |
Assemblers |
Newbler version 2.5.3 |
| MIGS-32 |
Gene calling method |
Prodigal |
| |
GenBank ID |
CANJ00000000 |
| |
GenBank Date of Release |
April 15, 2013 |
| MIGS-13 | Project relevance | Biodiversity of the Ornithodoros sonrai tick microbial flora |
Growth conditions and DNA isolation
O. massiliensis gen. nov., sp. nov., strain OS18T (CSUR = P764, DSM = 24860) was grown aerobically in BME/CTVM2 cell line at 28°C. Infected cells were harvested from 20 culture flasks. Bacterial purification using a renografin gradient was performed as previously described [22]. A total of 200 μL of bacterial suspension was diluted in 1 ml TE buffer for lysis treatment. After incubation with 2.5 μg/μL lysozyme for 30 minutes at 37°C, the lysis was performed with 1% laurylsarcosyl and 50 μg/μL RNAse A for 1 hr at 37°C, followed by an overnight incubation at 37°C with proteinase K. The DNA was purified three times by phenol-chloroform extraction and then precipitated by addition of ethanol at −20°C overnight. After centrifugation, the DNA was resuspended in 199 μL TE buffer. The DNA concentration was measured by the Quant-it Picogreen kit (Invitrogen) on the Genios-Tecan fluorometer at 69.12 ng/μl.
Genome sequencing and assembly
A 3 kb paired-end sequencing strategy (454 GS FLX Titanium, Roche) was selected. DNA (5 μg) was mechanically fragmented on the Covaris device (KBioScience-LGC Genomics, Teddington, UK) through miniTube-Red 5 kb with an enrichment size of 3–4 kb. The DNA fragmentation was visualized using the Agilent 2100 BioAnalyzer on a DNA labchip 7500 with an average size of 3.2 kb. Circularization and nebulization were performed and generated a pattern with an average size of 580 bp. After PCR amplification over 17 cycles followed by double size selection, the single-stranded paired-end library was then quantified on the Genios-Tecan fluorometer with the Quant-iT ribogreen (Invitrogen) at 1,120 pg/μL. The library concentration equivalence was calculated as 3.55 × 109 molecules/μL. The library was stored at −20°C until further use.
The library was clonally amplified with 0.7 cpb in 4 emPCR reactions, with the GS Titanium SV emPCR Kit (Lib-L) v2 (Roche). The yield was calculated at 9.16%, within the recommended yield range of between 5 and 20% from the Roche procedure. After amplification, 790,000 beads from the emPCR reaction were loaded on a ¼ region on the GS Titanium PicoTiterPlate PTP Kit 70 × 75 and sequenced with the GS FLX Titanium Sequencing Kit XLR70 (Roche). The run was analyzed on the cluster through the gsRunBrowser and Newbler assembler (Roche). A total of 103,355 passed filter wells were obtained and generated 34.1 Mb of DNA sequence with an average read length of 330 bp.
The passed filter sequences were assembled using Newbler with 90% identity and 40 bp for overlap requirements. The final assembly identified 18 scaffolds and 47 large contigs (>1.5 kb) generating a genome size of 1,47 Mb which corresponds to a coverage of 23.2×.
Genome annotation
Open reading frames were predicted using PRODIGAL with default parameters [23], but predicted ORFs were excluded if they spanned a sequencing gap region. The functional annotation of protein sequences was performed using BLASTP against the GenBank and Clusters of Orthologous Groups (COG) databases [24]. The prediction of tRNAs and rRNAs was carried out using the tRNAScan-SE [25] and RNAmmer [26] tools, respectively. Lipoprotein signal peptides and numbers of transmembrane helices were predicted using SignalP [27] and TMHMM [28], respectively. ORFans were identified if their BLASTP E-value was lower than 1e-03 for alignment length greater than 80 amino acids. If alignment lengths were smaller than 80 amino acids, we used an E-value of 1e-05. Such parameter thresholds have already been used in previous works to define ORFans. To estimate the mean level of nucleotide sequence similarity at the genome level between O. massiliensis and another 4 members of the family Rickettsiaceae (Table 3), we used the Average Genomic Identity Of gene Sequences (AGIOS) home-made software [29]. Briefly, this software combines the Proteinortho software [30] for detecting orthologous proteins in pairwise comparisons of genomes, then retrieves the corresponding genes and determines the mean percentage of nucleotide sequence identity among orthologous ORFs using the Needleman-Wunsch global alignment algorithm.
Table 3.
Nucleotide content and gene count levels of the genome
| Attribute | Value | % of totala |
|---|---|---|
| Size (bp) |
1,469,252 |
100 |
| DNA G + C content (bp) |
426,780 |
29.05 |
| Total genes |
1,670 |
100 |
| RNA genes |
41 |
2.66 |
| Protein-coding genes |
1,502 |
97.34 |
| Protein with predicted function (COGs + NR) |
1,099 |
73.17 |
| Genes assigned to COG |
1,062 |
70.71 |
| Genes with peptide signal |
131 |
8.72 |
| Genes with transmembrane helices (≥3) | 331 | 22.04 |
aThe total is based on either the size of the genome in base pairs or the total number of protein coding genes in the annotated genome.
Genome properties
The genome is 1,469,252 bp long (one chromosome, no plasmid) with a 29.05% GC content (Table 4). It is composed of 301 contigs (18 scaffolds). Of the 1,543 predicted genes, 1,502 were protein-coding genes, and 41 were RNAs (1 rRNA operon and 38 tRNA genes).
Table 4.
Number of genes associated with the 25 general COG functional categories
| Code | Value | %age | Description |
|---|---|---|---|
| J |
158 |
10.52 |
Translation |
| A |
0 |
0 |
RNA processing and modification |
| K |
40 |
2.66 |
Transcription |
| L |
107 |
7.12 |
Replication, recombination and repair |
| B |
0 |
0 |
Chromatin structure and dynamics |
| D |
20 |
1.33 |
Cell cycle control, mitosis and meiosis |
| Y |
0 |
0 |
Nuclear structure |
| V |
17 |
1.13 |
Defense mechanisms |
| T |
32 |
2.13 |
Signal transduction mechanisms |
| M |
114 |
7.59 |
Cell wall/membrane biogenesis |
| N |
2 |
0.13 |
Cell motility |
| Z |
0 |
0 |
Cytoskeleton |
| W |
0 |
0 |
Extracellular structures |
| U |
64 |
4.26 |
Intracellular trafficking and secretion |
| O |
72 |
4.79 |
Posttranslational modification, protein turnover and chaperones |
| C |
94 |
6.26 |
Energy production and conversion |
| G |
45 |
3 |
Carbohydrate transport and metabolism |
| E |
90 |
5.99 |
Amino acid transport and metabolism |
| F |
20 |
1.33 |
Nucleotide transport and metabolism |
| H |
40 |
2.66 |
Coenzyme transport and metabolism |
| I |
29 |
1.93 |
Lipid transport and metabolism |
| P |
52 |
3.46 |
Inorganic ion transport and metabolism |
| Q |
14 |
0.93 |
Secondary metabolites biosynthesis, transport and catabolism |
| R |
146 |
9.72 |
General function prediction only |
| S |
53 |
3.52 |
Function unknown |
| X | 440 | 29.29 | Not in COGs |
The total is based on the total number of protein-coding genes in the annotated genome.
A total of 1,099 genes (73.17%) were assigned a putative function (by COG or by NR blast), and 185 genes were identified as ORFans (12.32%). The remaining genes were annotated as hypothetical proteins (165 genes = > 10.99%). The distribution of genes into COGs functional categories is presented in Table 5 and Figure 6. The properties and the statistics of the genome are summarized in Tables 4 and 5.
Table 5.
Bacterial genomes used for the genomic comparison
| Species | Numberof proteins | Genome Size (Mb) | G + C content |
|---|---|---|---|
|
O. massiliensis strain OS18T |
1502 |
1.47 |
29.0 |
|
O. tsutsugamushi strain Boryong |
1182 |
2.13 |
30.5 |
|
O. tsutsugamushi strain Ikeda |
1967 |
2.01 |
30.5 |
|
R. bellii strain RML369-C |
1428 |
1.52 |
31.6 |
| R. prowazekii strain Madrid E | 842 | 1.1 | 29.0 |
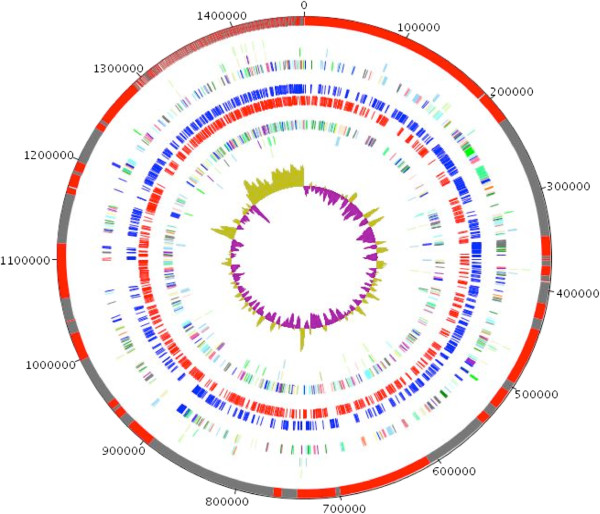
Figure 6.
Graphical circular map of the chromosome. From outside in: contigs (red / grey), COG category of genes on the forward strand (three circles), genes on forward strand (blue circle), genes on the reverse strand (red circle) and COG category of genes on the reverse strand (three circles). The inner-most circle shows the G + C skew, with purple and olive indicating negative and positive values, respectively.
Genomic comparison of O. massiliensis and other members of the family Rickettsiaceae
We compared the genome sequence of O. massiliensis strain OS18T to those of O. tsutsugamushi strains Boryong (GenBank accession number NC_009488) and Ikeda (NC_010793), and R. prowazekii strain Madrid E (NC_000963) and R. bellii strain RML369-C (NC_007940). Occidentia massiliensis strain OS18T had a much smaller genome (1,469,252 bp, 2,127,051 bp and 2,008,987 bp, respectively), fewer genes (1,670, 2,216 and 2,005 genes, respectively) and a lower G + C content (29.05%, 30.5%, 30.5% than O. tsutsugamushi strains Boryong and Ikeda (Table 6). However, when compared to Rickettsia species, O. massiliensis had a larger genome than R. prowazekii (1,469,252 bp and, 1,111,523 bp, respectively), but smaller than R. bellii (1,522,076 bp). In contrast, the G + C content of O. massiliensis was identical to that of R. prowazekii (29%) but lower than R. bellii (31.6%). In addition, O. massiliensis exhibited AGIOS values of 73.58 and 73.62% when compared to O. tsutsugamushi strains Boryong and Ikeda, respectively, higher than those obtained by comparison with R. bellii and R. prowazekii (68.7 and 69.45%, respectively, Table 3). However, these values were lower than those obtained between O. tsutsugamushi strains (97.49%) and Rickettsia species (81.57%), but similar to those obtained by comparison of Orientia and Rickettsia genomes (67.48 to 68.08%, Table 3), thus confirming the new genus status of O. massiliensis.
Table 6.
Genomic Comparison of O. massiliensis strain OS18T with other members of the family Rickettsiaceae
| Species | O. massiliensis | O. tsutsugamushi Boryong | O. tsutsugamushi Ikeda | R. bellii | R. prowazekii |
|---|---|---|---|---|---|
|
O. massiliensis |
1,502 |
73.58 |
73.62 |
68.70 |
69.45 |
|
O. tsutsugamushi Boryong |
564 |
1,182 |
97.49 |
67.48 |
68.06 |
|
O. tsutsugamushi Ikeda |
572 |
592 |
1,967 |
67.58 |
68.08 |
|
R. bellii |
603 |
502 |
507 |
1,428 |
81.57 |
| R. prowazekii | 588 | 482 | 489 | 598 | 842 |
Upper right, AGIOS values; lower left, number of orthologous proteins; bold number indicate the numbers of protein-coding genes.
Conclusions
Strain OS18 shares a maximum 93.76% 16S rRNA identity with O. tsutsugamushi, its closest phylogenetic neighbor, and 91.61% with R. prowazekii. These values are lower than the 95% threshold proposed by Stackebrandt and Ebers to delineate genera [21]. In addition, the genomic comparison of O. massiliensis and members of two genera from the family Rickettsiaceae demonstrated that the former species exhibits AGIOS values similar to those obtained by comparison of genera, but much lower than those obtained by intra-genus strain comparison.
Moreover, the morphological (monotrichous flagellum, weak coloration by Gimenez staining, concentration around the mitochondria inside infected cell, large periplasmatic space) and epidemiological (association with soft ticks, inability to grow in L929 and XTC cell lines) evidence also differentiates strain OS18 from other members of the family Rickettsiaceae.
On the basis of phenotypic, phylogenetic and genomic analyses, we formally propose the creation of Occidentia massiliensis gen. nov., sp. nov., that contains strain OS18T. This bacterium has been isolated in France from a tick collected in Senegal.
Description of Occidentia gen. nov
Occidentia (oc.ci.den’tia N.L. fem. Adj. occidentia, of the occident, for the western part of Africa where the tick from which the type strain was isolated, was collected, and in contrast with Orientia, the name of its phylogenetically closest relative, distributed in Asia).
Gimenez positive and weakly Gram-negative rods. Strictly intracellular. Non-spore-forming. Grows in BME/CTVM2 tick cells at 28°C. The bacteria multiply freely in the cytoplasm, but not the nucleus, of host cells. Monotrichous flagellum. Habitat: Ornithodoros sonrai. Type species: Occidentia massiliensis.
Description of Occidentia massiliensis gen. nov., sp. nov.
Occidentia massiliensis (mas.si.li.en’sis. L. gen. fem. n. massiliensis, of Massilia, the Latin name of Marseille, France, where strain OS18T was first grown, identified and characterized).
Gimenez positive and weakly Gram-negative rods. Strictly intracellular. Non-spore-forming. Grows in BME/CTVM2 tick cells at 28°C. The bacteria multiply freely in the cytoplasm, but not the nucleus, of host cells. Monotrichous flagellum. The mean length and width of the bacteria are 1.23 ± 0.19 μm and 0.42 ± 0.06 μm, respectively. Bacteria exhibit a large periplasmic space of 0.028 ± 0.007 μm. The genome is 1,469,252-bp long and contains 1,502 protein-coding and 41 RNA genes. The 16S rRNA and genomic sequences are deposited in GenBank under accession numbers GU937608 and CANJ00000000, respectively. The genomic G + C content is 29.05%. The type strain OS18T (CSUR = P764, DSM = 24860) was isolated from an Ornithodoros sonrai soft tick collected in Senegal.
Competing interests
The authors declare that they have no competing interests.
Authors’ contribution
OM isolated the bacterium, performed the photographies and electron micrography, phylogenetic analysis and drafted the manuscript, TTN carried out the genome sequencing, LBS participated in the cell culture, RP performed the genome comparison, PEF supervised the genomic studies and drafted the manuscript, DR initiated and organised the study and drafted the manuscript.
Contributor Information
Oleg Mediannikov, Email: olegusss1@gmail.com.
Thi-Thien Nguyen, Email: evenguyen1@yahoo.fr.
Lesley Bell-Sakyi, Email: lesley.sakyi@pirbright.ac.uk.
Roshan Padmanabhan, Email: rosaak@gmail.com.
Pierre-Edouard Fournier, Email: pierre-edouard.fournier@univ-amu.fr.
Didier Raoult, Email: didier.raoult@gmail.com.
Acknowledgements
We are grateful to Marie-Laure Birg, Nathalie Duclos, Audrey Borg and Denis Pyak for their technical help. The present work was funded by the Agence Nationale de Recherche grant 2010 MALEMAF (research on emergent pathogens in Africa) and the Mediterranée-Infection Foundation. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- Pinkerton H. Criteria for the accurate classification of the rickettsial disease (Rickettsioses) and their etiological agents. Parasitology. 1936;28:172–189. doi: 10.1017/S003118200002237X. http://dx.doi.org/10.1017/S003118200002237X. [DOI] [Google Scholar]
- Tamura A, Ohashi N, Urakami H, Miyamura S. Classification of Rickettsia tsutsugamushi in a new genus, Orientia gen. nov., as Orientia tsutsugamushi comb. nov. Int J Syst Bacteriol. 1995;45:589–591. doi: 10.1099/00207713-45-3-589. PubMed http://dx.doi.org/10.1099/00207713-45-3-589. [DOI] [PubMed] [Google Scholar]
- LPSN. http://www.bacterio.net/rickettsiaceae.html.
- Mediannikov O, Audoly G, Diatta G, Trape JF, Raoult D. New Rickettsia sp. in tsetse flies from Senegal. Comp Immunol Microbiol Infect Dis. 2012;35:145–150. doi: 10.1016/j.cimid.2011.12.011. PubMed http://dx.doi.org/10.1016/j.cimid.2011.12.011. [DOI] [PubMed] [Google Scholar]
- Shpynov SN, Fournier PE, Rudakov NV, Raoult D. "Candidatus Rickettsia tarasevichiae" in Ixodes persulcatus ticks collected in Russia. Ann N Y Acad Sci. 2003;990:162–172. doi: 10.1111/j.1749-6632.2003.tb07358.x. PubMed http://dx.doi.org/10.1111/j.1749-6632.2003.tb07358.x. [DOI] [PubMed] [Google Scholar]
- Dumler JS, Barbet AF, Bekker CPJ, Dasch GA, Palmer GH, Ray SC, Rikihisa Y, Rurangirwa FR. Reorganization of genera in the families Rickettsiaceae and Anaplasmataceae in the order Rickettsiales: unification of some species of Ehrlichia with Anaplasma, Cowdria with Ehrlichia and Ehrlichia with Neorickettsia, description of six new species combinations and designation of Ehrlichia equi and 'HGE agent' as subjective synonyms of Ehrlichia phagocytophila. Int J Syst Evol Microbiol. 2001;51:2145–2165. doi: 10.1099/00207713-51-6-2145. PubMed http://dx.doi.org/10.1099/00207713-51-6-2145. [DOI] [PubMed] [Google Scholar]
- Yu XJ, Walker DH. In: Bergeys's Manual of systematic Bacteriology, 2second edition, vol.2 (The Proteobacteria), part C (the Alpha-, Beta-, Delta-, and Epsilonproteobacteria) 2. Brenner DJ, Krieg NR, Staley JT, Garrity GM, editor. New York: Springer; 2005. Family I. Rickettsiaceae; pp. 1–324. [Google Scholar]
- Silverman DJ, Wisseman CL Jr. Comparative ultrastructural study on the cell envelopes of Rickettsia prowazekii, Rickettsia rickettsii, and Rickettsia tsutsugamushi. Infect Immun. 1978;21:1020–1023. doi: 10.1128/iai.21.3.1020-1023.1978. PubMed. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fournier PE, Dumler JS, Greub G, Zhang J, Wu Y, Raoult D. Gene sequence-based criteria for identification of new rickettsia isolates and description of Rickettsia heilongjiangensis sp. nov. J Clin Microbiol. 2003;41:5456–5465. doi: 10.1128/JCM.41.12.5456-5465.2003. PubMed http://dx.doi.org/10.1128/JCM.41.12.5456-5465.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19:1572–1574. doi: 10.1093/bioinformatics/btg180. PubMed http://dx.doi.org/10.1093/bioinformatics/btg180. [DOI] [PubMed] [Google Scholar]
- Mr Bayes software. http://mrbayes.csit.fsu.edu/
- Bell-Sakyi L. Ehrlichia ruminantium grows in cell lines from four ixodid tick genera. J Comp Pathol. 2004;103:285–293. doi: 10.1016/j.jcpa.2003.12.002. [DOI] [PubMed] [Google Scholar]
- Mediannikov O, Makarova V, Tarasevich I, Sidelnikov Y, Raoult D. Isolation of Rickettsia heilongjiangensis strains from humans and ticks and its multispacer typing. Clin Microbiol Infect. 2009;15:288–289. doi: 10.1111/j.1469-0691.2008.02239.x. PubMed http://dx.doi.org/10.1111/j.1469-0691.2008.02239.x. [DOI] [PubMed] [Google Scholar]
- Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, Ashburner M, Axelrod N, Baldauf S, Ballard S, Boore J, Cochrane G, Cole J, Dawyndt P, De Vos P, DePamphilis C, Edwards R, Faruque N, Feldman R, Gilbert J, Gilna P, Glöckner FO, Goldstein P, Guralnick R, Haft D, Hancock D. et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol. 2008;26:541–547. doi: 10.1038/nbt1360. PubMed http://dx.doi.org/10.1038/nbt1360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archae, Bacteria, and Eukarya. Proc Natl Acad Sci U S A. 1990;87:4576–4579. doi: 10.1073/pnas.87.12.4576. PubMed http://dx.doi.org/10.1073/pnas.87.12.4576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garrity GM, Bell JA, Lilburn T. In: The Proteobacteria), part B (the Gammaproteobacteria. 2. Brenner DJ, Krieg NR, Staley JT, Garrity GM, editor. New York: Springer; 2005. Phylum XIV. Proteobacteria phyl. nov; p. 1. [Google Scholar]
- Garrity GM, Bell JA, Lilburn T. In: Bergeys's Manual of systematic Bacteriology, second edition, vol.2 (The Proteobacteria), part C (the Alpha-, Beta-, Delta-, and Epsilonproteobacteria) Brenner DJ, Krieg NR, Staley JT, Garrity GM, editor. New York: Springer; 2005. pp. 1–324. [Google Scholar]
- Gieszczykiewicz M. Zagadniene systematihki w bakteriologii - Zür Frage der Bakterien-Systematic. Bull Acad Pol Sci, Ser Sci Biol. 1939;1:9–27. [Google Scholar]
- Brenner DJ, O'Connor SP, Winkler HH, Steigerwalt AG. Proposals to unify the genera Bartonella and Rochalimaea, with descriptions of Bartonella quintana comb. nov., Bartonella vinsonii comb. nov., Bartonella henselae comb. nov., and Bartonella elizabethae comb. nov., and to remove the family Bartonellaceae from the order Rickettsiales. Int J Syst Bacteriol. 1993;43:777–786. doi: 10.1099/00207713-43-4-777. PubMed http://dx.doi.org/10.1099/00207713-43-4-777. [DOI] [PubMed] [Google Scholar]
- Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–29. doi: 10.1038/75556. PubMed http://dx.doi.org/10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stackebrandt E, Ebers J. Taxonomic parameters revisited: tarnished gold standards. Microbiol Today. 2006;33:152–155. [Google Scholar]
- Ogata H, Renesto P, Audic S, Robert C, Blanc G, Fournier PE, Parinello H, Claverie JM, Raoult D. The genome sequence of Rickettsia felis identifies the first putative conjugative plasmid in an obligate intracellular parasite. PLoS Biol. 2005;3:e248. doi: 10.1371/journal.pbio.0030248. PubMed http://dx.doi.org/10.1371/journal.pbio.0030248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prokaryotic Dynamic Programming Genefinding Algorithm (PRODIGAL) http://prodigal.ornl.gov/. 13-9-2000.
- Tatusov RL, Galperin MY, Natale DA, Koonin EV. The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000;28:33–36. doi: 10.1093/nar/28.1.33. PubMed http://dx.doi.org/10.1093/nar/28.1.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25:955–964. doi: 10.1093/nar/25.5.0955. PubMed http://dx.doi.org/10.1093/nar/25.5.0955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lagesen K, Hallin P, Rodland EA, Staerfeldt HH, Rognes T, Ussery DW. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35:3100–3108. doi: 10.1093/nar/gkm160. PubMed http://dx.doi.org/10.1093/nar/gkm160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petersen TN, Brunak S, Von HG, Nielsen H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods. 2011;8:785–786. doi: 10.1038/nmeth.1701. PubMed http://dx.doi.org/10.1038/nmeth.1701. [DOI] [PubMed] [Google Scholar]
- TMHMM software. TMHMM software. http://www.cbs.dtu.dk/services/TMHMM/
- Ramasamy D, Mishra AK, Lagier JC, Padhmanabhan R, Rossi-Tamisier M, Sentausa E, Raoult D, Fournier PE. A polyphasic strategy incorporating genomic data for the taxonomic description of new bacterial species. Int J Syst Evol Microbiol. 2014;64:384–391. doi: 10.1099/ijs.0.057091-0. PubMed http://dx.doi.org/10.1099/ijs.0.057091-0. [DOI] [PubMed] [Google Scholar]
- Lechner M, Findeiss S, Steiner L, Marz M, Stadler PF, Prohaska SJ. Proteinortho: detection of (co-)orthologs in large-scale analysis. BMC Bioinformatics. 2011;12:124. doi: 10.1186/1471-2105-12-124. PubMed http://dx.doi.org/10.1186/1471-2105-12-124. [DOI] [PMC free article] [PubMed] [Google Scholar]